Methods and compositions for selecting siRNA of improved functionality
a sirna and functionality technology, applied in the field of rna interference, can solve the problems of inability to induce rnai with limited success, inhibiting protein synthesis, and long dsrna being not a viable option for rnai in mammalian systems, and achieve the effect of increasing the efficiency of rnai and increasing the efficacy of sirna
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Benefits of technology
Problems solved by technology
Method used
Image
Examples
examples
General Techniques and Nomenclatures
[0418] siRNA nomenclature. All siRNA duplexes are referred to by sense strand. The first nucleotide of the 5′-end of the sense strand is position 1, which corresponds to position 19 of the antisense strand for a 19-mer. In most cases, to compare results from different experiments, silencing was determined by measuring specific transcript mRNA levels or enzymatic activity associated with specific transcript levels, 24 hours post-transfection, with siRNA concentrations held constant at 100 nM. For all experiments, unless otherwise specified, transfection efficiency was ensured to be over 95%, and no detectable cellular toxicity was observed. The following system of nomenclature was used to compare and report siRNA-silencing functionality: “F” followed by the degree of minimal knockdown. For example, F50 signifies at least 50% knockdown, F80 means at least 80%, and so forth. For this study, all sub-F50 siRNAs were considered non-functional.
[0419] ...
example i
Sequences Used to Develop the Algorithm
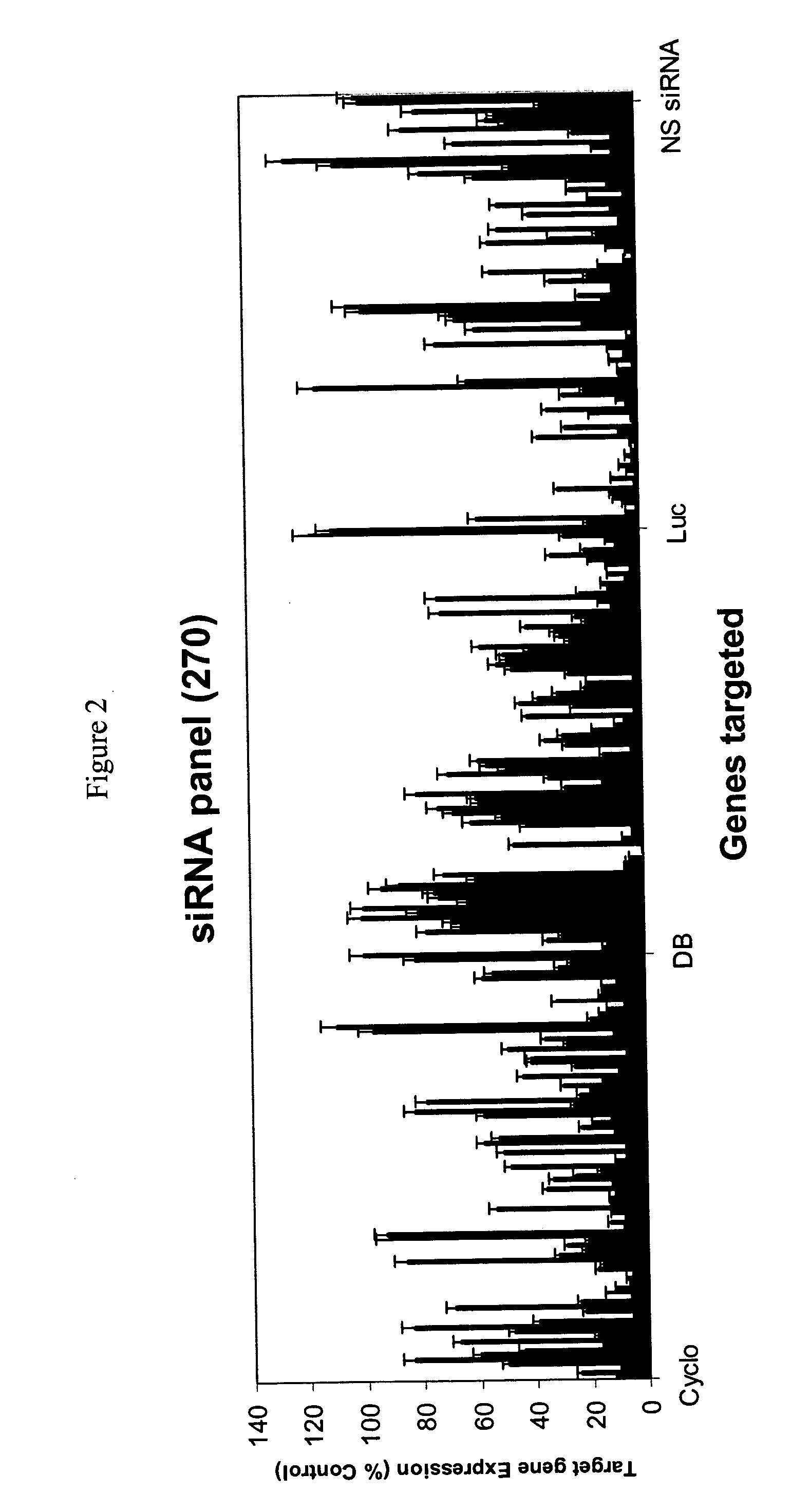
[0421] Anti-Firefly and anti-Cyclophilin siRNAs panels (FIG. 5a, b) sorted according to using Formula VIII predicted values. All siRNAs scoring more than 0 (formula VIII) and more then 20 (formula IX) are fully functional. All ninety sequences for each gene (and DBI) appear below in Table III.
TABLE IIICyclo1SEQ. ID 0032GUUCCAAAAACAGUGGAUACyclo2SEQ. ID 0033UCCAAAAACAGUGGAUAAUCyclo3SEQ. ID 0034CAAAAACAGUGGAUAAUUUCyclo4SEQ. ID 0035AAAACAGUGGAUAAUUUUGCyclo5SEQ. ID 0036AACAGUGGAUAAUUUUGUGCyclo6SEQ. ID 0037CAGUGGAUAAUUUUGUGGCCyclo7SEQ. ID 0038GUGGAUAAUUUUGUGGCCUCyclo8SEQ. ID 0039GGAUAAUUUUGUGGCCUUACyclo9SEQ. ID 0040AUAAUUUUGUGGCCUUAGCCyclo10SEQ. ID 0041AAUUUUGUGGCCUUAGCUACyclo11SEQ. ID 0042UUUUGUGGCCUUAGCUACACyclo12SEQ. ID 0043UUGUGGCCUUAGCUACAGGCyclo13SEQ. ID 0044GUGGCCUUAGCUACAGGAGCyclo14SEQ. ID 0045GGCCUUAGCUACAGGAGAGCyclo15SEQ. ID 0046CCUUAGCUACAGGAGAGAACyclo16SEQ. ID 0047UUAGCUACAGGAGAGAAAGCyclo17SEQ. ID 0048AGCUACAGGAGAGAAAGGACyclo18SEQ. ID ...
example ii
Validation of the Algorithm Using DBI, Luciferase, PLK, EGFR, and SEAP
[0422] The algorithm (Formula VIII) identified siRNAs for five genes, human DBI, firefly luciferase (fLuc), renilla luciferase (rLuc), human PLK, and human secreted alkaline phosphatase (SEAP). Four individual siRNAs were selected on the basis of their SMARTscores™ derived by analysis of their sequence using Formula VIII (all of the siRNAs would be selected with Formula IX as well) and analyzed for their ability to silence their targets' expression. In addition to the scoring, a BLAST search was conducted for each siRNA. To minimize the potential for off-target silencing effects, only those target sequences with more than three mismatches against un-related sequences were selected. Semizarov, et al, Specificity of short interfering RNA determined through gene expression signatures. Proc. Natl. Acad. Sci. U.S.A. 2003, 100:6347. These duplexes were analyzed individually and in pools of 4 and compared with several s...
PUM
| Property | Measurement | Unit |
|---|---|---|
| length | aaaaa | aaaaa |
| internal stability | aaaaa | aaaaa |
| cellular stress | aaaaa | aaaaa |
Abstract
Description
Claims
Application Information
 Login to View More
Login to View More