Expression profile of prostate cancer
a prostate cancer and expression profile technology, applied in the field of cancer diagnostics, can solve the problems of prostate specific antigen failure risk, prognosis comprises a risk of prostate cancer, and prognosis comprises a risk of prostate cancer, and achieve the effect of inhibiting ezh2 expression and reducing cell proliferation
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Benefits of technology
Problems solved by technology
Method used
Image
Examples
example 1
Preparation of Total RNA and Reference Pools
[0319]The prostate surgical specimens were obtained from The University of Michigan Specialized Research Program in Prostate Cancer (S.P.O.R.E.) Tumor Bank with Institutional Review Board approval. Tumors samples were derived from patients with clinically localized and advanced hormone refractory prostate cancer. Table 1 shows the samples used in the present studies. All patients were operated on between 1993 and 1998 for clinically localized prostate cancer as determined by preoperative PSA, digital-rectal examination, and prostate needle biopsy. In addition, a subset of patients received bone and CAT scans to evaluate the possibility of metastatic spread. All patients received radical prostatectomy as a monotherapy (i.e., no hormonal or radiation therapy). The advanced prostate tumors were collected from a series of 12 rapid autopsies performed at the University of Michigan on men who died of hormone refractory prostate cancer. In brief,...
example 2
Microarray Analysis
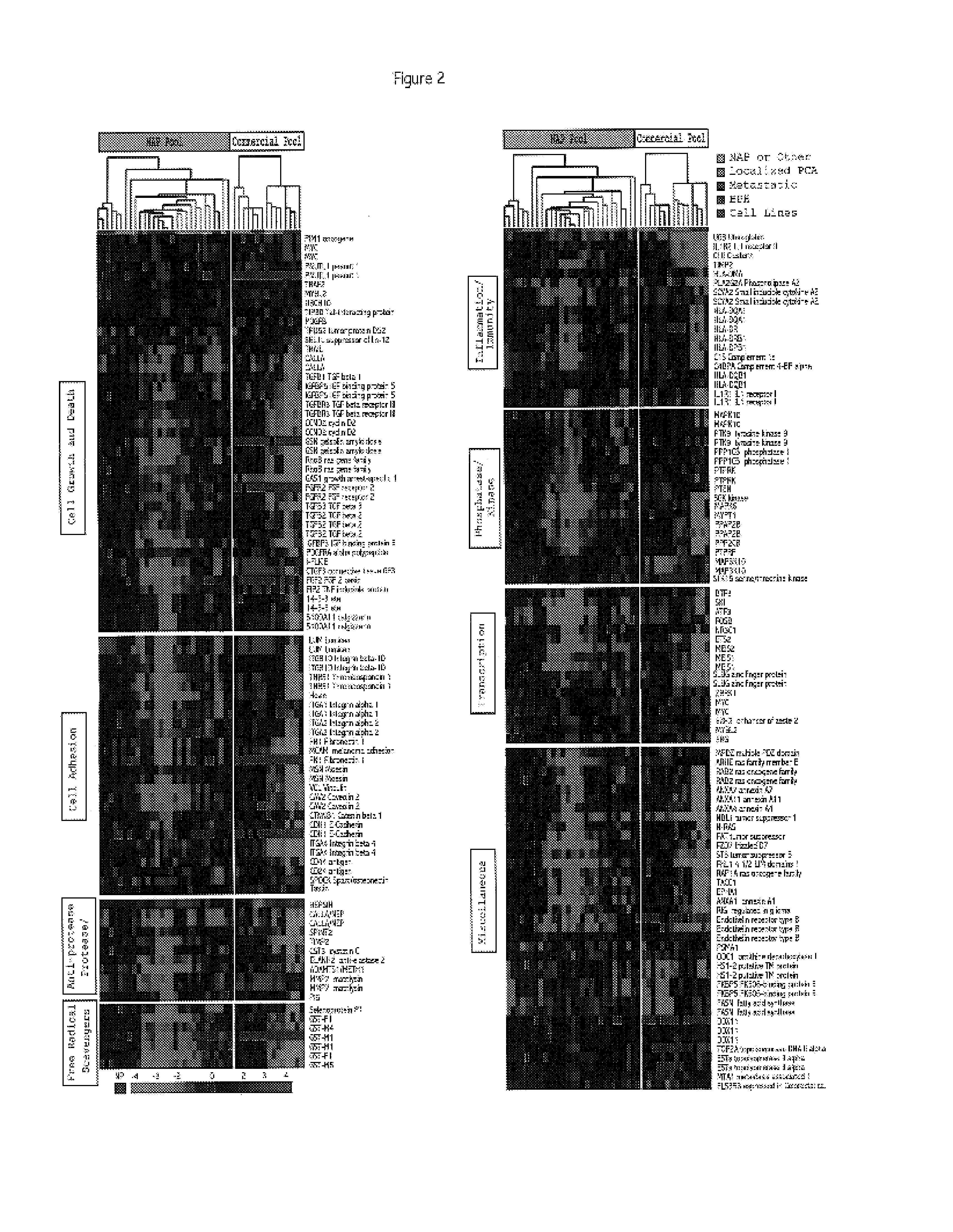
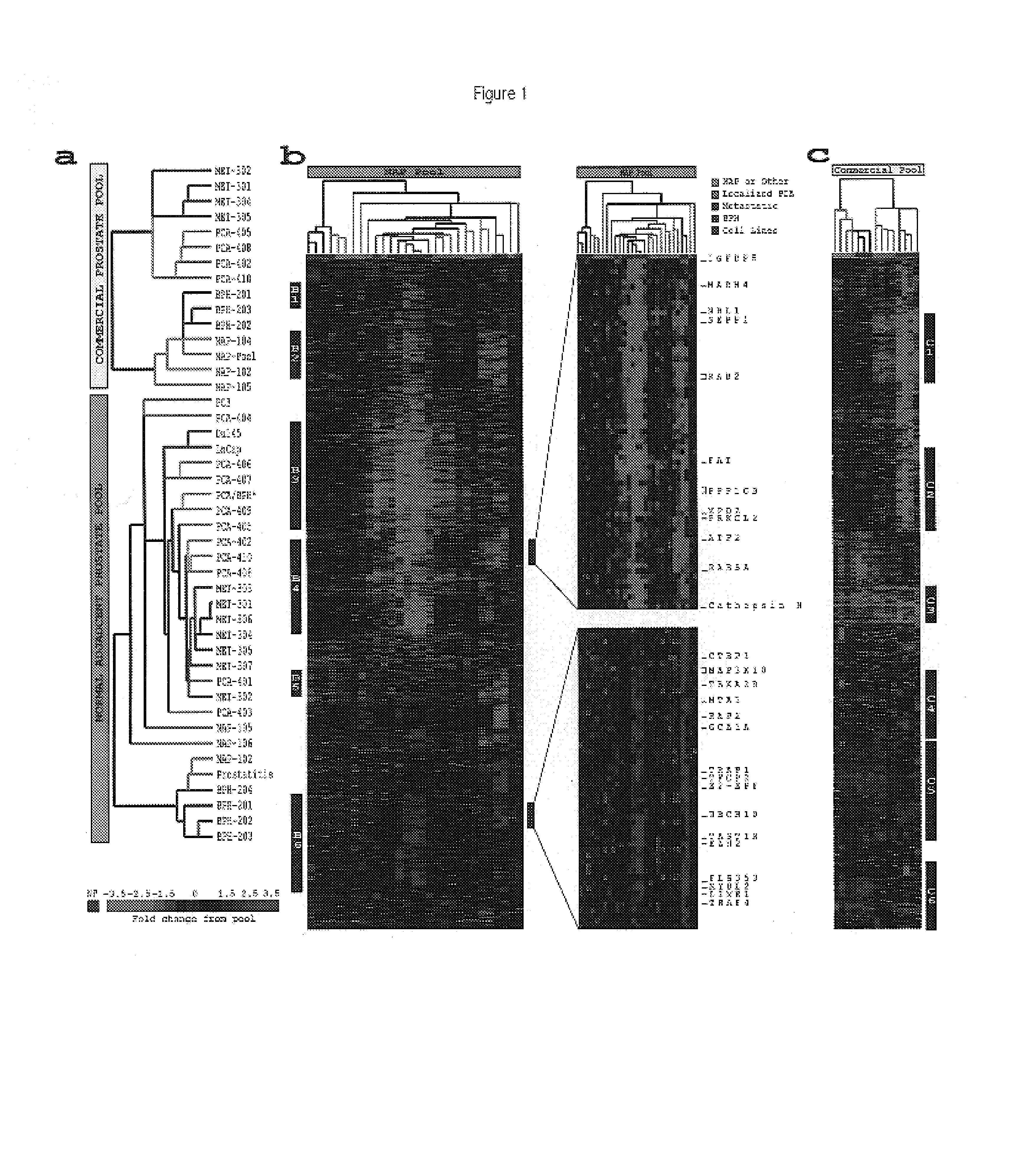
[0322]This example describes the use of microarray analysis to identify genes that demonstrate an altered level of expression in cancerous or benign prostate tissues.
A. Experimental Methods
[0323]Microarray analysis of gene expression was conducted essentially as described by the Brown and Derisi Labs (available at the Internet site www.microarrays.org). The sequence-verified cDNA clones on the human cDNA microarray are available from the web site of Research Genetics. Based on the latest Unigene build, the 10K human cDNA microarray used covers approximately 5520 known, named genes and 4464 ESTs. All chips have various control elements that include human, rat, and yeast genomic DNAs, SSC, yeast genes and “housekeeping genes,” among others. In addition, 500 cancer- and apoptosis-related cDNAs from Research Genetics were used to serve as independent controls for clone tracking and function as duplicates for quality control. Three metastatic prostate cancer cell lines...
example 3
Northern Blot Analysis
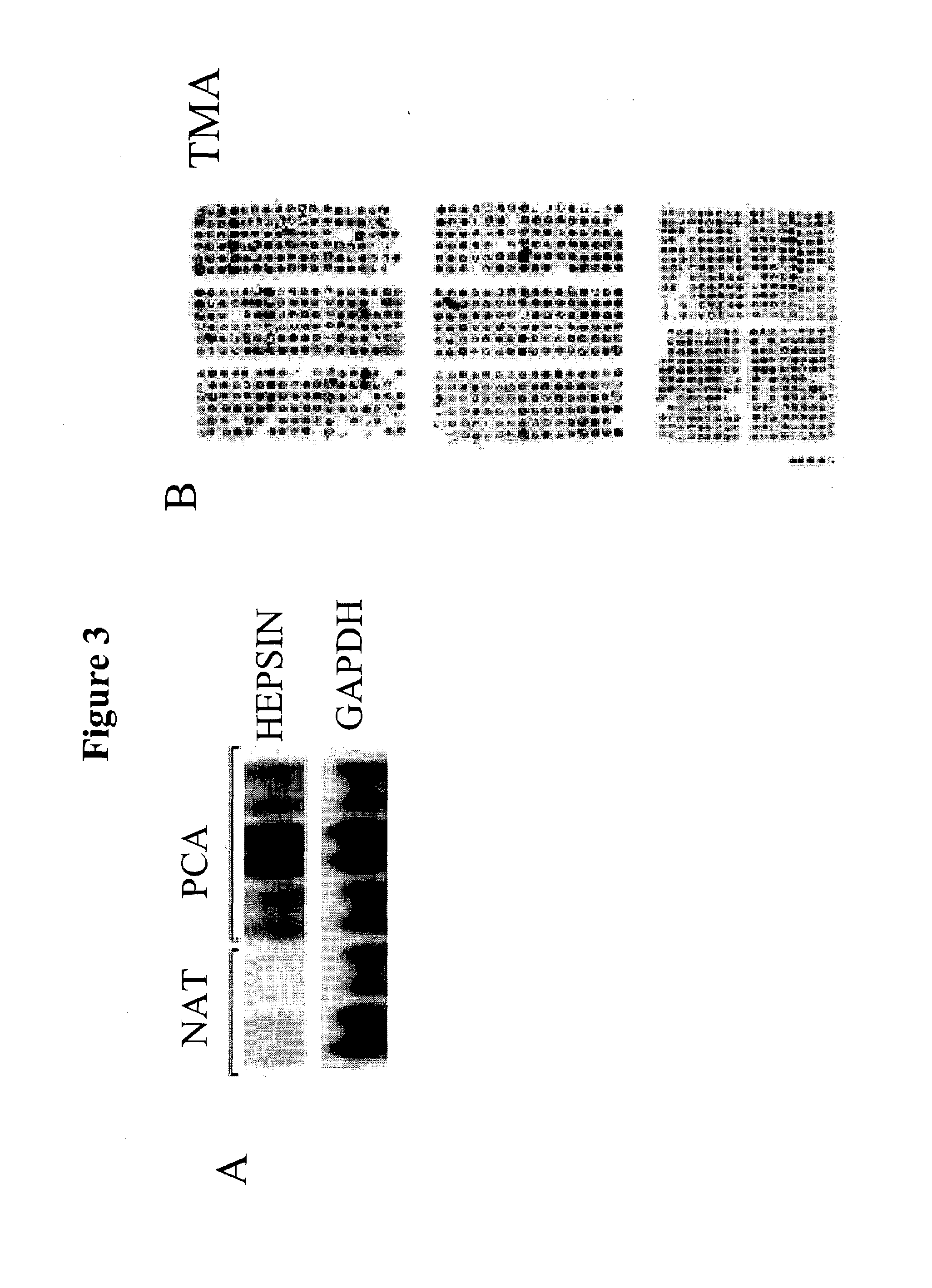
[0347]Thirty micrograms of total RNA was resolved by denaturing formaldehyde agarose gel and transferred onto Hybond membrane (Amersham) by a capillary transfer set up. Hybridizations were performed by the method described by Church and Gilbert, 1984. Signal was visualized and quantitated by phosphorimager. For relative fold estimation, the ratio between the signals obtained from hepsin and GAPDH probes was calculated.
[0348]Selected genes identified by microarray analysis were corroborated by Northern analysis. For example, hevin, 4 1 / 2 LIM domain protein and gelsolin were shown to be 3.2-, 3.2- and 1.9-fold down-regulated, respectively by microarray and 8.8-, 4.5-, and 3.5-fold down-regulated by Northern analysis. Similarly, hepsin was 4.3-fold up-regulated by microarray and 11.3-fold up-regulated by Northern analysis (FIG. 3a). As bepsin is a cell-surface serine protease with transcript expression precisely restricted to localized and metastatic PCA, its expr...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More