Method and system for realizing large-scale database clustering by double-buffer model
A technology of model implementation and clustering method, applied in the field of data processing, can solve the problem of time-consuming data reading, achieve the effect of low utilization rate and improve parallelism
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
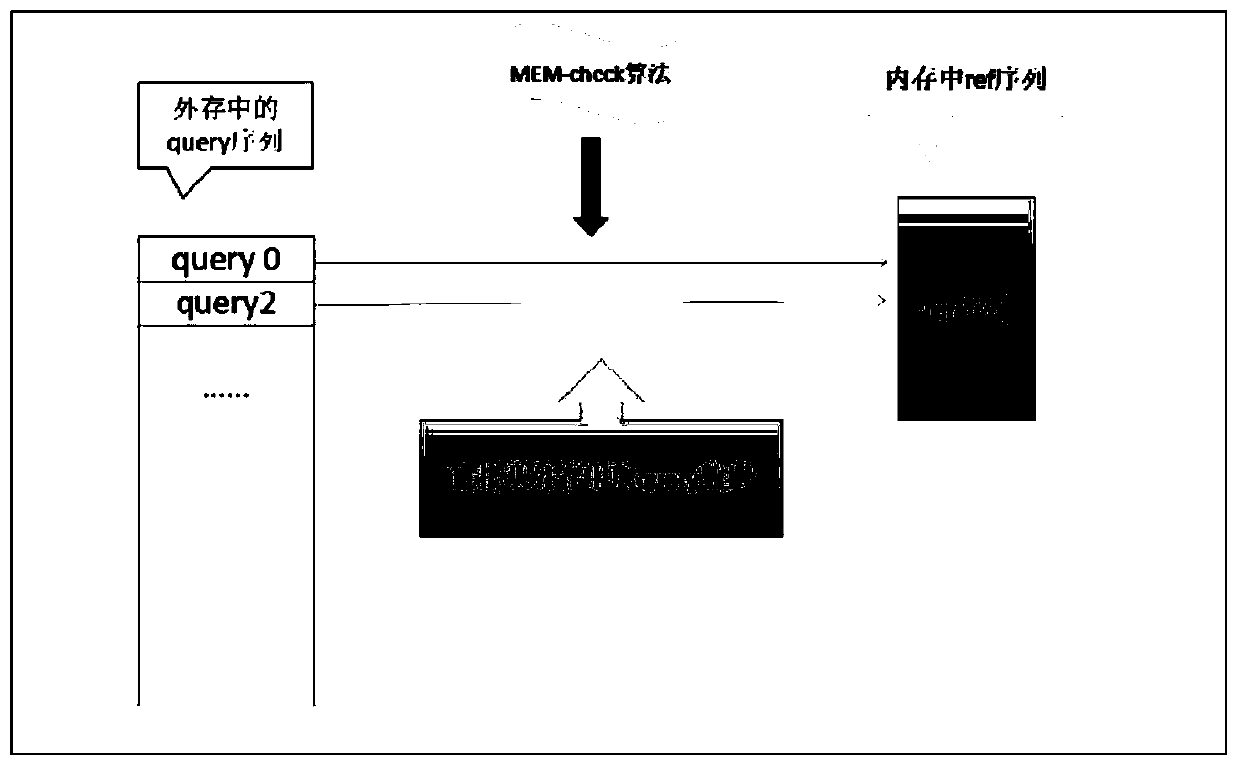
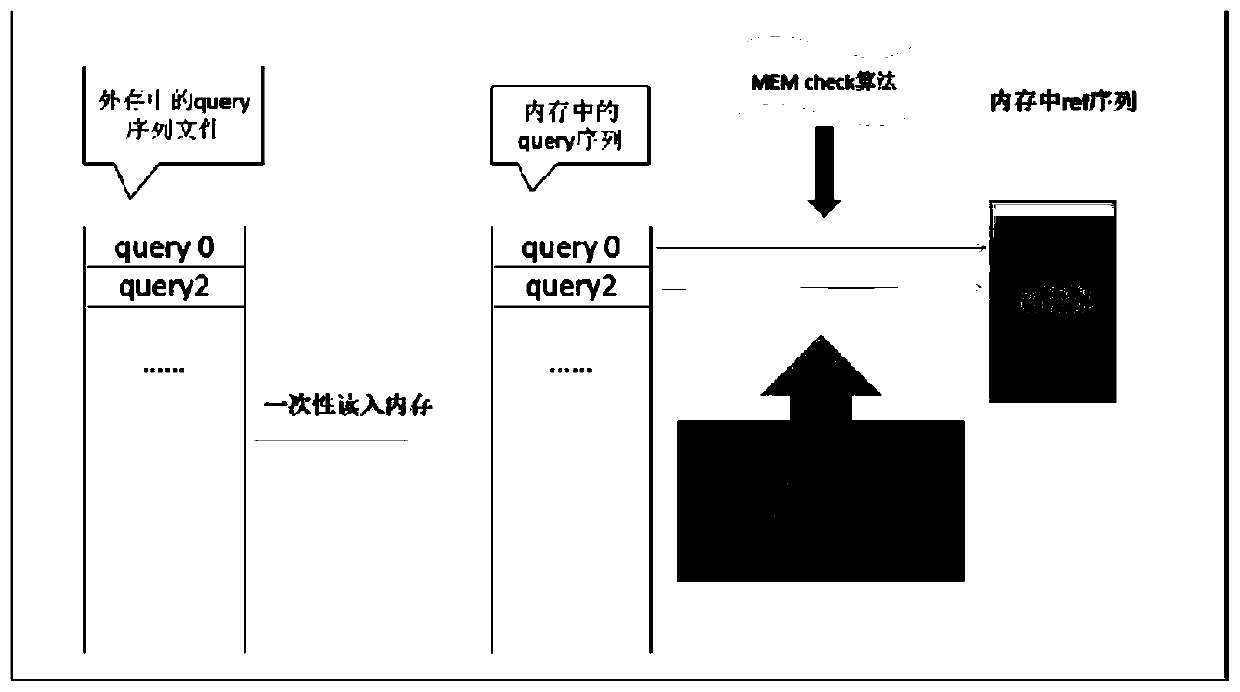
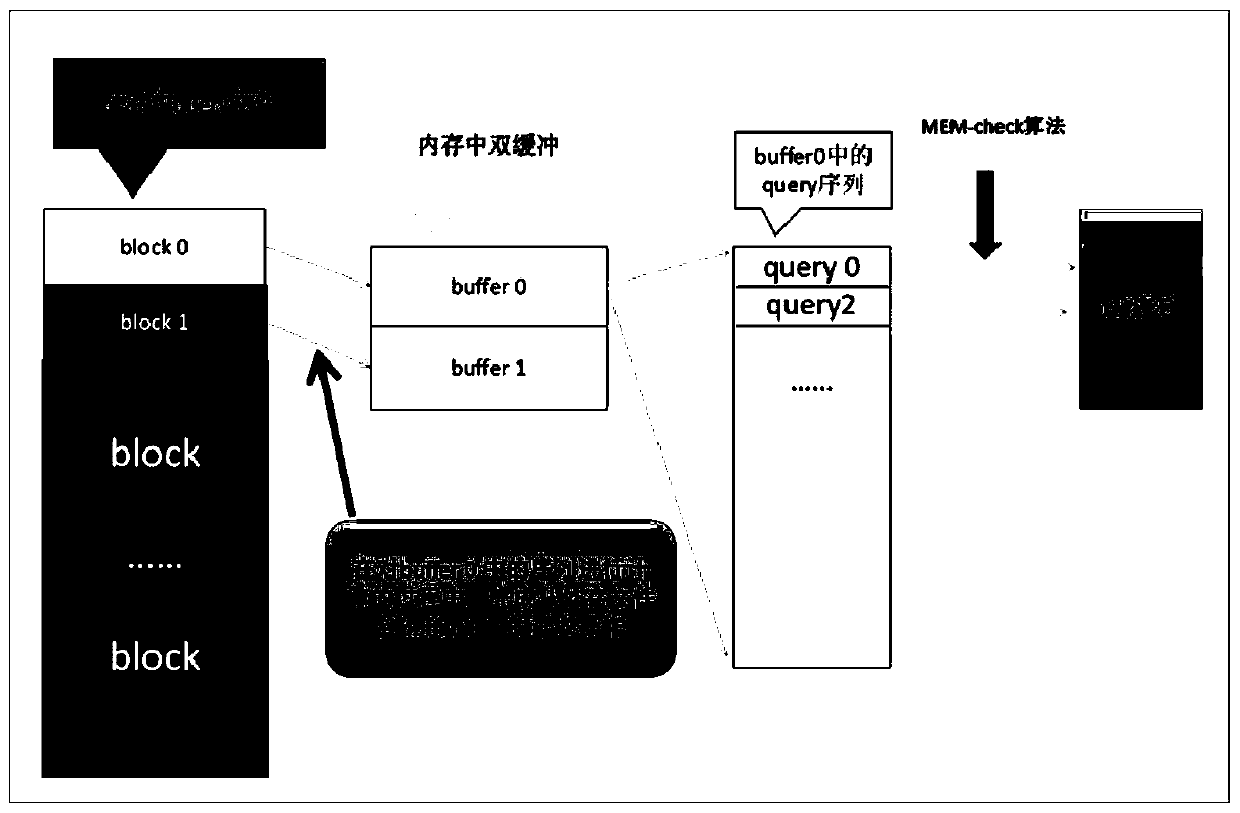
[0046] This embodiment discloses the general flow of the double buffering model to realize the large-scale database clustering algorithm and the MEMs maximum exact matching algorithm:
[0047] Sorting in descending length against gene sequence databases. When performing similarity matching on two sequences, the default longer one is the representative sequence, and the shorter one is the redundant sequence. Therefore, the first item after sorting must be a representative sequence, and the lower one whose similarity reaches the threshold is marked as its redundant sequence.
[0048]The implementation of the algorithm needs to build a matching dictionary first. The specific implementation method is Sparse Suffix Array (Sparse SuffixArray, SSA). A gene sequence is constructed as a sparse suffix array as a dictionary, and other gene sequences are matched with the dictionary suffix array. During the matching process, the query A certain position of the sequence adopts binary searc...
Embodiment 2
[0068] The purpose of this embodiment is to provide a double-buffering model to realize a large-scale database clustering system, including a memory, a processor, and a computer program stored on the memory and operable on the processor, and the processor implements the following when executing the program. steps, including:
[0069] Sorting in descending length for the gene sequence database;
[0070] Build a matching dictionary: sparse suffix array, build a sparse suffix array with a gene sequence as a dictionary, and match other gene sequences with the dictionary suffix array. During the matching process, a binary search is used at a certain position of the query sequence to search, and an inverse suffix is used The array, the minimum common sub-prefix array, and the suffix link are optimized and upgraded. After the calculated matching value reaches the threshold, it is determined to be a redundant sequence.
PUM
 Login to view more
Login to view more Abstract
Description
Claims
Application Information
 Login to view more
Login to view more - R&D Engineer
- R&D Manager
- IP Professional
- Industry Leading Data Capabilities
- Powerful AI technology
- Patent DNA Extraction
Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic.
© 2024 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap