Ipomoea batatas metallothionein gene (IbMT1) and application thereof in improving salt resistance and drought resistance of plants
A technology of metallothionein and gene, applied in the field of molecular biology and biology, can solve the problems of poor tolerance and limit the extensive growth of plants, so as to improve the resistance to drought, improve the salt tolerance and drought resistance of plants, and improve the salt tolerance of plants Effect
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
specific Embodiment approach
[0018] The specific embodiment: the present invention will be described in further detail below in conjunction with accompanying drawing:
[0019] The methods used in the following examples are conventional methods unless otherwise specified.
[0020] experiment material
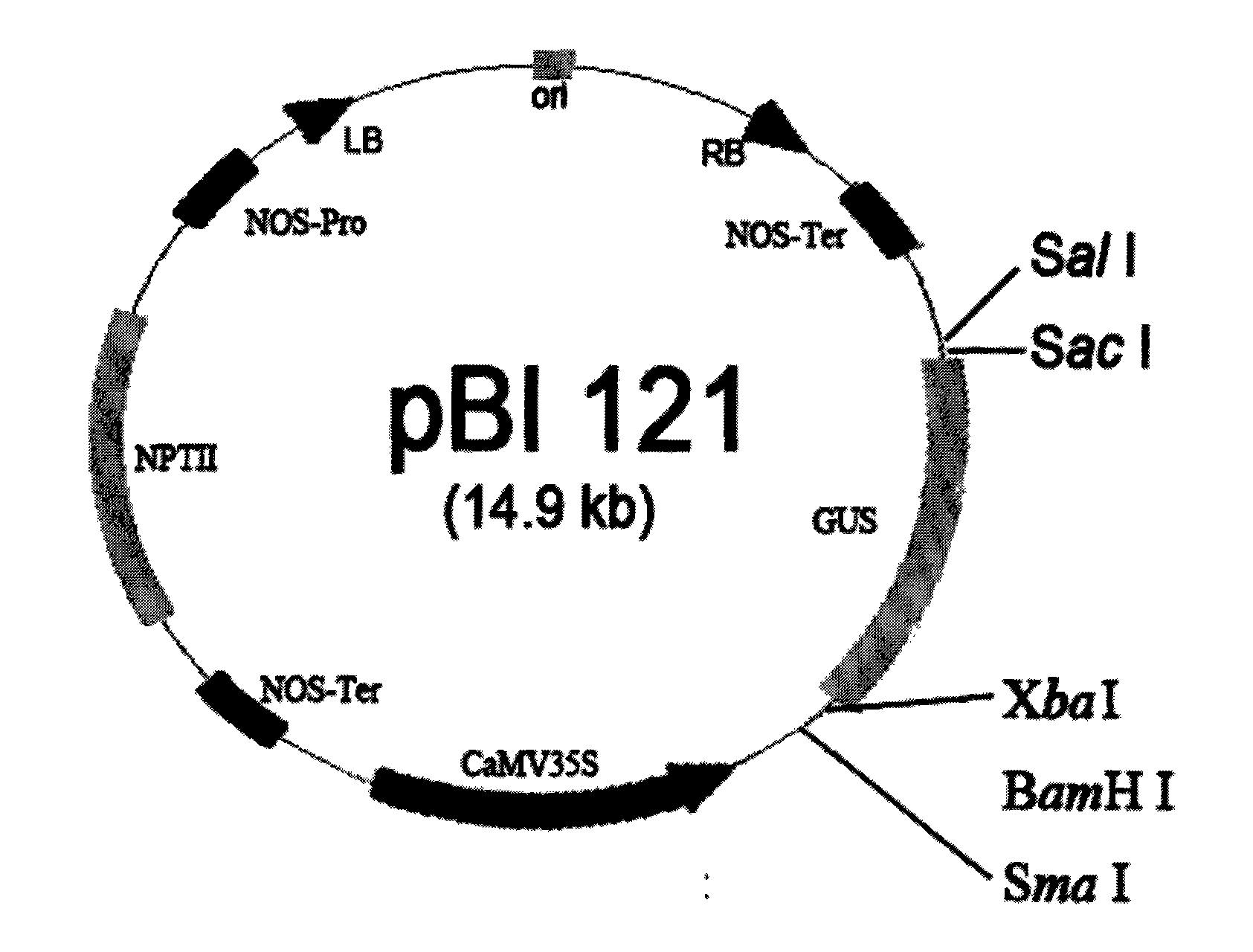
[0021] 1. Strains and vectors: Escherichia coli strain DHα and Agrobacterium strain EHA105 were purchased from Huamei Biological Company; expression vector pBI121 was purchased from Clontech Company.
[0022] 2. Main reagent: SMART TM PCR cDNA Synthesis Kit and ClontechPCR-Select cDNASubtraction Kit were purchased from Clontech Company; Taq DNA polymerase was purchased from Beijing Quanshijin Company, DNA standard molecular weight was purchased from Takara Company; restriction endonuclease, T4DNA ligase, M-MLV inversion Recording enzymes were purchased from Promega; RNeasy Plant Mini Kit was purchased from Qiagen; other chemical reagents were of domestic analytical grade.
[0023] 3. The primer sequence ...
Embodiment 1
[0036] Embodiment 1: the acquisition of sweet potato metallothionein gene IbMT1
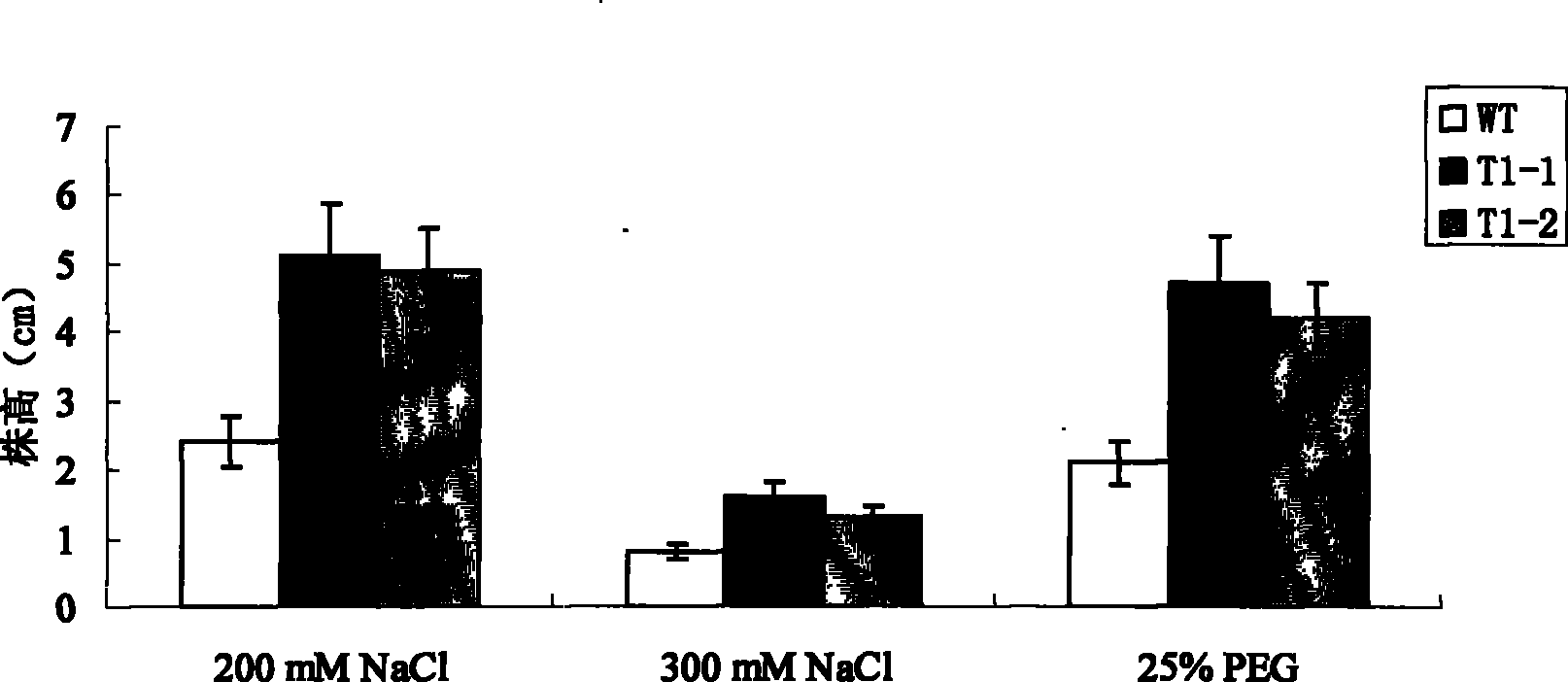
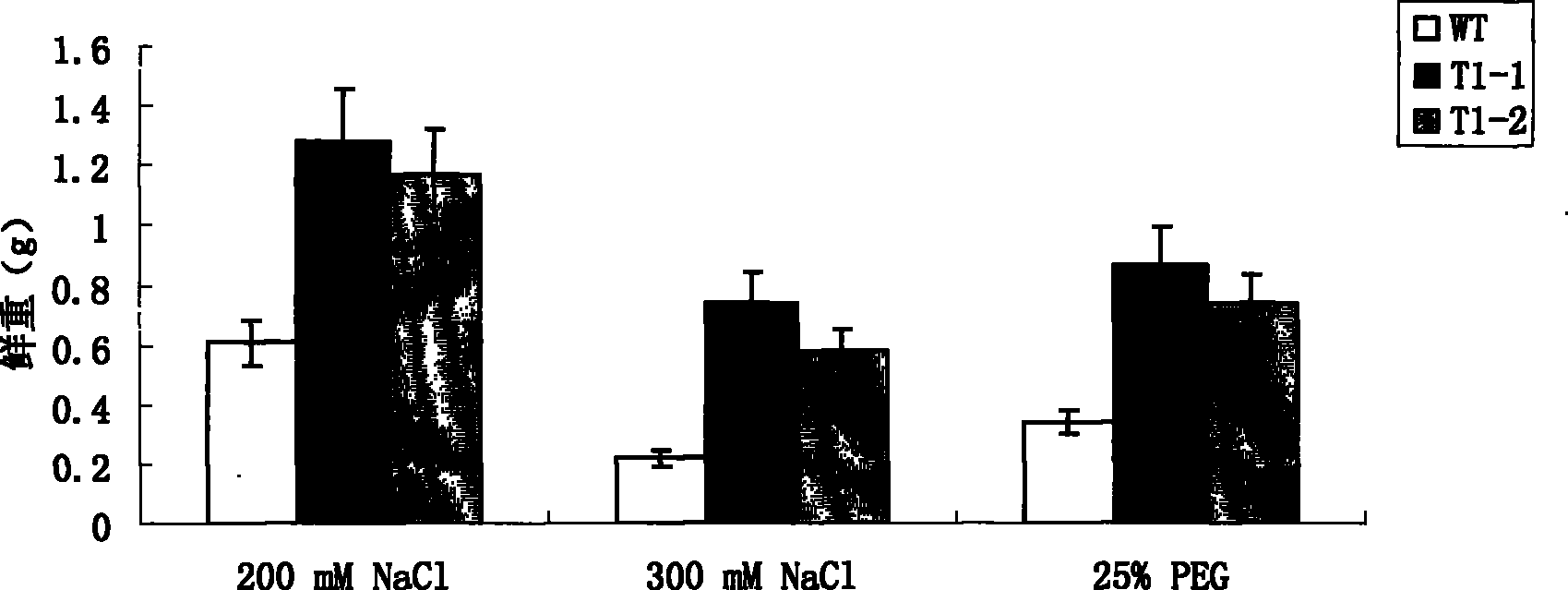
[0037] 1. Treatment of sweet potato seedlings: Select stem tips or undifferentiated axillary buds of sweet potato plants, after disinfection, cut them into sections and inoculate them on MS solid medium for cultivation. The cultivation temperature is 27±2°C and the light is about 2000 Lux. When the aseptic seedlings grow to a height of about 3 cm, select vigorously growing healthy aseptic seedlings and transfer them to MS solid medium containing 200 mM NaCl for induction and culture for 24 hours.
[0038] 2. Extraction of total RNA: RNeasy Plant Mini Kit (Qiagen Company) was used to extract total RNA of NaCl-treated and untreated sweet potato seedlings, and the specific operation was carried out according to the kit instructions.
[0039] 3. Construction of a salt stress suppression subtractive library: 2 μg of total RNA from NaCl-treated and untreated sweet potato seedlings was reverse-transcrib...
Embodiment 2
[0042] Embodiment 2: Construction of PBI121-IbMT1 plant expression vector
[0043] 1. Amplification of the coding region of IbMT1 cDNA: Design primers IbMT-BamH I and IbMT-Sac I according to the sequences at both ends of the coding region of IbMT1 cDNA, and carry out PCR amplification with the bacterial solution containing IbMT1 as a template. Into the 25μL reaction system, add: 10×buffer (Mg 2+ ) 2.5 μL, dNTP (10 mM) 1 μL, each primer 1 μL (10 μM), template 1 μL, Taq enzyme 0.2 μL, and ddH for the rest 2 O make up.
[0044] The conditions of PCR amplification were as follows: 94° C. for 5 minutes, followed by 1 minute at 94° C., 1 minute at 56° C., and 1 minute at 72° C., for a total of 35 cycles. The results of agarose gel electrophoresis showed that a specific band with a size of about 200 bp was amplified, which was sent to Beijing Huada Gene Research Center for sequencing and confirmed to be the complete sequence of the IbMT1 cDNA coding region.
[0045] 2. Digestion a...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More