Dengue hemorrhagic fever virus II nucleic acid molecular characteristic standard sample and preparation method thereof
A dengue hemorrhagic fever and standard sample technology, which is applied in the field of dengue hemorrhagic fever virus type II nucleic acid molecular characteristic standard samples and its preparation, can solve the problems of shortened inspection process time limit, difficulty in ensuring laboratory quality control, etc., and achieve uniformity Good, strong stability, good purity
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0022] Embodiment 1: the preparation of standard sample
[0023] 1. Synthetic sequence: use the software Clone Manager 7.0 to download the publicly released sequence from GenBank, perform homology analysis and primer matching, and analyze and determine the fitted viral sequence: ATG AAT AAC CAA CGA AAA AAG GCG AGA AGT ACG CCT TTC AAT ATG CTG AAA CGC GAG AGA AAC CGC GTG TCA ACT GTG CAA CAG CTG ACA AAG AGA TTC TCA CTT GGA ATG CTG CAA GGA CGC GGA CCA TTA AAA CTG TTC ATG GCC CTT GTG GCG TTC CTT CGT TTC CTA ACA ATC CCA CCA ACA GCA GGG ATA CTA AAA AGA TGG GGA ACG ATC AAA AAA TCA AAA GCT ATC AAT GTC TTG AGA GGG TTC AGG AA, the sequence synthesis is entrusted to the company; in order to facilitate the in vitro transcription of the viral nucleic acid sequence into RNA, a T7 promoter is added in front of each sequence, Sequence 20bp: TAA TAC GAC TCA CTA TAG GG;
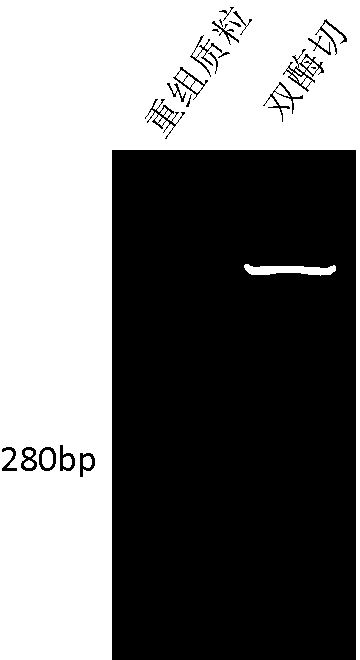
[0024] 2. Clone the synthetic sequence bluntly into the pUC19 plasmid and identify it by enzyme digestion and gel purifica...
Embodiment 2
[0045] Embodiment 2: the homogeneity inspection of standard sample
[0046] Samples were randomly selected for homogeneity analysis according to the random sampling method:
[0047] 1. Uniformity in bottle
[0048] Randomly select 1 bottle, add 200 μL of deionized water, take three parts, each part is 50 μL, numbered 101-103, each part is tested three times, the results are shown in Table 2; the quality data of the three groups were analyzed by variance, and the quality was not significant The difference meets the requirement of uniformity, and the standard deviation is 0.0006. The purity is about 1.80, meeting the purity requirement.
[0049] Table 2 Experimental results of homogeneity in the bottle (mass unit: μg)
[0050]
[0051] 2. Uniformity between bottles
[0052] 20 bottles, numbered 201-220, were selected by simple random sampling method, and each bottle was tested 3 times. Measurement sequence: first time, 201→220; second time, 220→201; third time, odd num...
Embodiment 3
[0055] Embodiment 3: the stability check of standard sample
[0056] Samples were randomly selected for stability analysis according to the random sampling method:
[0057] 1. short-term stability
[0058] Freeze-dried samples were randomly selected and placed at -20°C, 0°C, 4°C, 25°C, and 37°C for 2 weeks. Place 8 bottles at each temperature, 40 bottles in total, numbered from 301-350, and measure the quality 3 times for each bottle. Refer to the statistical analysis requirements of GB / T 15000.3-2008 for sample stability, and use SPSS 20.0 software to analyze the variance of the data;
[0059] The 5 groups of data (Table 4) were analyzed by variance, and there were significant statistical differences among the groups. Further comparisons were made among the groups. There was no statistical difference between the -20°C, 0°C, and 4°C groups. There is no statistical difference between ℃, but there is statistical difference between the three groups of -20 ℃, 0 ℃ and 4 ℃ and t...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More - R&D
- Intellectual Property
- Life Sciences
- Materials
- Tech Scout
- Unparalleled Data Quality
- Higher Quality Content
- 60% Fewer Hallucinations
Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic, Popular Technical Reports.
© 2025 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap|About US| Contact US: help@patsnap.com