Myocardial infarction biomarker miR-1283
A technology of mir-1283 and myocardial infarction, applied in the field of application of myocardial infarction biomarker miR-1283, miR-1283 and its target genes in the diagnosis and treatment of myocardial infarction
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0047] Example 1 Collection of samples and extraction of total RNA
[0048] The peripheral blood of patients with myocardial infarction and 6 cases of healthy controls were collected in the hospital from March 2015 to September 2016. Diagnostic criteria for acute myocardial infarction: developed according to the diagnostic criteria recommended by the third "Global Unified Definition of Myocardial Infarction" in 2012. An increase and / or decrease in the level of myocardial necrosis markers (mainly troponin) was detected, exceeding the 99th percentile value of the upper limit of the reference value at least once, and accompanied by at least one of the following ischemic symptoms.
[0049] 1) Symptoms of myocardial ischemia;
[0050] 2) New ischemic ECG changes;
[0051] 3) Pathological Q waves appear in the electrocardiogram;
[0052] 4) Imaging evidence shows new loss of myocardial activity or new local wall motion abnormality;
[0053] 5) Coronary angiography or autopsy fou...
Embodiment 2
[0055] Example 2 Sequencing, data analysis and electronic verification
[0056] Sequencing: miRNA and mRNA were sequenced using llumina Hiseq2500 / Miseq second-generation high-throughput sequencing technology, and data processing was completed by removing adapters, removing low-quality, and decontaminating processes to obtain final data. The miRNA and mRNA sequencing raw data were background-corrected by the transcriptome data analysis software, and then the t-test was performed to obtain the P value, and then the Fisher test was used to combine the P values to screen the differentially expressed miRNA and mRNA. Use algorithms including miRanda, miRDB, miRWalk and Targetscan to predict the target genes of differentially expressed miRNAs, select ≥ 4 target genes predicted by the algorithms, and search for verified target genes of differentially expressed miRNAs in the miRWalk database.
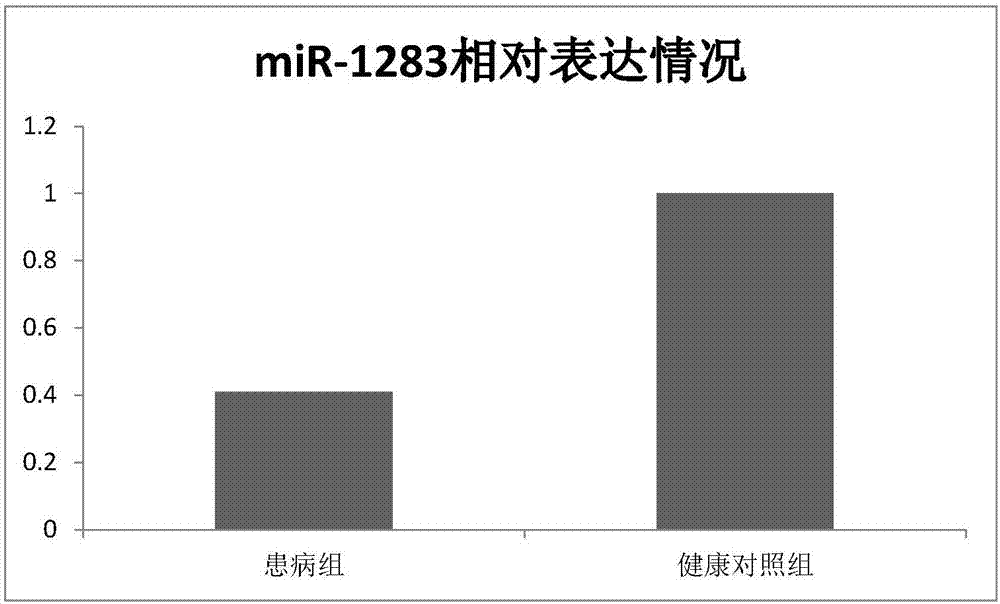
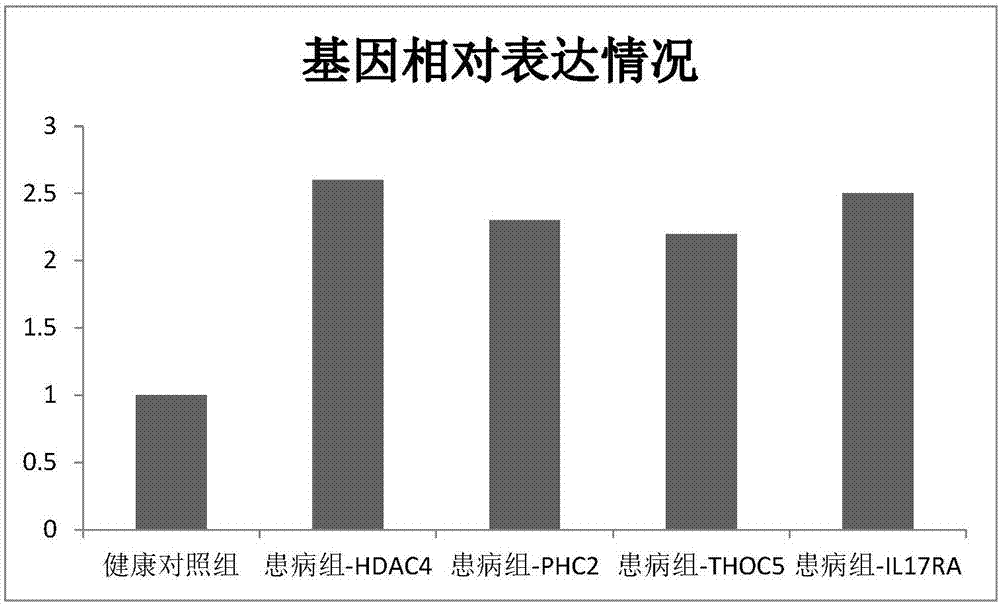
[0057] Finally, miR-1283 and its target genes HDAC4, PHC2, THOC5, and IL17RA were selected...
Embodiment 3
[0058] Embodiment 3 Electronic Verification
[0059] Electronic verification: 3 sets of mRNA data sets (GSE48060, GSE34198 and GSE61145-GPL6884) and 2 sets of miRNA data sets (GSE61741, GSE31568) were screened from the GEO (Gene Expression Omnibus) database, and 3 sets of mRNA data sets (GSE48060, GSE34198 and GSE61145 -GPL6884) has a total of 13,680 genes. We calculated and merged the effect value by using the metaMA package and using the combined P-value method to obtain 612 (FDR1) were found. The results showed that the electronic verification results were consistent with the expression trend of the sequencing results.
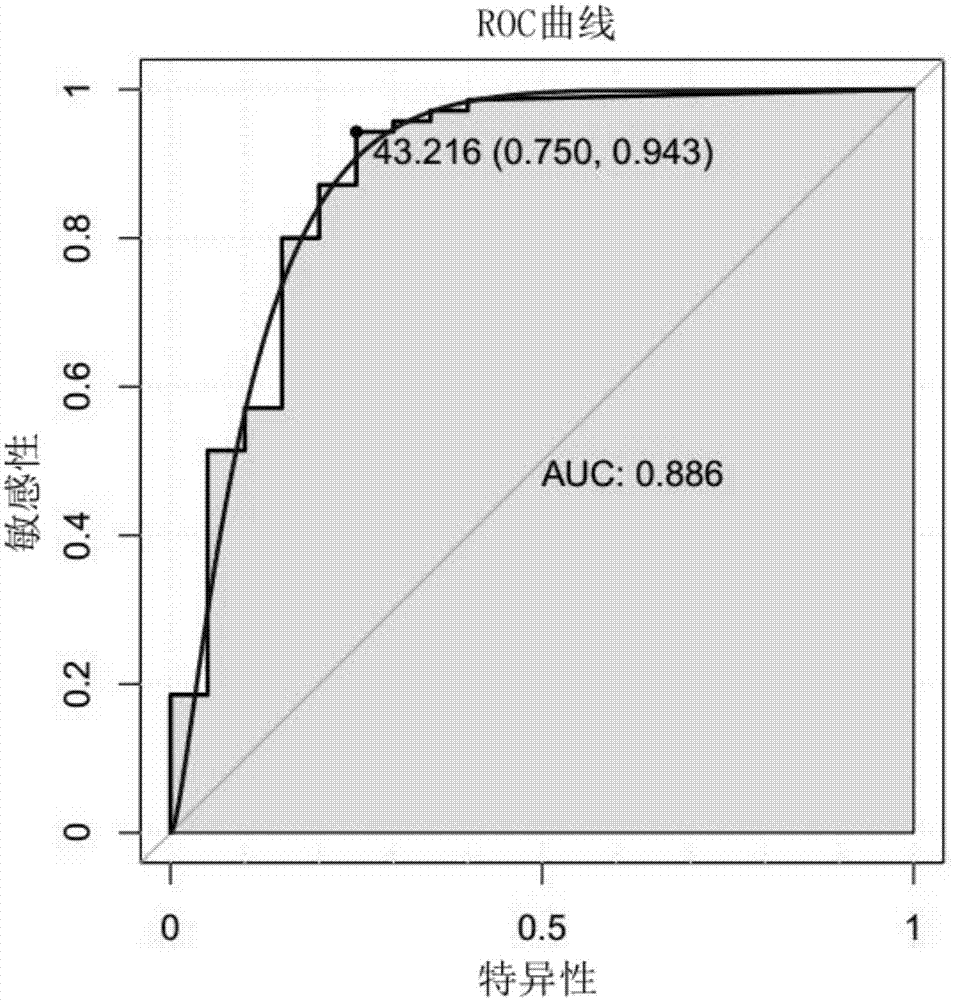
[0060] The method for evaluating the efficacy of a single miRNA molecule or a diagnostic model is to establish a receiver operating characteristic (ROC) curve, and judge the ability of diagnosis by calculating the area under the curve (Area UnderCurve). The value of the area under the ROC curve is between 1.0 and 0.5. In the case of AUC>0.5, the closer the...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More