Fusion protein for cellulose accessibility measurement and application thereof
A fusion protein and cellulose technology, which is applied in the direction of fusion with spectrum/fluorescence detection, measurement devices, enzymes, etc., can solve the problems of large measurement deviation, poor repeatability, time-consuming and laborious measurement process, etc., to improve accuracy and avoid changes Effect
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0035] Construction of recombinant plasmid pET28a-GFP2-CBM in Escherichia coli
[0036] According to the sequences of GFP gene and CBM gene in Genebank, two gene sequences of GFP and CBM fusion protein were designed and synthesized on the pUC57-simple vector. According to the gene sequence of the fusion protein GFP2-CBM and the characteristics of the multiple cloning site on the Escherichia coli expression vector pET28a, the following two primers were designed and synthesized:
[0037] GFP_Fe: 5'-GACgaattcATGCGTAAAGGCGAAGAGCTGTTC-3'
[0038] CBM_Rh:5'- GACaagcttTTACAAGCACTGAGAGTAGTAAGG-3'
[0039] EcoRI and HindIII restriction sites were designed at the two ends of GFP_Fe and CBM_Rh, respectively, and are indicated in lowercase fonts. Using GFP_Fe, CBM_Rh primers, using the recombinant plasmid pUC57-simple-GFP2-CBM constructed in our laboratory as a template, the GFP2-CBM gene was amplified by PCR. PCR conditions: pre-denaturation at 95°C for 5 minutes; then 30 cycles of de...
Embodiment 2
[0042] Expression of fusion protein GFP2-CBM
[0043] The recombinant plasmid pET28a-GFP2-CBM was transformed into Escherichia coli expression strain BL21, and the bacterial solution was spread on LB plates containing 50 μg / ml Kanamycin for resistance screening, single clones were picked for PCR detection, and positive clones were selected.
[0044] Pick the recombinant Escherichia coli expression strain BL21-pET28a-GFP2-CBM, inoculate it in 5 ml LB medium, and cultivate it overnight at 37°C on a shaker at 200 rpm. Then take 1 ml of the bacterial liquid and transfer it to 100 ml of LB medium, and culture it on a shaker at 37°C and 200 rpm to OD 600 0.6, add IPTG to a final concentration of 1 mM, and culture at 30°C for 6 h on a shaker at 200 rpm. The bacteria were collected by centrifugation, supernatant was obtained by ultrasonic crushing, purified by Ni-NTA medium column, and dialyzed to remove salt. The final protein yield was 100 mg / L bacterial liquid. After SDS-PAGE ele...
Embodiment 3
[0046] Example 3 Recognition of pure cellulose by fusion protein
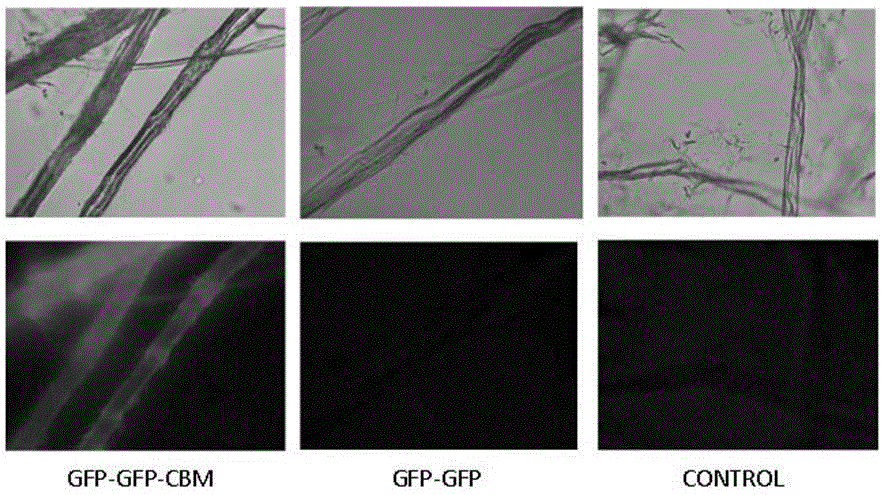
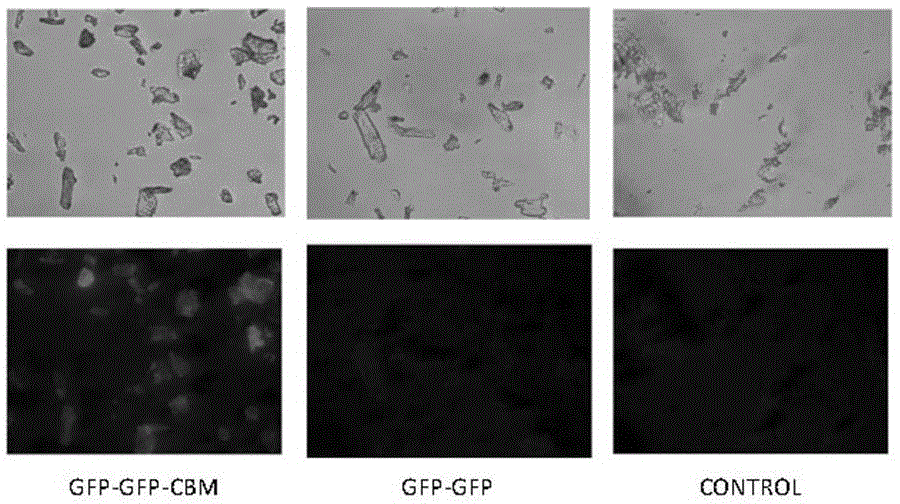
[0047] The fusion protein GFP-GFP-CBM was obtained according to the steps described in Examples 1 and 2. And use similar steps to obtain the fluorescent protein GFP-GFP without CBM. Two proteins, GFP-GFP-CBM and GFP-GFP, were respectively adsorbed on filter paper and microcrystalline cellulose as substrates at 37 °C and 200 rpm for 2 h, and the adsorbed filter paper cellulose was observed under a fluorescence microscope (attached figure 1 ) and microcrystalline cellulose (with figure 2 ), it can be found that due to the recognition of CBM on cellulose, the GFP-GFP-CBM fusion protein can be effectively adsorbed on the cellulose, while the GFP-GFP fusion protein does not have this function, so it is not obvious under the fluorescence microscope fluorescence phenomenon.
[0048] Result analysis: The results of this example show that the fusion protein provided by the present invention can obtain visual analysi...
PUM
| Property | Measurement | Unit |
|---|---|---|
| Molecular weight | aaaaa | aaaaa |
| Molecular weight | aaaaa | aaaaa |
Abstract
Description
Claims
Application Information
 Login to View More
Login to View More