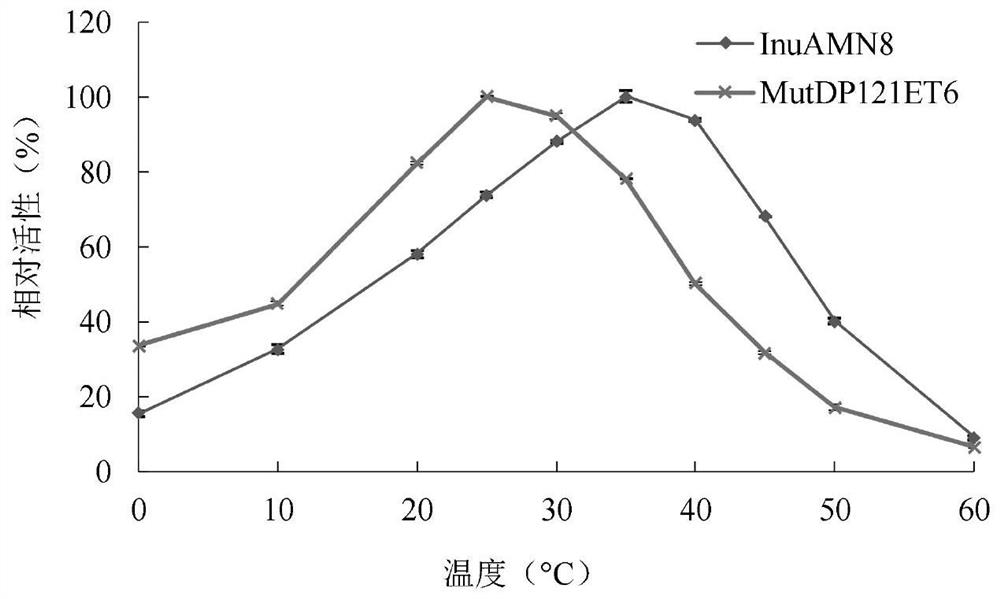
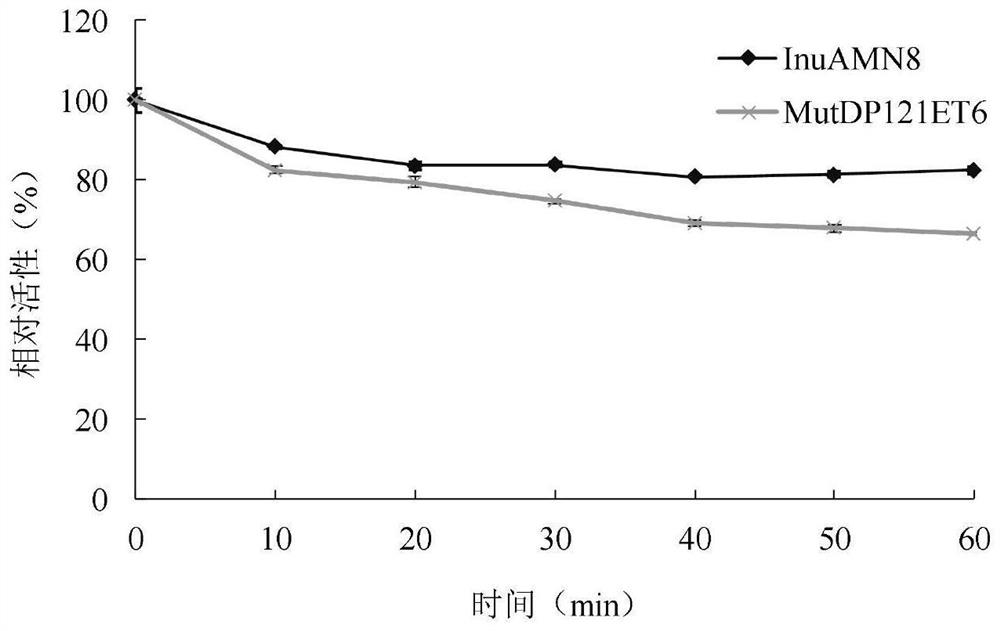
Exo-inulinase mutant MutDP121ET6 with improved low-temperature activity
An exo-inulinase and mutant technology, applied in the direction of enzymes, hydrolases, glycosylases, etc., can solve the problems of easy exposure of protease cleavage sites and easy degradation of enzymes
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0033] Construction and Transformation of Embodiment 1 Wild Enzyme InuAMN8 Expression Vector
[0034] (1) Extraction of Arthrobacter genomic DNA: Centrifuge the bacterial liquid cultured for 2 days to get the bacterial cells, add 1mL lysozyme, treat at 37°C for 60min, and then follow the bacterial genomic DNA extraction kit (Tiangen Biochemical Technology (Beijing) Co., Ltd.) According to the instructions, Arthrobacter genomic DNA was extracted and stored at -20°C for later use.
[0035](2) According to the exo-inulinase nucleotide sequence JQ863111 (SEQ ID NO.4) recorded in GenBank, primers 5'-ATGAATTCATTGACGACGGC-3'(SEQ ID NO.5) and 5'-TCAACGGCCGACGACGTCGA-3'( SEQ ID NO.6), using Arthrobacter genomic DNA as a template for PCR amplification, the PCR reaction parameters are: denaturation at 95°C for 5min; then denaturation at 95°C for 30sec, annealing at 58°C for 30sec, extension at 72°C for 1min 30sec, and after 30 cycles, 72 ℃ for 5 minutes. The PCR results obtained the co...
Embodiment 2
[0038] Construction and transformation of embodiment 2 mutant enzyme MutDP121ET6 expression vector
[0039] (1) Design primers 5'-TGAAGAAGACCGAAAGACGGGCCGGCAGGCGCAGTCG-3'(SEQ ID NO.7) and 5'-CCGTCTTTCGGTCTTTTTCACTGTAGGCACTG-3'(SEQ ID NO.8), use plasmid pEasy-E1-inuAMN8 as template for PCR amplification, PCR The reaction parameters were: denaturation at 95°C for 30 sec; then denaturation at 95°C for 15 sec, annealing at 70°C for 15 sec, extension at 72°C for 3 min and 30 sec, and after 30 cycles, incubation at 72°C for 5 min. As a result of PCR, the recombinant expression linearized plasmid pEasy-E1-mutDP121ET6 containing mutDP121ET6 (SEQ ID NO.2) was obtained. mutDP121ET6 and pEasy-E1-mutDP121ET6 can also be obtained by gene synthesis.
[0040] (2) Add 1 μL of DpnI enzyme to 50 μL of the PCR product of the linearized plasmid pEasy-E1-mutDP121ET6, and digest it at 37° C. for 1 hour.
[0041] (3) Using Mut II Fast Mutagenesis Kit, place the digested product in step (2) at 37...
Embodiment 3
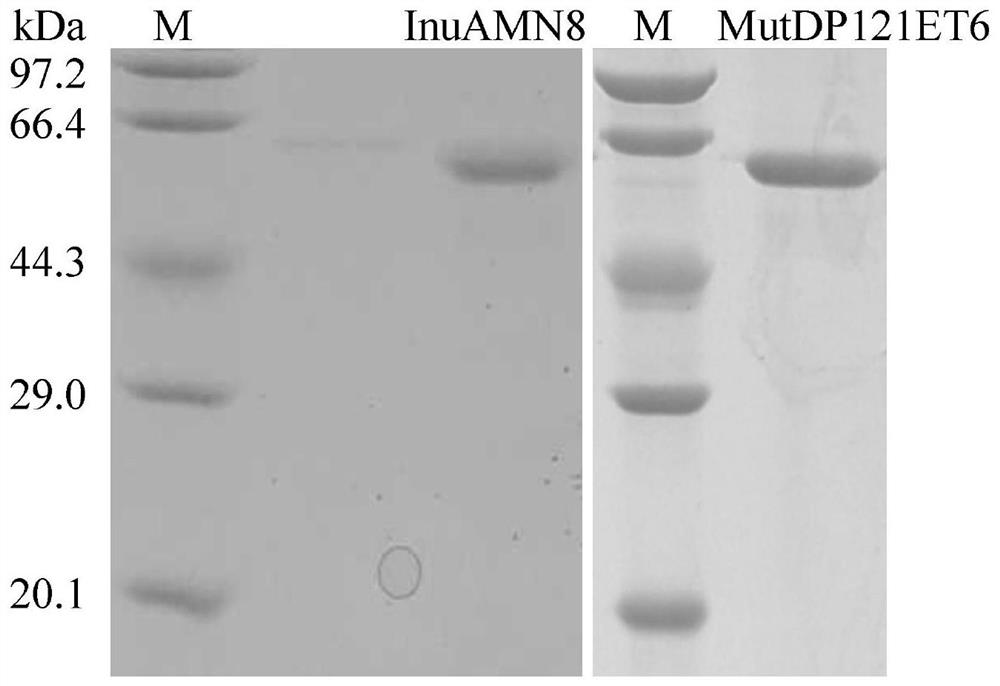
[0043] Embodiment 3 Preparation of recombinant wild enzyme InuAMN8 and mutant enzyme MutDP121ET6
[0044] (1) The recombinant strains BL21(DE3) / inuAMN8 and BL21(DE3) / mutDP121ET6 were inoculated in LB (containing 100 μg mL -1 Amp) medium, shake rapidly at 37°C for 16h.
[0045] (2) Then the activated bacterial solution was inoculated into fresh LB (containing 100 μg·mL -1 Amp) culture medium, rapid shaking culture for about 2–3h (OD 600 After reaching 0.6-1.0), add IPTG with a final concentration of 0.7mM for induction, and continue shaking culture at 20°C for about 20h. Centrifuge at 12000rpm for 5min to collect the bacteria. After suspending the cells with an appropriate amount of McIlvainebuffer with a pH of 7.0, the cells were disrupted ultrasonically in a low-temperature water bath. The crude enzyme solution concentrated in the cells above was centrifuged at 13,000rpm for 10min, the supernatant was aspirated and the target protein was affinity and purified with Nickel-...
PUM
 Login to View More
Login to View More Abstract
Description
Claims
Application Information
 Login to View More
Login to View More