Artificial antibody polypeptides
a technology of artificial antibodies and polypeptides, which is applied in the direction of library member identification, directed macromolecular evolution, fungi, etc., can solve the problems of limiting the level of uptake, the size of antibodies and the complexity of six loops is a major design hurdle, and the antibody fragments are not suitable for structural analysis using nmr spectroscopy, etc., to achieve the effect of increasing the melting point and increasing the melting poin
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Benefits of technology
Problems solved by technology
Method used
Image
Examples
example i
Construction of the Fn3 Gene
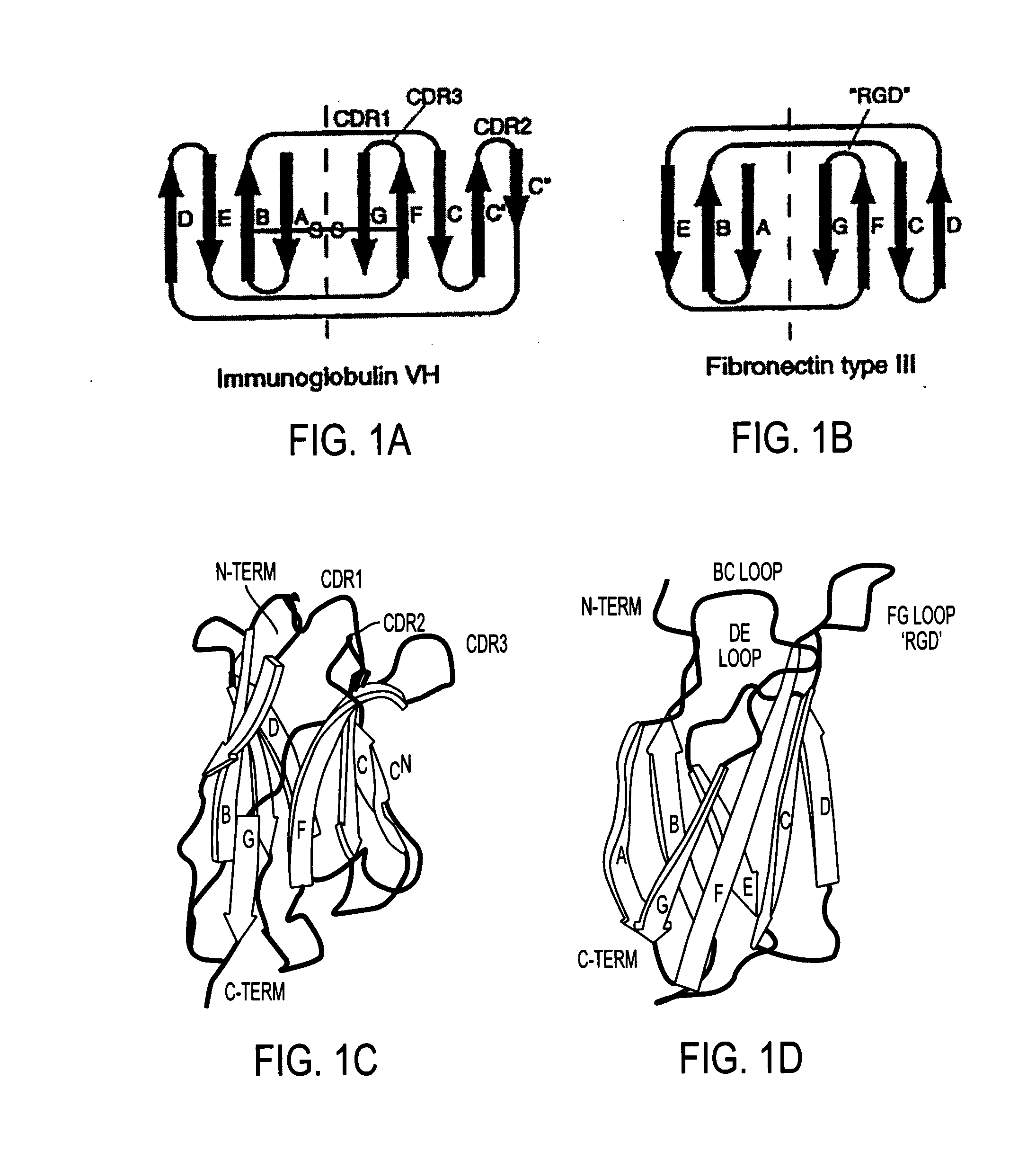
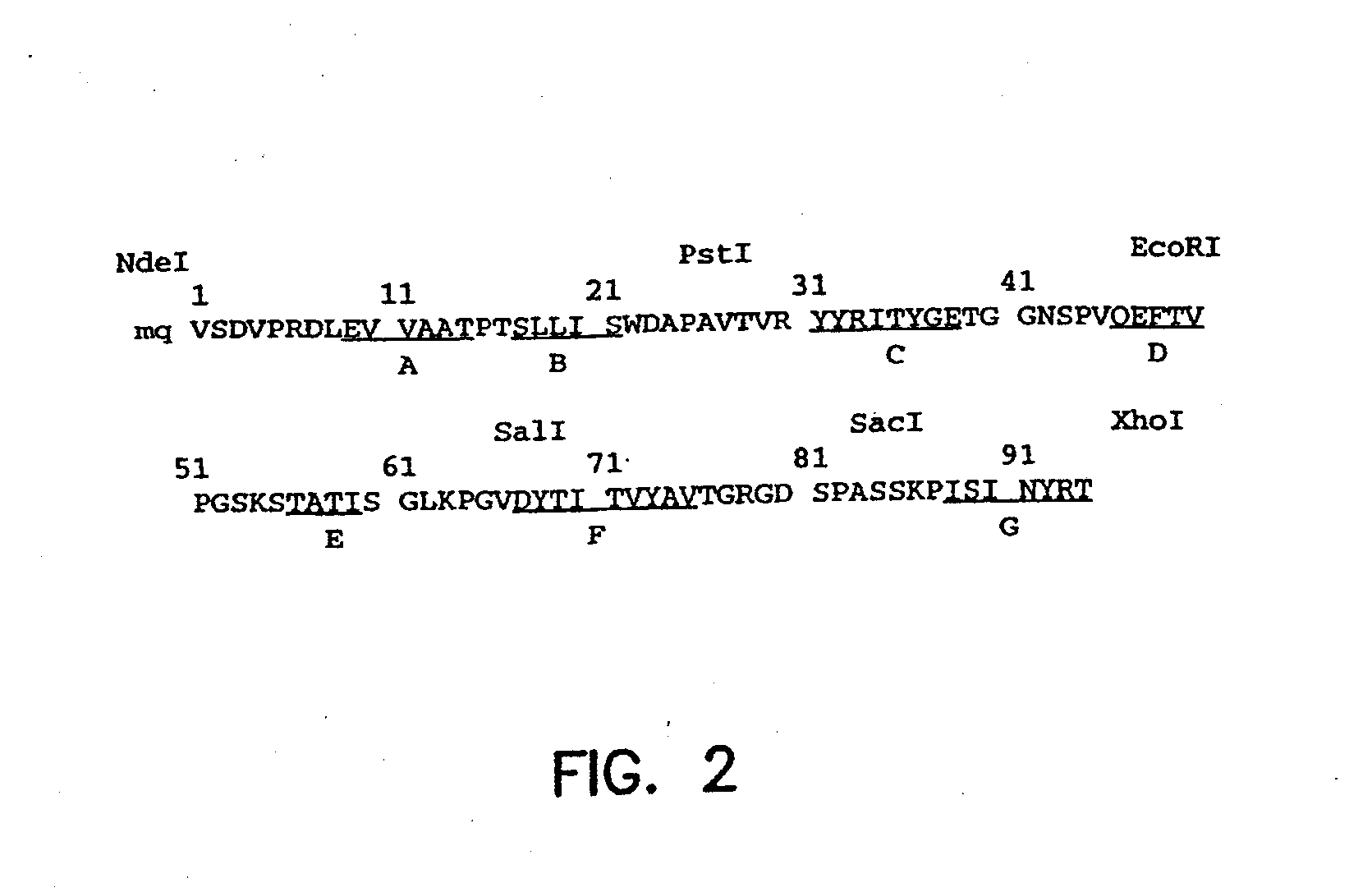
[0145]A synthetic gene for tenth Fn3 of fibronectin (FIG. 1) was designed on the basis of amino acid residue 1416-1509 of human fibronectin (Kornblihtt, et al., 1985) and its three dimensional structure (Main, et al., 1992). The gene was engineered to include convenient restriction sites for mutagenesis and the so-called “preferred codons” for high level protein expression (Gribskov, et al., 1984) were used. In addition, a glutamine residue was inserted after the N-terminal methionine in order to avoid partial processing of the N-terminal methionine which often degrades NMR spectra (Smith, et al., 1994). Chemical reagents were of the analytical grade or better and purchased from Sigma Chemical Company and J. T. Baker, unless otherwise noted. Recombinant DNA procedures were performed as described in “Molecular Cloning” (Sambrook, et al., 1989), unless otherwise stated. Custom oligonucleotides were purchased from Operon Technologies. Restriction and modific...
example ii
Modifications to Include Restriction Sites in the Fn3 Gene
[0151]The restriction sites were incorporated in the synthetic Fn3 gene without changing the amino acid sequence Fn3. The positions of the restriction sites were chosen so that the gene construction could be completed without synthesizing long (>60 bases) oligonucleotides and so that two loop regions could be mutated (including by randomization) by the cassette mutagenesis method (i.e., swapping a fragment with another synthetic fragment containing mutations). In addition, the restriction sites were chosen so that most sites were unique in the vector for phage display. Unique restriction sites allow one to recombine monobody clones which have been already selected in order to supply a larger sequence space.
example iii
Construction of M13 Phage Display Libraries
[0152]A vector for phage display, pAS38 (for its map, see FIG. 8) was constructed as follows. The XbaI-BamHI fragment of pET12a encoding the signal peptide of OmpT was cloned at the 5′ end of the Fn3 gene. The C-terminal region (from the FN5F and FN5R oligonucleotides, see Table 2) of the Fn3 gene was replaced with a new fragment consisting of the FN5F and FN5R′ oligonucleotides (Table 2) which introduced a MluI site and a linker sequence for making a fusion protein with the pIII protein of bacteriophage M13. A gene fragment coding the C-terminal domain of M13 pIII was prepared from the wild-type gene III of M13 mp18 using PCR (Corey, et al., 1993) and the fragment was inserted at the 3′ end of the OmpT-Fn3 fusion gene using the MluI and HindIII sites.
[0153]Phages were produced and purified using a helper phage, M13K07, according to a standard method (Sambrook, et al., 1989) except that phage particles were purified by a second polyethylene...
PUM
| Property | Measurement | Unit |
|---|---|---|
| dissociation constant | aaaaa | aaaaa |
| melting point | aaaaa | aaaaa |
| melting point | aaaaa | aaaaa |
Abstract
Description
Claims
Application Information
 Login to View More
Login to View More