Primer for detecting salmonella enteritidis and detecting reagent kit containing primer
A technology of Salmonella enteritidis and detection kit, which is applied in the direction of microbial determination/inspection, resistance to vector-borne diseases, biochemical equipment and methods, etc., can solve the problems of no specificity and inability to directly distinguish serotypes, etc., and achieve Strong specificity, easy to promote and use, and easy to grasp the effect
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0019] Embodiment 1 adopts specific primer of the present invention to carry out PCR detection
[0020] 1.1 Primers
[0021] Prot6E-F: 5'-ACAGGGGCACAATAACCGTA-3' and
[0022] Prot6E-R: 5'-TGCATCCCTGTCACAACATT-3'
[0023] 1.2 Bacteria enrichment
[0024] After sampling, add buffered peptone water (BPW) at a ratio of 1:9 and shake for 18 hours, draw 1 ml of the culture solution and add it to 9 ml of selenite cystine selective enrichment solution (SC), and shake for 18 hours.
[0025] 1.3 DNA extraction
[0026] Take 2 mL of the selective enrichment solution and place it in an Ep tube, centrifuge at 12000 r / min for 5 min, wash with ultrapure water twice, suspend with 0.2 mL of ultrapure water, boil in water for 15 min, insert into ice to cool for 2 min, and centrifuge at 12000 r / min for 15 min , and the supernatant was used as a DNA template.
[0027] 1.4PCR amplification
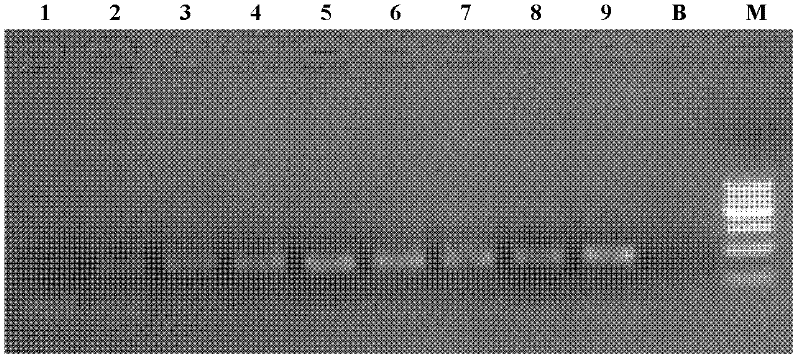
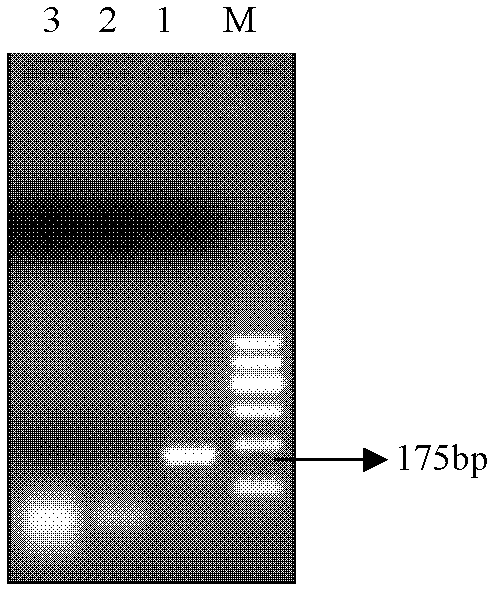
[0028] In the PCR reaction, the standard strain DNA of Salmonella enteritidis was used as a positive ...
Embodiment 2
[0030] Embodiment 2 adopts specific primer of the present invention to carry out the sensitivity analysis of PCR detection
[0031] The standard strain DNA of Salmonella enteritidis was extracted according to the method of 1.3 in Example 1, and the DNA concentration was determined to be 138 ng / μl by a spectrophotometer, and it was diluted in proportion and the accurate concentration after dilution was determined. Take 138ng / μl, 55.5ng / μl, 20.5ng / μl, 13ng / μl, 4.2ng / μl, 1.3ng / μl, 0.65ng / μl, 0.2ng / μl, 0.05ng / μl DNA at 9 concentrations Each 1 μl was used as a template for PCR amplification, and the amplification conditions were the same as 1.4 in Example 1. The result is as figure 2 As shown, the results show that the PCR method can detect template DNA with a lower limit of 0.2ng, so as long as the template DNA content in the sample is not less than 0.2ng, this method can be used for effective amplification detection.
Embodiment 3
[0032] Embodiment 3 adopts specific primer of the present invention to carry out PCR detection and adopts traditional separation method to detect the comparison of Salmonella enteritidis
[0033] Get 62 4-week-old white legion laying hens, and inoculate 10 by eating feed 4 After CFU of Salmonella Enteritidis, on the 7th and 14th days, the detection method of Example 1 and the traditional isolation and culture method were used to detect the contents of the cecum, wherein the traditional isolation and culture methods included pre-enrichment, selective enrichment, and isolation and culture , biochemical identification, serological typing a total of 5 steps.
[0034] The contents of the cecum contained many bacteria, mixed strains, and many uncertain interfering factors, which could test the sensitivity and accuracy of the detection method. The results show that the detection rate of the PCR method is obviously higher than that of the traditional culture separation method, and th...
PUM
 Login to view more
Login to view more Abstract
Description
Claims
Application Information
 Login to view more
Login to view more - R&D Engineer
- R&D Manager
- IP Professional
- Industry Leading Data Capabilities
- Powerful AI technology
- Patent DNA Extraction
Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic.
© 2024 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap