Carboxyesterase and application thereof to production of cyclohexene formic acid by kinetic resolution of cyclohexene formate
A kinetic separation and cyclohexenecarboxylic acid technology, applied in the field of enzyme engineering, can solve the problems of poor substrate tolerance and insufficient product concentration selectivity, etc.
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0032] Embodiment 1: Cloning of carboxylesterase gene
[0033] According to the gene sequence (Genebank accession number: AXY61874.1) of Acinetobacter sp.WCHAc010052 carboxylesterase recorded in Genbank, the PCR primers were designed as follows:
[0034] Upstream primer: 5'-gtgccgcgcggcagc catatg ATGGTTGCGTTTAATACAAAGATTCA-3'
[0035] Downstream primer: 5'-gtggtggtggtggtg ctcgag TTAGCGCTGGCGATCCCA-3';
[0036] Wherein, the underlined part of the upstream primer (nucleotide sequence shown in SEQ ID NO: 5) is the NdeI restriction site, and the underlined part of the downstream primer (the nucleotide sequence is shown in SEQ ID NO: 6) is the XhoI restriction site point.
[0037] The genomic DNA of Acinetobacter sp.WCHAc010052 was used as a template for PCR amplification. The PCR system is: 10 μL of 2×Taq PCR MasterMix, 1 μL (0.3 μmol / L) of each upstream primer and downstream primer, 1 μL (0.1 μg) of DNA template and ddH 2 O 7 μL. The PCR amplification program is: (1) Pre-d...
Embodiment 2
[0038] Embodiment 2: Preparation of carboxylesterase recombinant plasmid and recombinant expression transformant
[0039] Digest the carboxylesterase gene DNA fragment obtained in Example 1 and pET28a empty plasmid with restriction endonucleases NdeI and XhoI at 37°C for 2 h, purify by agarose gel electrophoresis, and use an agarose gel DNA recovery kit Recycle the target fragment. The target fragment was ligated overnight at 4°C under the action of T4 DNA ligase to obtain the recombinant expression plasmid pET28a-CarEst3.
[0040] The above recombinant expression plasmids were transformed into Escherichia coli E.coli DH5α competent cells, positive recombinants were screened on a resistance plate containing kanamycin, single clones were picked, and positive clones were verified by colony PCR. Cultivate the recombinant bacteria, extract the plasmid after the amplification of the plasmid, and retransform into E. coli BL21 (DE3) competent cells. The transformation solution is sp...
Embodiment 3
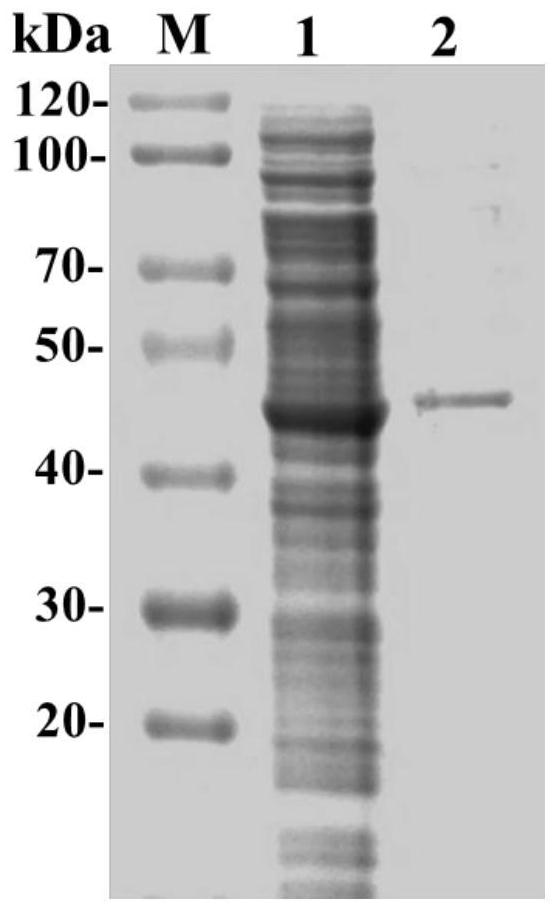
[0041] Example 3: Expression of Carboxylesterase
[0042] The recombinant Escherichia coli obtained in Example 2 was inoculated into LB medium containing kanamycin (peptone 10g / L, yeast extract 5g / L, NaCl 10g / L, pH 7.0), cultured with shaking at 37°C overnight, and 1% (v / v) inoculum was placed in a 500mL Erlenmeyer flask containing 100mL LB medium, placed at 37°C and shaken at 180rpm for culture, when the OD of the culture solution 600 When it reaches 0.6, add IPTG with a final concentration of 0.2mmol / L as an inducer, and after induction at 25°C for 12 hours, centrifuge the culture medium, collect the cells, and wash twice with normal saline to obtain resting cells, which can be obtained by freeze-drying for 24 hours Cells were freeze-dried and stored at 4°C after collection. The resulting resting cells can also be suspended in a pH 7.0 buffer, ultrasonically disrupted in an ice bath, and the supernatant collected by centrifugation is the crude enzyme solution of the recombi...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More