Non-specific nucleases from the genus pseudomonas
A non-specific, nuclease technology, applied in enzyme stabilization, hydrolytic enzymes, instruments, etc., can solve problems such as increasing solution viscosity and clogging
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment approach
[0127] In one embodiment of the invention, a polypeptide having NSN activity disclosed herein is used to degrade and thereby remove nucleic acids from solution. These solutions may contain proteins, cells, tissues or other biological material. In one example, a polypeptide having NSN activity is used to degrade nucleic acids, such as DNA and / or RNA, present in crude cell extracts prepared for isolation and purification of recombinant or native proteins from host cells. Since nucleic acids contaminate the target protein, it is of interest to remove as much nucleic acid as possible. Cells including the protein of interest are lysed using a buffer containing a polypeptide having NSN activity (eg, pH 7-8 based phosphate buffer). The buffer may contain a cationic complexing agent. This solution is incubated at a defined temperature for a defined time (eg, at about 37°C for 10 min or at about 4°C for 2 hours). During incubation, nucleic acids are degraded. The protein is then is...
Embodiment 1
[0163] Example 1: Non-specific nucleic acids from Pseudomonas syringae and from phylogenetically related Pseudomonas Identification of the enzyme PsNuc
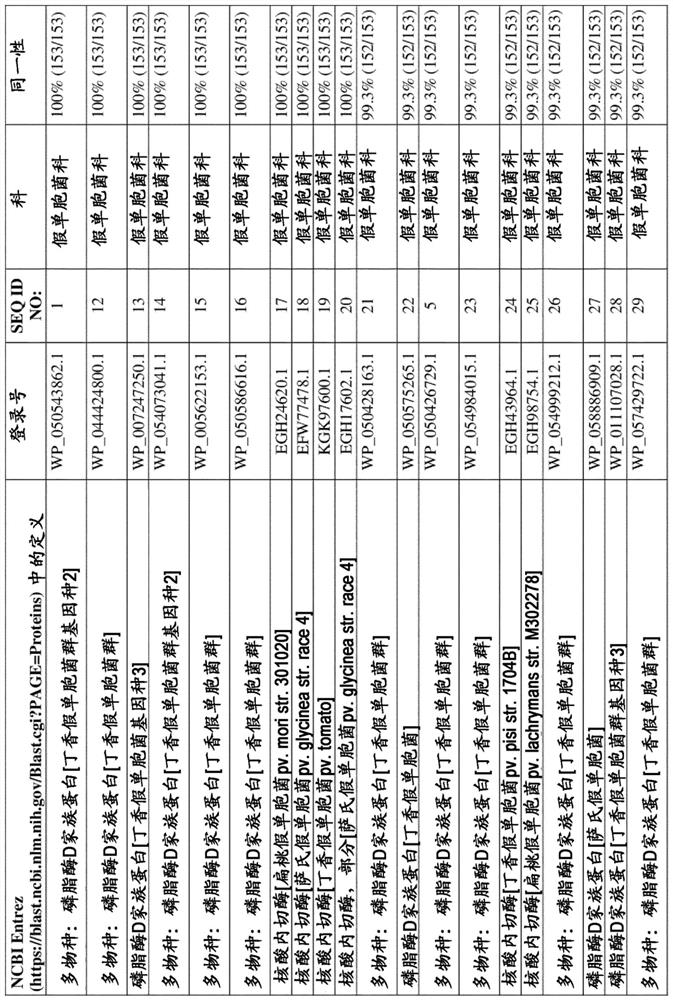
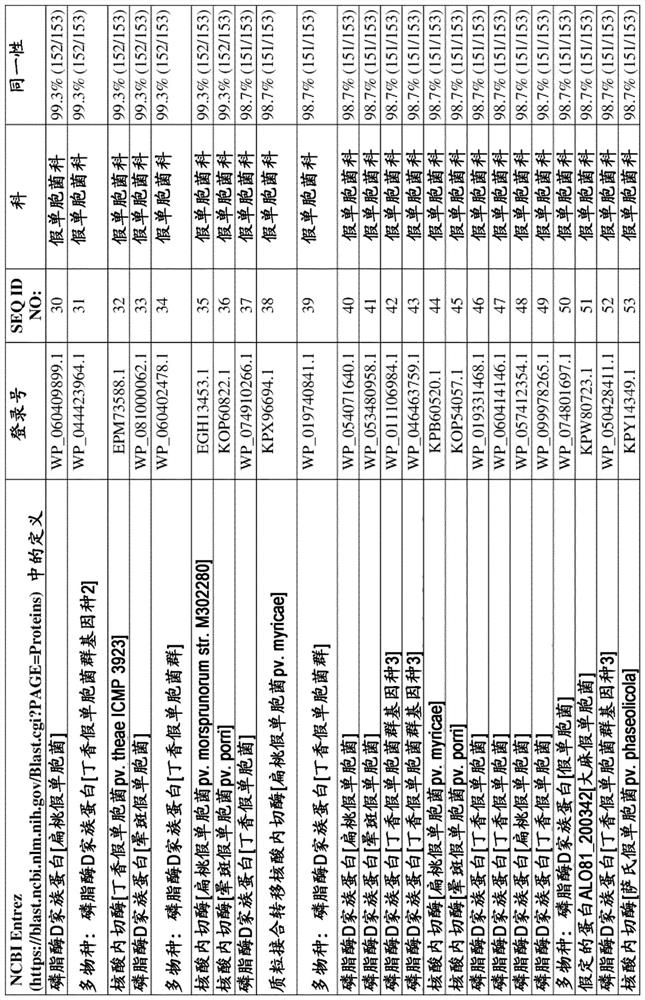
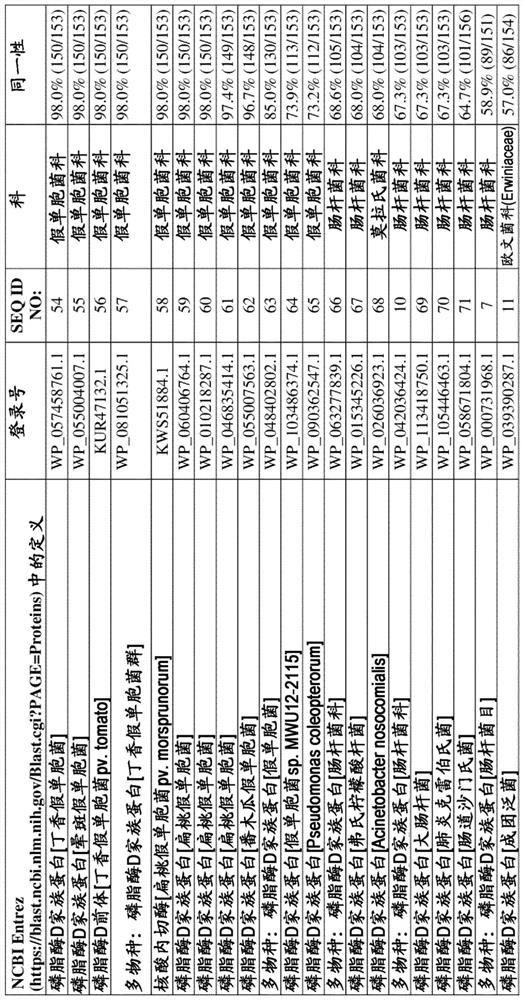
[0164] To identify putative non-specific bacterial endonucleases, BLAST analysis was performed using StNuc from Salmonella entericasubsp. enterica serovar Typhimurium as interrogator sequence (NCBI entry WP_000731968.1, SEQ ID NO:7). NCBI Entrez's BLASTP server (https: / / blast.ncbi.nlm.nih.gov / Blast.cgi?PAGE=Proteins) was used to screen putative endonucleases derived from cold-active bacterial species. A putative endonuclease from the PLD family of Pseudomonas syringae genomosp. 2 (WP_050543862.1, SEQ ID NO: 1 ) was retrieved which, in a local alignment omitting the putative secretion signal, was shown to be compatible with 57.2% amino acid sequence identity compared to StNuc (WP_000731968.1, SEQ ID NO:7). The gene encoding PsNuc consists of 552 nucleotides, including a stop (STOP) codon sequence, and the deduced amino ac...
Embodiment 2
[0165] Example 2: Cloning of genes encoding PsNuc and PsNuc variants and transformation of E. coli
[0166] The DNA sequences were derived from the mature amino acid sequences of PsNuc and its variants without predicted signal peptide sequences (SEQ ID NO:2, SEQ ID NO:3, SEQ ID NO:4, SEQ ID NO:5 and SEQ ID NO: 6) Derivation. The gene was synthesized as a codon-optimized sequence for expression in Escherichia coli (ATUM, Inc.) and cloned into the pET24d(+) vector (Merck Millipore) as a fusion protein containing an N-terminal affinity tag, which and tags allow affinity purification of recombinant PsNuc proteins ( image 3 ). E. coli DH5α cells (Thermo Fisher Scientific) were transformed with the corresponding pET24d:: / JvNuc plasmids by heat shock. Therefore, cells were thawed on ice for 10 min and supplemented with 10-100 ng of plasmid DNA. After an additional 20 min incubation on ice, a heat shock of 42 °C was applied for 45 sec. Thereafter, the cells were incubated on i...
PUM
| Property | Measurement | Unit |
|---|---|---|
| purity | aaaaa | aaaaa |
Abstract
Description
Claims
Application Information
 Login to View More
Login to View More