Design method of oligonucleotide probe and application thereof
A technology for oligonucleotide probes and design methods, which can be used in biochemical equipment and methods, microbial determination/inspection, DNA/RNA fragments, etc. The increase of density chip preparation and other problems can achieve the effect of strong discrimination ability, excellent specificity, and reduced screening workload.
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0034] Example 1 Probe design of gene chip for detecting MDRI gene SNPs
[0035] The multi-drug resistance gene (MDRI) is located on human chromosome 7 q21.1, taking C-4T as an example to illustrate the probe design method. The following is the sequence of MDRIC-4T site (LOCUS: AY910577):
[0036] 1 CTCTGGCTTC GACGGGGGAC TAGAGGTTAG TCTCACCTCC AGCGCGCCTG AGGCTCATGC
[0037] 61 ATTTGGCTAA TGAGCTGCGG TTTCTCTTCA GGT GGGATG GATCTTGAAG GGGACCGCAA
[0038] 121 TGGAGGAGCA AAGAAGAAGA ACTTTTTTAA ACTGAACAAT AAAAGGTAAC TAGCTTGTTT
[0039] 181 CATTTTCATA GTTTACATAG TTGCGAGATT TGAGTAATTT ATTTCTAGCC TCCAGCTCTG
[0040] 241 AAATAAATGA CATGTTGTTG TTTTTAATTA TTTTTAAGAA ACGCAAGCTA GCCTTTGGAA
[0041] 301 TCAATATCCC TGCTTAGAGC AGAAGTTTGT TGGCTGAGTG GAGCACAGCA TATGCATTTT
[0042] In the above sequence, It is the site where the mutation occurs, that is, the wild type (Normal) is C, and the mutant type (Mutation) is T. During probe design, mutation sites and artificial insertion sites are distributed at both...
Embodiment 2
[0043] Example 2 Detection of C-4T and A61G sites of MDRI gene by gene chip of slide carrier
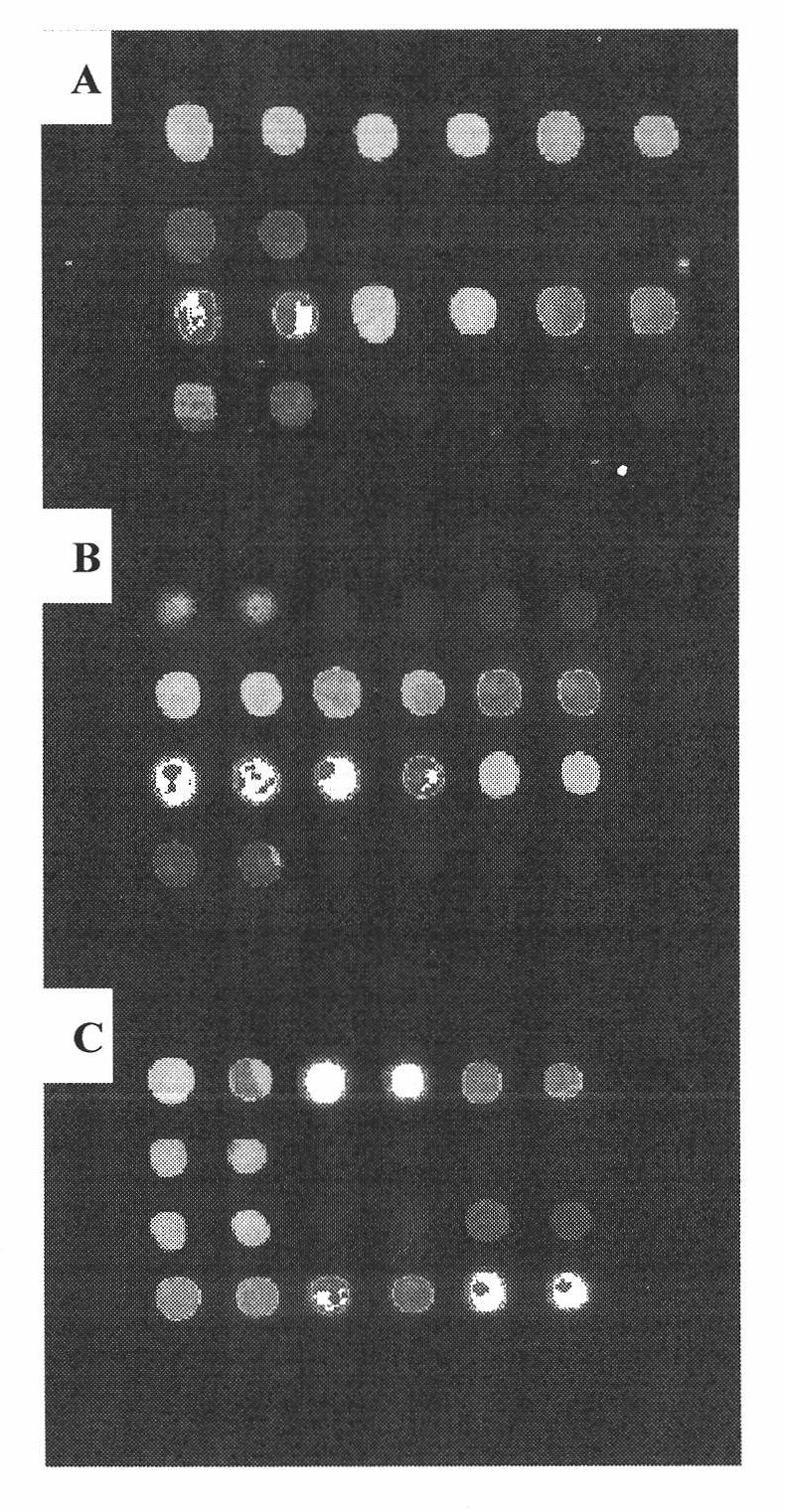
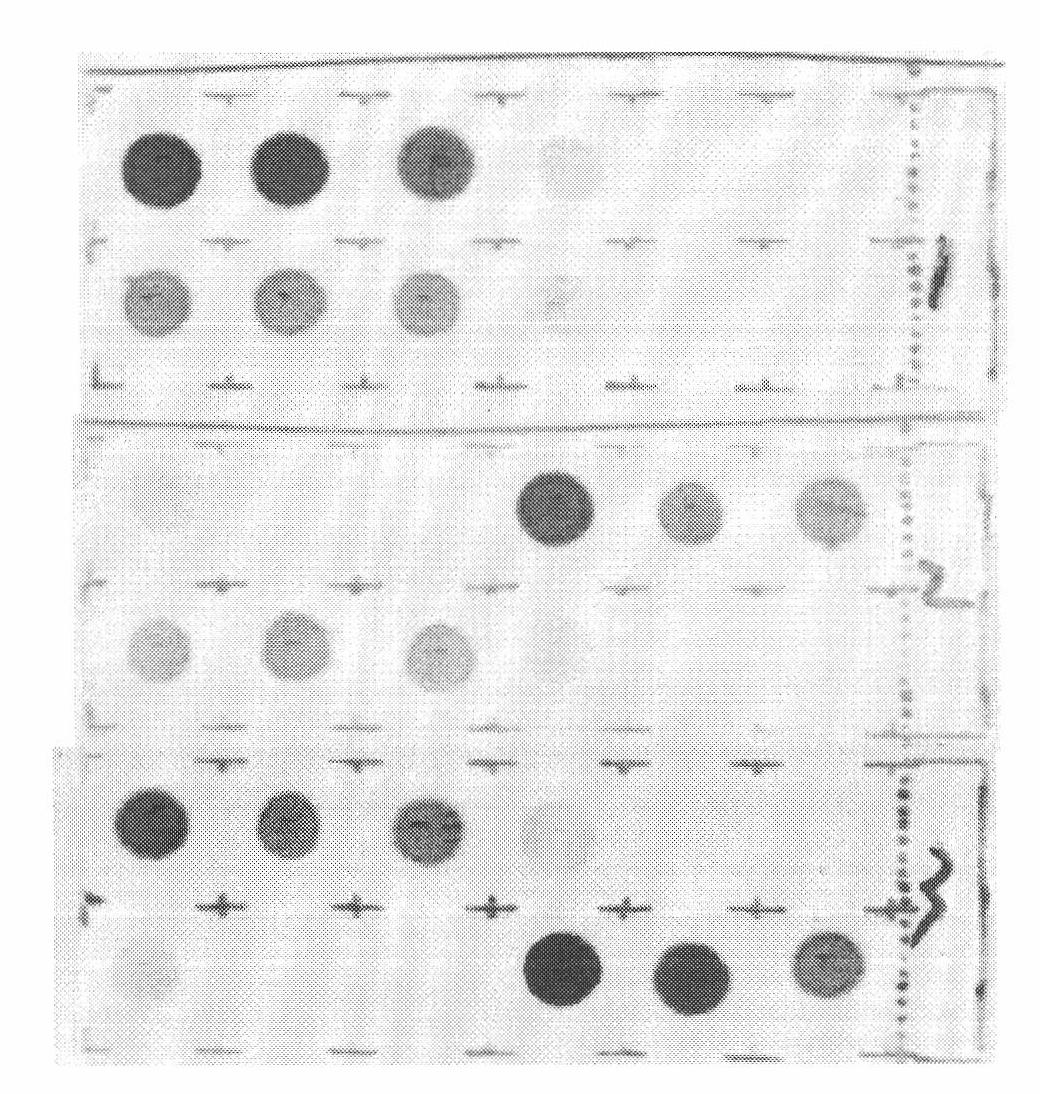
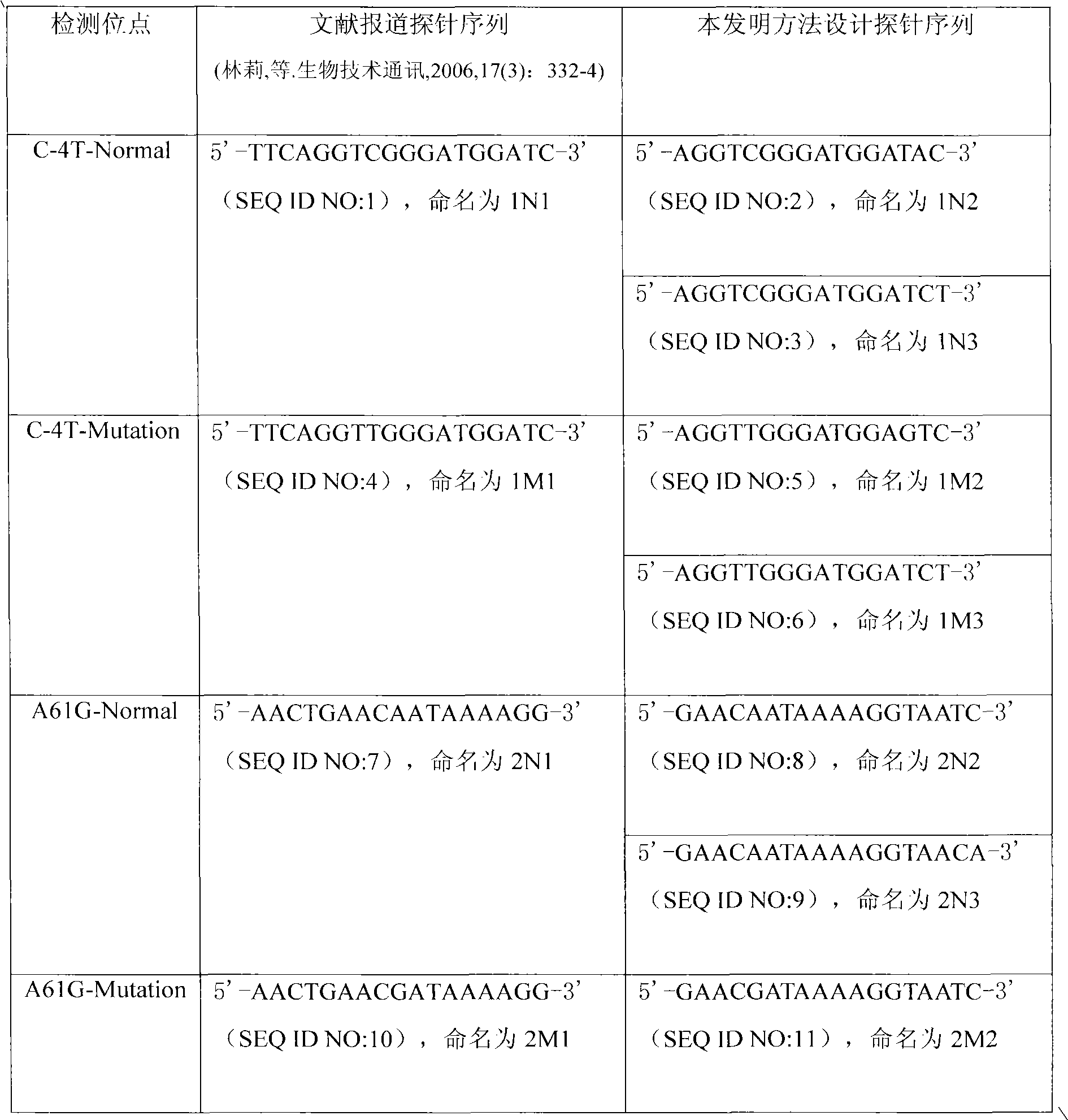
[0044] The probe designed in Example 1 was used to prepare a slide carrier gene chip to detect MDRI gene SNPs, and compared with the gene chip prepared by the probe reported in the literature.
[0045] Multi-drug resistance gene (MDRI) is located on human chromosome 7 q21.1. Its protein product p-GP is an important member of the ABC transport system and belongs to the ABCB1 family. MDRI gene expression level can be used as a reference indicator for predicting the effect of chemotherapy. By detecting MDRI gene polymorphisms to predict the efficacy of chemotherapy in cancer patients has become a feasible clinical detection method. The PCR primer sequences used are:
[0046] 1F: 5'TTGGCTAATGAGCTGCGGTTTCTC 3'(SEQ ID NO: 25);
[0047] 1R: 5'cy3-GATTCCAAAGGCAGCTTGCGTTTC 3'(SEQ ID NO: 26).
[0048] The amplified fragment of the above sequence is 240bp, and the two SNPs sites C-4T and A61G of exon 2...
Embodiment 3
[0058] Example 3 Probe design of gene chip for detecting β-thalassemia gene mutation
[0059] The human β-Globin gene (β-Globin) cluster is located at 11p15.5. Beta thalassemia (abbreviated as beta thalassaemia) occurs mainly due to point mutations in genes, and a few are gene deletions. The following is the sequence of β-Globin CD26 (G→A) (LOCUS: NC 000011)
[0060] 1 TGACTCCTGA GGAGAAGTCT GCCGTTACTG CCCTGTGGGG CAAGGTGAAC GTGGATGAAG
[0061] 61 TTGGTGGT A GGCCCTGGGC AGGTTGGTAT CAAGGTTACA AGACAGGTTT AAGGAGACCA
[0062] 121 ATAGAAACTG GGCATGTGGA GACAGAGAAG ACTCTTGGGT TTCTGATAGG CACTGACTCT
[0063] 181 CTCTGCCTAT TGGTCTATTT TCCCACCCTT AGGCTGCTGG TGGTCTACCC TTGGACCCAG
[0064] 241 AGGTTCTTTG AGTCCTTTGG GGATCTGTCC ACTCCTGATG CTGTTATGGG CAACCCTAAG
[0065] 301 GTGAAGGCTC ATGGCAAGAA AGTGCTCGGT GCCTTTAGTG ATGGCCTGGC TCACCTGGAC
[0066] In the above sequence, It is the site where the mutation occurs, that is, the wild type (Normal) is G, and the mutant type (Mutation) is A. During probe desig...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More