Protein marker combination for gastric cancer diagnosis and application thereof
A protein marker and gastric cancer technology, applied in the field of medical diagnosis, can solve the problems of low diagnostic sensitivity, high false positive rate, and poor prognosis, achieve high diagnostic sensitivity and specificity, promote accurate diagnosis and treatment, and improve accuracy
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0035] 1. Download gastric cancer (stomach) proteomics data and clinical information from the CPTAC database, and the sample size is gastric cancer tissue: normal tissue = 80:80.
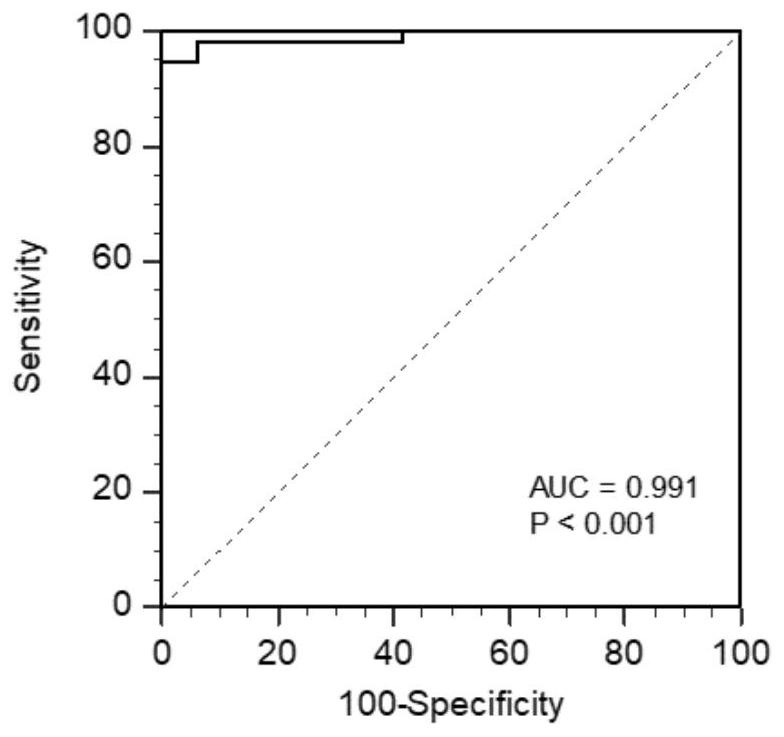
[0036] 1.1 Extract the concentration of protein markers COL10A1, GKN1, GKN2, LIPF, PGI, PGII, and G-17 involved in the present invention in gastric cancer tissues and paracancerous tissues. Further, the concentration of the above-mentioned related protein markers was converted to natural logarithm, and the regression equation was obtained through logistic regression analysis: Logit(P)=C+α1*Ln(COL10A1)+α2*Ln(GKN1)+α3*Ln (GKN2)+α4*Ln(LIPF)+α5*Ln(PGI / II)+α6*Ln(G17). Substitute the detected protein marker concentration of each sample into the regression equation to calculate the probability of each sample suffering from gastric cancer, determine the probability cut-off value (cut-off) through the maximum point of Youden index, and finally determine whether each sample suffers from gastric cancer situat...
Embodiment 2
[0052] 2. Analysis of diagnostic performance based on gastric cancer blood samples
[0053] 2.1 Use the purchased enzyme-linked immunoassay kit to test the protein markers (COL10A1, GKN1, GKN2, LIPF, PGI, PGII, G-17) in clinical serum samples (40 cases of control, 60 cases of gastric cancer, and 60 cases of chronic gastritis). ), and register the clinicopathological information of the cases at the same time.
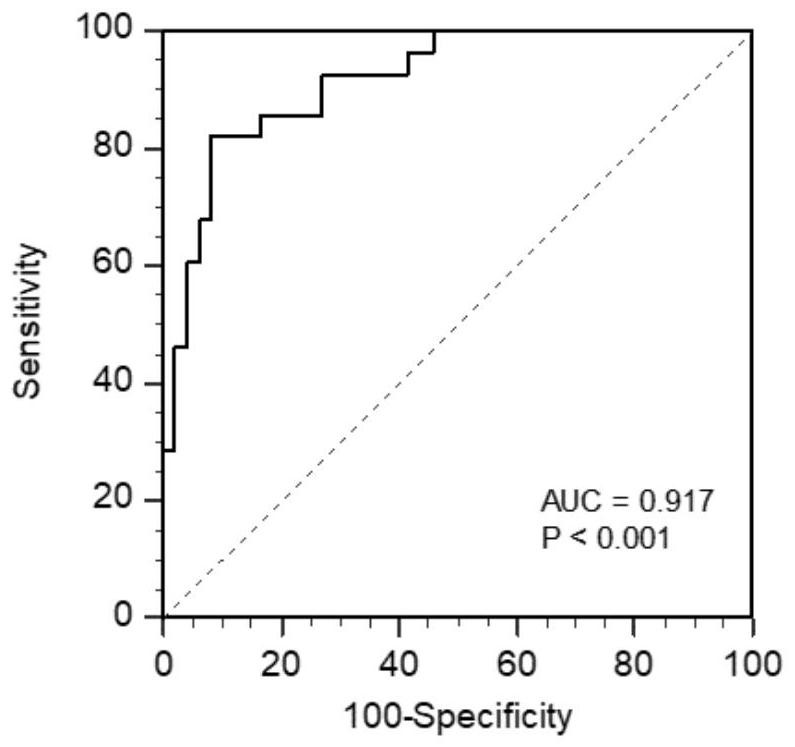
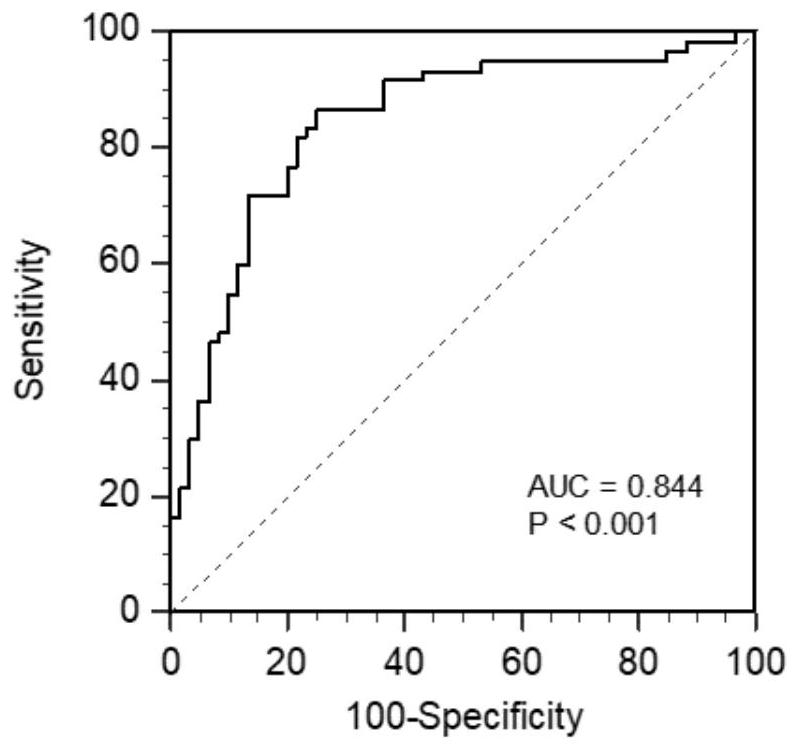
[0054] Further, the concentrations of the protein markers in the above three groups of cases were transformed by natural logarithm, and after logistic regression analysis, after removing the non-contributing markers, the regression equation was obtained: Logit(P)=C+α1*Ln(COL10A1 )+α2*Ln(GKN1)+α3*Ln(GKN2)+α4*Ln(LIPF)+α5*Ln(PGI / II)+α6*Ln(G17). Substitute the detected protein marker concentration of each sample into the regression equation to calculate the probability of each sample suffering from gastric cancer, determine the probability cut-off value (cut-off) through th...
PUM
 Login to view more
Login to view more Abstract
Description
Claims
Application Information
 Login to view more
Login to view more - R&D Engineer
- R&D Manager
- IP Professional
- Industry Leading Data Capabilities
- Powerful AI technology
- Patent DNA Extraction
Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic.
© 2024 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap