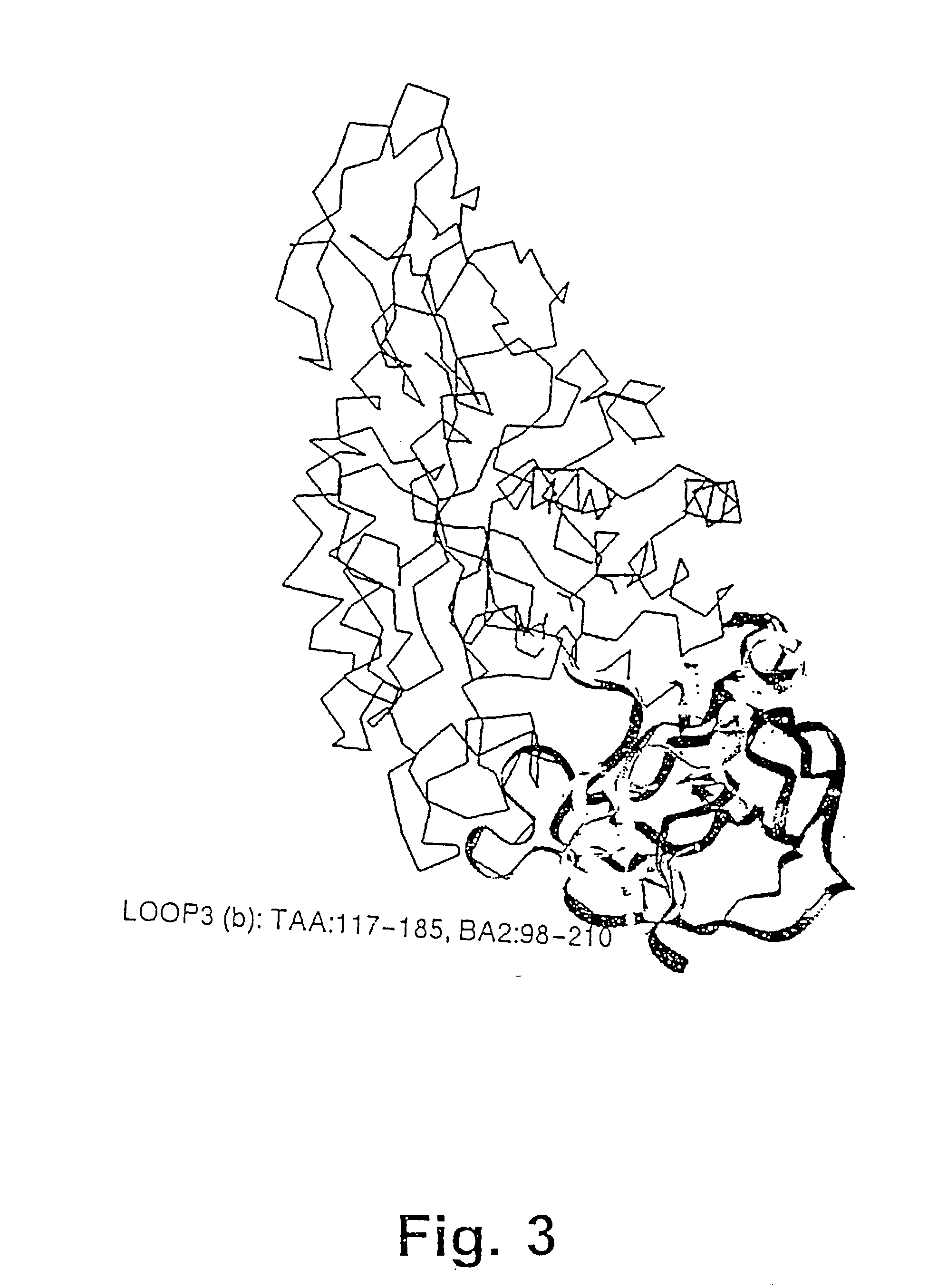Alpha-amylase mutants
a technology of alpha-amylase and mutants, which is applied in the field of alpha-amylase mutants, can solve the problems of difficult to obtain .alpha-amylase crystals suitable for solving structure problems, and it is difficult to determine the three-dimensional structure of all alpha-amylases, so as to improve specific activity, increase stability, and improve stability
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
example 1
[0480] Example on Homology Building of TERM
[0481] The overall homology of the B. licheniformis .alpha.-amylase (in the following referred to as TERM) to other Termamyl-like .alpha.-amylases is high and the percent similarity is extremely high. The similarity calculated for TERM to BSG (the B. stearothermophilus .alpha.-amylase with SEQ ID NO: 6), and BAN (the B. amyloliquefaciens .alpha.-amylase with SEQ ID NO: 4) using the University of Wisconsin Genetics Computer Group's program GCG gave 89% and 78%, respectively. TERM has a deletion of 2 residues between residue G180 and K181 compared to BAN and BSG. BSG has a deletion of 3 residues between G371 and I372 in comparison with BAN and TERM. Further BSG has a C-terminal extension of more than 20 residues compared to BAN and TERM. BAN has 2 residues less and TERM has one residue less in the N-terminal compared to BSG.
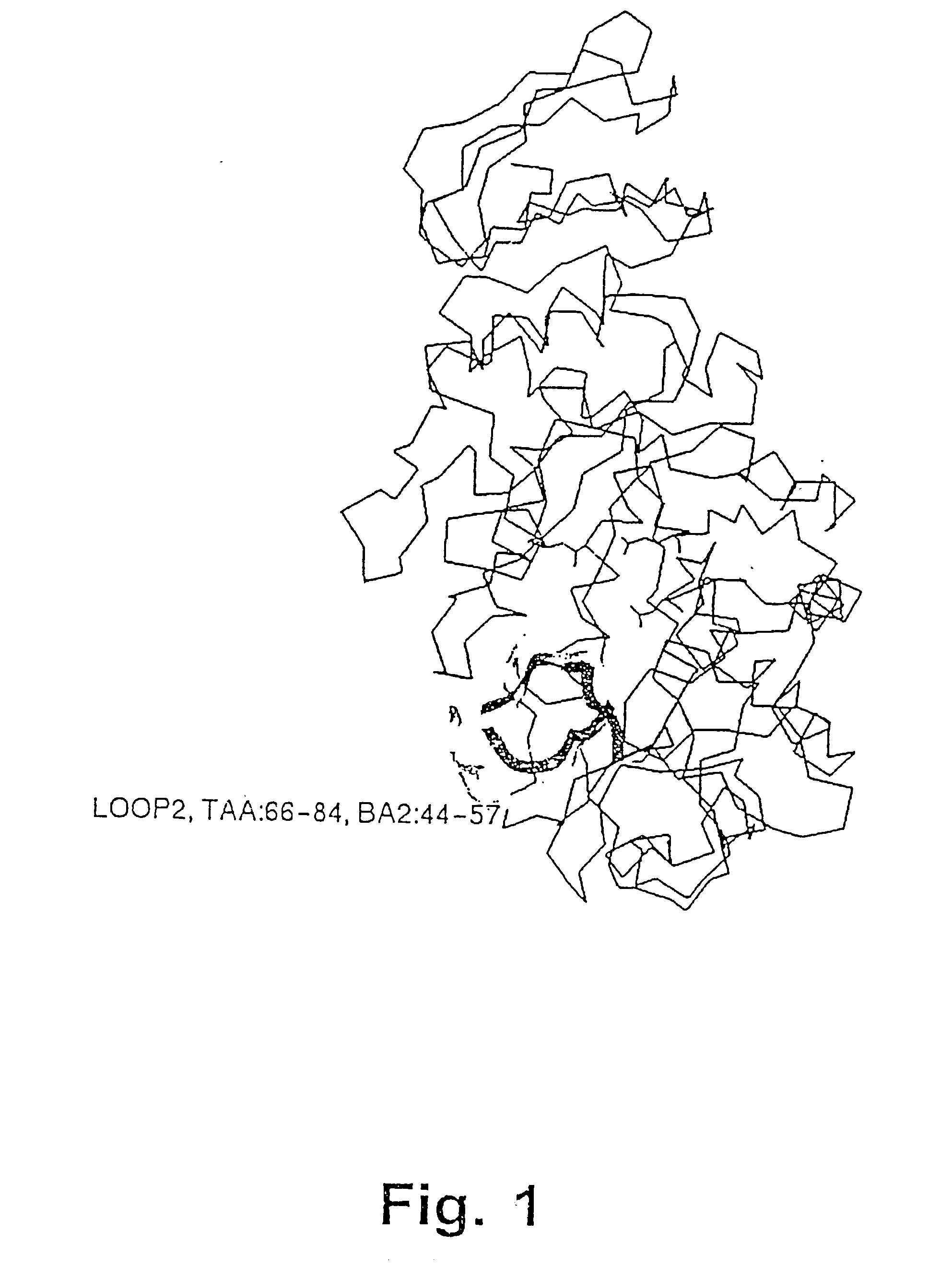
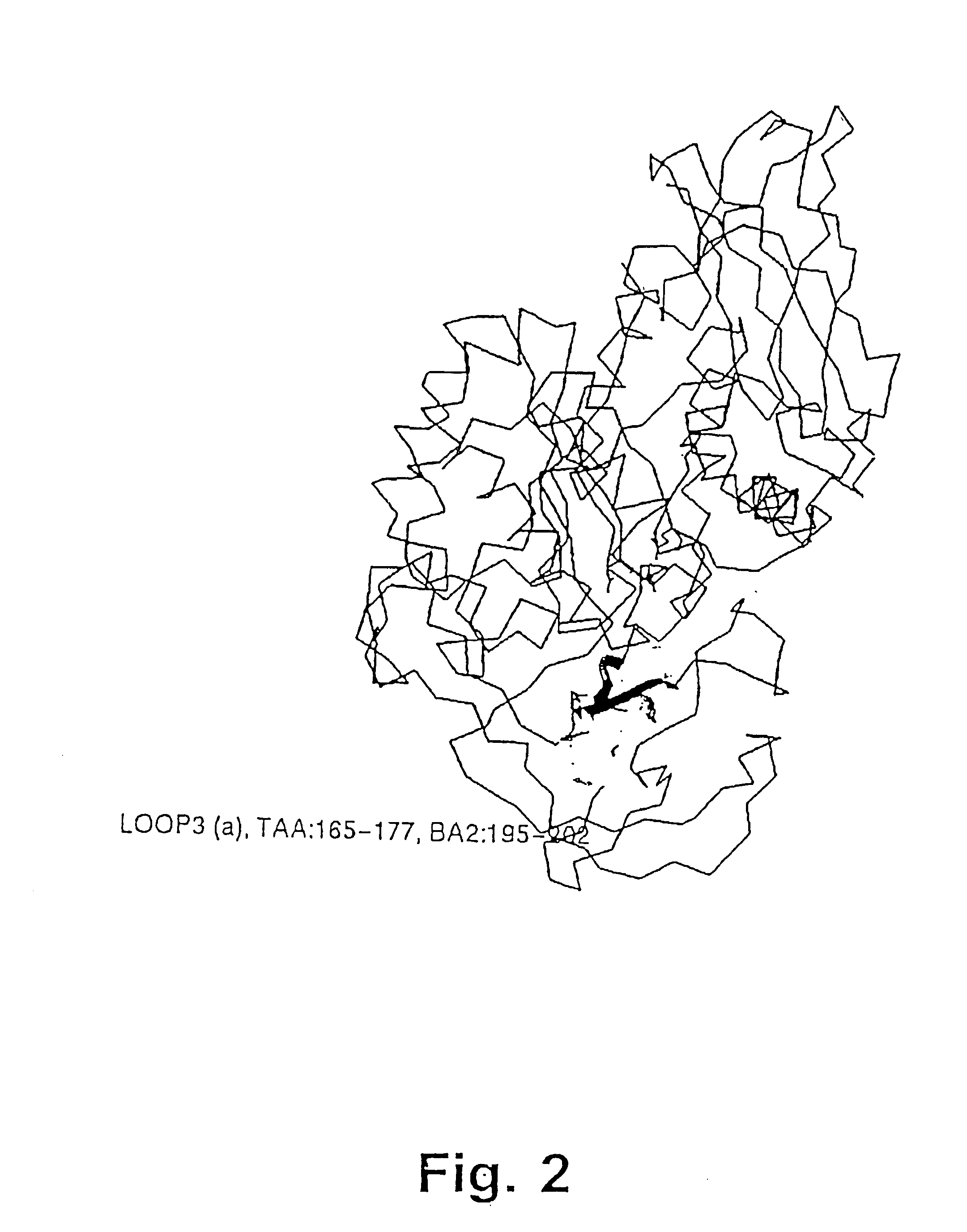
[0482] The structure of the B. licheniformis (TERM) and of the B. amyloliquefaciens .alpha.-amylase (BAN), respectively,...
example 2
[0485] Determination of Residues within 10 .ANG. from the Ions Present in the Solved Structure
[0486] The coordinates of Appendix 1 are read into the INSIGHT program provided by BIOSYM technologies. The spatial coordinates are presented showing the bonds between the atoms. The ions are presented as well as the water atoms. The program package part of creating subset are used to create a 10 .ANG. subset around the Calcium and the Sodium ions in the structure using the command ZONE. All residues having an atom within the 1 .ANG. are compiled and written out by the LIST MOLECULE command. By giving the ions the name ium in the coordinate file a 10 .ANG. sphere around all atoms called ium is compiled. The specific residues identified in this manner are given further above in the section entitled "Ca.sup.2+ dependency".
example 3
[0487] Determination of Cavities in the Solved Structure (Appendix 1)
[0488] The solved structure exhibits many internal holes and cavities. When analysing for such cavities the Connolly program is normally used (Lee, B. and Richards, F. M. (1971) J. Mol. Biol. 55,p. 379-400). The program uses a probe with radius to search the external and internal surface of the protein. The smallest hole observable in this way has the probe radius.
[0489] To analyse the solved structure a modified version of the Connolly program included in the program of INSIGHT were used. First the water molecules and the ions were removed by unmerging these atoms from the solved structure. By using the command MOLECULE SURFACE SOLVENT the solvent accessible surface area were calculated for all atoms and residues using a probe radius of 1.4 .ANG., and displayed on the graphics screen together with the model of the solved structure. The internal cavities where then seen as dot surfaces with no connections to extern...
PUM
| Property | Measurement | Unit |
|---|---|---|
| pH | aaaaa | aaaaa |
| distance | aaaaa | aaaaa |
| distance | aaaaa | aaaaa |
Abstract
Description
Claims
Application Information
 Login to View More
Login to View More