Polycyclic macrocyclic lactam compound as well as preparation method and application thereof
A technology of macrocyclic lactams and compounds, applied in the field of natural medicinal chemistry and microbiology, can solve the problems of weak cytotoxic activity, no report on the activity of inhibiting bacterial virulence protein secretion, and no anti-bacterial involvement, etc., and achieve the effect of strong inhibitory activity
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
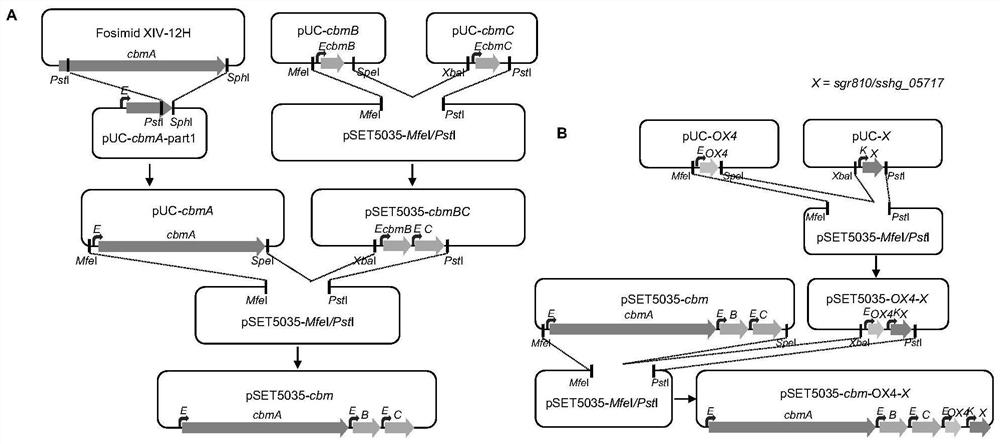
[0057] Construction of embodiment 1 plasmid pUC-cbmA
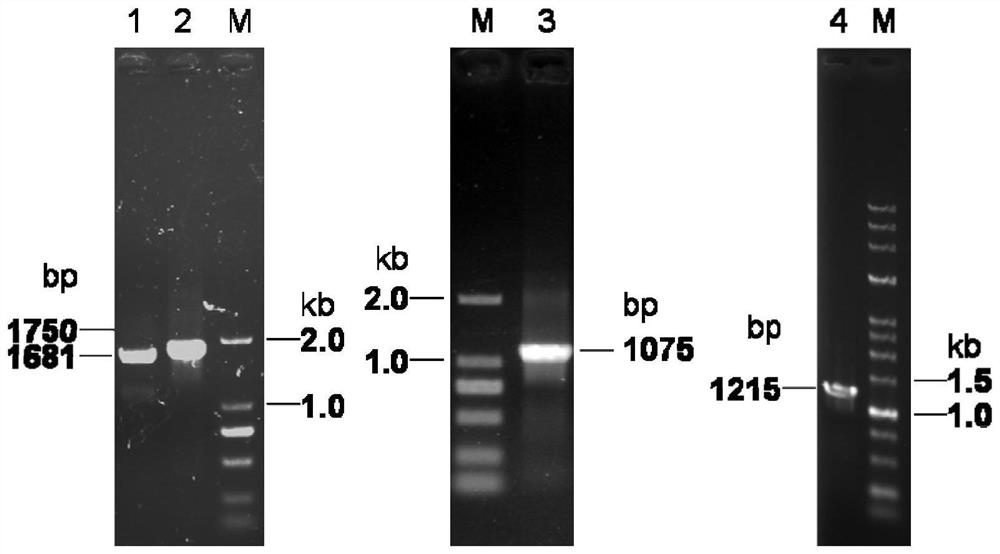
[0058] Using the fosmid XIV-12H containing the polyketide-nonribosomal peptide synthase gene cbmA as a template, the cbmA-part1 fragment was amplified by PCR, and the primers were designed using bioinformatics software to introduce NdeI and SpeI single enzyme cleavage sites at both ends of the sequence , the primer sequences are as follows:
[0059] NdeI-cbmA-LF:5'-cggacatatgtcggacctatgcaaggtc-3'
[0060] SpeI-cbmA-LR:5'-attactagtctacgccgcatgcgagattgatgtggggtggga-3'
[0061] The PCR amplification system was prepared according to the kit instructions;
[0062] PCR amplification conditions: pre-denaturation, 95°C 5min; denaturation, 95°C 30sec, annealing, 65°C 30sec, extension, 72°C 2min, 30 cycles; final extension, 72°C 5min.
[0063]The obtained PCR product was recovered by a gel recovery kit, and the PCR product and the vector plasmid pUC-kasOp* / ermEp* were digested with NdeI and SpeI respectively, and the obtained cbm...
Embodiment 2
[0064] Example 2 Construction of plasmids pUC-cbmB, pUC-cbmC, pUC-OX4, pUC-sgr810 and pUC-sshg_05717
[0065] Streptomyces sp.S10 genomic DNA was used as a template to amplify the cbmB fragment by PCR, and bioinformatics software was used to design primers to introduce NdeI and SpeI single enzyme cleavage sites at both ends of the sequence respectively. The primer sequences are as follows:
[0066] NdeI-cbmB-F:5'-cggacatatgcggcgtagtaacagggtg-3'
[0067] SpeI-cbmB-R:5'-attactagttcacctcatgctcttgtcg-3'
[0068] Streptomyces sp.S10 genomic DNA was used as a template to amplify the cbmC fragment by PCR, and bioinformatics software was used to design primers to introduce NdeI and SpeI single enzyme cleavage sites at both ends of the sequence respectively. The primer sequences are as follows:
[0069] NdeI-cbmC-F:5'-cggacatatgcctcgcaagccccg-3'
[0070] SpeI-cbmC-R:5'-attactagttcagcggacgtgaacgg-3'
[0071] The OX4 fragment was amplified by PCR using Lysobacter enzymogenes C3 genomi...
Embodiment 3
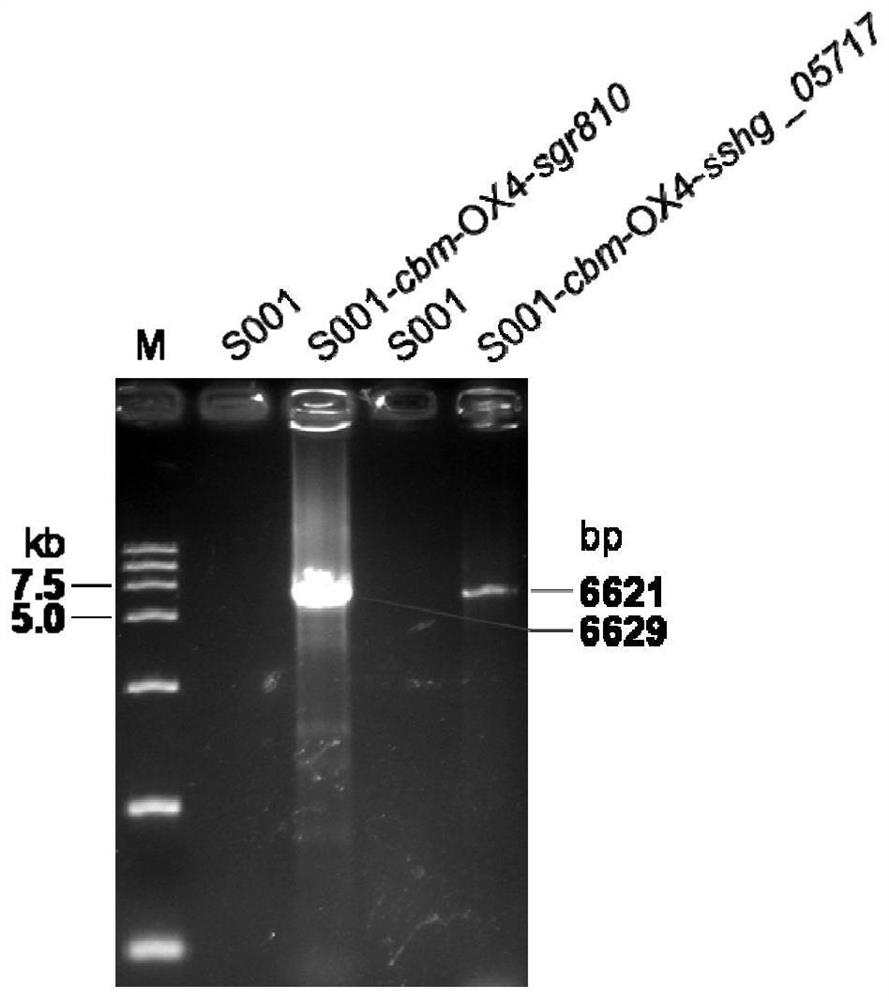
[0081] Example 3: Construction of expression vectors pSET5035-cbm-OX4-sgr810 and pSET5035-cbm-OX4-sshg_05717
[0082] Plasmids pUC-cbmA, pUC-cbmB, pUC-OX4 and vector plasmid pSET5035-KT were digested with MfeI and SpeI respectively to obtain cbmA-MfeI / SpeI, cbmB-MfeI / SpeI, OX4-MfeI / SpeI fragments and vector fragment pSET5035 -MfeI / PstI.
[0083] The cbmC-XbaI / PstI, sgr810-XbaI / PstI and sshg_05717-XbaI / PstI fragments were obtained from the plasmids pUC-cbmC, pUC-sgr810 and pUC-sshg_05717 respectively after double digestion with XbaI and PstI.
[0084] cbmB-MfeI / SpeI, cbmC-XbaI / PstI and vector fragment pSET5035-MfeI / PstI were ligated by ligase to obtain plasmid pSET5035-cbmB-C.
[0085] Plasmid pSET5035-cbmB-C was digested with XbaI and PstI to obtain cbmB-C-XbaI / PstI fragment; cbmA-MfeI / SpeI, cbmB-C-XbaI / PstI and vector fragment pSET5035-MfeI / PstI were ligated by ligase, Plasmid pSET5035-cbmA-B-C was obtained.
[0086] OX4-MfeI / SpeI, sgr810-XbaI / PstI and carrier plasmid pSET...
PUM
 Login to view more
Login to view more Abstract
Description
Claims
Application Information
 Login to view more
Login to view more - R&D Engineer
- R&D Manager
- IP Professional
- Industry Leading Data Capabilities
- Powerful AI technology
- Patent DNA Extraction
Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic.
© 2024 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap