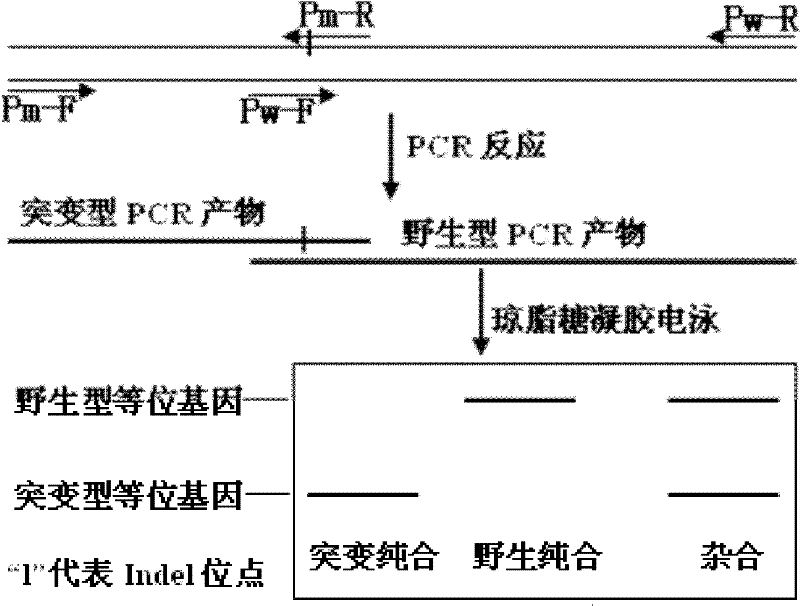
Method for detecting single base Indel mutation
A single-base, base-based technology, applied in the field of detection of single-base Indel mutations, can solve the problems of false negatives, long experimental process, cumbersome operation, etc., and achieve the effect of preventing false positives and improving specificity
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0054] Example 1 Detection of Indel polymorphism of goat weaver gene 5'UTR
[0055] 1) There is a base "A" Indel polymorphism in the 5'UTR of the goat weaver gene as shown in SEQ ID No.1 at position 118;
[0056] Design primer pair Pw as:
[0057] The wild-type chain was used as a template to design the forward primer at position 1 at the 3'end of the forward primer which is the same as the base at position 3 downstream of the Indel allele locus, that is: "C at position 121 of SEQ ID No. 1 in the sequence listing. "Base, and then the same DNA sequence upstream of this base is the forward primer. The length of the primer depends on the annealing temperature and GC content of the primer. The reverse primer is designed with an interval of 348 bp. The specific primer pair Pw is:
[0058] Forward primer Pw-F: ctacgttatc tccgggcaaa gcc 23;
[0059] Reverse primer Pw-R: tgggcacgaa tgcaaatac 19;
[0060] The amplified fragment length of this primer pair is 367bp.
[0061] Design primer pair and ...
Embodiment 2
[0080] Example 2 Indel diagnostic application in detection of polymorphism in different goat populations
[0081] 1. Indel diagnosis in population polymorphism
[0082] Three goat breeds were tested using the method described in Example 1. Among them, Xinong Saanen dairy goats (268 DNA samples) and Guanzhong dairy goats (440 DNA samples) belong to the dairy type; Xinjiang cashmere goats are the cashmere type (119 DNA samples).
[0083] 2. Statistical analysis of frequency of Indel sites
[0084] Genotype frequency refers to the ratio of the number of individuals with a certain genotype for a certain trait to the total number of individuals in a population. PAA=NAA / N, where PAA represents the frequency of AA genotype at a certain site; NAA represents the number of individuals with AA genotype in the population; N is the total number of the tested population.
[0085] Gene frequency refers to the relative ratio of the number of a gene to the total number of alleles in a population. The...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More