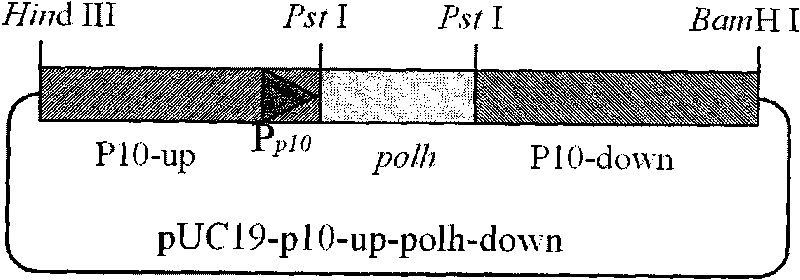Patents
Literature
212 results about "BamHI" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
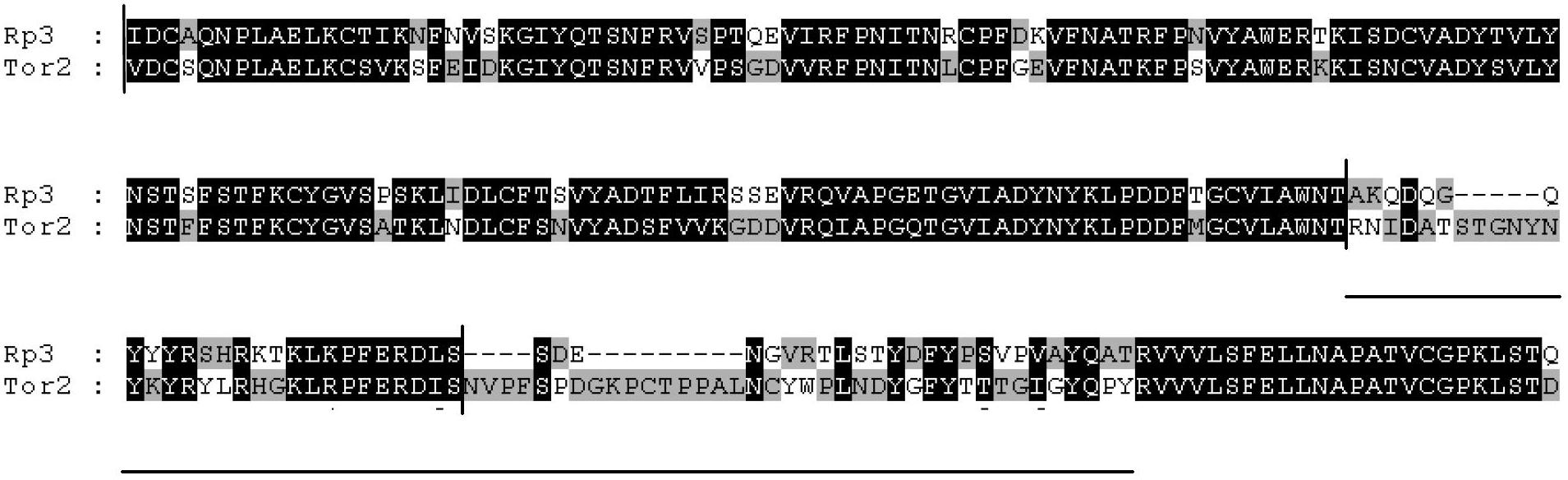
BamH I (from Bacillus amyloliquefaciens) is a type II restriction endonuclease, having the capacity for recognizing short sequences (6 b.p.) of DNA and specifically cleaving them at a target site. This exhibit focuses on the structure-function relations of BamH I as described by Newman, et al. (1995). BamH I binds at the recognition sequence 5'-GGATCC-3', and cleaves these sequences just after the 5'-guanine on each strand. This cleavage results in sticky ends which are 4 b.p. long. In its unbound form, BamH I displays a central b sheet, which resides in between α-helices. BamH I is an extraordinarily unique molecule in that it undergoes a series of unconventional conformational changes upon DNA recognition. This allows the DNA to maintain its normal B-DNA conformation without distorting to facilitate enzyme binding. BamH I is a symmetric dimer. DNA is bound in a large cleft that is formed between dimers; the enzyme binds in a "crossover" manner. Each BamH I subunit makes the majority of its backbone contacts with the phosphates of a DNA half site but base pair contacts are made between each BamH I subunit and nitrogenous bases in the major groove of the opposite DNA half site. The protein binds the bases through either direct hydrogen bonds or water-mediated H-bonds between the protein and every H-bond donor/acceptor group in the major groove. Major groove contacts are formed by atoms residing on the amino-terminus of a parallel 4 helix bundle. This bundle marks the BamH I dimer interface, and it is thought that the dipole moments of the NH2-terminal atoms on this bundle may contribute to electrostatic stabilization.
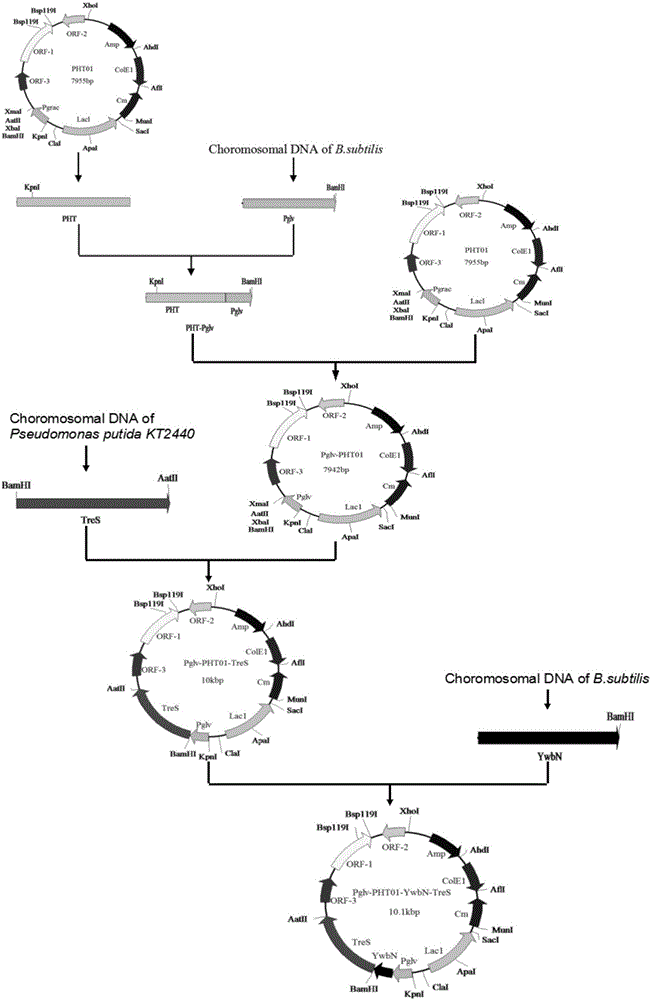
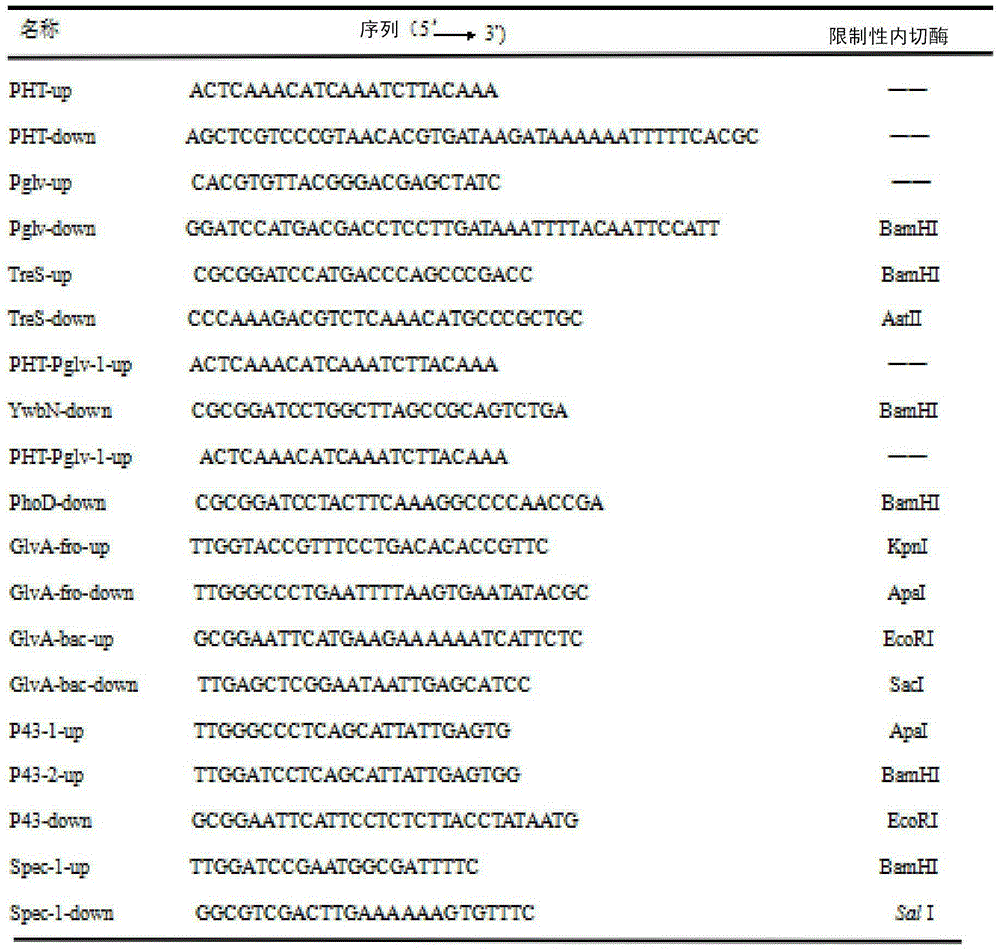
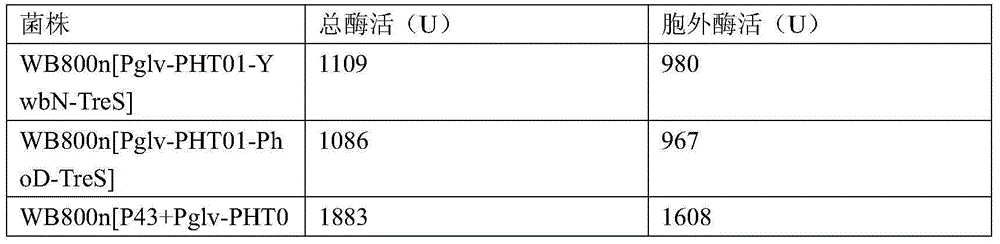
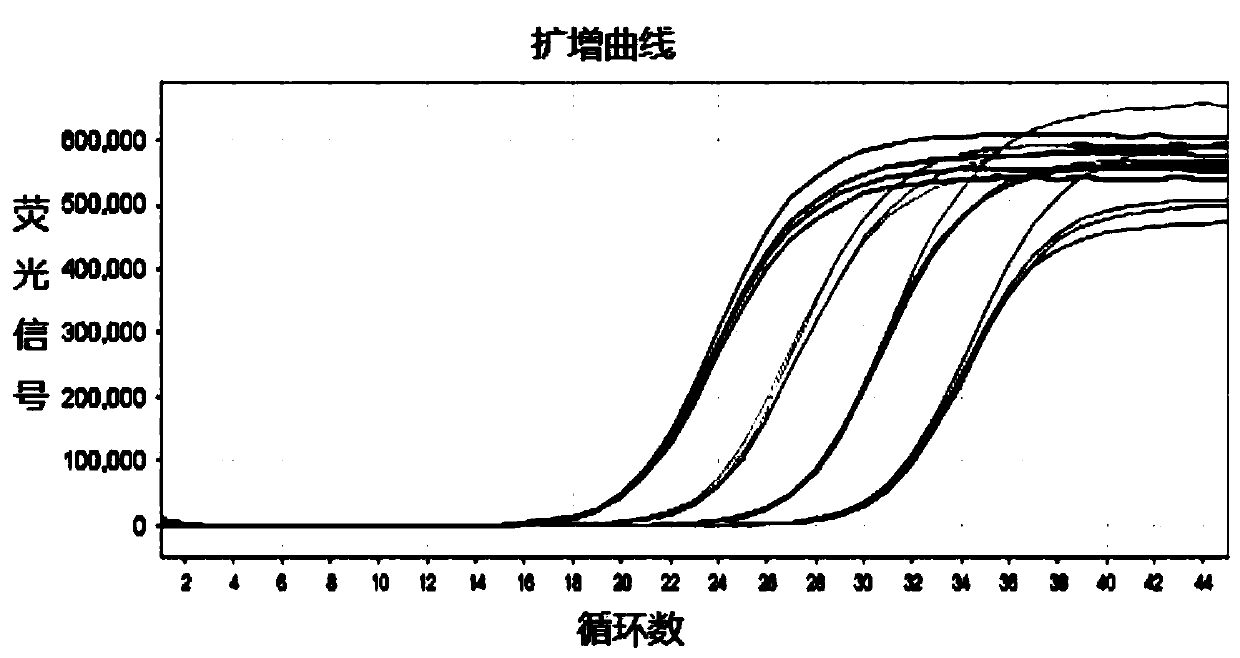
Maltose inducible trehalose synthase synthesis engineering bacterium, method for preparing same and application
The invention relates to a maltose inducible trehalose synthase synthesis engineering bacterium, a method for preparing the same and application. The maltose inducible trehalose synthase synthesis engineering bacterium is characterized in that maltose inducible promoters are inserted in the fronts of BamHI cleavage sites of PHT01 plasmids of recombinant plasmid vectors instead of Pgrac promoters on the PHT01 plasmids, expression genes of Tat type signal peptides are inserted in the fronts of the BamHI cleavage sites, and expression genes of trehalose synthase are inserted in the rears of the BamHI cleavage sites. The maltose inducible trehalose synthase synthesis engineering bacterium, the method and the application have the advantage that expression effects realized after the maltose inducible promoters and the trehalose synthase are fused with one another are obviously superior to other inducible expression effects.
Owner:山东开盾生物科技有限公司
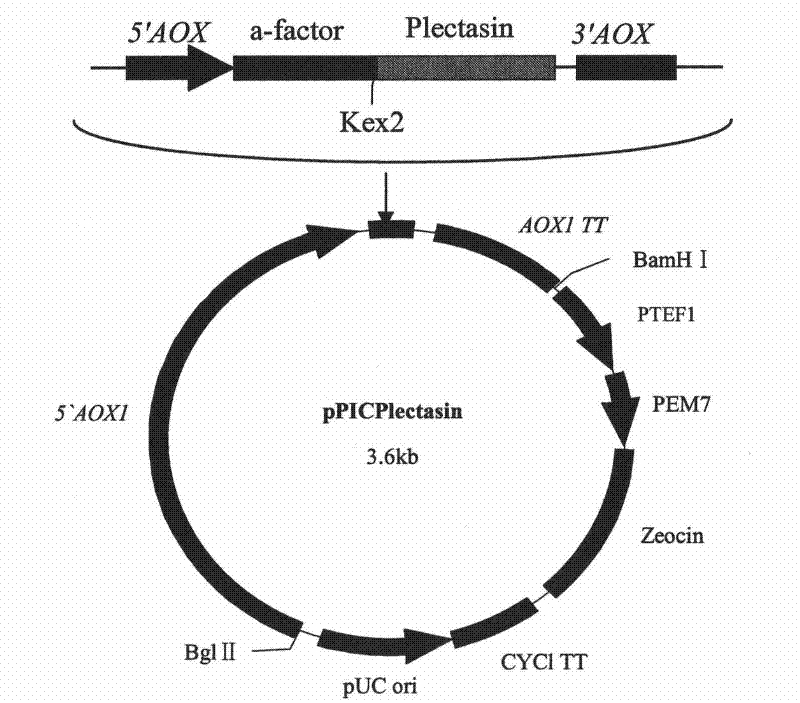
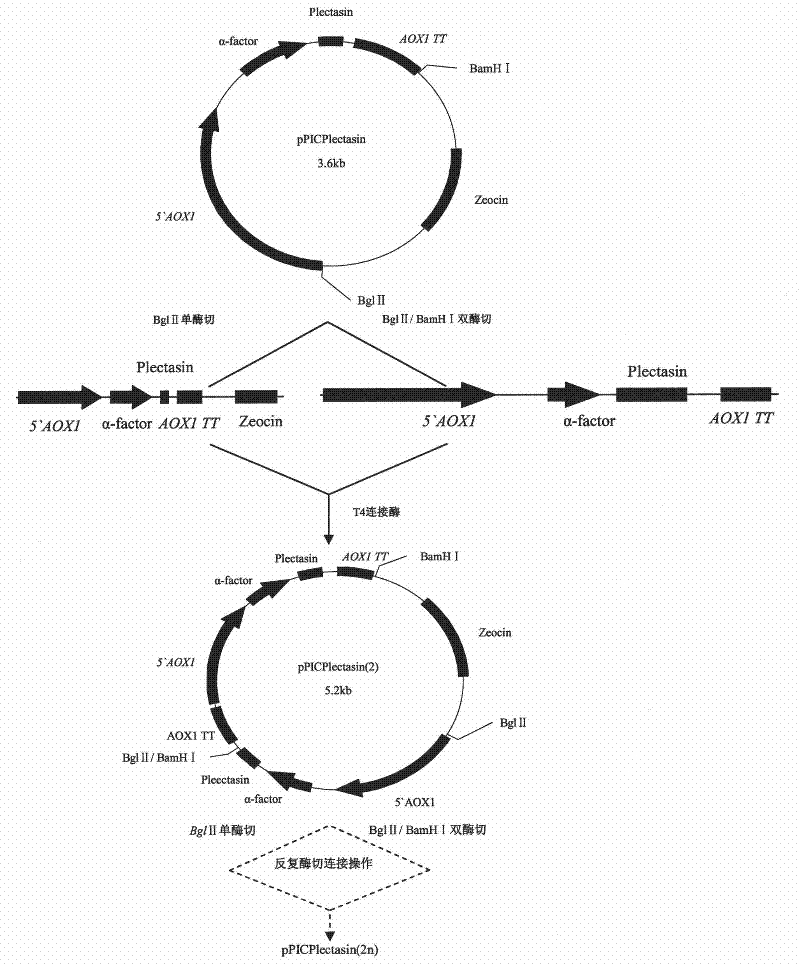
Multi-copy high expressed recombined plectasin by pichia pastoris
ActiveCN102409003AIncrease expression abundanceFungiAntibody mimetics/scaffoldsPichia pastorisPlectasin
The invention discloses a preparation method of multi-copy high expressed recombined plectasin by pichia pastoris. The method comprises the following steps: a plectasin expressing gene sequence is designed according to preference performance to codon translated by pichia pastoris; the optimized plectasin gene is fused on an alpha-factor signal peptide C terminus of an expression vector pPICZalphaA to construct a single-copy expression vector, the vector comprises a plectasin expression cassette containing a start signal element alcohol oxygen dehydrogenase strong promoter (AOX), alpha-factor signal peptide gene and a plectasin gene fused in C terminus, a stop signal element AOX (TT) and the like. A complementation principle of restriction endonuclease Bg1II and BamHI cohesive end is used to obtain plectasin gene-containing recombinant plasmid of different copy cascade expression cassettes, pichia pastoris is electrotransformed and secreted and expressed plectasin with high efficiency under the methanol induction. The expression level and plectasin gene copy number exist a linear relation. The constructed multi-copy high expressed yeast cells can be used for raising the output and reducing the cost, and is adapted to large scale production of plectasin.
Owner:FEED RESEARCH INSTITUTE CHINESE ACADEMY OF AGRICULTURAL SCIENCES
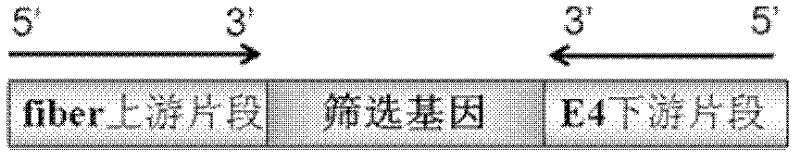
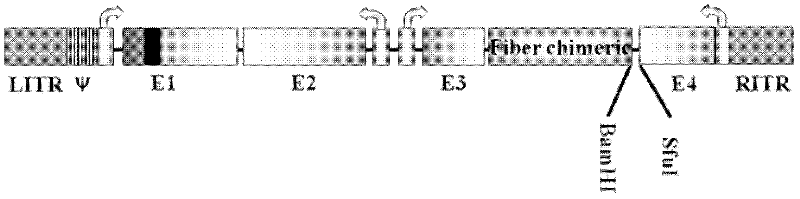
D24 fiber protein modified conditionally replicating adenovirus carrier with exogenous gene by one-step method and application of carrier
InactiveCN102586327AGood treatment effectGenetic material ingredientsFermentationFiberEnzyme digestion
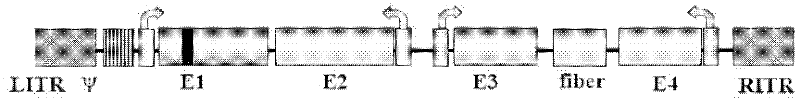
The invention discloses a D24 fiber protein modified conditionally replicating adenovirus carrier with an exogenous gene by a one-step method and an application of the carrier. on the basis of an Ad56 D24 conditionally replicating adenovirus carrier, the construction efficiency of the carrier for inserting into the exogenous gene can be improved by two aspects: (1) a BamHi locus on an adenovirus gene group is mutated; and (2) BamHi and SfuI are introduced between fiber and E4 by a homologous recombination method so as to introduce the exogenous gene into the adenovirus gene group in one step by an enzyme digestion ligation method. The obtained D24 fiber protein modified conditionally replicating adenovirus carrier with an exogenous gene is subjected to the pesticide effect experiment to prove that the D24 fiber protein modified conditionally replicating adenovirus carrier with an exogenous gene can improve a treatment effect on turmor.
Owner:SHAANXI NORMAL UNIV
Preparation method of HRPII protein monoclonal antibody of plasmodium falciparum
ActiveCN101659975AGood repeatabilityAchieve serial expressionMicroorganism based processesFermentationChemical synthesisEscherichia coli
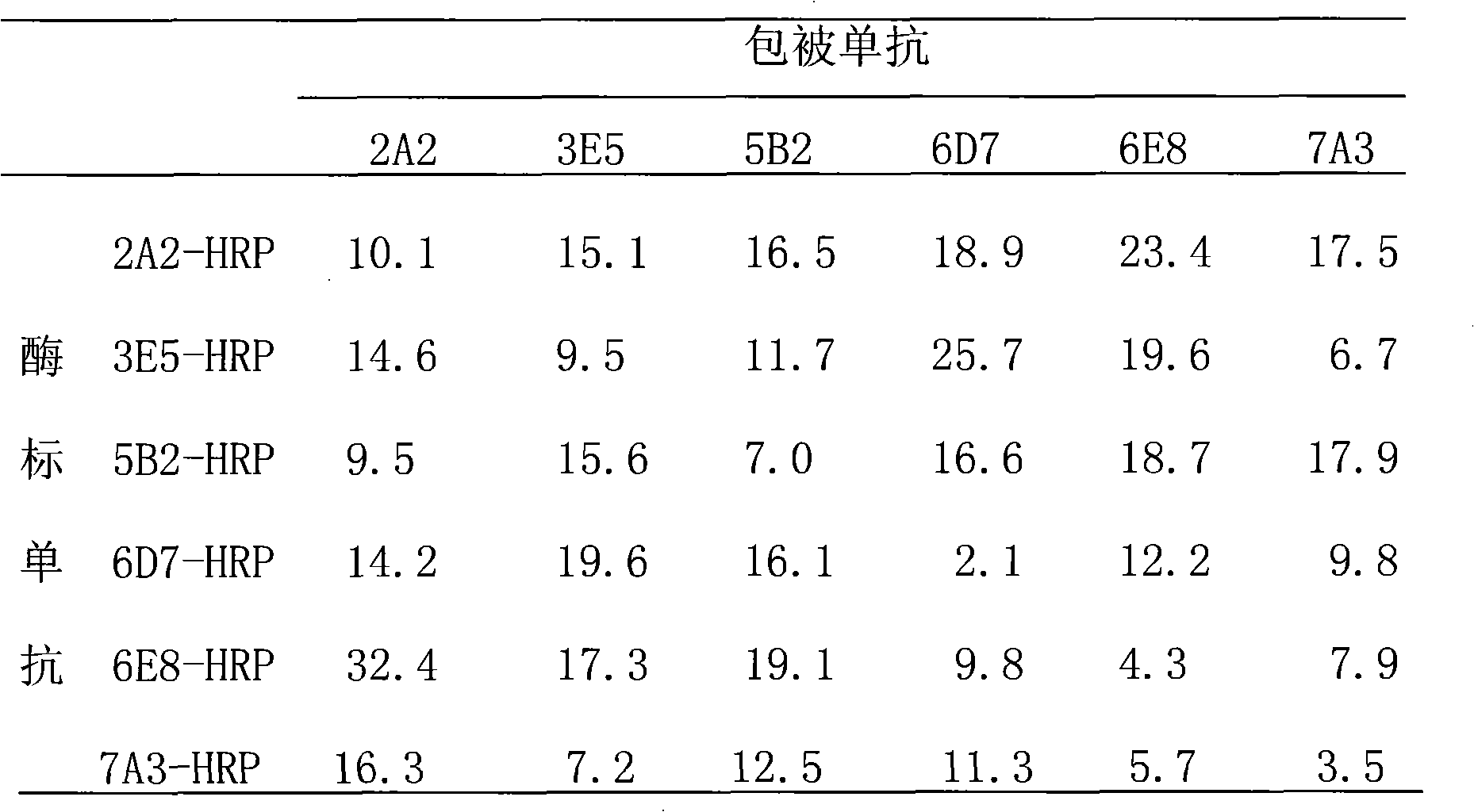
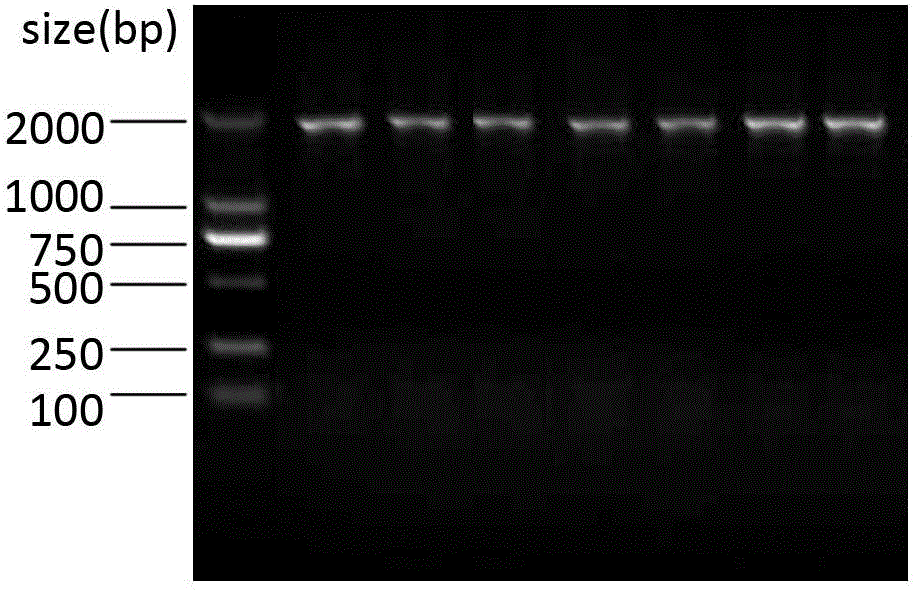
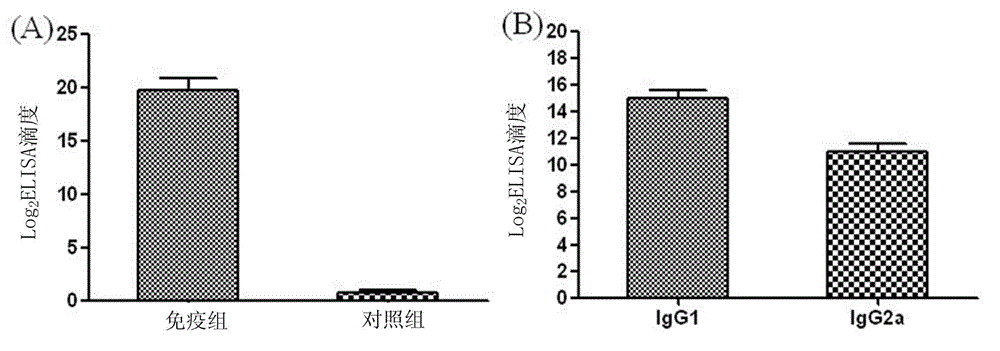
The invention relates to a preparation method of HRPII protein monoclonal antibody of plasmodium falciparum. The preparation method comprises the following steps of: adopting HRPII protein of plasmodium falciparum as target antigen and respectively analyzing and selecting two dominant antigen epitopes of A and B; respectively repeating the two dominant antigen epitopes of A and B, then continuously connecting four glycine and forming recombinant protein C; adopting most securest code of escherichia coli and converting the amino acid sequence of the recombinant protein C into corresponding nucleotide sequence; carrying out chemical synthesis to the former step to obtain the nucleotide sequence, and respectively adding enzyme cutting sites BamHI and EcoRI at the upstream and downstream thereof; inserting nucleotide fragment obtained by the former step into expression carrier PET-28a(+), constructing recombinant protein C expression carrier and inducing to express the recombinant proteinC in the escherichia coli BL21 (DE3); carrying out ultrasonic bacteria breaking and low-temperature centrifugation, then taking supernatant of the solution, affining a chromatographic column by nickel-agarose, eluting and obtaining purified recombinant protein C; after immunizing Balb / c mouse with the recombinant protein C for a plurality of times, taking and fusing spleen cells with sp2 / 0 myelomacells, and obtaining six hybridoma cell lines by multiple rounds of screening; and purifying monoclonal antibody, respectively marking horse radish peroxidase and prorating matching and combination of optimum monoclonal antibody by ELISA orthogonal experiment.
Owner:杭州新脉生物科技有限公司
Streptococcus protective antigen C5a and preparation method thereof
InactiveCN102746388AStrong immune responseSignificant passive immune protectionAntibacterial agentsBacterial antigen ingredientsForward primerProtective antigen
The present invention relates to a streptococcus antigen C5a and a preparation method thereof. The streptococcus protective antigen C5a is an SEZC5a recombinant protein, which consists of 571 amino acids and has a molecular weight of 60.3kDa; a forward primer has one BamHI restriction enzyme cutting site, and a reverse primer has one EcoRI restriction enzyme cutting site. According to the preparation method of the streptococcus protective antigen C5a, the SEZ C5a recombinant protein is treated with cloning, expression and purification; and a series of biological engineering technologies and experiments on mice are applied to conduct system analysis on an rSCPZ. After vaccination, the rSCPZ can provide high protective efficacy; an anti-rSCPZ mice double-immunized serum has significant passive immune protection on mice; and the mice immunized by the rSCPZ show high level of antibody titer in serum. The anti-rSCPZ antibody can induce high level of bactericidal capability; an scpZ gene has a transcription level in SEZ (Streptococcus zooepidemicus) infected mice higher than that of culture in vitro; and the rSCPZ can adhere to hep-2 cells and inhibit cell infection ability of SEZ.
Owner:广东艾佩克科技有限公司
Plasmid for expressing plutella xylostella arginine kinase genes dsRNA (double-stranded ribonucleic acid) and application
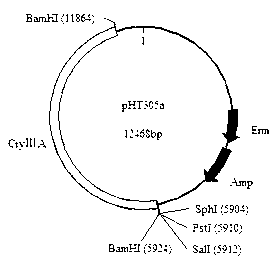
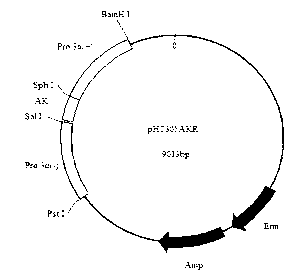
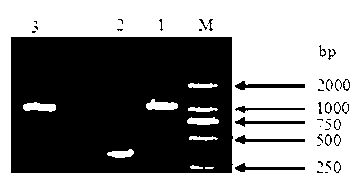
The invention relates to a plasmid for expressing plutella xylostella arginine kinase genes dsRNA (double-stranded ribonucleic acid) and an application, belonging to the field of gene engineering. The plasmid pHT305AKR is a recombinant plasmid obtained by inserting deoxyribonucleic acid (DNA) shown by SEQ ID NO.1 between BamHI and PstI enzyme cutting sites of the plasmid pHT305a. The plasmid pHT305AKR not only realizes the research of expressing dsRNA in bacillus thuringiensis, but also provides a powerful means for applying RNAi (RNA interference) technique to identifying the function of bacillus thuringiensis genes and researching a functional genome of the bacillus thuringiensis, and can also be used for synthesizing dsRNA by utilizing host bacterium, so that the cost for obtaining the dsRNA is reduced. The invention also provides a direction for exogenous transformation of the bacillus thuringiensis and research of bacillus thuringiensis biopesticide, so that the plasmid has a promising application prospect. Meanwhile, a novel way is provided for controlling the plutella xylostella, and a new field is created for controlling the pests.
Owner:FUJIAN AGRI & FORESTRY UNIV
Escherichia coli and preparation method thereof
InactiveCN104004698AGood immune effectBroad prospects for industrial developmentBacteriaMicroorganism based processesEscherichia coliMicroorganism
The invention belongs to microbial strains and preparation methods of the microbial strains, and particularly relates to Escherichia coli and a preparation method of the Escherichia coli. A cloned iss gene segment in outer membrane protein of Escherichia coli pathogenic bacteria of a coded bird is regrouped in an expression plasmid pEGX-6p-1 after BamHI and SalI double enzyme digestion, and a pronucleus expression plasmid pEGX-6p-1 / iss is built; the pronucleus expression plasmid pEGX-6p-1 / iss is transformed into Escherichia coli BL21-DE3, and regrouped Escherichia coli BL21-DE3 / pEGX-6p-1 / iss is built. The technical problem that the vaccine broad-spectrum performance is poor in the prior art is solved. The Escherichia coli has the advantages of being good in broad spectrum performance, good in immune effect, wide in industrialized development prospect and the like.
Owner:HEBEI KEXING PHARMA
Starch induction type recombinant bacillus subtilis as well as preparation method and application thereof
ActiveCN105039374AHigh expressionNo toxicityBacteriaMicroorganism based processesSequence signalRestriction Enzyme Cut Site
The invention relates to starch induction type recombinant bacillus subtilis as well as a preparation method and application thereof. The starch induction type recombinant bacillus subtilis contains a recombinant vector, wherein the recombinant vector is characterized in that in front of a BamHI restriction enzyme cutting site of a PHT43 plasmid, a Pgrac promoter is replaced with an alpha-amylase promoter PamyQ by performing for three times a mode of overlapping PCR continuously, and then an MTSase-MTHase fusion enzyme gene is inserted behind the BamHI restriction enzyme cutting site. When the used alpha-amylase promoter and an amylase signal peptide are combined with the expression gene, i.e., the MTSase-MTHase fusion enzyme gene, the expression effect of the starch induction type is better than that of other induction types.
Owner:QILU UNIV OF TECH
Chrysanthemum stress-resistance transcription factor CmMYB2 as well as plant expression vector, construction method and application thereof
InactiveCN102161697AImprove stress resistanceImprove salt tolerancePlant peptidesFermentationBiotechnologyEnzyme digestion
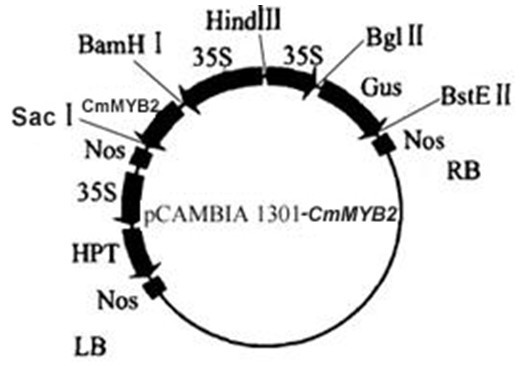
The invention discloses a chrysanthemum stress-resistance transcription factor CmMYB2 as well as a plant expression vector, a construction method and an application thereof, belonging to the field of molecular biology. The expression vector pCAMBIA1301-CmMYB2 is constructed with the construction method which comprises the following steps: inserting the CmMYB2 gene into a colon vector pMD19-Tsimplevector (Takara); and after carrying out enzyme digestion on the CmMYB2 gene by BamHI (Takara) and SacI (Takara), connecting to the BamHI locus and SacI locus of an expression vector pCAMBIA1301. The plant expression vector is used for the genetic transformation of plants, the CmMYB2 gene is subjected to over expression under the starting of CaMV35S to synthesize a great quantity of MYB2 proteins, regulate the expression of a downstream target gene and improve plant stress resistance.
Owner:NANJING AGRICULTURAL UNIVERSITY
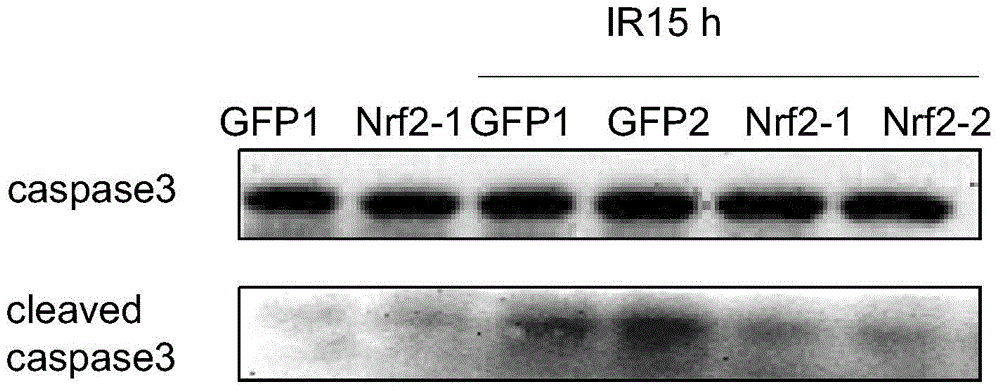
Cell strain MSCs for overexpression of Nrf2 gene as well as preparation method and application of cell strain MSCs
InactiveCN104877967AStrong anti-apoptotic propertiesMicrobiological testing/measurementUnknown materialsEnzyme digestionHuc mscs
The invention discloses a cell strain MSCs for overexpression of an Nrf2 gene as well as a preparation method and application of the cell strain MSCs. The preparation method comprises the following steps: performing PCR amplification on a human-derived Nrf2 gene ORF; performing purification and double enzyme digestion on the PCR product obtained in the step (1), then connecting the PCR product with the framework plasmid pLV-CMV-XbaI-BamHI-GFP to construct an Nrf2 slow virus recombinant vector, and after conversion, selecting clone and determining positive clone; sequencing the positive clone, and after no mutation is determined, performing amplification massively; leading the recombinant vector and an auxiliary plasmid in a 293 FT cell to obtain a virus; using the virus to infect the 3-5 th generation hUC-MSCs cells so as to obtain the cell strain MSCs for overexpression of the Nrf2 gene. According to the invention, the overexpression of Nrf2 in MSCs is stably performed to increase the cell activity, and a higher anti-apoptosis feature is also realized under the anoxic and oxidative stress conditions; Nrf2-MSCs with the treatment level amount is obtained in vitro and can serve as an excellent tool cell for preclinical study of MSCs and can be used for preparing a medicine for enhancing preclinical study of MSCs transplanting.
Owner:THE THIRD AFFILIATED HOSPITAL OF SUN YAT SEN UNIV
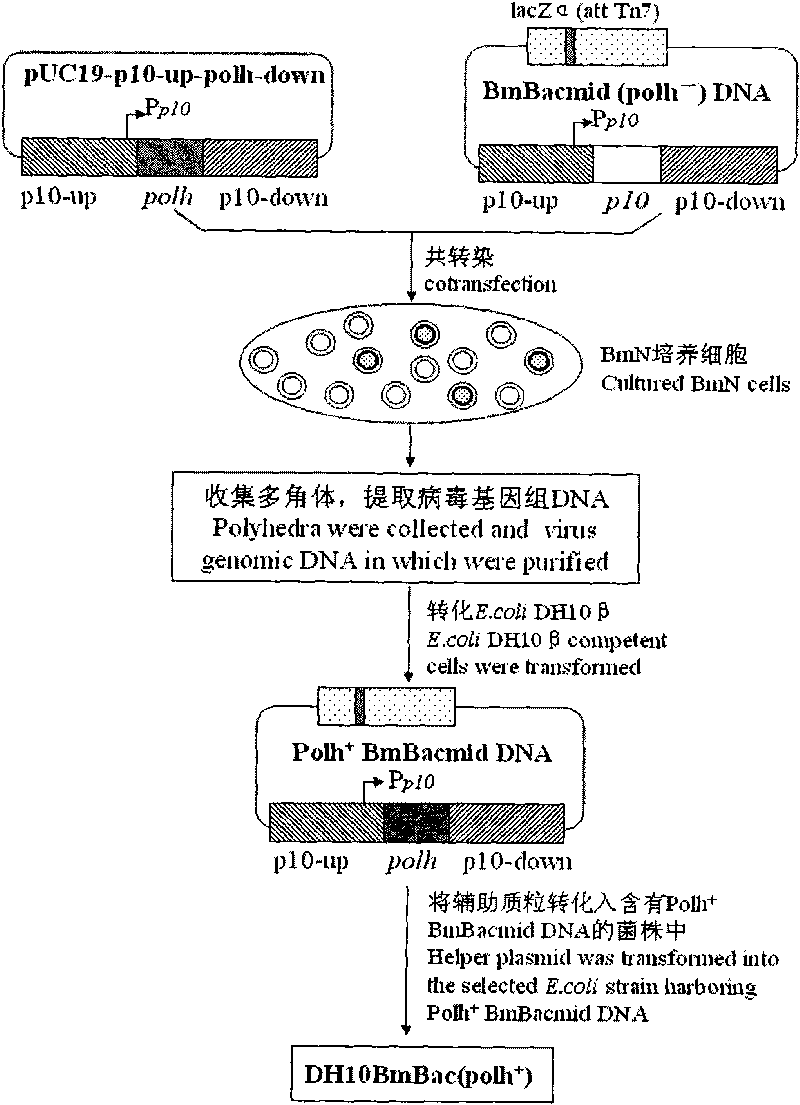
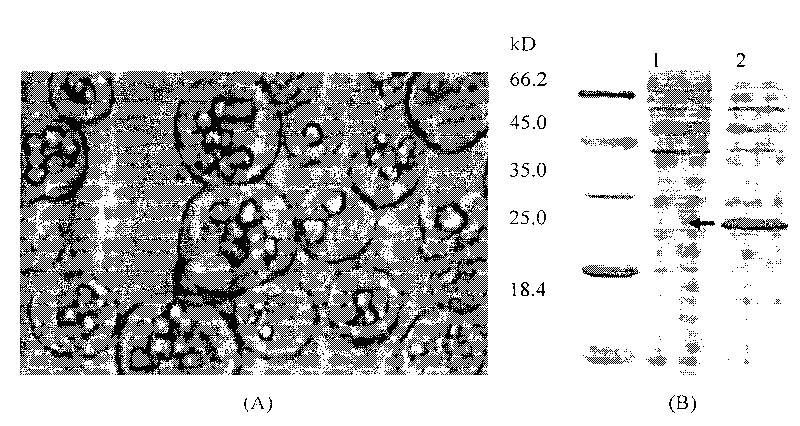
Building method of silkworm BmNPV Polh+Bac-to-Bac rhabdovirus expression system
The invention discloses a building method of a silkworm BmNPV Polh+Bac-to-Bac rhabdovirus expression system, comprising: taking silkworm wild type BmNPV genome DNA as the template; respectively taking P10-upF / P10-upB and P10-downF / P10-down B as a primer PCR for amplification to obtain p10-up and p10-down; after processed, inserting into BamHI-PstI-HindIII locus in pUC 19 to obtain recombinant plasmid pUC19-p10-up-down; using pUC-19-p10-up-polh-down, liposome lipofectin and MilliQ H2O to prepare DNA-Lipofectin mixed liquor; adding the mixed liquor into BmN cells for cultivating; collecting transfection cell supernatant; inoculating BmN cells, and recovering a polyhedral body; extracting virus DNA from the polyhedral body and electrically converting DH10 beta competent cells; screening locus ceruleus and cultivating; after cultivating PCR positive bacterial plaque, extracting macro-molecular DNA to transfect to the BmN cells; separating to obtain helper plasmids from DH10Bac culture bacteria of AcMNPV Bac-to-Bac; converting the helper plasmids into E.coli DH10 beta containg Ploh+BmBacmid; and screening the DH10 beta bacterial strain of the helper plasmid containg Ploh+BmBacmid. The invention can produce recombinant virus capable of infecting by eating with mouth, and recombinant virus does not need to infect by intracutaneous inoculation, thus improving the production efficiency of the silkworm rhabdovirus expression system.
Owner:ZHEJIANG UNIV
Method for constructing laryngeal cancer cell line for stably expressing green fluorescent protein
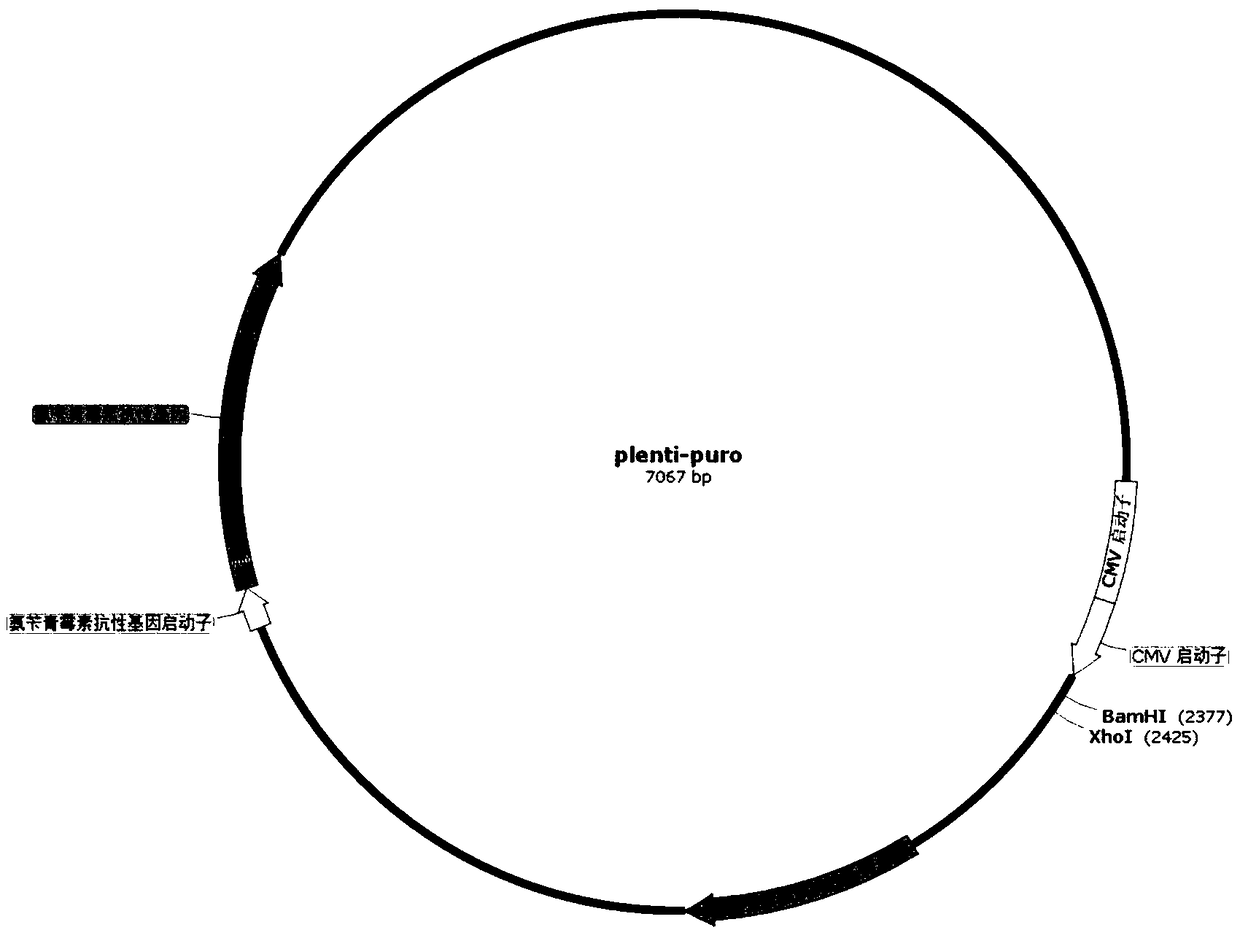
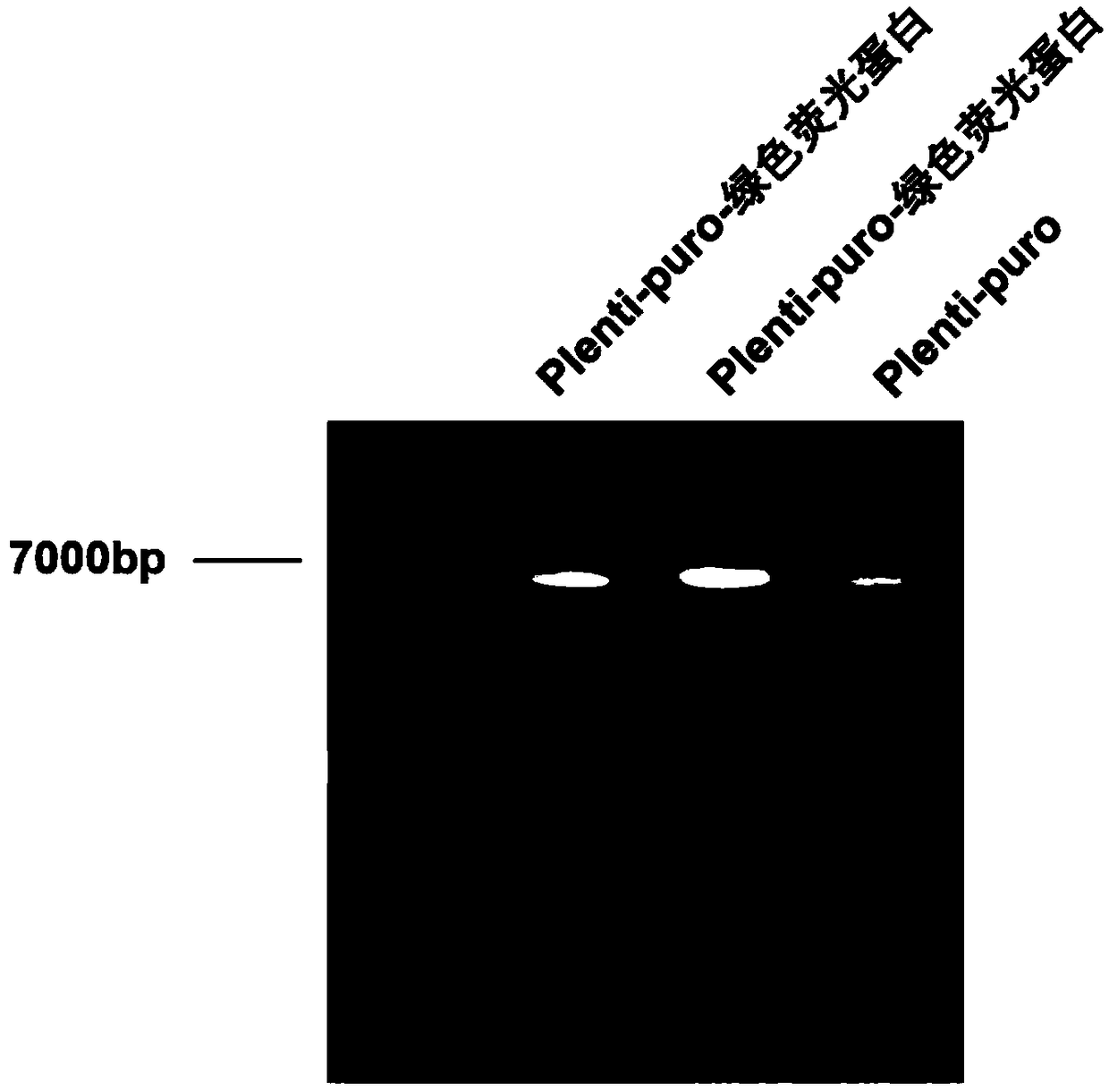
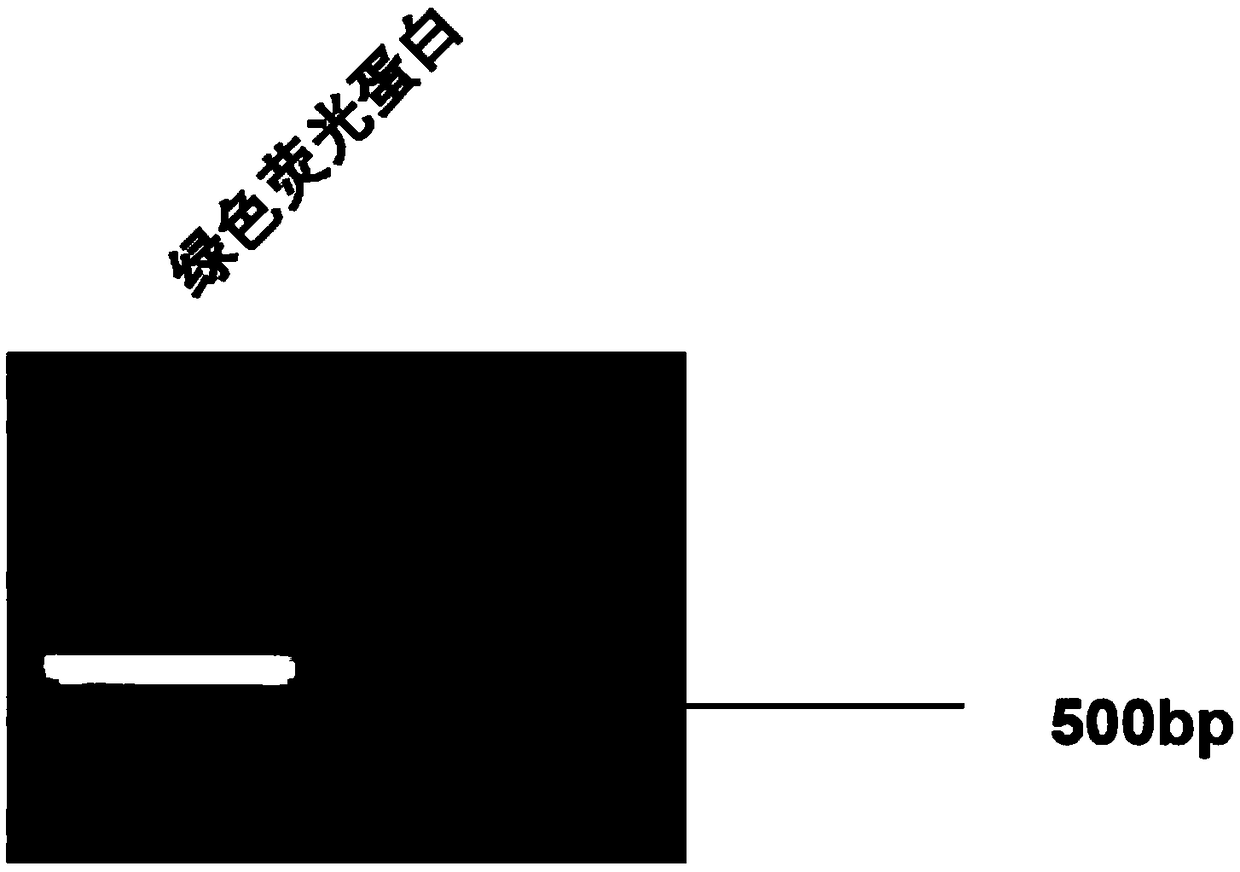
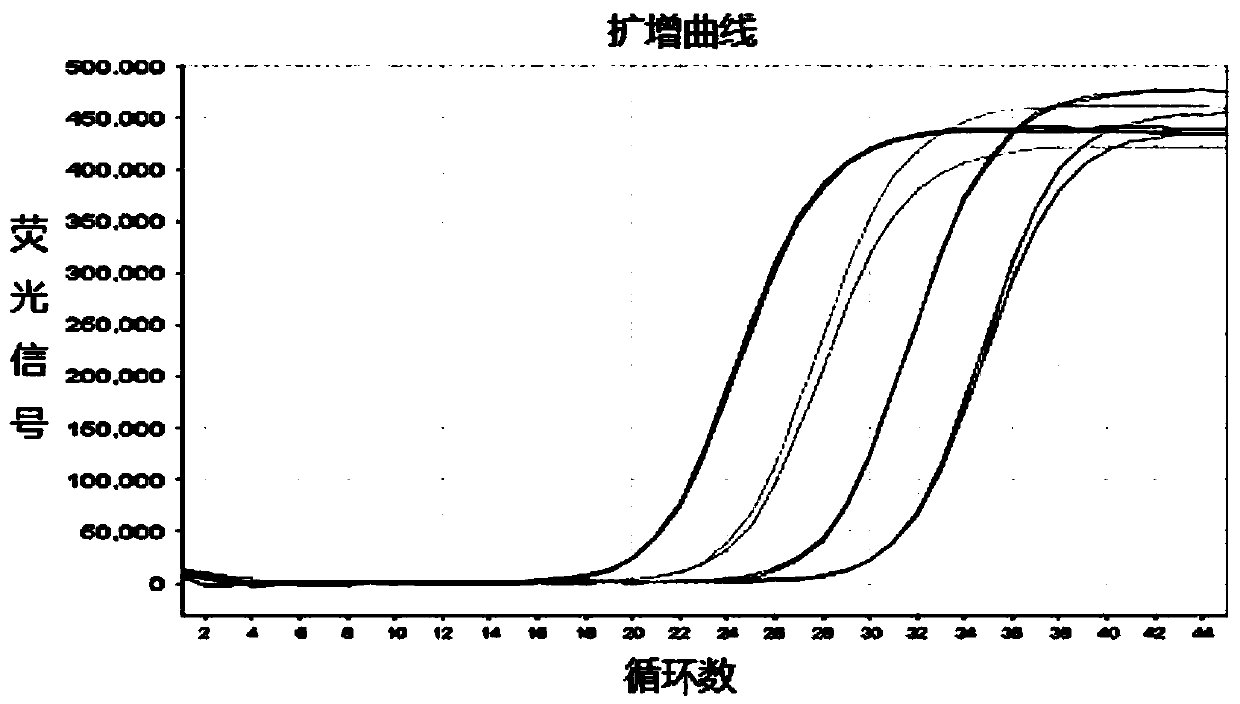
InactiveCN108949697AShort screening cycleMeet different throughput needsGenetically modified cellsPeptidesProtein CBamHI
The invention belongs to the technical field of cell research, and particularly relates to a method for constructing a laryngeal cancer cell line for stably expressing the green fluorescent protein. The method comprises the following steps: (1) designing and synthesizing a primer; (2) digesting the gene sequence of pLenti-puro carrier plasmids with BamHI and XhoI according to the gene sequence andpLenti-puro sequence of the green fluorescent protein, and connecting digested products to obtain a target carrier pLenti-puro-GFP; (3) verifying the carrier pLenti-puro-GFP with endotoxin extraction, digesting the pLenti-puro-GFP plasmids with BamHI and XhoI, and detecting the gene sequence by agarose gel electrophoresis, and sequencing the carrier pLenti-puro-GFP plasmids to detect the sequencesituation of the GFP-LC3; and (4) co-transfecting a 293T cell with the carrier pLenti-puro-GFP plasmids and a lentivirus package carrier, collecting virus supernatant, and transfecting a laryngeal cancer cell Hep-2 with the virus supernatant, infecting for 24 hours, then adding a culture medium containing puromycin, screening the cell, and observing under high content until a stable strain is obtained.
Owner:FIRST HOSPITAL OF SHANXI MEDICAL UNIV
Preparation method of artificial dual false virus particle comprising HCV (hepatitis C virus) and HIV (human immunodeficiency virus) nucleic acid fragments
InactiveCN103789277AHigh clone purityImprove stabilityInactivation/attenuationFermentationHuman immunodeficiencyCompanion animal
Owner:东北制药集团辽宁生物医药有限公司
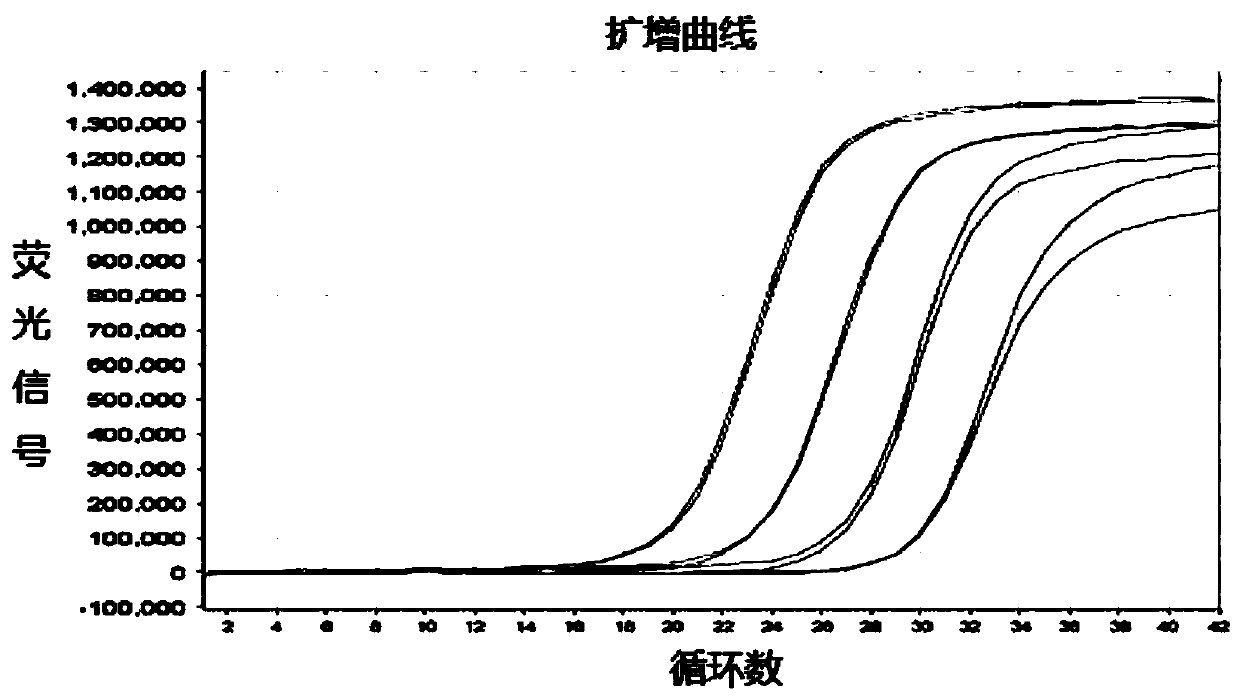
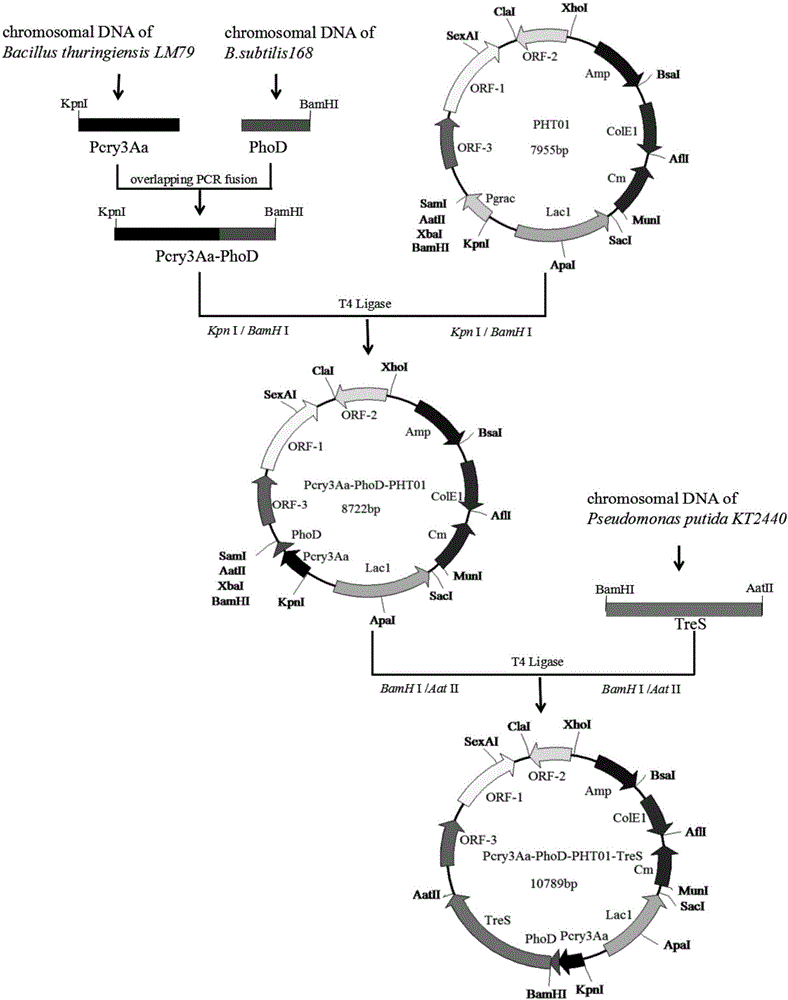
Method for constructing highly expressed trehalose synthase engineering bacteria by using Pcry3Aa promoter
ActiveCN105779489AGood synthesis effectImprove stabilityBacteriaTransferasesSequence signalProtein target
The invention relates to a method for constructing highly expressed trehalose synthase engineering bacteria by using a Pcry3Aa promoter. For a recombinant carrier, a Pcry3Aa-PhoD fragment by which a Pcry3Aa promoter fragment and a PhoD signal peptide fragment are connected by an overlap PCR (Polymerase Chain Reaction) is inserted at the upstream of a restriction enzyme cutting site BamHI of a shuttle plasmid PHT01, and a target protein trehalose synthase TreS fragment is inserted between two restriction enzyme cutting site, i.e., BamHI and AatII. The invention further relates to a method for constructing the highly expressed trehalose synthase engineering bacteria by using the recombinant carrier. According to the method disclosed by the invention, the Pcry3Aa promoter is adopted to naturally induce the synthesis of trehalose synthase; because the Pcry3Aa promoter contains a special STAB-SD structure, the stability of the Pcry3Aa promoter to transcribe mRNA is improved, the half-life period of mRNA is prolonged, the mRNA translation level of a downstream target gene is improved, and therefore the trehalose synthase is highly expressed.
Owner:山东开盾生物科技有限公司
Double-LAMP (loop-mediated isothermal amplification) method for simultaneously detecting vibrio parahaemolyticus and vibrio vulnificus
ActiveCN105219845AMicrobiological testing/measurementMicroorganism based processesMetalloprotease GeneLoop-mediated isothermal amplification
The invention discloses a double-LAMP (loop-mediated isothermal amplification) detecting method, and belongs to the technical field of molecular biology. By the aid of the method, vibrio parahaemolyticus and vibrio vulnificus can be simultaneously detected. An LAMP detection system comprises primer groups for detecting vibrio parahaemolyticus genes OmpA and vibrio vulnificus metalloprotease genes, and each primer group contains a pair of outer primers F3 and B3 for the OmpA genes and the metalloprotease genes and a pair of inner primers FIP and BIP; PstI and BamHI restriction enzyme cutting sites are respectively arranged on the inner primers BIP. The double-LAMP method has the advantages of high sensitivity and detection speeds, good specificity, simplicity in operation and visual and clear result observation.
Owner:QINGDAO AGRI UNIV
Construction method for sandwiched antibody chip detection system based on single chain antibody fusion protein
InactiveCN1866022AGuaranteed specificityMaintain affinityBiological testingFluorescence/phosphorescenceSingle-Chain AntibodiesElisa method
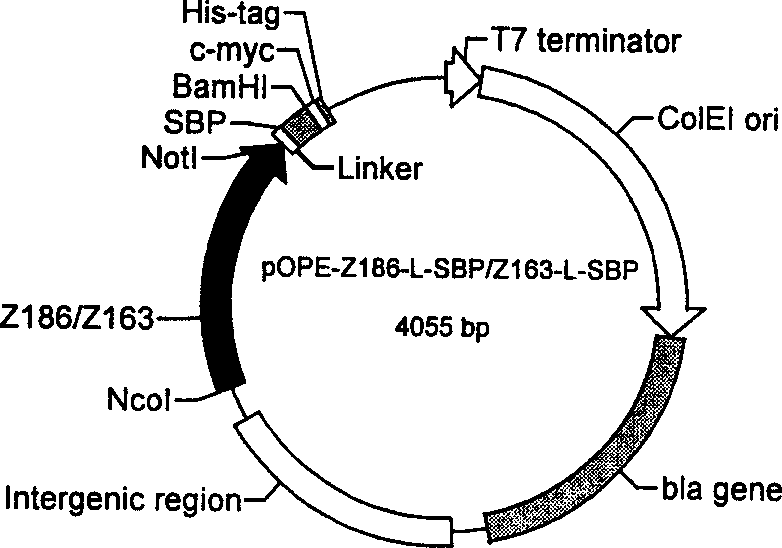
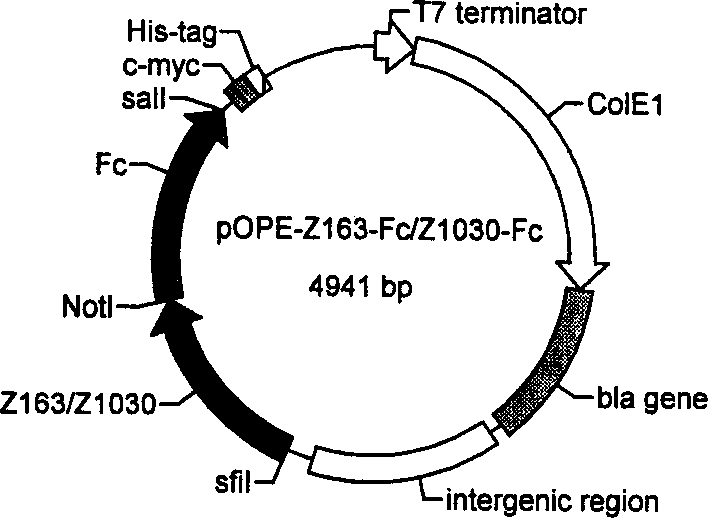
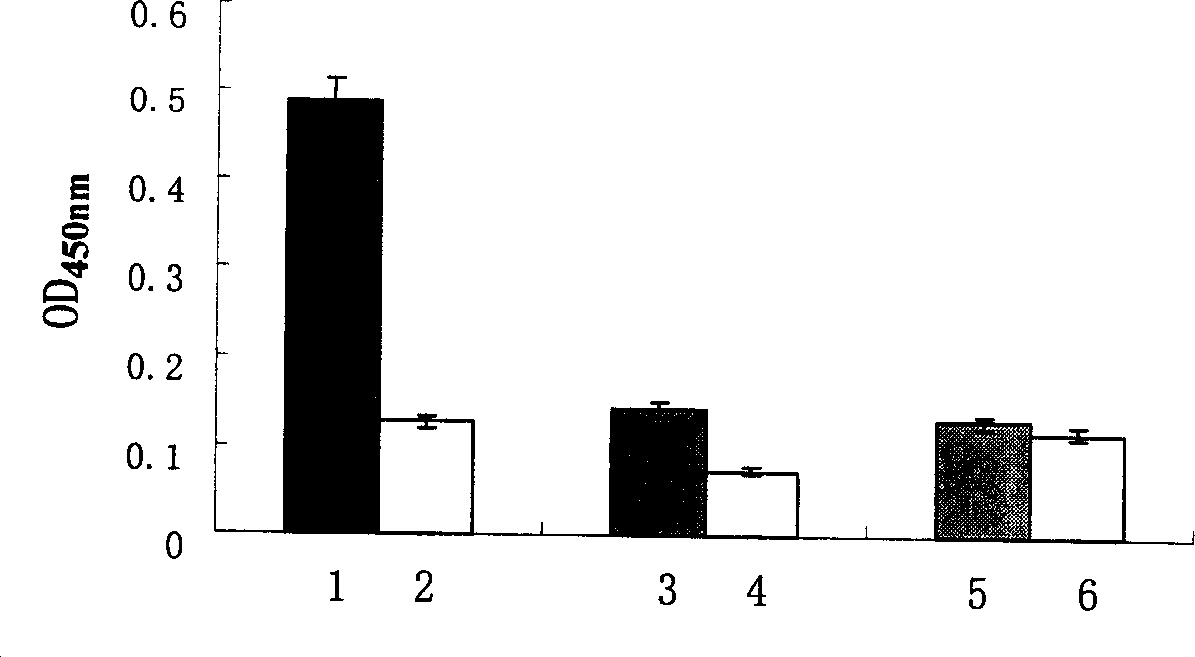
The disclosed construction method for sandwich antibody chip detection system based on single-chain anti-body fused protein comprises: constructing the express carrier for Z186-L-SBP and Z163-L-SBP, designing upper and downstream primer, connecting double-chain ends as NotI and BamHI limited enzyme sites to the product; constructing the express carrier for Z163-Fc, and screening the single-chain antibody fused protein by ELISA method. This invention has wide application.
Owner:WUHAN INST OF VIROLOGY CHINESE ACADEMY OF SCI
Primer, kit and detection method for detecting chicken green shin character linkage SNP locus genotype
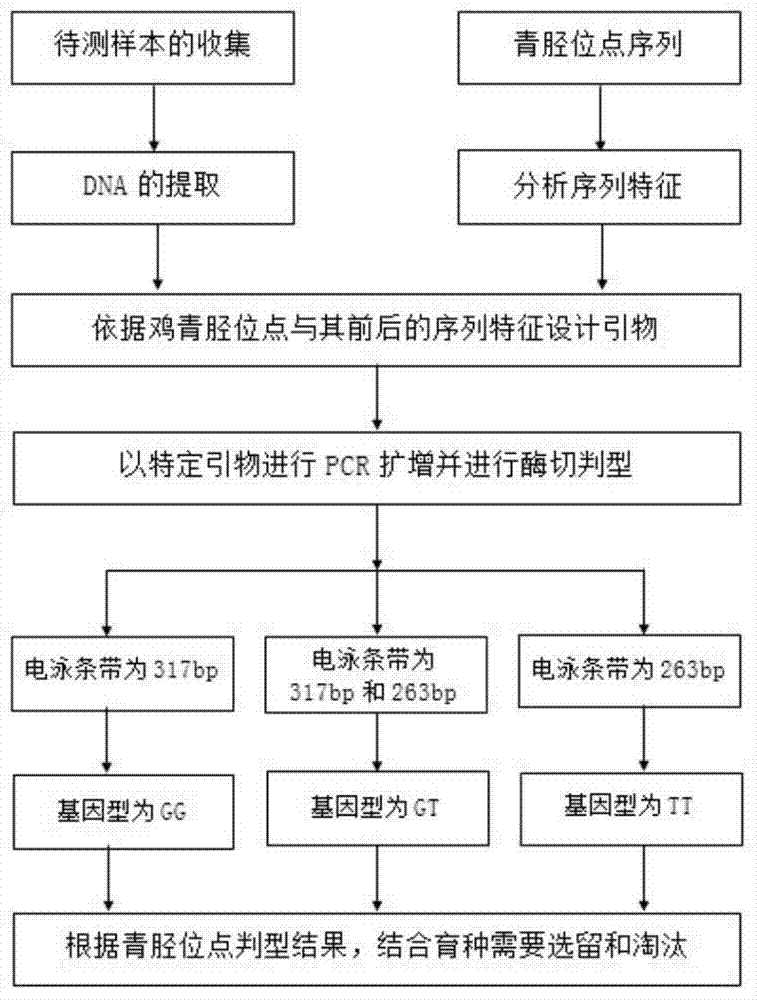
ActiveCN104293905ASimple and fast operationLow costMicrobiological testing/measurementDNA/RNA fragmentationMarker-assisted selectionEnzyme digestion
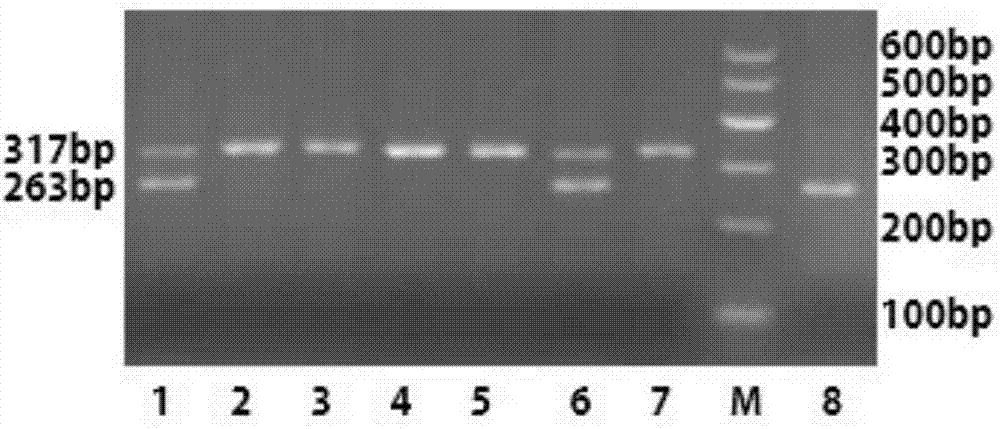
The invention discloses a primer, a kit and a detection method for detecting chicken green shin character linkage SNP locus genotype, and belongs to the technical field of biological detection. Aiming at an SNP locus specific for the chicken green shin character, a primer pair (P) is designed at DNA segment near the SNP locus; and BamHI enzyme digestion is performed on an amplified product after PCR amplification. If the mutation site is G and a BamHI enzyme digestion sequence GGATCC presents, the segment of the PCR product after enzyme digestion is 263bp; and if the mutation site is T and no BamHI enzyme digestion sequence GGATCC presents, the segment of the PCR product after enzyme digestion is 317 bp. Compared with SSCP and a direct sequencing method, the detection method is simple to operate, low in cost and short in cycle, can greatly increase accuracy for determining the SNP locus genotype, is in no need of special instruments, and can be popularized easily. Experiments demonstrate that the method can effectively determine the genotype of the chicken green shin character SNP locus, can be used for marker-assisted selection of the chicken green shin characters, and provide effective molecular marker for establishing green shin chickens.
Owner:HENAN AGRICULTURAL UNIVERSITY
Method for preparing recombined human amyloid A beta 42 and application thereof
InactiveCN101270359AIncrease productionLow costNervous disorderPeptide/protein ingredientsEscherichia coliEukaryotic plasmids
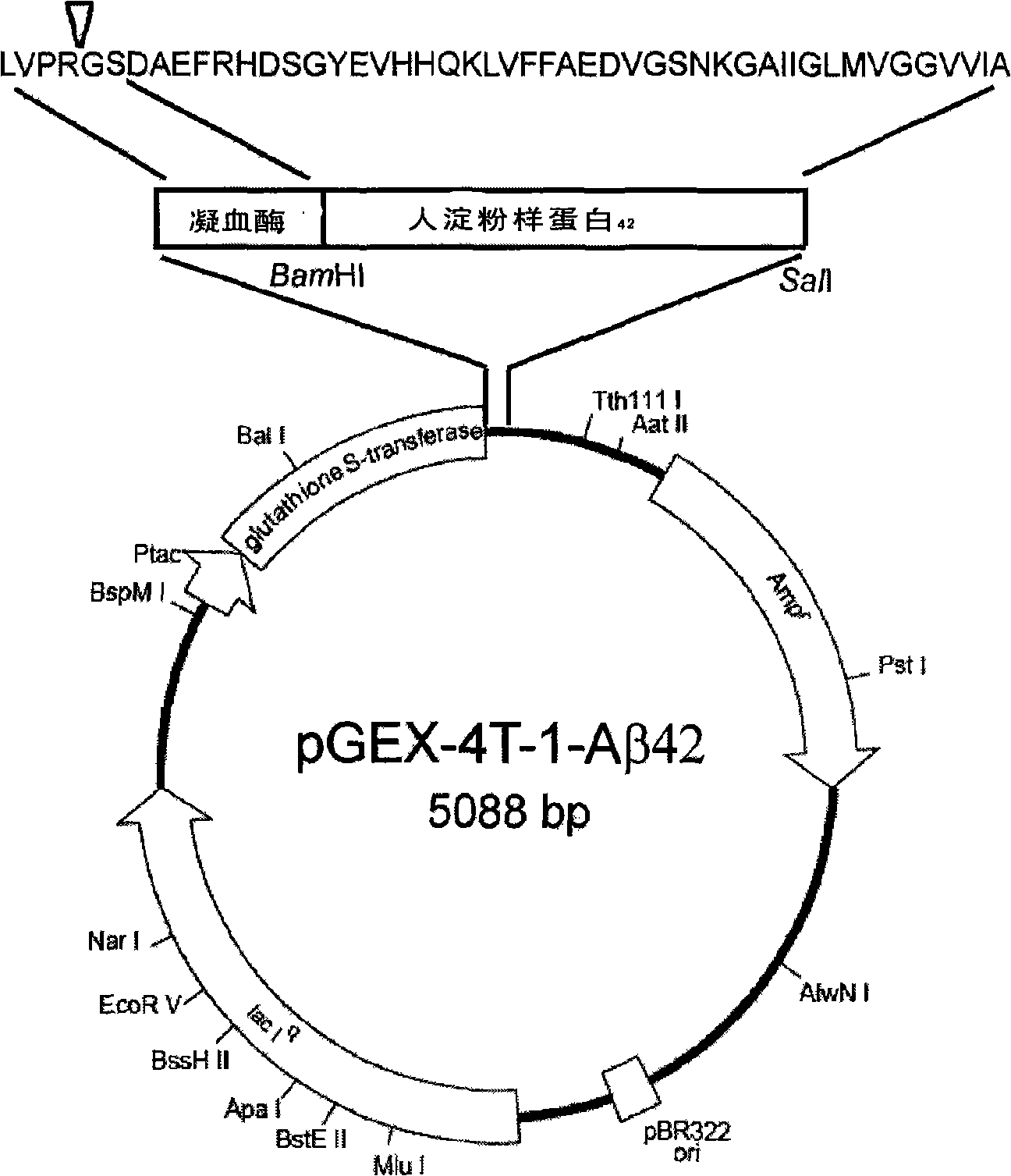
The invention provides a method for preparing recombinant human amyloid protein A Beta42 and application of the recombinant human amyloid protein, which belongs to the technical field of fusion protein. In the method, PCR sense primer P1 of the human amyloid A Beta42 is designed; a BamHI restriction enzyme cutting site, an anti-sense primers P2, a Sall restriction enzyme cutting site and a terminator codon TAG are introduced according to the sequence of the precusor protein APP of the human amyloid and multiple cloning sites of the cloning vector pGEX-4T-1. cDNA of human SH-SY5Y cells is taken as a templet for amplifing PCR; the length of the fragment of the product is 147bp. The Beta42 fragment of human APP from 672 to 713 is encoded. The A Beta42 gene fragment sequence is combined into a prokaryotic expression vector pGEX-4T-1 to form the prokaryotic expression plasmids pGEX-4T-1 / A Beta42 of human A Beta42. PGEX-4T-1 / A Beta42 is transformed into colibacillus BL21 (DE3), and purified activated recombinant human amyloid A Beta42 is got through induced expression, separation and enzyme cutting.
Owner:JIANGSU INST OF NUCLEAR MEDICINE
Bat SARS-like coronavirus spike protein immunity determining area and preparation method and application thereof
InactiveCN102690336AImproving immunogenicitySimple methodDepsipeptidesBiological testingMicrochiropteraEnzyme digestion
The invention discloses a bat SARS-like coronavirus spike protein immunity determining area and a preparation method and application thereof. The method comprises the following steps of: A, amplifying by taking the bat SARS-like coronavirus S gene as a template design primer; B, performing enzyme digestion of the fragment obtained by the amplification by use of BamHI and XhoI, connecting to an expression vector pET32a, and sequencing to assure correctness; and C, purifying the recombinant plasmid, transforming the BL21 competent cell, culturing the monoclonal antibody, and performing 30-degree induction in a culture medium with the final concentration of 0.3mMIPTG; collecting the thalli, performing ultrasonic breaking, and then purifying with a HisTag purification kit; and after the purification, detecting the concentration of protein in SDS-PAGE to obtain the target protein. The method is simple and easy to implement, convenient to operate and easy in production; the peptide has the best immunogenicity; and a method of applying the protein to the identification of mice S monoclonal antibody epitope has high specificity, and is simple to operate and easy to repeat.
Owner:WUHAN INST OF VIROLOGY CHINESE ACADEMY OF SCI
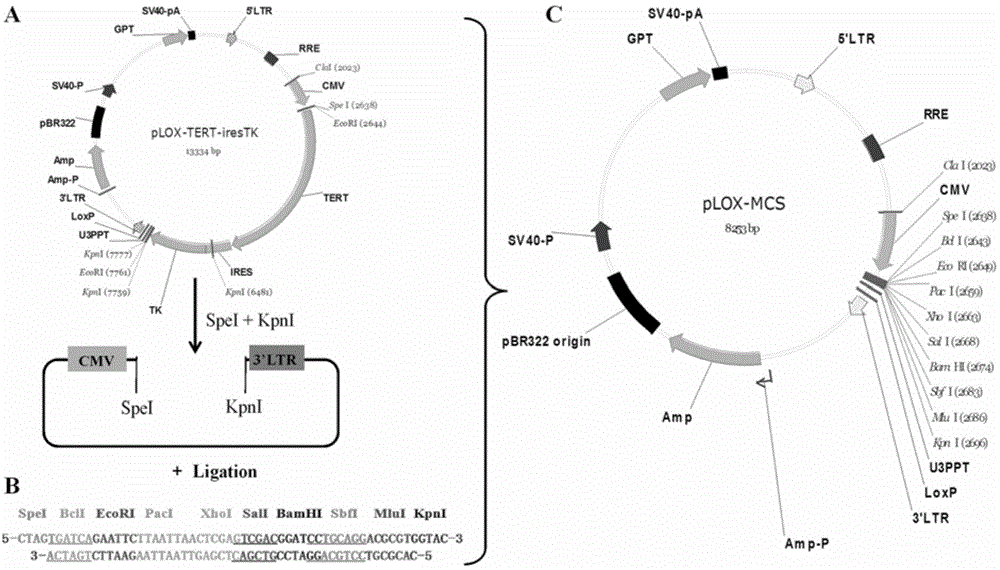
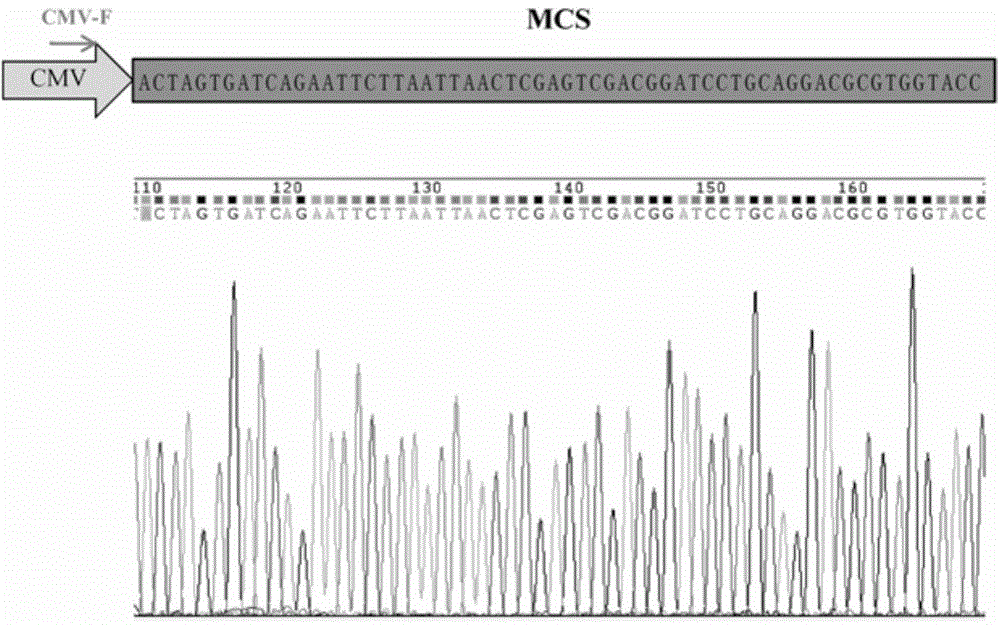
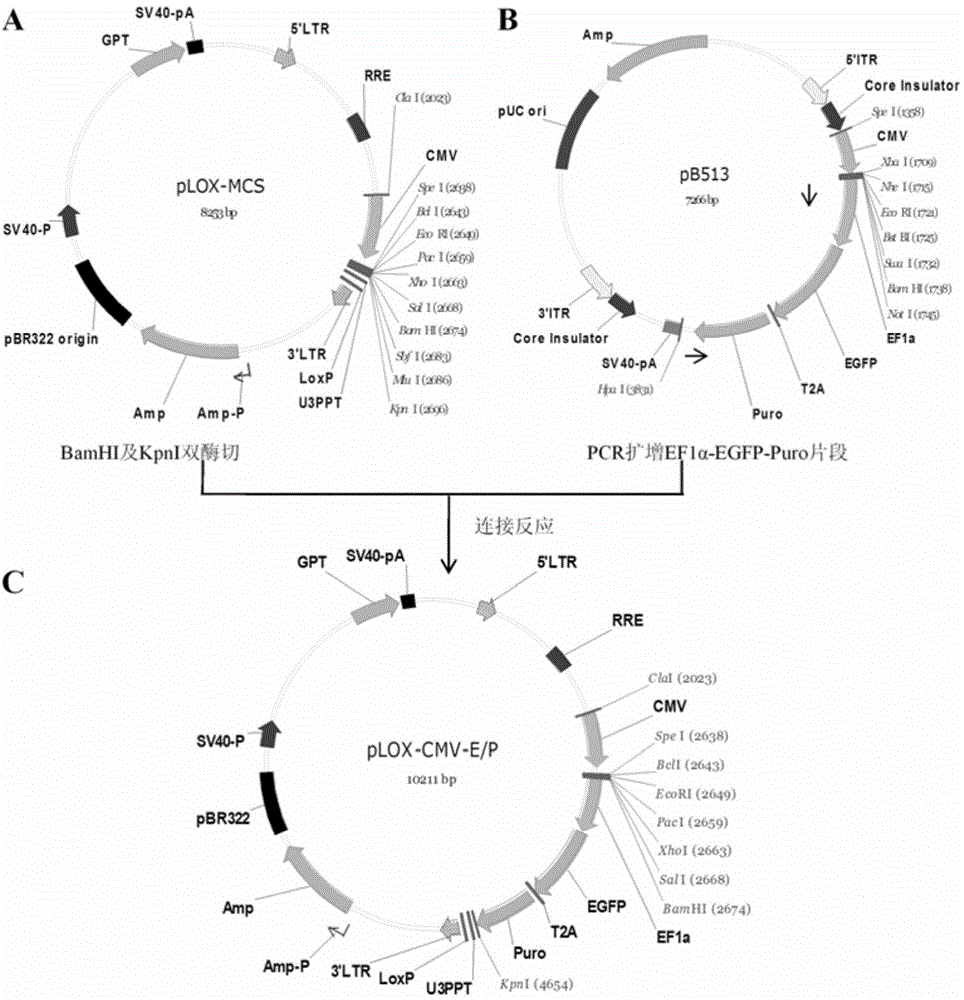
Lentiviral expression vector pLOX-CMV-E/P and construction and application thereof
InactiveCN104372027AStable expressionFermentationVector-based foreign material introductionAgricultural scienceFluorescence
The invention discloses a lentiviral expression vector pLOX-CMV-E / P and construction and application thereof. Multiple cloning site which is inserted into the downstream part of a CMV promoter of the vector can be recognized by 7 restriction endonucleases, including SpeI, BclI, EcoRI, Paci, XhoI, SalI and BamHI. The application of the vector comprises the following steps: inserting a target gene to the multiple cloning site to realize the stable expression of the target gene; using Puromycin resistance gene and EGFP which is driven by EF1alpha promoter to perform fluorescence tracing and drug resistance screening; based on LoxP site inserted in 3'LTR region and the transcription and reverse transcription characteristic of lentivirus, realizing effective deleting of an exogenous gene and a reporter gene under the condition of Cre recombinase expression. The vector disclosed by the invention can realize the package of the lentiviral particles by commonly transfecting 293T cell together with pCMVR8.74 and PMD2.G vector.
Owner:JINAN UNIVERSITY
Recombinant baculovirus transfer vector containing porcine pseudorabies virus gD protein gene, recombinant baculovirus and preparation method and application thereof
ActiveCN109943592AHigh protection rateIncrease production capacityAntiviralsViruses/bacteriophagesSequence signalTransfer vector
The invention provides a recombinant baculovirus transfer vector containing a porcine pseudorabies virus gD protein gene, a recombinant baculovirus and a preparation method and application thereof, and belongs to the technical field of veterinary vaccines. The recombinant baculovirus transfer vector containing porcine pseudorabies virus gD protein gene is prepared by inserting HBM signal peptide-gD-His tagor HBM signal peptide-gD-IgGFc-His tag as an exogenous gene into an enzyme cutting site between BamHI and EcoRI and an enzyme cutting site between SpeI and HindIII of the baculovirus transfervector respectively; and the gD is a gD protein gene deleted from a encoding transmembrane region fragment. The vector promotes the secretory expression of the porcine pseudorabies virus gD protein and the gD-IgGFc fusion protein. The subunit vaccine prepared by using the gD protein and the gD-IgGFc fusion protein has the advantages of good safety, strong specificity and high virus attack protection rate.
Owner:HUAZHONG AGRI UNIV
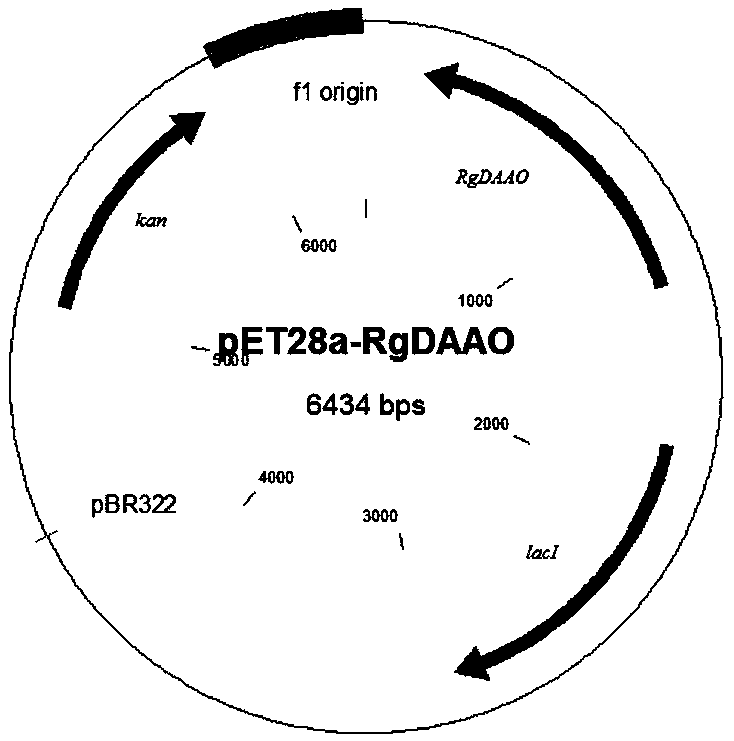
Bacterial strain of producing pyruvic acid and construction method of bacterial strain
The invention discloses a bacterial strain of producing pyruvic acid and a construction method of the bacterial strain. The construction method comprises the following steps: cloning a D-amino acid oxidase gene DAAO (D Amino Acid Oxidase) into the pUC57 plasmids through EcoRI or BamHI to obtain a plasmid pUC57-DAAO; carrying out gene amplification by taking plasmids pUC57-DAAO as a template; cloning a product obtained by gene DAAO amplification to a plasmid pET28A or pET21a; selecting and culturing positive clone plasmids connected to a DNA segment containing DAAO from the plasmids; converting the constructed positive clone plasmids to an expression host bacterium BL21(DE3); and selecting and culturing to obtain a first bacterial strain which can convert D-alanine to pyruvic acid, wherein the fragment size of the DAAO gene is 1124bp, and the gene contains cleavage sites of endonuclease NdeI and EcoRI. The bacterial strain constructed by the construction method can directly convert D-alanine into pyruvic acid through a biotransfer approach.
Owner:HEFEI BAIMAI BIOTECH CO LTD
Swing breeding and respiratory syndrome suicide DNA vaccine and use
InactiveCN1640497AViral antigen ingredientsGenetic material ingredientsMultiple cloning siteRespiratory syndrome virus
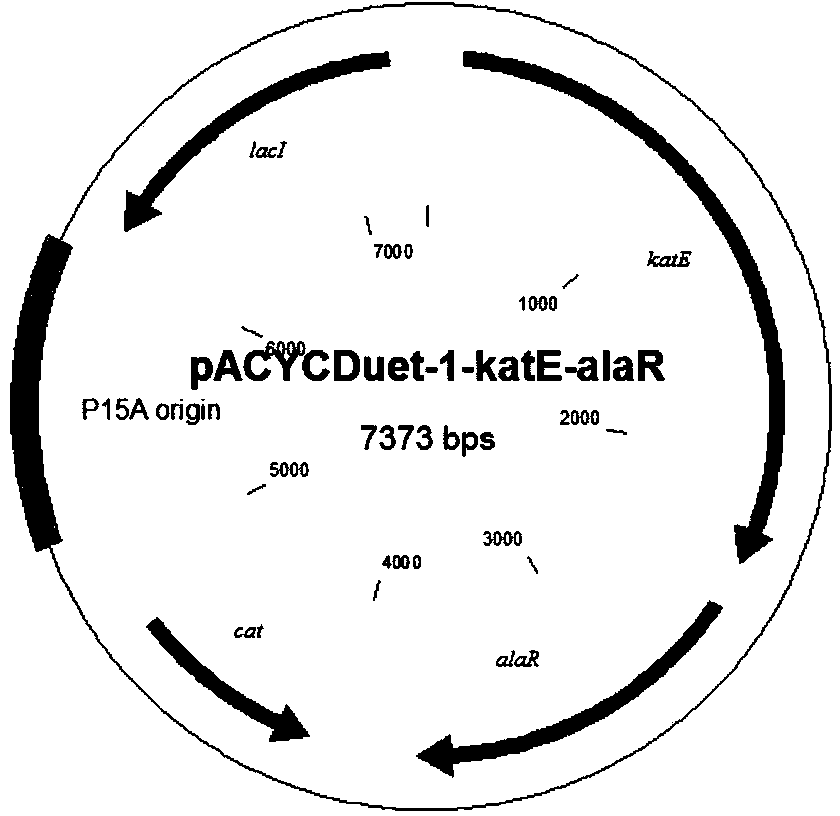
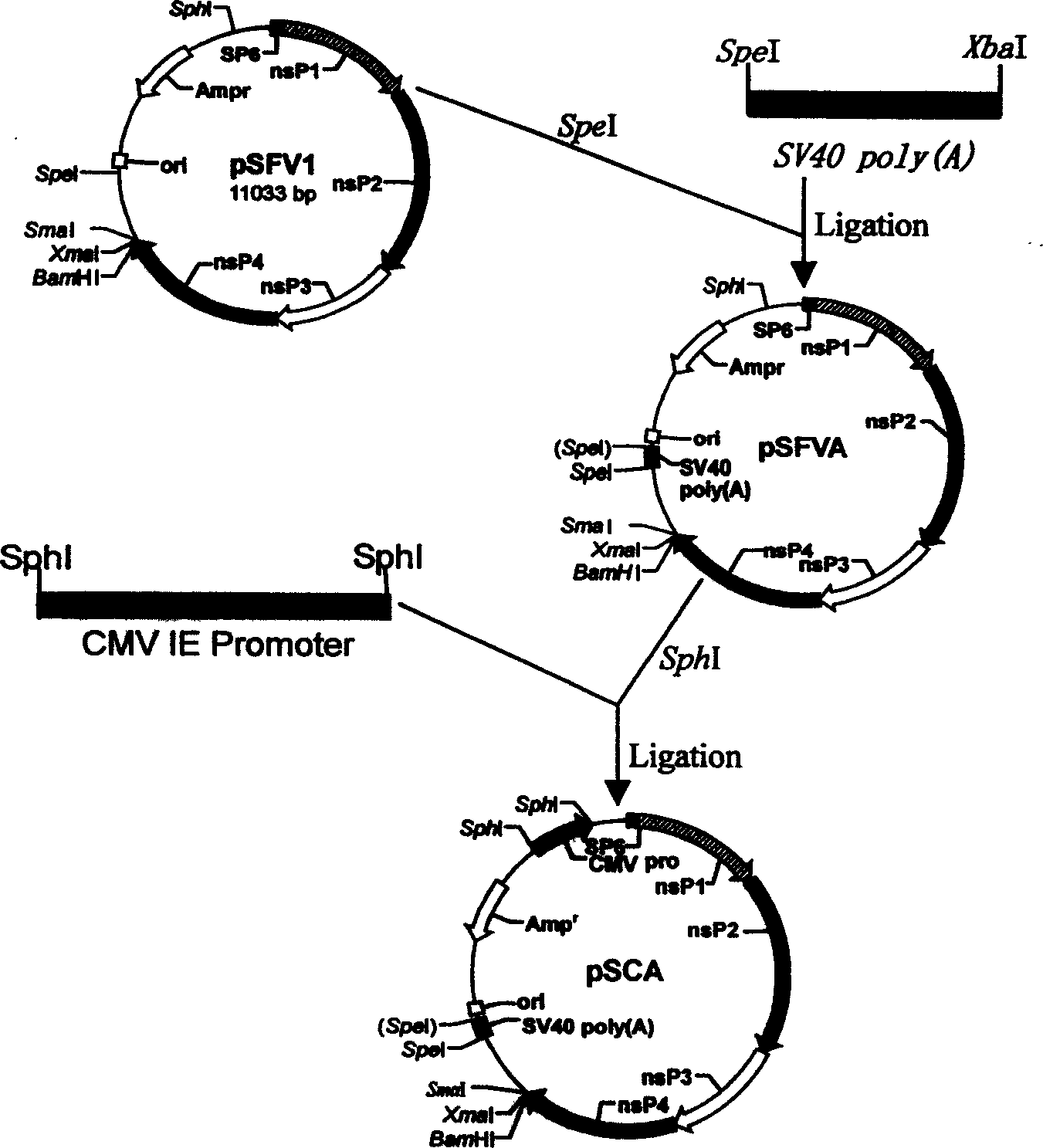
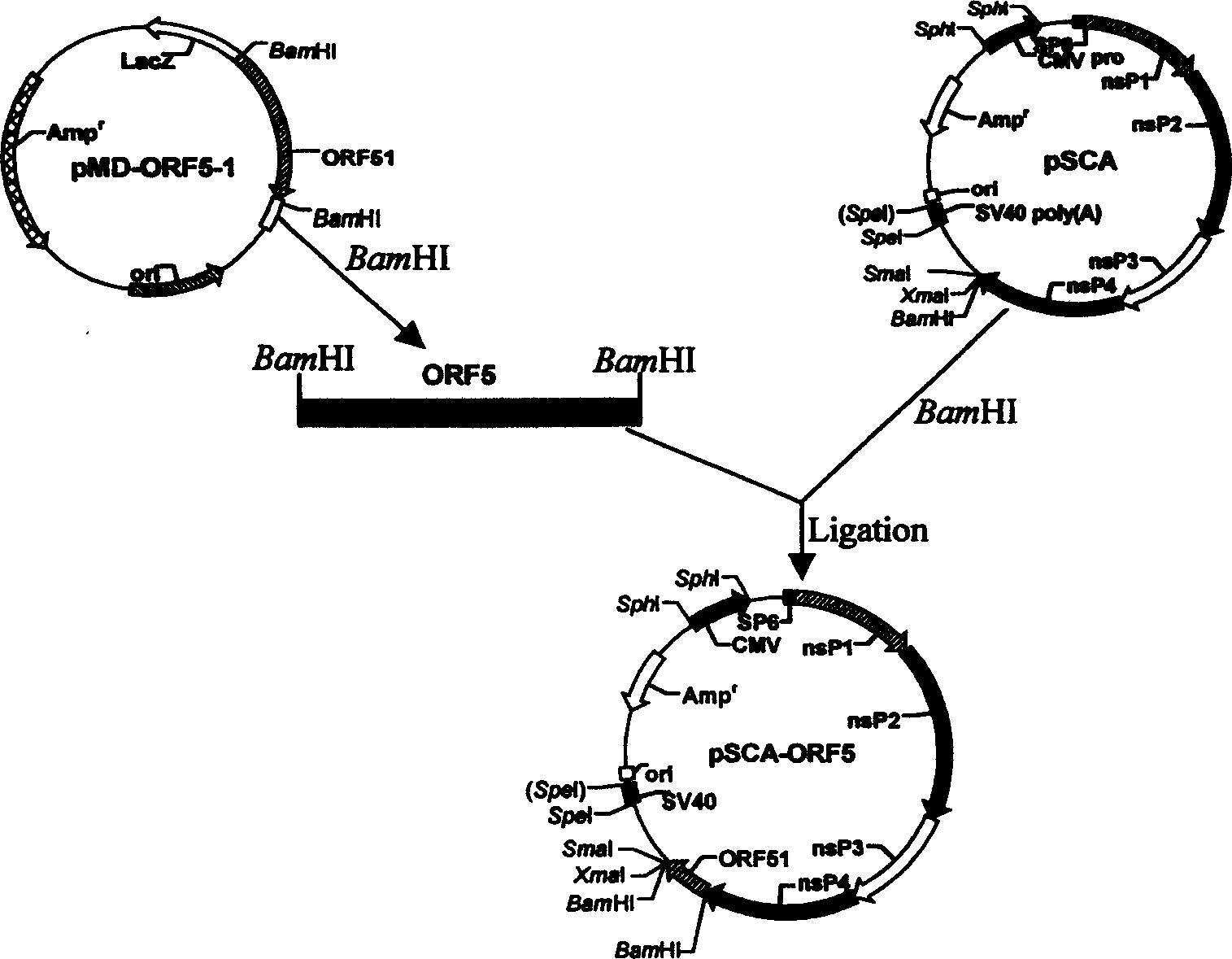
The present invention discloses a 'suicidal' DNA vaccine and its application for preventing and curing swine reproduction and respiration syndrome. It is characterized by that it is constructed by inserting the main immunogenic gene ORF5 of swine reproduction and respiration syndrome virus into BamHI site of 'suicidal' DNA vaccine expression vector pSCA, at the same time the expression vector pSCA suitable for preparing said 'suicidal' DNA vaccine also is constructed. Besides said invention also provides a method for preparing said expression vector pSCA.
Owner:HUAZHONG AGRI UNIV
Porcine reproductive and respiratory syndrome virus antibody detection kit
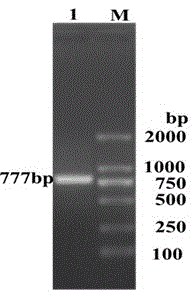
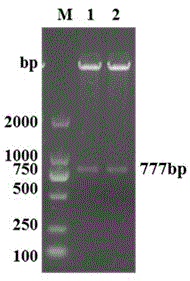
The invention discloses a porcine reproductive and respiratory syndrome virus antibody detection kit and belongs to the technical field of biological detection. The detection kit comprises a porcine reproductive and respiratory syndrome virus ELISA antigen coating plate, and the antigen in the ELISA antigen coating plate is protein expressed and purified by a recombinant expression plasmid pET777 obtained through double digestion of the gene segment NSP7-777 of a PRRSV DY strain and pET-32a by means of restriction enzymes BamHI and SacI. An antibody can be detected within three days after PRRSV DY strain NSP7 protein is immunized, and a high antibody level can still be realized within 248 days. The coincident rate of the detection method and an IDEXX kit can reach 99% or above, however, the cost is only one eighth that of the IDEXX kit. The porcine reproductive and respiratory syndrome virus antibody detection kit can replace imported kits to be used and popularized.
Owner:SHANDONG BINZHOU ANIMAL SCI & VETERINARY MEDICINE ACADEMY
Multifunctional shuttle vector new pBE2, construction method thereof and method for constructing alkali protease mutation library by using same
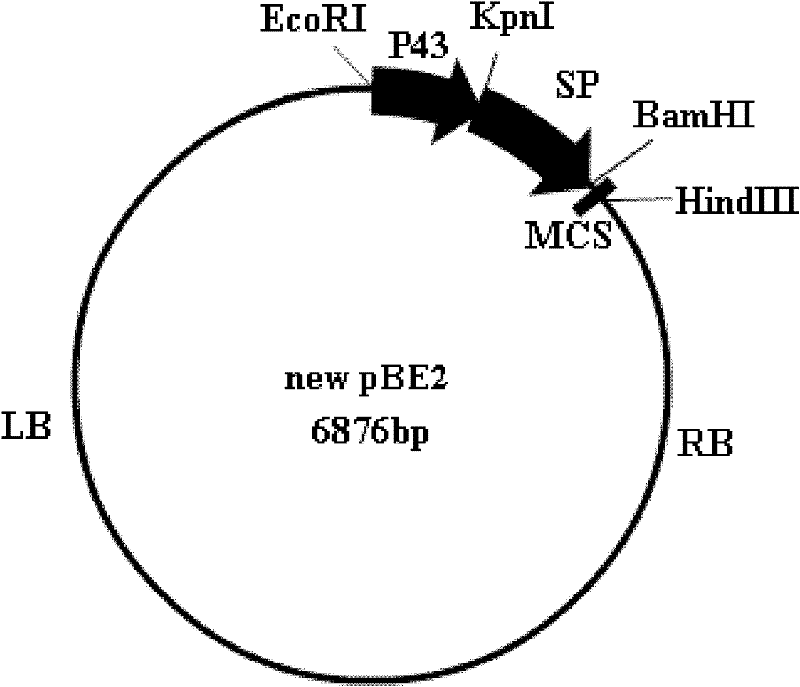
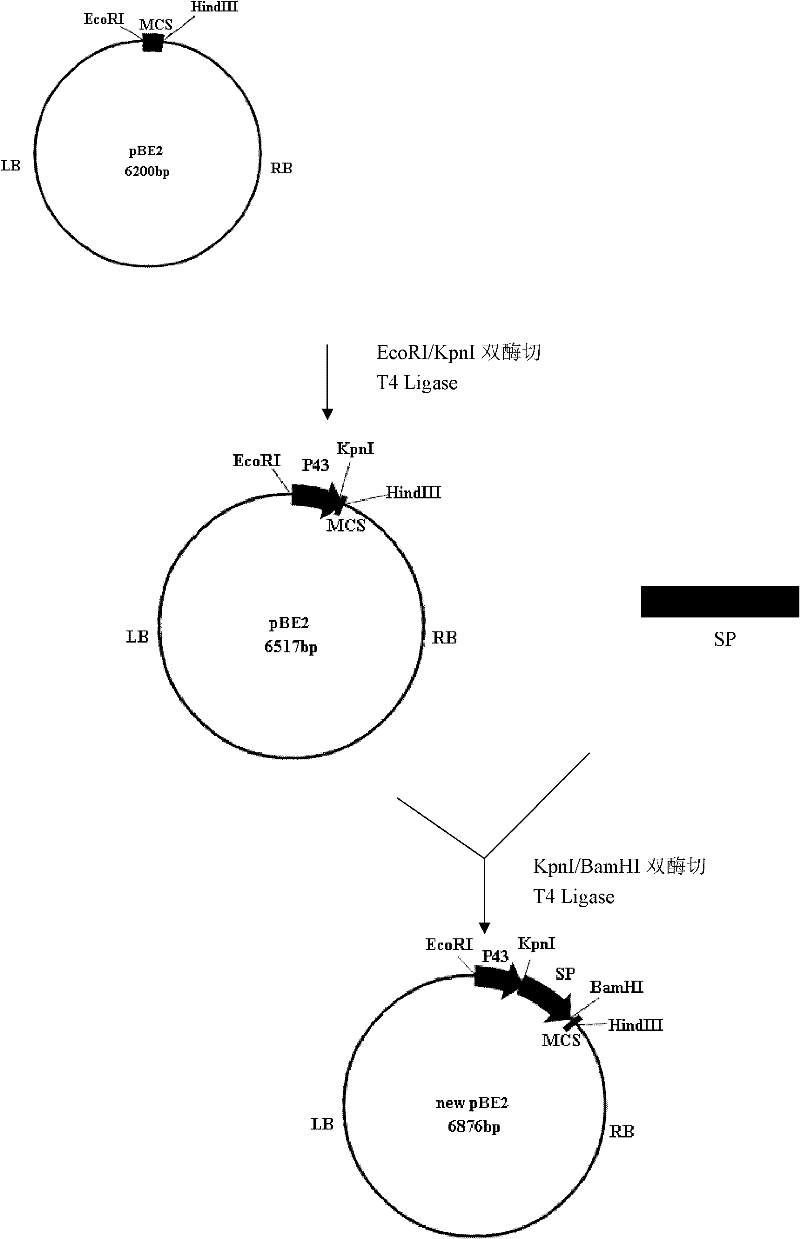
InactiveCN102229942AImprove screening efficiencyReduce workloadMicroorganism based processesVector-based foreign material introductionEscherichia coliShuttle vector
The invention relates to a multifunctional shuttle vector new pBE2, a construction method thereof and a method for constructing an alkali protease mutation library by using the same. The main structure of the multifunctional shuttle vector is LB-P43-SP-MCS-RB. The construction method mainly comprises the following steps: connecting a big fragment of the pBE2 vector with a P43 strong promoter and the product of the amplification of surfactant protein (SP) and introducing a BamHI enzyme cutting site. The method for constructing the alkali protease mutation library mainly comprises: connecting a mature peptide fragment of an alkali protease gene of bacillus alcalophilus to the new pBE2, transferring into Escherichia coli to obtain a positive clone, transferring the positive clone into bacillus subtilis WB600 and performing several circles of high-flux screening. When the method for constructing the alkali protease mutation library, which is provided by the invention, is used, the pertinence to mutation of an alkali protease gene is increased, mutation is performed according to the mature peptide part of the alkali protease, and the workload required for oriented evolution study on the alkali protease is reduced greatly.
Owner:TIANJIN UNIV OF SCI & TECH
Detection and Quantification of Target Nucleic Acid Sequence of a Microorganism
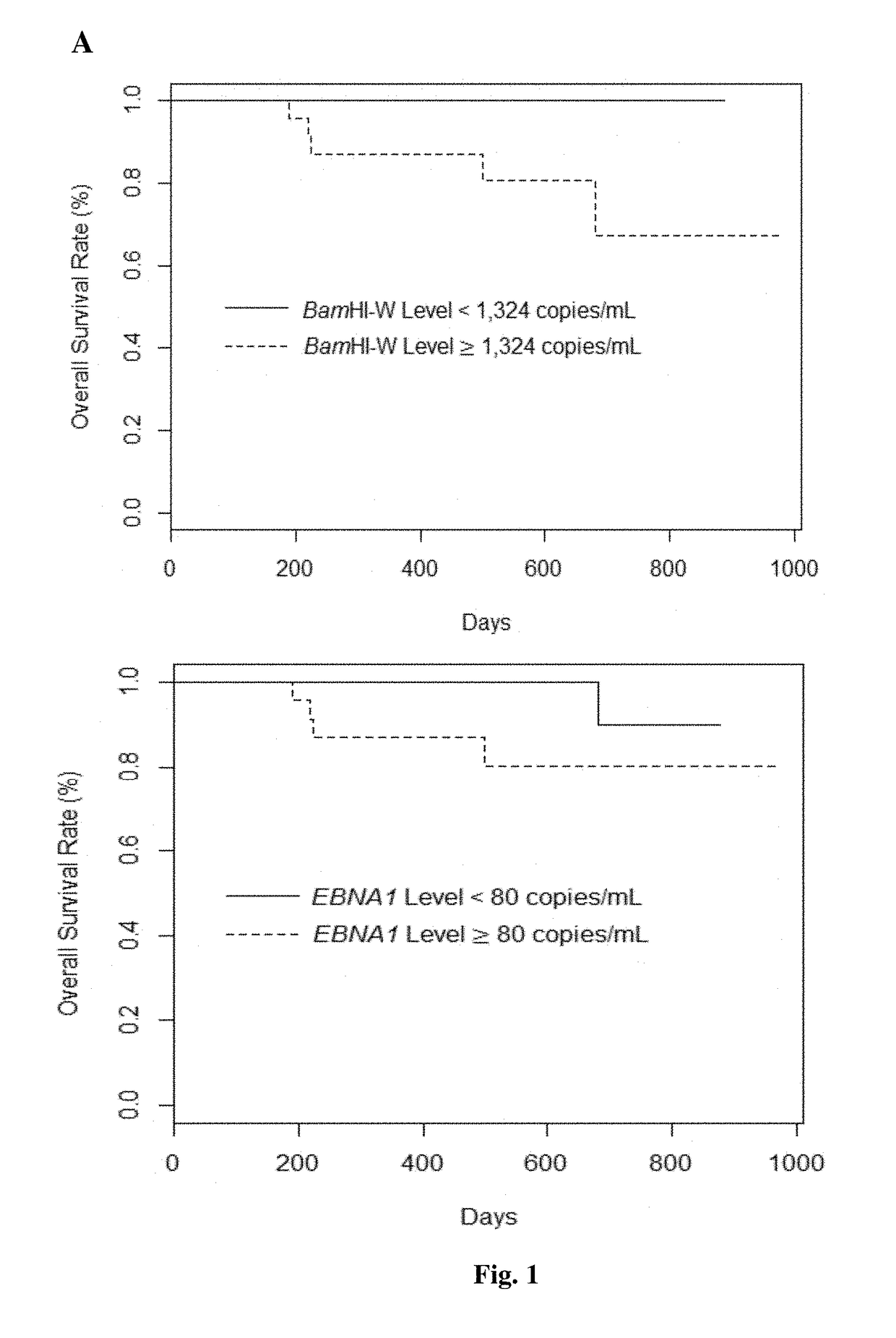
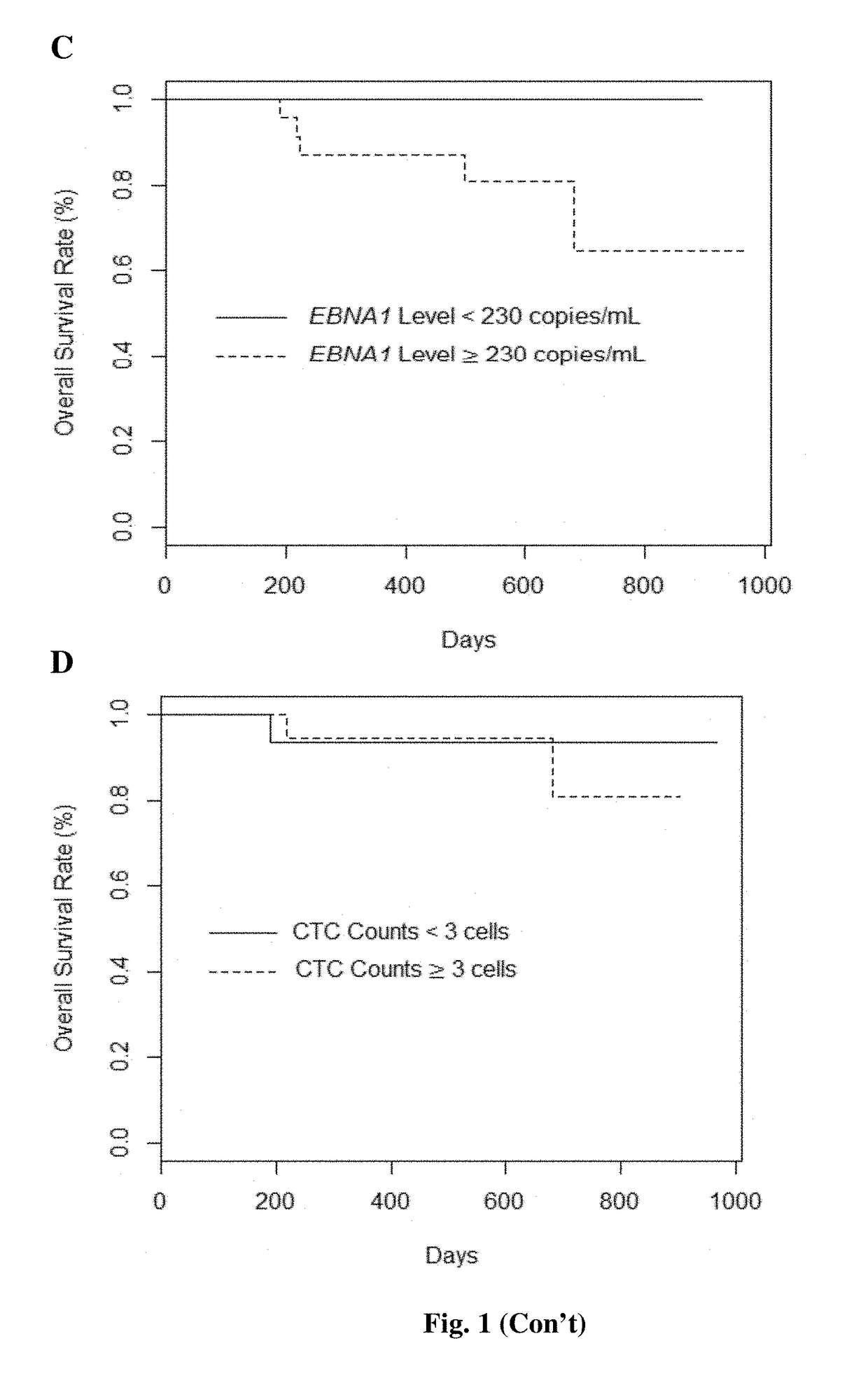
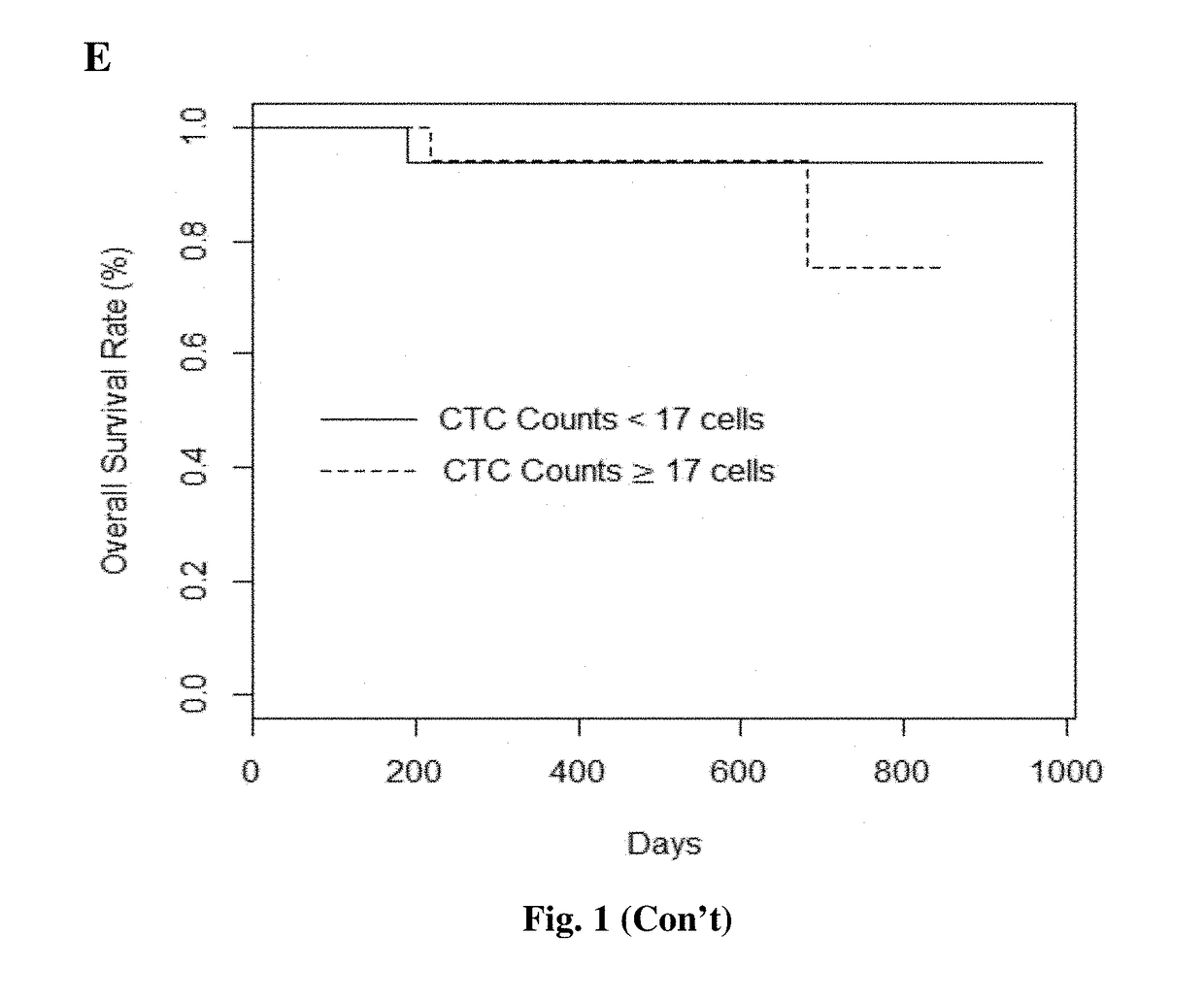
The present invention provides a method for detecting and / or quantifying the presence of a target nucleic acid sequence of a microorganism in a sample obtained from a subject, including amplifying the target sequence in a CpG island of the nucleic acid of the microorganism, irrespective of the methylation status of the CpG island. The invention is embodies by a method for detecting and / or quantifying the presence of a target nucleic acid sequence of Epstein-Barr virus (EBV) by amplifying a target sequence in the BamHI-W region of EBV in cell free DNAs (cfDNAs) obtained from a subject. The present invention also provides a kit to be used for the method of the invention.
Owner:LUCENCE LIFE SCI PTE LTD
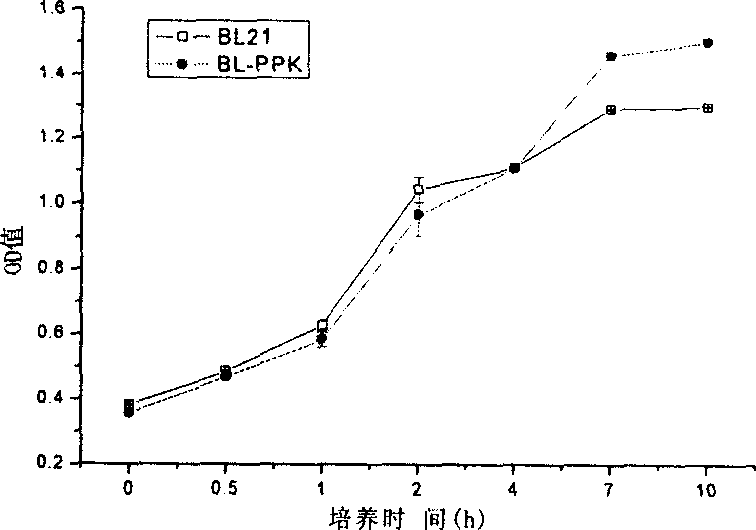
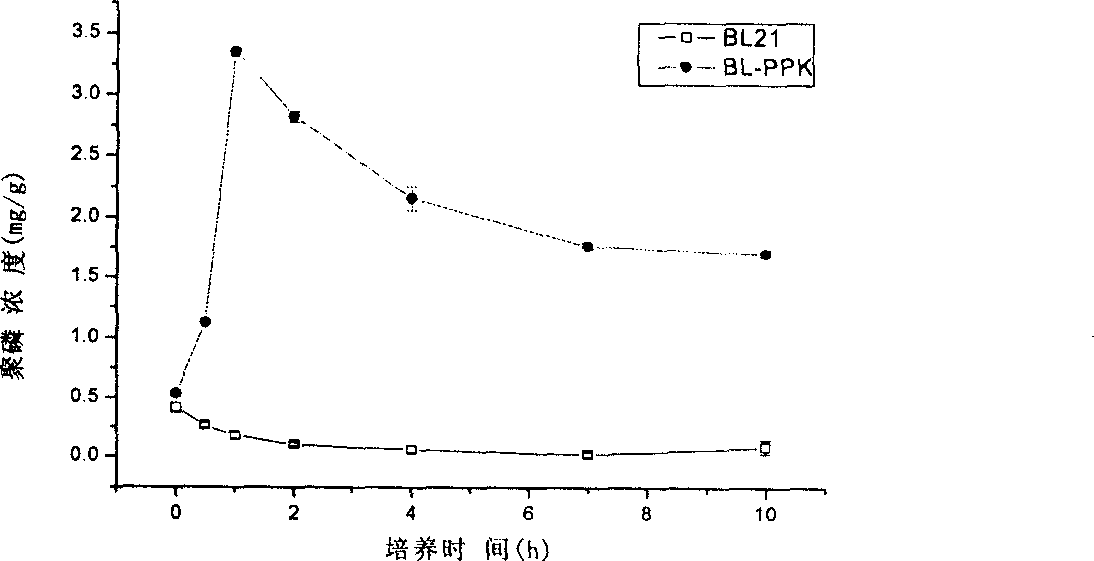
Construction method for polyphosphate kinase gene transformed Escherichia coli
InactiveCN1876809AProof of feasibilityEfficient removalBacteriaMicroorganism based processesBamHIGene clone
The invention discloses a construction approach for bacillus coli of transpolyphosphokinase gene, in which the said method includes gene-clone and carrier-construction; the clone includes extracting master DNA of bacillus coli; designing primers; augmenting ppk gene; restructuring the augmented object gene into clonic carrier pMD18-T and converting bacillus coli DH5 alpha; the construction includes adopting restriction enzymes of BamHI and HindIII bisenzyme to cut pMD18-T plasmid with ppk object gene and idle expression carrier pET-28a(+); directionally connecting the object gene by T4 to the expression carrier pET-28a(+), then converting the DH5 alpha; by the PCR and bisenzyme pressure methods screening and verifying the positive recon; extracting the recombination plasmid Pet28a-PPK and converting the acceptor strain BL21(DE3). The invention is of simple process and the obtained gene has efficient phosphorous removal ability.
Owner:NANJING UNIV
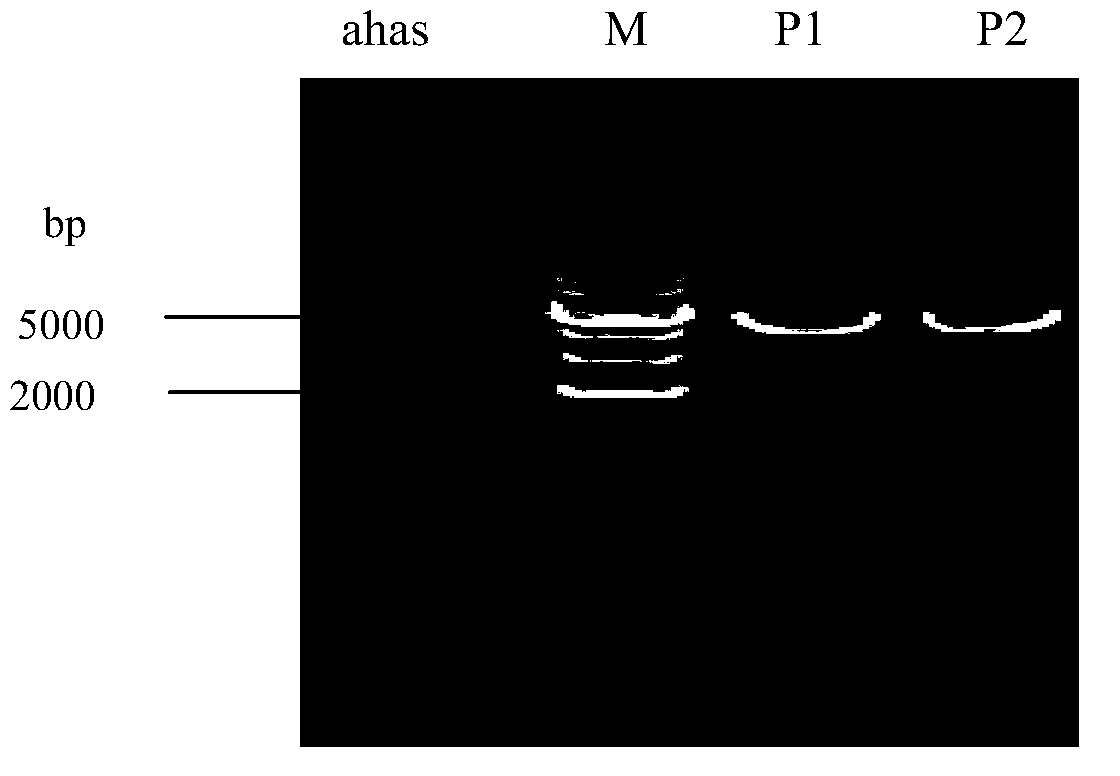
Preparation and preservation method for colon bacillus acetohydroxyacid synthase
InactiveCN103710328AHigh purityHigh activityTransferasesMicroorganism based processesEscherichia coliEnzyme digestion
The invention provides a method for preparing AHAS (Acetohydroxyacid Synthase) which has good purity and high activity and can exist stably. The method comprises the following steps: upstream and downstream primers are designed according to a gene sequence of a colon bacillus AHAS catalytic subunit, wherein the designed upstream primer carries an Xho I enzyme digestion site and the downstream primer carries a BamHI enzyme digestion site; a colon bacillus BL21 strain genomic DNA (Deoxyribonucleic Acid) is used as a template and a target segment ahas is obtained by a PCR (Polymerase Chain Reaction) amplification technology; the target segment ahas is connected onto an expression carrier PGEX-AT-1 to obtain a recombinant plasmid PGEX-4T-1-ahas; inducible expression is realized in a prokaryotic expression system; an expressed enzyme is purified by agarose gel resin modified by glutathione sulfur transferase (GST) so as to obtain the acetohydroxyacid synthase with good activity.
Owner:NORTHWEST UNIV
Method for expressing hABCG2 in insect cell sf9
InactiveCN104745632ARealize fully automatic high-throughput screeningIncrease flexibilityGenetic engineeringFermentationEscherichia coliEnzyme digestion
The invention relates to a method for expressing hABCG2 in an insect cell sf9. The method comprises the following steps: cloning a human ABCG2 gene with BamHI and HindIII enzyme digestions at two ends into BamHI and HindIII sites of a pFastBac1 carrier respectively; converting an obtained recombinant plasmid pFastbac1-hABCG2 into a DH10Bac escherichia coli competent cell to obtain a recombinant baculovirus shuttle vector Bacmid-hABCG2 which is inserted into the hABCG2 gene; mediating the recombinant baculovirus shuttle vector Bacmid-hABCG2 with lipidosome to transfect an sf9 insect cell to obtain hABCG2 recombinant virus, and further culturing to obtain the expressed hABCG2 protein. According to the method disclosed by the invention, a novel way is provided for obtaining hABCG2 proteins on a large scale, and a working foundation is laid for the application and development of the hABCG2 proteins.
Owner:苏州杰诺曼博生物科技有限公司
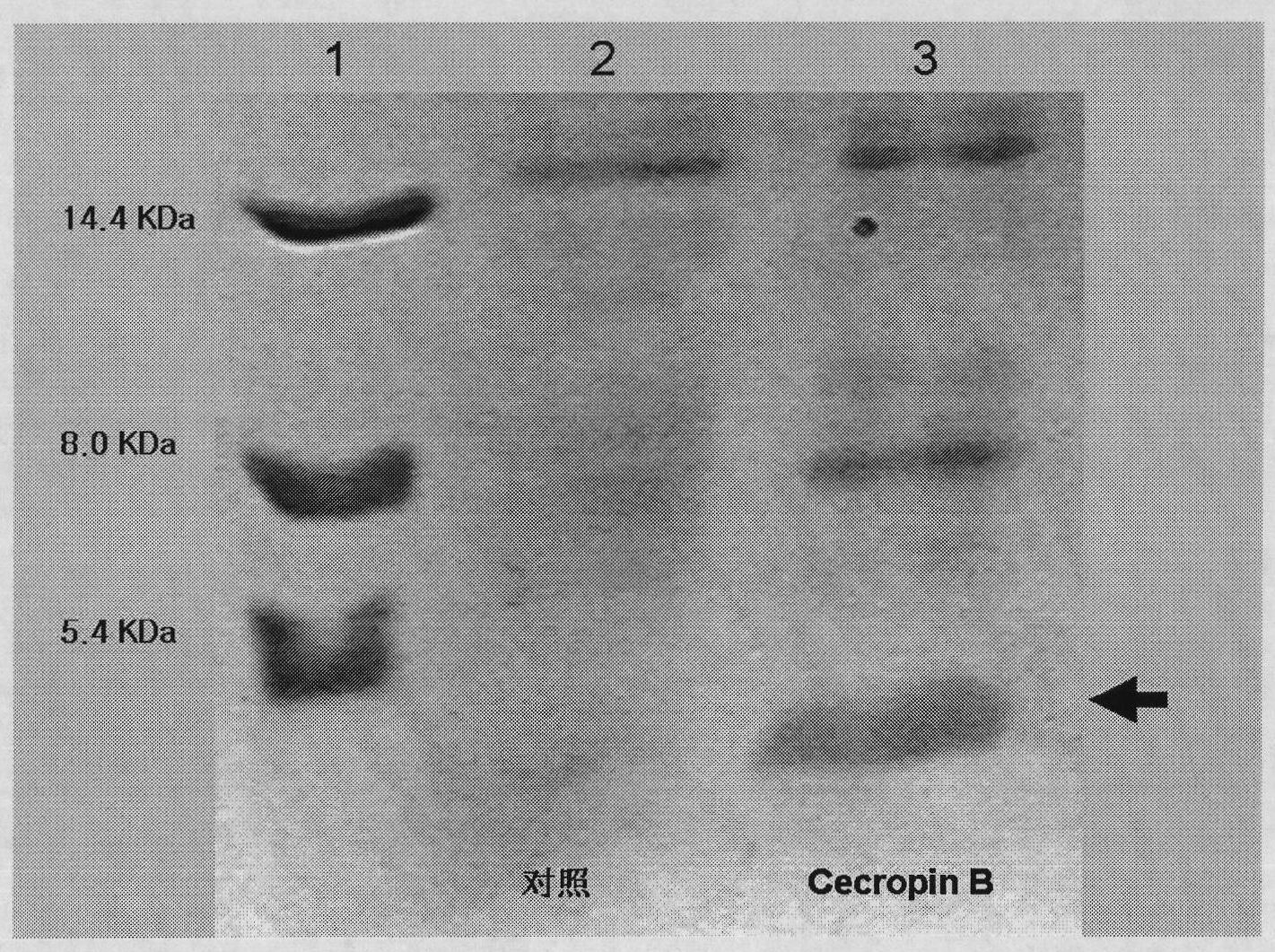
Method for using silkworm cultured cell to express antibacterial peptide Cecropin B
InactiveCN101845439ASolve operational problemsFix stability issuesViruses/bacteriophagesGenetic engineeringDideoxynucleotide TriphosphatesTotal rna
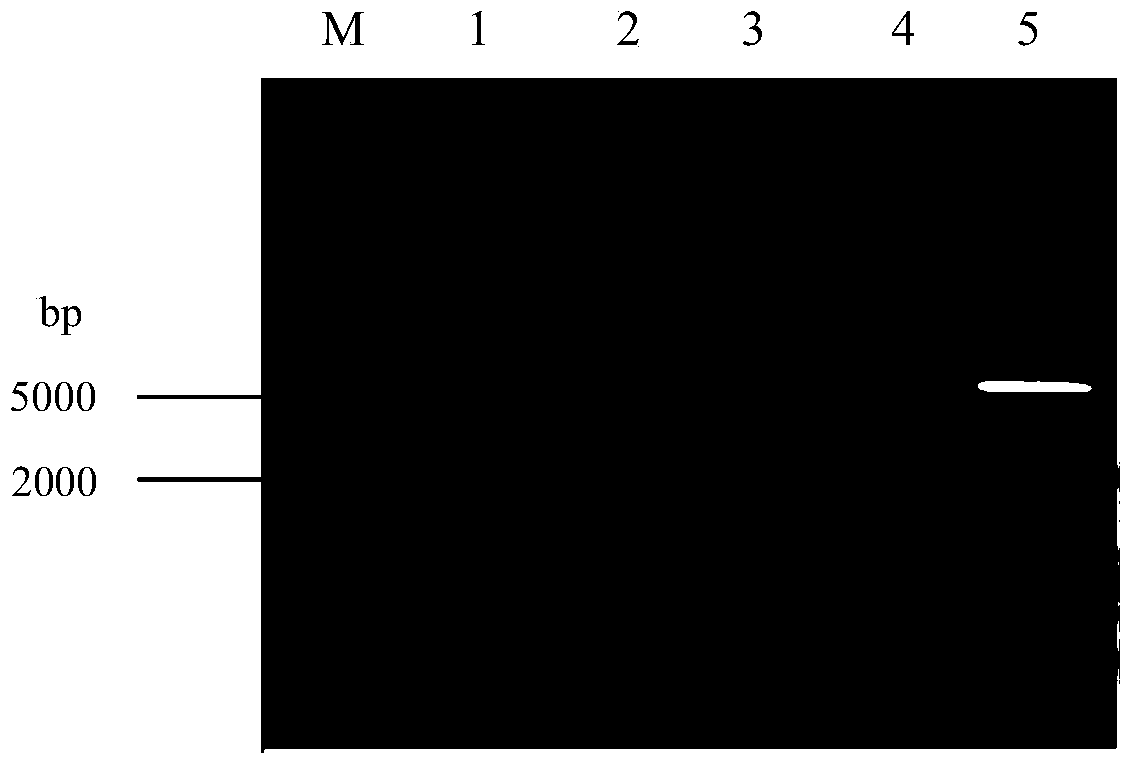
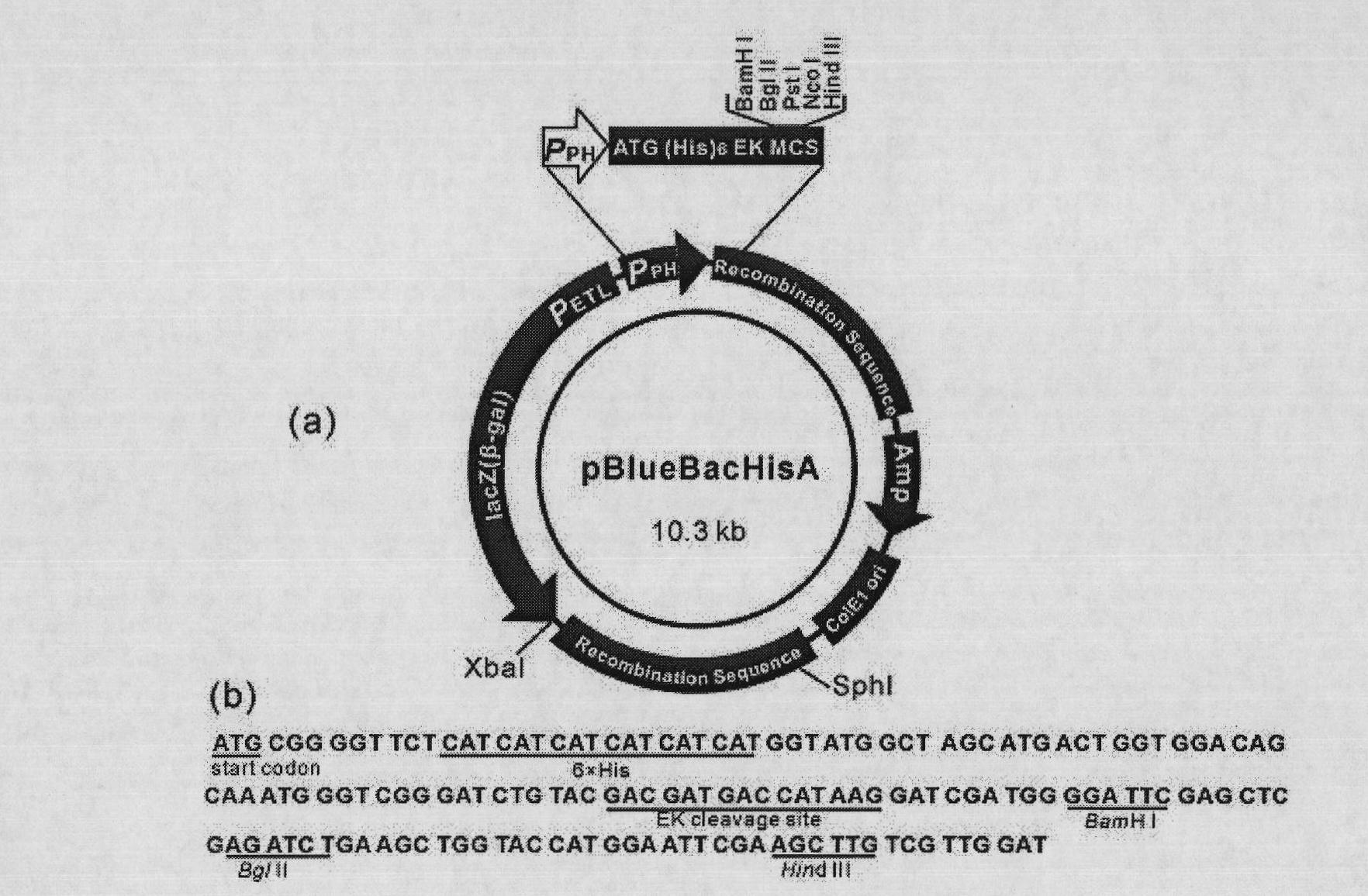
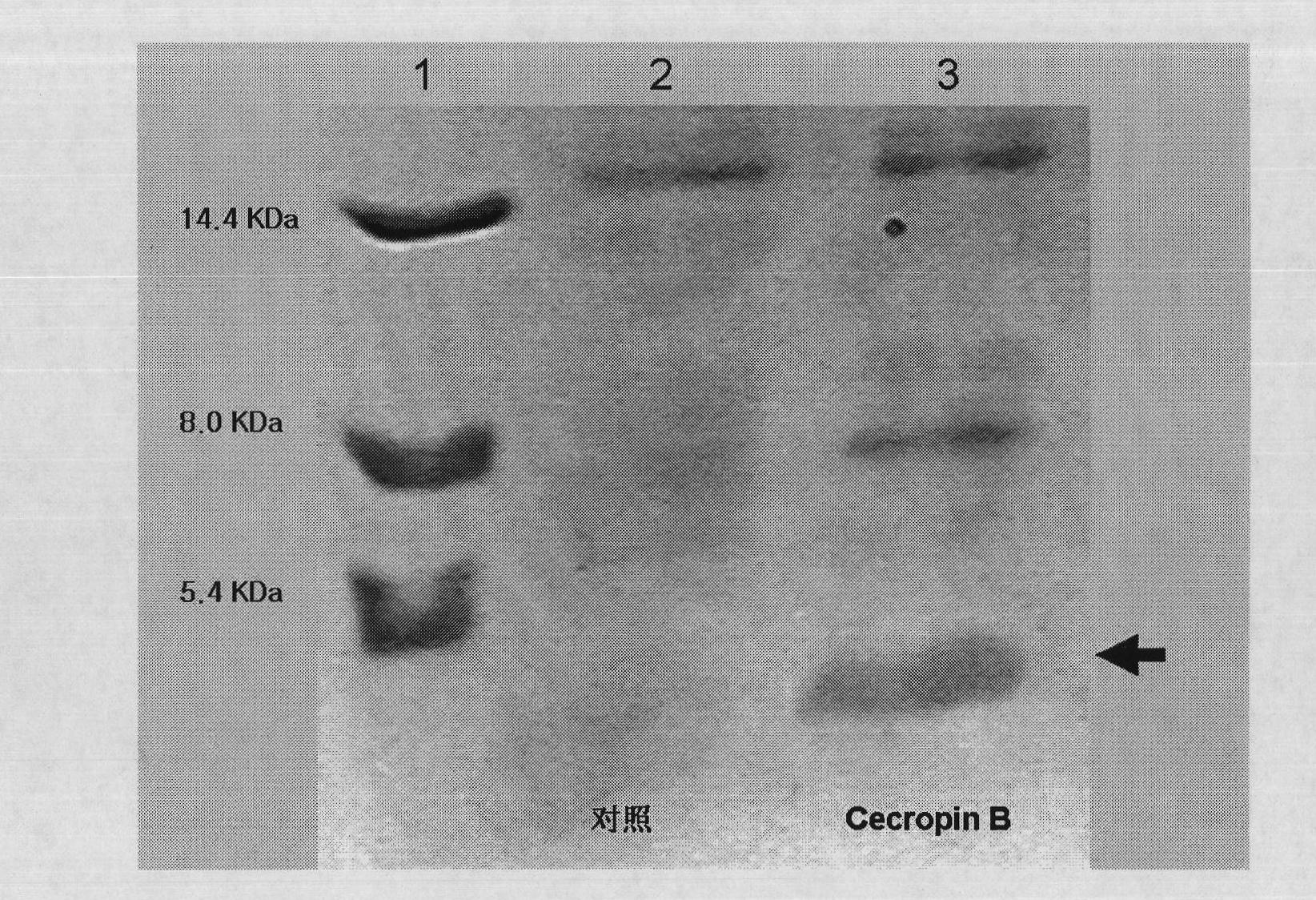
The invention discloses a method for using silkworm cultured cell to express antibacterial peptide Cecropin B. The method comprises the following steps: (1) using the total RNA extracted from the fat body cells of wild silkworm chrysalis as template, adopting RT-PCR amplification to obtain wild silkworm antibacterial peptide Cecropin B gene; using 1% agarose gel electrophoresis to perform PCR product analysis to the antibacterial peptide Cecropin B gene, using a PCR purification kit of Qiagene for purification, then cloning the purified PCR product to TA vector pCR2.1 to obtain pCR2.1-Cecropin B, utilizing the dideoxynucleotide chain termination to confirm the correctness of cloned gene order; using restriction endonuclease BamHI and HindIII to digest pCR2.1-Cecropin B and obtain Cecropin B genetic fragments, then cloning rhabdovirus transfer vector pBlueBacHisa in the genetic fragments to obtain reconstituted transfer vector; performing cotransfection of the reconstituted transfer vector and silkworm wild-type nuclear polyhedrosis virus BmNPV DNA in silkworm cultured cell, performing homologous recombination to generate recombinant virus; and (3) inoculating the recombinant virus containing wild silkworm antibacterial peptide Cecropin B gene in the silkworm cultured cell, infecting at 27 DEG C for three days to express the infection, and centrifuging to collect silkworm cultured cell.
Owner:ZHEJIANG UNIV