Single-domain antibody resistant to human beta2-microglobulin as well as preparation method and application of single-domain antibody
A β2 microglobulin and single domain antibody technology, applied in the field of anti-β2 microglobulin antibodies, can solve the problems of increased sensitivity, high production cost, limited antibody immobilization density, etc., to promote diagnosis and treatment, high specificity Binding capacity, improved diagnosis and treatment outcomes
- Summary
- Abstract
- Description
- Claims
- Application Information
AI Technical Summary
Problems solved by technology
Method used
Image
Examples
Embodiment 1
[0037] Example 1 Screening of monoclonal phage antibodies
[0038] Using phage display technology to carry out multiple rounds of enrichment and screening of the alpaca phage antibody library, the purified β 2 Microglobulin has undergone four rounds of "adsorption-elution-amplification" enrichment screening for antigens, and the recovery rate of each round of screening shows an increasing trend, indicating that after screening, there is β 2 Microglobulin-specific phages were highly enriched.
[0039] The specific method is as follows:
[0040] 1. Establishment of alpaca phage antibody library
[0041] The names and sequences of the primers used are shown in Table 1, and the underlined part is the enzyme cutting site.
[0042] Table 1. Primer names and their sequences
[0043]
[0044]
[0045] (1) Separation of lymphocytes and isolation of total RNA
[0046] Take 100ml of peripheral blood from a 3-year-old female alpaca, put it into an anticoagulant tube, add an equ...
Embodiment 2
[0084] Example 2 Detection of monoclonal phage antibodies:
[0085] (1) Randomly pick 80 single bacterial clones from the TY-AG plate cultured overnight after the fourth round of screening, mix BLT5615 with phage preservation solution / lysate (1‰ to 0.1‰), and culture on a shaker at 37°C until After lysing and lysing the bacteria, immediately add NaCl to a concentration of 0.5M in the solution, centrifuge at 10000rpm for 10min, take the supernatant, and recover the phage liquid. ELISA method for detection: Antigen β 2 Microglobulin was coated on the microtiter plate (0.4 μg / well), and after blocking, 100 μl of monoclonal phage solution was added to each well, and incubated at 37° C. for 2 hours. After washing, 1:5000 HRP-labeled mouse anti-M13 antibody was added and incubated at 37°C for 1h. The positive control was the primary antibody (mouse anti-human β 2 Microglobulin), secondary antibody (goat anti-mouse IgG); negative control only added 2% MPBS. O-phenylenediamine was...
Embodiment 3
[0087] Example 3 DNA sequence determination of positive clones:
[0088] The phagemids of A and B positive clones were extracted respectively, and the sdAb DNA was sequenced by the dideoxy terminal termination method using sdAb DNA sequencing primers (upstream primers and downstream primers were from T7select cloning kit).
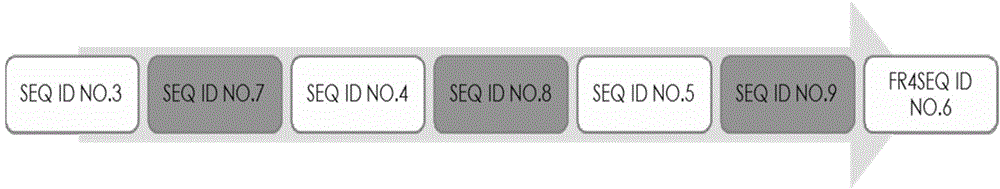
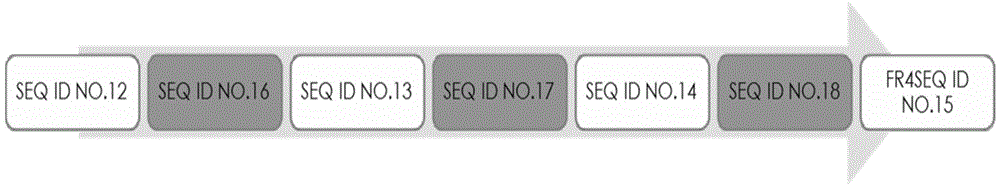
[0089] The nucleotide sequences of clones A and B were obtained, and the obtained sequences were compared with the single domain antibody variable region genes in Gene Bank using Blast software. The analysis results showed that the variable region genes of the obtained A and B clones were consistent with the SdAb antibody gene, the deduced amino acid sequence had a typical antibody variable region structure, and the protein sequence determined the start and end of the CDR according to the blast and Kabat codes parts. The amino acid composition and sequence of the complementarity determining regions (CDRs) are highly variable. The three CDRs together form...
PUM
| Property | Measurement | Unit |
|---|---|---|
| Titer | aaaaa | aaaaa |
Abstract
Description
Claims
Application Information
 Login to View More
Login to View More