Methods and systems for identifying crispr/cas off-target sites
a technology of off-target sites and methods, applied in the field of bioinformatics, can solve the problems that none of these bioinformatics search tools has considered off-target sites, nor provide application-specific primers, etc., and achieve the effects of improving the speed of search, facilitating experimental confirmation of off-target activity, and increasing run times
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Benefits of technology
Problems solved by technology
Method used
Image
Examples
example
Example 1
CRISPR Guide Strands can Exhibit Off-Target Activity at Similar Levels as On-Target Activity, Even with Mismatches within First 12 Nucleotides
Materials and Methods
[0324]CRISPR Design and Testing
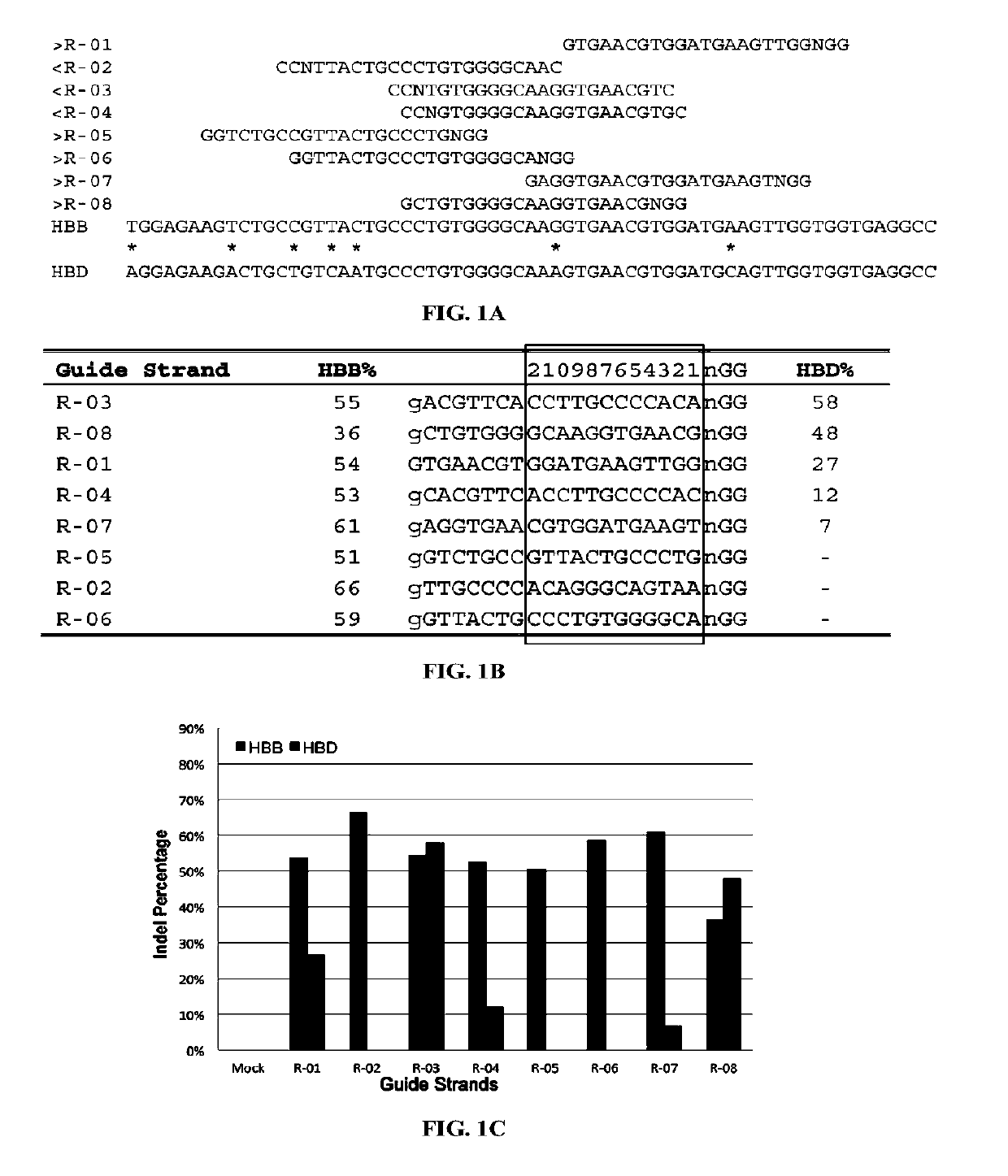
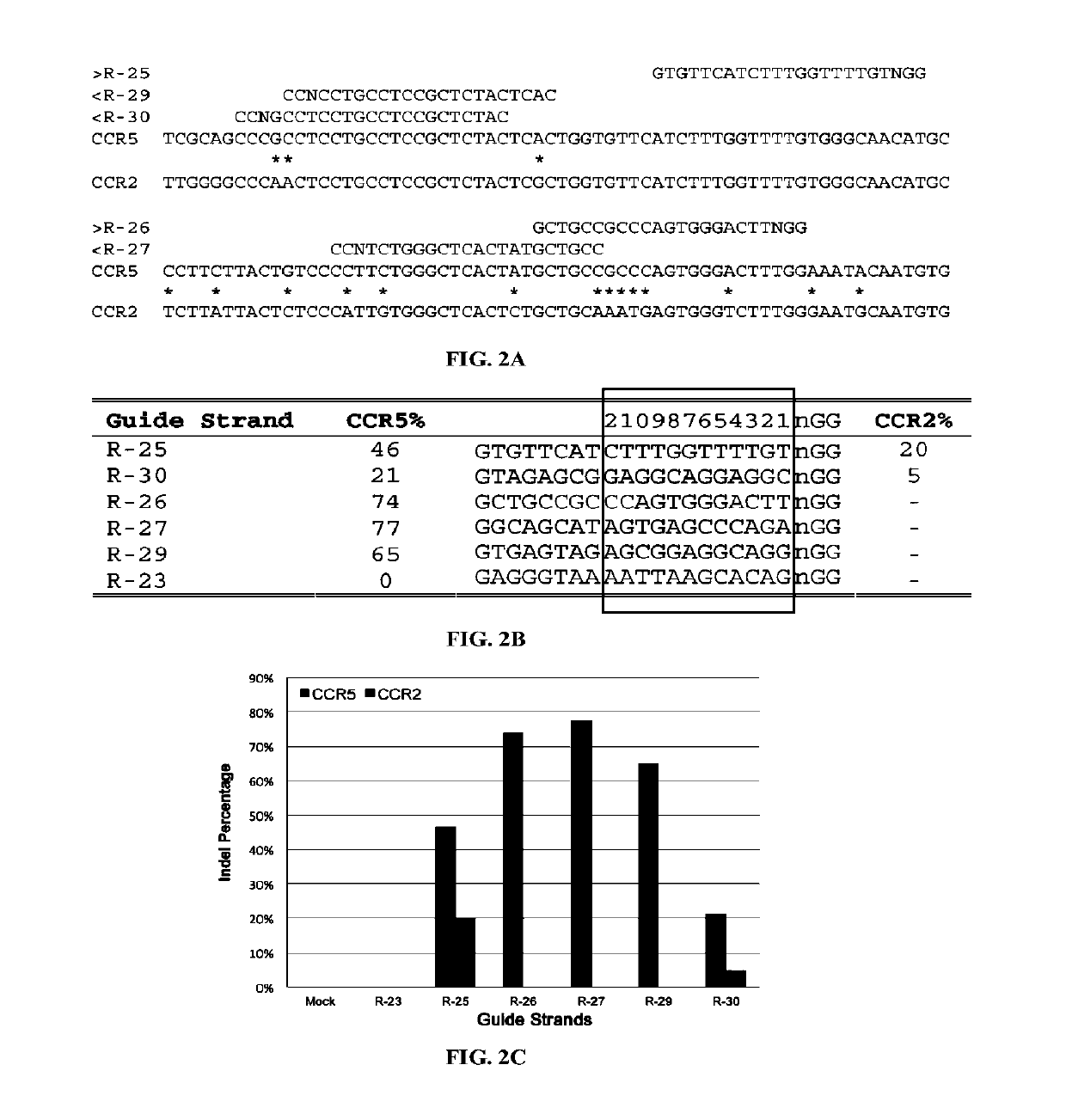
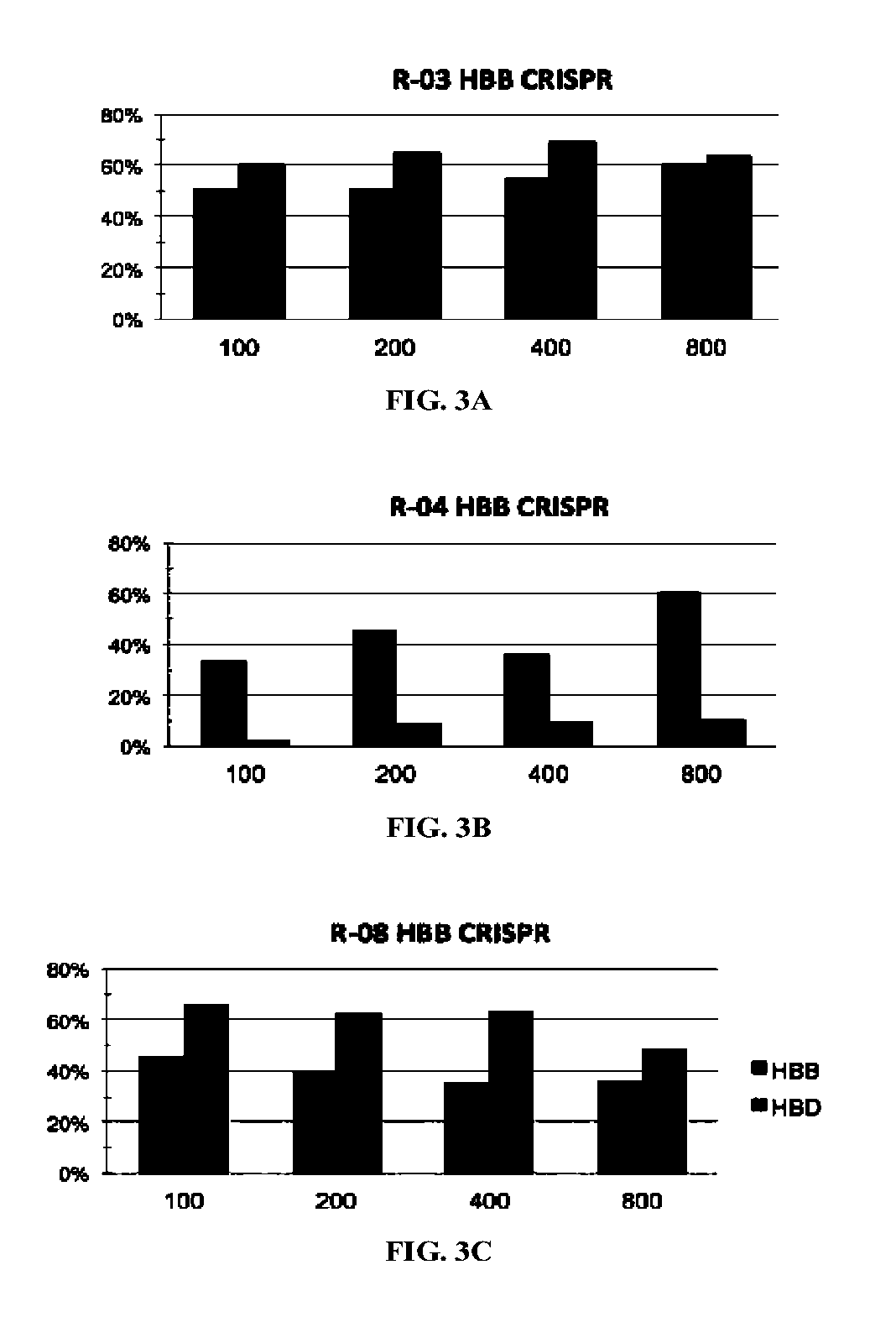
[0325]There were no CRISPR target sites in the human HBB gene sequence with their proximal 12 bases unique in the human genome (Cong, et al., Science, 339:819-823 (2013)); therefore, CRISPR / Cas9 guide strands targeting HBB were chosen by comparing the similar regions in the human hemoglobin δ (HBD) gene. Eight 20-base guide strands were designed to target sites near the sickle mutation in the HBB gene (FIG. 1A), each adjacent to a PAM sequence that contains the canonical trinucleotide NGG. Five guide strands were also designed to target two segments in the human CCRS gene (FIG. 2A), and tested the corresponding CRISPR / Cas9 systems to determine their on-target cleavage and potential off-target activity at the human C-C chemokine receptor type 2 (CCR2) gene. Herein the name of the guid...
example 2
CRISPR-Targeted Loci Showed a Wide Variety of Insertions, Deletions and Point Mutations
Materials and Methods
[0339]Chromosomal Deletion Analysis
[0340]To assay for gross chromosomal deletions, genomic DNA from cells transfected with R-03 was amplified using the HBD forward primer and the reverse primer downstream of the HBB site. Genomic DNA from cells transfected with R-25 or R-30 were similarly amplified using the CCR2 forward and the CCRS reverse primers. Agarose gels were used to confirm that the polymerase chain reaction (PCR) product sizes were consistent with chromosomal deletions between these sites. The R-03, R-25 and R-30 PCR products were cloned and the individual colonies Sanger sequenced and aligned.
[0341]Quantitative PCR
[0342]Quantitative PCR determination of the percentage of HBD-HBB chromosomal deletions. HEK-293 cells were transfected in triplicate with CRISPR plasmids containing guide strands R-02 or R-03, or mock transfected cells. Genomic DNA was harvested using Qu...
example 3
sgRNA Variants Containing Single-Base DNA Bulges Induce Cas9 Cleavage
Materials and Methods
[0351]CRISPR / Cas9 Plasmid Assembly
[0352]DNA oligonucleotides containing a G followed by a 19-nt guide sequence (Table 3) were kinased, annealed to create sticky ends and ligated into the pX330 plasmid that contains the +85 chimeric RNA under the U6 promoter and a Cas9 expression cassette under the CBh promoter (available at Addgene) (Hsu, et al., Nat Biotechnol, 31 (2013)).
TABLE 4Protospacer target sites for the sgRNAs used inExamples 3-8 (Table 4 discloses SEQ ID NOS 35-61,respectively, in order of appearance)StorageGeneIndexProtospacer Target (5′ to 3′) PAMHBBR-01GTGAACGTGGATGAAGTTGG TGGHBBR-03GACGTTCACCTTGCCCCACA GGGHBBR-04GCACGTTCACCTTGCCCCAC AGGHBBR-05GGTCTGCCGTTACTGCCCTG TGGHBBR-06GGTTACTGCCCTGTGGGGCA AGGHBBR-07GAGGTGAACGTGGATGAAGT TGGHBBR-08GCTGTGGGGCAAGGTGAACG TGGEGFPR-19GGTGGTGCAGATGAACTTCA GGGEGFPR-20GACCAGGATGGGCACCACCC CGGCCR5R-25GTGTTCATCTTTGGTTTTGT GGGCCR5R-26GCTGCCGCCCAGTGGGACTT ...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More