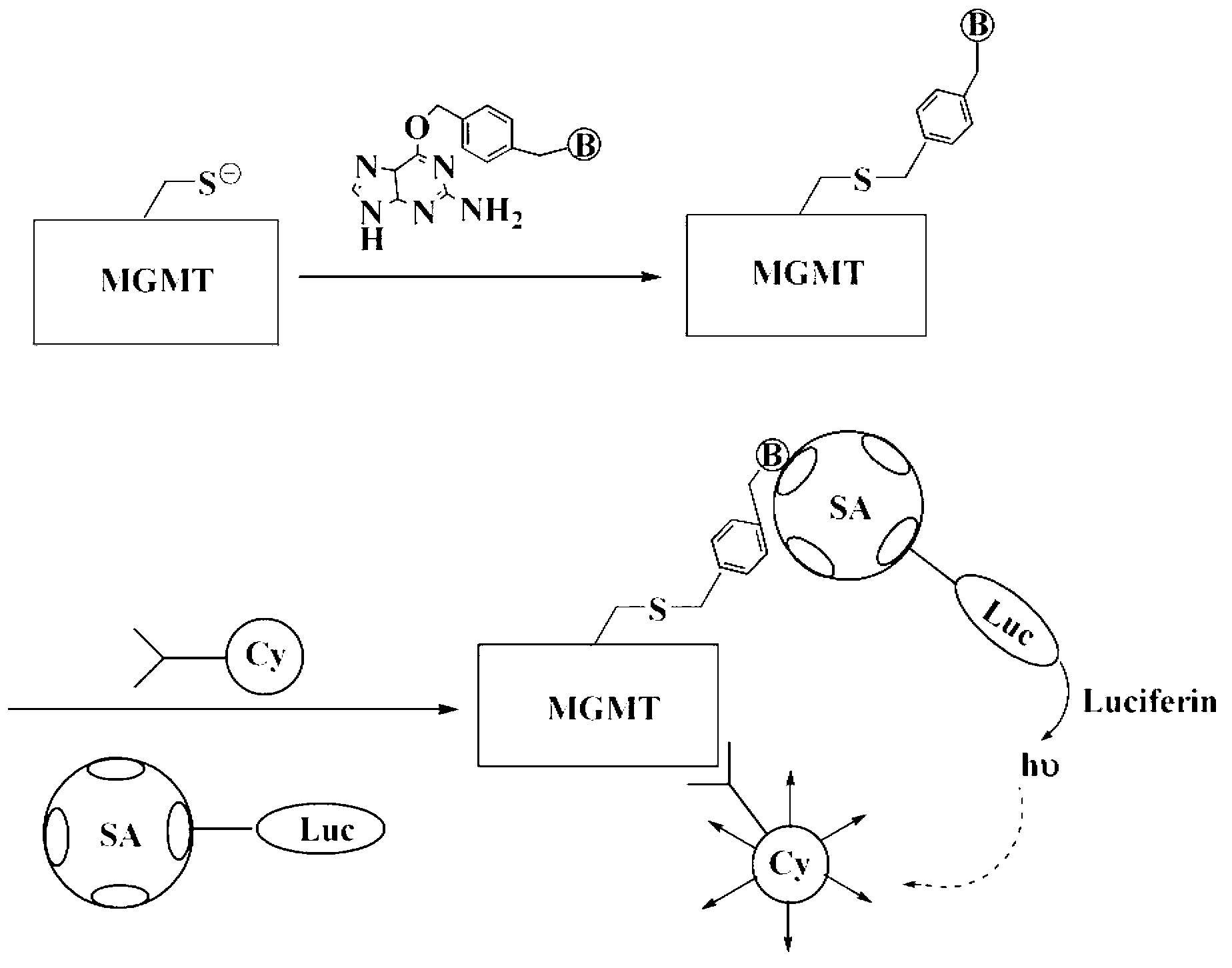
Method for detecting activity of O6-methylguanine-DNA (Deoxyribose Necleic Acid) methyltransferase
A technology of methylguanine and methyltransferase, used in measuring devices, instruments, scientific instruments, etc.
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0031] Example 1 O 6 -Biotin labeling of methylguanine-DNA methyltransferase (MGMT)
[0032] Add 1 ml of 2 mg of biotin-labeled O 6 - TBS buffer solution of methylguanine (pH 6.4), mix well and place in water bath at 37°C for 4 hours. After the reaction, add the reaction solution into a 14KD dialysis bag, and use TBS buffer solution with pH 6.4 as the dialysate Dialyze at 4°C, change the dialysate every 3 hours, and change three times in total, then filter the retentate with cross-linked agarose gel CL-6B, take the filtrate, freeze-dry at 4°C for 10 hours, and obtain biotin-labeled O 6 -Methylguanine-DNA methyltransferase.
Embodiment 2
[0033] Embodiment 2 detects MGMT activity
[0034] (1) Coating: The streptavidin-firefly luciferase fusion protein was prepared into 50ml of 1mg / mL coating solution with 0.1mol / L TBS buffer solution with pH 6.4. Add the coating solution to the sample wells of a 96-well white-bottomed polystyrene microwell plate, 200 μl per well, place at 37°C for 60 minutes, discard the coating solution, and add 200 μl of 10% bovine serum albumin (BSA) to each well ) above TBS buffer solution as blocking solution, blocked at 37°C for 1 hour, washed the plate with the above TBS buffer solution, soaked for 3 minutes each time, washed the plate three times, dried in vacuum at 30°C, put it into a sealed bag for vacuum storage, and prepared Microwell plates coated with streptavidin-firefly luciferase fusion protein;
[0035] (2) Adding samples: add the concentration of 1000, 500, 250 , 125, 62.5, 31.2, 15.6, 7.8, 3.4, 0ng / mL and unknown concentration (sample to be tested, taken from a tumor tissu...
Embodiment 3
[0040] Embodiment 3 detects MGMT activity
[0041] (1) Coating: The streptavidin-alkaline phosphatase fusion protein was prepared with 0.5mol / LTBS buffer solution with pH 6.4 to prepare 50ml of 3mg / mL coating solution. Add the coating solution to the sample wells of a 96-well white-bottomed polystyrene microwell plate, 200 μl per well, place at 25°C for 1 hour, discard the coating solution, and add 200 μl of 5% bovine serum albumin ( The above-mentioned TBS buffer solution of BSA) was used as the blocking solution, blocked at 37°C for 1 hour, washed with the above-mentioned TBS buffer solution, soaked for 3 minutes each time, washed the plate three times, dried in vacuum at 25°C, and stored in a sealed bag in vacuum to obtain streptavidin-alkaline phosphatase fusion protein;
[0042] (2) Adding samples: Add concentrations of 1000, 500, 250, 125, 62.5, 31.2, 15.6, 7.8, 3.4, 0 ng / mL and unknown concentration (sample to be tested, taken from a tumor tissue sample) 50 μl of TBS ...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More