Methods and compositions for amplification of RNA sequences
a technology of rna sequence and composition, applied in the field of polynucleotide amplification, can solve the problems of prone to generate pools of products which do not, large amount of starting mrna, and limited sample mrna availabl
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Benefits of technology
Problems solved by technology
Method used
Image
Examples
example 1
Amplification of Total Poly-A mRNA
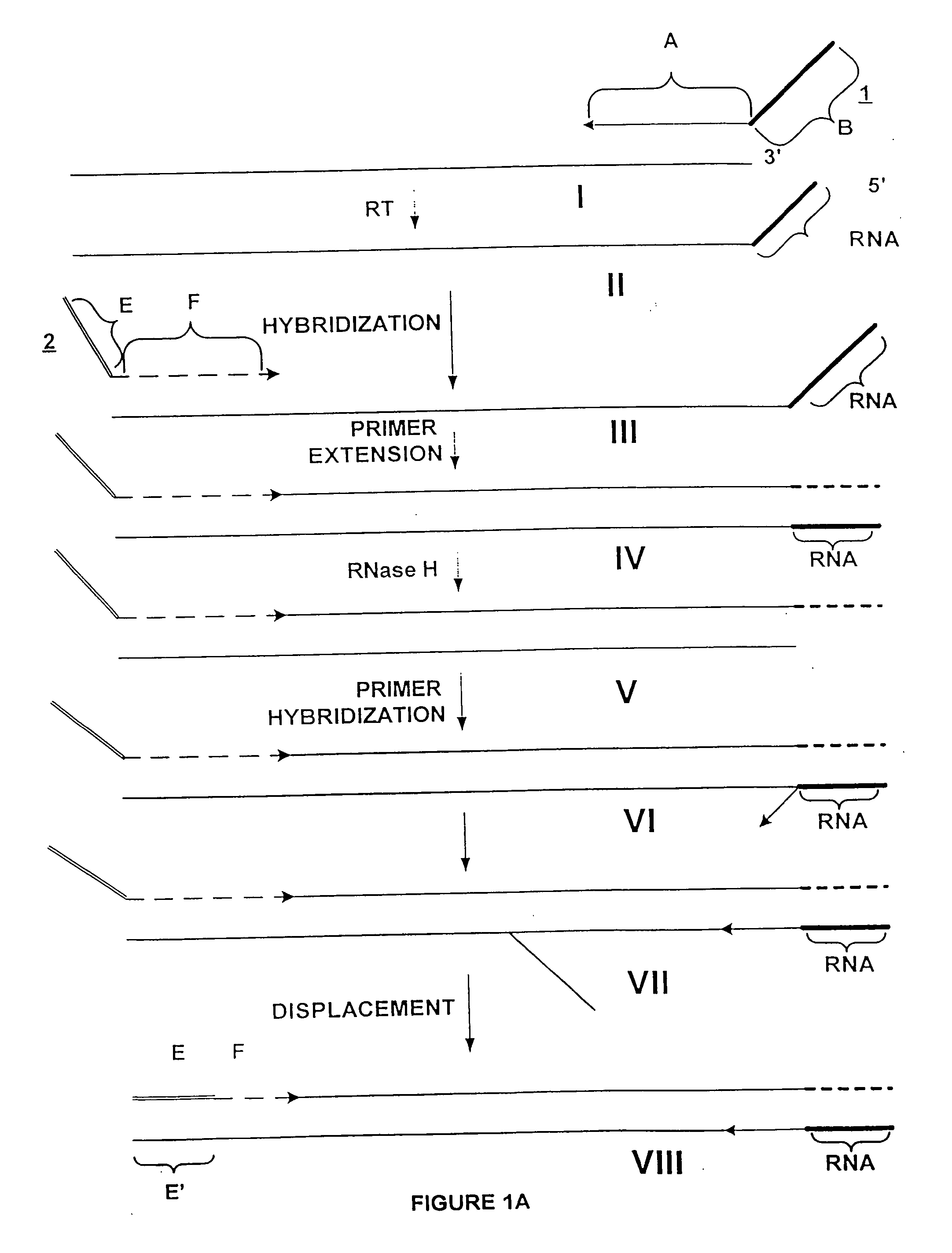
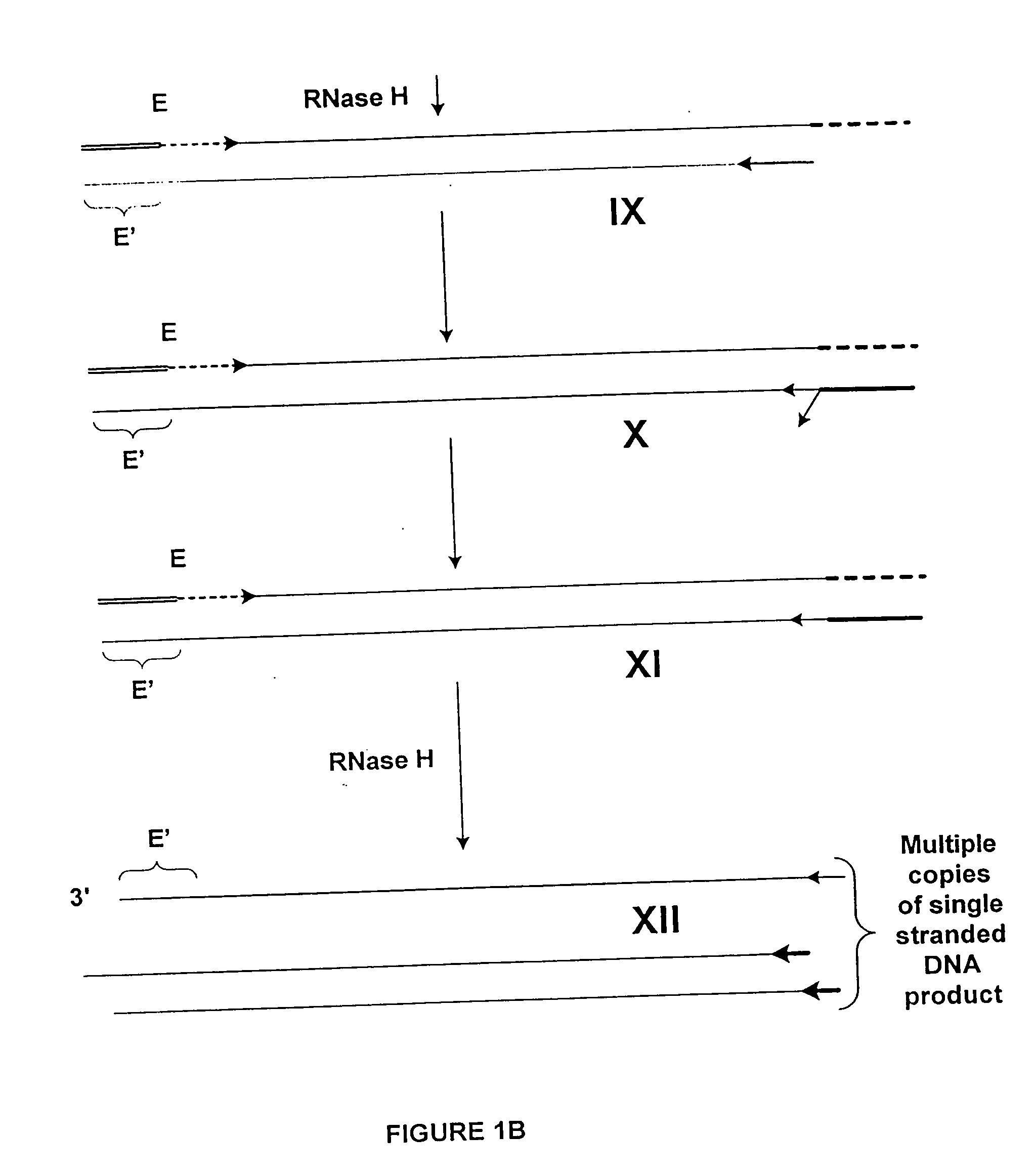
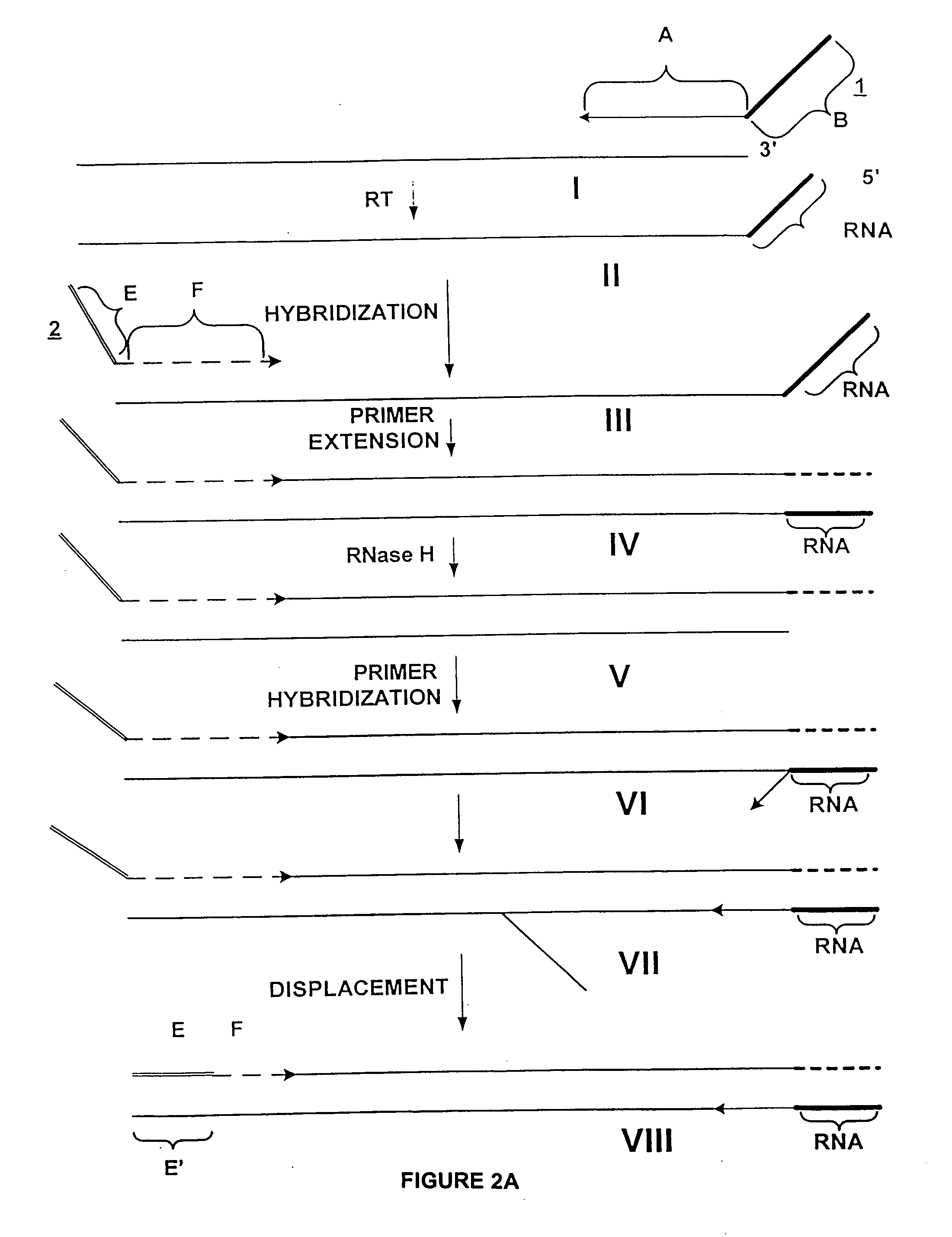
Poly-A mRNA from MOLT4 cell line (CLONTECH 6587-1) was used as a target for amplification. The process of amplification was in three steps: 1) synthesis of first cDNA strand; 2) synthesis of second cDNA strand to produce a double stranded cDNA from the total mRNA of the sample; and 3) amplification of the total mRNA. The double stranded cDNA product comprises at one end an RNA / DNA heteroduplex, which is a substrate for RNase H. The sequence of the two strands of this heteroduplex portion is not related to the target, and is incorporated through utilization of a composite (first) primer.
Primer sequences:
MTA1:GACGGAUGCGGUCUTTTTTTTMTA2:GACGGAUGCGGUCUTTTTTTTNMTA3:GACGGAUGCGGUCUTTTTTTTNN
wherein italicized nucleotides denote ribonucleotides and ‘N’ denotes a degenerate nucleotide (i.e., it can be A, T, C or G).
Step 1: Synthesis of the first strand cDNA from poly A mRNA 0.1 μg of total poly-A mRNA was mixed with the following reagents in a total ...
example 2
Characterization of Products of Step 2 and Step 3 Reactions of Example 1
In the amplification reactions of Example 1, a “unique” sequence (i.e., a sequence not hybridizable to the RNA template) is expected to be created at the 3′-end of the second strand cDNA due to the “unique” sequence of the 5′ RNA portion of the composite primer used. This sequence (of the 3′-end of the second strand cDNA) is complementary to the 5′-RNA portion of the composite primer and is not related to sequences in the target RNA. To determine the presence of this sequence in the second strand cDNA that is obtained, PCR amplification of the reaction products (as found in reaction mix of step 2 of Example 1) was performed using a primer which is complementary to the expected sequence at the 3′-end of the second strand cDNA, as a forward primer, and a G3PDH-specific primer as a reverse PCR primer. This primer pair would be expected to amplify a specific product from a double stranded cDNA that has the “unique...
example 3
Amplification of Total mRNA Starting With a Total RNA Preparation
The ability to amplify total mRNA from a preparation of total RNA greatly simplifies the process by eliminating the mRNA purification step. Experimental demonstration of amplifying total mRNA from a total RNA preparation using methods of the invention was carried out using commercial total RNA preparation from breast cancer tumor (CLONTECH; cat. no.: 64015-1). The process of amplification of total mRNA was carried out in three steps as described in the following.
Primer sequence:
MTB2:GAC GGA UGC GGU CUTTTTTTTTTTTTTTNNBA5:AAC TAC CTT CAA CTC CAT CABA3:GGA CTC GTC ATA CTC CTG C
wherein italicized nucleotides denote ribonucleotides and ‘N’ denotes a degenerate nucleotide (i.e., it can be A, T, C or G).
Step 1: First Strand cDNA Synthesis
Each reaction mixture comprised the following: 4 μl of a SX buffer (250 mM Tris-HCl, pH 8.3; 375 mM KCl, 15 mM MgCl2) MTB2 primer @1 μM 25 mM dNTPs 0.2 μl RNasin Ribonuclease I...
PUM
 Login to view more
Login to view more Abstract
Description
Claims
Application Information
 Login to view more
Login to view more - R&D Engineer
- R&D Manager
- IP Professional
- Industry Leading Data Capabilities
- Powerful AI technology
- Patent DNA Extraction
Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic.
© 2024 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap