Diagnostic marker, scoring model and diagnostic product for dilated cardiomyopathy
A technology for dilated cardiomyopathy and diagnostic markers, applied in the field of biomedicine, can solve problems such as difficulty in judging dilated cardiomyopathy, and achieve the effect of accurate judgment
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0103] The gene combination used for the diagnosis of human dilated cardiomyopathy described in this embodiment includes 13 characteristic genes (biomarkers), and the selection of these characteristic genes mainly includes the following steps:
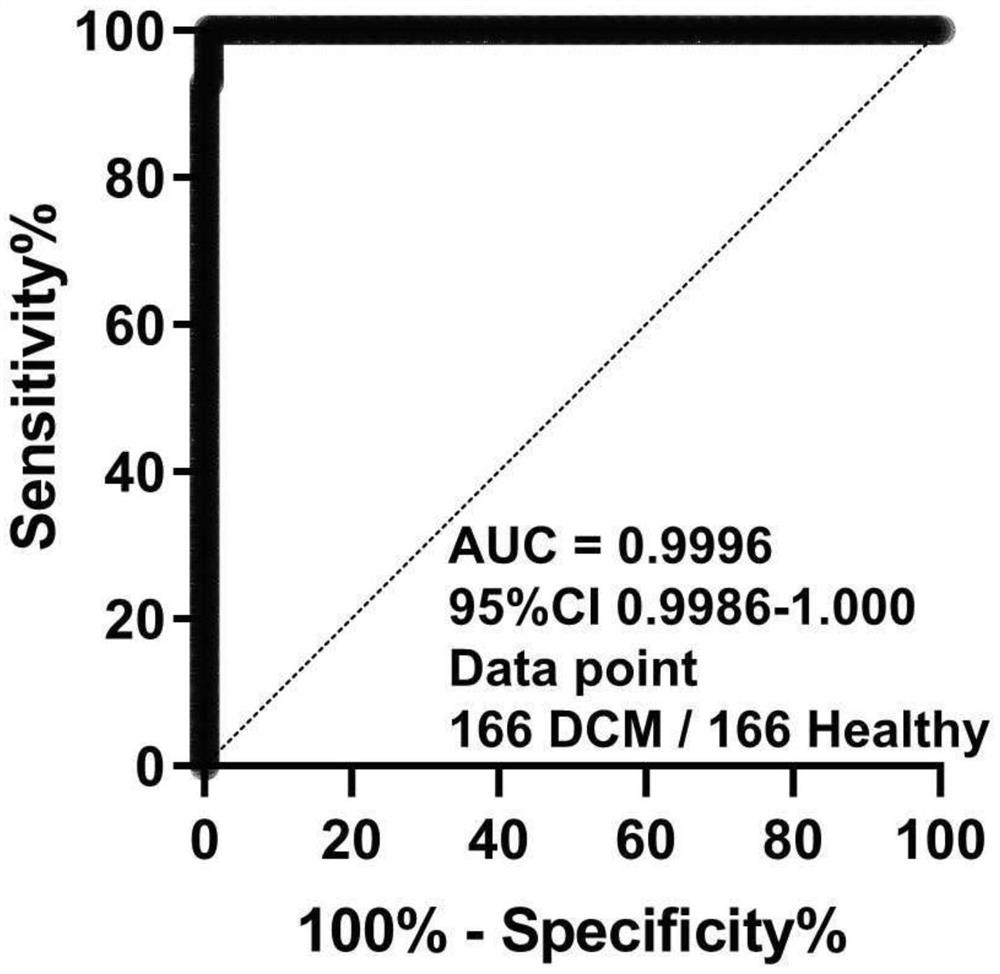
[0104] (1) Collection and processing of training set samples: The inventor of the application in this case analyzed a large sample of clinical data and biological sample data of patients with dilated cardiomyopathy and healthy people, a total of 332 cases, including 166 cases of dilated cardiomyopathy Corresponding RNA-seq expression profiling data and clinical data from patients and 166 healthy samples.
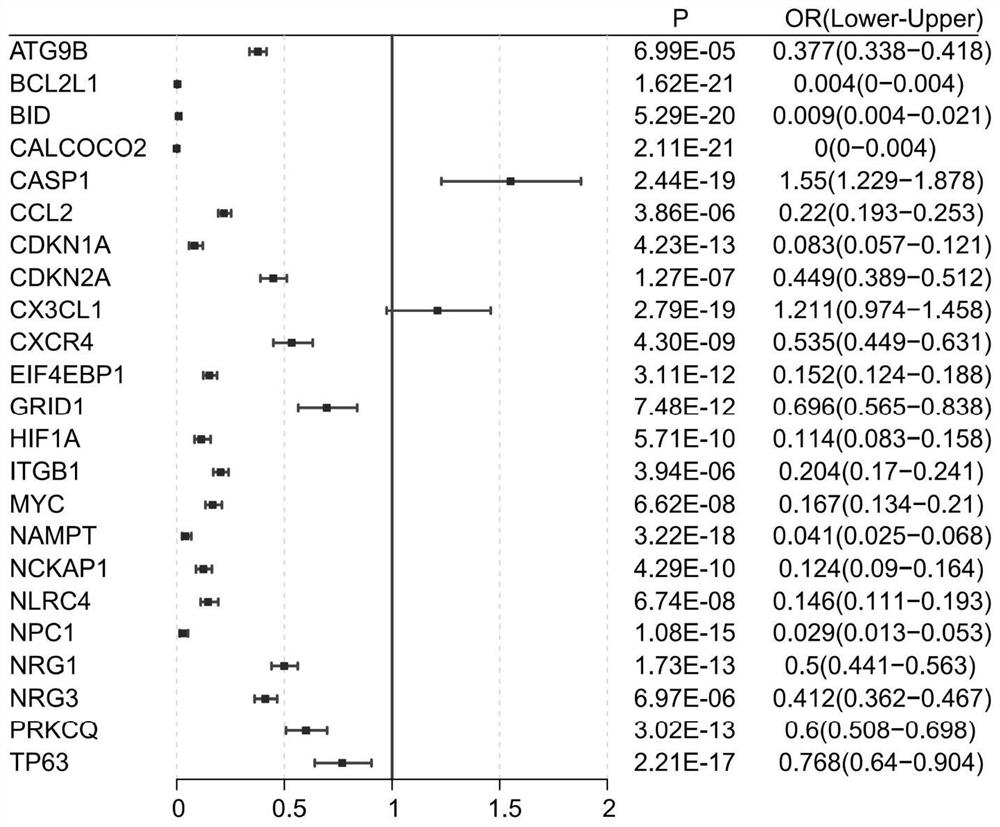
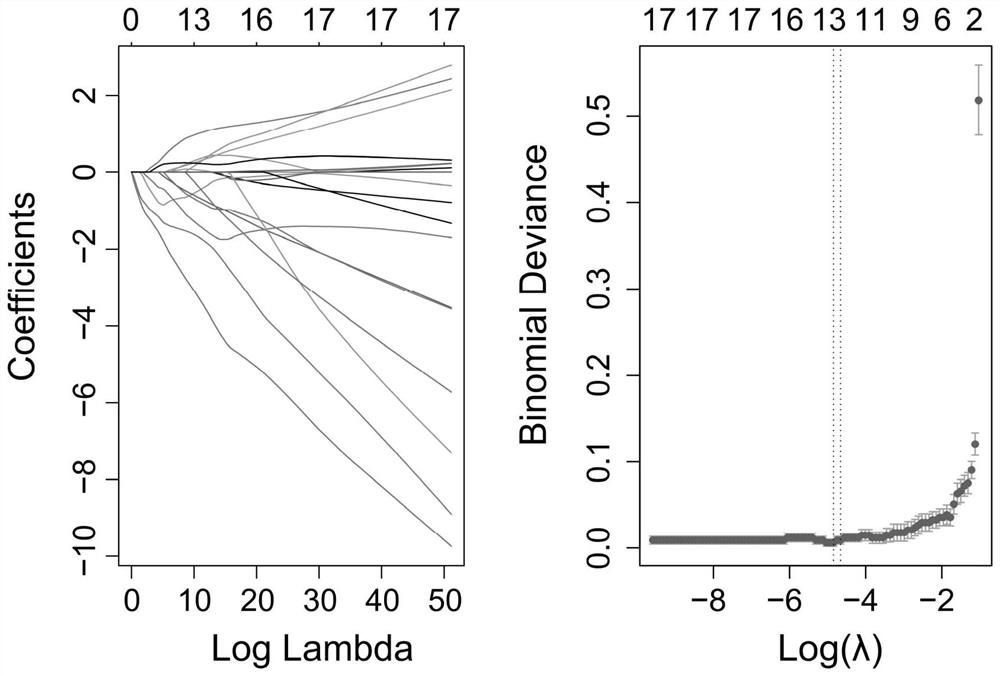
[0105] (2) Screening of 13 specific genes:
[0106] Autophagy is a process of cell self-phagocytosis and waste recycling. Under normal circumstances, autophagy maintains cellular homeostasis. In recent years, studies have reported that autophagy is closely related to the occurrence of dilated cardiomyopathy. Therefore, the inventor...
Embodiment 2
[0118] In this example, 10 left ventricular biopsy specimens were used to collect RNA-seq data:
[0119] Step 1. Extraction and preservation of human left ventricular tissue samples.
[0120] The myocardial biopsy tissue waste (left ventricular specimen) obtained from heart disease patients during surgery was obtained from the operating room of the hospital, and paraffin sections of 4um×10 were made by the pathology department and stored at room temperature.
[0121] Step 2. RNA-seq to determine the gene expression data of left ventricular tissue.
[0122] Specifically, it can be divided into two steps: sample preparation and sequencing library construction:
[0123] (1) First, extract total RNA from paraffin sections. The specific steps are: take paraffin sections, add wax remover, incubate at 56 °C, add protein PKD, add enzyme K, incubate at 56 °C and 80 °C, add DNase I, and use anhydrous Ethanol washing, and finally RNA elution to obtain total RNA.
[0124] (2) When buil...
Embodiment 3
[0174] This example uses the gene chip data of 10 left ventricular biopsy specimens, and the chip platform is GPL3050.
[0175] Step 1. Human left ventricular tissue extraction and preservation.
[0176] The myocardial biopsy tissue waste (left ventricular specimen) obtained from heart disease patients during surgery was obtained from the operating room of the hospital, and paraffin sections of 4um×10 were made by the pathology department and stored at room temperature.
[0177] Step 2: Gene chip directly measures gene expression. The specific steps can be divided into three steps:
[0178] (1) Sample preparation and labeling. The preparation of the samples is consistent with the total RNA extraction procedure of the "Determination of Gene Expression by RNA-seq" technical process. After the total RNA is obtained, the mRNA needs to be enriched and labeled after amplification. Labeling methods include biotin labeling, fluorescein labeling, isotope labeling and the like.
[0...
PUM
 Login to view more
Login to view more Abstract
Description
Claims
Application Information
 Login to view more
Login to view more - R&D Engineer
- R&D Manager
- IP Professional
- Industry Leading Data Capabilities
- Powerful AI technology
- Patent DNA Extraction
Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic.
© 2024 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap