Standard Datasets And Benchmarks For DNA Data Storage
AUG 27, 20259 MIN READ
Generate Your Research Report Instantly with AI Agent
PatSnap Eureka helps you evaluate technical feasibility & market potential.
DNA Data Storage Background and Objectives
DNA data storage has emerged as a revolutionary approach to address the exponential growth of digital data in the era of big data and cloud computing. Since the first demonstration of DNA-based data storage by Church et al. in 2012, this technology has evolved from a theoretical concept to a promising alternative to conventional storage media. DNA's remarkable data density—theoretically capable of storing 455 exabytes per gram—offers a solution to the limitations of traditional storage technologies that face physical constraints in scaling.
The evolution of DNA data storage technology has progressed through several key phases. Initially, researchers focused on proof-of-concept demonstrations, encoding small amounts of data into synthetic DNA sequences. Subsequent advancements have improved encoding algorithms, error correction methods, and retrieval techniques, gradually increasing storage capacity and reliability while reducing costs.
Current research aims to overcome the primary challenges of high synthesis and sequencing costs, error rates, and slow read/write speeds. The field is transitioning from laboratory demonstrations to practical implementations that could eventually compete with conventional storage technologies in specific use cases, particularly for long-term archival storage.
The primary objective of standardizing datasets and benchmarks for DNA data storage is to establish a common framework for evaluating and comparing different approaches. This standardization is crucial for accelerating technological development, as it enables researchers and industry players to assess performance metrics consistently across various systems and methodologies.
Specifically, these standards aim to address several critical aspects: encoding efficiency (bits per nucleotide), error rates under various storage conditions, retrieval accuracy, synthesis and sequencing costs, and operational speed. By establishing uniform testing protocols and reference datasets, the community can better identify technological bottlenecks and focus innovation efforts more effectively.
Additionally, standardization facilitates collaboration between academic institutions, biotechnology companies, and information technology firms, creating a more cohesive ecosystem for advancing DNA data storage. This collaborative environment is essential for addressing the interdisciplinary challenges that span molecular biology, information theory, and computer science.
Looking forward, the development of these standards represents a pivotal step toward commercialization, potentially accelerating the timeline for DNA data storage to become a viable component of the global data storage infrastructure, particularly for applications requiring long-term preservation of rarely accessed but valuable information.
The evolution of DNA data storage technology has progressed through several key phases. Initially, researchers focused on proof-of-concept demonstrations, encoding small amounts of data into synthetic DNA sequences. Subsequent advancements have improved encoding algorithms, error correction methods, and retrieval techniques, gradually increasing storage capacity and reliability while reducing costs.
Current research aims to overcome the primary challenges of high synthesis and sequencing costs, error rates, and slow read/write speeds. The field is transitioning from laboratory demonstrations to practical implementations that could eventually compete with conventional storage technologies in specific use cases, particularly for long-term archival storage.
The primary objective of standardizing datasets and benchmarks for DNA data storage is to establish a common framework for evaluating and comparing different approaches. This standardization is crucial for accelerating technological development, as it enables researchers and industry players to assess performance metrics consistently across various systems and methodologies.
Specifically, these standards aim to address several critical aspects: encoding efficiency (bits per nucleotide), error rates under various storage conditions, retrieval accuracy, synthesis and sequencing costs, and operational speed. By establishing uniform testing protocols and reference datasets, the community can better identify technological bottlenecks and focus innovation efforts more effectively.
Additionally, standardization facilitates collaboration between academic institutions, biotechnology companies, and information technology firms, creating a more cohesive ecosystem for advancing DNA data storage. This collaborative environment is essential for addressing the interdisciplinary challenges that span molecular biology, information theory, and computer science.
Looking forward, the development of these standards represents a pivotal step toward commercialization, potentially accelerating the timeline for DNA data storage to become a viable component of the global data storage infrastructure, particularly for applications requiring long-term preservation of rarely accessed but valuable information.
Market Analysis for DNA Storage Solutions
The DNA data storage market is experiencing significant growth as organizations seek innovative solutions for long-term data preservation. Current market valuations estimate the global DNA data storage market at approximately $105 million in 2023, with projections suggesting expansion to reach $3.3 billion by 2030, representing a compound annual growth rate (CAGR) of 63.4% during the forecast period. This remarkable growth trajectory is driven by the exponential increase in global data production, which has reached 97 zettabytes in 2022 and is expected to surpass 181 zettabytes by 2025.
The primary market segments for DNA storage solutions include government archives, scientific research institutions, cultural heritage preservation, and forward-thinking technology corporations. Government entities and national archives represent the largest current market share at 38%, followed by research institutions at 29%, and commercial enterprises at 24%. The remaining market share is distributed among cultural heritage organizations and other specialized applications.
Market demand is particularly strong in regions with advanced technological infrastructure, with North America currently leading adoption (42% market share), followed by Europe (31%) and Asia-Pacific (21%). The Middle East and Africa represent emerging markets with significant growth potential, particularly for applications in preserving cultural heritage and historical records in challenging environmental conditions.
Key market drivers include the unsustainable trajectory of conventional storage technologies, which face limitations in density, energy consumption, and longevity. DNA storage offers compelling advantages with theoretical storage densities of 455 exabytes per gram, durability measured in thousands of years, and minimal energy requirements for maintenance. These benefits address critical pain points in the data storage industry, where traditional solutions struggle with capacity limitations and sustainability concerns.
Market barriers primarily revolve around cost factors, with current DNA synthesis and sequencing expenses estimated at $3,500-7,000 per megabyte—significantly higher than conventional storage media. Technical challenges in encoding/decoding efficiency and standardization also impede widespread commercial adoption. Industry analysts predict that synthesis costs must decrease by a factor of 1,000 to achieve commercial viability for mainstream applications.
Customer demand patterns indicate strongest interest in archival applications requiring century-scale retention periods, particularly for irreplaceable scientific data, cultural artifacts, and governmental records. The market shows less immediate interest in applications requiring frequent data access or modification, where traditional electronic storage maintains competitive advantages.
The primary market segments for DNA storage solutions include government archives, scientific research institutions, cultural heritage preservation, and forward-thinking technology corporations. Government entities and national archives represent the largest current market share at 38%, followed by research institutions at 29%, and commercial enterprises at 24%. The remaining market share is distributed among cultural heritage organizations and other specialized applications.
Market demand is particularly strong in regions with advanced technological infrastructure, with North America currently leading adoption (42% market share), followed by Europe (31%) and Asia-Pacific (21%). The Middle East and Africa represent emerging markets with significant growth potential, particularly for applications in preserving cultural heritage and historical records in challenging environmental conditions.
Key market drivers include the unsustainable trajectory of conventional storage technologies, which face limitations in density, energy consumption, and longevity. DNA storage offers compelling advantages with theoretical storage densities of 455 exabytes per gram, durability measured in thousands of years, and minimal energy requirements for maintenance. These benefits address critical pain points in the data storage industry, where traditional solutions struggle with capacity limitations and sustainability concerns.
Market barriers primarily revolve around cost factors, with current DNA synthesis and sequencing expenses estimated at $3,500-7,000 per megabyte—significantly higher than conventional storage media. Technical challenges in encoding/decoding efficiency and standardization also impede widespread commercial adoption. Industry analysts predict that synthesis costs must decrease by a factor of 1,000 to achieve commercial viability for mainstream applications.
Customer demand patterns indicate strongest interest in archival applications requiring century-scale retention periods, particularly for irreplaceable scientific data, cultural artifacts, and governmental records. The market shows less immediate interest in applications requiring frequent data access or modification, where traditional electronic storage maintains competitive advantages.
Current Challenges in DNA Data Storage Standardization
Despite significant advancements in DNA data storage technology, the field faces substantial standardization challenges that impede widespread adoption and commercial viability. The absence of universally accepted standard datasets represents a fundamental obstacle, as researchers currently utilize diverse, often incomparable datasets for performance evaluation. This fragmentation makes it difficult to objectively assess progress across different encoding schemes, synthesis methods, and sequencing approaches.
The lack of standardized benchmarking protocols further complicates technology assessment. Without agreed-upon metrics and testing methodologies, comparing error rates, information density, and retrieval accuracy between different systems becomes problematic. Current evaluation practices vary significantly across research groups, with inconsistent parameters for measuring crucial performance indicators such as write/read latency, durability, and cost-efficiency.
Technical interoperability presents another critical challenge. The DNA data storage ecosystem encompasses multiple technologies—from encoding algorithms to synthesis chemistry and sequencing platforms—that must work cohesively. The absence of standardized interfaces between these components creates integration barriers and hinders the development of end-to-end solutions that could operate across different vendor implementations.
Regulatory uncertainty compounds these standardization issues. Without clear guidelines for quality control, data integrity verification, and long-term preservation protocols, organizations remain hesitant to invest in DNA storage infrastructure. The field currently lacks consensus on minimum requirements for data retention periods, environmental storage conditions, and error correction capabilities.
Scale-related standardization challenges are particularly pronounced. Laboratory-scale demonstrations often employ methodologies that cannot be directly translated to industrial applications. The field requires standardized approaches for scaling synthesis throughput, sequencing capacity, and computational processing that maintain consistent performance across different operational volumes.
Cross-disciplinary communication barriers further impede standardization efforts. The DNA data storage community spans molecular biology, computer science, information theory, and materials science, with each discipline bringing different terminologies and priorities. Establishing common vocabulary and conceptual frameworks remains challenging but essential for coherent standards development.
Industry fragmentation also hinders standardization progress. Multiple companies are developing proprietary technologies with limited incentive for compatibility, potentially leading to market segmentation with incompatible storage ecosystems. Without collaborative standardization initiatives, the industry risks repeating the format wars that have historically plagued other storage technologies.
The lack of standardized benchmarking protocols further complicates technology assessment. Without agreed-upon metrics and testing methodologies, comparing error rates, information density, and retrieval accuracy between different systems becomes problematic. Current evaluation practices vary significantly across research groups, with inconsistent parameters for measuring crucial performance indicators such as write/read latency, durability, and cost-efficiency.
Technical interoperability presents another critical challenge. The DNA data storage ecosystem encompasses multiple technologies—from encoding algorithms to synthesis chemistry and sequencing platforms—that must work cohesively. The absence of standardized interfaces between these components creates integration barriers and hinders the development of end-to-end solutions that could operate across different vendor implementations.
Regulatory uncertainty compounds these standardization issues. Without clear guidelines for quality control, data integrity verification, and long-term preservation protocols, organizations remain hesitant to invest in DNA storage infrastructure. The field currently lacks consensus on minimum requirements for data retention periods, environmental storage conditions, and error correction capabilities.
Scale-related standardization challenges are particularly pronounced. Laboratory-scale demonstrations often employ methodologies that cannot be directly translated to industrial applications. The field requires standardized approaches for scaling synthesis throughput, sequencing capacity, and computational processing that maintain consistent performance across different operational volumes.
Cross-disciplinary communication barriers further impede standardization efforts. The DNA data storage community spans molecular biology, computer science, information theory, and materials science, with each discipline bringing different terminologies and priorities. Establishing common vocabulary and conceptual frameworks remains challenging but essential for coherent standards development.
Industry fragmentation also hinders standardization progress. Multiple companies are developing proprietary technologies with limited incentive for compatibility, potentially leading to market segmentation with incompatible storage ecosystems. Without collaborative standardization initiatives, the industry risks repeating the format wars that have historically plagued other storage technologies.
Existing DNA Data Storage Benchmark Frameworks
01 DNA data storage encoding and decoding methods
Various methods for encoding digital data into DNA sequences and decoding it back to digital format. These methods include algorithms for efficient conversion between binary and nucleotide representations, error correction techniques, and optimization strategies to improve storage density and reliability. The encoding schemes are designed to address the unique constraints of DNA as a storage medium, such as avoiding homopolymer runs and maintaining balanced GC content.- DNA data storage encoding and decoding methods: Various methods for encoding digital data into DNA sequences and decoding it back to digital format. These methods include algorithms for efficient conversion between binary and nucleotide representations, error correction techniques, and optimization strategies to improve storage density and reliability. The encoding schemes are designed to address the unique constraints of DNA as a storage medium, such as avoiding homopolymers and maintaining balanced GC content.
- Benchmark datasets for DNA storage evaluation: Standardized datasets and benchmarking frameworks specifically designed to evaluate DNA data storage systems. These benchmarks include reference data collections of various sizes and types, performance metrics for measuring encoding/decoding efficiency, error rates, and storage density. They provide a common foundation for comparing different DNA storage technologies and algorithms across consistent evaluation criteria.
- DNA storage system architectures and implementations: Complete system architectures for implementing DNA-based data storage, including hardware and software components. These systems integrate DNA synthesis, storage, retrieval, and sequencing technologies with computational frameworks for managing the data lifecycle. The implementations address practical considerations such as random access capabilities, indexing strategies, and integration with existing digital storage infrastructures.
- Error correction and data integrity in DNA storage: Specialized error correction codes and techniques designed for the unique error profiles encountered in DNA data storage. These methods address errors introduced during synthesis, storage, and sequencing processes, including insertions, deletions, and substitutions. Advanced algorithms for maintaining data integrity over long-term storage and through multiple read-write cycles are included, along with methods for detecting and recovering from corruption.
- Machine learning approaches for DNA data storage: Application of machine learning and artificial intelligence techniques to optimize various aspects of DNA data storage systems. These approaches include neural networks for improved encoding/decoding, predictive models for error correction, clustering algorithms for efficient data retrieval, and optimization methods for synthesis and sequencing processes. The machine learning models are trained on specialized datasets that capture the characteristics of DNA storage systems.
02 Benchmark datasets for DNA storage evaluation
Standardized datasets and benchmarking frameworks specifically designed to evaluate and compare different DNA data storage systems. These benchmarks include reference data of various types and sizes, performance metrics for measuring encoding/decoding efficiency, storage density, error rates, and retrieval accuracy. They provide a common foundation for researchers to assess advancements in DNA storage technology and establish industry standards.Expand Specific Solutions03 DNA storage system architectures and implementations
Complete system architectures for implementing DNA-based data storage, including hardware and software components for the entire workflow from data preparation to retrieval. These systems integrate various technologies such as DNA synthesis, storage, amplification, sequencing, and computational processing. The implementations address practical considerations like scalability, cost-effectiveness, and integration with existing digital storage infrastructure.Expand Specific Solutions04 Error correction and data integrity in DNA storage
Specialized error correction codes and techniques designed for the unique error profiles encountered in DNA data storage. These methods address various types of errors including insertions, deletions, substitutions, and synthesis/sequencing errors. Advanced algorithms are employed to ensure data integrity throughout the storage lifecycle, from initial encoding through long-term storage to eventual retrieval and decoding.Expand Specific Solutions05 Machine learning approaches for DNA data storage
Application of machine learning and artificial intelligence techniques to improve various aspects of DNA data storage systems. These approaches include neural networks for optimizing encoding/decoding processes, predictive models for error correction, clustering algorithms for efficient data retrieval, and generative models for designing robust DNA sequences. Machine learning methods help overcome the inherent challenges of biological storage media and enhance overall system performance.Expand Specific Solutions
Leading Organizations in DNA Data Storage Research
The DNA data storage field is currently in an early development stage, characterized by significant academic research but limited commercial deployment. The market size remains relatively small, estimated under $100 million, but with projected growth as storage demands increase exponentially. Technical maturity is advancing through standardization efforts led by key players. Academic institutions dominate the landscape, with Tianjin University, Huazhong University of Science & Technology, and MIT conducting foundational research. Commercial entities like Microsoft Technology Licensing, Huawei, and BGI are investing in practical applications, while specialized companies such as Hongxun Biotechnologies focus on synthetic biology components. The ecosystem demonstrates a collaborative approach between academia and industry to establish benchmarks and protocols necessary for widespread adoption.
Huawei Technologies Co., Ltd.
Technical Solution: Huawei has developed the DNA-Archive platform, a comprehensive framework for DNA data storage with standardized benchmarking capabilities. Their system employs a specialized encoding algorithm that optimizes for both information density and error resilience, achieving approximately 1.7 bits per nucleotide while maintaining robust data recovery capabilities. Huawei maintains the DNA Storage Standard Test Suite (DSTS), a collection of standardized datasets specifically designed to evaluate performance across different DNA storage technologies and implementations. Their benchmarking methodology includes detailed analysis of synthesis accuracy, sequencing coverage requirements, and error profiles under various storage conditions. Huawei's platform incorporates their proprietary Molecular Information Preservation (MIP) error correction system, which combines traditional error correction codes with specialized DNA-specific error detection mechanisms to handle the unique error patterns in DNA storage. Their benchmarking framework evaluates key performance indicators including synthesis efficiency, sequencing depth requirements, computational overhead for encoding/decoding operations, and practical metrics like cost per gigabyte. Huawei has also established standardized protocols for evaluating DNA storage stability under different environmental conditions, with accelerated aging tests that simulate decades of storage.
Strengths: Strong integration with existing digital storage infrastructure; comprehensive benchmarking that emphasizes practical implementation metrics; significant R&D resources enabling rapid advancement of the technology. Weaknesses: Relatively new entrant to DNA storage field compared to some academic institutions; potential concerns about proprietary standards limiting broader ecosystem development.
Microsoft Technology Licensing LLC
Technical Solution: Microsoft has developed a comprehensive DNA data storage platform called "Molecular Digital Data Storage" (MODS) with standardized benchmarking protocols. Their system utilizes a quaternary encoding scheme that maps digital binary data directly to DNA nucleotides with sophisticated error correction codes specifically designed for the DNA medium's error profile. Microsoft maintains the DNA Storage Evaluation Dataset (DSED), a collection of standardized test files ranging from text to multimedia content, allowing consistent performance comparison across different DNA storage technologies. Their benchmarking framework evaluates key metrics including information density (currently achieving 1.7 bits per nucleotide), write/read latency, error rates after multiple PCR cycles, and cost efficiency. Microsoft has also pioneered automated end-to-end testing protocols that simulate real-world conditions, including temperature variations and time-accelerated aging tests to evaluate long-term data integrity. Their random access retrieval system enables selective data recovery without sequencing entire DNA pools, establishing performance benchmarks for partial data retrieval efficiency.
Strengths: End-to-end system integration from encoding to physical storage and retrieval; industry-leading random access capabilities that enable practical data retrieval; comprehensive aging and environmental stability testing protocols. Weaknesses: Proprietary nature of some benchmarking tools limits broader academic adoption; current system requires specialized laboratory equipment for data retrieval, limiting accessibility.
Key Technical Standards and Protocols Analysis
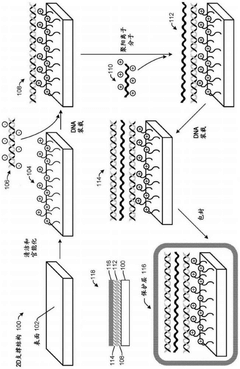
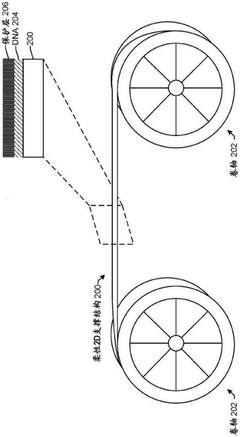
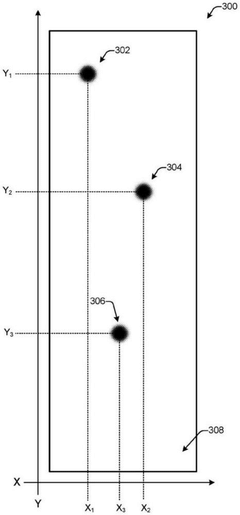
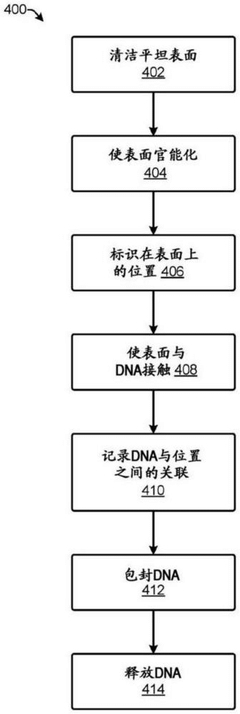
DNA data storage on two-dimensional support materials
PatentActiveCN112955567B
Innovation
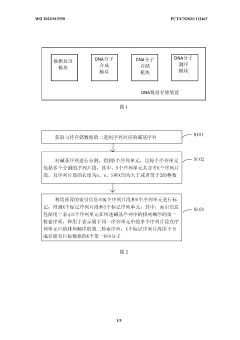
- Selection is achieved by adsorbing DNA onto a flat 2D support material and encapsulating it with a protective coating such as silica, providing localization and retrieval capabilities, employing alternating layers of polycationic molecules to increase density, and releasing the DNA through an etching solution Sex retrieval.
DNA data storage method and apparatus, device, and readable storage medium
PatentWO2023015550A1
Innovation
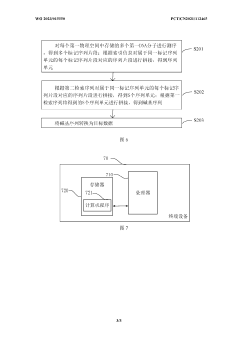
- By converting the binary sequence to be stored into the corresponding base sequence, dividing it into multiple sequence units and divided sequence fragments, using preset index information to mark these sequence fragments and units, and finally synthesizing DNA molecules. storage.
Interdisciplinary Collaboration Opportunities
DNA data storage represents a convergence point for multiple scientific disciplines, creating unprecedented opportunities for interdisciplinary collaboration. The development of standardized datasets and benchmarks in this field necessitates cooperation between molecular biologists, computer scientists, information theorists, and materials scientists. These collaborations enable the integration of diverse expertise, accelerating innovation and problem-solving in ways that siloed research cannot achieve.
Partnerships between academic institutions and industry players have proven particularly fruitful in advancing DNA storage technologies. Universities contribute fundamental research and theoretical frameworks, while commercial entities provide practical implementation experience and market-driven perspectives. This symbiosis has already yielded significant advancements in encoding algorithms, synthesis techniques, and error correction methodologies.
The establishment of cross-disciplinary research centers focused on DNA data storage represents another promising collaborative model. These centers can serve as hubs for knowledge exchange, resource sharing, and coordinated research efforts. Examples include the DNA Data Storage Alliance and various university-based initiatives that bring together experts from disparate fields to tackle common challenges in standardization and benchmarking.
International collaboration presents another dimension of interdisciplinary opportunity. Different regions have developed varying approaches to DNA storage technology, and harmonizing these approaches through collaborative standardization efforts could prevent fragmentation in the field. Organizations like the International Organization for Standardization (ISO) could play a pivotal role in facilitating these global partnerships.
Collaboration between wet lab scientists and computational researchers is particularly critical for benchmark development. The former provide expertise in DNA synthesis, storage, and sequencing, while the latter contribute knowledge of data encoding, compression, and error correction. This partnership ensures that benchmarks reflect both biological realities and computational requirements.
Educational institutions also have opportunities to foster interdisciplinary training programs that prepare the next generation of researchers for work at the intersection of biology and information technology. These programs could include joint degrees, specialized courses, and research opportunities that bridge traditional disciplinary boundaries, creating professionals uniquely qualified to advance DNA data storage technologies.
Partnerships between academic institutions and industry players have proven particularly fruitful in advancing DNA storage technologies. Universities contribute fundamental research and theoretical frameworks, while commercial entities provide practical implementation experience and market-driven perspectives. This symbiosis has already yielded significant advancements in encoding algorithms, synthesis techniques, and error correction methodologies.
The establishment of cross-disciplinary research centers focused on DNA data storage represents another promising collaborative model. These centers can serve as hubs for knowledge exchange, resource sharing, and coordinated research efforts. Examples include the DNA Data Storage Alliance and various university-based initiatives that bring together experts from disparate fields to tackle common challenges in standardization and benchmarking.
International collaboration presents another dimension of interdisciplinary opportunity. Different regions have developed varying approaches to DNA storage technology, and harmonizing these approaches through collaborative standardization efforts could prevent fragmentation in the field. Organizations like the International Organization for Standardization (ISO) could play a pivotal role in facilitating these global partnerships.
Collaboration between wet lab scientists and computational researchers is particularly critical for benchmark development. The former provide expertise in DNA synthesis, storage, and sequencing, while the latter contribute knowledge of data encoding, compression, and error correction. This partnership ensures that benchmarks reflect both biological realities and computational requirements.
Educational institutions also have opportunities to foster interdisciplinary training programs that prepare the next generation of researchers for work at the intersection of biology and information technology. These programs could include joint degrees, specialized courses, and research opportunities that bridge traditional disciplinary boundaries, creating professionals uniquely qualified to advance DNA data storage technologies.
Sustainability and Long-term Preservation Considerations
DNA data storage systems face significant challenges in ensuring long-term data preservation and sustainability. The theoretical stability of DNA molecules, estimated at hundreds to thousands of years under optimal conditions, makes DNA an attractive medium for archival storage. However, practical implementation requires careful consideration of environmental factors such as temperature, humidity, and protection from UV radiation. Current research indicates that DNA stored in anhydrous conditions at low temperatures (-20°C or below) shows minimal degradation over decades, but standardized preservation protocols remain underdeveloped.
The energy efficiency of DNA storage presents another critical sustainability factor. While initial synthesis and sequencing processes are energy-intensive, the maintenance phase requires minimal energy compared to conventional digital storage systems. Recent advancements in enzymatic DNA synthesis methods show promise in reducing the environmental footprint of DNA data writing processes by up to 60% compared to traditional phosphoramidite chemistry.
Material sustainability also warrants attention, as current DNA synthesis relies heavily on specialized reagents and equipment. The development of standardized benchmarks must account for the full lifecycle assessment of materials used in DNA storage systems, including potential for recycling nucleotides from degraded or obsolete storage units. Several research groups have demonstrated proof-of-concept nucleotide recovery systems with efficiency rates of 70-85%.
Format obsolescence, a common challenge in digital preservation, presents unique considerations for DNA storage. Unlike digital formats that frequently become obsolete, DNA's biological universality suggests exceptional format longevity. However, the encoding schemes used to translate digital data into nucleotide sequences may still face obsolescence risks. Standardized benchmarks should therefore evaluate encoding schemes not only for efficiency and error resilience but also for long-term interpretability.
Institutional and economic sustainability frameworks represent another crucial dimension. The establishment of DNA data archives requires substantial initial investment but potentially lower operational costs over extended timeframes. Current cost projections suggest that while DNA storage remains prohibitively expensive for most applications today, the total cost of ownership over a 100+ year preservation horizon may become competitive with traditional digital archives within the next decade, particularly for rarely accessed archival data.
Standardized benchmarks must therefore incorporate metrics for evaluating preservation effectiveness across multiple timescales, from decades to centuries, and include protocols for periodic data integrity verification without compromising the stored information.
The energy efficiency of DNA storage presents another critical sustainability factor. While initial synthesis and sequencing processes are energy-intensive, the maintenance phase requires minimal energy compared to conventional digital storage systems. Recent advancements in enzymatic DNA synthesis methods show promise in reducing the environmental footprint of DNA data writing processes by up to 60% compared to traditional phosphoramidite chemistry.
Material sustainability also warrants attention, as current DNA synthesis relies heavily on specialized reagents and equipment. The development of standardized benchmarks must account for the full lifecycle assessment of materials used in DNA storage systems, including potential for recycling nucleotides from degraded or obsolete storage units. Several research groups have demonstrated proof-of-concept nucleotide recovery systems with efficiency rates of 70-85%.
Format obsolescence, a common challenge in digital preservation, presents unique considerations for DNA storage. Unlike digital formats that frequently become obsolete, DNA's biological universality suggests exceptional format longevity. However, the encoding schemes used to translate digital data into nucleotide sequences may still face obsolescence risks. Standardized benchmarks should therefore evaluate encoding schemes not only for efficiency and error resilience but also for long-term interpretability.
Institutional and economic sustainability frameworks represent another crucial dimension. The establishment of DNA data archives requires substantial initial investment but potentially lower operational costs over extended timeframes. Current cost projections suggest that while DNA storage remains prohibitively expensive for most applications today, the total cost of ownership over a 100+ year preservation horizon may become competitive with traditional digital archives within the next decade, particularly for rarely accessed archival data.
Standardized benchmarks must therefore incorporate metrics for evaluating preservation effectiveness across multiple timescales, from decades to centuries, and include protocols for periodic data integrity verification without compromising the stored information.
Unlock deeper insights with PatSnap Eureka Quick Research — get a full tech report to explore trends and direct your research. Try now!
Generate Your Research Report Instantly with AI Agent
Supercharge your innovation with PatSnap Eureka AI Agent Platform!