Plant chromosome compositions and methods
a technology of chromosome composition and composition, applied in the field of molecular biology, can solve the problems of achieve the effect of severe limiting the use of genetic analysis of plants
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Benefits of technology
Problems solved by technology
Method used
Image
Examples
example 1
Generation of an Arabidopsis thaliana Mapping Population
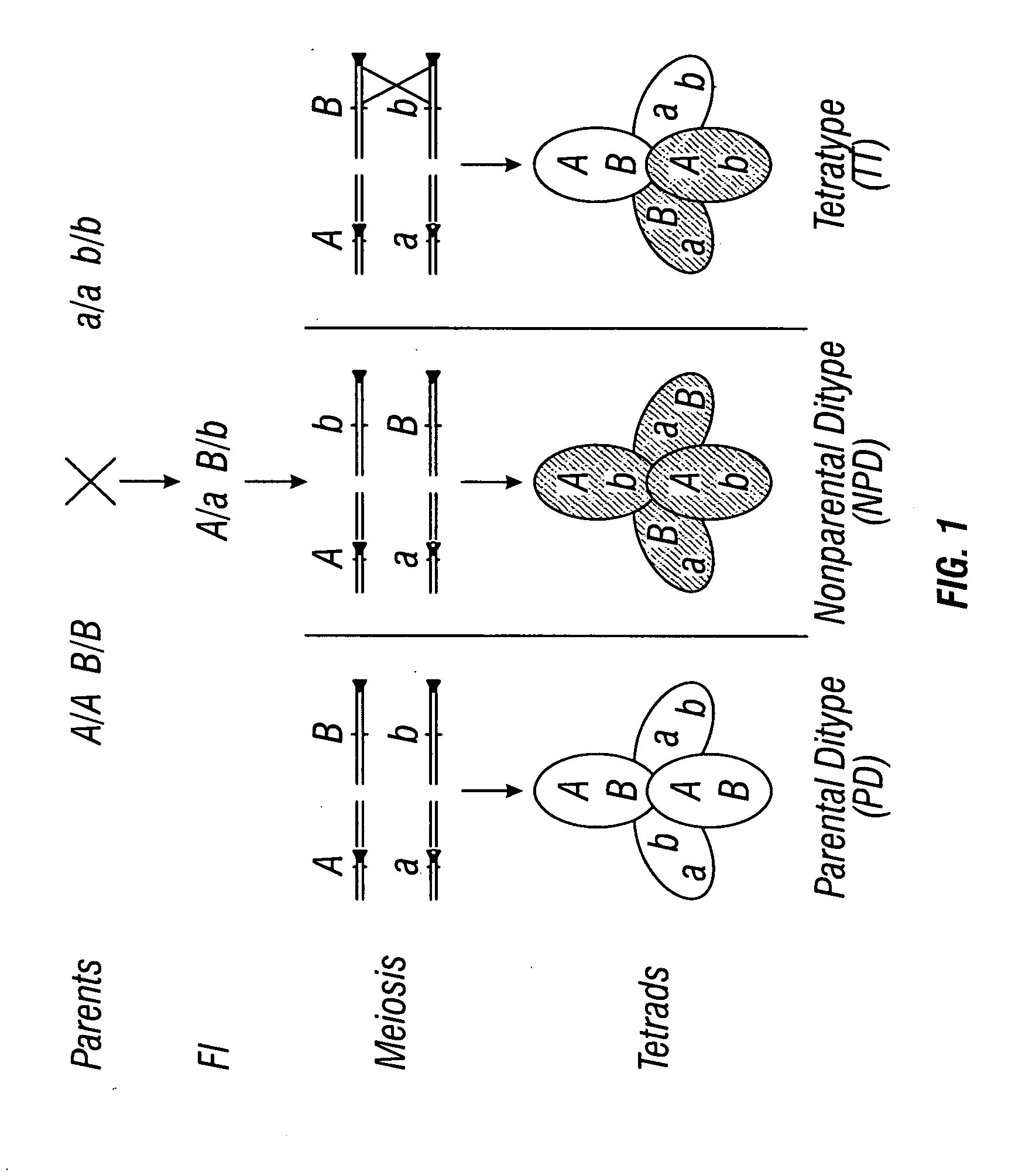
[0350] To generate a pollen donor plant, two parental lines carrying qrtl were crossed to one another. The qrtl-1 allele was in the Landsberg ecotype background and the qrtl-2 allele was in the Columbia ecotype background. The Landsberg ecotype was readily discernible from the Columbia ecotype because it carries a recessive mutation, erecta, which causes the stems to thicken, infloresences to be more compact, and the leaves to be more rounded and small than wildtype. To utilize this as a marker of a donor plant, qrtl-2 pollen was crossed onto a qrtl-1 female stigma. The F1 progeny were heterozygous at all molecular markers yet the progeny retain the quartet phenotype of a tetrad of fused pollen grains. In addition, progeny display the ERECTA phenotype of the Columbia plant. This visible marker serves as an indication that the crossing was successful in generating plants segregating ecotype specific markers. Further testing was...
example 2
Tetrad Pollinations
[0352] Tetrad pollinations were carried out as follows. A mature flower was removed from the donor plant and tapped upon a glass microscope slide to release mature tetrad pollen grains. This slide was then placed under a 20-40× Zeiss dissecting microscope. To isolate individual tetrad pollen grains, a small wooden dowel was used to which an eyebrow hair with rubber cement was mounted. Using the light microscope, a tetrad pollen unit was chosen and touched to the eyebrow hair. The tetrad preferentially adhered to the eyebrow hair and was thus lifted from the microscope slide and transported the recipient plant stigmatic surface. The transfer was carried out without the use of the microscope, and the eyebrow hair with adhering tetrad was then placed against the recipient stigmatic surface and the hair was manually dragged across the stigma surface. The tetrad then preferentially adhered to the stigma of the recipient and the cross pollination was completed.
[0353] ...
example 3
Preparation and Analysis of Centromere-Spanning Contigs
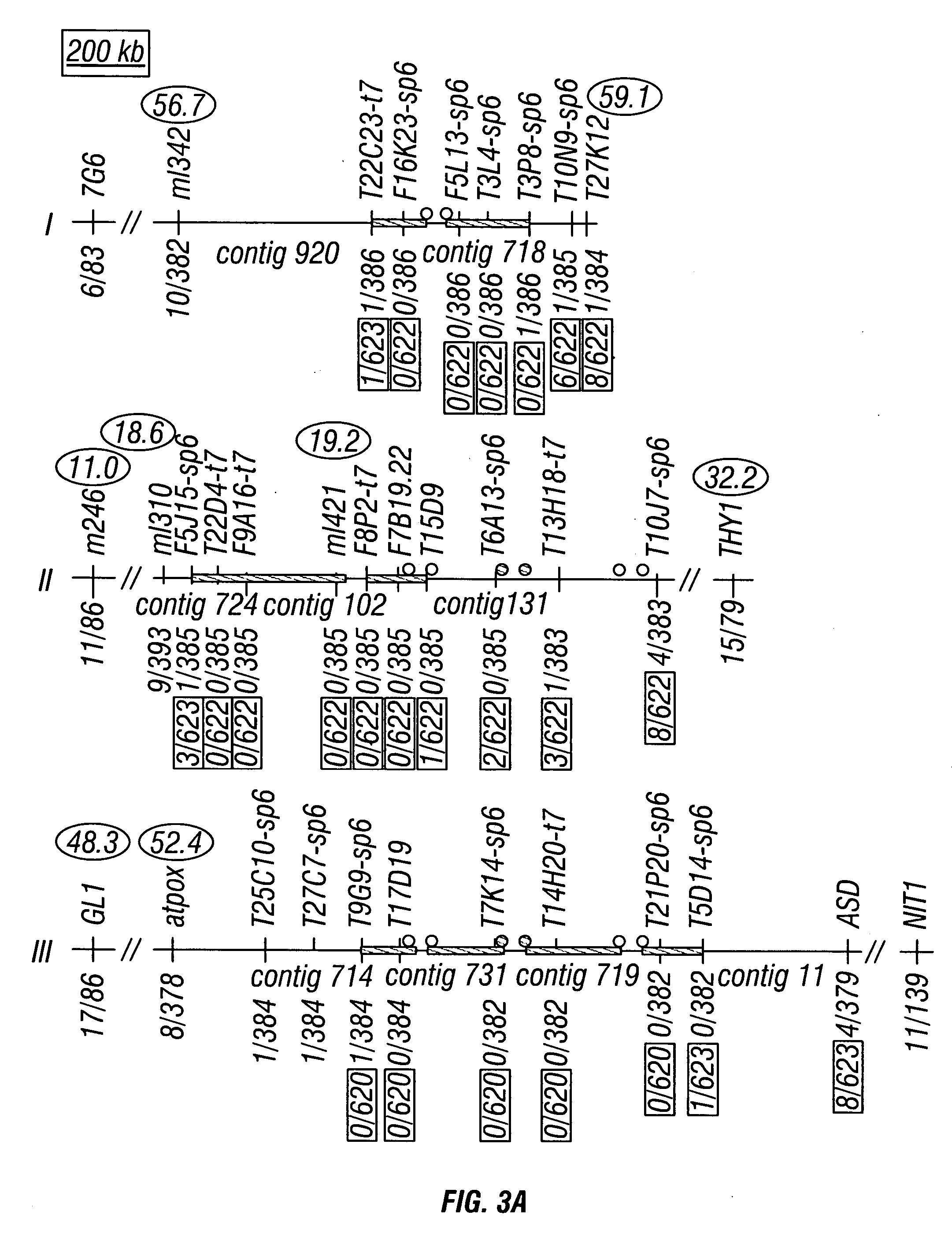
[0354] Previously, DNA fingerprint and hybridization analysis of two bacterial artificial chromosome (BAC) libraries led to the assembly of physical maps covering nearly all single-copy portions of the Arabidopsis genome (Marra et al., 1999). However, the presence of repetitive DNA near the Arabidopsis centromeres, including 180 bp repeats, retroelements, and middle repetitive sequences complicated efforts to anchor centromeric BAC contigs to particular chromosomes (Murata et al., 1997; Heslop-Harrison et al., 1999; Brandes et al., 1997; Franz et al., 1998; Wright et al., 1996; Konieczny et al., 1991; Pelissier et al, 1995; Voytas and Ausubel, 1988; Chye et al., 1997; Tsay et al., 1993; Richards et al., 1991; Simoens et al., 1988; Thompson et al., 1996; Pelissier et al., 1996). The inventors used genetic mapping to unambiguously assign these unanchored contigs to ±20 specific centromeres, scoring polymorphic markers in 48 plant...
PUM
| Property | Measurement | Unit |
|---|---|---|
| diameter | aaaaa | aaaaa |
| temperatures | aaaaa | aaaaa |
| temperatures | aaaaa | aaaaa |
Abstract
Description
Claims
Application Information
 Login to View More
Login to View More