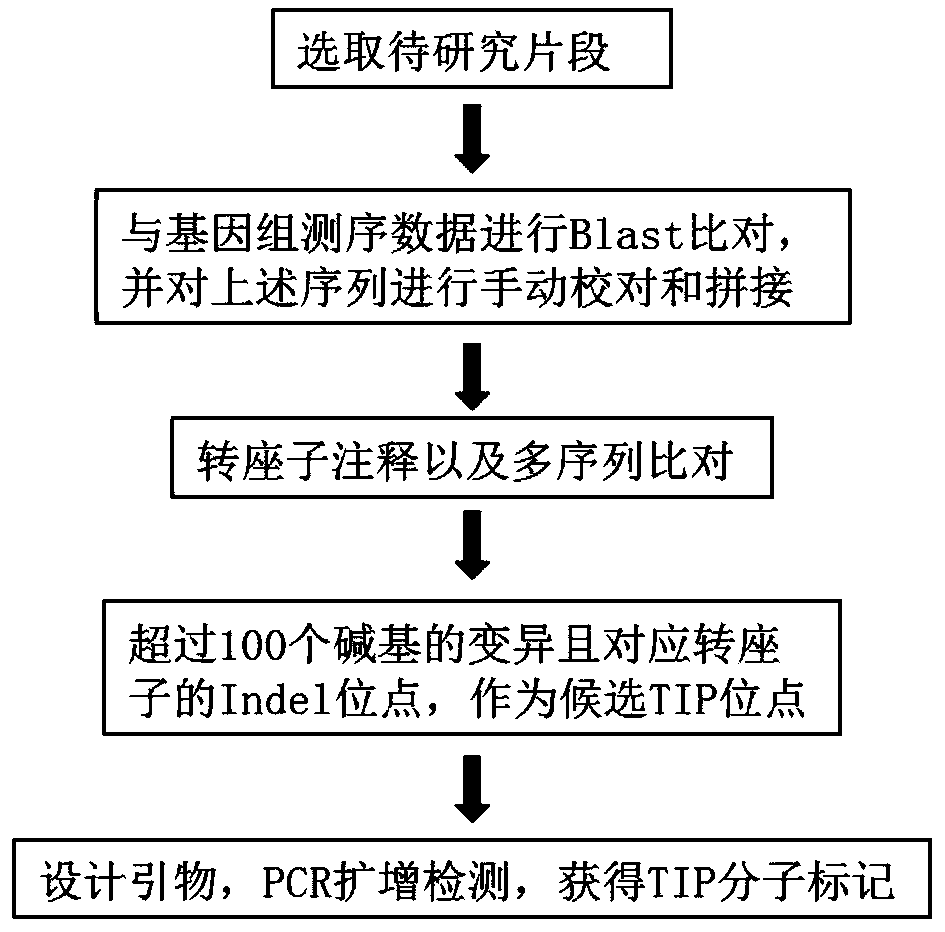
Method for mining transposon insertion polymorphisim TIP molecular markers based on comparative genomics
A comparative genomics and molecular marker technology, applied in the field of animal genetics and breeding, can solve the problem of lack of molecular markers in fine positioning, and achieve the effect of clear and easy judgment of test results, clear results, and convenient marker detection.
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment Construction
[0028] Before further describing the specific embodiments of the present invention, it should be understood that the protection scope of the present invention is not limited to the following specific specific embodiments; it should also be understood that the terms used in the examples of the present invention are to describe specific specific embodiments, It is not intended to limit the protection scope of the present invention.
[0029] The following is a detailed description in conjunction with the development of the TIP molecular marker in the porcine GHR gene.
[0030] 1. Sequence acquisition
[0031]The reference sequence of the GHR gene (Gene ID: 397488) was obtained from the NCBI database (https: / / www.ncbi.nlm.nih.gov / ). Then use the reference sequence to compare with the genome sequencing data of different breeds of pigs in NCBI's WGS database, so as to obtain the sequences of GHR genes (including reference sequences) in 15 sequenced genomes, and the GHR sequences of...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More