Directed evolution of microorganisms
a technology of microorganisms and evolution, applied in the direction of microorganisms, peptides, bacteria based processes, etc., can solve the problems of limiting the use of microorganisms in industrial biotechnology, toxicity of solvents, and general toxicity of organic solvents, so as to reduce reduce the burden on cells , the effect of reducing the expression of mutator genes
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Benefits of technology
Problems solved by technology
Method used
Image
Examples
example 1
Construction of mutD and mutD5′ Plasmids and Testing in 3 Bacterial Strains
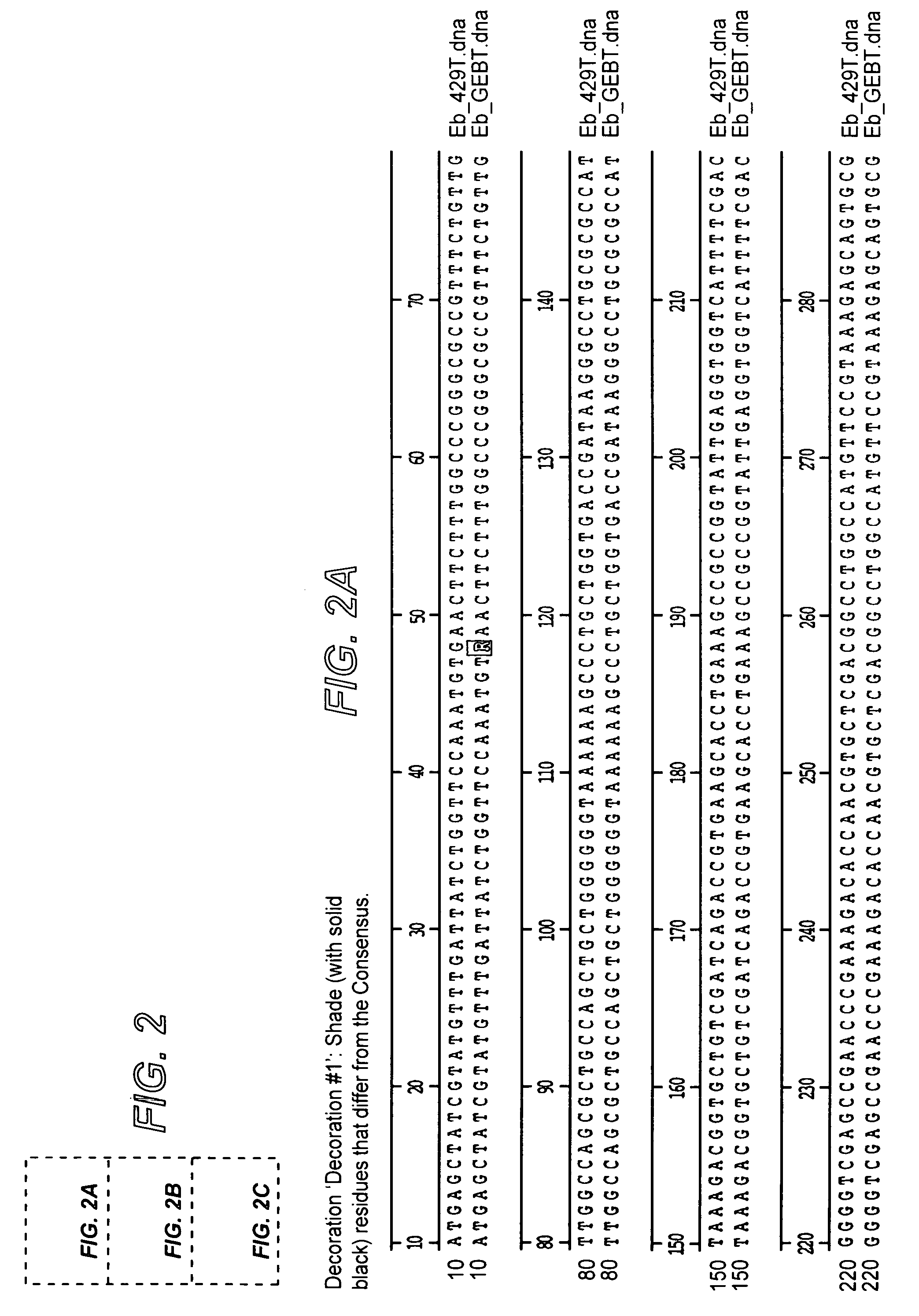
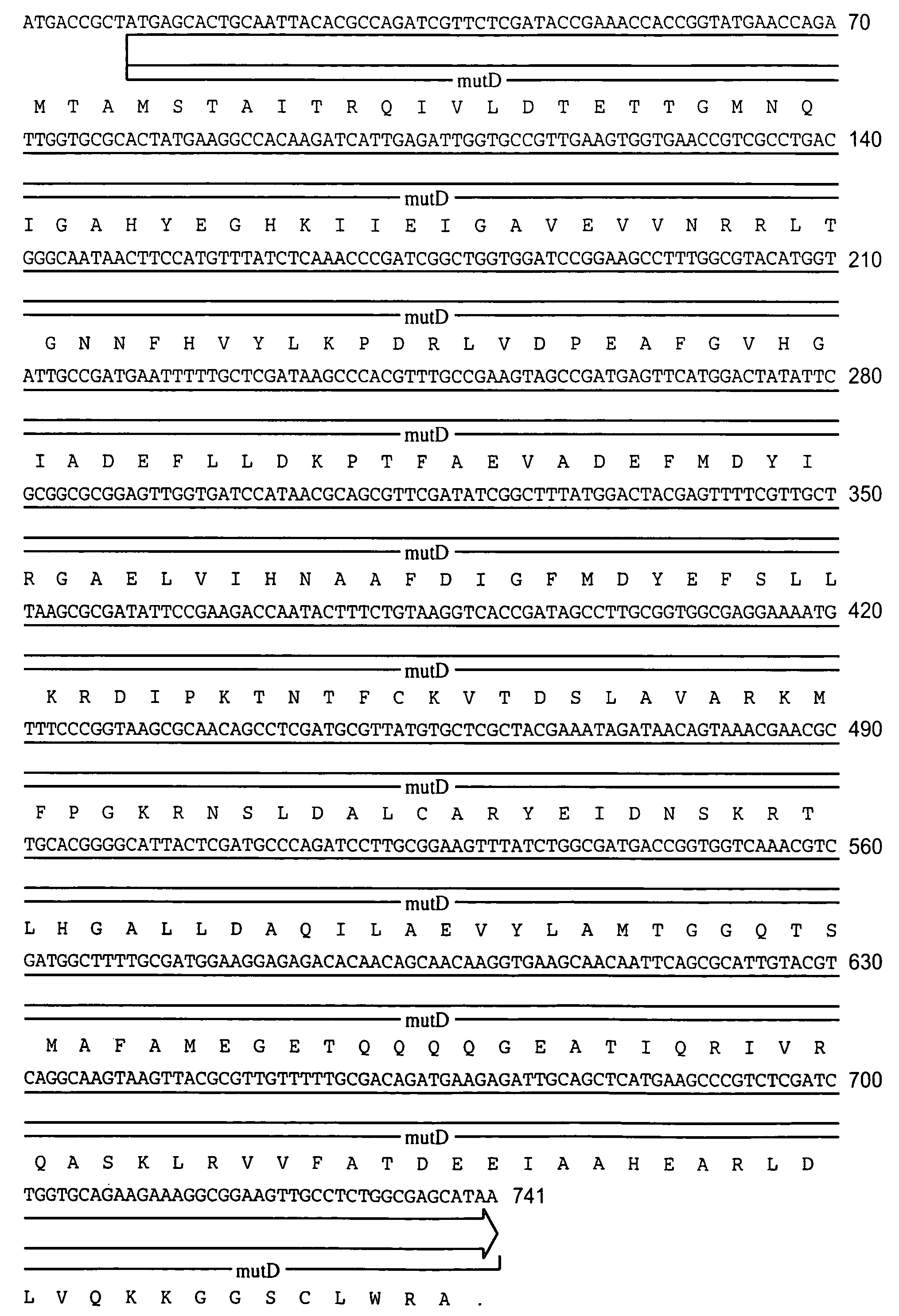
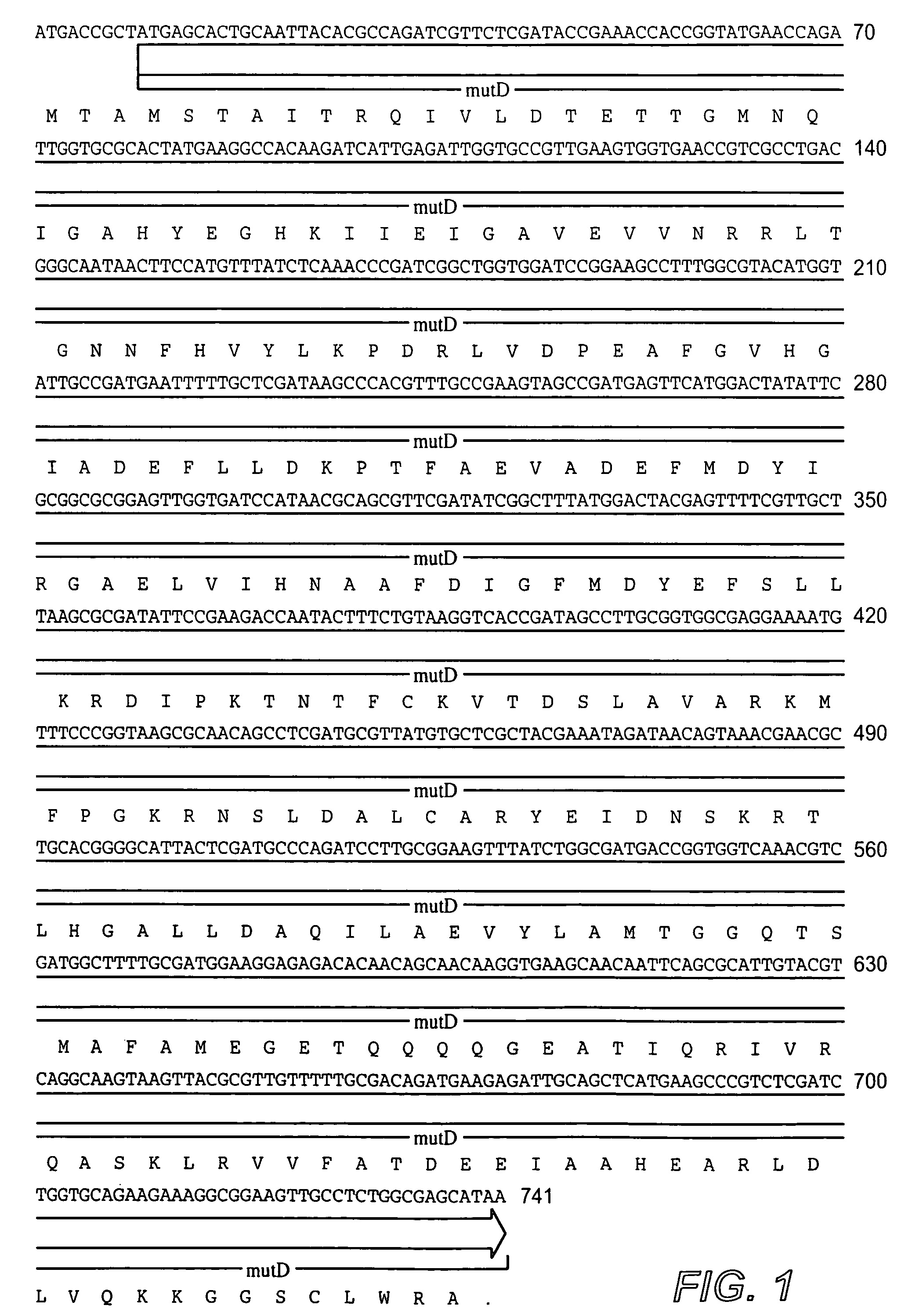
[0055]The following example illustrates the construction of plasmids comprising the mutator gene, mutD5′.
[0056]mutD and mutD5′ genes were amplified by PCR using mutd1 (5′-CGCCTCCAGCGCGACAATAGCGGCCATC-3′) and mutd2 (5′-CCGACTGAACTACCGCTCCGCGTTGTG-3′) primers from genomic DNA of E. coli and E. coli CSH116 (Miller 1992), respectively. The PCR products were cloned into pCR-Blunt vector (Invitrogen, Carlsbad, Calif.). Plasmids from clones with the correct orientation were isolated and digested with SmaI-HindIII restriction enzymes. The overhang ends were filled using T4 polymerase and cloned into pMAK705 plasmid digested with SmaI-PvuII. The ligation products were transformed into JM101 competent cells. The resulted plasmids had the temperature-sensitive origin of replication, carried kanamycin resistance marker and were named pMutD-71 (control plasmid, wild type genotype) and pMutD5-61 (mutator plasmid).
[0057]The...
example 2
Evolution of Solvent Tolerance
[0058]The following example illustrates the evolution of solvent tolerant microorganisms using the mutator plasmids constructed as in Example 1.
[0059]In order to make evolution experiments quantitative, LB agar plates supplemented with 50, 60, 70, 80 and 90 g / l DMF and 25 ug / ml kanamycin were used. The size of every evolving population was limited to 106 cells. After each plating, the number of raised colonies was counted and 10 were selected for the next plating. Cells from selected colonies were mixed together and aliquots containing 106 cells were plated on fresh plates containing the same and higher concentrations of DMF. After 2 consequent platings the cells were cured of the plasmids by growth at elevated temperatures. E. blattae 33429 and E. coli MM294 were cured at 41° C. and 43° C., respectively. Three to four subculturings at indicated temperatures were sufficient for 87-100% curing. Individual cured clones were selected by parallel growth of ...
example 3
Evolution of High Temperature Strains
[0066]Example 3 illustrates high temperature evolution under conditions of continuous fermentation in the mode of turbidostat which allows for fermentation wherein the cell density is stabilized. Two independent experiments were run with the strains: A: W1485 (ATCC12435) (=non mutator); B: W1485 / pBRmutD68 (same strain but comprises mutator plasmid). Both strains were gown in continuous culture in a turbidostat in LB medium for about 1800 doublings. The temperature was controlled by a computer based on the measured growth rate to maintain a doubling time of about 1 h. Whenever the culture grew faster the temperature was raised and vice versa. The time course of both cultures is shown in FIG. 4. Initially, the culture started from W1485 / pBRmutD68 evolved faster than the culture started from W1485. This indicates the advantage of the mutator plasmid. However, After about 400 doublings W1485 reached the higher temperature. We also measured the mutati...
PUM
| Property | Measurement | Unit |
|---|---|---|
| temperature | aaaaa | aaaaa |
| temperatures | aaaaa | aaaaa |
| concentration | aaaaa | aaaaa |
Abstract
Description
Claims
Application Information
 Login to View More
Login to View More