Compare Phosphorylation Effects Across Cell Types
SEP 23, 202510 MIN READ
Generate Your Research Report Instantly with AI Agent
PatSnap Eureka helps you evaluate technical feasibility & market potential.
Phosphorylation Mechanisms and Research Objectives
Phosphorylation represents one of the most critical post-translational modifications in cellular biology, serving as a fundamental regulatory mechanism across diverse biological processes. This reversible biochemical process involves the addition of a phosphate group to proteins, primarily on serine, threonine, or tyrosine residues, catalyzed by protein kinases and reversed by protein phosphatases. The historical trajectory of phosphorylation research began with pioneering work by Edmond Fischer and Edwin Krebs in the 1950s, who discovered phosphorylase kinase, establishing the foundation for our understanding of enzymatic phosphorylation.
The evolution of phosphorylation research has progressed through several distinct phases, from initial biochemical characterization to current systems biology approaches. Early research focused on individual phosphorylation events, while contemporary studies employ high-throughput phosphoproteomics to map complex phosphorylation networks across different cell types. Recent technological advancements, particularly in mass spectrometry and computational biology, have revolutionized our ability to detect and quantify phosphorylation events with unprecedented precision and scale.
Cell-type specific phosphorylation patterns represent an emerging frontier in molecular biology research. Different cell types exhibit unique phosphorylation signatures that reflect their specialized functions and regulatory requirements. These distinctive phosphorylation profiles arise from variations in kinase and phosphatase expression, subcellular localization, and activation states across different cellular contexts. Understanding these cell-type specific differences is crucial for deciphering how identical signaling pathways can produce diverse outcomes in different tissues.
The primary objective of comparative phosphorylation research is to systematically characterize phosphorylation dynamics across diverse cell types under both normal and pathological conditions. This includes mapping phosphorylation sites, identifying responsible kinases and phosphatases, and elucidating downstream functional consequences. By comparing phosphorylation networks across cell types, researchers aim to identify both conserved regulatory mechanisms and cell-specific adaptations that contribute to specialized cellular functions.
Technical goals in this field include developing standardized methodologies for phosphoproteomic analysis that enable direct comparison between cell types, creating comprehensive phosphorylation atlases across human tissues, and establishing computational frameworks for predicting cell-type specific phosphorylation outcomes. Additionally, researchers seek to integrate phosphorylation data with other omics datasets to construct holistic models of cellular regulation that account for cell-type variability.
The ultimate translational objective is to leverage cell-type specific phosphorylation knowledge to develop more precise therapeutic strategies. By understanding how phosphorylation networks differ between normal and diseased cells, and between different cell types, researchers can identify novel drug targets with enhanced specificity and reduced off-target effects, potentially revolutionizing treatment approaches for cancer, neurodegenerative disorders, and other diseases characterized by dysregulated phosphorylation.
The evolution of phosphorylation research has progressed through several distinct phases, from initial biochemical characterization to current systems biology approaches. Early research focused on individual phosphorylation events, while contemporary studies employ high-throughput phosphoproteomics to map complex phosphorylation networks across different cell types. Recent technological advancements, particularly in mass spectrometry and computational biology, have revolutionized our ability to detect and quantify phosphorylation events with unprecedented precision and scale.
Cell-type specific phosphorylation patterns represent an emerging frontier in molecular biology research. Different cell types exhibit unique phosphorylation signatures that reflect their specialized functions and regulatory requirements. These distinctive phosphorylation profiles arise from variations in kinase and phosphatase expression, subcellular localization, and activation states across different cellular contexts. Understanding these cell-type specific differences is crucial for deciphering how identical signaling pathways can produce diverse outcomes in different tissues.
The primary objective of comparative phosphorylation research is to systematically characterize phosphorylation dynamics across diverse cell types under both normal and pathological conditions. This includes mapping phosphorylation sites, identifying responsible kinases and phosphatases, and elucidating downstream functional consequences. By comparing phosphorylation networks across cell types, researchers aim to identify both conserved regulatory mechanisms and cell-specific adaptations that contribute to specialized cellular functions.
Technical goals in this field include developing standardized methodologies for phosphoproteomic analysis that enable direct comparison between cell types, creating comprehensive phosphorylation atlases across human tissues, and establishing computational frameworks for predicting cell-type specific phosphorylation outcomes. Additionally, researchers seek to integrate phosphorylation data with other omics datasets to construct holistic models of cellular regulation that account for cell-type variability.
The ultimate translational objective is to leverage cell-type specific phosphorylation knowledge to develop more precise therapeutic strategies. By understanding how phosphorylation networks differ between normal and diseased cells, and between different cell types, researchers can identify novel drug targets with enhanced specificity and reduced off-target effects, potentially revolutionizing treatment approaches for cancer, neurodegenerative disorders, and other diseases characterized by dysregulated phosphorylation.
Market Applications of Cell-Specific Phosphorylation Research
The market for cell-specific phosphorylation research is experiencing significant growth, driven by increasing demand for precision medicine and personalized therapeutic approaches. Pharmaceutical companies are heavily investing in technologies that can identify cell type-specific phosphorylation patterns to develop targeted drugs with reduced side effects. This market segment is projected to reach several billion dollars by 2028, with an annual growth rate exceeding the broader biotechnology sector.
Diagnostic applications represent another substantial market opportunity. The ability to detect abnormal phosphorylation patterns in specific cell types offers powerful biomarkers for early disease detection, particularly in cancer, neurodegenerative disorders, and autoimmune conditions. Companies developing diagnostic platforms based on cell-specific phosphorylation signatures are attracting significant venture capital, with several successful funding rounds reported in the past two years.
The agricultural biotechnology sector has begun exploring cell-specific phosphorylation research to develop crops with enhanced stress resistance and yield. By understanding how phosphorylation differs across plant cell types under various environmental conditions, companies are creating more resilient crop varieties. This application area is expected to grow substantially as climate change intensifies the need for adaptive agricultural solutions.
Research tools and reagents for studying cell-specific phosphorylation constitute a rapidly expanding market segment. Suppliers of antibodies, phospho-specific dyes, mass spectrometry equipment, and single-cell analysis platforms are experiencing increased demand from both academic and industrial laboratories. The market for these specialized research tools is becoming increasingly competitive, with several companies developing proprietary technologies to gain market advantage.
Contract research organizations (CROs) specializing in phosphoproteomics across different cell types are emerging as key service providers in the biotechnology ecosystem. These organizations offer expertise and high-throughput capabilities that many smaller companies and academic institutions cannot maintain in-house. The global market for phosphoproteomics services is growing at approximately twice the rate of general proteomics services.
Bioinformatics and artificial intelligence applications for analyzing cell-specific phosphorylation data represent a high-growth market niche. Software platforms capable of integrating multi-omics data to predict cell type-specific responses to drugs based on phosphorylation patterns are becoming essential tools in drug discovery pipelines. Several technology startups focused exclusively on this analytical challenge have secured significant funding in recent funding rounds.
Diagnostic applications represent another substantial market opportunity. The ability to detect abnormal phosphorylation patterns in specific cell types offers powerful biomarkers for early disease detection, particularly in cancer, neurodegenerative disorders, and autoimmune conditions. Companies developing diagnostic platforms based on cell-specific phosphorylation signatures are attracting significant venture capital, with several successful funding rounds reported in the past two years.
The agricultural biotechnology sector has begun exploring cell-specific phosphorylation research to develop crops with enhanced stress resistance and yield. By understanding how phosphorylation differs across plant cell types under various environmental conditions, companies are creating more resilient crop varieties. This application area is expected to grow substantially as climate change intensifies the need for adaptive agricultural solutions.
Research tools and reagents for studying cell-specific phosphorylation constitute a rapidly expanding market segment. Suppliers of antibodies, phospho-specific dyes, mass spectrometry equipment, and single-cell analysis platforms are experiencing increased demand from both academic and industrial laboratories. The market for these specialized research tools is becoming increasingly competitive, with several companies developing proprietary technologies to gain market advantage.
Contract research organizations (CROs) specializing in phosphoproteomics across different cell types are emerging as key service providers in the biotechnology ecosystem. These organizations offer expertise and high-throughput capabilities that many smaller companies and academic institutions cannot maintain in-house. The global market for phosphoproteomics services is growing at approximately twice the rate of general proteomics services.
Bioinformatics and artificial intelligence applications for analyzing cell-specific phosphorylation data represent a high-growth market niche. Software platforms capable of integrating multi-omics data to predict cell type-specific responses to drugs based on phosphorylation patterns are becoming essential tools in drug discovery pipelines. Several technology startups focused exclusively on this analytical challenge have secured significant funding in recent funding rounds.
Current Challenges in Cross-Cell Phosphorylation Analysis
Despite significant advancements in phosphoproteomics technologies, cross-cell type comparison of phosphorylation events remains fraught with methodological and analytical challenges. One of the primary obstacles is the inherent biological variability between different cell types, which can mask or confound true phosphorylation differences related to specific cellular functions or disease states. Even within the same tissue, heterogeneous cell populations exhibit distinct phosphorylation profiles that reflect their specialized roles and microenvironments.
Sample preparation inconsistencies represent another major hurdle in cross-cell phosphorylation analysis. Different cell types often require specialized lysis conditions and phosphopeptide enrichment protocols, introducing technical variability that complicates direct comparisons. The efficiency of phosphopeptide enrichment can vary dramatically depending on cell type-specific protein composition and abundance, leading to systematic biases in phosphoproteomic datasets.
Mass spectrometry-based detection faces sensitivity limitations when dealing with low-abundance phosphorylation events, which are often biologically significant but technically challenging to quantify consistently across different cell types. This challenge is compounded by the dynamic nature of phosphorylation, where rapid turnover rates can result in temporal variations that are difficult to control for in experimental designs spanning multiple cell types.
Computational analysis of cross-cell phosphoproteomics data presents its own set of challenges. Current normalization methods struggle to account for cell type-specific differences in total phosphorylation levels and kinase activity profiles. Statistical approaches developed for single-cell type analyses often perform poorly when applied to heterogeneous datasets, leading to high false discovery rates or missed biologically relevant differences.
The contextual interpretation of phosphorylation events across cell types remains particularly problematic. A phosphorylation site may have dramatically different functional consequences depending on the cellular context, yet current databases and annotation systems lack the granularity to capture these context-dependent effects. This limitation severely hampers our ability to translate phosphoproteomic findings into meaningful biological insights.
Integration of phosphorylation data with other omics layers (transcriptomics, metabolomics) across cell types introduces additional complexity. Current multi-omics integration frameworks typically assume consistent relationships between different molecular layers, an assumption that often breaks down when comparing diverse cell types with fundamentally different regulatory architectures.
Standardization efforts in the field remain insufficient, with inconsistent reporting of experimental conditions, analytical parameters, and validation approaches across studies. This lack of standardization impedes meta-analyses and the development of robust computational models that could potentially overcome some of the aforementioned challenges in cross-cell phosphorylation analysis.
Sample preparation inconsistencies represent another major hurdle in cross-cell phosphorylation analysis. Different cell types often require specialized lysis conditions and phosphopeptide enrichment protocols, introducing technical variability that complicates direct comparisons. The efficiency of phosphopeptide enrichment can vary dramatically depending on cell type-specific protein composition and abundance, leading to systematic biases in phosphoproteomic datasets.
Mass spectrometry-based detection faces sensitivity limitations when dealing with low-abundance phosphorylation events, which are often biologically significant but technically challenging to quantify consistently across different cell types. This challenge is compounded by the dynamic nature of phosphorylation, where rapid turnover rates can result in temporal variations that are difficult to control for in experimental designs spanning multiple cell types.
Computational analysis of cross-cell phosphoproteomics data presents its own set of challenges. Current normalization methods struggle to account for cell type-specific differences in total phosphorylation levels and kinase activity profiles. Statistical approaches developed for single-cell type analyses often perform poorly when applied to heterogeneous datasets, leading to high false discovery rates or missed biologically relevant differences.
The contextual interpretation of phosphorylation events across cell types remains particularly problematic. A phosphorylation site may have dramatically different functional consequences depending on the cellular context, yet current databases and annotation systems lack the granularity to capture these context-dependent effects. This limitation severely hampers our ability to translate phosphoproteomic findings into meaningful biological insights.
Integration of phosphorylation data with other omics layers (transcriptomics, metabolomics) across cell types introduces additional complexity. Current multi-omics integration frameworks typically assume consistent relationships between different molecular layers, an assumption that often breaks down when comparing diverse cell types with fundamentally different regulatory architectures.
Standardization efforts in the field remain insufficient, with inconsistent reporting of experimental conditions, analytical parameters, and validation approaches across studies. This lack of standardization impedes meta-analyses and the development of robust computational models that could potentially overcome some of the aforementioned challenges in cross-cell phosphorylation analysis.
Established Protocols for Comparative Phosphorylation Studies
01 Phosphorylation in signal transduction pathways
Phosphorylation plays a crucial role in cellular signal transduction pathways, regulating various biological processes. The addition of phosphate groups to proteins by kinases can activate or deactivate signaling cascades, affecting cell growth, differentiation, and survival. These phosphorylation events serve as molecular switches that control the flow of information within cells and between cells, enabling appropriate responses to environmental stimuli.- Phosphorylation in signal transduction pathways: Phosphorylation plays a crucial role in cellular signal transduction pathways, regulating various biological processes. The addition of phosphate groups to proteins by kinases can activate or deactivate enzymes, alter protein-protein interactions, and trigger downstream signaling cascades. These phosphorylation events are essential for normal cell function and are often dysregulated in disease states, making them important targets for therapeutic intervention.
- Detection methods for protein phosphorylation: Various analytical techniques have been developed to detect and quantify protein phosphorylation states. These include antibody-based methods such as Western blotting and ELISA, mass spectrometry approaches, and phospho-specific fluorescent probes. These detection methods enable researchers to monitor phosphorylation events in real-time, identify novel phosphorylation sites, and understand the dynamics of phosphorylation in different cellular contexts.
- Phosphorylation in disease mechanisms: Aberrant phosphorylation patterns are implicated in numerous diseases, including cancer, neurodegenerative disorders, and metabolic syndromes. Hyperphosphorylation or hypophosphorylation of key proteins can disrupt normal cellular functions, leading to pathological conditions. Understanding these disease-associated phosphorylation events provides insights into disease mechanisms and identifies potential biomarkers for diagnosis and prognosis.
- Kinase inhibitors targeting phosphorylation: Kinase inhibitors are a class of therapeutic compounds designed to block specific phosphorylation events by targeting the enzymes responsible for adding phosphate groups to proteins. These inhibitors have emerged as important drugs for treating various diseases, particularly cancer, where dysregulated kinase activity drives tumor growth and progression. The development of selective kinase inhibitors requires understanding the structural and functional aspects of phosphorylation mechanisms.
- Phosphorylation in cellular metabolism regulation: Phosphorylation serves as a key regulatory mechanism in cellular metabolism, controlling enzyme activity, substrate availability, and metabolic flux. Reversible phosphorylation of metabolic enzymes allows cells to rapidly respond to changing energy demands and nutrient availability. This regulatory system is particularly important in processes such as glycolysis, gluconeogenesis, and lipid metabolism, where phosphorylation events can switch metabolic pathways on or off in response to hormonal and environmental signals.
02 Detection methods for protein phosphorylation
Various analytical techniques have been developed to detect and quantify protein phosphorylation states. These methods include phospho-specific antibodies, mass spectrometry, phospho-proteomics, and fluorescence-based assays. Such detection systems allow researchers to monitor changes in phosphorylation patterns in response to different stimuli or disease states, providing valuable insights into cellular regulation mechanisms and potential therapeutic targets.Expand Specific Solutions03 Phosphorylation in disease mechanisms
Abnormal phosphorylation patterns are implicated in various disease states, including cancer, neurodegenerative disorders, and metabolic diseases. Dysregulation of kinase activity or phosphatase function can lead to aberrant signaling, contributing to disease progression. Understanding these phosphorylation-related mechanisms provides opportunities for developing targeted therapies that can modulate specific phosphorylation events to restore normal cellular function.Expand Specific Solutions04 Therapeutic targeting of phosphorylation pathways
Phosphorylation pathways represent important therapeutic targets for drug development. Kinase inhibitors and phosphatase modulators can be designed to specifically alter phosphorylation states of key proteins involved in disease processes. These targeted approaches aim to normalize aberrant signaling pathways while minimizing effects on normal cellular functions. The development of such therapeutics requires detailed understanding of phosphorylation networks and their regulation in both healthy and diseased states.Expand Specific Solutions05 Phosphorylation in cellular metabolism and energy regulation
Phosphorylation is fundamental to cellular metabolism and energy regulation. Key metabolic enzymes are controlled through phosphorylation events that respond to the cell's energy status. This regulation affects processes such as glycolysis, gluconeogenesis, and lipid metabolism. Additionally, phosphorylation of proteins involved in mitochondrial function influences cellular energy production. These phosphorylation-dependent regulatory mechanisms allow cells to adapt their metabolic activities to changing energy demands and nutrient availability.Expand Specific Solutions
Leading Research Institutions and Biotech Companies
Phosphorylation effects across cell types represent a maturing research field with significant market growth potential, currently estimated at $2-3 billion annually and expanding at 8-10% CAGR. The competitive landscape features established pharmaceutical leaders (Roche, Novartis, Vertex Pharmaceuticals) alongside specialized research institutions (University of California, University of Michigan) and emerging biotechnology firms. Technical maturity varies across applications, with companies like Revvity Health Sciences and Ionis Pharmaceuticals advancing proprietary phosphorylation analysis platforms, while academic-industry partnerships drive innovation. The field is transitioning from basic research to clinical applications, with increasing focus on cell-type specific phosphorylation signatures for precision medicine applications in oncology, neurodegenerative diseases, and metabolic disorders.
Revvity Health Sciences, Inc.
Technical Solution: Revvity Health Sciences has developed a comprehensive phosphoproteomics platform that enables high-throughput comparison of phosphorylation patterns across diverse cell types. Their technology combines mass spectrometry-based phosphopeptide enrichment with advanced bioinformatics tools to identify cell type-specific phosphorylation signatures. The platform utilizes their proprietary PhosphoSignature™ technology that can simultaneously quantify thousands of phosphorylation sites with high sensitivity and reproducibility. Their approach incorporates stable isotope labeling with amino acids in cell culture (SILAC) methodology for accurate quantification and comparison between different cellular contexts. Revvity's system is particularly notable for its ability to detect low-abundance phosphoproteins and transient phosphorylation events that are often missed by conventional techniques.
Strengths: Industry-leading sensitivity and reproducibility in phosphoprotein detection; comprehensive coverage of phosphoproteome; integrated bioinformatics solutions for data interpretation. Weaknesses: High cost of implementation; requires specialized expertise; limited accessibility for smaller research institutions.
Ionis Pharmaceuticals, Inc.
Technical Solution: Ionis has developed PhosphoScan™, a specialized platform for analyzing phosphorylation differences across cell types with particular emphasis on antisense oligonucleotide (ASO) effects on cellular signaling. Their technology combines targeted phosphoproteomics with RNA sequencing to correlate changes in gene expression with alterations in phosphorylation patterns. The platform employs a unique approach to phosphopeptide enrichment that enhances detection of low-abundance regulatory phosphorylation events. Ionis has applied this technology to characterize cell type-specific responses to their antisense therapeutics, enabling more precise targeting of disease-relevant pathways. Their system includes a proprietary database of phosphorylation signatures across neuronal, hepatic, and muscle cell types, providing valuable context for therapeutic development. The platform's integration with Ionis' antisense technology has accelerated the development of targeted therapeutics for previously undruggable targets by identifying cell type-specific vulnerabilities in signaling networks.
Strengths: Unique integration with antisense technology; excellent capability for correlating RNA and protein phosphorylation changes; specialized focus on therapeutic-relevant phosphorylation events. Weaknesses: Narrower application scope compared to broader phosphoproteomics platforms; primarily optimized for antisense drug development.
Key Technologies in Phosphoproteomics
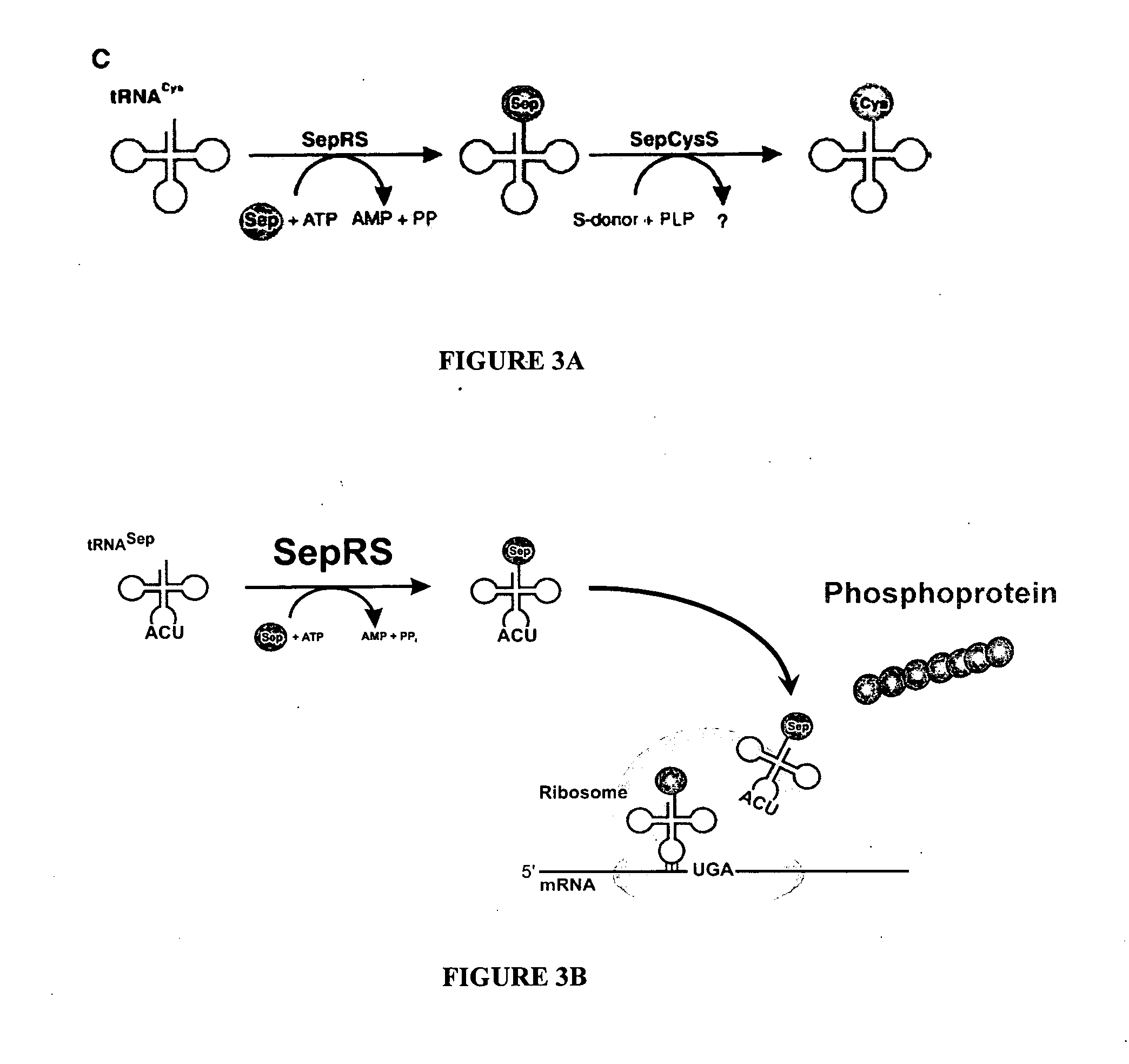
Site specific incorporation of phosphoserine into polypeptides using phosphoseryl-tRNA synthetase
PatentActiveUS20070077620A1
Innovation
- The use of nucleic acids encoding genes with SepRS and tRNASep activity, specifically the cysteinyl-tRNA from Methanocaldococcus jannaschii and the class II-type O-phosphoseryl-tRNA synthetase, to preferentially aminoacylate tRNASep with O-phosphoserine, allowing for the targeted incorporation of phosphoserine into proteins during translation.
Methods for target molecule detection using siderophores and related compositions
PatentInactiveEP1766076A2
Innovation
- The use of siderophores coordinated with transition metal cations as capture agents to selectively bind and isolate these molecules from samples, allowing for their detection and isolation at or near neutral pH, using various analytical methods such as fluorescence resonance energy transfer and inductively-coupled plasma mass spectrometry.
Regulatory Considerations in Phosphorylation-Based Therapeutics
The regulatory landscape for phosphorylation-based therapeutics presents unique challenges due to the complex and cell-type specific nature of phosphorylation events. Regulatory agencies, including the FDA and EMA, have established specific guidelines for the development and approval of drugs targeting phosphorylation pathways, recognizing their potential for both efficacy and off-target effects across different cell types.
When comparing phosphorylation effects across cell types, developers must address regulatory requirements for demonstrating cell-type specificity in preclinical studies. This includes comprehensive phosphoproteomic profiling across multiple tissue types to identify potential off-target phosphorylation events that could lead to adverse effects. The FDA's guidance on kinase inhibitor development specifically requires cross-cell type analysis to establish a safety profile.
Biomarker development and validation represents another critical regulatory consideration. Phosphorylation-based biomarkers must be validated across relevant cell types to ensure they accurately reflect drug activity and patient response. Regulatory bodies increasingly require companion diagnostics for phosphorylation-targeting therapeutics to identify appropriate patient populations based on cell-type specific phosphorylation profiles.
Manufacturing consistency presents unique challenges for phosphorylation-based therapeutics. Regulatory agencies require robust analytical methods to ensure batch-to-batch consistency in the drug's ability to modulate phosphorylation across different cell types. This includes validated assays that can detect subtle changes in phosphorylation patterns that might impact efficacy or safety.
Clinical trial design for phosphorylation-targeting drugs must account for cell-type variability. Regulatory guidelines recommend stratification of patient populations based on tissue-specific phosphorylation profiles and inclusion of pharmacodynamic endpoints that measure phosphorylation changes across multiple tissue types when feasible.
Post-market surveillance requirements are typically more stringent for phosphorylation-targeting therapeutics due to the potential for delayed off-target effects in specific cell populations. Regulatory agencies often mandate long-term monitoring programs to detect rare adverse events that might emerge from altered phosphorylation in non-target tissues.
International regulatory harmonization efforts are underway to standardize approaches to phosphorylation-based therapeutic development. The International Council for Harmonisation (ICH) has initiated working groups focused on establishing consistent requirements for demonstrating phosphorylation specificity across cell types, with particular emphasis on oncology applications where tissue-specific effects are critical for both efficacy and safety.
When comparing phosphorylation effects across cell types, developers must address regulatory requirements for demonstrating cell-type specificity in preclinical studies. This includes comprehensive phosphoproteomic profiling across multiple tissue types to identify potential off-target phosphorylation events that could lead to adverse effects. The FDA's guidance on kinase inhibitor development specifically requires cross-cell type analysis to establish a safety profile.
Biomarker development and validation represents another critical regulatory consideration. Phosphorylation-based biomarkers must be validated across relevant cell types to ensure they accurately reflect drug activity and patient response. Regulatory bodies increasingly require companion diagnostics for phosphorylation-targeting therapeutics to identify appropriate patient populations based on cell-type specific phosphorylation profiles.
Manufacturing consistency presents unique challenges for phosphorylation-based therapeutics. Regulatory agencies require robust analytical methods to ensure batch-to-batch consistency in the drug's ability to modulate phosphorylation across different cell types. This includes validated assays that can detect subtle changes in phosphorylation patterns that might impact efficacy or safety.
Clinical trial design for phosphorylation-targeting drugs must account for cell-type variability. Regulatory guidelines recommend stratification of patient populations based on tissue-specific phosphorylation profiles and inclusion of pharmacodynamic endpoints that measure phosphorylation changes across multiple tissue types when feasible.
Post-market surveillance requirements are typically more stringent for phosphorylation-targeting therapeutics due to the potential for delayed off-target effects in specific cell populations. Regulatory agencies often mandate long-term monitoring programs to detect rare adverse events that might emerge from altered phosphorylation in non-target tissues.
International regulatory harmonization efforts are underway to standardize approaches to phosphorylation-based therapeutic development. The International Council for Harmonisation (ICH) has initiated working groups focused on establishing consistent requirements for demonstrating phosphorylation specificity across cell types, with particular emphasis on oncology applications where tissue-specific effects are critical for both efficacy and safety.
Data Integration Strategies for Multi-Cell Phosphorylation Analysis
Integrating phosphorylation data across multiple cell types presents significant challenges due to the inherent biological variability and technical differences in data collection methodologies. Effective integration strategies must address these challenges while preserving the biological significance of phosphorylation events in different cellular contexts.
Mass spectrometry-based phosphoproteomics generates vast datasets that require sophisticated computational approaches for meaningful integration. Normalization techniques specifically designed for phosphorylation data are essential, as standard RNA-seq or proteomics normalization methods often fail to account for the unique characteristics of phosphorylation dynamics. Quantile normalization and variance stabilization transformations have shown promise when adapted for phosphoproteomics data from diverse cell types.
Batch effect correction represents another critical component of multi-cell phosphorylation analysis. Methods such as ComBat and Surrogate Variable Analysis (SVA) have been successfully applied to phosphoproteomics data, enabling the identification of technical variations while preserving biological differences between cell types. Recent advances in deep learning approaches, particularly variational autoencoders, demonstrate superior performance in handling non-linear batch effects common in phosphoproteomics data.
Dimensionality reduction techniques facilitate the visualization and interpretation of integrated phosphorylation datasets. While Principal Component Analysis (PCA) remains widely used, t-SNE and UMAP have gained popularity for their ability to preserve local structures in high-dimensional phosphorylation data. These methods help identify cell type-specific phosphorylation signatures and common regulatory patterns across different cellular contexts.
Network-based integration approaches leverage protein-protein interaction networks and kinase-substrate relationships to contextualize phosphorylation events across cell types. Algorithms such as NetworKIN and KinomeXplorer incorporate prior knowledge of signaling pathways to predict kinase activities from phosphoproteomics data, enabling comparative analysis of signaling networks across different cell types. These approaches have proven particularly valuable for identifying conserved and divergent signaling mechanisms.
Machine learning frameworks increasingly support multi-cell phosphorylation analysis through transfer learning paradigms. Models trained on phosphorylation data from well-characterized cell types can be adapted to predict phosphorylation patterns in less-studied cell types, maximizing the utility of existing datasets. Ensemble methods combining multiple integration strategies have demonstrated superior performance in benchmarking studies, suggesting that hybrid approaches may offer the most robust solution for comprehensive phosphorylation analysis across diverse cellular contexts.
Mass spectrometry-based phosphoproteomics generates vast datasets that require sophisticated computational approaches for meaningful integration. Normalization techniques specifically designed for phosphorylation data are essential, as standard RNA-seq or proteomics normalization methods often fail to account for the unique characteristics of phosphorylation dynamics. Quantile normalization and variance stabilization transformations have shown promise when adapted for phosphoproteomics data from diverse cell types.
Batch effect correction represents another critical component of multi-cell phosphorylation analysis. Methods such as ComBat and Surrogate Variable Analysis (SVA) have been successfully applied to phosphoproteomics data, enabling the identification of technical variations while preserving biological differences between cell types. Recent advances in deep learning approaches, particularly variational autoencoders, demonstrate superior performance in handling non-linear batch effects common in phosphoproteomics data.
Dimensionality reduction techniques facilitate the visualization and interpretation of integrated phosphorylation datasets. While Principal Component Analysis (PCA) remains widely used, t-SNE and UMAP have gained popularity for their ability to preserve local structures in high-dimensional phosphorylation data. These methods help identify cell type-specific phosphorylation signatures and common regulatory patterns across different cellular contexts.
Network-based integration approaches leverage protein-protein interaction networks and kinase-substrate relationships to contextualize phosphorylation events across cell types. Algorithms such as NetworKIN and KinomeXplorer incorporate prior knowledge of signaling pathways to predict kinase activities from phosphoproteomics data, enabling comparative analysis of signaling networks across different cell types. These approaches have proven particularly valuable for identifying conserved and divergent signaling mechanisms.
Machine learning frameworks increasingly support multi-cell phosphorylation analysis through transfer learning paradigms. Models trained on phosphorylation data from well-characterized cell types can be adapted to predict phosphorylation patterns in less-studied cell types, maximizing the utility of existing datasets. Ensemble methods combining multiple integration strategies have demonstrated superior performance in benchmarking studies, suggesting that hybrid approaches may offer the most robust solution for comprehensive phosphorylation analysis across diverse cellular contexts.
Unlock deeper insights with PatSnap Eureka Quick Research — get a full tech report to explore trends and direct your research. Try now!
Generate Your Research Report Instantly with AI Agent
Supercharge your innovation with PatSnap Eureka AI Agent Platform!