How Geometric Isomerism Affects Protein Solvation Shell Structures
AUG 4, 20259 MIN READ
Generate Your Research Report Instantly with AI Agent
PatSnap Eureka helps you evaluate technical feasibility & market potential.
Geometric Isomerism and Protein Solvation Background
Geometric isomerism and protein solvation are fundamental concepts in biochemistry and biophysics that play crucial roles in understanding protein structure, function, and interactions. Geometric isomerism refers to the spatial arrangement of atoms or groups within a molecule, where different configurations can exist without breaking or forming new chemical bonds. In the context of proteins, this phenomenon is particularly relevant for amino acid residues and their side chains.
Protein solvation, on the other hand, describes the interaction between proteins and the surrounding solvent molecules, typically water. The solvation shell, also known as the hydration shell, is the layer of solvent molecules that surrounds and interacts with the protein surface. This shell is critical for maintaining protein stability, influencing protein dynamics, and mediating protein-protein interactions.
The interplay between geometric isomerism and protein solvation is a complex and fascinating area of study. Geometric isomers of amino acid residues can significantly affect the local structure and properties of the protein surface, which in turn influences the organization and behavior of the solvation shell. This relationship is bidirectional, as the solvation environment can also impact the conformational preferences of geometric isomers within the protein structure.
Understanding how geometric isomerism affects protein solvation shell structures is crucial for several reasons. Firstly, it provides insights into protein folding and stability, as the solvation shell plays a vital role in these processes. Secondly, it helps elucidate the mechanisms of protein-ligand interactions and drug binding, which are often mediated by the solvation shell. Lastly, this knowledge is essential for developing accurate computational models of protein behavior in solution, which have applications in drug design, protein engineering, and understanding disease-related protein mutations.
The study of geometric isomerism and protein solvation has evolved significantly over the past few decades, driven by advancements in experimental techniques and computational methods. X-ray crystallography, NMR spectroscopy, and cryo-electron microscopy have provided high-resolution structural information about proteins and their solvation shells. Complementing these experimental approaches, molecular dynamics simulations and quantum mechanical calculations have offered detailed insights into the dynamic behavior of proteins and their interactions with solvent molecules at the atomic level.
Recent research has focused on understanding the subtle effects of geometric isomerism on local hydration patterns, water dynamics, and the thermodynamics of protein-solvent interactions. These studies have revealed that even small changes in the geometric configuration of amino acid side chains can lead to significant alterations in the structure and properties of the surrounding solvation shell, highlighting the importance of considering geometric isomerism in protein solvation studies.
Protein solvation, on the other hand, describes the interaction between proteins and the surrounding solvent molecules, typically water. The solvation shell, also known as the hydration shell, is the layer of solvent molecules that surrounds and interacts with the protein surface. This shell is critical for maintaining protein stability, influencing protein dynamics, and mediating protein-protein interactions.
The interplay between geometric isomerism and protein solvation is a complex and fascinating area of study. Geometric isomers of amino acid residues can significantly affect the local structure and properties of the protein surface, which in turn influences the organization and behavior of the solvation shell. This relationship is bidirectional, as the solvation environment can also impact the conformational preferences of geometric isomers within the protein structure.
Understanding how geometric isomerism affects protein solvation shell structures is crucial for several reasons. Firstly, it provides insights into protein folding and stability, as the solvation shell plays a vital role in these processes. Secondly, it helps elucidate the mechanisms of protein-ligand interactions and drug binding, which are often mediated by the solvation shell. Lastly, this knowledge is essential for developing accurate computational models of protein behavior in solution, which have applications in drug design, protein engineering, and understanding disease-related protein mutations.
The study of geometric isomerism and protein solvation has evolved significantly over the past few decades, driven by advancements in experimental techniques and computational methods. X-ray crystallography, NMR spectroscopy, and cryo-electron microscopy have provided high-resolution structural information about proteins and their solvation shells. Complementing these experimental approaches, molecular dynamics simulations and quantum mechanical calculations have offered detailed insights into the dynamic behavior of proteins and their interactions with solvent molecules at the atomic level.
Recent research has focused on understanding the subtle effects of geometric isomerism on local hydration patterns, water dynamics, and the thermodynamics of protein-solvent interactions. These studies have revealed that even small changes in the geometric configuration of amino acid side chains can lead to significant alterations in the structure and properties of the surrounding solvation shell, highlighting the importance of considering geometric isomerism in protein solvation studies.
Market Demand for Protein Solvation Research
The market demand for protein solvation research has been steadily growing, driven by its critical importance in various fields, including drug discovery, protein engineering, and biotechnology. The pharmaceutical industry, in particular, has shown a keen interest in understanding protein solvation shell structures and how geometric isomerism affects them. This knowledge is crucial for developing more effective and targeted drugs, as it helps predict protein-ligand interactions and drug efficacy.
In the biotechnology sector, protein solvation research is essential for optimizing protein production and purification processes. Companies involved in the production of biopharmaceuticals, enzymes, and other protein-based products are investing heavily in this area to improve their manufacturing efficiency and product quality. The food industry is also showing increased interest in protein solvation research, particularly in the development of novel food additives and protein-based ingredients with enhanced stability and functionality.
The academic research community continues to be a significant driver of demand for protein solvation studies. Universities and research institutions worldwide are conducting extensive research in this field, contributing to the overall market growth. This academic interest is further fueled by the availability of advanced computational tools and experimental techniques that allow for more detailed investigations of protein solvation phenomena.
The market for protein solvation research tools and services is expanding rapidly. Manufacturers of analytical instruments, such as spectroscopy and calorimetry equipment, are developing specialized tools for studying protein solvation. Software companies are also capitalizing on this trend by offering advanced molecular dynamics simulation packages and data analysis tools tailored for protein solvation research.
Environmental concerns and the push for sustainable technologies have created new opportunities in the protein solvation research market. Industries such as bioremediation and green chemistry are increasingly relying on protein solvation studies to develop more efficient and environmentally friendly processes. This trend is expected to continue as global efforts to reduce environmental impact intensify.
The growing interest in personalized medicine has further boosted the demand for protein solvation research. Understanding how individual genetic variations affect protein solvation and function is crucial for developing tailored therapeutic approaches. This has led to increased collaboration between pharmaceutical companies, research institutions, and healthcare providers, driving further investment in this field.
In the biotechnology sector, protein solvation research is essential for optimizing protein production and purification processes. Companies involved in the production of biopharmaceuticals, enzymes, and other protein-based products are investing heavily in this area to improve their manufacturing efficiency and product quality. The food industry is also showing increased interest in protein solvation research, particularly in the development of novel food additives and protein-based ingredients with enhanced stability and functionality.
The academic research community continues to be a significant driver of demand for protein solvation studies. Universities and research institutions worldwide are conducting extensive research in this field, contributing to the overall market growth. This academic interest is further fueled by the availability of advanced computational tools and experimental techniques that allow for more detailed investigations of protein solvation phenomena.
The market for protein solvation research tools and services is expanding rapidly. Manufacturers of analytical instruments, such as spectroscopy and calorimetry equipment, are developing specialized tools for studying protein solvation. Software companies are also capitalizing on this trend by offering advanced molecular dynamics simulation packages and data analysis tools tailored for protein solvation research.
Environmental concerns and the push for sustainable technologies have created new opportunities in the protein solvation research market. Industries such as bioremediation and green chemistry are increasingly relying on protein solvation studies to develop more efficient and environmentally friendly processes. This trend is expected to continue as global efforts to reduce environmental impact intensify.
The growing interest in personalized medicine has further boosted the demand for protein solvation research. Understanding how individual genetic variations affect protein solvation and function is crucial for developing tailored therapeutic approaches. This has led to increased collaboration between pharmaceutical companies, research institutions, and healthcare providers, driving further investment in this field.
Current Challenges in Isomer-Solvation Studies
The study of geometric isomerism and its effects on protein solvation shell structures faces several significant challenges in current research. One of the primary obstacles is the complexity of protein structures and their interactions with solvent molecules. Proteins are large, dynamic macromolecules with intricate three-dimensional conformations that can change in response to environmental factors, including the presence of different isomers.
The high-resolution characterization of solvation shells around proteins, particularly in the context of geometric isomers, requires advanced experimental techniques that are often limited in their spatial and temporal resolution. Traditional methods such as X-ray crystallography and NMR spectroscopy, while powerful, may not capture the full dynamic nature of solvation processes influenced by isomerism.
Another challenge lies in the computational modeling of isomer-solvation interactions. Current molecular dynamics simulations and quantum mechanical calculations struggle to accurately represent the subtle differences in solvation patterns caused by geometric isomers. The development of force fields and simulation parameters that can precisely account for isomeric effects on solvation remains an ongoing area of research.
The heterogeneity of protein surfaces further complicates the study of isomer-solvation interactions. Different regions of a protein may interact differently with solvent molecules and isomers, making it difficult to generalize findings across diverse protein systems. This variability necessitates case-by-case studies, which can be time-consuming and resource-intensive.
Moreover, the influence of geometric isomerism on solvation shell structures can be subtle and easily masked by other factors such as temperature, pH, and ionic strength. Isolating the specific effects of isomerism from these confounding variables presents a significant experimental and analytical challenge.
The dynamic nature of solvation shells adds another layer of complexity. Solvation structures are not static but constantly fluctuating, making it challenging to capture and analyze the transient effects of geometric isomers on these structures. Advanced time-resolved spectroscopic techniques are needed to probe these fast-occurring phenomena.
Lastly, the integration of experimental data with theoretical models remains a significant hurdle. Bridging the gap between empirical observations and computational predictions is crucial for developing a comprehensive understanding of how geometric isomerism affects protein solvation shell structures. This integration requires interdisciplinary collaboration and the development of new analytical frameworks.
The high-resolution characterization of solvation shells around proteins, particularly in the context of geometric isomers, requires advanced experimental techniques that are often limited in their spatial and temporal resolution. Traditional methods such as X-ray crystallography and NMR spectroscopy, while powerful, may not capture the full dynamic nature of solvation processes influenced by isomerism.
Another challenge lies in the computational modeling of isomer-solvation interactions. Current molecular dynamics simulations and quantum mechanical calculations struggle to accurately represent the subtle differences in solvation patterns caused by geometric isomers. The development of force fields and simulation parameters that can precisely account for isomeric effects on solvation remains an ongoing area of research.
The heterogeneity of protein surfaces further complicates the study of isomer-solvation interactions. Different regions of a protein may interact differently with solvent molecules and isomers, making it difficult to generalize findings across diverse protein systems. This variability necessitates case-by-case studies, which can be time-consuming and resource-intensive.
Moreover, the influence of geometric isomerism on solvation shell structures can be subtle and easily masked by other factors such as temperature, pH, and ionic strength. Isolating the specific effects of isomerism from these confounding variables presents a significant experimental and analytical challenge.
The dynamic nature of solvation shells adds another layer of complexity. Solvation structures are not static but constantly fluctuating, making it challenging to capture and analyze the transient effects of geometric isomers on these structures. Advanced time-resolved spectroscopic techniques are needed to probe these fast-occurring phenomena.
Lastly, the integration of experimental data with theoretical models remains a significant hurdle. Bridging the gap between empirical observations and computational predictions is crucial for developing a comprehensive understanding of how geometric isomerism affects protein solvation shell structures. This integration requires interdisciplinary collaboration and the development of new analytical frameworks.
Existing Methods for Studying Geometric Isomers
01 Computational methods for analyzing protein solvation shell structures
Advanced computational techniques are employed to analyze and model protein solvation shell structures. These methods involve simulations and algorithms to predict the arrangement of water molecules around proteins, considering geometric isomerism. Such approaches help in understanding protein-solvent interactions and their impact on protein function and stability.- Computational methods for analyzing protein solvation shells: Advanced computational techniques are employed to study the structure and dynamics of protein solvation shells. These methods involve molecular dynamics simulations, quantum mechanical calculations, and statistical analysis to understand the interactions between proteins and surrounding water molecules. The analysis helps in predicting protein behavior and stability in various environments.
- Geometric isomerism in protein-ligand interactions: The study of geometric isomerism plays a crucial role in understanding protein-ligand interactions. Researchers investigate how different spatial arrangements of atoms in ligands affect their binding to protein targets. This knowledge is essential for drug design and optimization, as it helps predict the efficacy and specificity of potential therapeutic compounds.
- Solvation effects on protein structure and function: The solvation environment significantly influences protein structure and function. Researchers examine how different solvents and solvent compositions affect protein folding, stability, and activity. This includes studying the role of water molecules in protein-protein interactions and enzyme catalysis, which is crucial for understanding biological processes at the molecular level.
- Machine learning approaches for predicting protein solvation properties: Machine learning algorithms are increasingly used to predict and analyze protein solvation properties. These methods can process large datasets of protein structures and their corresponding solvation characteristics to identify patterns and make accurate predictions. This approach accelerates research in protein engineering, drug discovery, and understanding protein-solvent interactions.
- Experimental techniques for studying protein solvation shells: Various experimental methods are employed to investigate protein solvation shells and their structural properties. These include spectroscopic techniques such as NMR and X-ray crystallography, as well as advanced microscopy methods. These experimental approaches provide valuable data on the arrangement of water molecules around proteins and help validate computational models.
02 Experimental techniques for studying protein solvation shells
Various experimental methods are used to investigate protein solvation shell structures and their geometric isomerism. These techniques may include spectroscopic methods, X-ray crystallography, and nuclear magnetic resonance (NMR) spectroscopy. These approaches provide valuable insights into the organization of water molecules around proteins and how different isomers affect solvation.Expand Specific Solutions03 Influence of geometric isomerism on protein solvation
Geometric isomerism plays a crucial role in determining the structure and properties of protein solvation shells. Different isomers can lead to variations in the arrangement of water molecules around proteins, affecting their solubility, stability, and function. Understanding these relationships is essential for predicting protein behavior in solution and designing protein-based therapeutics.Expand Specific Solutions04 Solvation effects on protein structure and function
The solvation shell structure significantly impacts protein structure and function. Geometric isomerism in the solvation shell can influence protein folding, conformational changes, and interactions with other molecules. This understanding is crucial for predicting protein behavior in different environments and designing proteins with specific properties.Expand Specific Solutions05 Applications in drug design and protein engineering
Knowledge of protein solvation shell structures and geometric isomerism is applied in drug design and protein engineering. This information helps in predicting drug-protein interactions, optimizing protein stability, and designing novel proteins with desired properties. Such applications have significant implications in pharmaceutical development and biotechnology.Expand Specific Solutions
Key Players in Protein Structure Research
The field of geometric isomerism and its impact on protein solvation shell structures is in a relatively mature stage of development, with ongoing research refining our understanding of these complex interactions. The market for related technologies and applications is substantial, particularly in pharmaceutical and biotechnology sectors. Companies like Abbott Laboratories, Vertex Pharmaceuticals, and Novo Nordisk are at the forefront, leveraging this knowledge for drug development and protein engineering. Academic institutions such as Oxford University and McMaster University contribute significantly to fundamental research. The technology's maturity is evident in its widespread application, but there's still room for innovation, especially in computational modeling and experimental techniques for studying protein-solvent interactions at the molecular level.
Abbott Laboratories
Technical Solution: Abbott Laboratories has focused on developing novel analytical techniques to study the effects of geometric isomerism on protein solvation shell structures. They have pioneered the use of advanced nuclear magnetic resonance (NMR) spectroscopy methods, including 2D and 3D heteronuclear experiments, to probe the local environment around specific isomers within proteins[5]. This approach allows for atomic-level resolution of solvation shell structures and dynamics. Additionally, Abbott has developed custom software tools for analyzing and visualizing NMR data in the context of protein isomerism and solvation[6].
Strengths: High-resolution experimental techniques, custom data analysis tools. Weaknesses: Limited to NMR-amenable systems, potentially time-consuming experiments.
Novo Nordisk A/S
Technical Solution: Novo Nordisk has developed a comprehensive platform for studying the effects of geometric isomerism on protein solvation, with a particular focus on therapeutic proteins and peptides. Their approach combines high-resolution X-ray crystallography and cryo-electron microscopy to obtain structural information about different isomeric states of proteins[7]. This is complemented by molecular dynamics simulations that model the behavior of water molecules around these structures. Novo Nordisk has also implemented advanced biophysical techniques, such as isothermal titration calorimetry, to quantify the thermodynamic consequences of isomer-specific solvation effects[8].
Strengths: Integration of multiple structural biology techniques, focus on therapeutic relevance. Weaknesses: Reliance on static structural data, potential challenges in capturing dynamic solvation effects.
Core Innovations in Solvation Shell Modeling
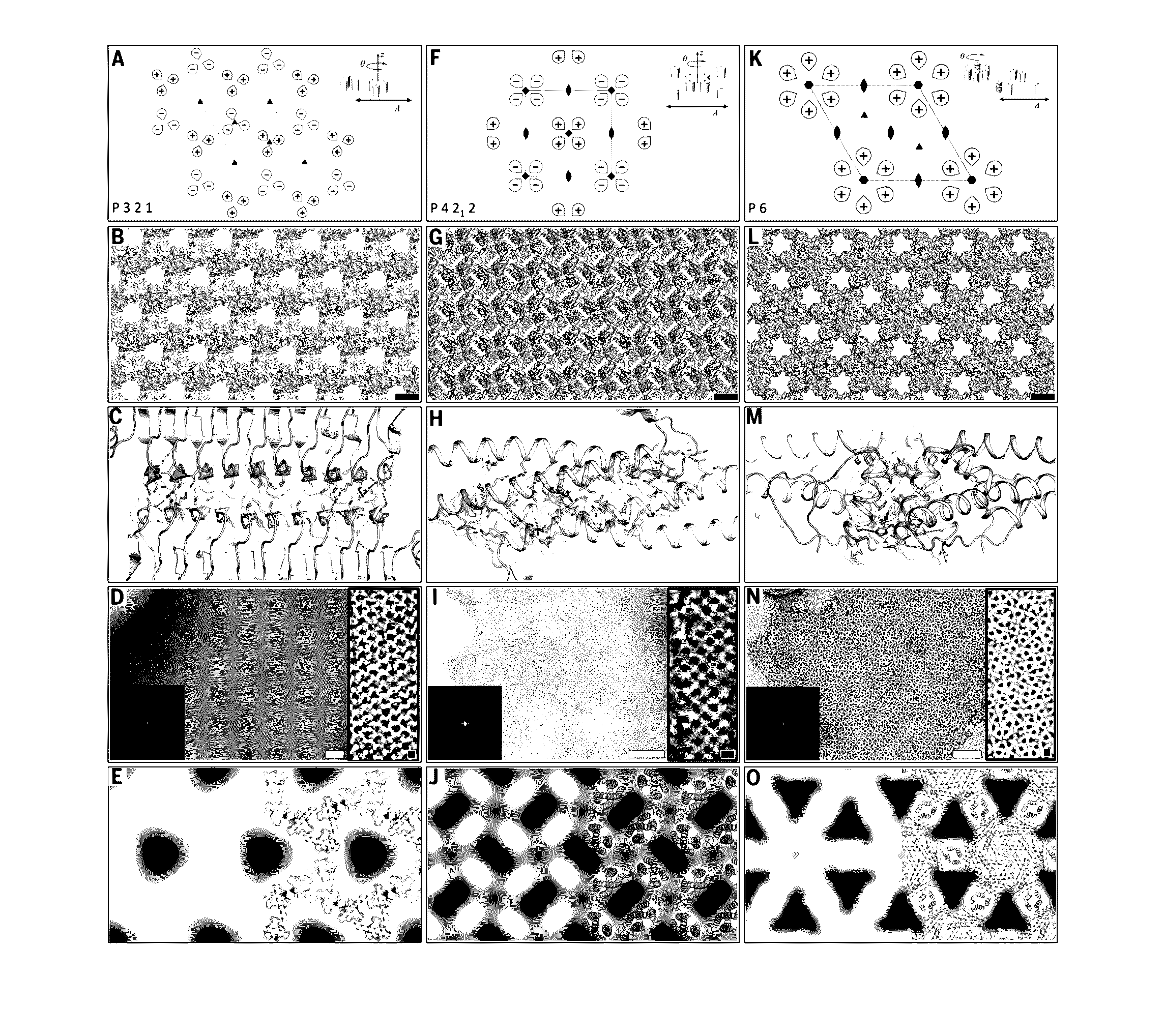
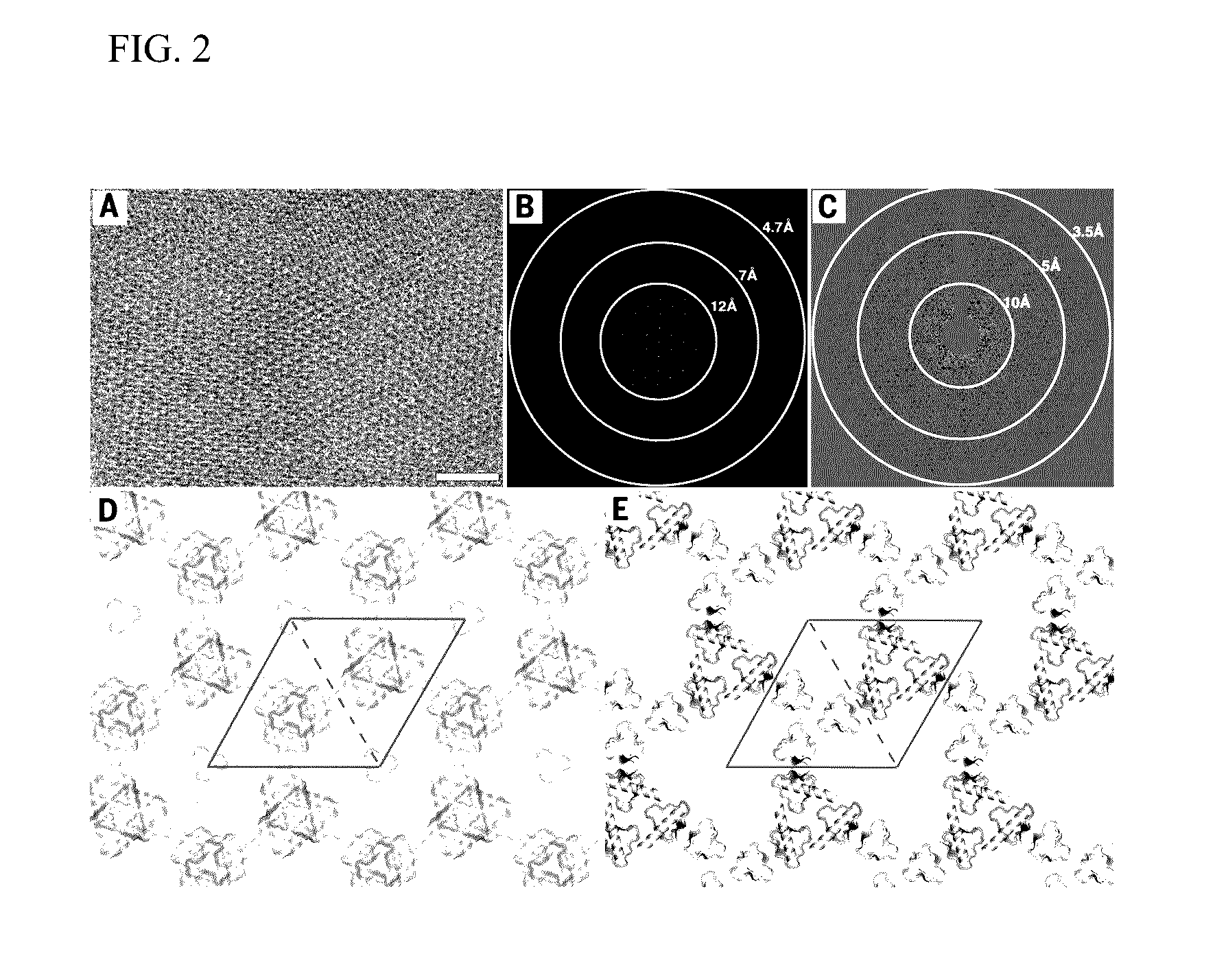
Self-assembling two-dimensional protein arrays
PatentInactiveUS20160369264A1
Innovation
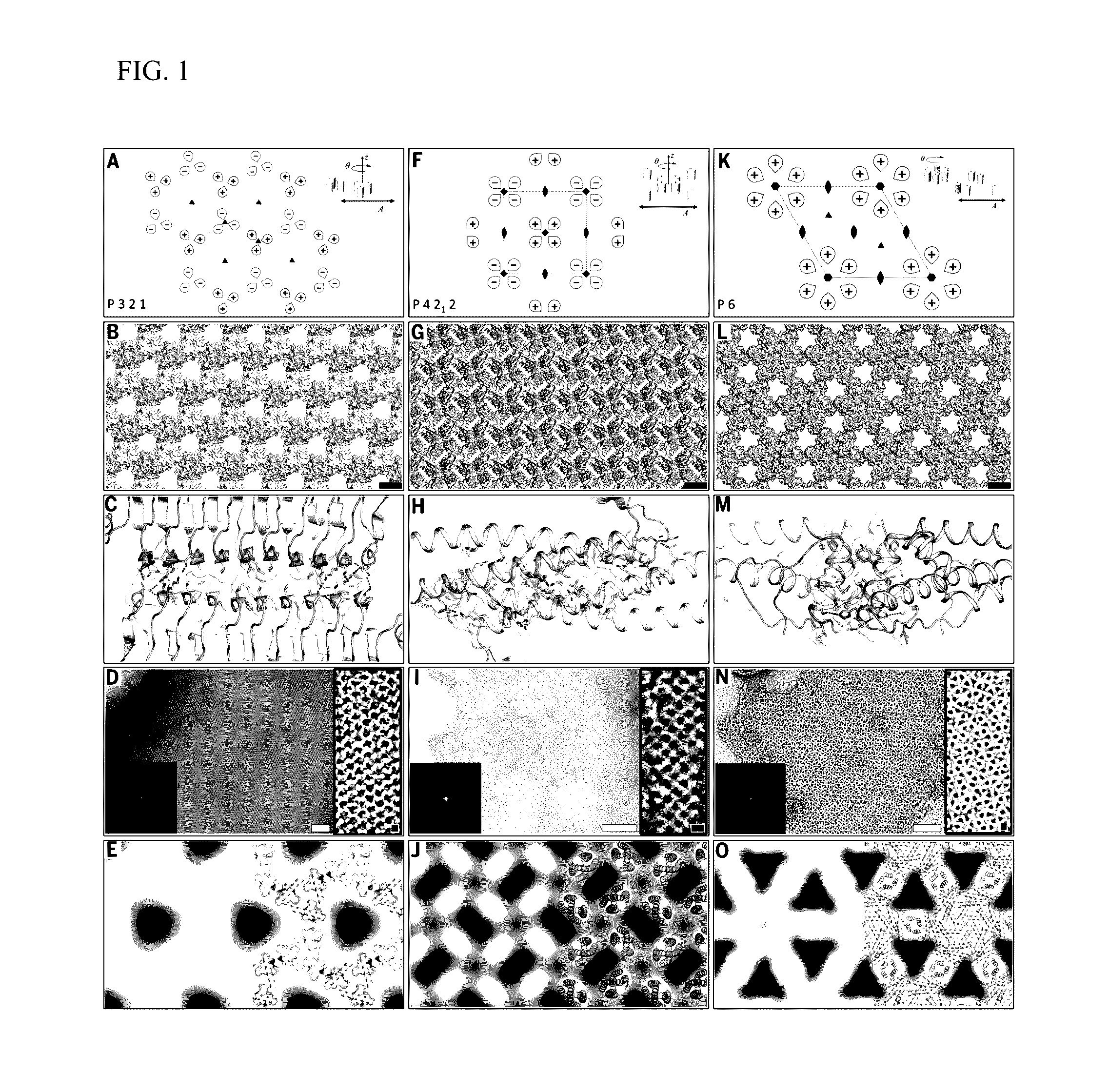
- The development of 2D protein arrays composed of oligomeric protein unit cells with rotational symmetry, mediated by noncovalent protein-protein interfaces, allowing for the formation of ordered structures that can diffract electrons or X-rays at higher resolutions, using self-assembling proteins like p3Z_11, p3Z_42, p4Z_9, and p6_9H, which can be expressed in Escherichia coli and assembled intracellularly or extracellularly.
Computational Tools for Solvation Analysis
Computational tools play a crucial role in analyzing protein solvation shell structures and understanding the effects of geometric isomerism. These tools have evolved significantly over the years, offering researchers powerful means to investigate complex molecular interactions.
Molecular dynamics (MD) simulations are among the most widely used computational methods for studying protein solvation. These simulations allow researchers to model the behavior of proteins and surrounding water molecules over time, providing insights into the formation and dynamics of solvation shells. Advanced MD software packages, such as GROMACS, NAMD, and AMBER, offer specialized force fields and algorithms optimized for protein-solvent interactions.
Quantum mechanical (QM) methods, including density functional theory (DFT) and ab initio calculations, provide a more detailed understanding of electronic structures and interactions at the atomic level. These approaches are particularly useful for investigating the effects of geometric isomerism on local solvation structures. Software packages like Gaussian and ORCA enable researchers to perform high-level QM calculations on protein-solvent systems.
Hybrid quantum mechanics/molecular mechanics (QM/MM) methods combine the accuracy of QM calculations with the efficiency of MM simulations. This approach allows for the detailed study of specific regions of interest within a protein while treating the rest of the system at a lower level of theory. Tools such as CHARMM and CP2K implement QM/MM methods for solvation analysis.
Continuum solvation models, such as the Poisson-Boltzmann (PB) equation and generalized Born (GB) methods, offer computationally efficient alternatives for estimating solvation free energies. These models treat the solvent as a continuous medium rather than individual molecules. Software packages like DelPhi and APBS implement these methods for rapid solvation calculations.
Machine learning (ML) approaches have recently emerged as powerful tools for analyzing protein solvation. These methods can predict solvation properties based on large datasets of experimental and computational results. Packages like DeepSolv and AquaSol utilize neural networks and other ML algorithms to model protein-solvent interactions.
Visualization tools are essential for interpreting the results of solvation analyses. Software like VMD, PyMOL, and Chimera allow researchers to visualize protein structures, solvation shells, and isomeric conformations in three dimensions. These tools often integrate with simulation packages, enabling real-time visualization of MD trajectories.
Data analysis and statistical tools are crucial for processing the large amounts of data generated by solvation simulations. Packages like MDAnalysis and MDTraj provide specialized functions for analyzing MD trajectories, while general-purpose scientific computing environments like Python with NumPy and SciPy offer flexible data processing capabilities.
Molecular dynamics (MD) simulations are among the most widely used computational methods for studying protein solvation. These simulations allow researchers to model the behavior of proteins and surrounding water molecules over time, providing insights into the formation and dynamics of solvation shells. Advanced MD software packages, such as GROMACS, NAMD, and AMBER, offer specialized force fields and algorithms optimized for protein-solvent interactions.
Quantum mechanical (QM) methods, including density functional theory (DFT) and ab initio calculations, provide a more detailed understanding of electronic structures and interactions at the atomic level. These approaches are particularly useful for investigating the effects of geometric isomerism on local solvation structures. Software packages like Gaussian and ORCA enable researchers to perform high-level QM calculations on protein-solvent systems.
Hybrid quantum mechanics/molecular mechanics (QM/MM) methods combine the accuracy of QM calculations with the efficiency of MM simulations. This approach allows for the detailed study of specific regions of interest within a protein while treating the rest of the system at a lower level of theory. Tools such as CHARMM and CP2K implement QM/MM methods for solvation analysis.
Continuum solvation models, such as the Poisson-Boltzmann (PB) equation and generalized Born (GB) methods, offer computationally efficient alternatives for estimating solvation free energies. These models treat the solvent as a continuous medium rather than individual molecules. Software packages like DelPhi and APBS implement these methods for rapid solvation calculations.
Machine learning (ML) approaches have recently emerged as powerful tools for analyzing protein solvation. These methods can predict solvation properties based on large datasets of experimental and computational results. Packages like DeepSolv and AquaSol utilize neural networks and other ML algorithms to model protein-solvent interactions.
Visualization tools are essential for interpreting the results of solvation analyses. Software like VMD, PyMOL, and Chimera allow researchers to visualize protein structures, solvation shells, and isomeric conformations in three dimensions. These tools often integrate with simulation packages, enabling real-time visualization of MD trajectories.
Data analysis and statistical tools are crucial for processing the large amounts of data generated by solvation simulations. Packages like MDAnalysis and MDTraj provide specialized functions for analyzing MD trajectories, while general-purpose scientific computing environments like Python with NumPy and SciPy offer flexible data processing capabilities.
Implications for Drug Design and Delivery
The implications of geometric isomerism on protein solvation shell structures have significant ramifications for drug design and delivery. Understanding these effects can lead to more effective and targeted pharmaceutical interventions. The spatial arrangement of atoms in isomers directly influences their interactions with water molecules, which in turn affects the protein's solvation shell. This knowledge can be leveraged to optimize drug-protein interactions and enhance drug efficacy.
One key implication is the potential for improved drug binding affinity. By considering the geometric isomerism of both the drug molecule and the target protein, researchers can design compounds that better complement the protein's solvation shell structure. This approach may lead to more stable drug-protein complexes and potentially lower required dosages, reducing side effects and improving patient outcomes.
Furthermore, the impact of geometric isomerism on solvation shells can inform strategies for enhancing drug solubility and bioavailability. Isomers with favorable interactions with water molecules may exhibit improved solubility profiles, facilitating better absorption and distribution within the body. This knowledge can guide the development of more effective drug formulations and delivery systems.
The understanding of how geometric isomerism affects protein solvation shells also has implications for the design of prodrugs. By manipulating the isomeric form of a drug molecule, it may be possible to create compounds that remain inactive in the solvated state but become active upon reaching their target site. This approach could lead to more targeted drug delivery and reduced off-target effects.
Additionally, this knowledge can inform the development of novel drug delivery systems. By considering the solvation properties of different isomers, researchers can design carrier molecules or nanoparticles that better protect and transport drugs through the body. This could result in improved drug stability, controlled release mechanisms, and enhanced targeting to specific tissues or organs.
The implications extend to the field of personalized medicine as well. As our understanding of individual genetic variations in protein structures grows, the ability to tailor drugs based on geometric isomerism and solvation effects could lead to more personalized and effective treatments. This approach may help address issues of drug resistance and improve outcomes for patients with specific genetic profiles.
In conclusion, the insights gained from studying how geometric isomerism affects protein solvation shell structures have far-reaching implications for drug design and delivery. By incorporating this knowledge into pharmaceutical research and development processes, we can anticipate more efficient, targeted, and effective therapeutic interventions in the future.
One key implication is the potential for improved drug binding affinity. By considering the geometric isomerism of both the drug molecule and the target protein, researchers can design compounds that better complement the protein's solvation shell structure. This approach may lead to more stable drug-protein complexes and potentially lower required dosages, reducing side effects and improving patient outcomes.
Furthermore, the impact of geometric isomerism on solvation shells can inform strategies for enhancing drug solubility and bioavailability. Isomers with favorable interactions with water molecules may exhibit improved solubility profiles, facilitating better absorption and distribution within the body. This knowledge can guide the development of more effective drug formulations and delivery systems.
The understanding of how geometric isomerism affects protein solvation shells also has implications for the design of prodrugs. By manipulating the isomeric form of a drug molecule, it may be possible to create compounds that remain inactive in the solvated state but become active upon reaching their target site. This approach could lead to more targeted drug delivery and reduced off-target effects.
Additionally, this knowledge can inform the development of novel drug delivery systems. By considering the solvation properties of different isomers, researchers can design carrier molecules or nanoparticles that better protect and transport drugs through the body. This could result in improved drug stability, controlled release mechanisms, and enhanced targeting to specific tissues or organs.
The implications extend to the field of personalized medicine as well. As our understanding of individual genetic variations in protein structures grows, the ability to tailor drugs based on geometric isomerism and solvation effects could lead to more personalized and effective treatments. This approach may help address issues of drug resistance and improve outcomes for patients with specific genetic profiles.
In conclusion, the insights gained from studying how geometric isomerism affects protein solvation shell structures have far-reaching implications for drug design and delivery. By incorporating this knowledge into pharmaceutical research and development processes, we can anticipate more efficient, targeted, and effective therapeutic interventions in the future.
Unlock deeper insights with PatSnap Eureka Quick Research — get a full tech report to explore trends and direct your research. Try now!
Generate Your Research Report Instantly with AI Agent
Supercharge your innovation with PatSnap Eureka AI Agent Platform!