Patents
Literature
140results about "Cis-trans-isomerases" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
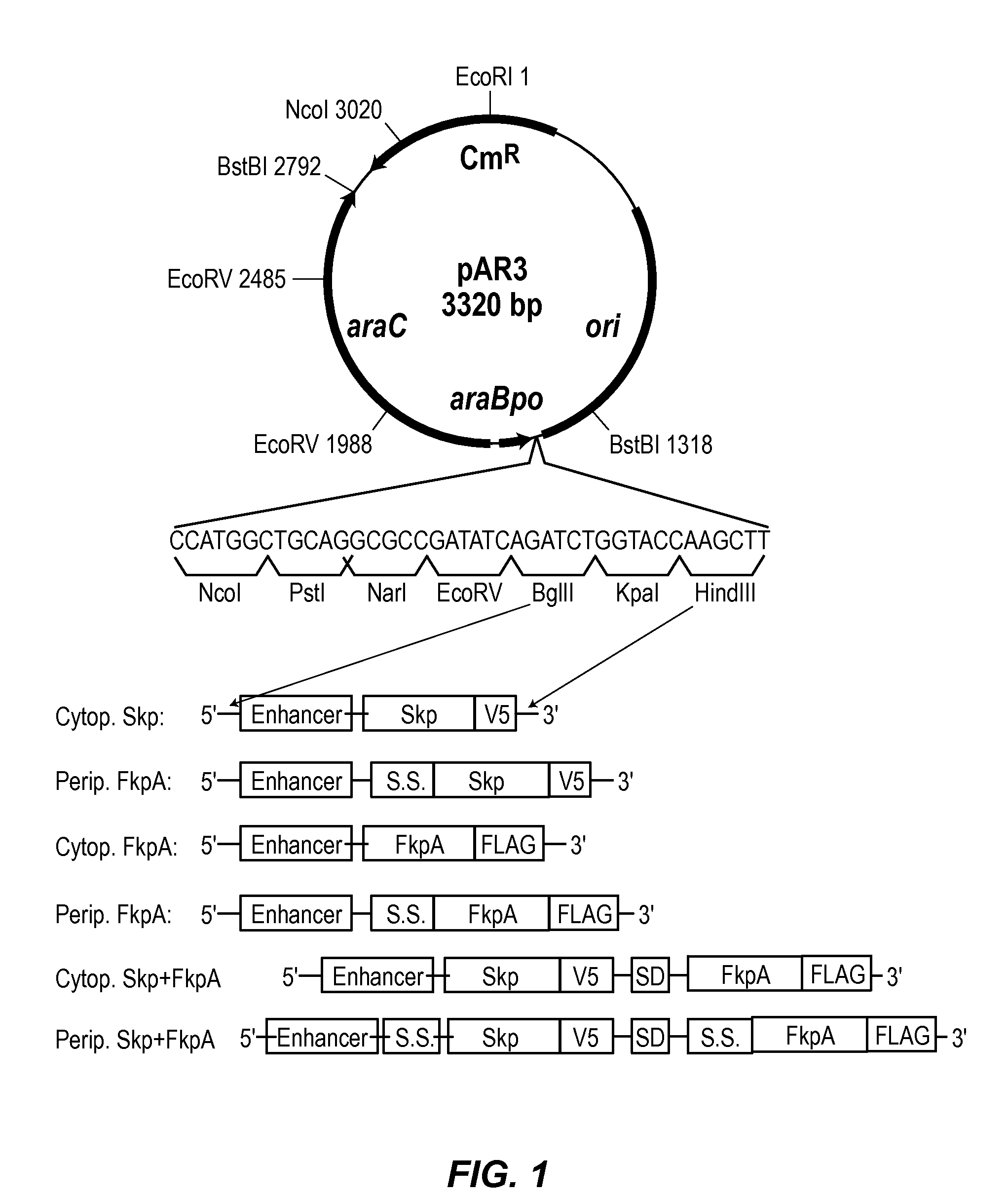
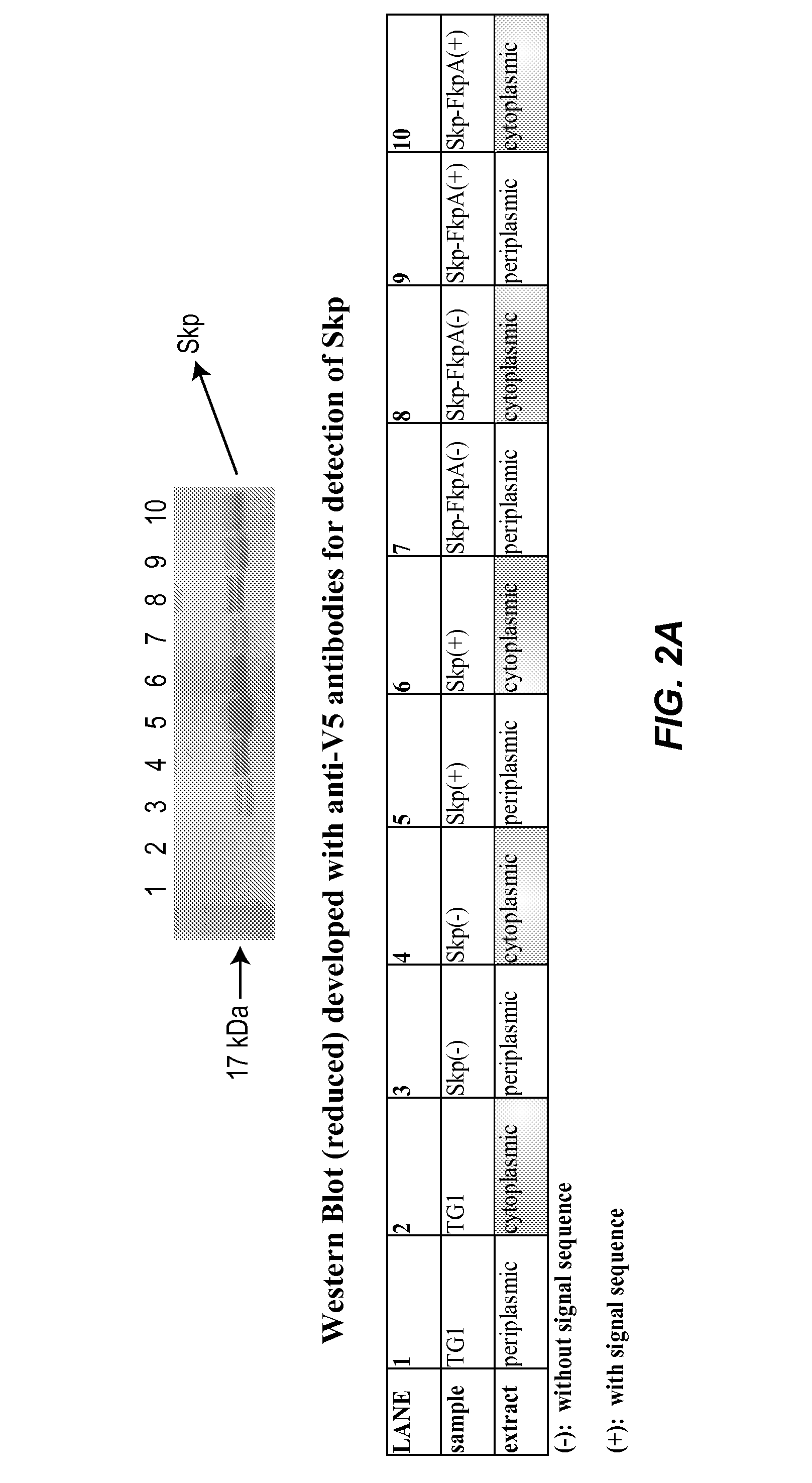
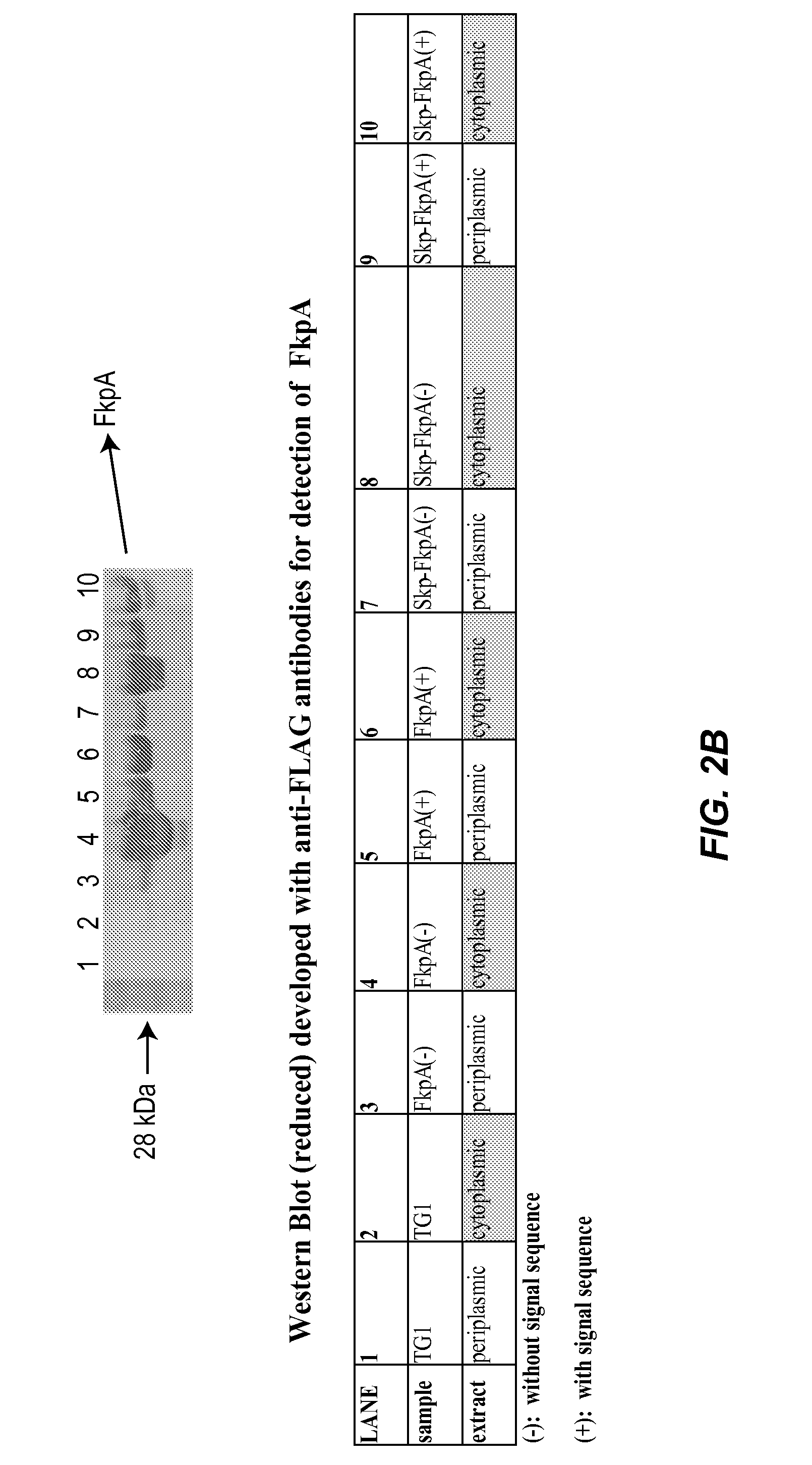
Methods and materials for enhancing functional protein expression in bacteria
ActiveUS20140154743A1Increase productionIncrease volumeBacteriaAntibody mimetics/scaffoldsHeterologousBiochemistry
Novel materials and methods useful for expressing heterologous proteins in prokaryotic cells are provided, including prokaryotic cells expressing FkpA and / or Skp.
Owner:XOMA TECH LTD
FK506-binding protein 7 related protein as a biomarker for neurodegenerative disease
InactiveUS20060115855A1Reduce expressionCell receptors/surface-antigens/surface-determinantsCis-trans-isomerasesFK-506-Binding ProteinProlyl isomerase
The present invention relates to a biomarker for neurodegenerative disease, including amyotrophic lateral sclerosis (ALS), Alzheimer's (AD), and Parkinson's (PD) disease. More particularly, the present invention relates to the identification of the FK 506-binding protein 7, or prolylisomerase, as a biomarker useful for the detection, diagnosis, and differentiation of neurodegenerative disease, including but not limited to ALS, AD, and PD.
Owner:BAYLOR COLLEGE OF MEDICINE +1
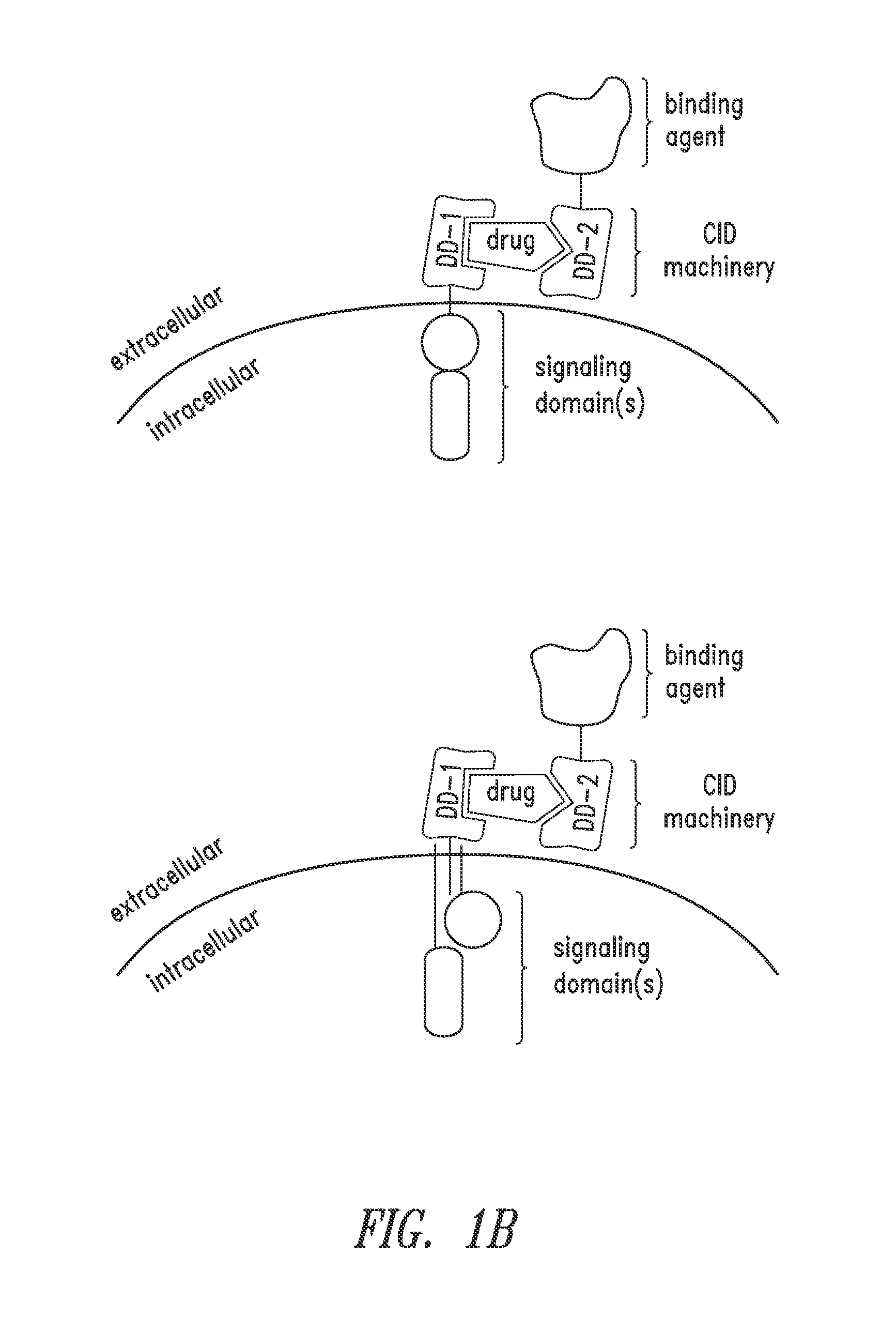
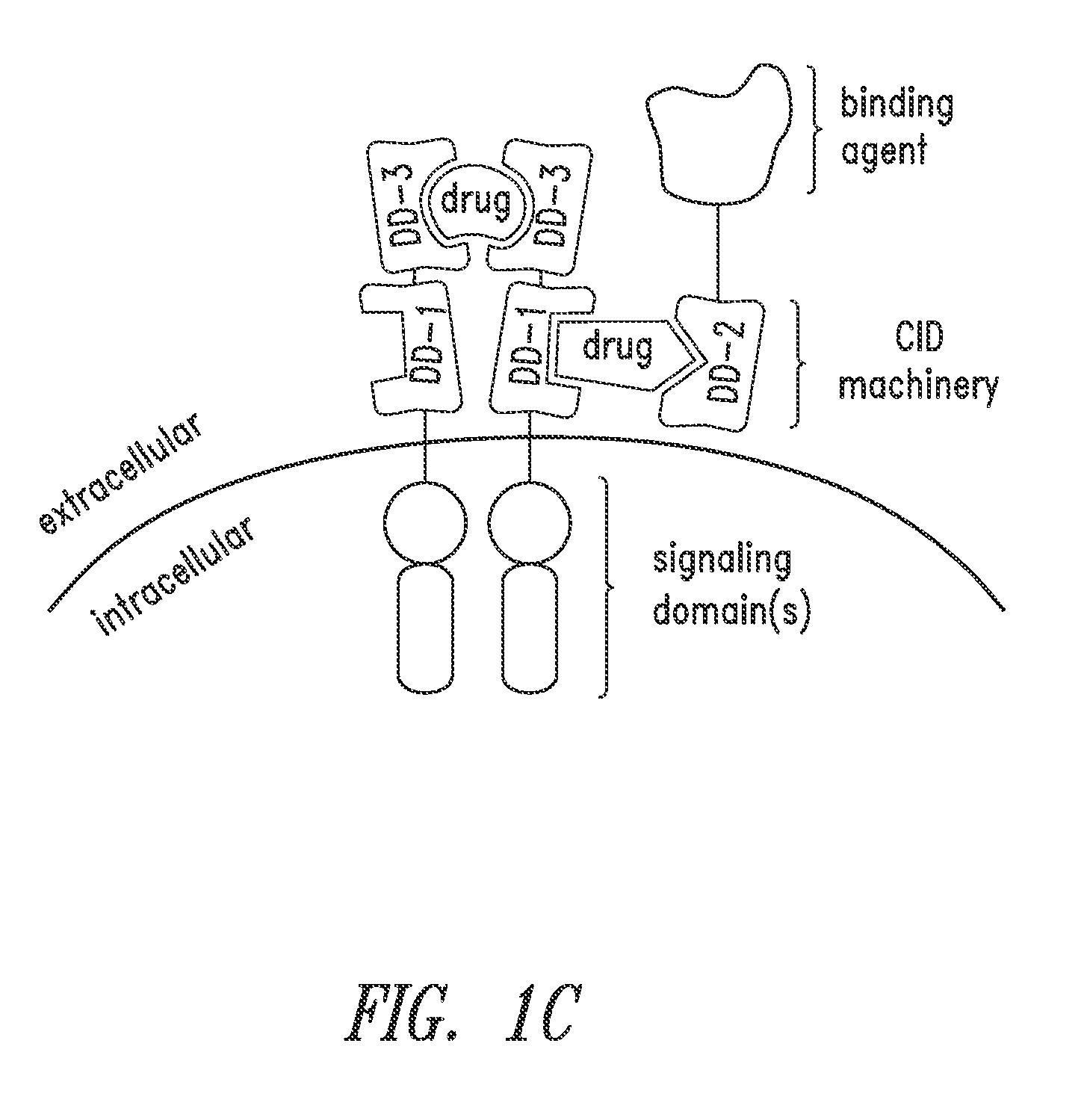
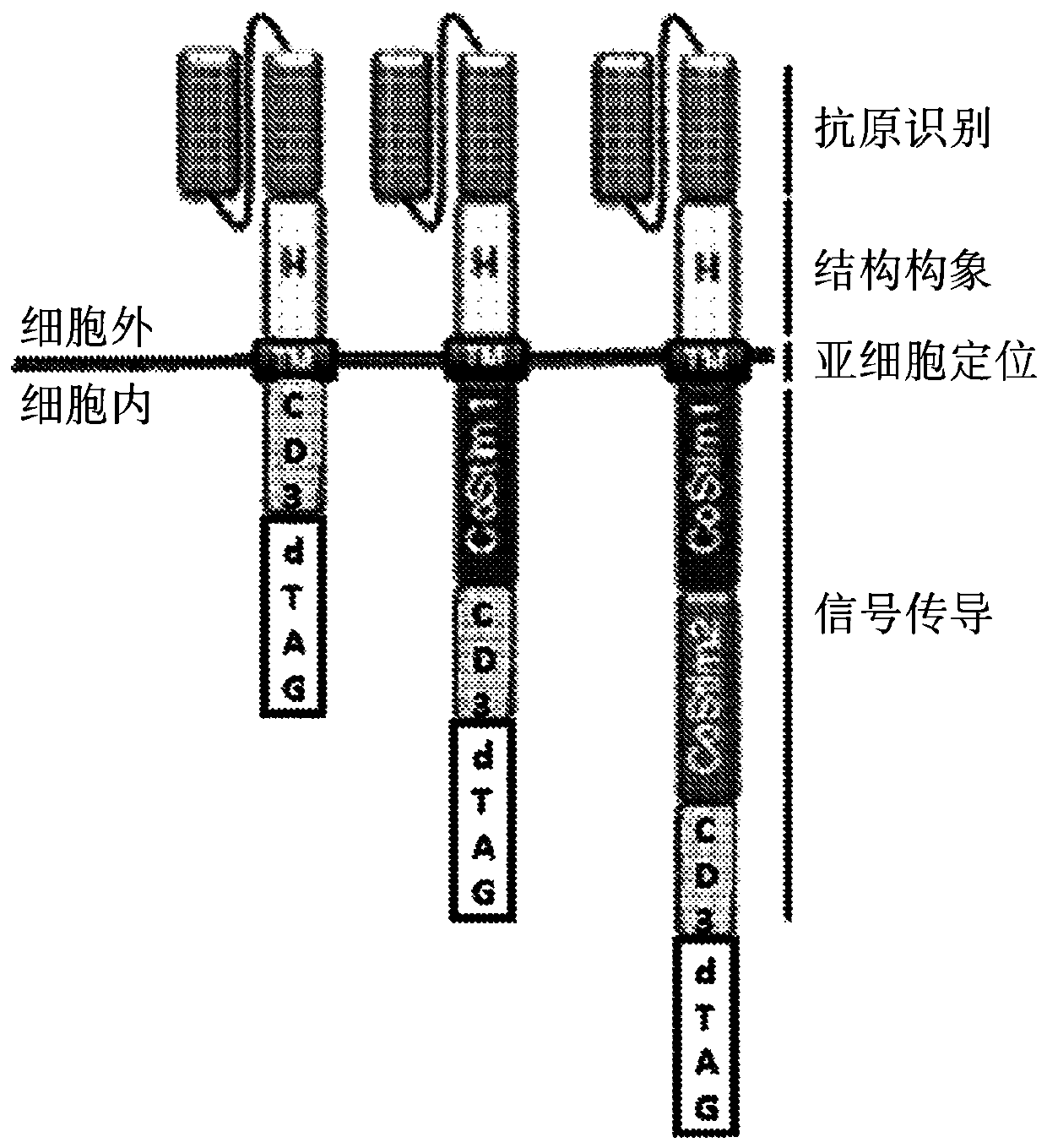
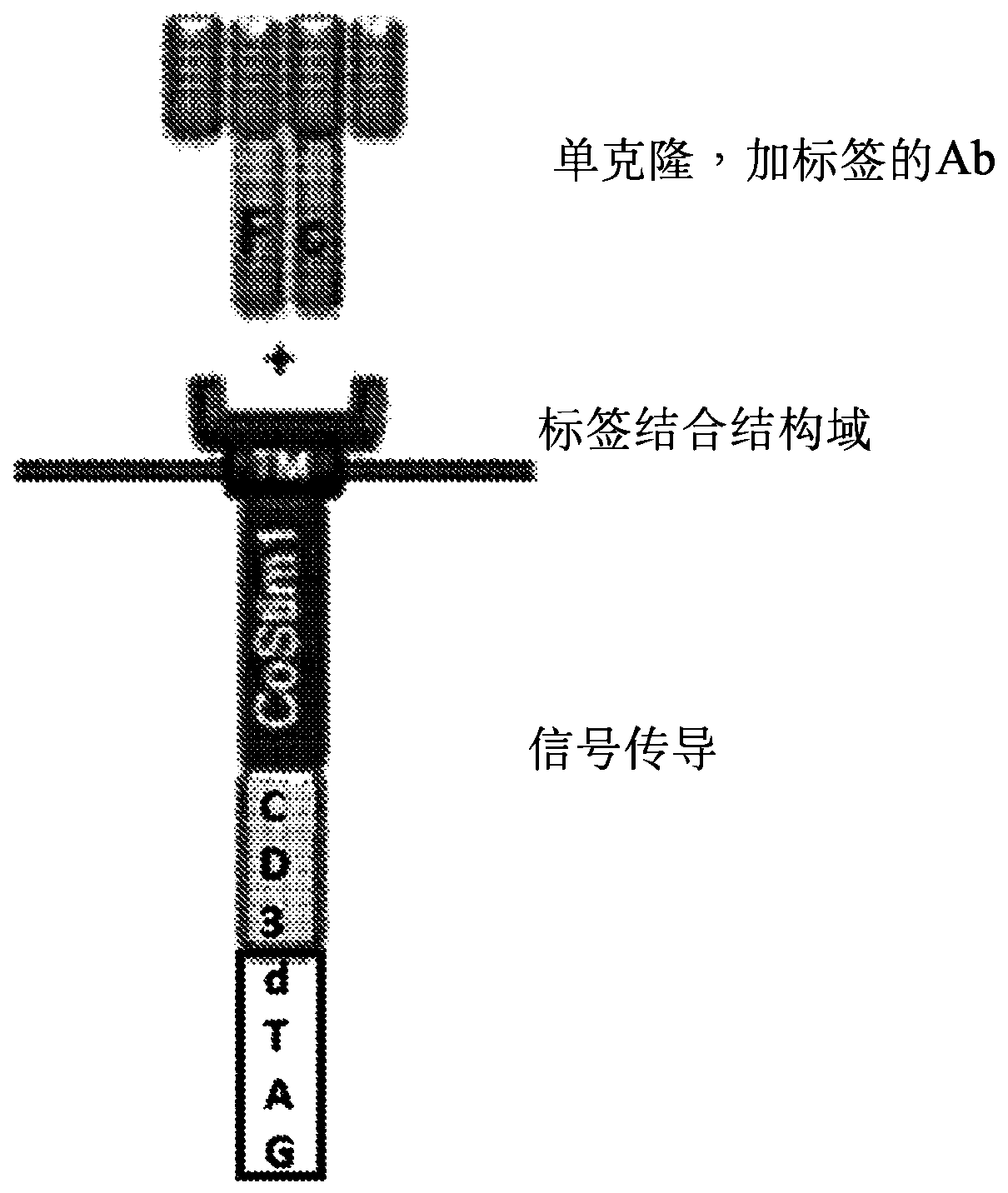
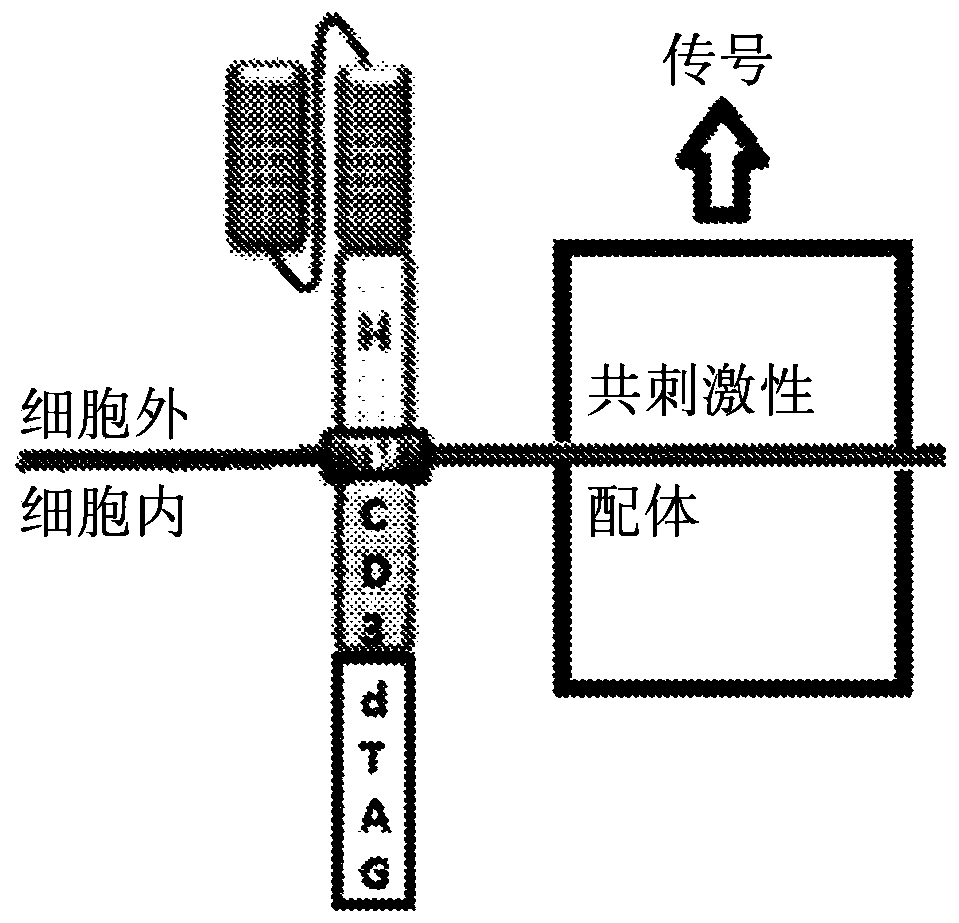
Multipartite signaling proteins and uses thereof
The present disclosure relates to compositions and methods for using cells having chemically-induced fusion protein complexes to spatially and temporally control immune cell signal initiation and downstream responses for treating disease.
Owner:2SEVENTY BIO INC
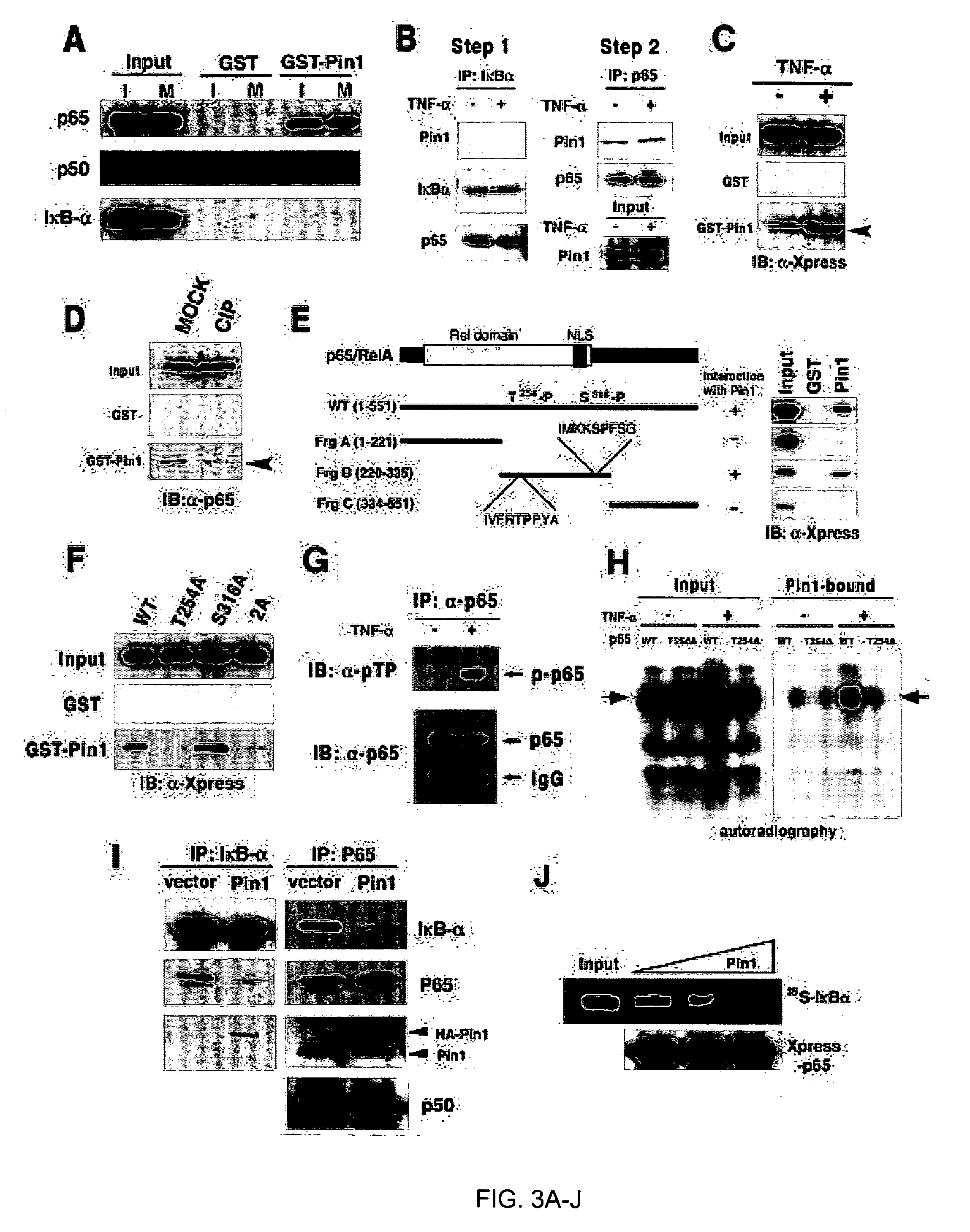
Novel regulatory mechanisms of NF-kappaB
InactiveUS20050147608A1Inhibiting degredation of NF-kBIncreasing nuclear accumulation and protein stabilityHydrolasesAntipyreticIsomerizationProteolysis
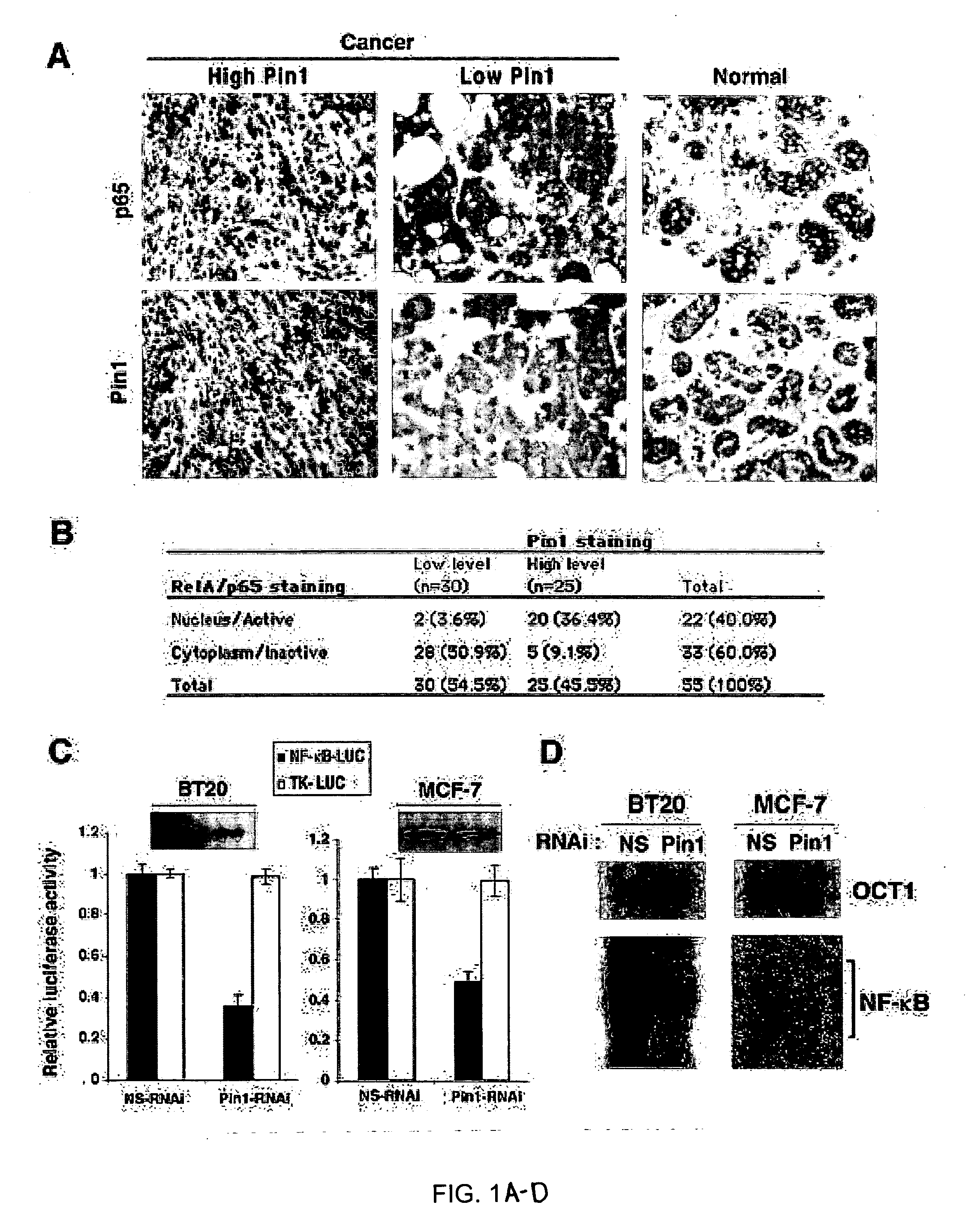
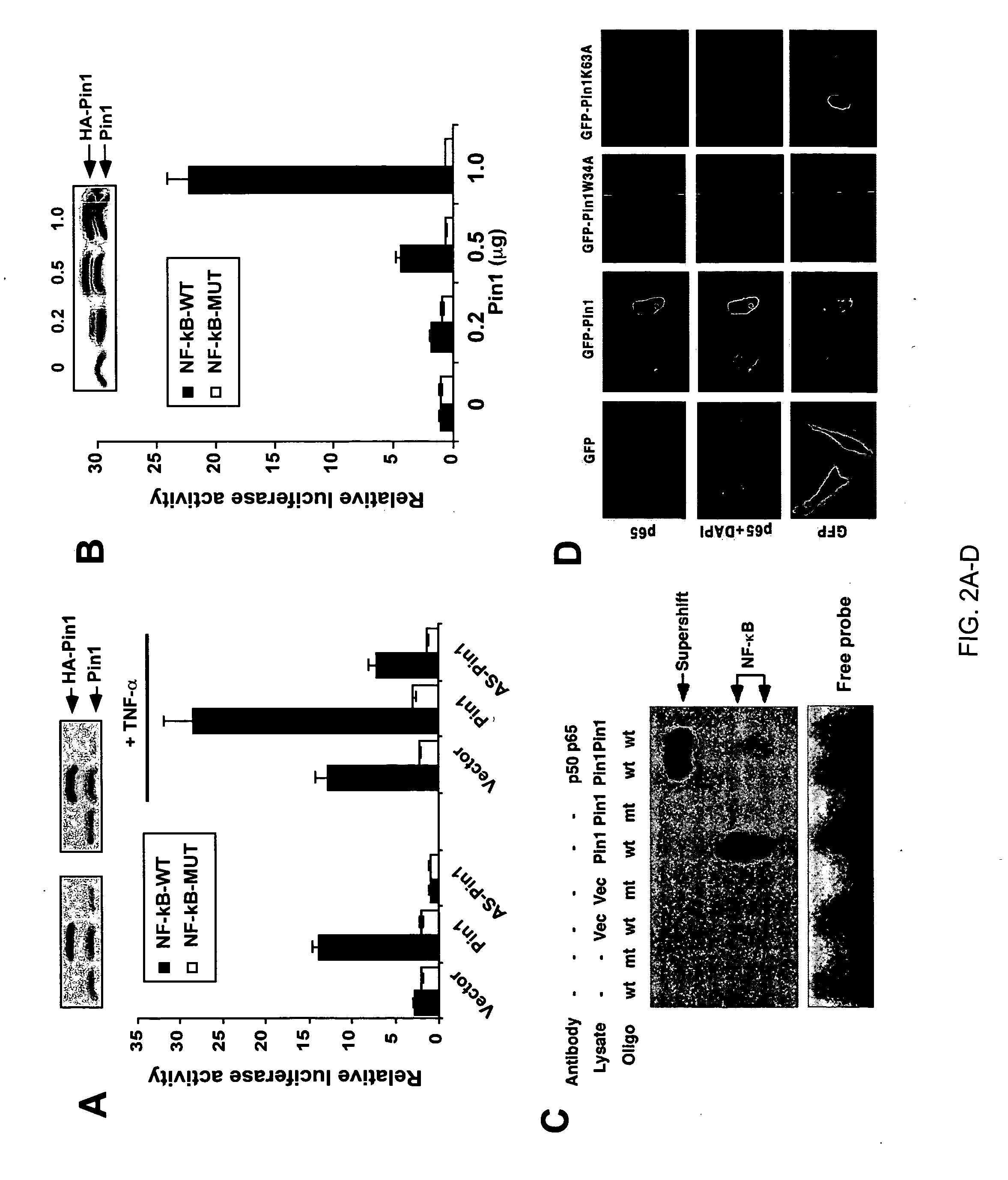
The instant invention pertains to the discovery of two novel regulatory mechanisms of NF-kB. The instant invention demonstrates that NF-kB is regulated by Pin1-catalyzed prolyl isomerization and ubiquitin-mediated proteolysis of p65. Accordingly, the instant invention provides methods for regulating NF-kB, and diseases and disorders associated with NF-kB. Further, the invention provides compositions capable of modulating the activity or expression of NF-kB, Pin1, and / or the proteolysis of p65.
Owner:BETH ISRAEL DEACONESS MEDICAL CENT INC
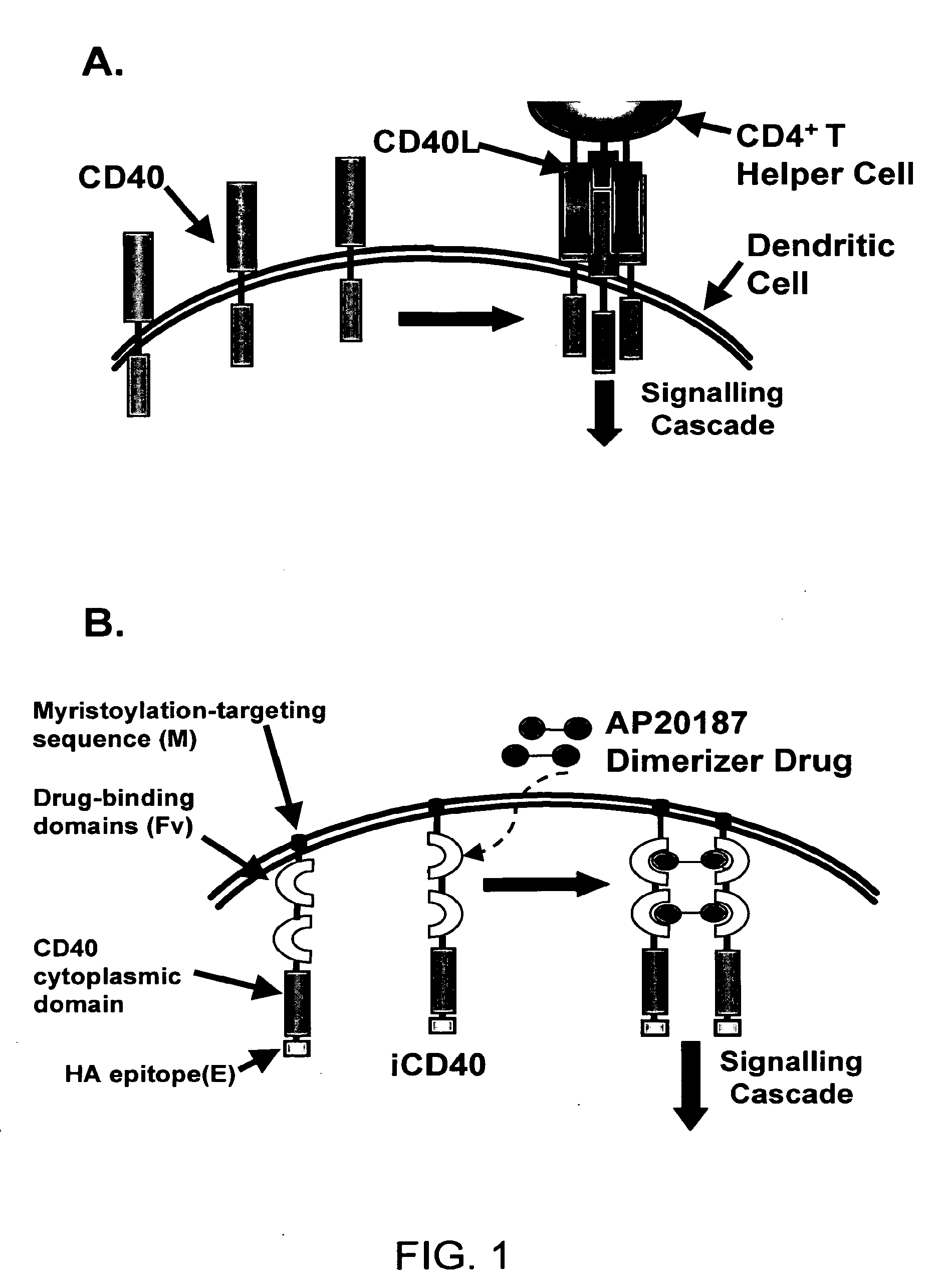
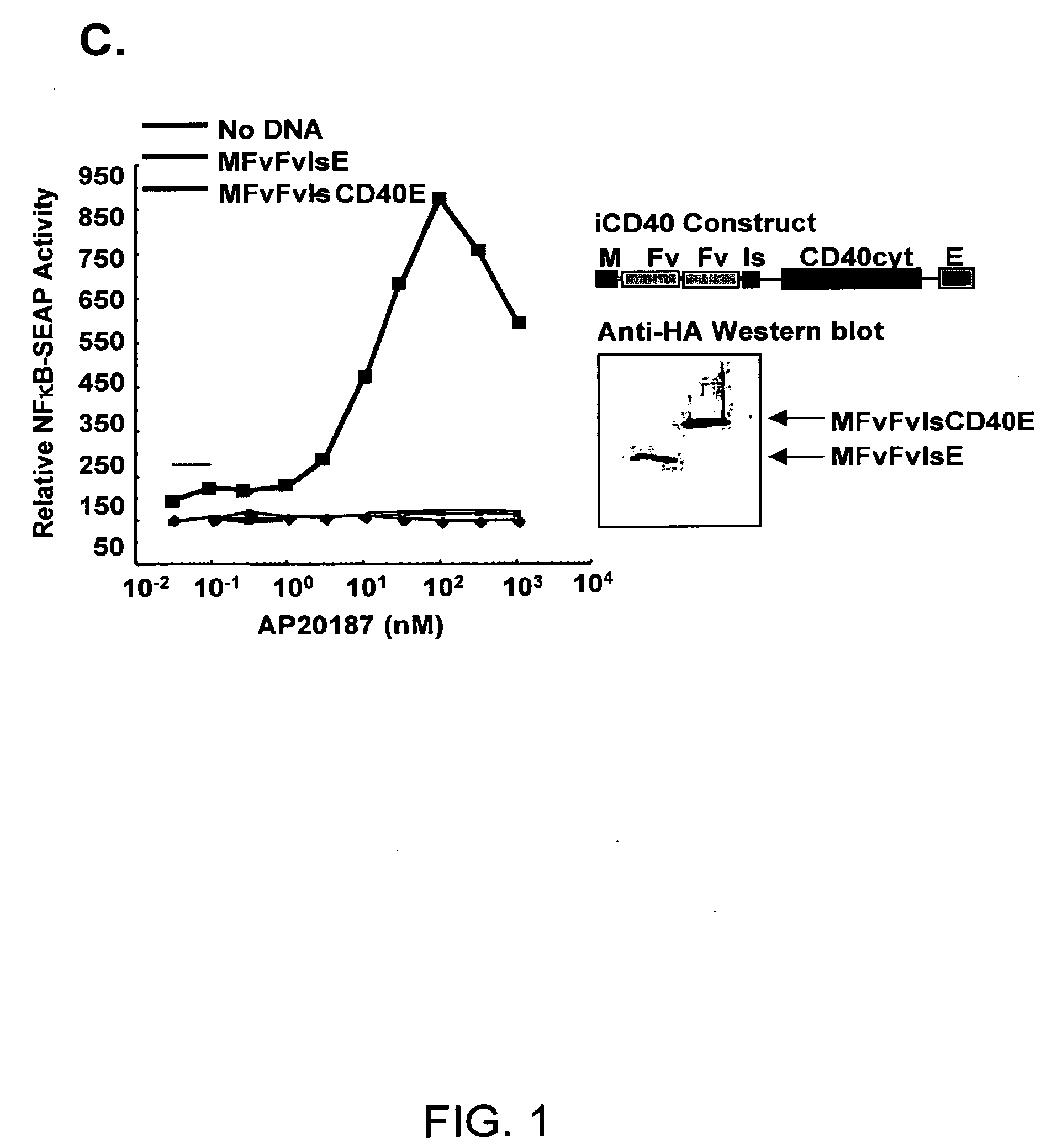
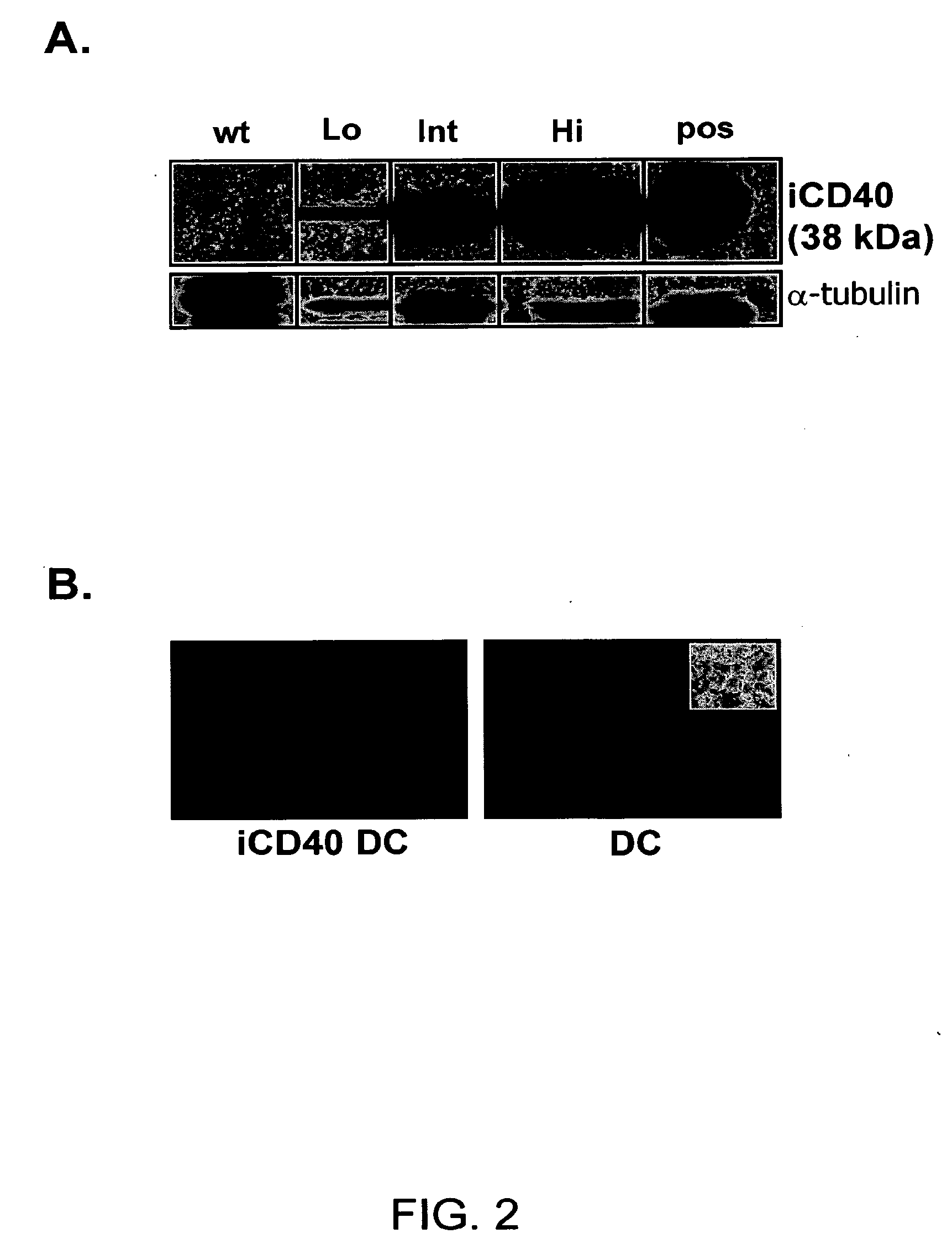
Induced activation in dendritic cells
Owner:BAYLOR COLLEGE OF MEDICINE
Modified gene resulting in parthenocarpic fruit set
ActiveUS20170335339A1Good effectMicrobiological testing/measurementCis-trans-isomerasesFruit setWild type
The invention relates to a modified PIN4 gene, the wild type of which is as identified in SEQ ID No. 1, encoding the protein of SEQ ID No. 5, or the wild type of which encodes a protein that has a sequence similarity of at least 80% to SEQ ID No. 5, which modified PIN4 gene encodes a protein that comprises an amino acid change as a result of the modification, and which modified protein is capable of inducing parthenocarpic fruit set when present in a plant. The invention also relates to plants and fruits that carry the gene. Such plants are capable of parthenocarpic fruit set and the fruits do not contain seeds. The invention further relates to use of the gene in breeding and producing plants capable of parthenocarpic fruit set.
Owner:RIJK ZWAAN ZAADTEELT & ZAADHANDEL BV
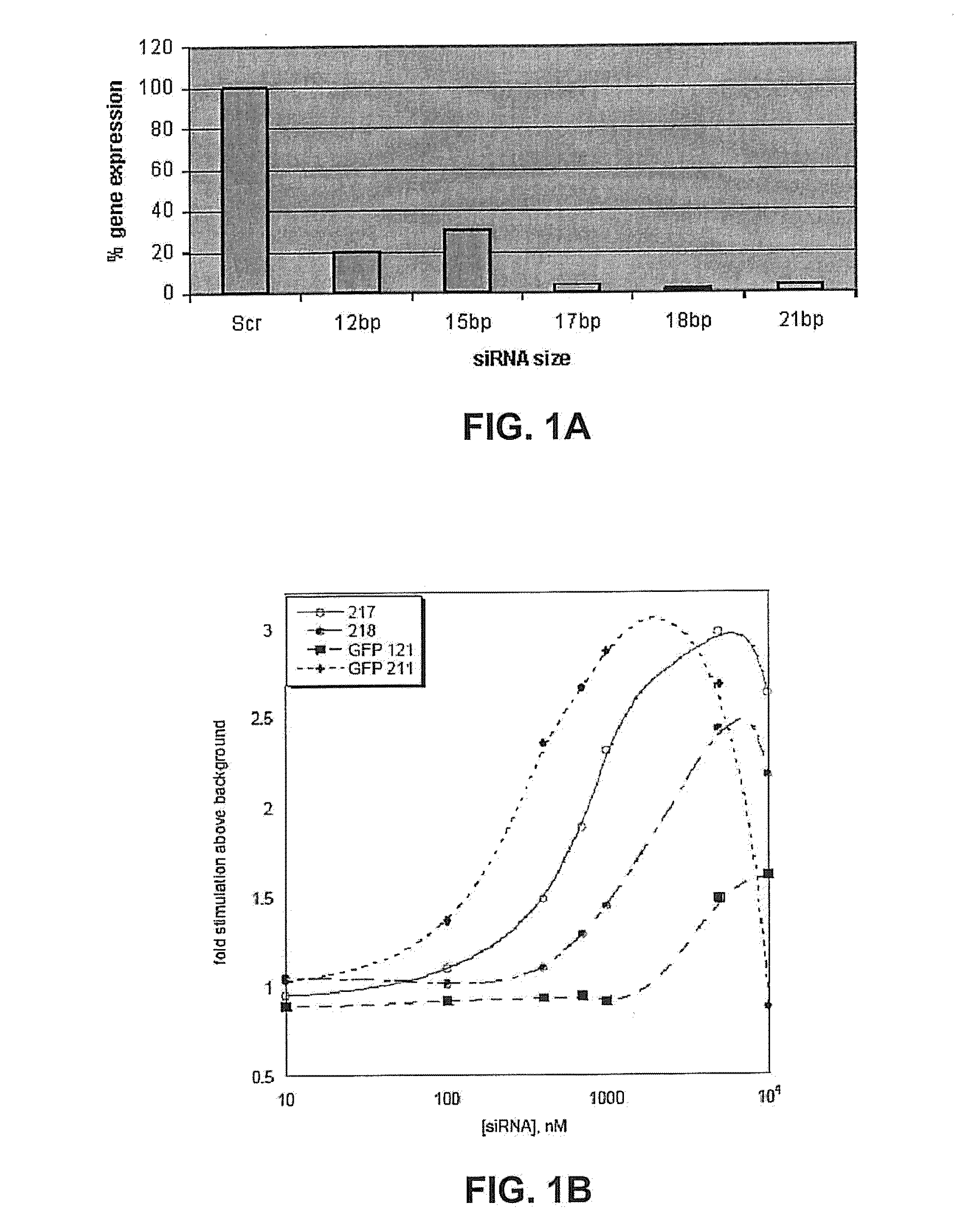
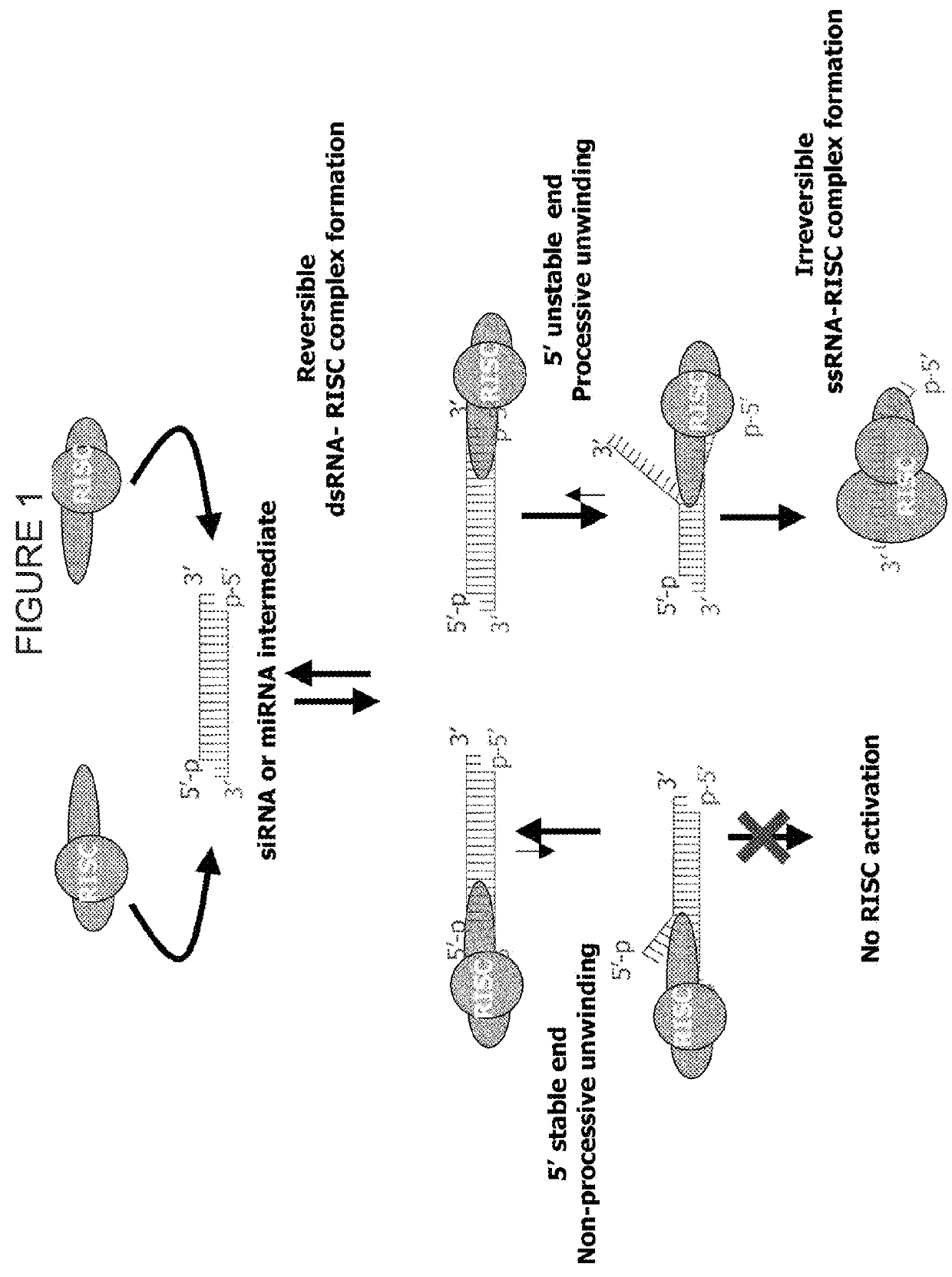
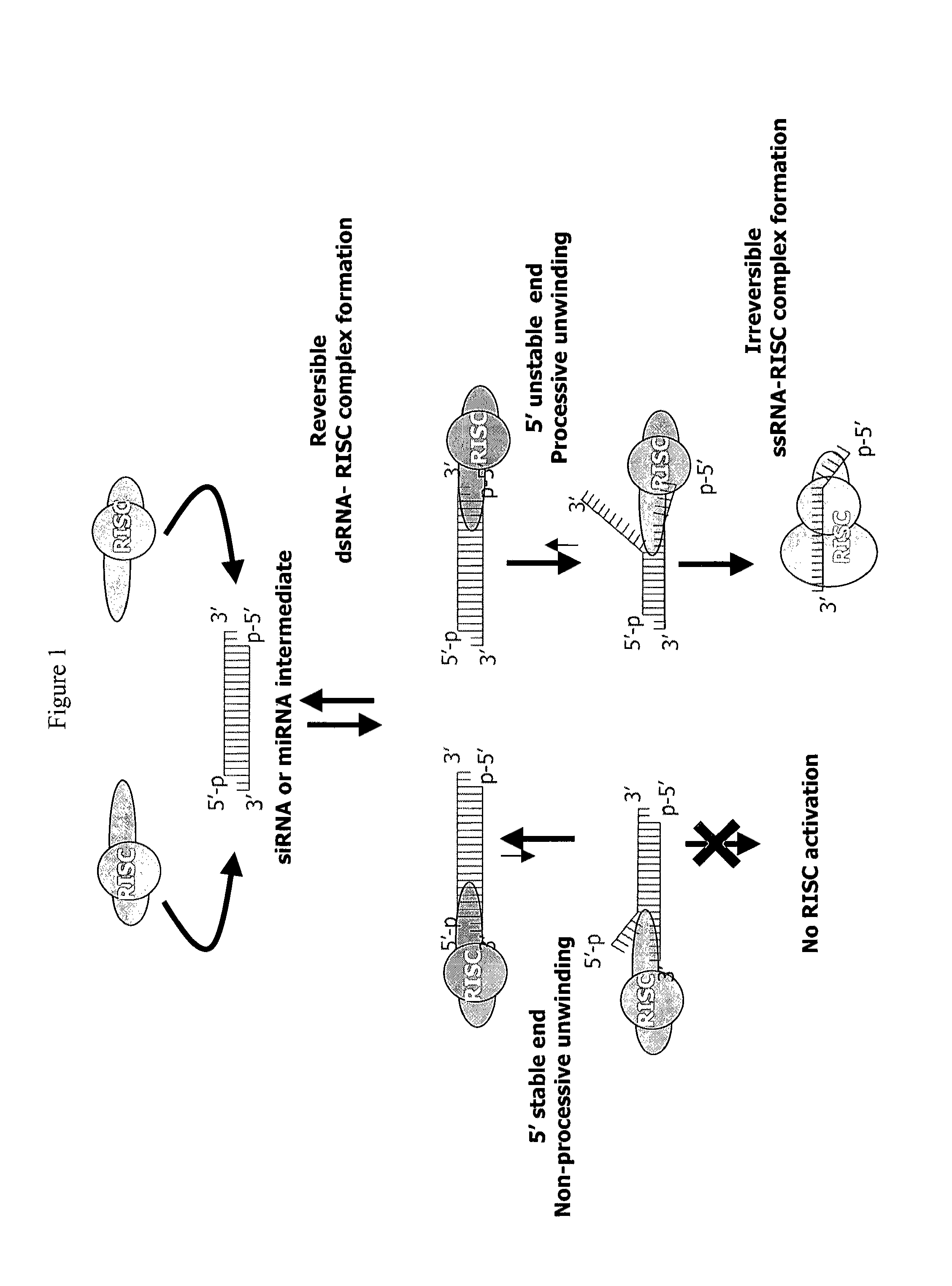
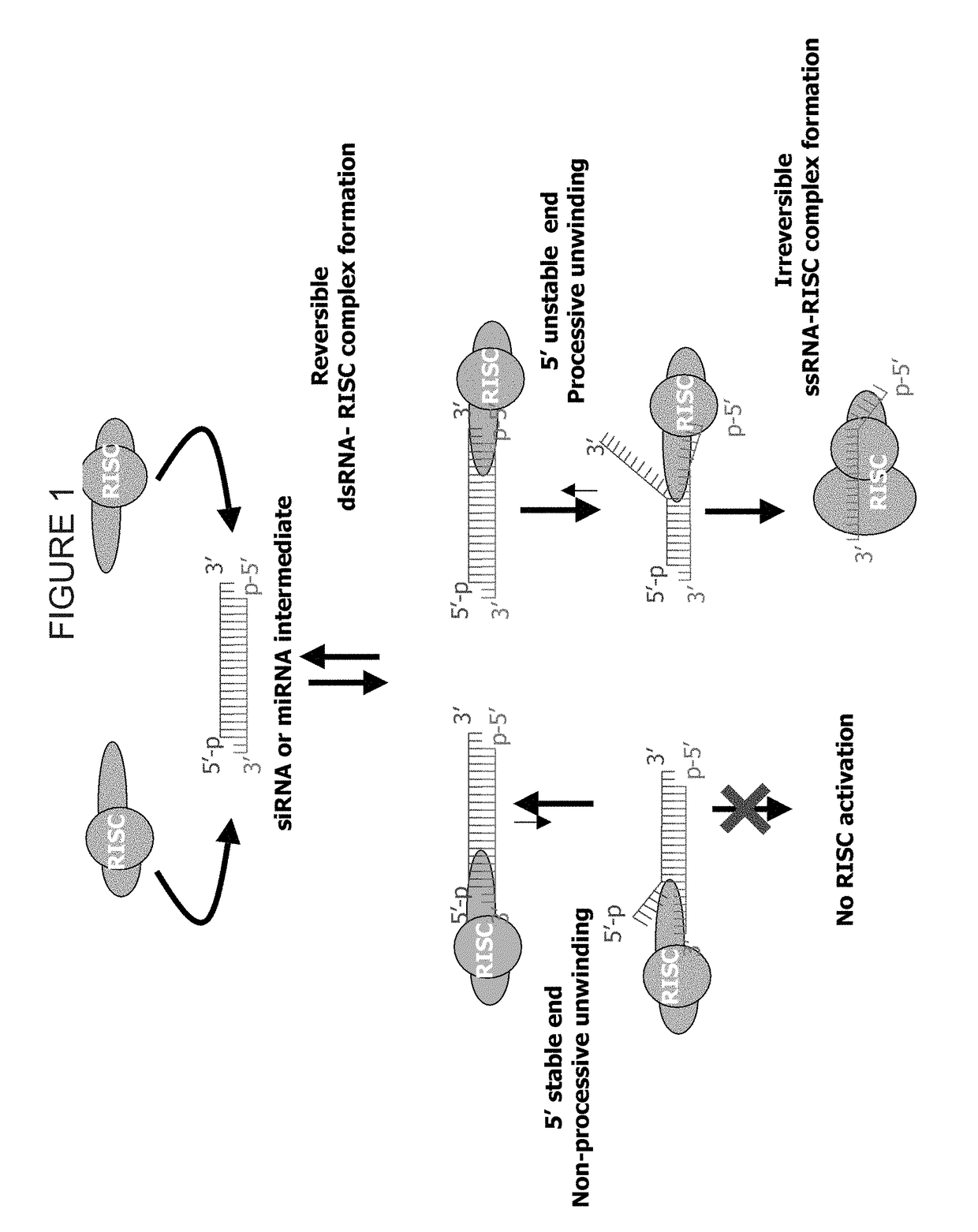
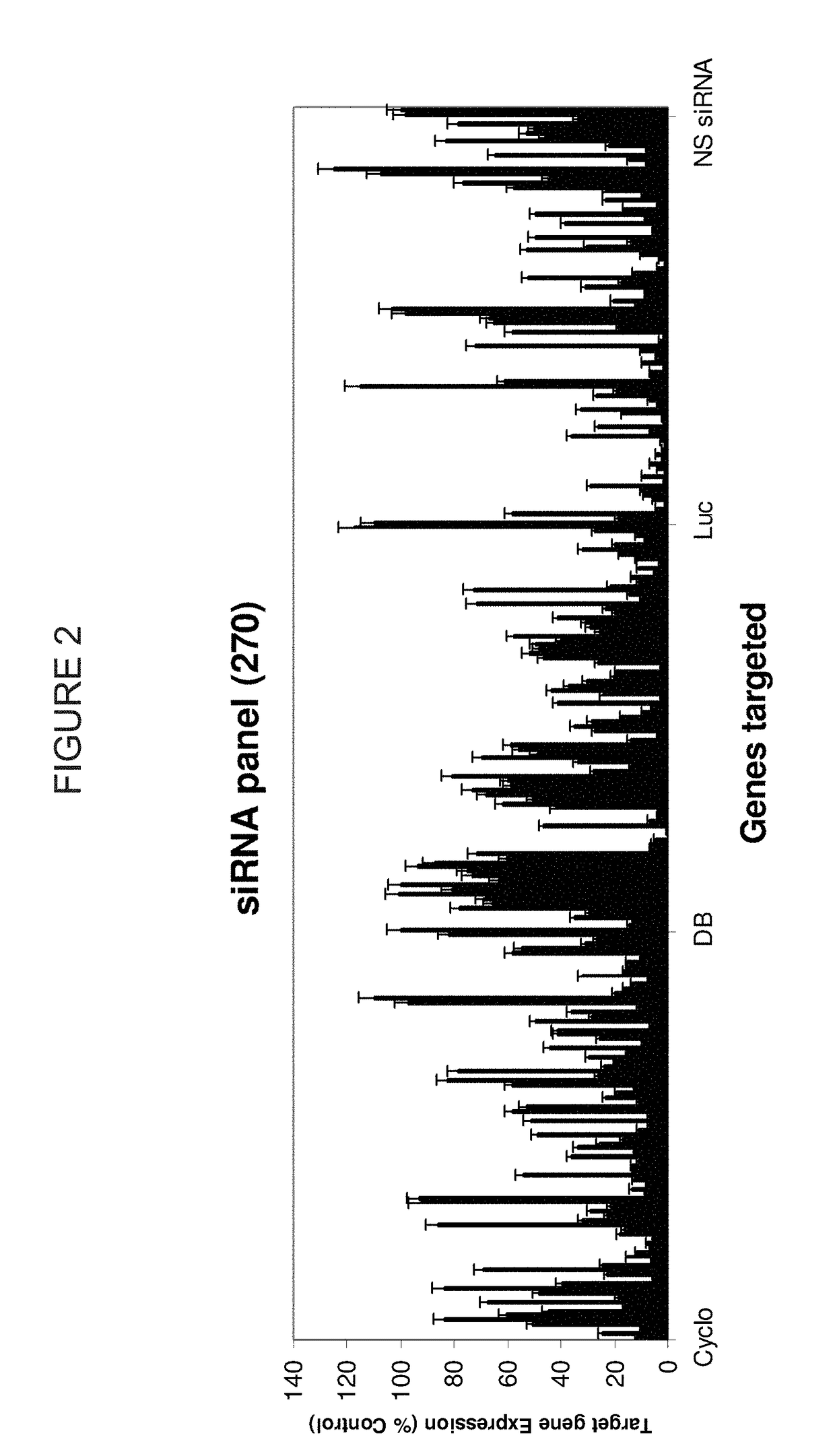
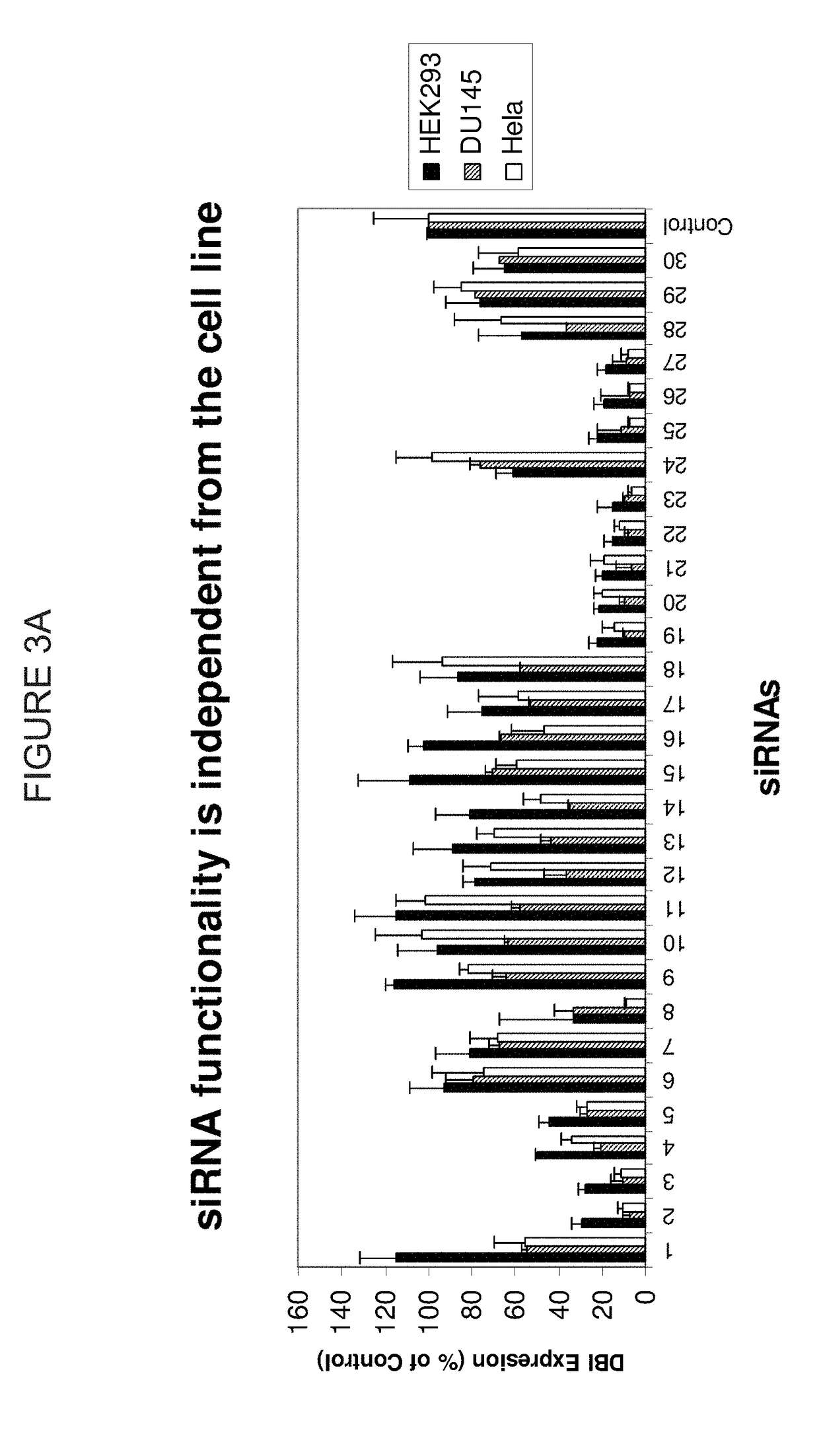
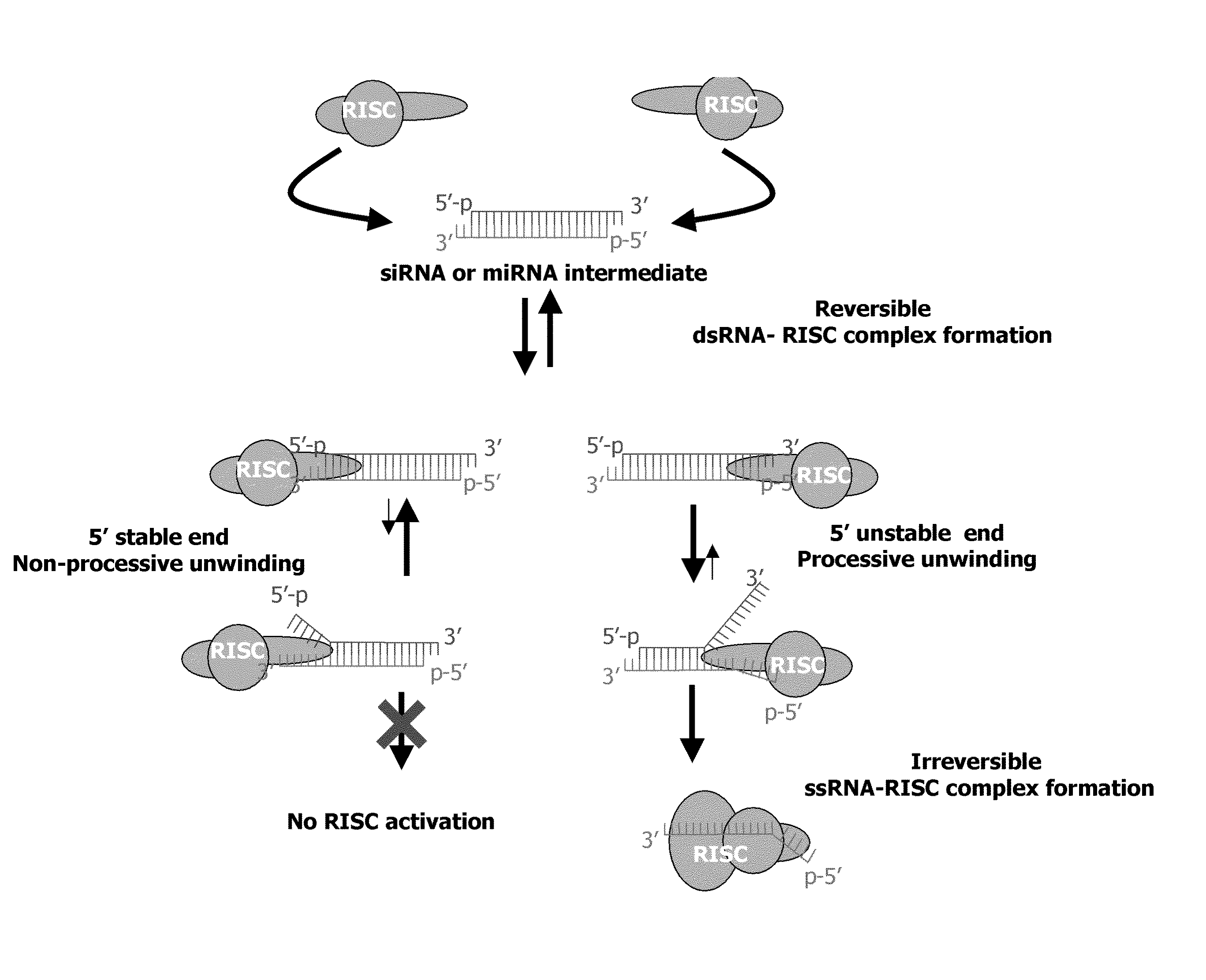
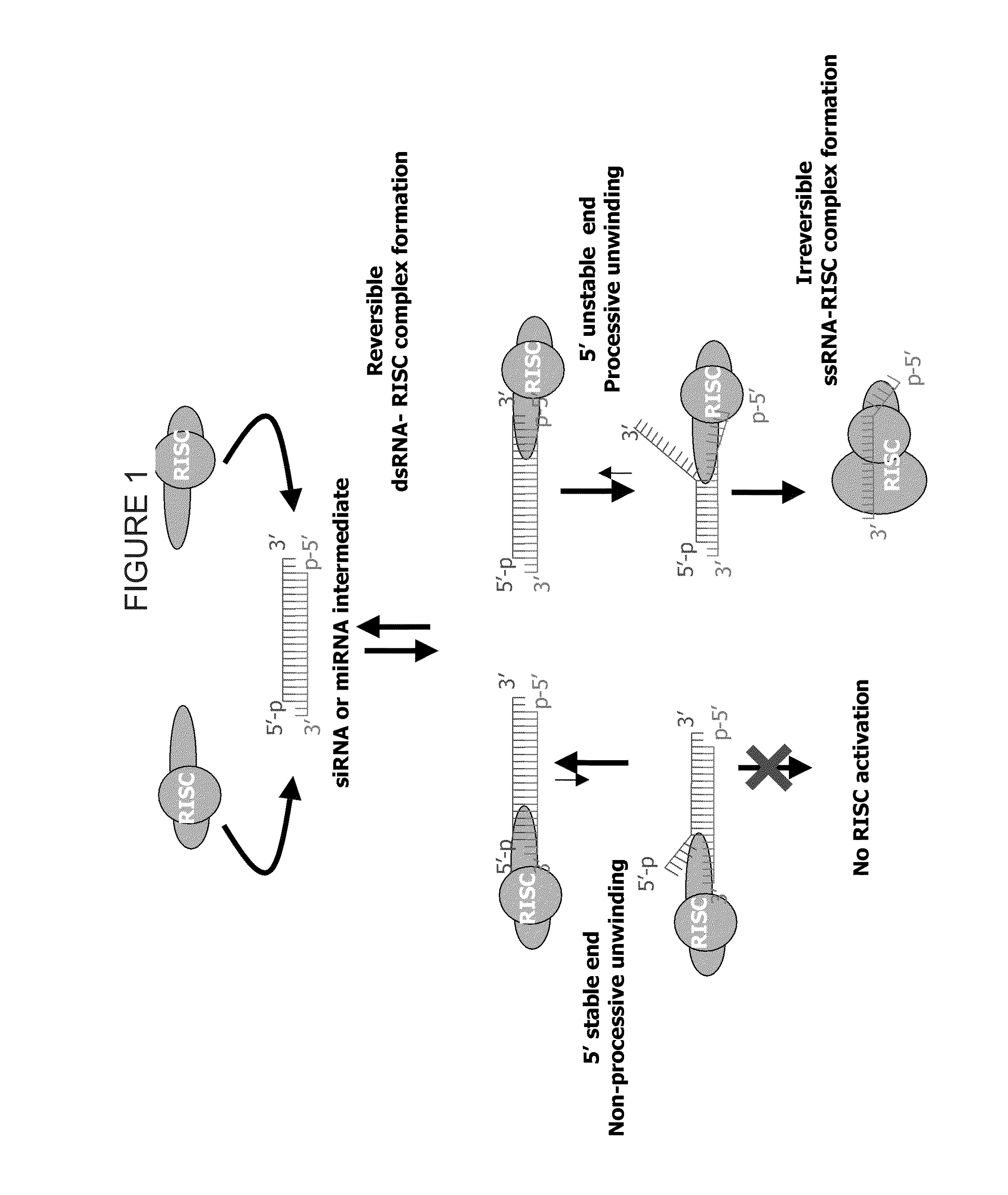
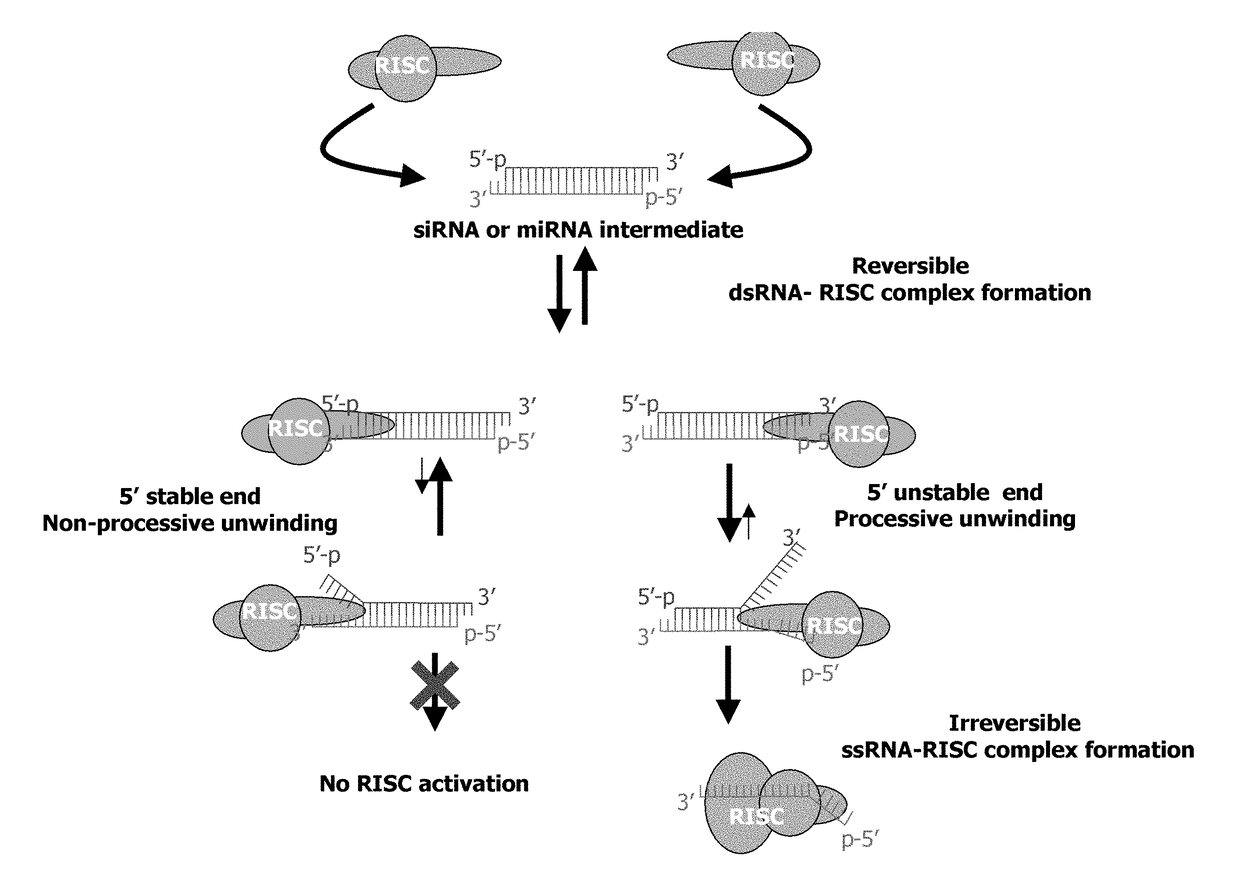
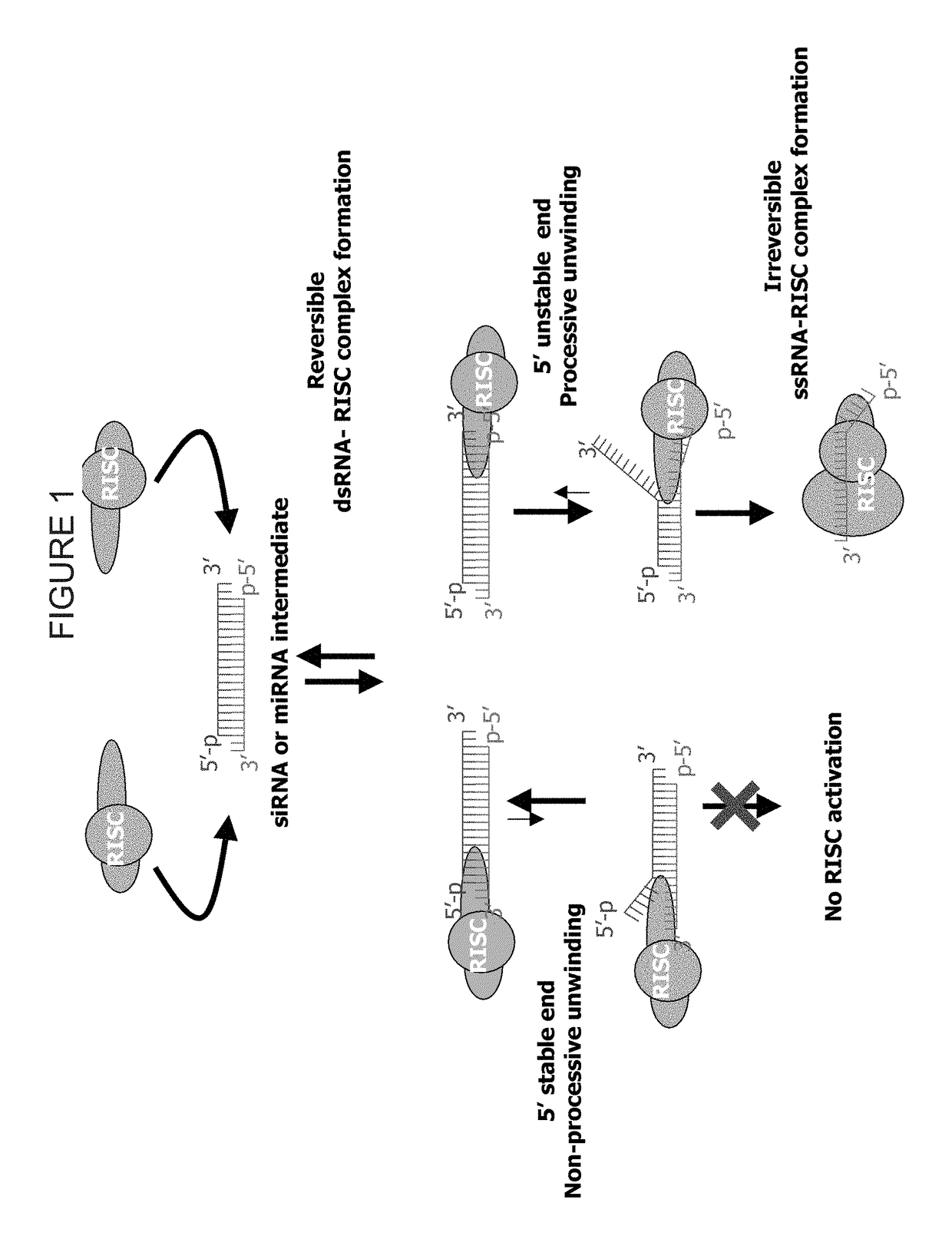
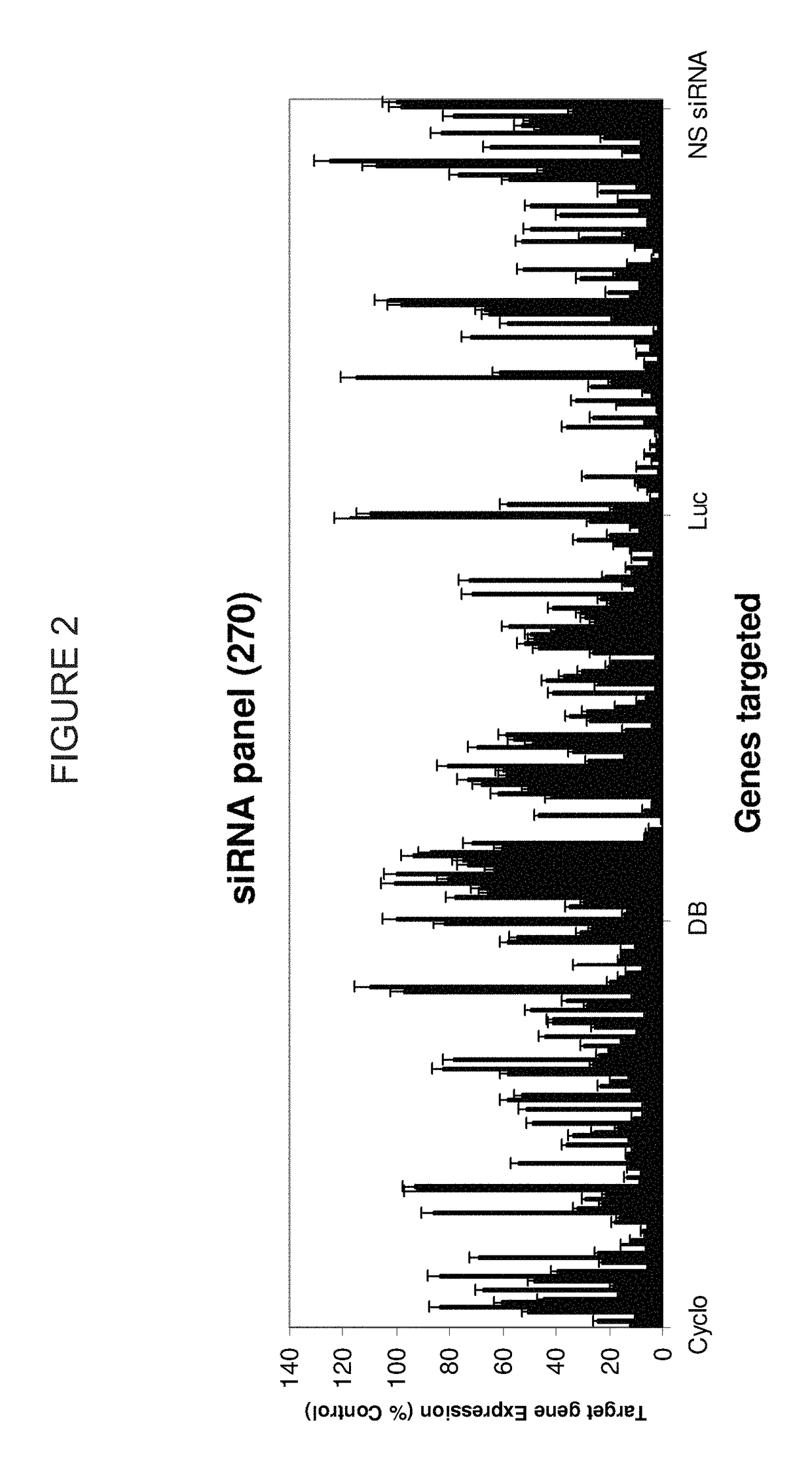
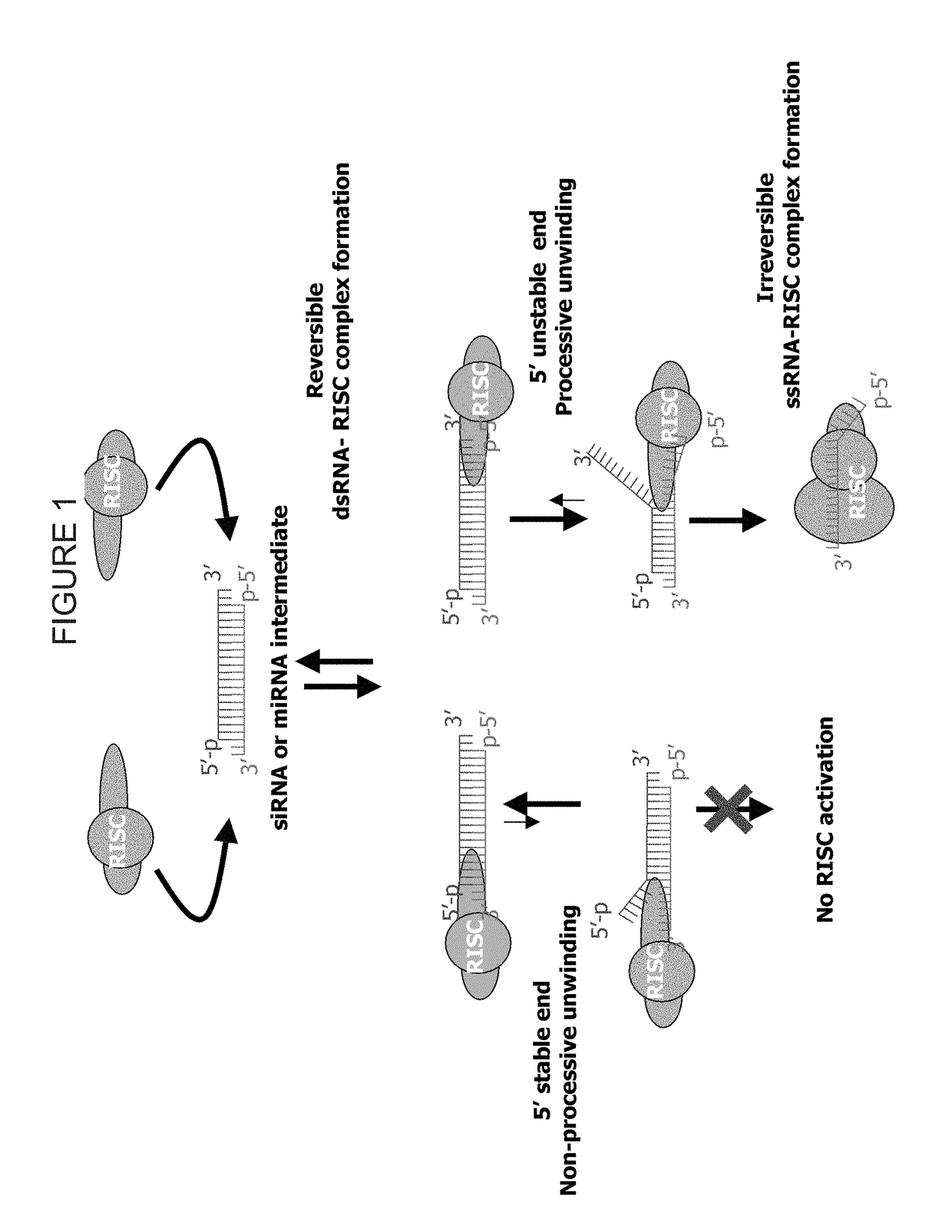
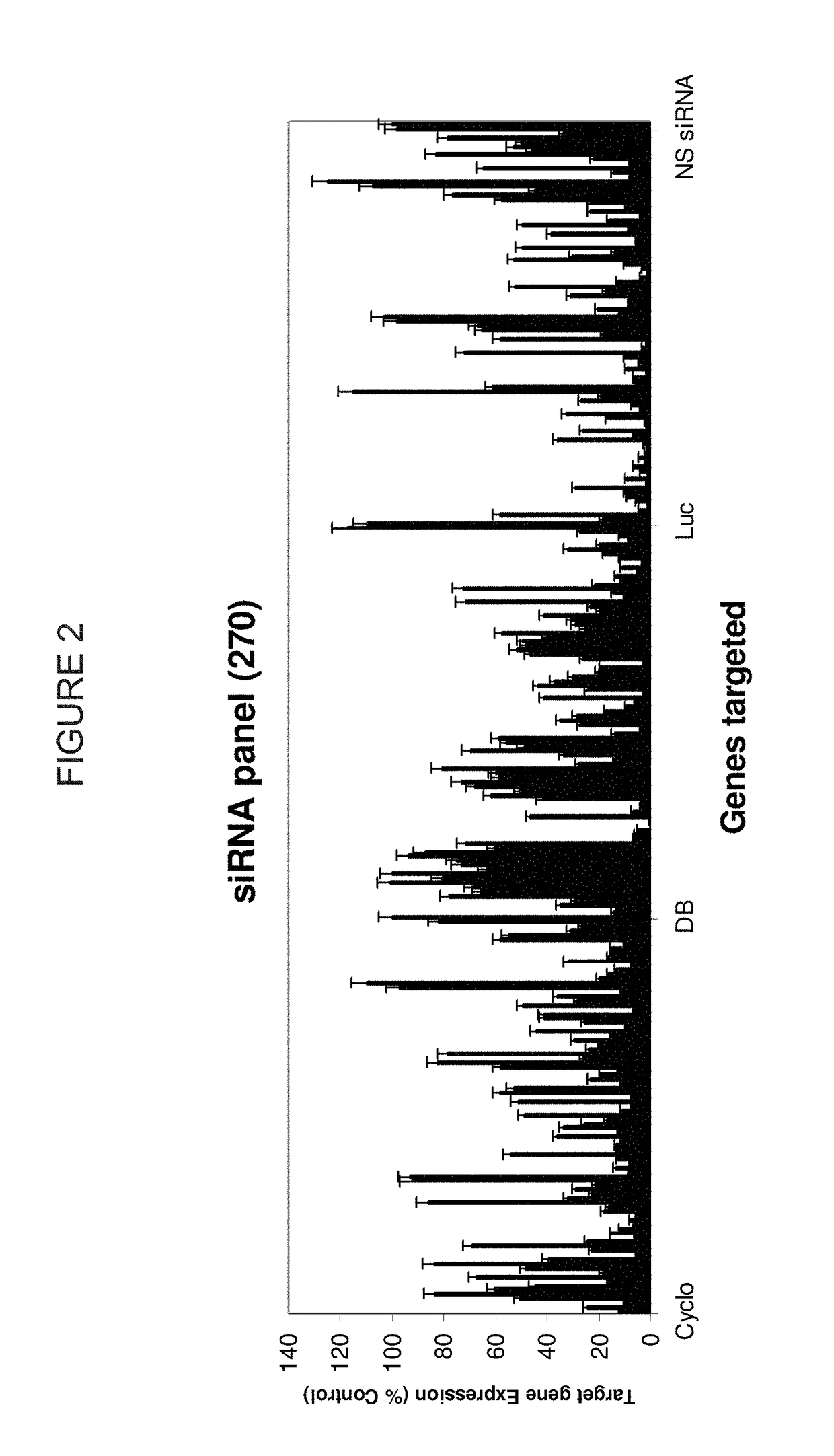
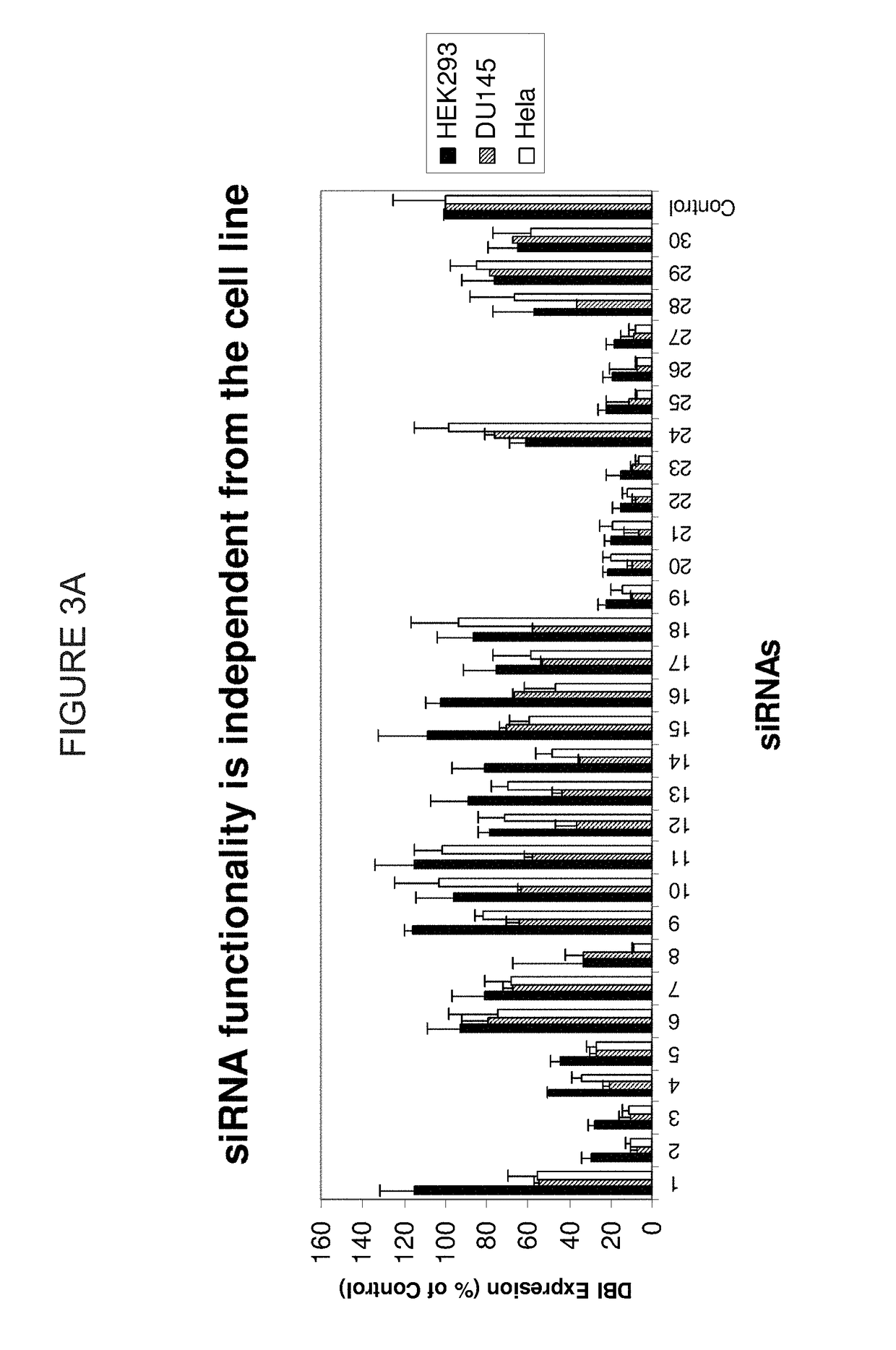
METHODS AND COMPOSITIONS CONCERNING siRNA'S AS MEDIATORS OF RNA INTERFERENCE
The present invention concerns an isolated siRNA of from about 5 to about 20 nucleotides that mediates RNA interference. Also disclosed are methods of reducing expression of a target gene in a cell comprising obtaining at least one siRNA of 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, or 20 basepairs in length; and delivering the siRNA into the cell. The siRNAs can be chemically synthesized RNA or an analog of a naturally occurring RNA.
Owner:APPL BIOSYSTEMS INC
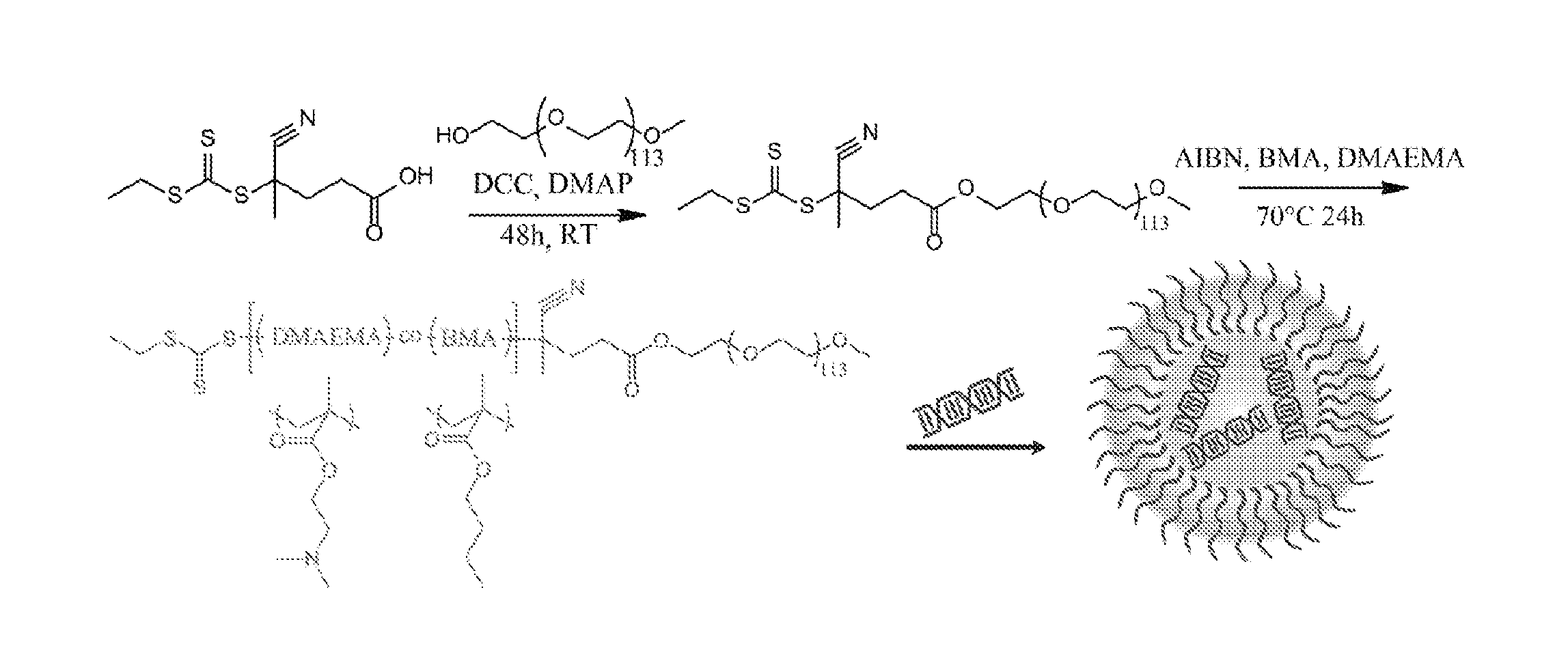
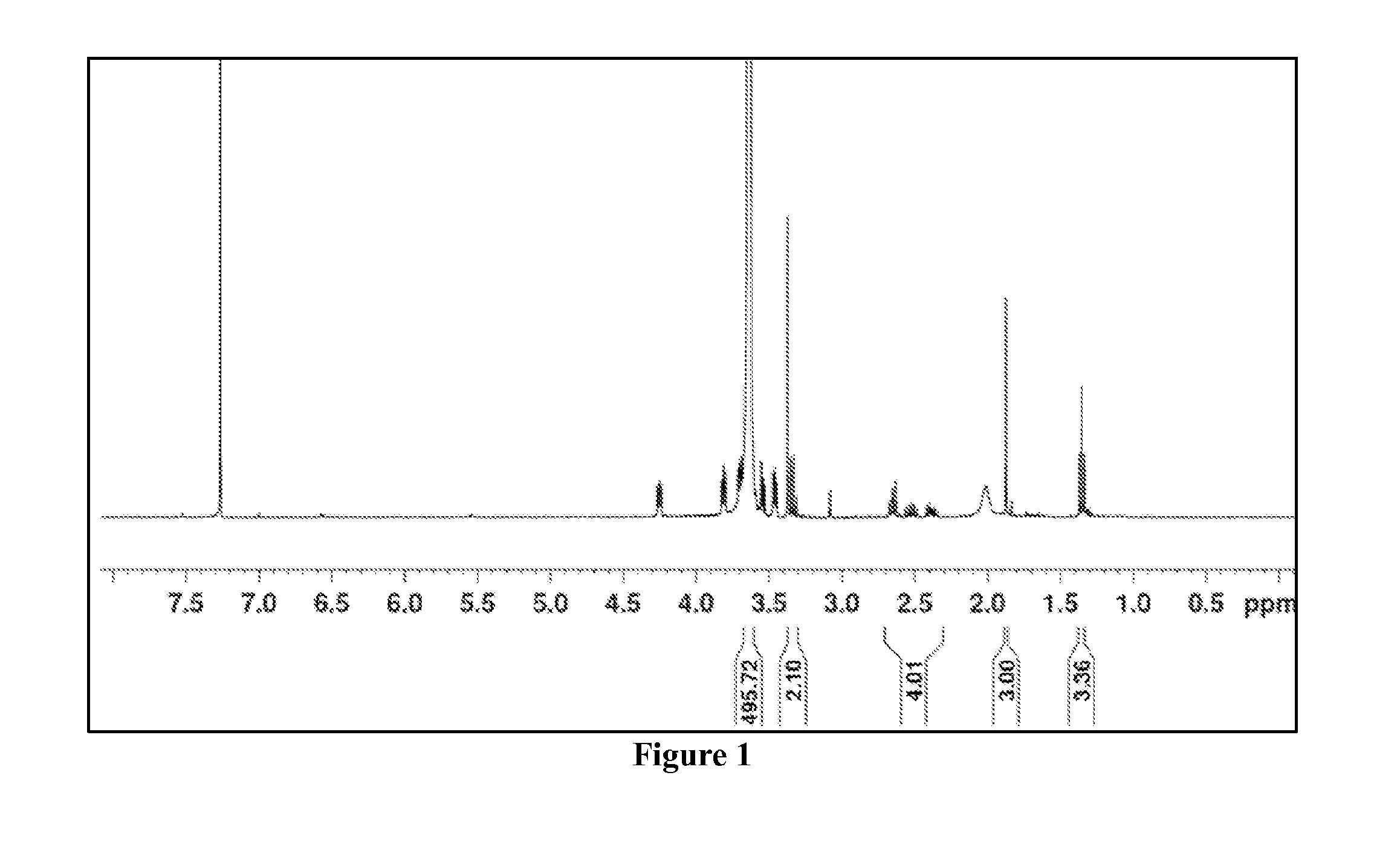
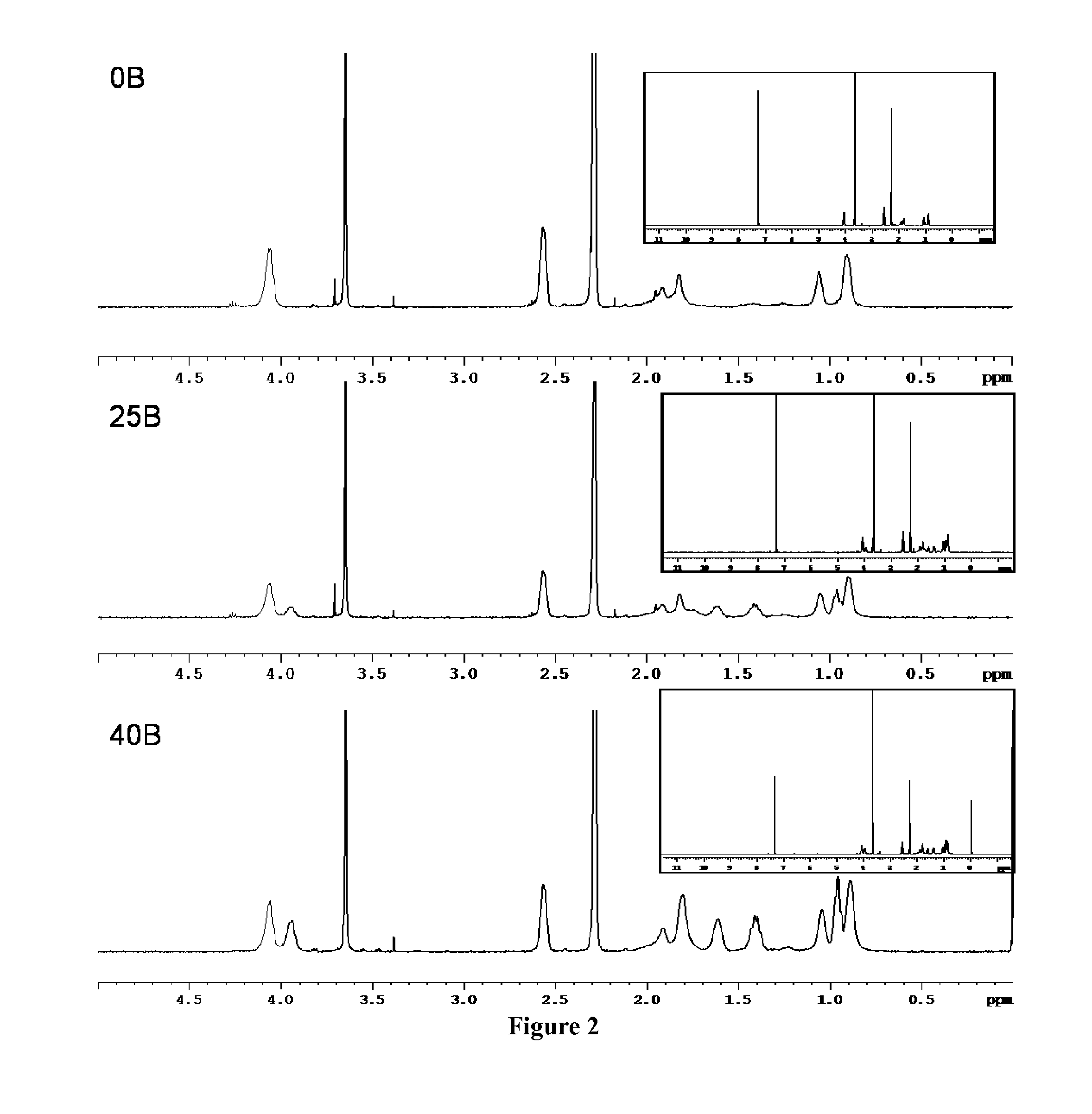
Polymeric nanoparticles
The presently-disclosed subject matter includes nanoparticles that comprise a plurality of assembled polymers. In some embodiments the polymers comprise a first block that includes hydrophilic monomers, the first block substantially forming an outer shell of the nanoparticle, and a second block that includes cationic monomers and hydrophobic monomers, the second block substantially forming a core of the nanoparticle. In some embodiments a polynucleotide is provided that is bound to the cationic monomers of the nanoparticle. The presently-disclosed subject matter also comprises methods for using the present nanoparticles to include RNAi in a cell as well as methods for making the present nanoparticles.
Owner:VANDERBILT UNIV
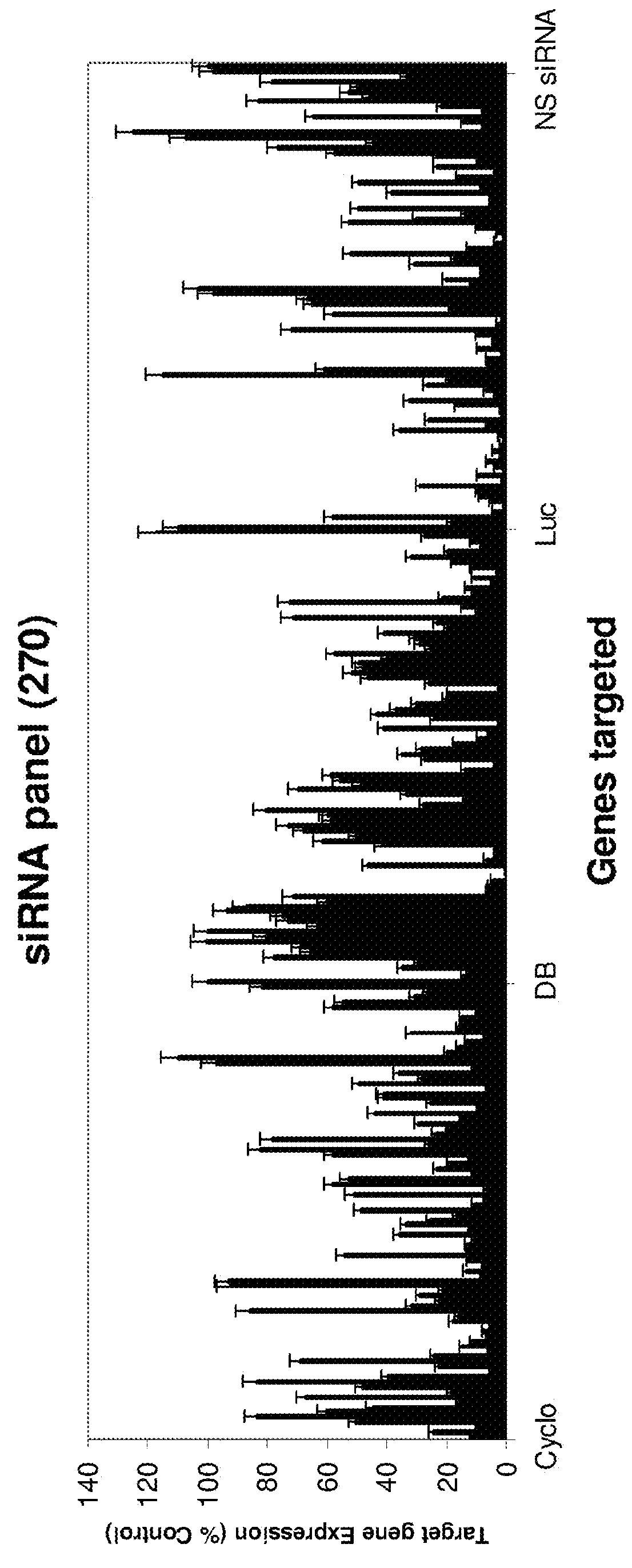
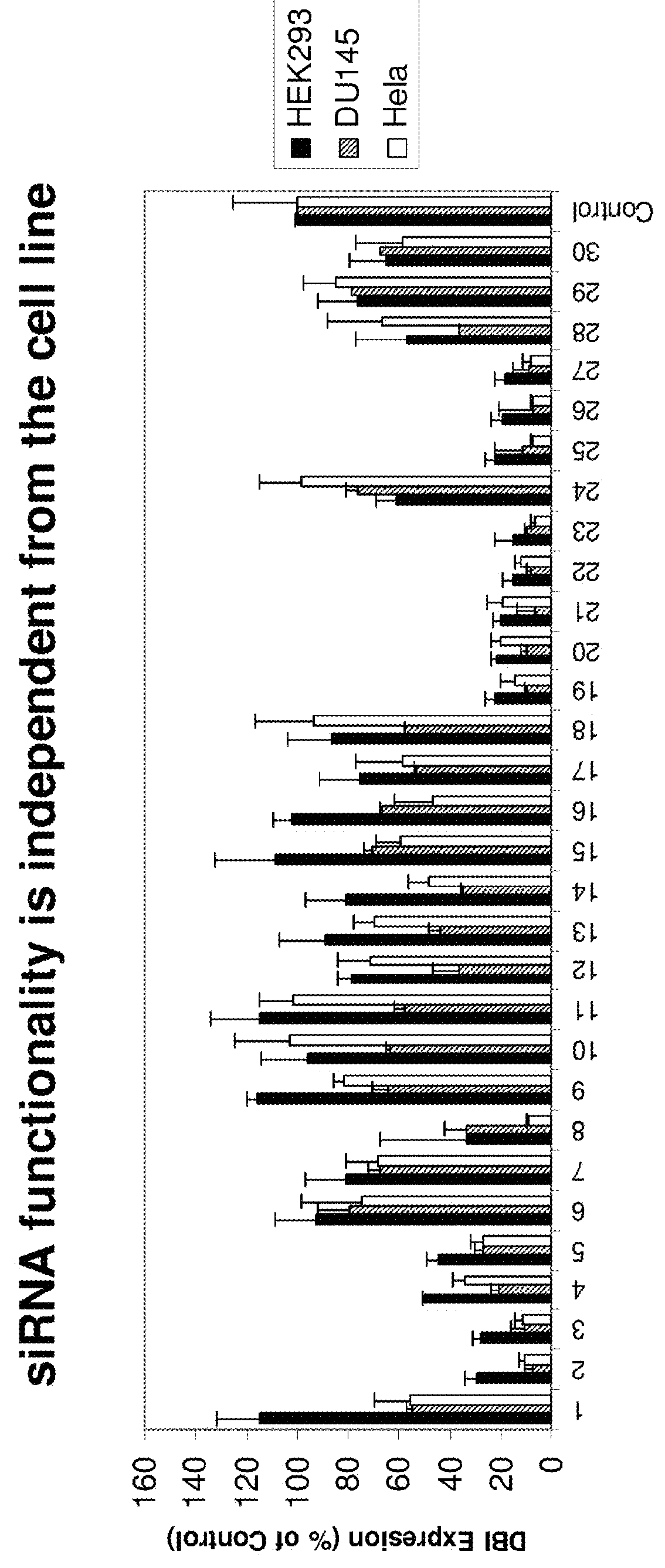
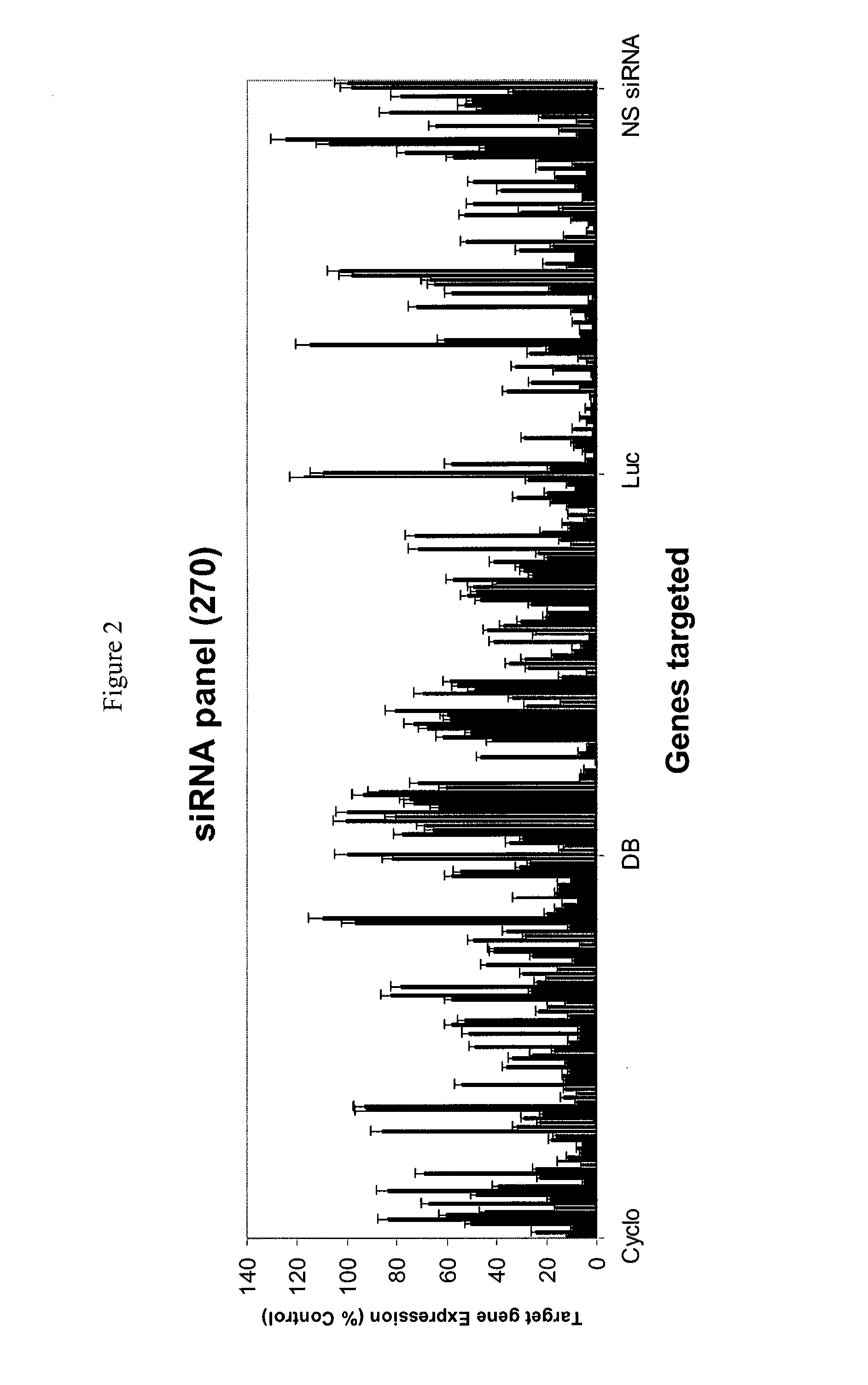
Methods and compositions for selecting siRNA of improved functionality
ActiveUS9228186B2Improve efficiencyGood curative effectOrganic active ingredientsSugar derivativesNucleotideGene silencing
Owner:THERMO FISHER SCIENTIFIC INC
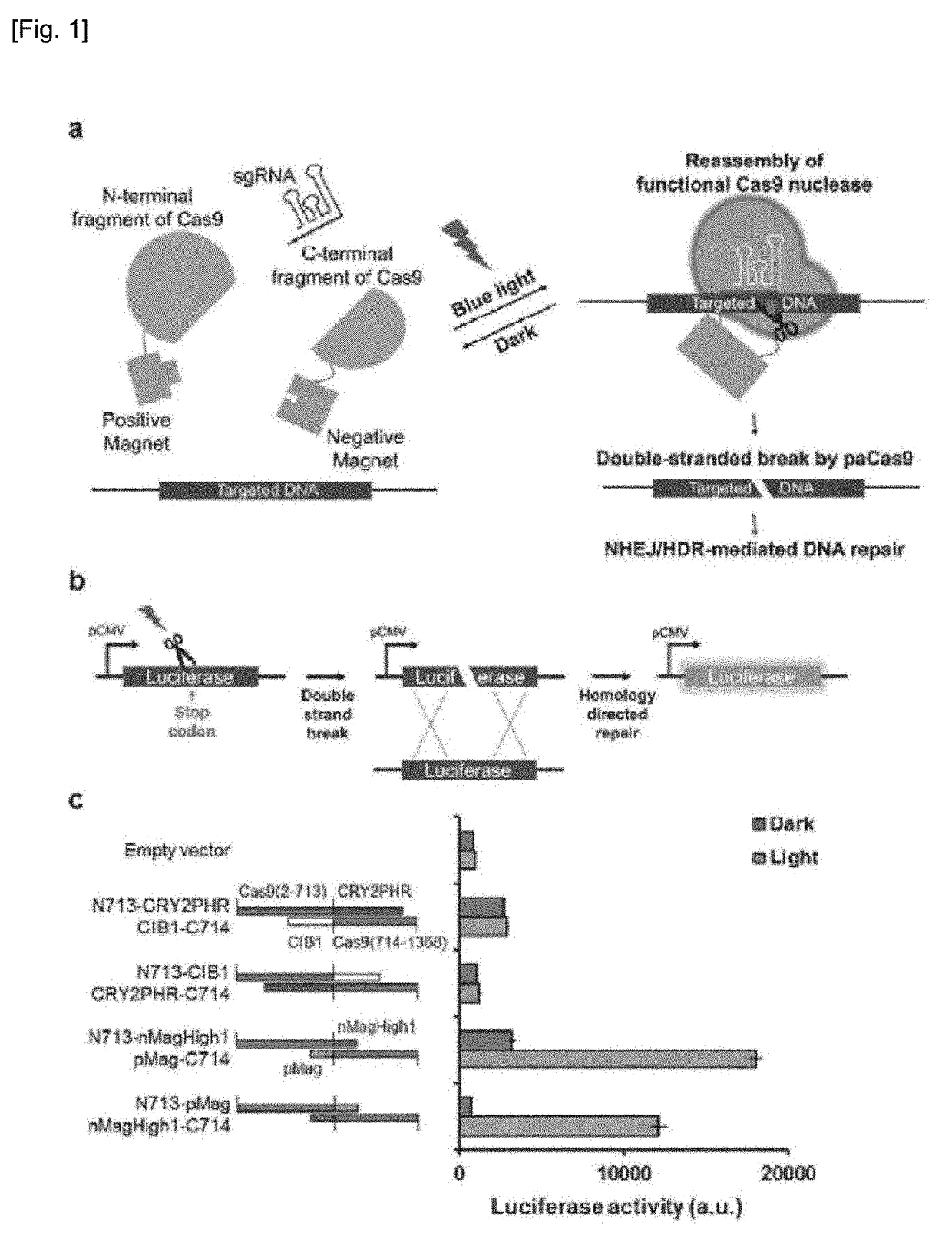
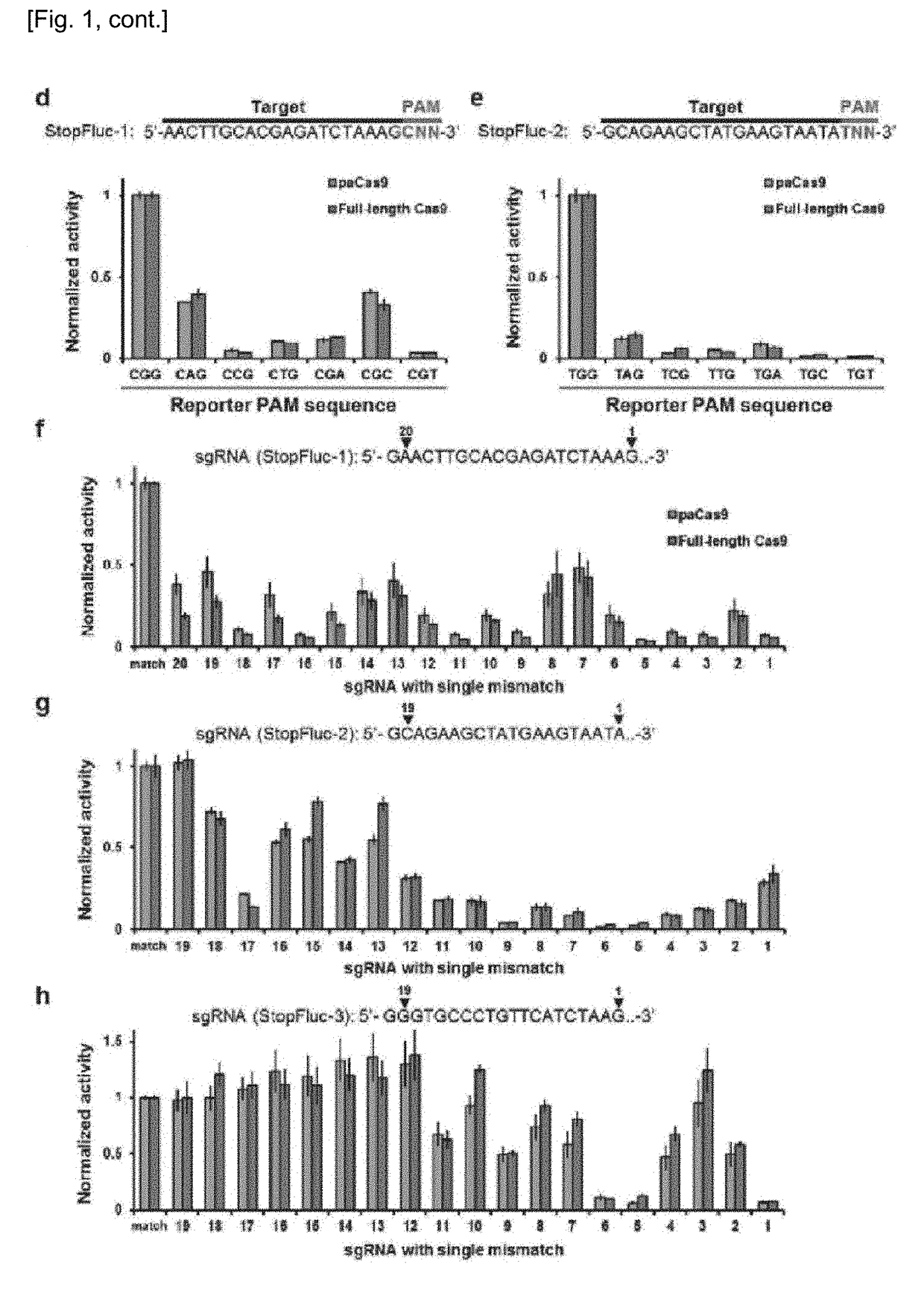
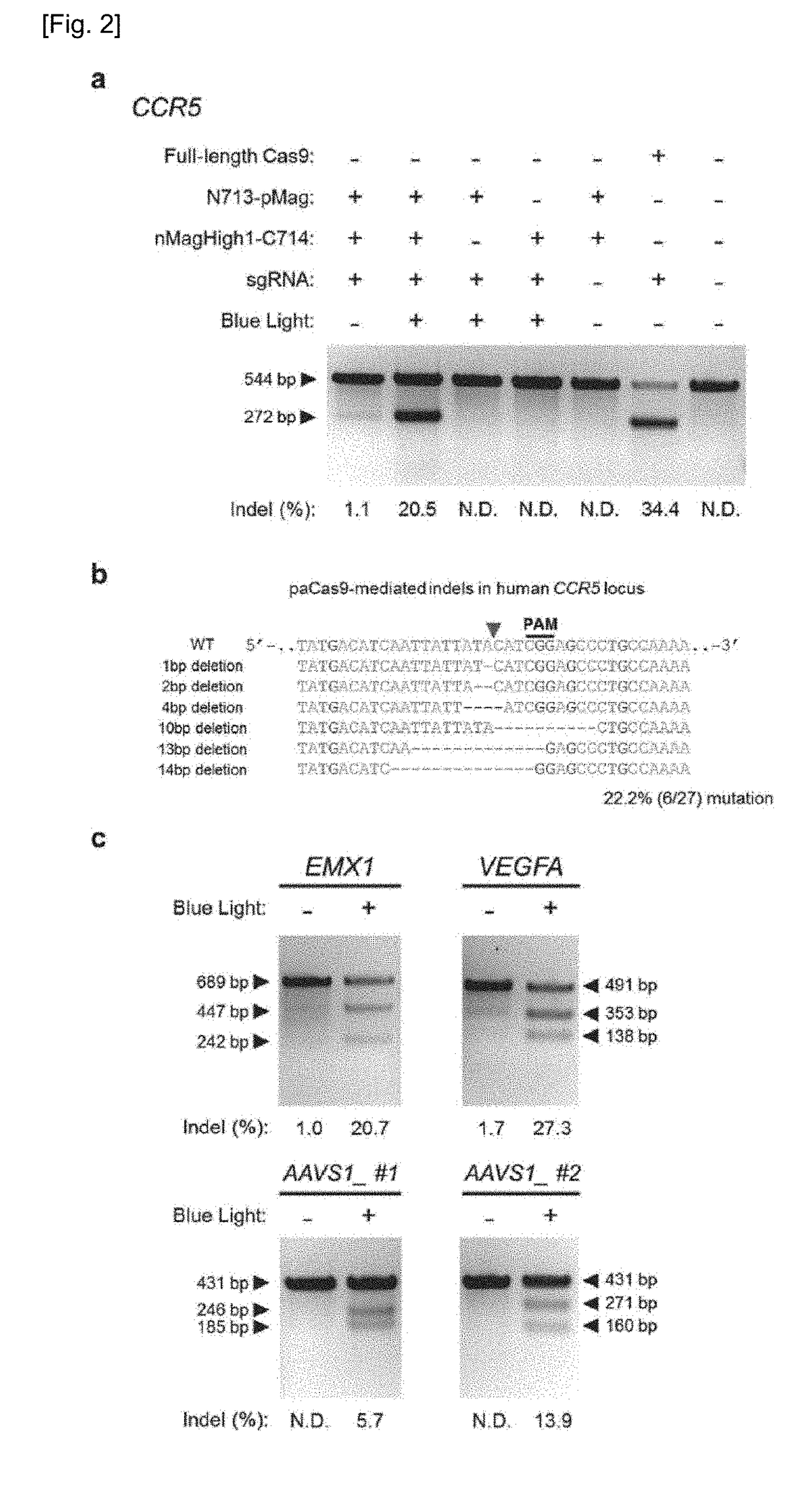
Set of Polypeptides Exhibiting Nuclease Activity or Nickase Activity with Dependence on Light or in Presence of Drug or Suppressing or Activating Expression of Target Gene
The present invention provides, for example, a set of two polypeptides exhibiting the nuclease activity with dependence on light or in the presence of a drug, in which an N-terminal side fragment and a C-terminal side fragment of a Cas9 protein are bound to each of two polypeptides which form a dimer with dependence on light or in the presence of a drug.
Owner:THE UNIV OF TOKYO
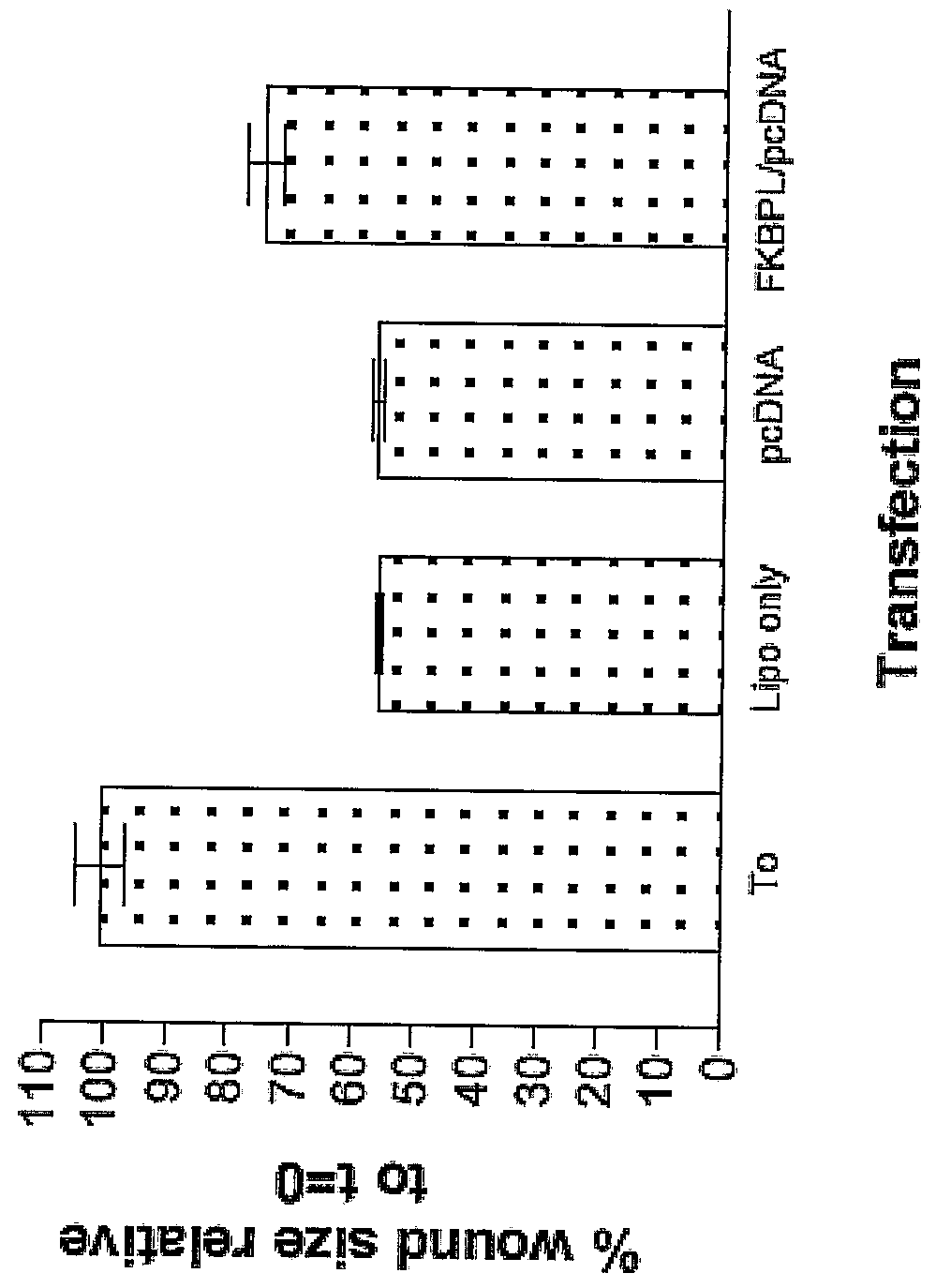
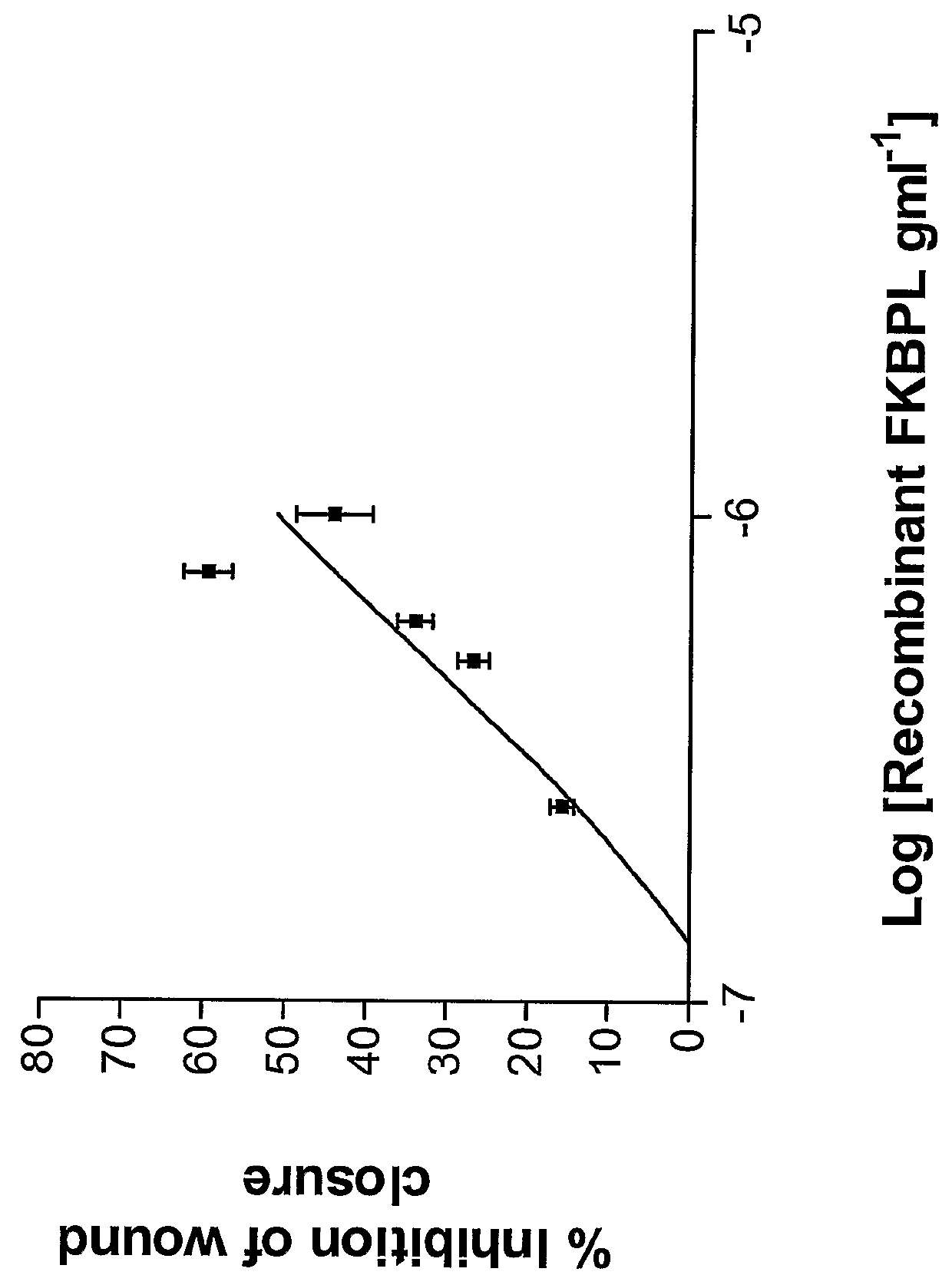
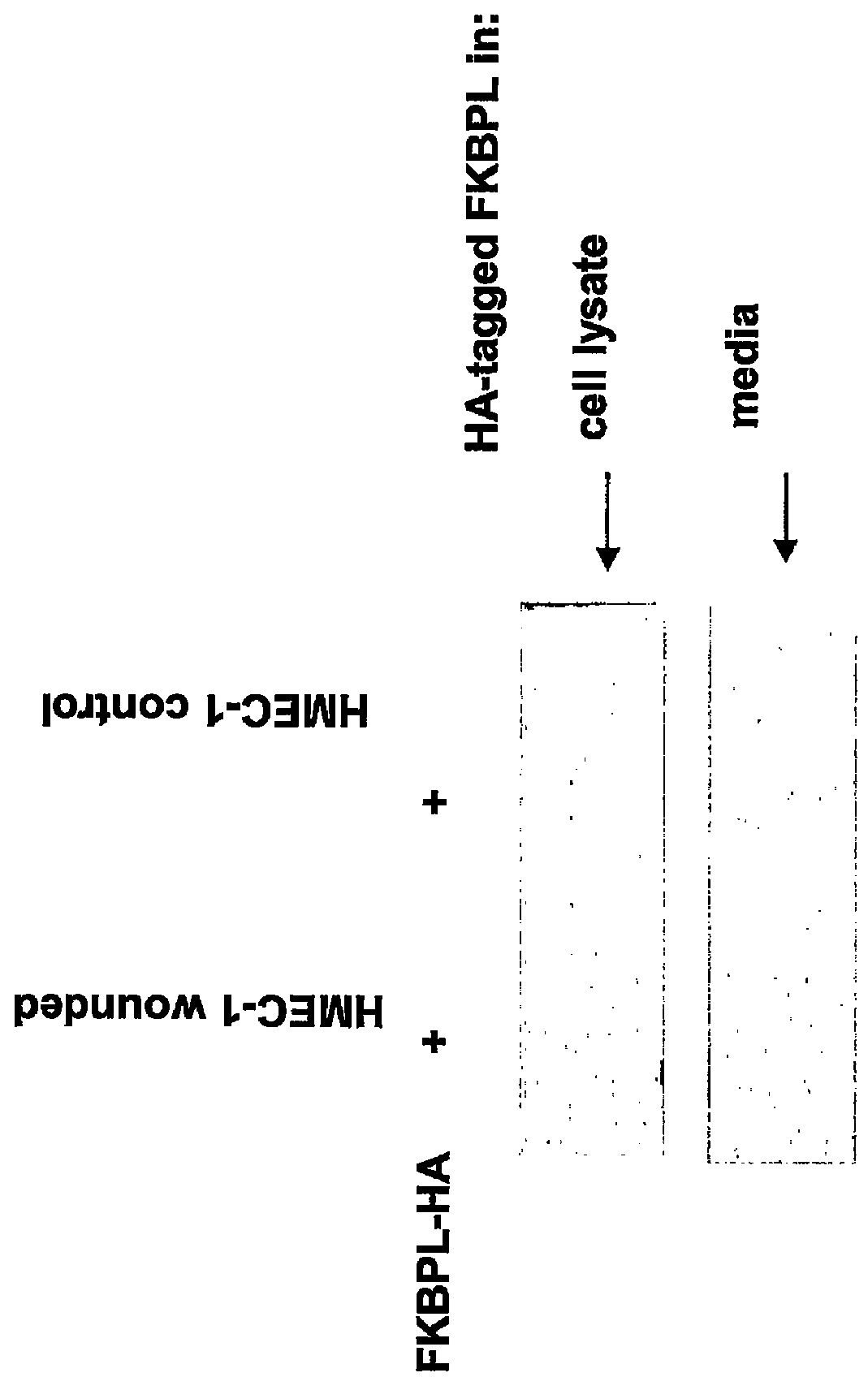
FKBP-L And Uses Thereof
Disclosed are methods and compositions that employ FKBP-L polypeptides for modulating angiogenesis and / or tumor metastasis. The FKBP-L polypeptides may be used for the treatment of disorders mediated by angiogenesis such as cancer.
Owner:ALMAC DISCOVERY LIMITED
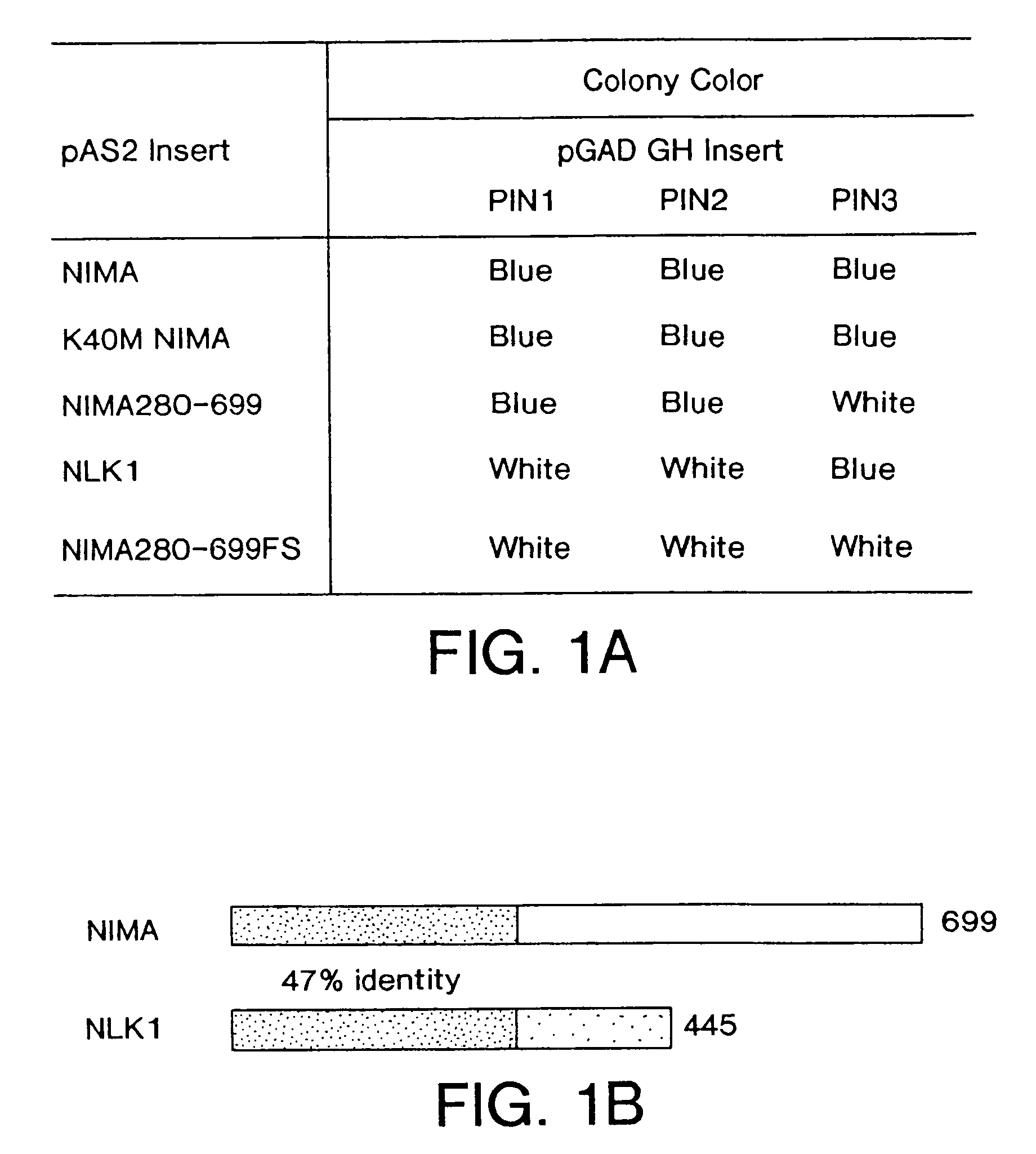
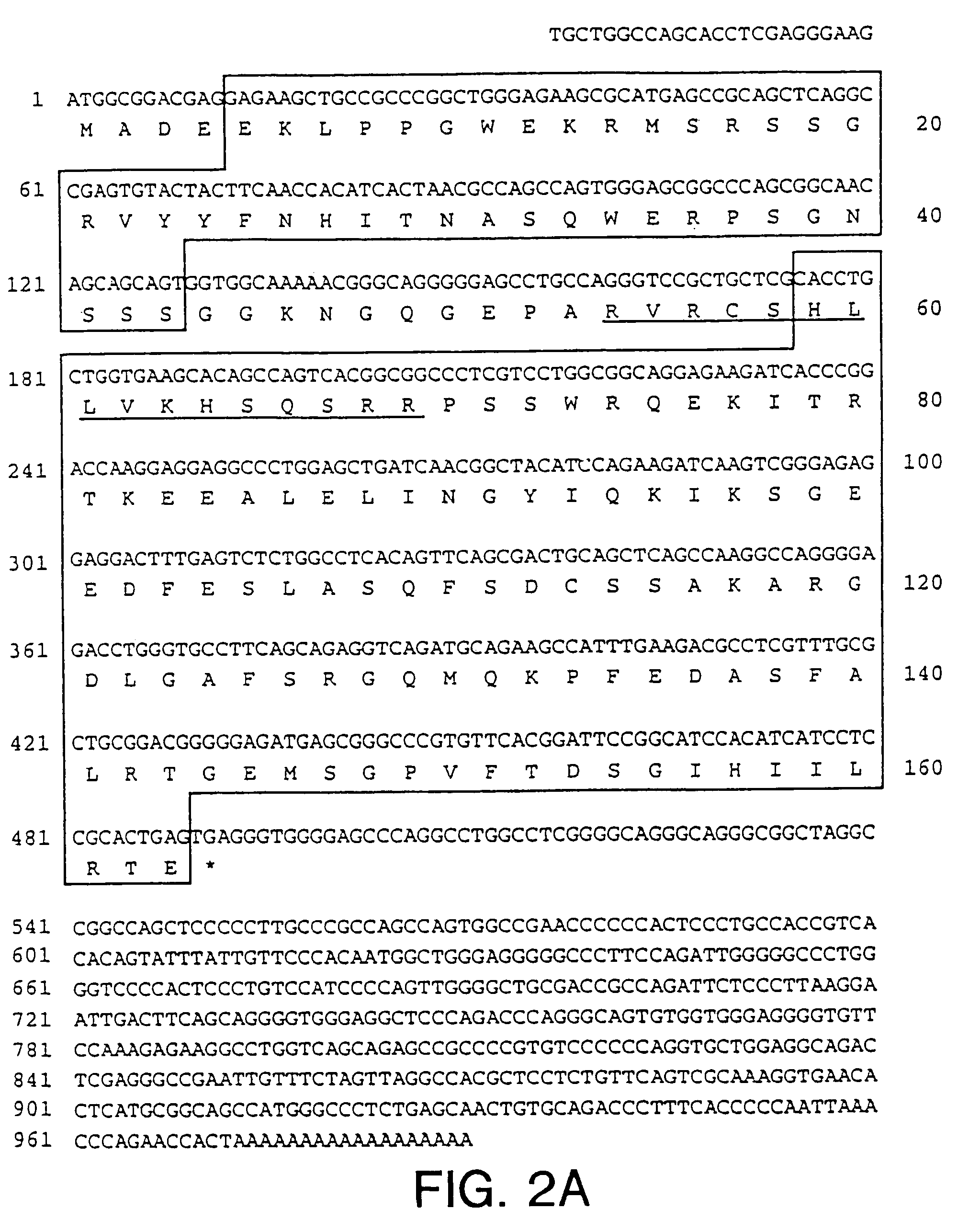
NIMA interacting proteins
InactiveUS7125677B2Inhibits mitosis promoting functionHigh activityPeptide/protein ingredientsMicrobiological testing/measurementG2 arrestDNA fragmentation
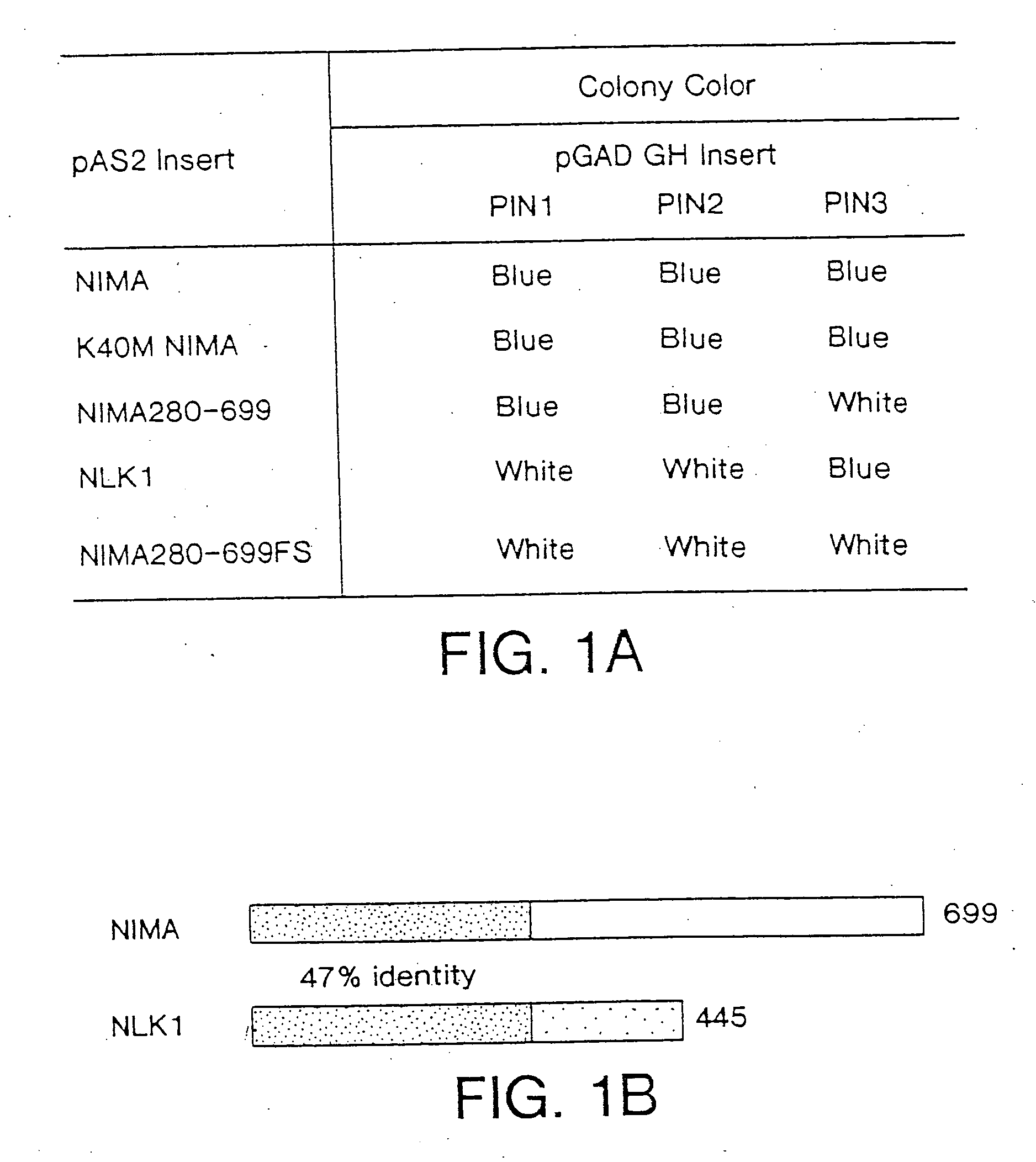
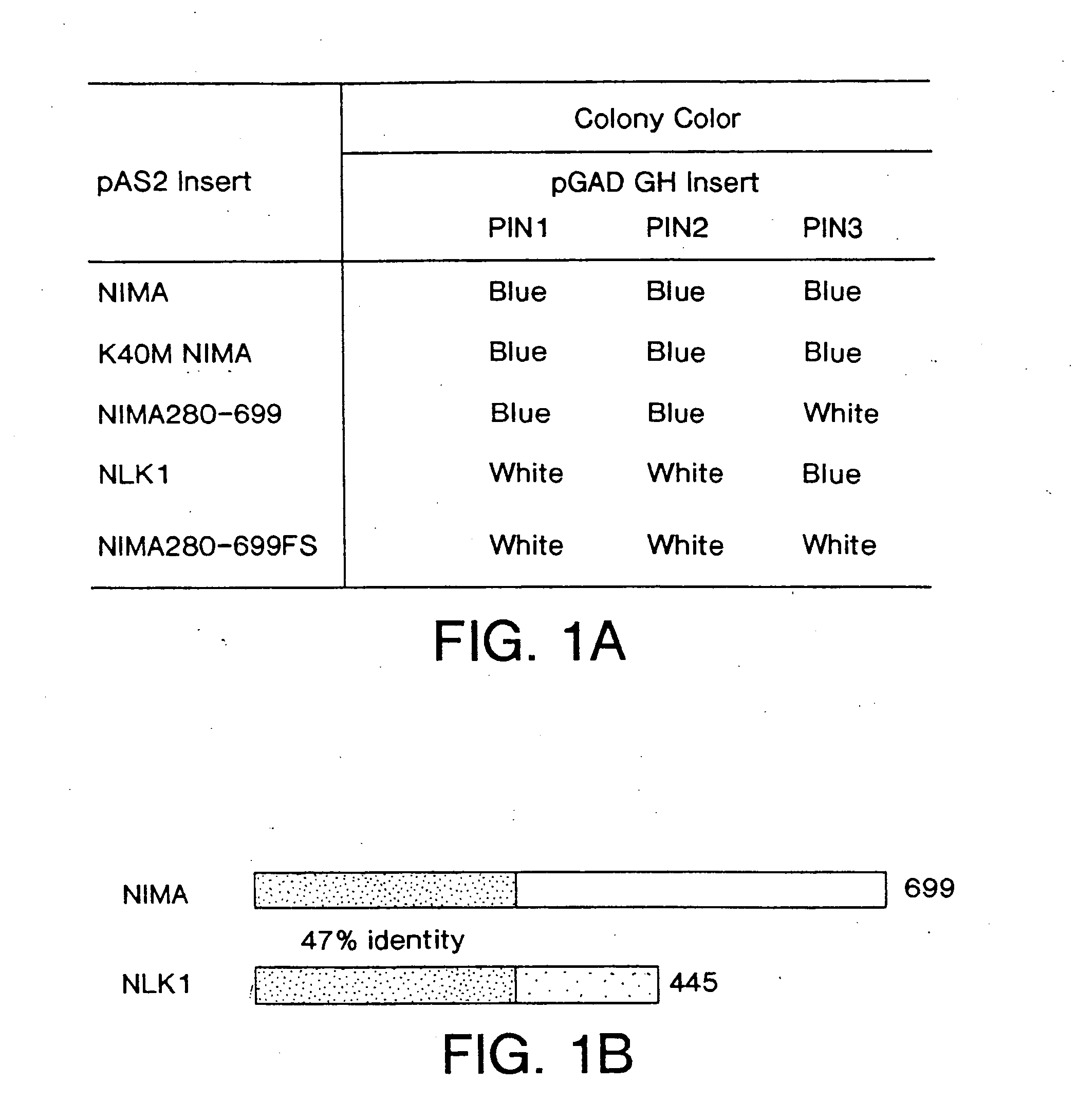
A novel class of NIMA interacting proteins (PIN), exemplified by Pin1, is provided. Pin1 induces a G2 arrest and delays NIMA-induced mitosis when overexpressed, and triggers mitotic arrest and DNA fragmentation when depleted. Methods of identifying other Pin proteins and Pin-interacting proteins and identifying compositions which affect Pin activity or expression are also provided.
Owner:SALK INST FOR BIOLOGICAL STUDIES
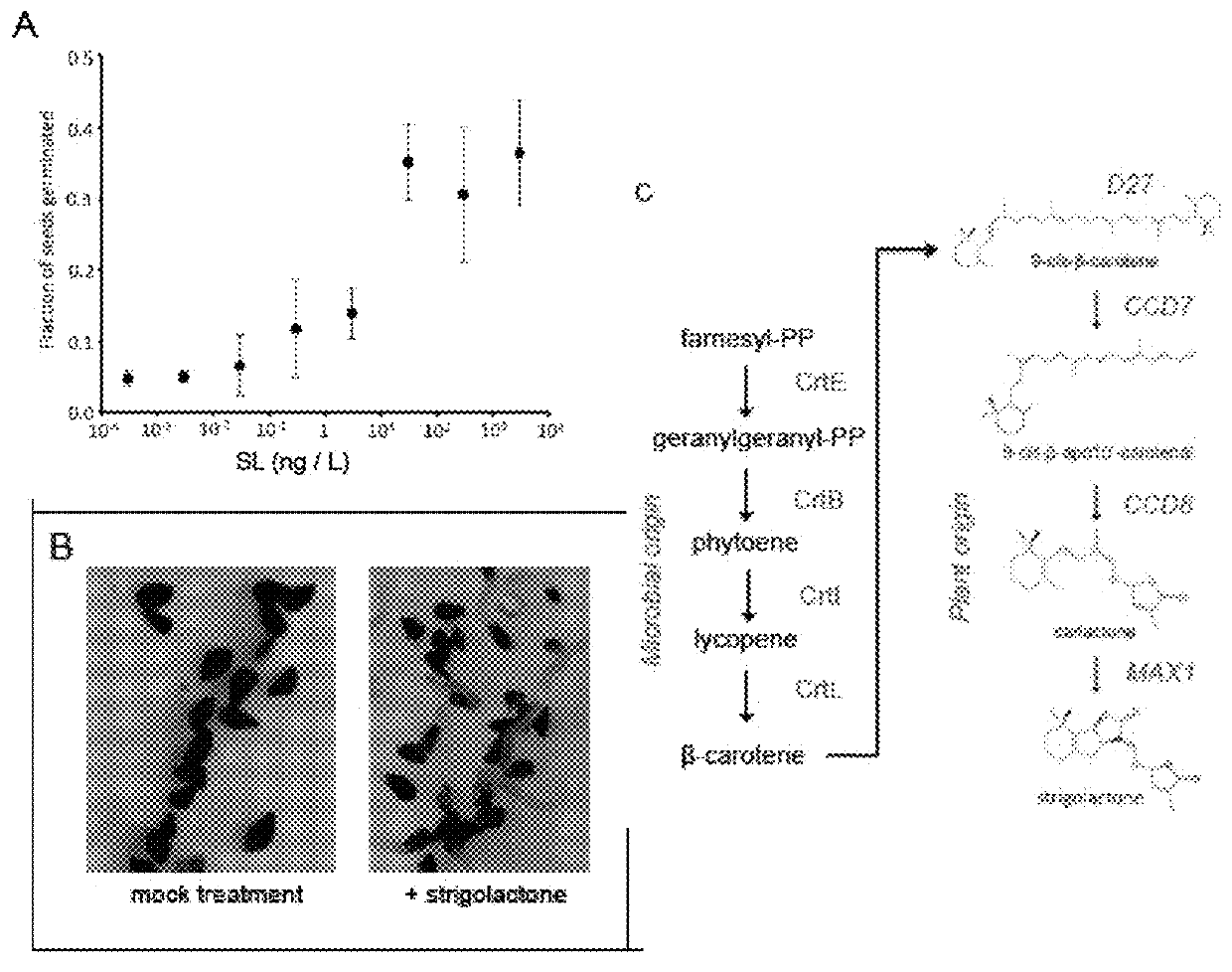
Strigolactone formulations and uses thereof
Disclosed herein plant propagation materials, methods of manufacturing, formulations and uses thereof. The plant propagation materials disclosed herein may comprise a strigolactone obtained by a biosynthetic process. The plant propagation material may comprise a chemical mimic of a strigolactone. The strigolactone may be 5-deoxystrigol. Methods of manufacturing the plant propagation materials may comprise a chemical process. Alternatively, methods of manufacturing the plant propagation material may comprise a biosynthetic process. The methods may comprise use of one or more polynucleotides. The polynucleotides may encode a metabolite. The polynucleotides may comprise one or more genes encoding one or more components of a strigolactone pathway.
Owner:SOUND AGRI CO
Regulating chimeric antigen receptors
PendingCN111386263APeptide/protein ingredientsAntibody mimetics/scaffoldsTumor lysis syndromeProtein target
This invention is in the area of compositions and methods for regulating chimeric antigen receptor immune effector cell, for example T-cell (CAR-T), therapy to modulate associated adverse inflammatoryresponses, for example, cytokine release syndrome and tumor lysis syndrome, using targeted protein degradation.
Owner:DANA FARBER CANCER INST INC
FKBP-L and uses thereof
Disclosed are methods and compositions that employ FKBP-L polypeptides for modulating angiogenesis and / or tumor metastasis. The FKBP-L polypeptides may be used for the treatment of disorders mediated by angiogenesis such as cancer.
Owner:ALMAC DISCOVERY LIMITED
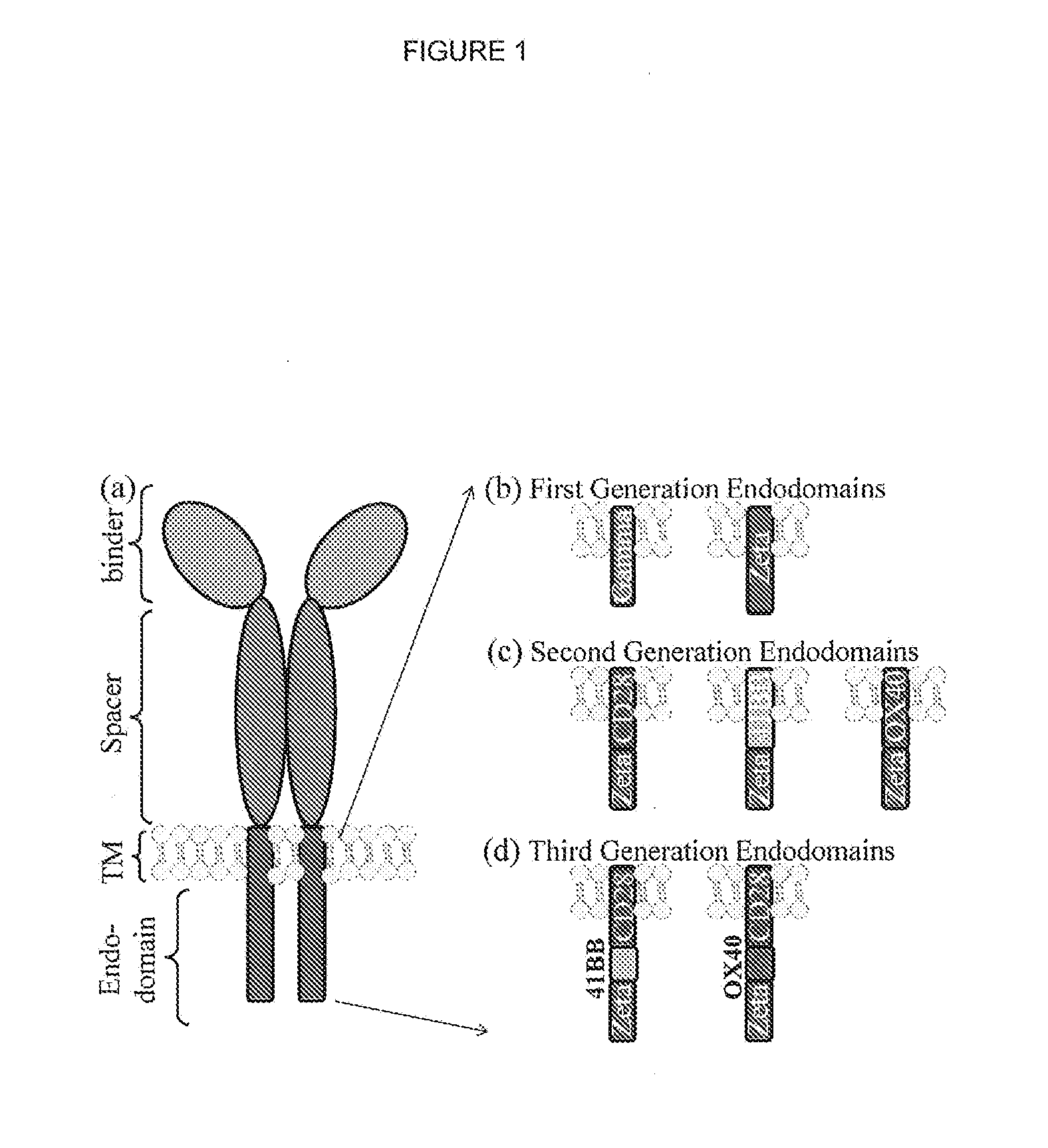
Chimeric antigen receptor (CAR) signalling system
ActiveUS20170014508A1Possible to separateAvoid toxicityOrganic active ingredientsPeptide/protein ingredientsHeterologousIntracellular signalling
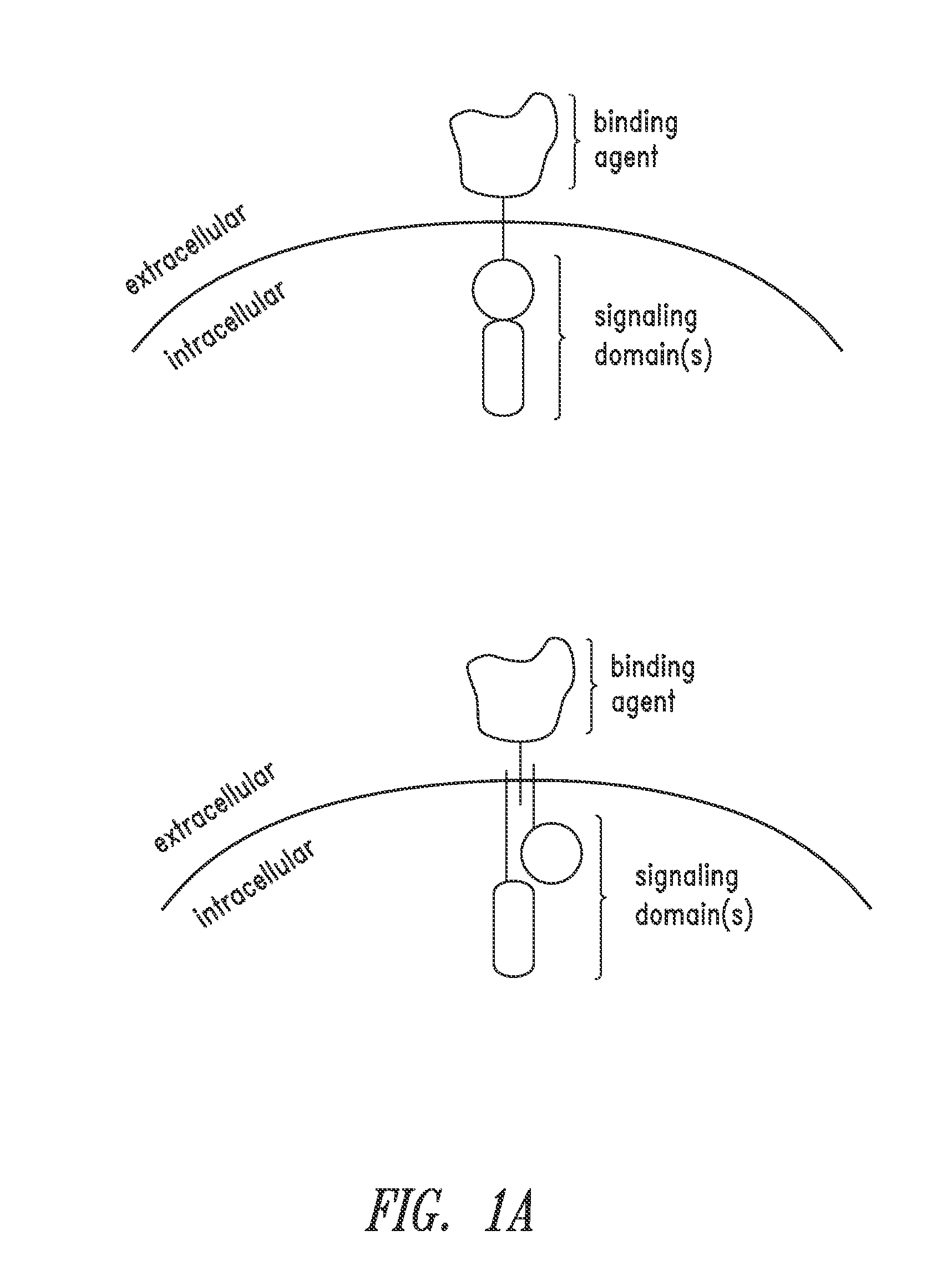
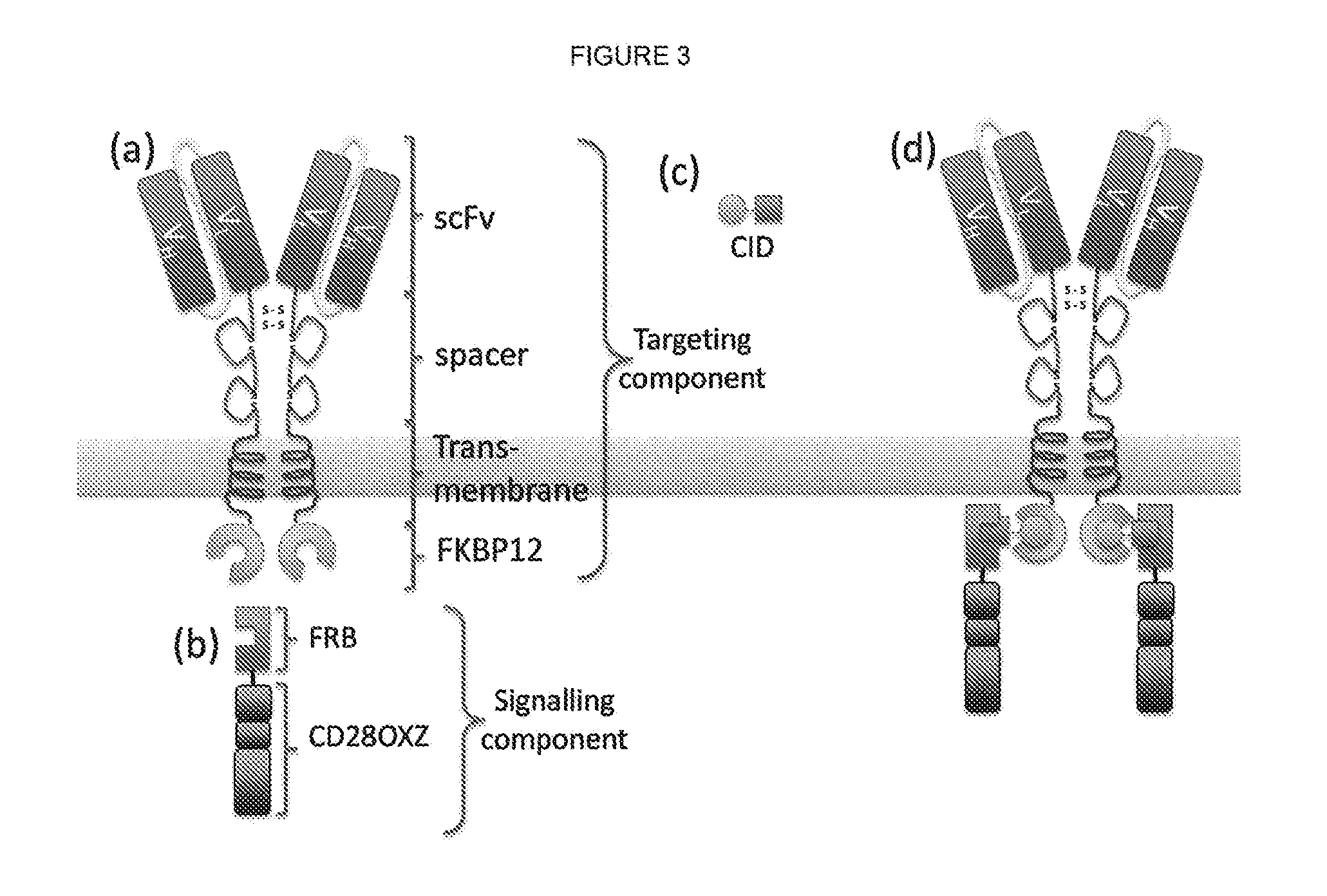
The present invention relates to a chimeric antigen receptor (CAR) signalling system comprising; (i) a receptor component comprising an extracellular antigen-binding domain, a transmembrane domain and a intracellular first chemical inducer of dimerization binding domain 1 (CBD1); and (ii) an intracellular signalling component comprising a signalling domain and a second chemical inducer of dimerization binding domain 2 (CBD2); wherein CBD1 and CBD2 are capable of simultaneously binding to a chemical inducer of dimerization (CID); wherein, in the absence of the CID, binding of the antigen-binding component to antigen does not result in signalling through the signalling component; whilst, in the presence of the CID, the receptor component and the signalling component heterodimerize and binding of the antigen-binding domain to antigen results in signalling through the signalling domain.
Owner:AUTOLUS LIMIED
Tunable endogenous protein degradation with heterobifunctional compounds
Owner:DANA FARBER CANCER INST INC
Apoptosis methods, genes and proteins
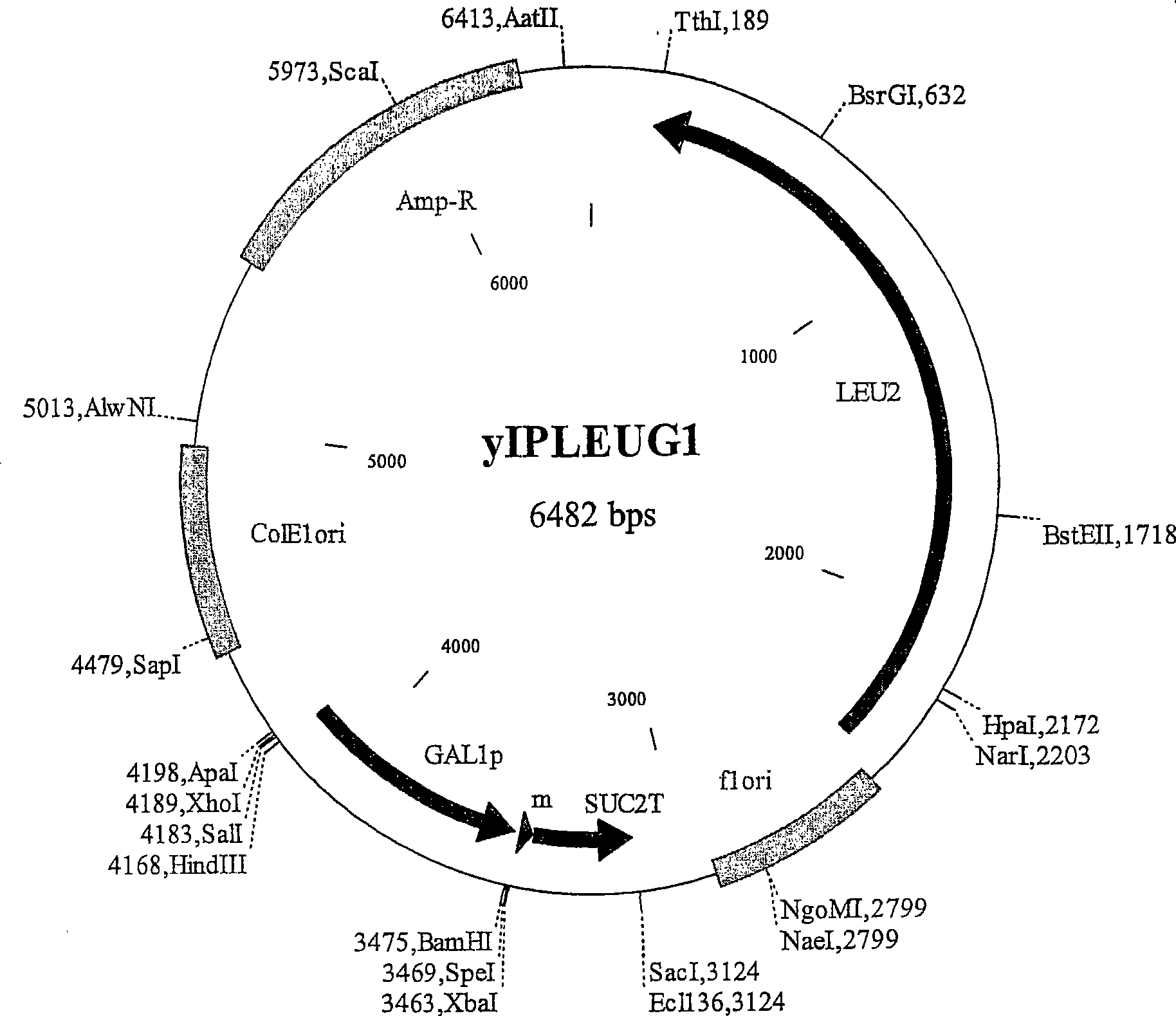
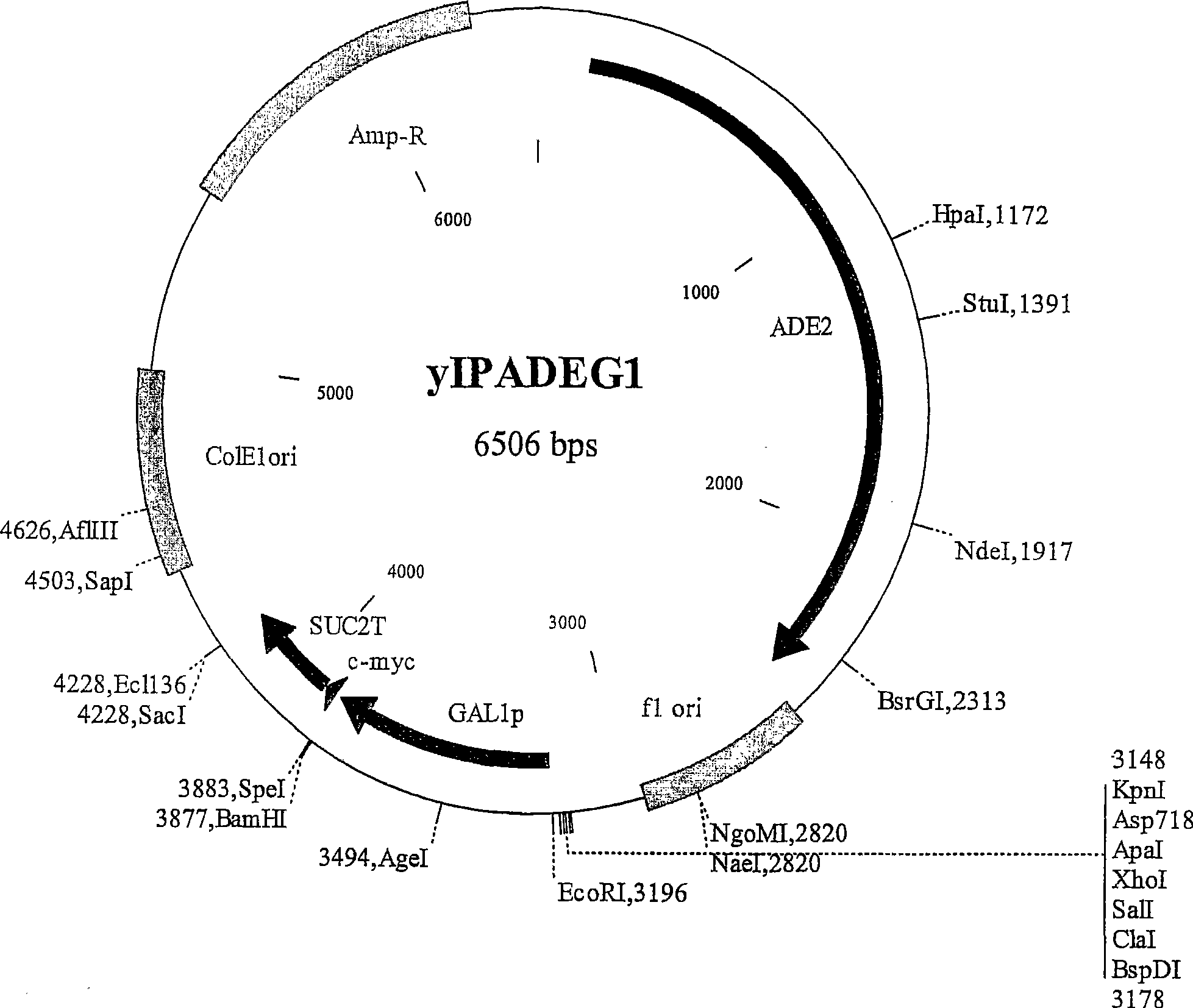
A W303a Saccharomyces cerevisiae yeast cell which contains a polynucleotide that encodes a functional Bax polypeptide under the control of a galactose- inducible promoter that is integrated at the LEU2 chromosomal locus. A kit of parts comprising the yeast cells and a yeast plasmid vector suitable for transforming a cDNA library into the yeast cells. Use of the yeast cell for screening a cDNA library for a polynucleotide that is or encodes an inhibitor of B ax-mediated apoptosis. Genes and polypeptides that inhibit Bax-mediated apoptosis and which were identified from a human hippocampus cDNA library screened in the yeast cells. A method of combating Bax-mediated apoptosis in a cell using an inhibitor of Bax-mediated apoptosis which was identified from a human hippocampus cDNA library screened in the yeast cells. A method of promoting Bax-mediated apoptosis in a cell using an inhibitor or antagonist of the anti-apoptotic polypeptides identified from a human hippocampus cDNA library screened in the yeast cells.
Owner:MORVUS TECH LTD
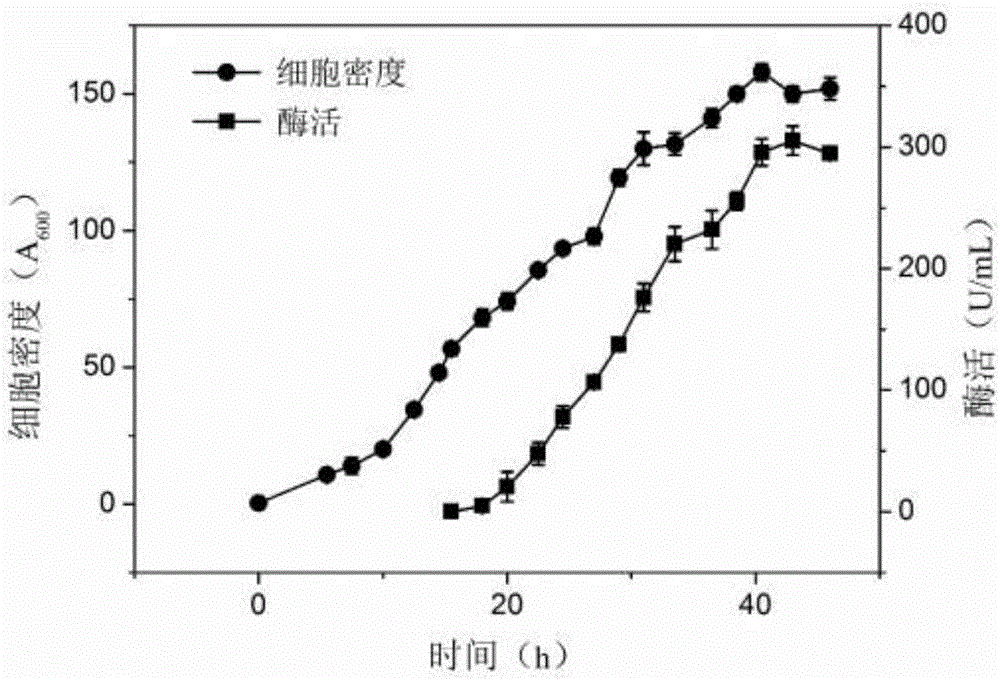
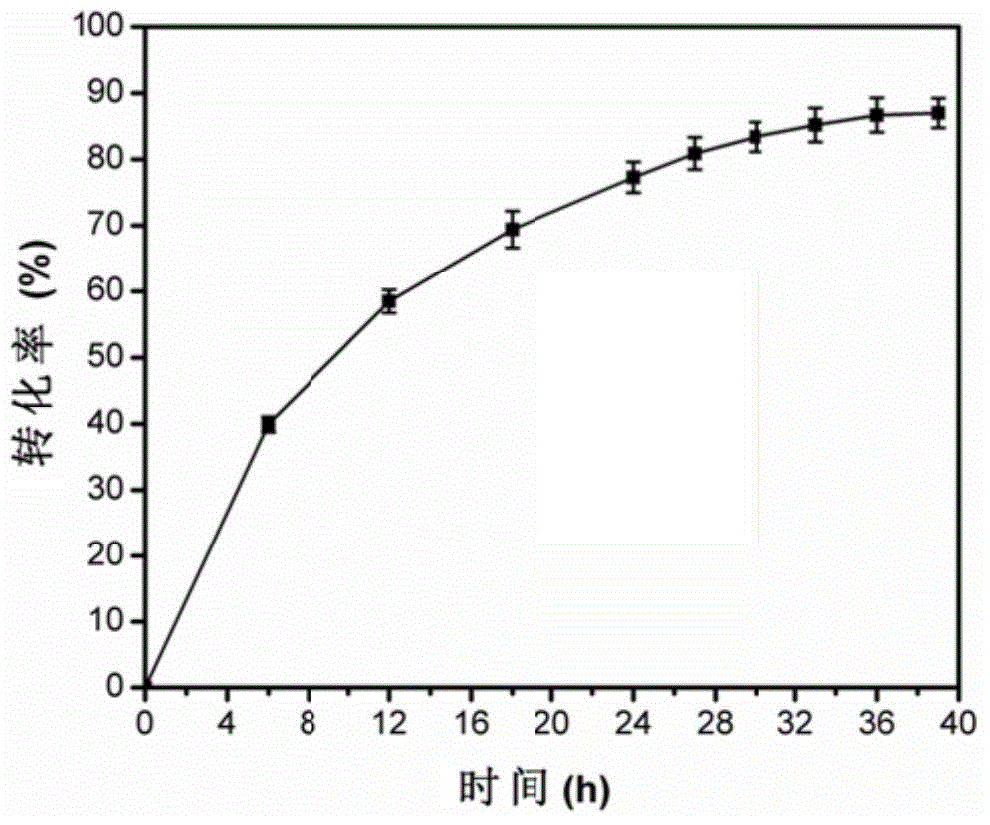
Engineered Escherichia coli and method of synthesis of catalyzing fumaric acid from maleic acid in presence of Engineered Escherichia coli
InactiveCN106222122AImprove expression efficiencyHigh densityBacteriaCis-trans-isomerasesFumaraseEscherichia coli
The invention relates to engineered Escherichia coli capable of high-yield production of fumaric acid; specifically, fumarase coded genes fumA and fumC of Escherichia coli are knocked off, and maleic cis-trans isomerase gene from Serratia marcescens is converted to obtain the engineered Escherichia coli. The invention also discloses a method of synthesis of catalyzing fumaric acid from maleic acid by using the engineered Escherichia coli. The method includes culturing the engineered Escherichia coli by fermenting, subjecting the engineered Escherichia coli to high-density inductive expression when OD600 reaches 40-80, and applying the obtained fermentation broth to biochemical catalysis of maleic acid in presence of Engineered Escherichia coli to obtain fumaric acid. The genes fumA and fumC in the Escherichia coli are knocked off by using Red homologous recombination, a metabolic pathway of its fumaric acid to L-malic acid is cut off, the maleic cis-trans isomerase from Serratia marcescens is then expressed, and the obtained genetically-engineered Escherichia coli can efficiently convert maleic matrix to synthesize high-purity fumaric acid, there is nearly no L-malic acid byproduct synthesized, and basis is provided for the industrialized production of fumaric acid by whole-cell process catalysis of maleic acid.
Owner:JIANGNAN UNIV
SiRNA targeting catenin, beta-1 (CTNNB1)
ActiveUS8198427B1Improve efficiencyGood curative effectCell receptors/surface-antigens/surface-determinantsSugar derivativesNucleotideNucleotide sequencing
Efficient sequence specific gene silencing is possible through the use of siRNA technology. By selecting particular siRNAs by rational design, one can maximize the generation of an effective gene silencing reagent, as well as methods for silencing genes. Methods, compositions, and kits generated through rational design of siRNAs are disclosed including those directed to nucleotide sequences for CTNNB1.
Owner:THERMO FISHER SCIENTIFIC INC
Methods and Compositions for Selecting siRNA of Improved Functionality
ActiveUS20180201934A1Improve efficiencyGood curative effectOrganic active ingredientsCis-trans-isomerasesNucleotideGene silencing
Efficient sequence specific gene silencing is possible through the use of siRNA technology. Be selecting particular siRNAs by rational design, one can maximize the generation of an effective gene silencing reagent, as well as methods for silencing genes. Methods compositions, and kits generated through rational design of siRNAs are disclosed, including those directed to the nucleotide sequences for HAO1.
Owner:THERMO FISHER SCIENTIFIC INC
Compositions and methods for immunotherapy
PendingCN110913954APrevent or reverse exhaustionReduce tumor burdenVirusesHydrolasesCancer researchEffector
The present invention provides biocircuit systems, effector modules and compositions for cancer immunotherapy. Methods for inducing anti-cancer immune responses in a subject are also provided.
Owner:OBSIDIAN THERAPEUTICS INC
NIMA interacting proteins
Owner:SALK INST FOR BIOLOGICAL STUDIES
NIMA interacting proteins
InactiveUS20050027107A1Peptide/protein ingredientsMicrobiological testing/measurementG2 arrestDNA fragmentation
A novel class of NIMA interacting proteins (PIN), exemplified by Pin1, is provided. Pin1 induces a G2 arrest and delays NIMA-induced mitosis when overexpressed, and triggers mitotic arrest and DNA fragmentation when depleted. Methods of identifying other Pin proteins and Pin-interacting proteins and identifying compositions which affect Pin activity or expression are also provided.
Owner:SALK INST FOR BIOLOGICAL STUDIES
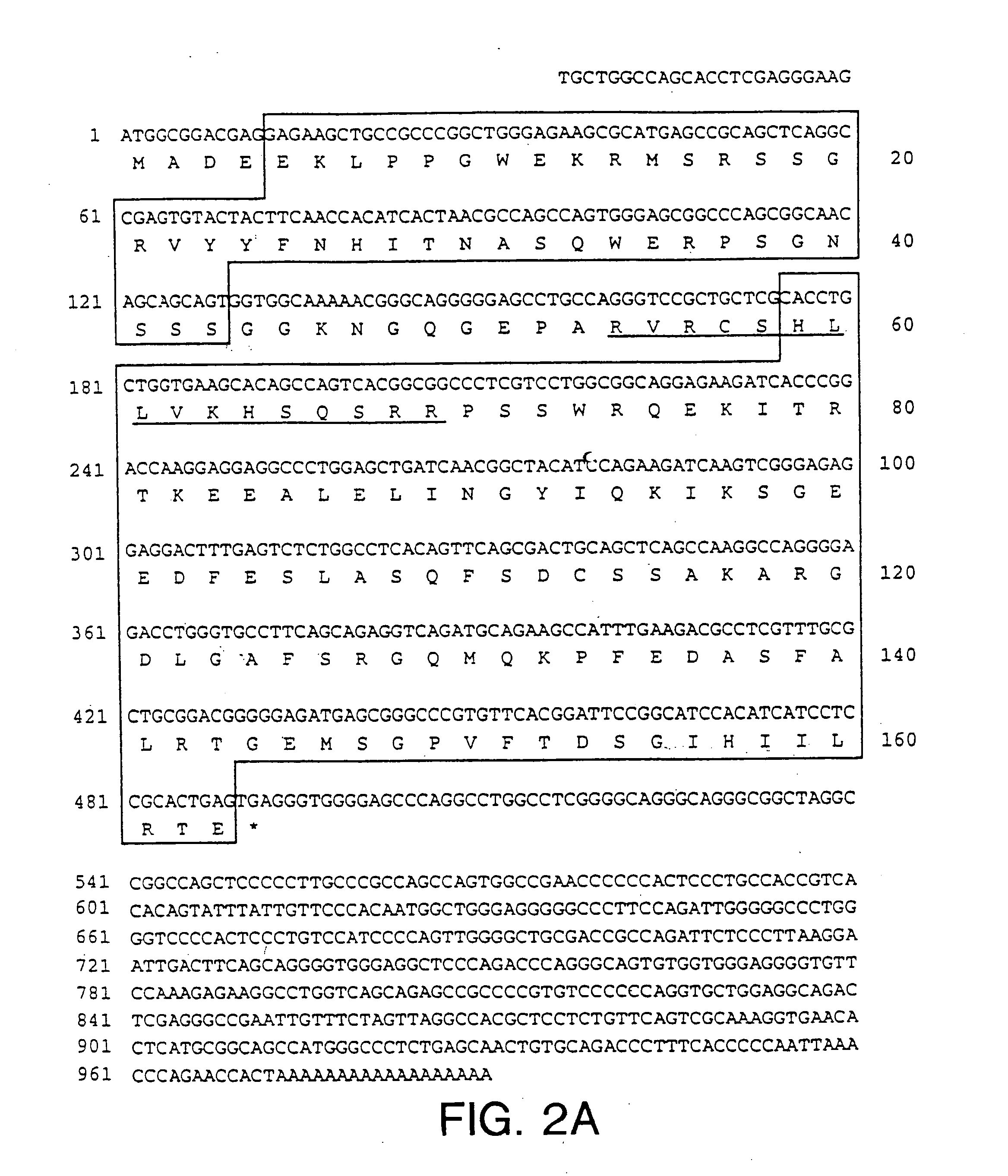
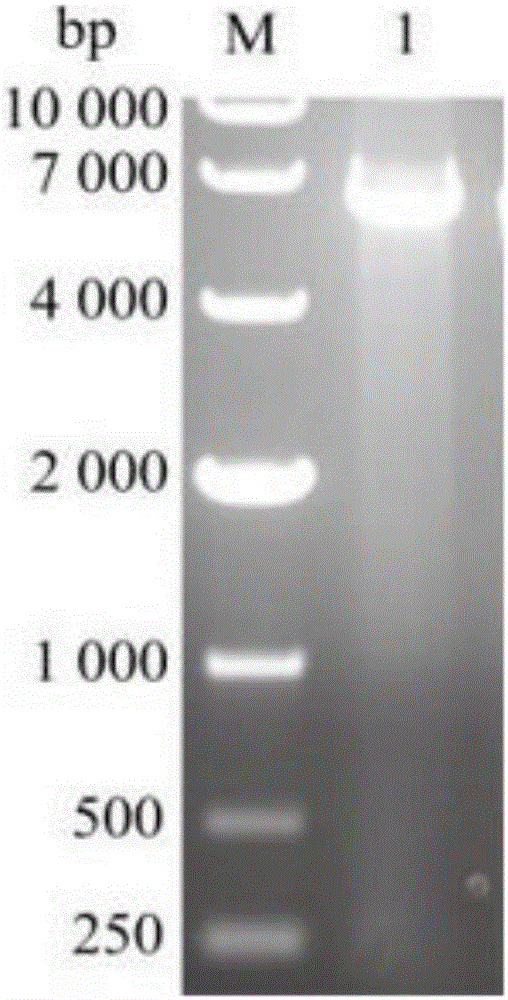
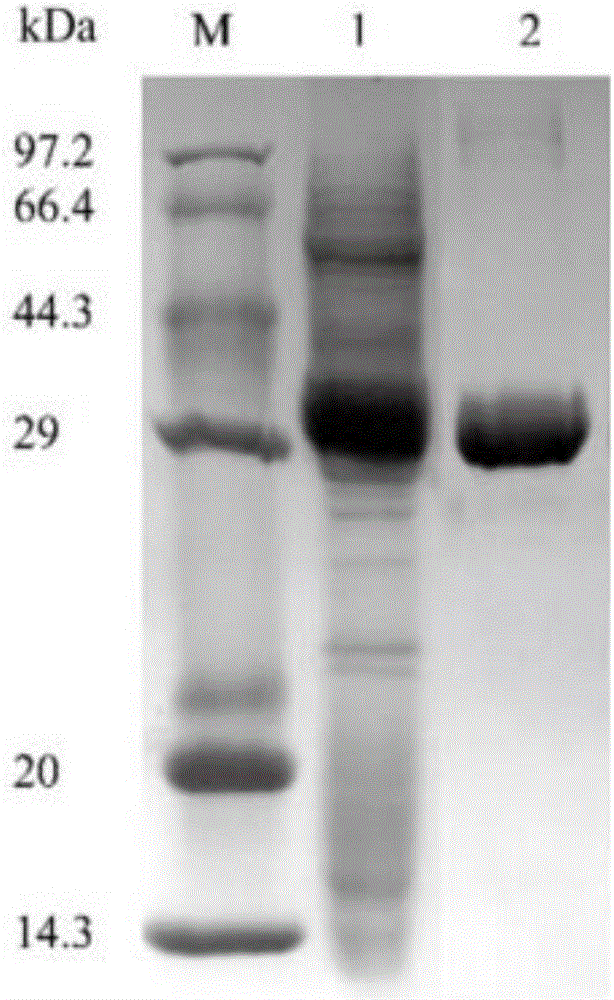
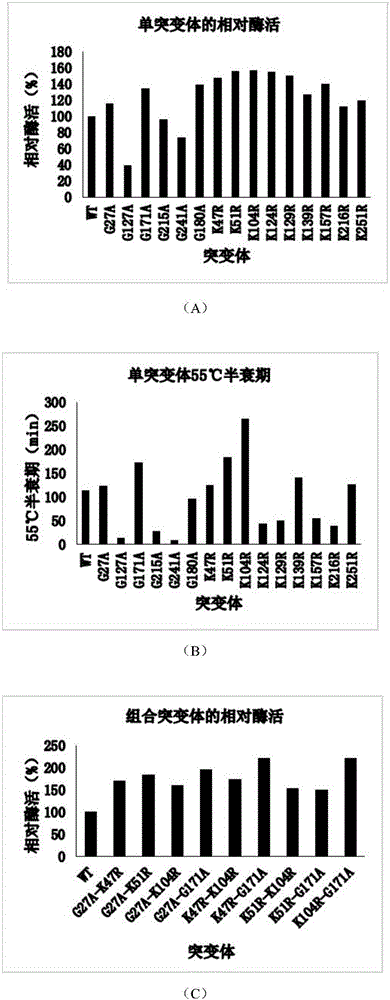
Thermostability transformation of maleic acid cis-trans isomerase and application thereof
ActiveCN106636052AImprove thermal stabilityIncrease enzyme activityBacteriaMicroorganism based processesHalf-lifeWild type
The invention discloses thermostability transformation of maleic acid cis-trans isomerase and application thereof, and belongs to the field of enzyme engineering. The maleic acid cis-trans isomerase gene maiA from serratia marcescens is subjected to thermostability transformation by a semi-rational design method. Compared with the wild maleic acid cis-trans isomerase, the obtained mutant has the advantages that the half-life period of the mutant G27A-G171A at 55 DEG C is 2.9 times of that of the wild type, and the enzyme activity is 1.96 times of that of the wild type; the half-life period of the mutant G27A-K104R is 4.18 times of the wild type at 55 DEG C, and the enzyme activity is 1.59 times of that of the wild type.
Owner:JIANGNAN UNIV
Methods and Compositions for Selecting siRNA of Improved Functionality
ActiveUS20140148362A1Improve efficiencyGood curative effectOrganic active ingredientsSugar derivativesNucleotideSilencing gene
Efficient sequence specific gene silencing is possible through the use of siRNA technology. By selecting particular siRNAs by rational design, one can maximize the generation of an effective gene silencing reagent, as well as methods for silencing genes. Methods, compositions, and kits generated through rational design of siRNAs are disclosed including those directed to nucleotide sequences for TTR.
Owner:THERMO FISHER SCIENTIFIC INC
Methods and compositions for selecting siRNA of improved functionality
ActiveUS9879266B2Improve efficiencyGood curative effectOrganic active ingredientsSugar derivativesNucleotide sequencingSilencing gene
Efficient sequence specific gene silencing is possible through the use of siRNA technology. Be selecting particular siRNAs by rational design, one can maximize the generation of an effective gene silencing reagent, as well as methods for silencing genes. Methods compositions, and kits generated through rational design of siRNAs are disclosed, including those directed to the nucleotide sequences for HAO1.
Owner:THERMO FISHER SCIENTIFIC INC
Methods and compositions for selecting siRNA of improved functionality
ActiveUS9839649B2Improve efficiencyGood curative effectOrganic active ingredientsSugar derivativesSilencing geneNucleotide sequencing
Efficient sequence specific gene silencing is possible through the use of siRNA technology. Be selecting particular siRNAs by rational design, one can maximize the generation of an effective gene silencing reagent, as well as methods for silencing genes. Methods compositions, and kits generated through rational design of siRNAs are disclosed, including those directed to the nucleotide sequences for AAT.
Owner:THERMO FISHER SCIENTIFIC INC
Propionibacterium acnes linoleate isomerase and application thereof
ActiveCN104531655AOvercoming the disadvantages of producing conjugated linoleic acidLow costBacteriaCis-trans-isomerasesIsomerizationPropanoic acid
Propionibacterium acnes linoleate isomerase and its application. An amino acid sequence is as shown in SEQ ID NO.1. Through concerted catalysis of compound enzyme, conjugated linoleic acid can be directly prepared from plant oil containing high content of linoleic acid, thus overcoming the defect that conjugated linoleic acid must be produced by using expensive free linoleic acid as a substrate and saving costs greatly. Compound enzyme is prepared by the utilization of recombinant bacteria, thus raising enzyme activity and increasing enzyme output. There is no need to purchase commercial enzyme, and cost is saved effectively. The isomerization product conjugated linoleic acid contains single component and has a plurality of physiological activities. The scheme has good practicality. Good economic benefit can be produced. The propionibacterium acnes linoleate isomerase has a good industrial prospect.
Owner:NANJING FORESTRY UNIV
Maleate cis-trans isomerase mutant, and coding gene and application thereof
The invention relates to a protein mutant, particularly a maleate cis-trans isomerase, and a coding gene and application thereof. The amino acid sequence of the maleate cis-trans isomerase is disclosed as SEQ ID NO.4, or an amino acid sequence with equal functions, which is derived from SEQ ID NO.4 and subjected to substitution, deletion or addition of one or more amino acids. Compared with the wild type maleate cis-trans isomerase, the changes of the specific amino acid sequence of the mutant as follows: K51I (the 51st lysine is changed into isoleucine), R177S (the 177th arginine is changed into serine), and A212G (the 212th alanine is changed into glycine). The test verifies that the enzyme activity of the mutant is enhanced by 2.71 times as compared with the wild type maleate cis-trans isomerase.
Owner:ANHUI BBCA FERMENTATION TECH ENG RES
Popular searches
Virus peptides Immunoglobulins Fusions for enhanced expression stability/folding Fermentation Vector-based foreign material introduction Immunoglobulins against cell receptors/antigens/surface-determinants Depsipeptides Peptide preparation methods Disease diagnosis Carrier-bound/immobilised peptides