Nucleic acid amplification
a technology of nucleic acid and amplification, applied in the field of nucleic acid amplification, can solve the problems of high cost or insufficient coverage of wga methods, limiting the dna yield of human samples, and insufficient average dna size, etc., and achieves less complicated amplification and more consistent output
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Benefits of technology
Problems solved by technology
Method used
Image
Examples
specific embodiments
[0332]Disclosed is a method of amplifying a target nucleic acid sequence where the method can comprise bringing into contact a set of primers, DNA polymerase, and a target sample, and incubating the target sample under conditions that promote replication of the target sequence. Replication of the target sequence results in replicated strands, where during replication at least one of the replicated strands is displaced from the target sequence by strand displacement replication of another replicated strand. The target sample is not subjected to denaturing conditions. The method can further comprise labeling the replicated strands using terminal deoxynucleotidyl transferase.
[0333]The method can further comprise diluting the replicated strands, bringing into contact a set of primers, DNA polymerase, and the diluted replicated strands, and incubating the replicated strands under conditions that promote replication of the target sequence. Replication of the target sequence results in add...
example 1
A. Example 1
Whole Genome Amplification Using Nuclease Resistant Hexamer Primers
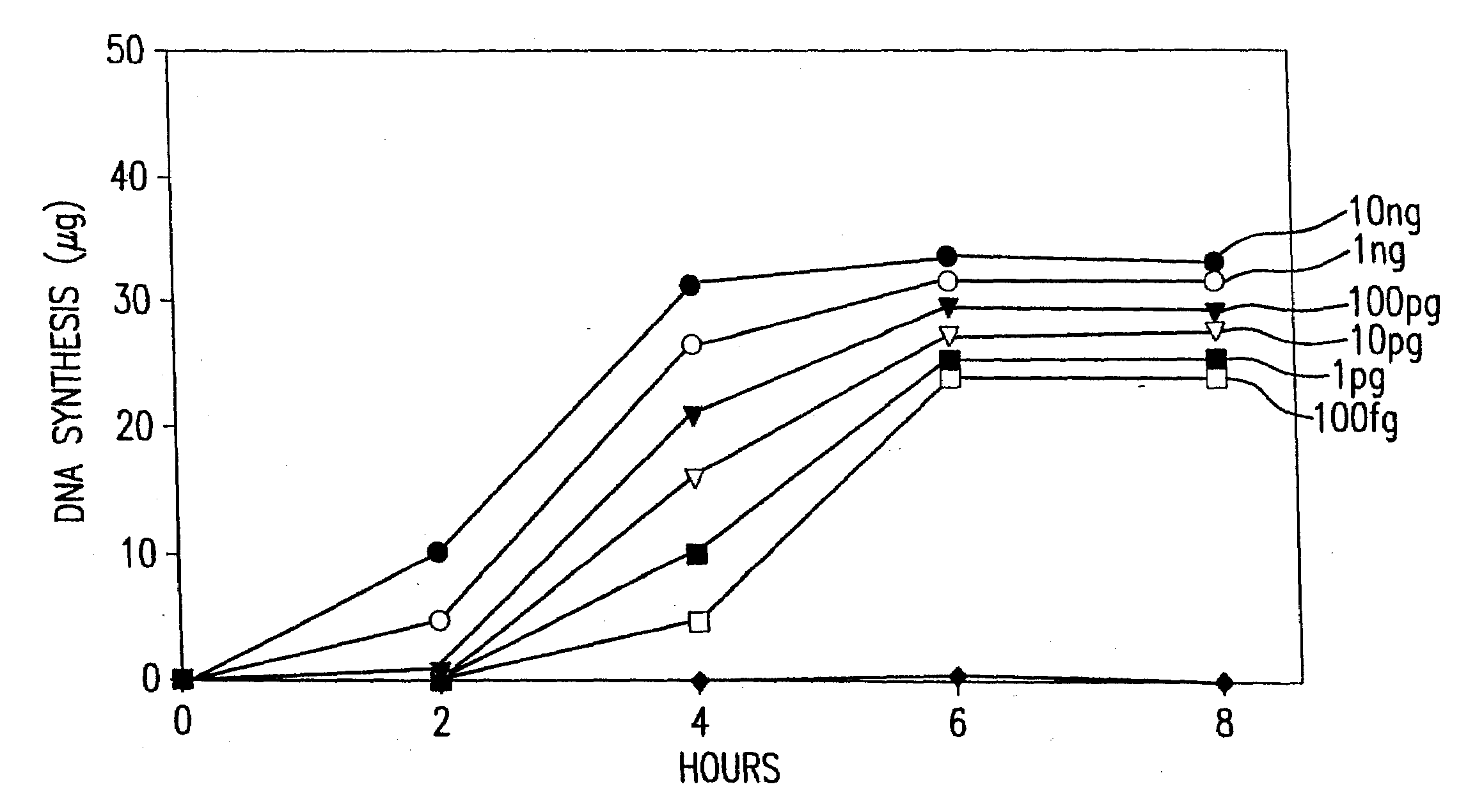
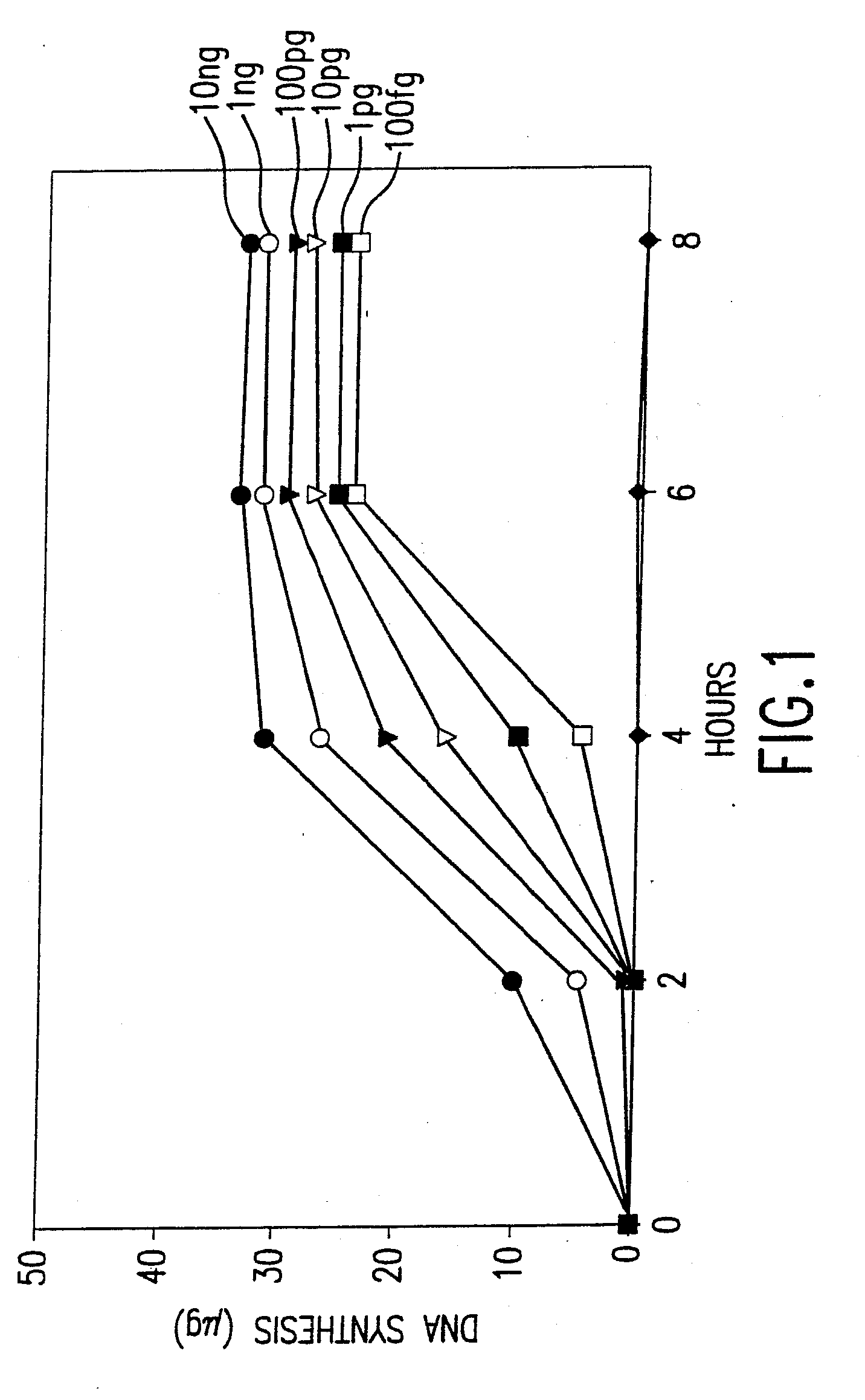
[0637]This example describes a demonstration of an embodiment of the disclosed method and analysis and comparison of the results. The exemplified method is the disclosed multiple displacement amplification form of whole genome amplification using nuclease resistant random hexamer primers. Some reactions in this example were performed without subjecting the sample to denaturing conditions, a preferred form of the disclosed method. In other reactions, the template DNA was subjected to denaturation prior to amplification. MDA was performed using φ29 DNA polymerase.
[0638]1. Materials and Methods
[0639]i. DNA and Enzymes.
[0640]A panel of human genomic DNA samples, the Human Variation Panel-Caucasian Panel of 100 (reference number HD100CAU) was obtained from Coriell Cell Repositories. Human genomic DNA was also obtained from Promega Corp. Thiophosphate-modified random hexamer (5′-NpNpNpNpSNpSN-3′) was synthesize...
example 2
B. Example 2
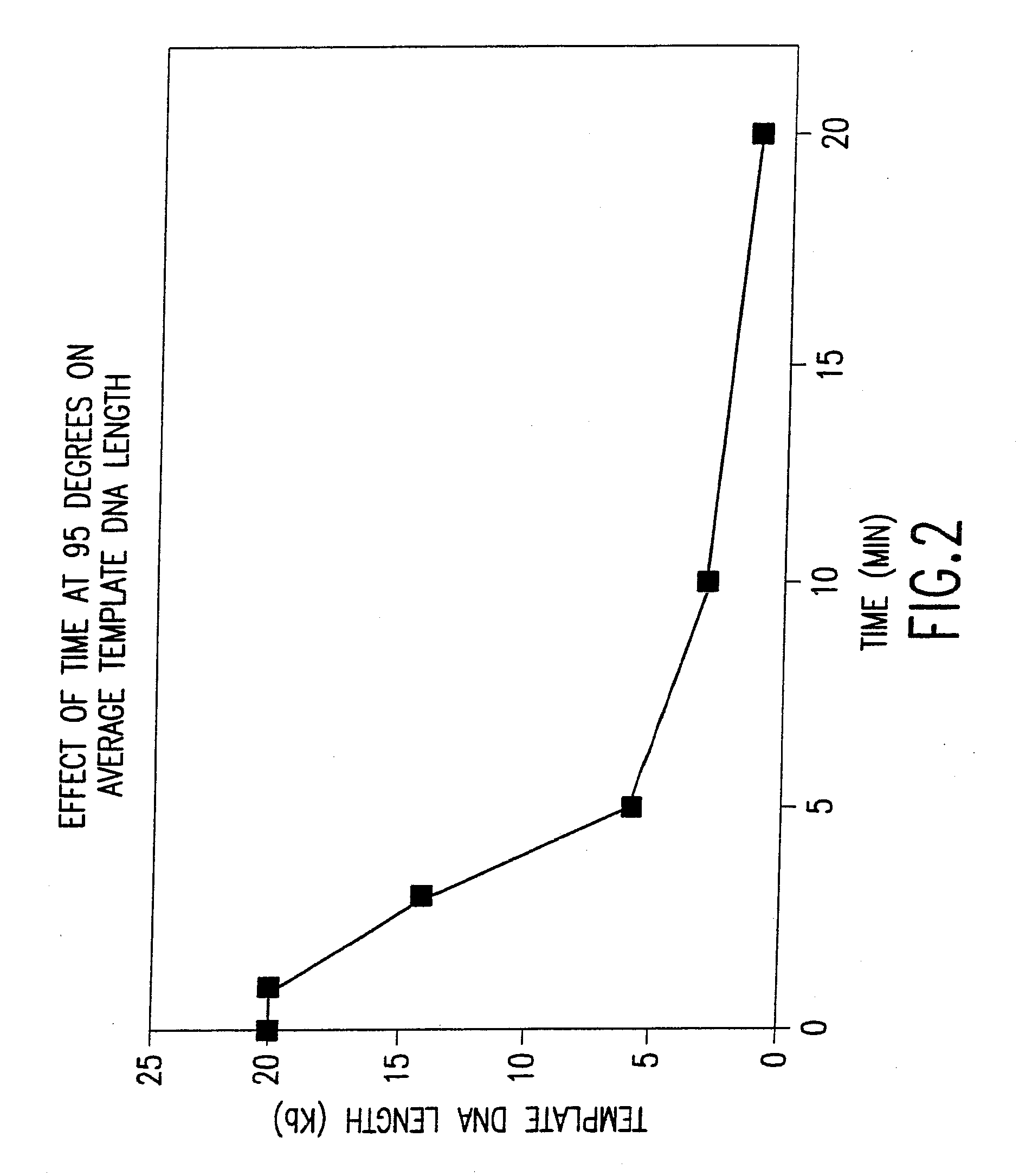
Increasing Time of Incubation at 95° C. Causes Increasing Template DNA Strand Breakage
[0668]This example demonstrates that significant template DNA strand breakage is generated by incubation at 95° C. (which is used in typical amplification reactions to denature the DNA), and that strand breakage is reduced by decreasing the duration of heat treatment. As with most nucleic acid amplification techniques, the integrity of the starting DNA template can have an important effect on the rate and yield of the amplified product. In reactions where the nucleic acid to be amplified is degraded, the yield of amplified product may be reduced both in quality and quantity. This example demonstrates the reduction of template DNA strand breakage by decreasing time of incubation at 95° C.
[0669]Six reactions were carried out under the conditions used for DNA template strand separation with heat-denaturation in order to illustrate the degradation of template DNA. Human genomic DNA (10 μg) ...
PUM
| Property | Measurement | Unit |
|---|---|---|
| pH | aaaaa | aaaaa |
| pH | aaaaa | aaaaa |
| pH | aaaaa | aaaaa |
Abstract
Description
Claims
Application Information
 Login to View More
Login to View More