Detection kit for salmonella and shigella in feeds and detection method thereof
A detection kit and salmonella detection technology, applied in biochemical equipment and methods, microorganism-based methods, material separation, etc., can solve the problems of easy pollution, different enrichment time, high detection cost, etc., to save labor and financial resources , The effect of short detection time and simple operation
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0038] Embodiment 1, establishment of Salmonella and Shigella detection kit and detection method thereof in feed
[0039] (1) Design and synthesis of primers and assembly of kits:
[0040]
[0041] The above primers were synthesized by Bao Biological Engineering (Dalian) Co., Ltd.
[0042] On this basis, design the kit that is used for PCR-DHPLC detection: Include a kind of detection solution in this kit, contain 10mM Tris Cl, 50mM KCl, 25mM MgCl in this detection solution 2 , 2.5 mM each of dNTP (dATP, dGTP, dCTP and dTTP), 5 U / μL of Taq DNA polymerase, and 10 μM each of Salmonella and Shigella primer pairs;
[0043] (2) PCR-DHPLC detection method
[0044] This detection method uses the PCR-DHPLC detection kit that the present embodiment establishes, comprises the following steps:
[0045] ① Preparation of the sample to be tested: the DNA genome of the sample to be tested is prepared by the kit extraction method
[0046] a. Weigh 25g of animal-derived feed sample, add ...
Embodiment 2
[0067] Embodiment 2, mPCR-DHPLC detection method specificity test
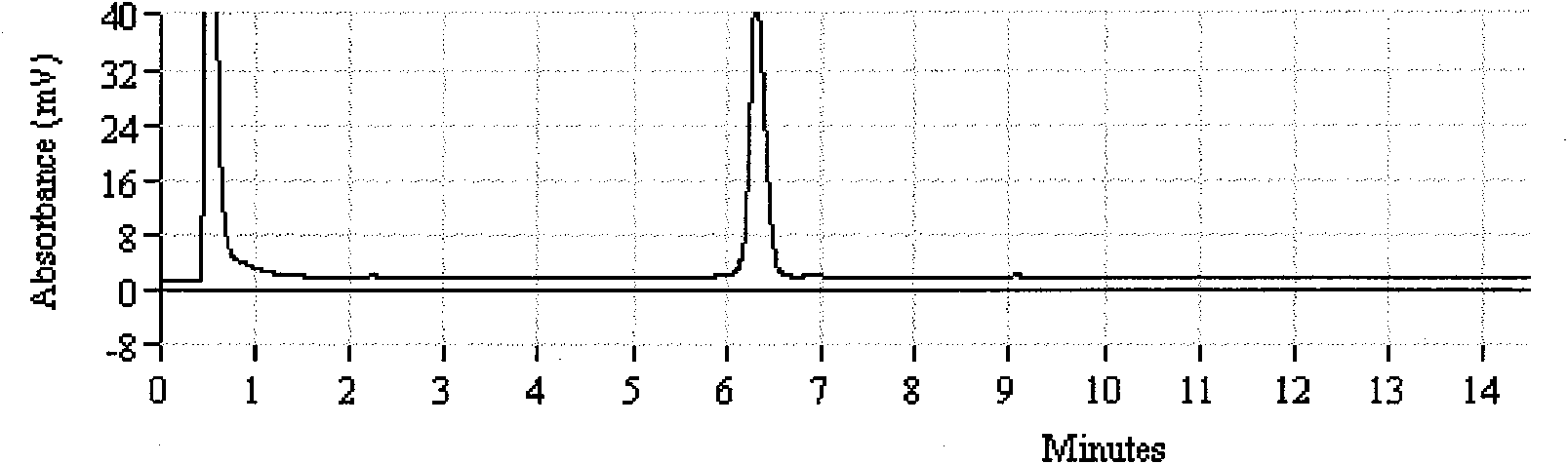
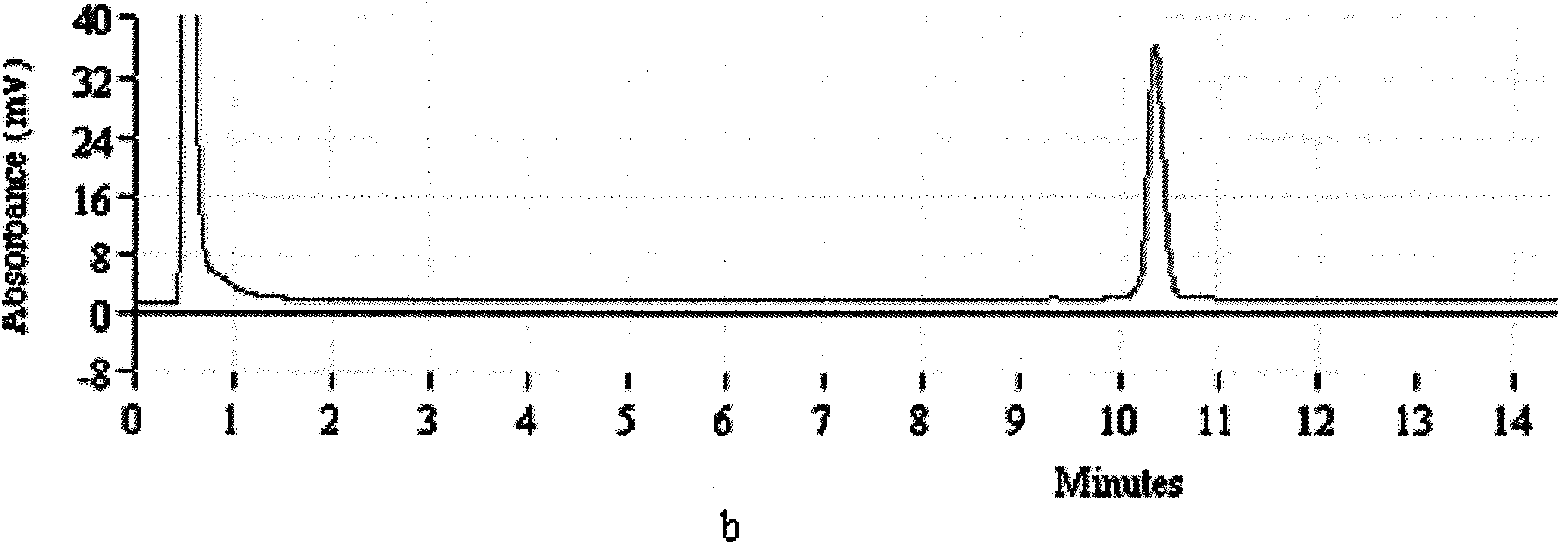
[0068] Take the test strains listed in Table 1, extract the genomic DNA after culture, and establish the template library. . Then, using these genomic DNAs as templates, PCR-DHPLC analysis and detection were carried out using the method established in the examples. The test results show that all Salmonella use have detected the Salmonella positive absorption peak of DHPLC (attached Figure 1.a ), the peak time is about 6.3min, and the non-Salmonella test results are all negative; Shigella have all detected the Shigella positive absorption peak of DHPLC (attached Figure 1.b ), the peak time was 10.4min, and the non-shigella test results were all negative.
[0069] The above results show that: the detection primers for Salmonella and Shigella used in this test have good identification specificity.
[0070] Table 1
[0071]
[0072]
[0073]
Embodiment 3
[0074] Embodiment 3, composite PCR-DHPLC method specificity test
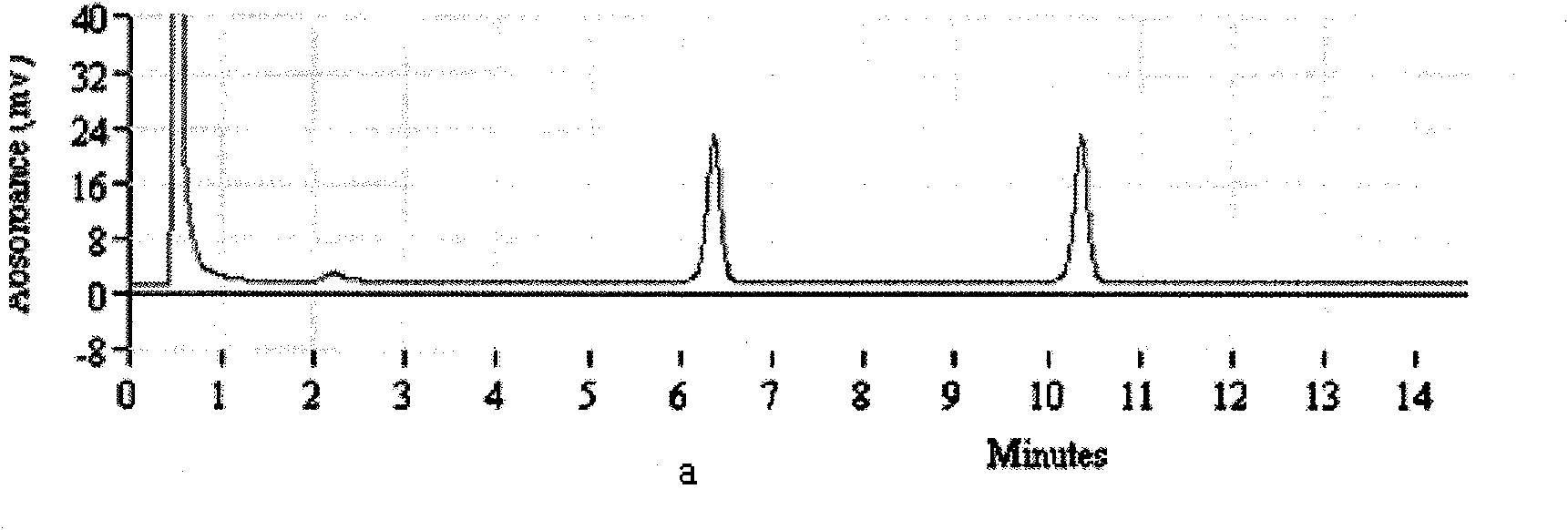
[0075] Take the compound universal enrichment culture solution of Salmonella enteritidis and Shigella flexneri, that is, the mixed culture enrichment solution of two standard strains to extract DNA, and perform PCR-DHPLC detection according to the method established in Example 1. The results are attached Figure 2.a As shown, the peaks of Salmonella and Shigella were detected at 6.3min and 10.4min, respectively.
[0076] As a comparison, in the embodiment, 2.5 μL of each single PCR amplified product of Salmonella enteritidis and Shigella flexneri was mixed and then detected by DHPLC under the same conditions. The results are shown in the attached Figure 2.b As shown, Salmonella and Shigella also detected their respective positive absorption peaks, and the peak time was the same as above.
[0077] In order to confirm that the positive absorption peaks detected by the composite PCR-DHPLC are the gene fragments...
PUM
 Login to view more
Login to view more Abstract
Description
Claims
Application Information
 Login to view more
Login to view more - R&D Engineer
- R&D Manager
- IP Professional
- Industry Leading Data Capabilities
- Powerful AI technology
- Patent DNA Extraction
Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic.
© 2024 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap