Phosphatases with improved phytase activity
A phytase and phosphatase technology, which can be used in plant genetic improvement, hydrolase, biochemical equipment and methods, etc., and can solve problems such as instability hindering practical application.
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment Embodiment 1
[0052] Example Example 1 Materials and methods used in Examples 2-6
[0053] Expression of r-AppA The appA gene was obtained from Escherichia coli BL21 strain transformed with the expression vector pAPPA1(20) (Genebank Accession No. M58708). A 1.35 kb DNA fragment containing the appA coding region was amplified by PCR according to the manufacturer's instructions (Perkin Elmer). Primers are derived from the 5' and 3' regions of this nucleotide sequence (18), and said primers include: E2 [forward: 242-252]: 5'GGAATTCCAGAGTGAGCCGGA3' (SEQ.ID.NO.2); and K2 [reverse: 1468-1490]: 5'GGGGTACCTTACAAACTGCACG3' (SEQ.ID.NO.3). Both primers were synthesized by the Cornell University Oligonucleotide Synthesis Facility (Ithaca, NY). The amplified product was excised from 1% agarose gel and eluted with GENECLEAN II kit (Bio101). The purified fragment was first cloned into pGEM T-easy vector (Promega), and then inserted into the yeast expression vector pPIcZαA (In...
Embodiment 2
[0057] In X33 transformed with the appA gene, more than 30 colonies expressed extracellular phytase activity that hydrolyzes sodium phytate. Colony 26 had the highest activity of this enzyme (88 U / ml) and was selected for further studies. After elution of r-PhyA and r-AppA samples from the DEAE-agarose column, 45 fractions of 4 ml each were collected for analysis of phytase activity. The phytase specific activities of r-PhyA and r-AppA of the fractions used for proteolysis were 9.6 U / mg protein and 8.1 U / mg protein, respectively. Example 3 Effects of Trypsin Digestion on the Phytase Activities of Two Enzymes
Embodiment 3
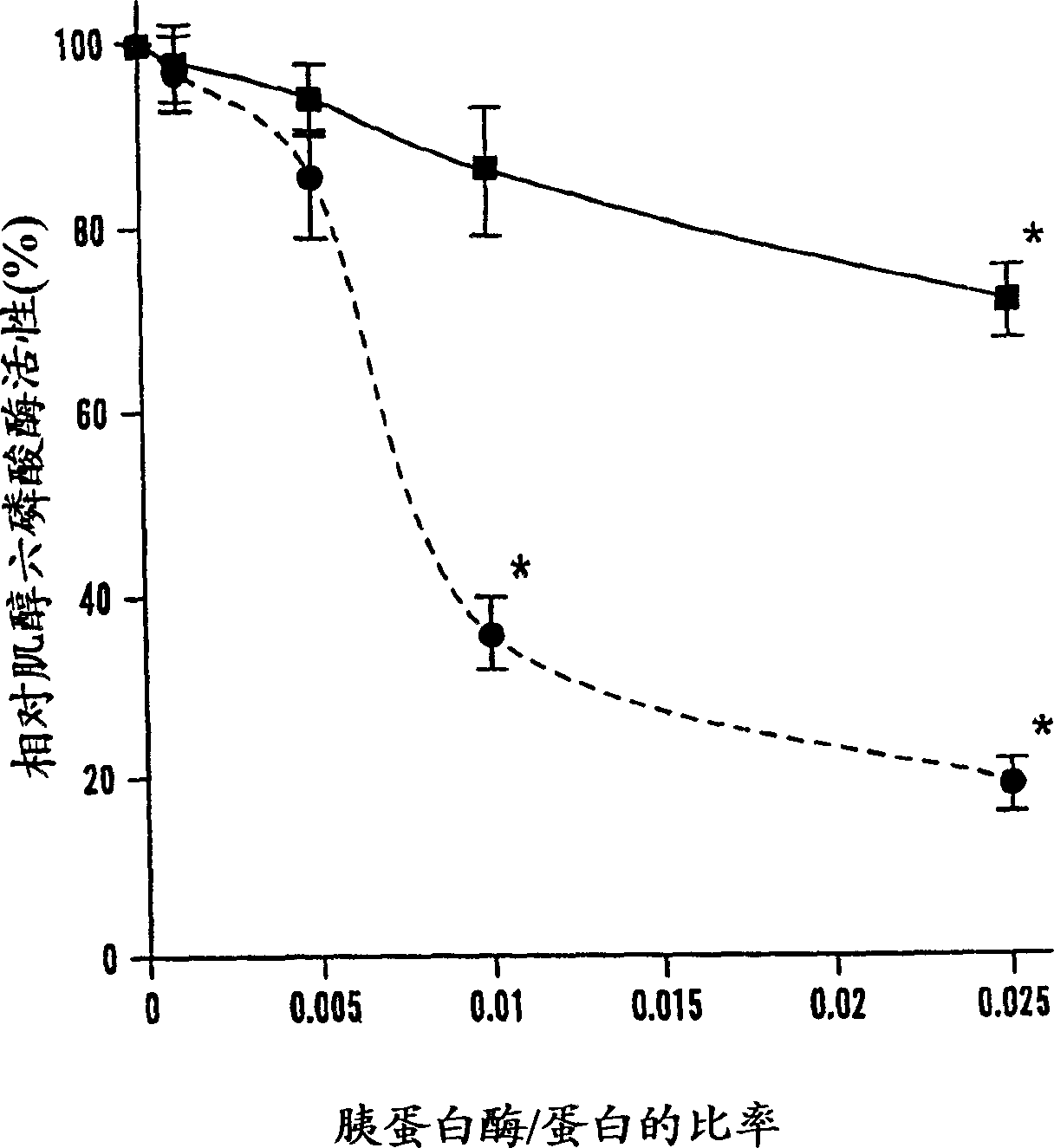
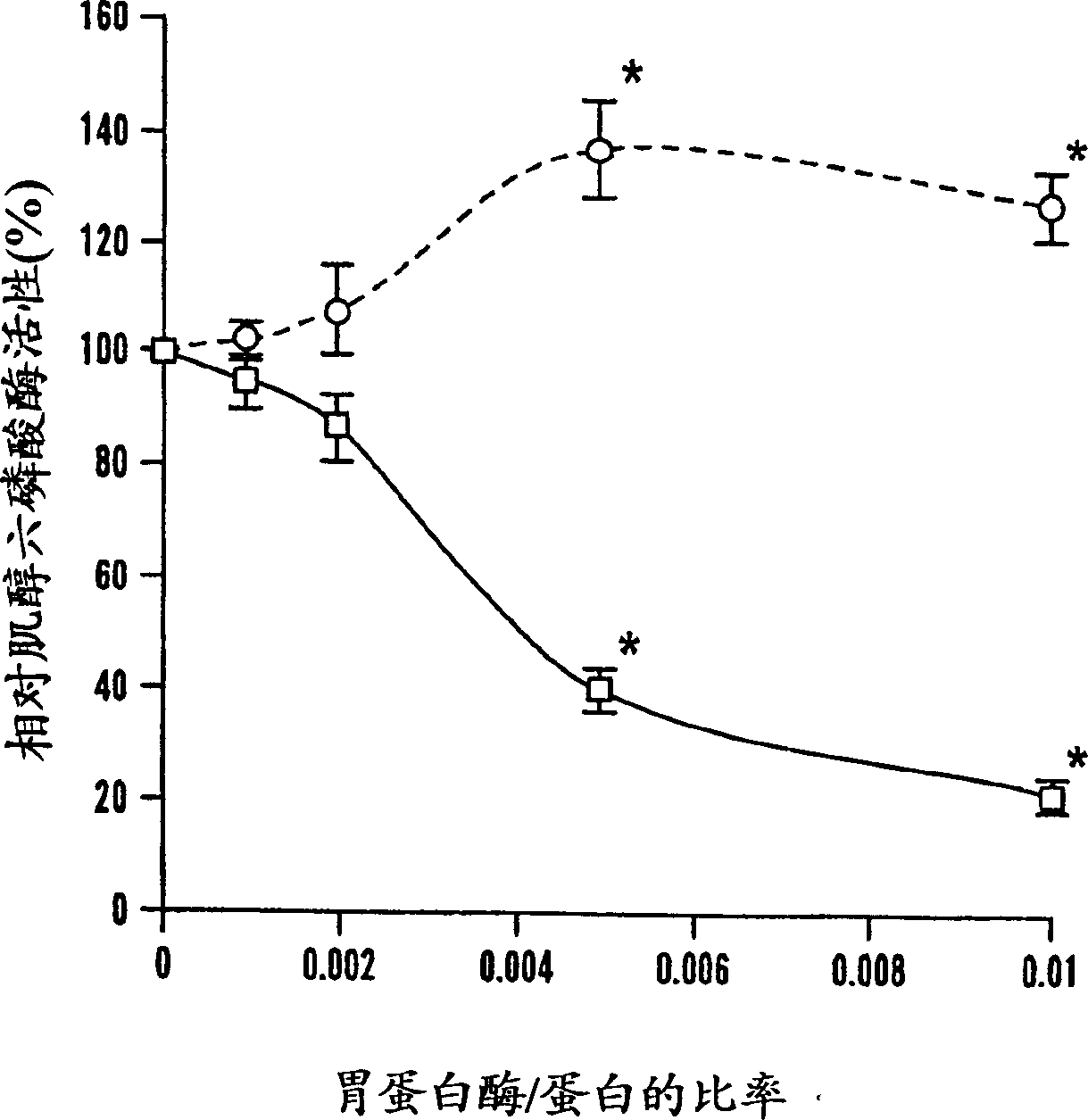
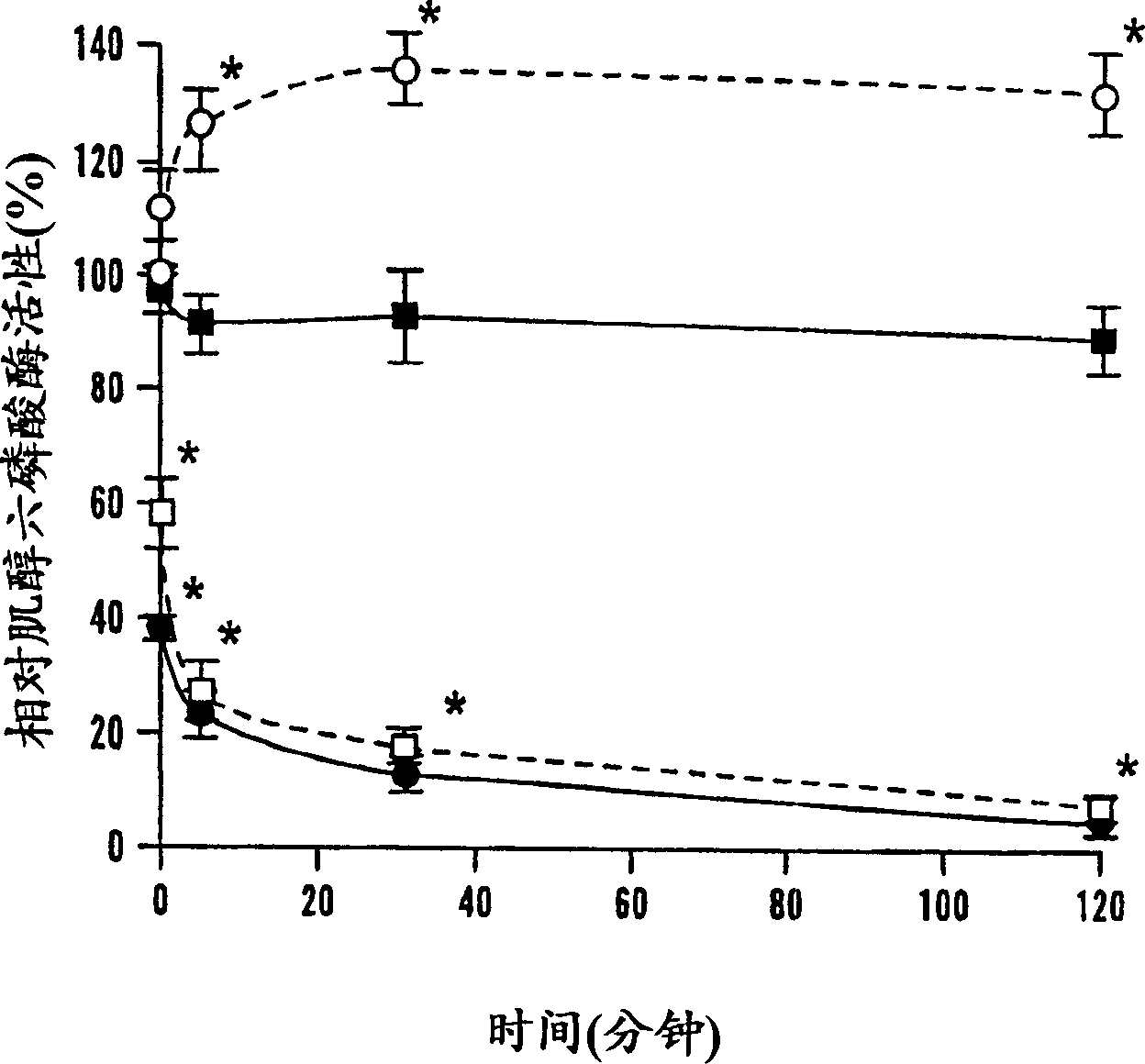
[0058] After 2 hours of trypsinization, there was a significant difference in remaining phytase activity between r-PhyA and r-AppA ( Figure 1A ). Although at trypsin / phytase ratios of 0.001 and 0.005, both enzymes retained more than 85% of their initial activity; at said ratios of 0.01 and 0.025, respectively, r-AppA lost its 64% and 74% of initial activity. At the same time, r-PhyA lost only 14% and 23% of its initial activity, respectively, correspondingly. Since the sensitivity of these two enzymes to trypsinization differs significantly at the ratio of 0.01, time course studies were performed at this ratio. Trypsinization up to 2 hours, r-PhyA still retained 90% of its original activity ( figure 2 ). In comparison, r-AppA lost 64%, 77%, 87% and 95% of its initial activity after 1 min, 5 min, 30 min and 120 min of digestion, respectively. Example 4 Effects of Pepsin Digestion on the Phytase Activities of Two Enzymes
PUM
| Property | Measurement | Unit |
|---|---|---|
| molecular weight | aaaaa | aaaaa |
Abstract
Description
Claims
Application Information
 Login to view more
Login to view more - R&D Engineer
- R&D Manager
- IP Professional
- Industry Leading Data Capabilities
- Powerful AI technology
- Patent DNA Extraction
Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic.
© 2024 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap