Detection methods for microarray mismarked samples
A technology for labeling samples and detection methods, applied in the field of computational biology, can solve problems such as low recall rate
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment Construction
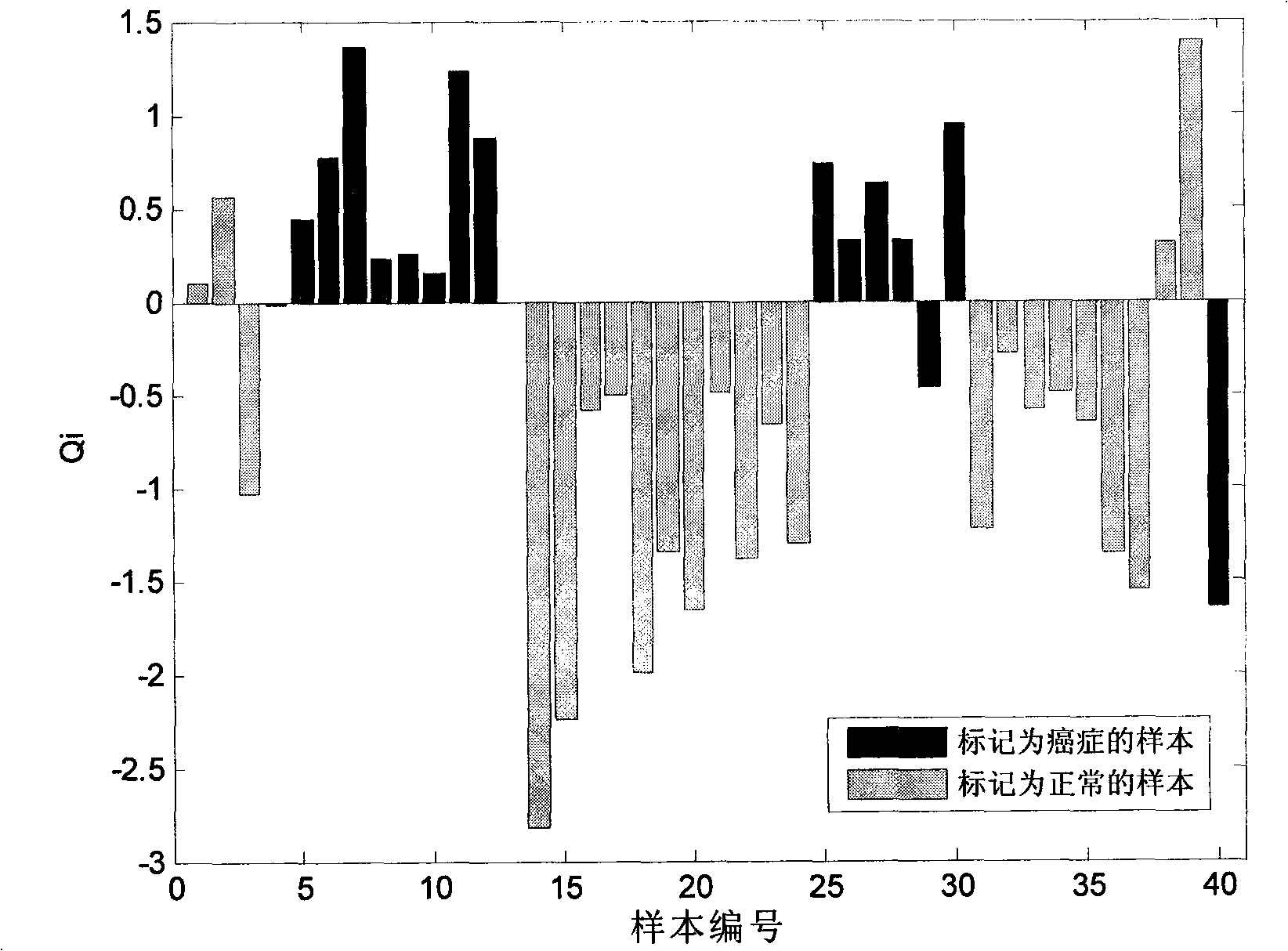
[0050] In the following, the present invention will be described in detail through the example of two-category gene chip data of breast cancer. The breast cancer (breast) gene expression profile data set of West et al. is a general data set, which contains 49 breast cancer samples, including 25 estrogen receptor positive (ER+) samples, estrogen receptor There were 24 negative (ER-) samples, and the gene chip contained 7129 genes. On this basis, suspicious samples 11, 14, 16, 31, 33, 45, 46, 40, 43 were removed, and then samples 1, 2, 3, 47, 48, 49 were manually flipped to make them mislabeled samples. The obtained data set is the instance data that will be used below.
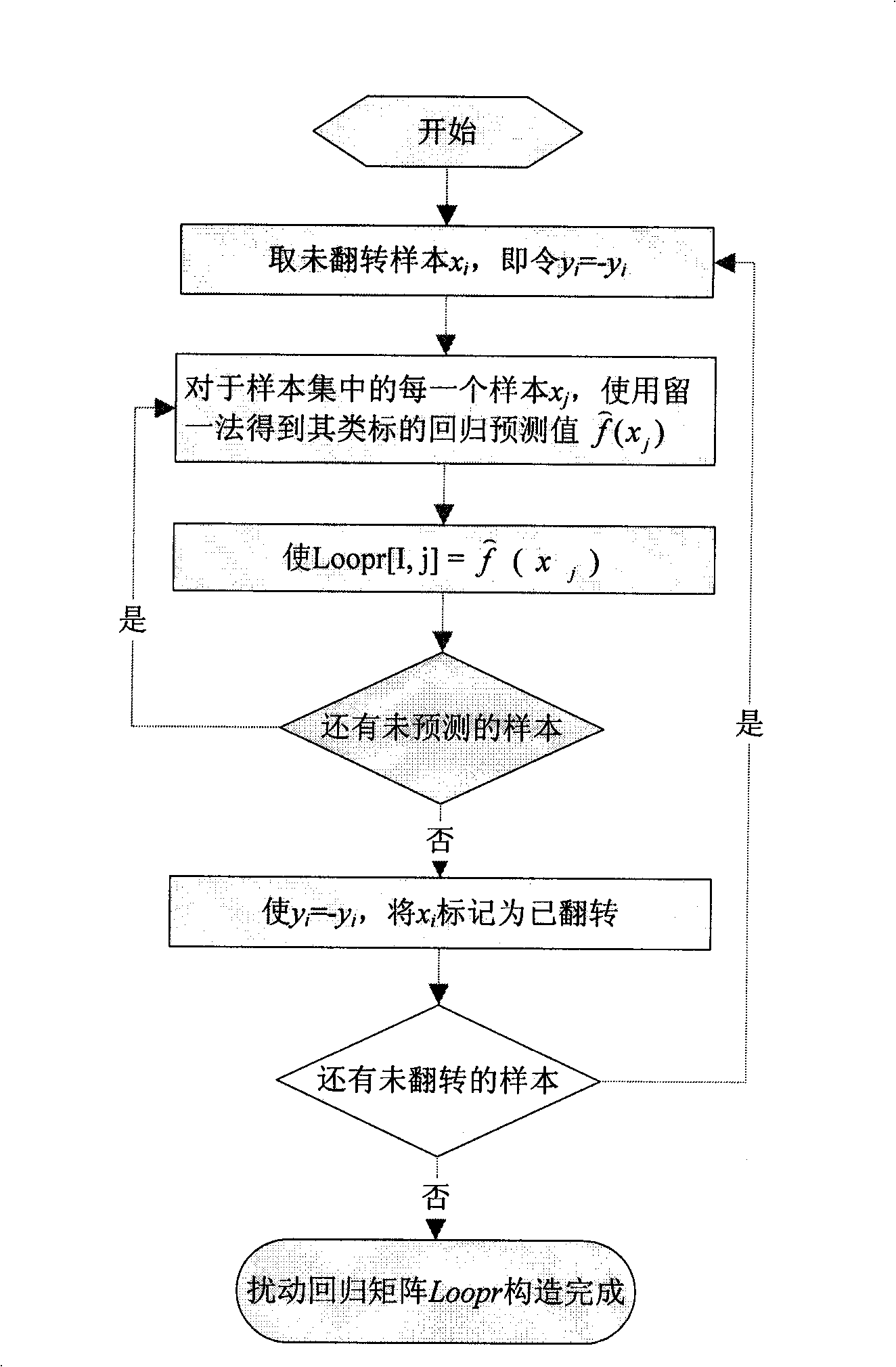
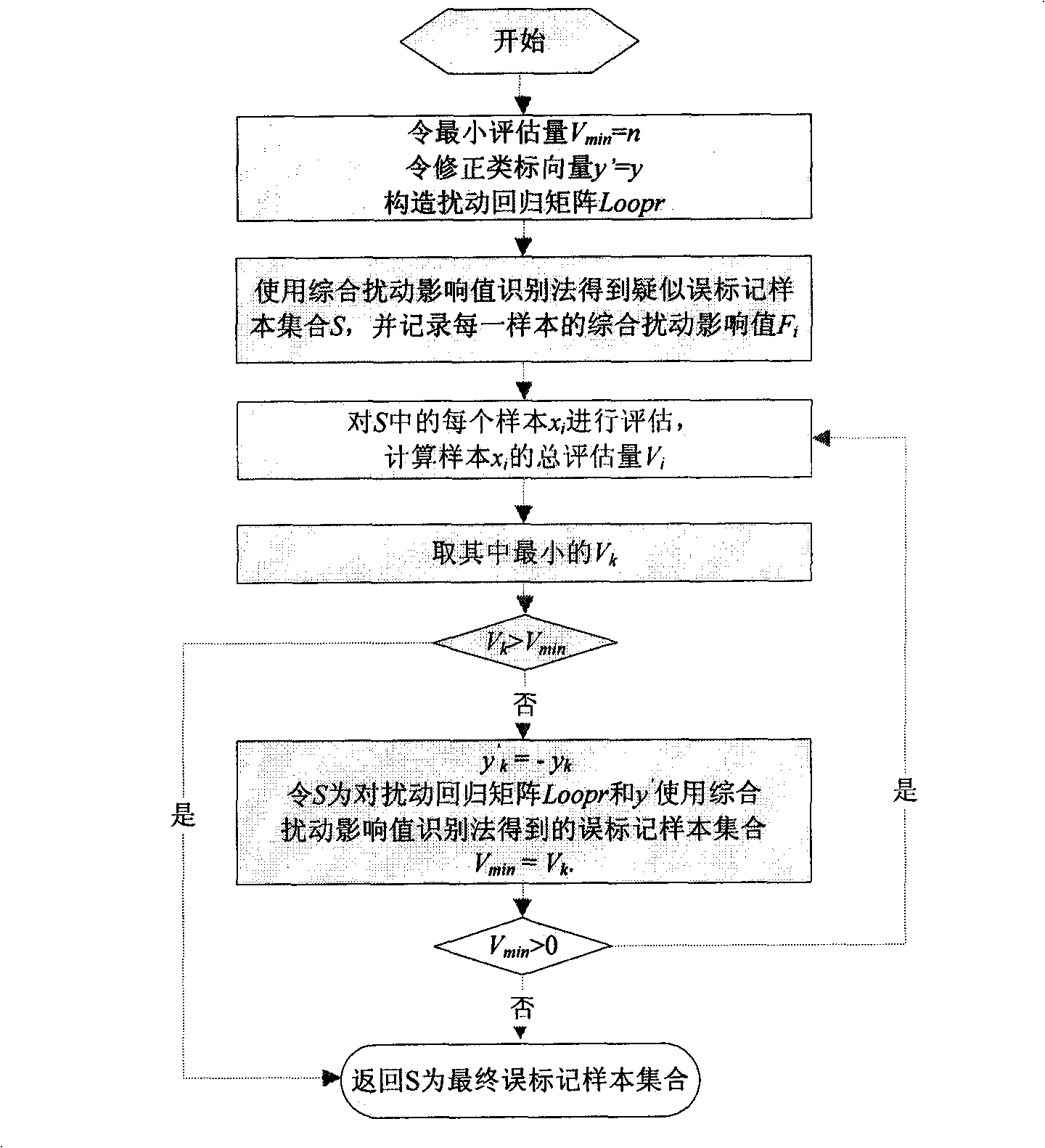
[0051] 1. Overall Disturbance Impact Recognition Method
[0052] Step 1: Take an unflipped sample x from the dataset i , making its class label y i =-y i ;
[0053] Step 2: For each sample x in the dataset j , put x j As the test sample, other samples as the training set, for the sample x j Regression ...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More