A pair of transcription activator-like effector nucleases and coding engines as well as application thereof
A transcriptional activation and effector technology, applied in the field of genetic engineering, to achieve strong specificity, high accuracy, and high targeting efficiency
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
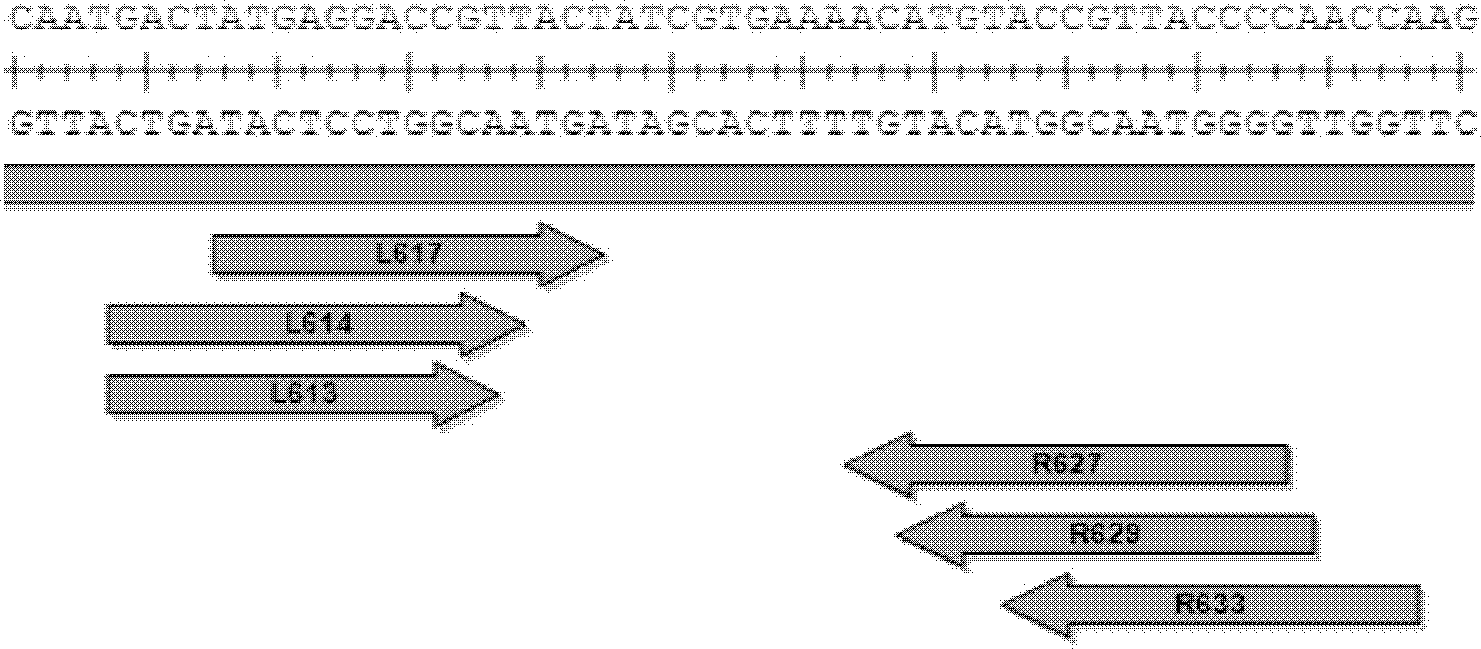
[0053] Embodiment 1 Design of TALENs target sequence
[0054] 1. Download goat and sheep PRNP genome sequences from NCBI (sheep Gene ID: 493887, goat PRNP gene sequence is the same as that of sheep)
[0055] 2. Design primers and PCR amplify the targeting site fragments on the genome, and sequence them. The PCR primers and sequencing primers are shown in Table 1;
[0056] Table 1
[0057]
[0058] 3. Design TALENs recognition sequence (target sequence):
[0059] According to the sequence obtained by sequencing, the recognition sequence of TALENs was determined according to the following principles:
[0060] (1) The 0th base is T (the base before the first in the recognition sequence is the 0th)
[0061] (2) The last base is T
[0062] (3) The length of the recognition sequence is between 13-19
[0063] (4) The length of the spacer sequence (Spacer) between the two recognition sequences is controlled between 13-21 (12 is also possible, but the efficiency may be lower) ...
Embodiment 2
[0067] Example 2 Connection between TALENs recognition modules and construction of recombinant vector
[0068] 1. Acquisition of TALENs identification module (modular)
[0069] (1) Synthesize four recognition modules NI, NG, HD, and NK that recognize bases A, T, C, and G respectively. The sequences are shown in Table 3.
[0070] table 3
[0071]
[0072]
[0073] (2) Connect the four fragments into the pEASY-B vector (purchased from Beijing Quanshijin Company), the connection method is:
[0074] ①Take 3 μl of PCR product;
[0075] ② Add 1 μl pEASY-B vector;
[0076] ③25℃, 7min;
[0077] ④Transform DH5a competent cells and spread kanamycin plate;
[0078] ⑤Pick clones, extract plasmids in a small amount, digest, and sequence, and finally obtain the recognition modules NI, NG, HD, and NK connected to the vector pEASY-B.
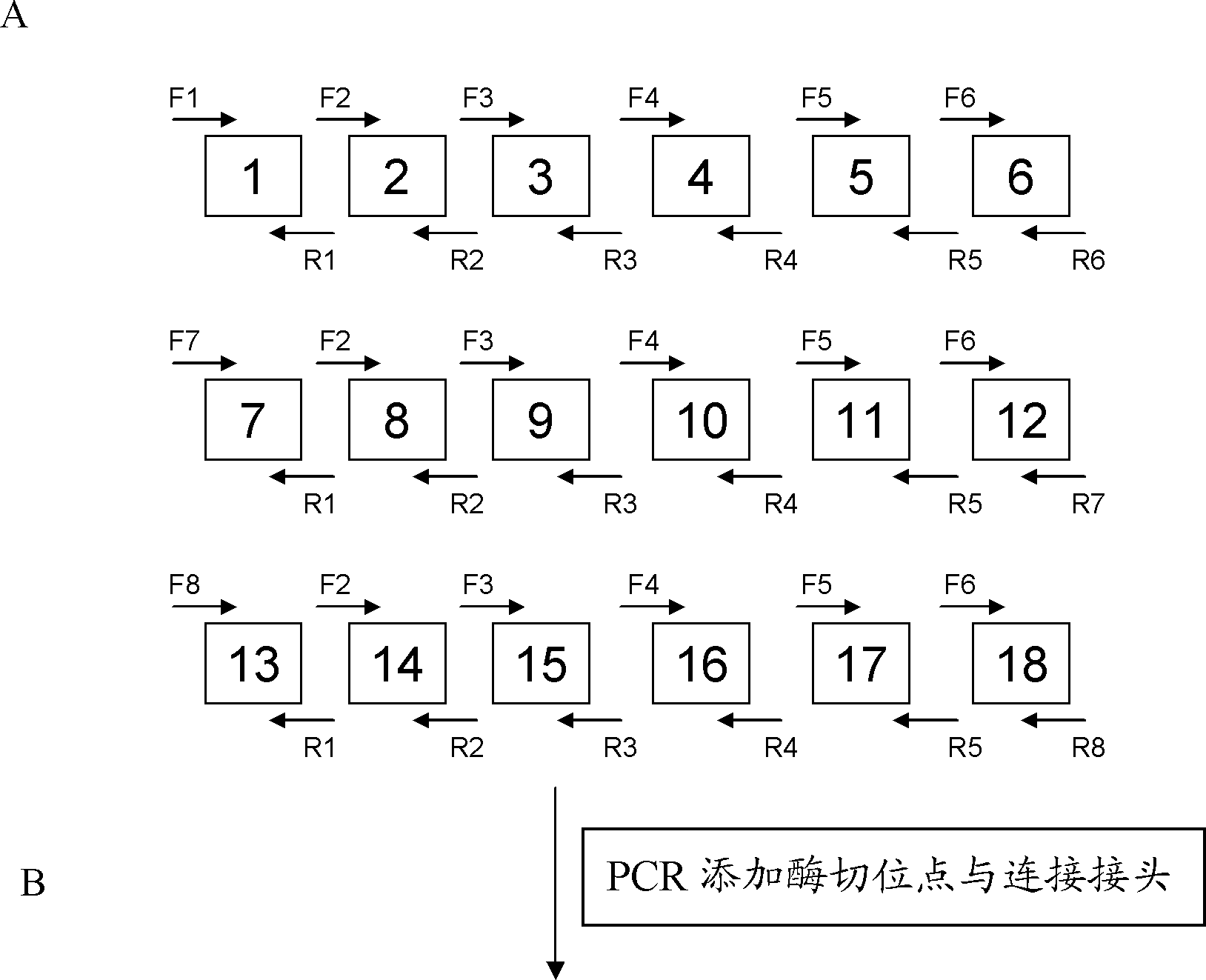
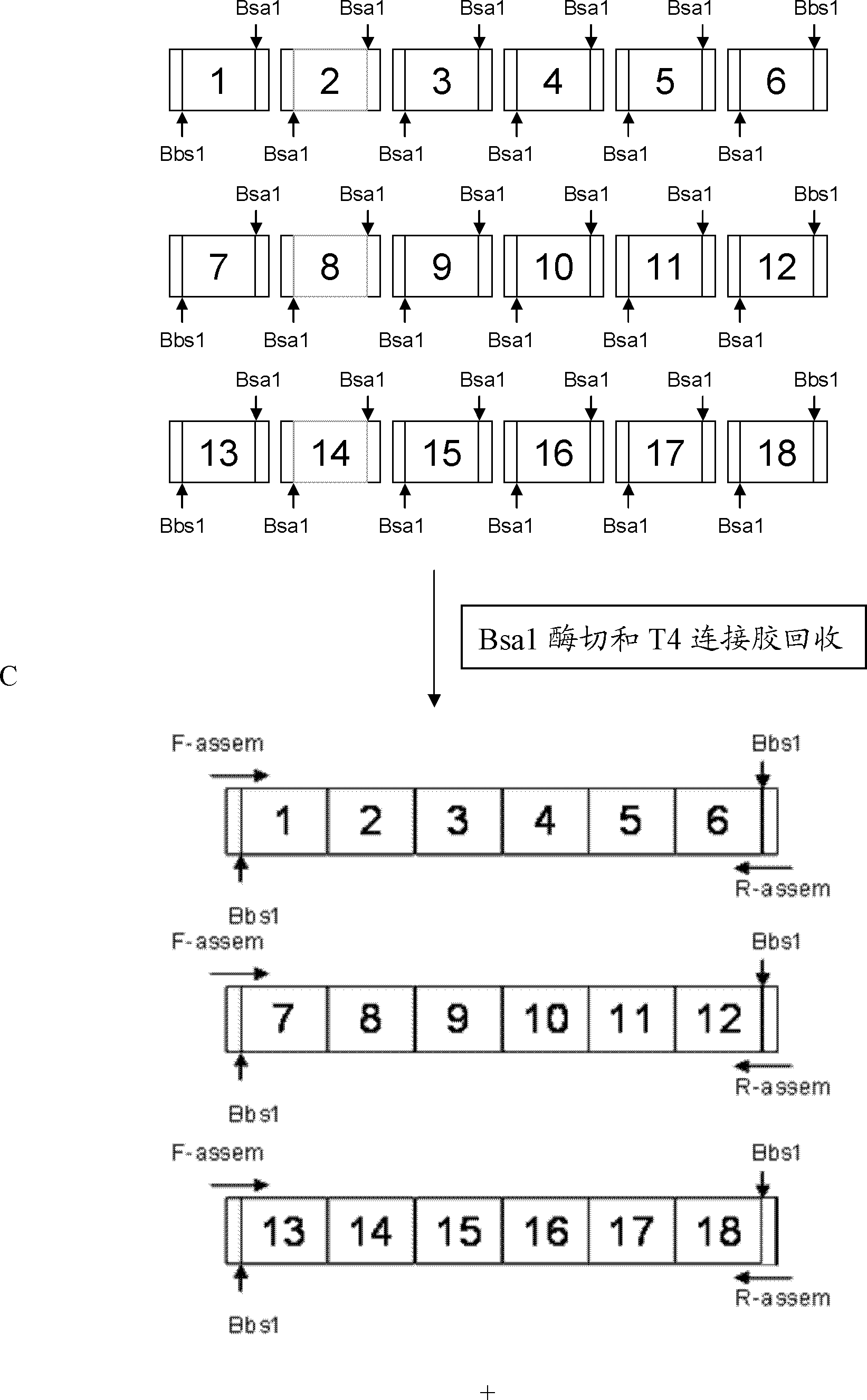
[0079] 2. Identify connections between modules
[0080] Connection strategy: Take the connection of 19 identification modules as an example to ill...
Embodiment 3
[0147] The transfection of embodiment 3 plasmids
[0148] 1. Add 100 μl Matrigel to each well of a 6-well plate, shake it back and forth to make it cover the bottom of the entire well, and place it in 5% CO2 30min in the incubator.
[0149] 2. Aspirate the culture medium in the T25 bottle of cultured IPS cells, and once in PBS, add 1mL of 0.25% trypsin, shake back and forth to make it evenly cover the bottom of the bottle, and place in 5% CO 2 5min in the incubator.
[0150] 3. After digestion, add 1ml 10% DMEM to neutralize trypsin, transfer the digested cells to a 15ml centrifuge tube, count the cells, and centrifuge at 1200rpm for 5min.
[0151] 4. Resuspend the cells with an appropriate amount of 4*Dox ES0, take 2 million IPS cells and place them in a 6-well plate that has been covered with Matrigel, and add 2ml of fresh 4*Dox ES0.
[0152] 5. Passage and transfect at the same time.
[0153] 6. Transfect the constructed PRNP-TALEN-L613, PRNP-TALEN-L614, PRNP-TALEN-L617,...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More - R&D
- Intellectual Property
- Life Sciences
- Materials
- Tech Scout
- Unparalleled Data Quality
- Higher Quality Content
- 60% Fewer Hallucinations
Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic, Popular Technical Reports.
© 2025 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap|About US| Contact US: help@patsnap.com