Reverse transcriptase mutant for nucleic acid detection
A mutant, reverse transcription technology, applied in the biological field, can solve problems such as the uncertainty of enzyme transformation
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0120] Calculation and screening of each site DDG value in embodiment 1
[0121] Input the MMLV protein sequence into the Rosetta algorithm software Cyrus Bench (Cyrus Biotechnology), and carry out the whole position of amino acid segments 0-100, 101-200, 201-300, 301-400, 401-500, 501-600, 601-671 Calculate the DDG value of the point full mutation, and the information of the mutation site with a significantly lower DDG value (DDG value <-2) is as follows:
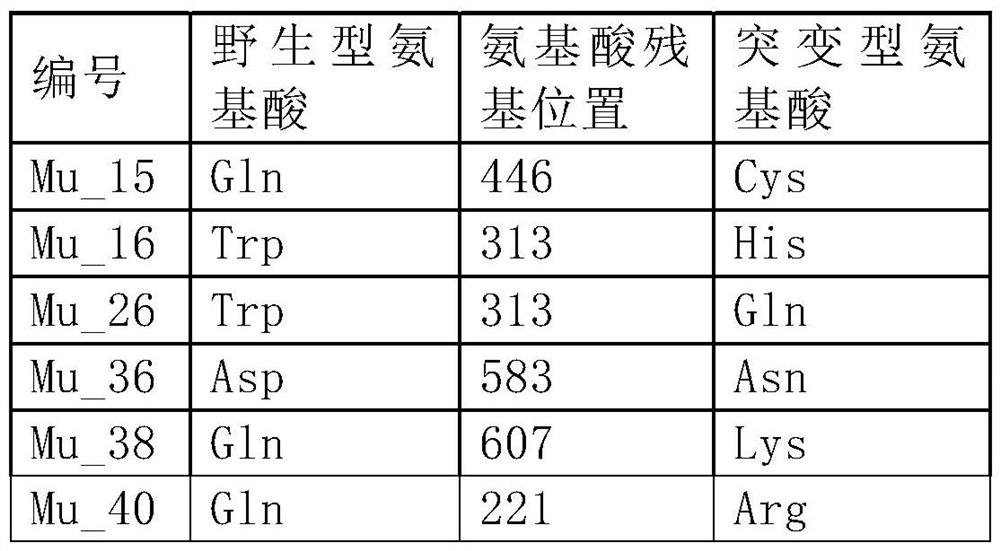
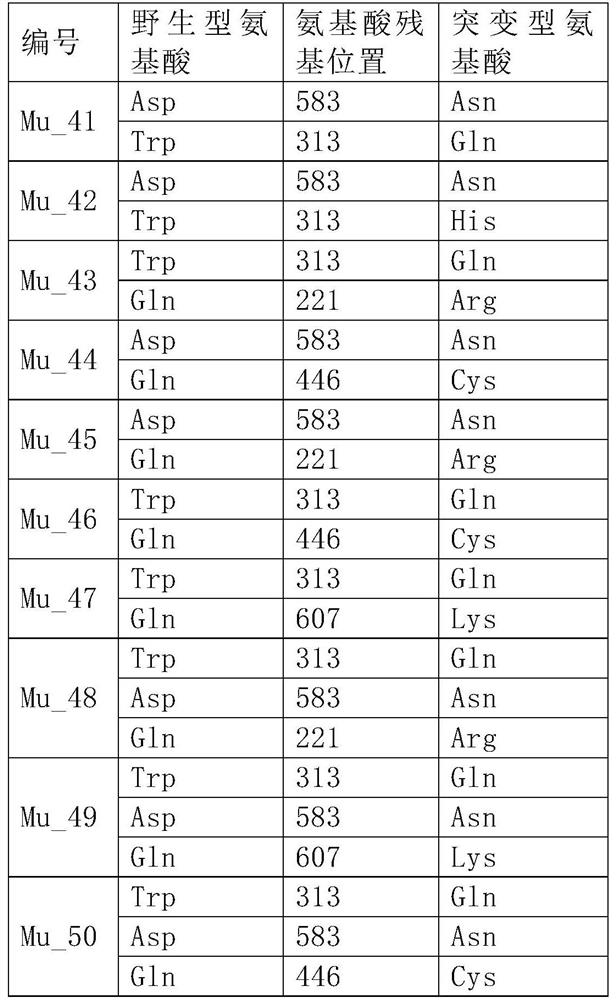
[0122] Table 1
[0123]
[0124]
Embodiment 2
[0125] Example 2 Construction of MMLV mutation library
[0126] According to the above protein sequence, codon optimization was performed by Suzhou Jinweizhi Biotechnology Co., Ltd., and the DNA sequence (SEQ ID NO.: 2) was compiled.
[0127] Suzhou Jinweizhi Biotechnology Co., Ltd. carried out gene synthesis based on the above DNA sequence, added 5' (NheI) and 3' (XhoI) restriction enzyme sites, and cloned the gene into the vector pET28a through 5'NheI and 3'XhoI, Construct plasmid WT-pET28a, prepare recombinant plasmid DNA and glycerol bacteria containing the recombinant plasmid, and perform site-directed mutation on plasmid WT-pET28a according to the mutation sites involved in Example 1, and construct mutation libraries Mu1-pET28a~Mu40-pET28a .
Embodiment 3
[0128] Example 3 Expression and purification of MMLV mutants
[0129] Transform WT-pET28a, Mu1~40-pET28a plasmids into BL21(DE3) competent cells to obtain 37 expression host bacteria, then transfer to 3ml LB medium, shake and culture at 37°C for 5 hours, then add 0.1Mm IPTG at 18°C Induce the culture overnight. Collect the induced cells, add lysate (50Mm Tris, 50Mm NaCl, pH7.5), ultrasonically lyse, and centrifuge to separate the supernatant. Take the supernatant and purify it with Ni NTA metal ion chelating filler to obtain wild type and 40 kinds of mutant MMLV proteins
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More