Quantitative sequencing method for genome DNA modifications in full genome range
A whole-genome, genome-wide technology, applied in the field of genome-wide DNA modification quantitative sequencing, can solve the problems of high data throughput requirements, inability to distinguish 5mC and 5hmC, costing money and labor, and achieve the effect of saving sequencing costs
Inactive Publication Date: 2012-01-25
FUDAN UNIV
View PDF1 Cites 4 Cited by
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
There is incomplete conversion of bisulfite conversion (bisulfite conversion is to convert unmethylated cytosine (C) into uracil (U) in the sequence, while methylated cytosine remains unchanged), data Disadvantages such as large throughput requirements, cumbersome operation, costly and laborious, etc.
Recent studies have also shown that sulfite conversion and methylation-specific cleavage techniques cannot distinguish 5mC from 5hmC
Existing genome-wide methods for detecting DNA methylation or hydroxymethylation modification are only suitable for qualitative analysis, not for quantitative analysis
Method used
the structure of the environmentally friendly knitted fabric provided by the present invention; figure 2 Flow chart of the yarn wrapping machine for environmentally friendly knitted fabrics and storage devices; image 3 Is the parameter map of the yarn covering machine
View moreImage
Smart Image Click on the blue labels to locate them in the text.
Smart ImageViewing Examples
Examples
Experimental program
Comparison scheme
Effect test
Embodiment Construction
[0045] The present invention is further specifically described below by specific examples.
[0046] 1. Extraction of cellular genomic DNA
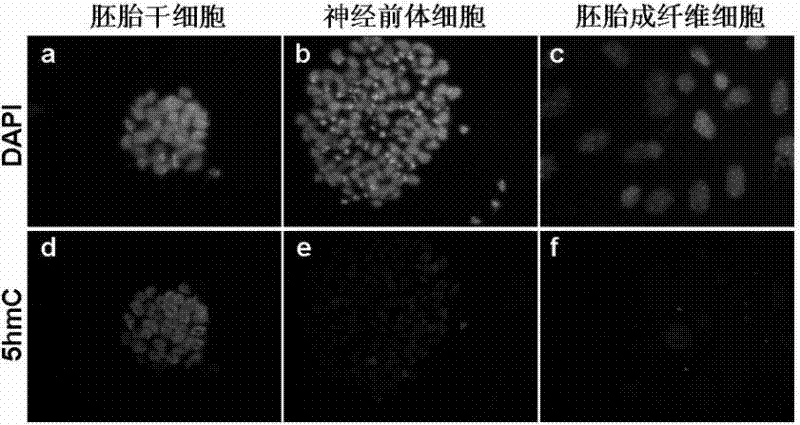
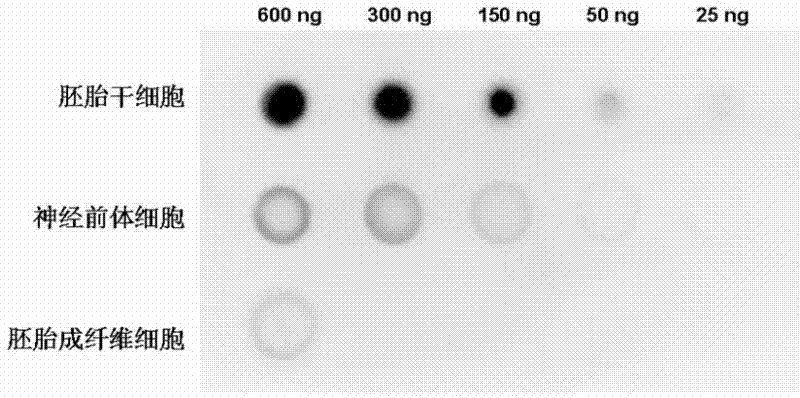
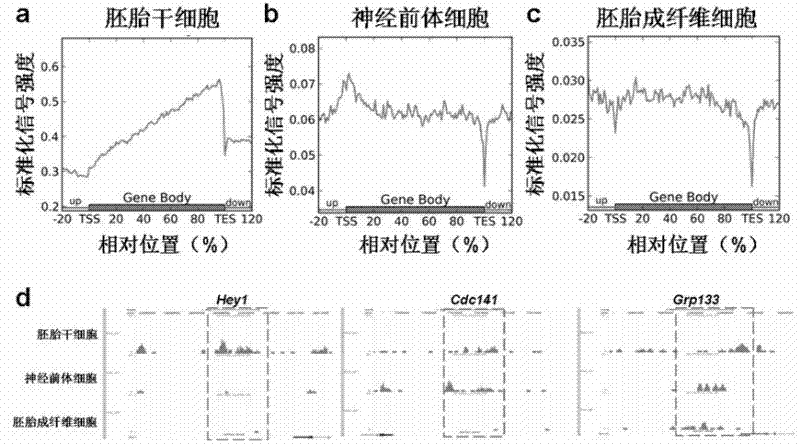
[0047] Collect three kinds of cells: mouse embryonic stem cells (mESCs), neural precursor cells (NPCs) and fibroblasts (MEFs) differentiated from mouse embryonic stem cells, and extract genomic DNA respectively (Blood & Cell Culture DNA Mini Kit, Qiagen).
[0048] 2. Barcode sample preparation
the structure of the environmentally friendly knitted fabric provided by the present invention; figure 2 Flow chart of the yarn wrapping machine for environmentally friendly knitted fabrics and storage devices; image 3 Is the parameter map of the yarn covering machine
Login to View More PUM
 Login to View More
Login to View More Abstract
The invention belongs to the technical field of second generation gene sequencing, in particular to a quantitative sequencing method for genome DNA modifications (such as 5-hydroxymethylcytosine (namely 5hmC), 5-methylation cytosine (namely 5mC), 5-carboxyl cytosine (namely 5caC), 5-formoxyl cytosine (namely 5fC) and the like) in a full genome range. Specific sequence tags (namely Barcode) are respectively added to multiple to-be-detected samples by adopting a tag method, and the samples are mixed together for DNA fragment immuno-precipitation) of corresponding modifications, so that quantitative sequencing of the genome DNA modifications in the full genome range is realized. The method comprises the following steps of: extraction of genome DNA, preparation of Barcode samples, DNA fragment immuno-precipitation experiments of the corresponding modifications, polymerase chain reaction (PCR) amplification of specific products, and sequencing on an illumina GA ll platform. The method can be used for quantitatively comparing the DNA of the corresponding modifications in multiple samples at the same time in the full genome range and accurately positioning the content difference of the DNA of different areas and different modifications.
Description
technical field [0001] The invention belongs to the technical field of second-generation gene sequencing, and in particular relates to a quantitative sequencing method for genome-wide DNA modification. Background technique [0002] Epigenetics is the study of heritable phenotypic changes that do not involve changes in DNA sequence. Epigenetic modifications mainly include DNA methylation / hydroxymethylation, covalent modification of histones, non-coding RNA, etc. Epigenetic modification plays an important role in embryonic development, cell differentiation, X chromosome inactivation, gene imprinting, etc. Abnormal epigenetic modification is associated with many diseases (Feinberg AP. Phenotypic plasticity and the epigenetics of human disease. Nature 2007; 447: 433-40. Ballestar E. Epigenetic alterations in autoimmune rheumatic diseases. Nat Rev Rheumatol 2011;7:263-71.), has important research significance. Genome-wide detection of epigenetic modification differences in biol...
Claims
the structure of the environmentally friendly knitted fabric provided by the present invention; figure 2 Flow chart of the yarn wrapping machine for environmentally friendly knitted fabrics and storage devices; image 3 Is the parameter map of the yarn covering machine
Login to View More Application Information
Patent Timeline
 Login to View More
Login to View More IPC IPC(8): C12Q1/68
Inventor 石雨江施扬谭理熊莉君吴飞珍
Owner FUDAN UNIV