Tumor marker for detecting human colorectal cancer and application thereof
A tumor marker, colorectal cancer technology, applied in the field of biological detection, can solve the problems of low specificity and high false positive rate
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
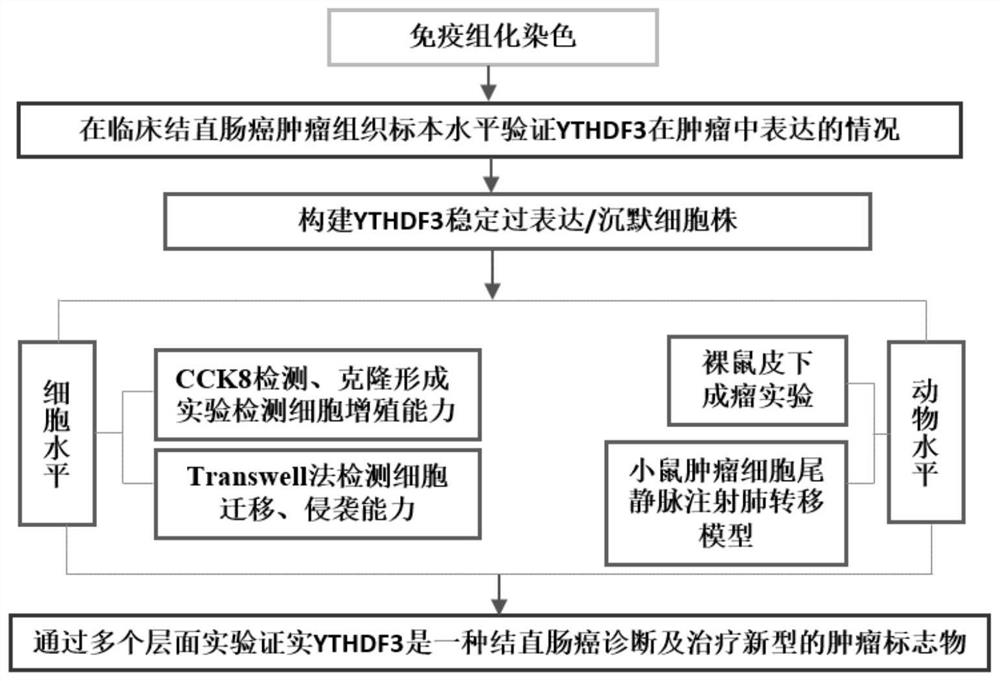
[0022] 200 paraffin samples of tumor tissue from colorectal cancer patients with complete follow-up information were collected, and an N6-methyladenine (m 6 A) The expression of the recognition enzyme YTHDF3 in colorectal cancer tumor tissues was significantly up-regulated compared with adjacent tissues, and the prognosis of colorectal cancer patients with high YTHDF3 expression was poor; The experimental model of tumor formation confirmed the ability of stable overexpression of YTHDF3 to promote the proliferation of colorectal cancer in vivo, and the proliferation of colorectal cancer cells in vivo and in vitro was slowed down by stably knocking out YTHDF3. Technical route such as figure 1 shown.
Embodiment 2
[0024] Immunohistochemistry (IHC): Routine staining steps: xylene I dewaxed for 15 min, xylene II dewaxed for 15 min, xylene III dewaxed for 15 min, 100% ethanol I hydrated for 5 min, 100% ethanol II hydrated for 5 min, 80% Ethanol hydration for 5 minutes, distilled water for 1 minute, hematoxylin semen staining for 5 minutes, 1% hydrochloric acid ethanol for 1-3 seconds, running water to remove hematoxylin semen for 15 minutes, 0.5% eosin solution for 1-3 minutes, distilled water rinse for 1-2 seconds, and 80% ethanol dehydration 1 min, 95% ethanol Ⅰ dehydration 1 min, 95% ethanol Ⅱ dehydration 1 min, absolute ethanol dehydration 1 min, xylene transparent 2 min, neutral gum seal, dry, and observe under a microscope. For IHC, add 0.3% hydrogen peroxide methanol solution for 30 minutes after rehydration to eliminate the influence of endogenous peroxidase, and then use PBS to wash 3 times, 5 minutes each time; act for 30 minutes with goat serum blocking solution, add The corresp...
Embodiment 3
[0027] qRT-PCR: (1) use TRIzol to extract the total RNA of colorectal cancer cells; (2) use the RNA reverse transcription kit to reverse transcribe the RNA into cDNA; (3) perform quantitative PCR detection.
[0028] (1) Use TRIzol to extract total RNA from colorectal cancer cells: collect the cell pellet, add 1ml TRIzol and vortex mix; add 0.2ml chloroform, shake vigorously for 15 seconds, and place at room temperature for 3 minutes. Centrifuge at 10,000g for 15 minutes at 2-8°C. The sample is divided into three layers: the bottom layer is a yellow organic phase, the upper layer is a colorless aqueous phase and an intermediate layer. RNA is mostly in the aqueous phase, imbibing the aqueous layer. Precipitate the RNA in the aqueous phase with isopropanol. Add 0.5ml of isopropanol for every 1ml of TRIzol used, and let stand at room temperature for 10 minutes. Centrifuge at 10,000g for 10 minutes at 2-8°C, and remove the supernatant. Wash the RNA pellet with 75% ethanol, cent...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More