Endpoint TAQMAN methods for determining zygosity of corn comprising TC1507 events
A TC1507, event technology, applied in the ment field, can solve problems such as not using reference genes
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0087] Example 1: Isolation and quantification of total genomic DNA and PCR primer amplification. Genomic DNA from Cry1F homozygous, hemizygous and wild-type samples was isolated from individual samples by punching 8 leaf discs per sample and grinding the discs to a fine powder using a Genogrinder 2000. use custom DNA was extracted using the gDNA Plant Kit (Invitrogen, Carlsbad, CA) or Qiagen DNeasy 96-well kit (Valencia, CA). Before PCR, use Quant-iT TM DNA samples were quantified using a quantification kit (Invitrogen, Carlsbad, CA) using the manufacturer's instructions.
[0088] Oligonucleotide primers and dual-labeled TaqMan probe with FAM and Black Hole Quencher 1 (BHQ1) were synthesized by MWG Biotech (High Point, NC) (Table 1b).
[0089] Table 1b: Sequences of primers and dual-labeled TaqMan probes
[0090]
[0091]
[0092] Dual-labeled TaqMan probes with Cy5 and BHQ2 were synthesized by IDT (Integrated DNA Technologies, Coralville, IA). All primers were ...
Embodiment 2
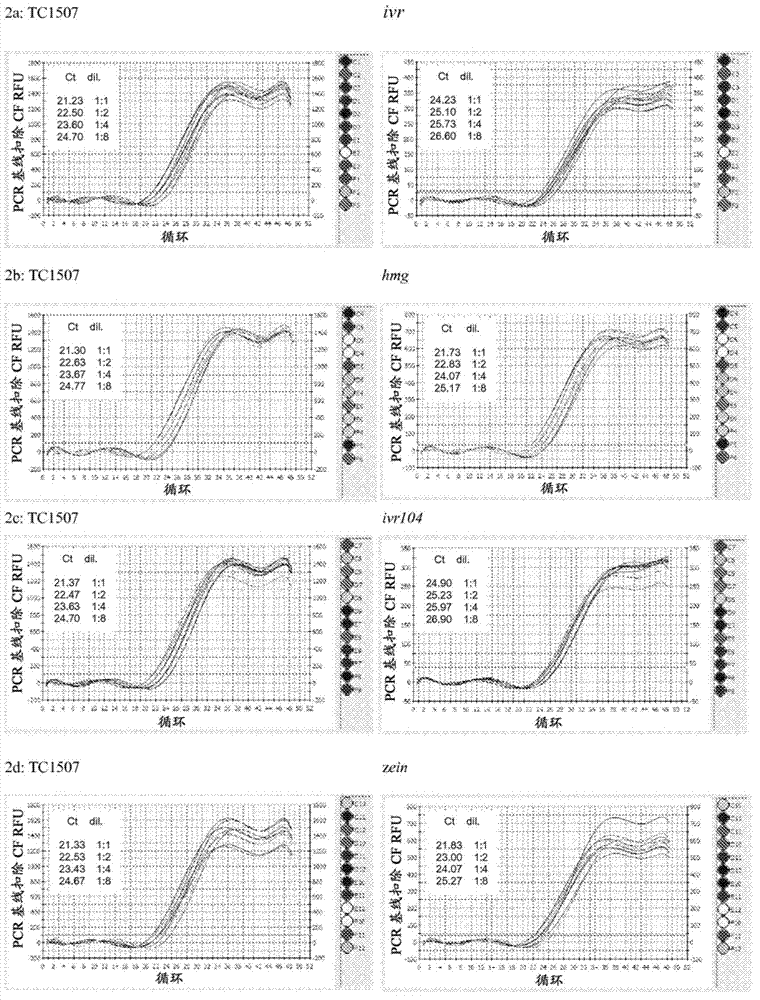
[0109] Example 2: PCR efficiency test of maize endogenous genes. One aspect of developing an endpoint TaqMan zygosity assay is the selection of the most appropriate endogenous gene as a reference gene. We selected species-specific invertases with low copy numbers in the genome, ie a suitable reference gene. Four maize endogenous genes were initially investigated among thousands of possibilities, namely alcohol dehydrogenase 1 (adh1), hypermobility group a (hmga), invertase (ivr), and zein (zein) As a possible reference gene for maize Cry1F in event TC1507.
[0110] The process of selecting an appropriate reference gene involved first combining 30 ng of extracted Cry1F homozygous, hemizygous and wild-type maize genomic DNA controls (extracted according to the isolation protocol described in this example) to assess the PCR efficiency of all primers. PCRs for TC1507 and the 5 initially selected reference genes (ivr, ivr104, adh, hmg, zein) were set up according to Table 2c. T...
Embodiment 3
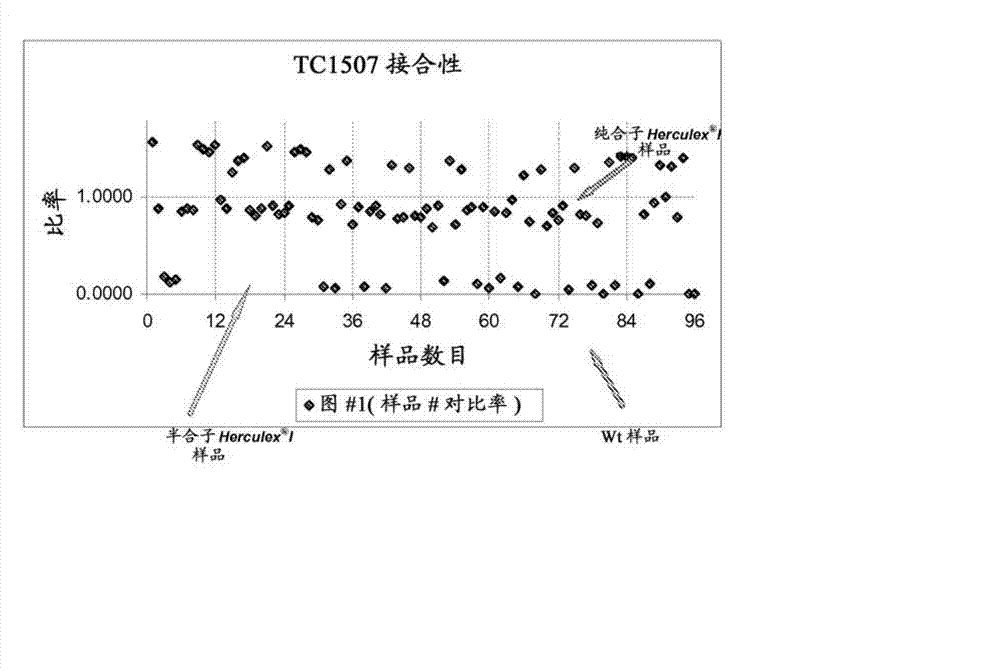
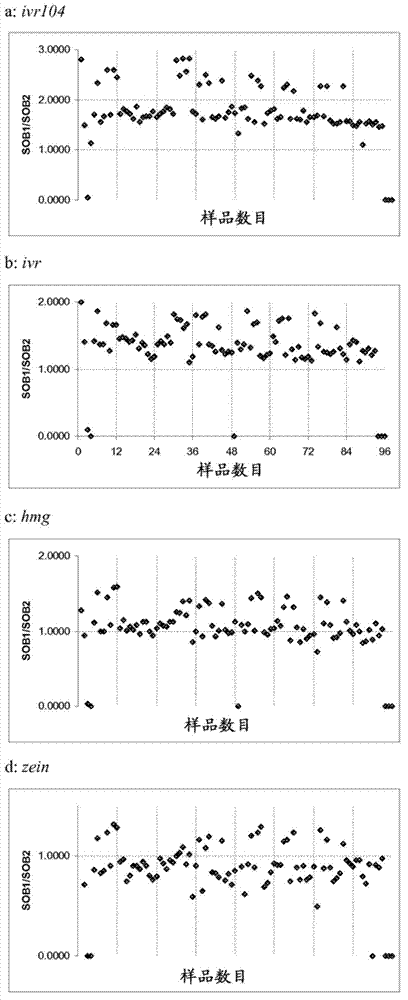
[0118] Example 3: Testing of an endpoint TaqMan assay for zygosity genotyping. Multiplexing of one reference gene (ivr, ivr104, hmg or zein) with TC1507 was tested using endpoint TaqMan PCR (Table 3) using a Cry1F single stack population with segregating TC1507 (Q:07K:PF04DS_ZYGO) . DNA was normalized to 10 ng / μl prior to endpoint TaqMand PCR. TaqMan PCR reactions were terminated at 28 cycles and then measured with a spectrofluorometer. Calculated above background (H 2 O) of FAM (TC1507) (as signal above background 1 (SOB1)), and the fluorescent signal of Cy5 (reference gene) 2 (SOB2) above background. Plot the SOB1 / SOB2 ratio as a scatter plot in Excel. In segregating populations, three clusters of data points should be obtained, allowing for visual determination of cut-off points. It was found that Ivr104 multiplexed with TC1507 only under the reaction conditions disclosed herein can produce unambiguous genotype calls. Other initially selected reference gene responses...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More