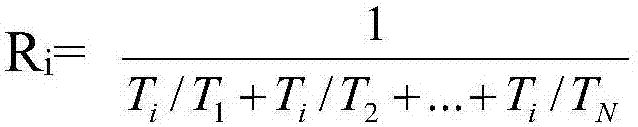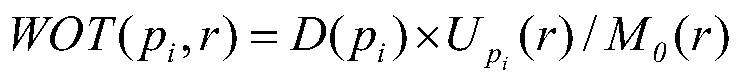Patents
Literature
47 results about "Multi core computing" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
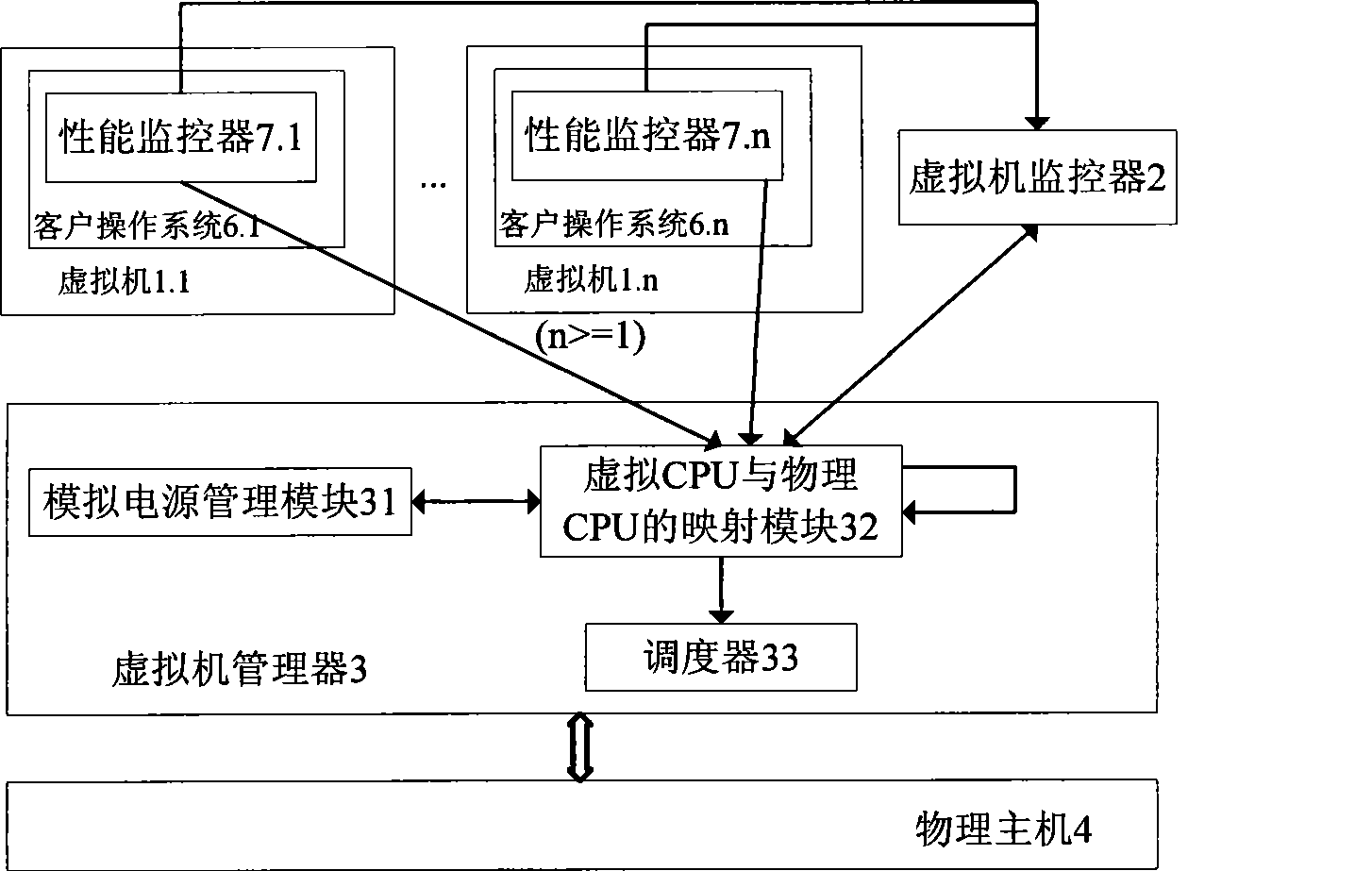
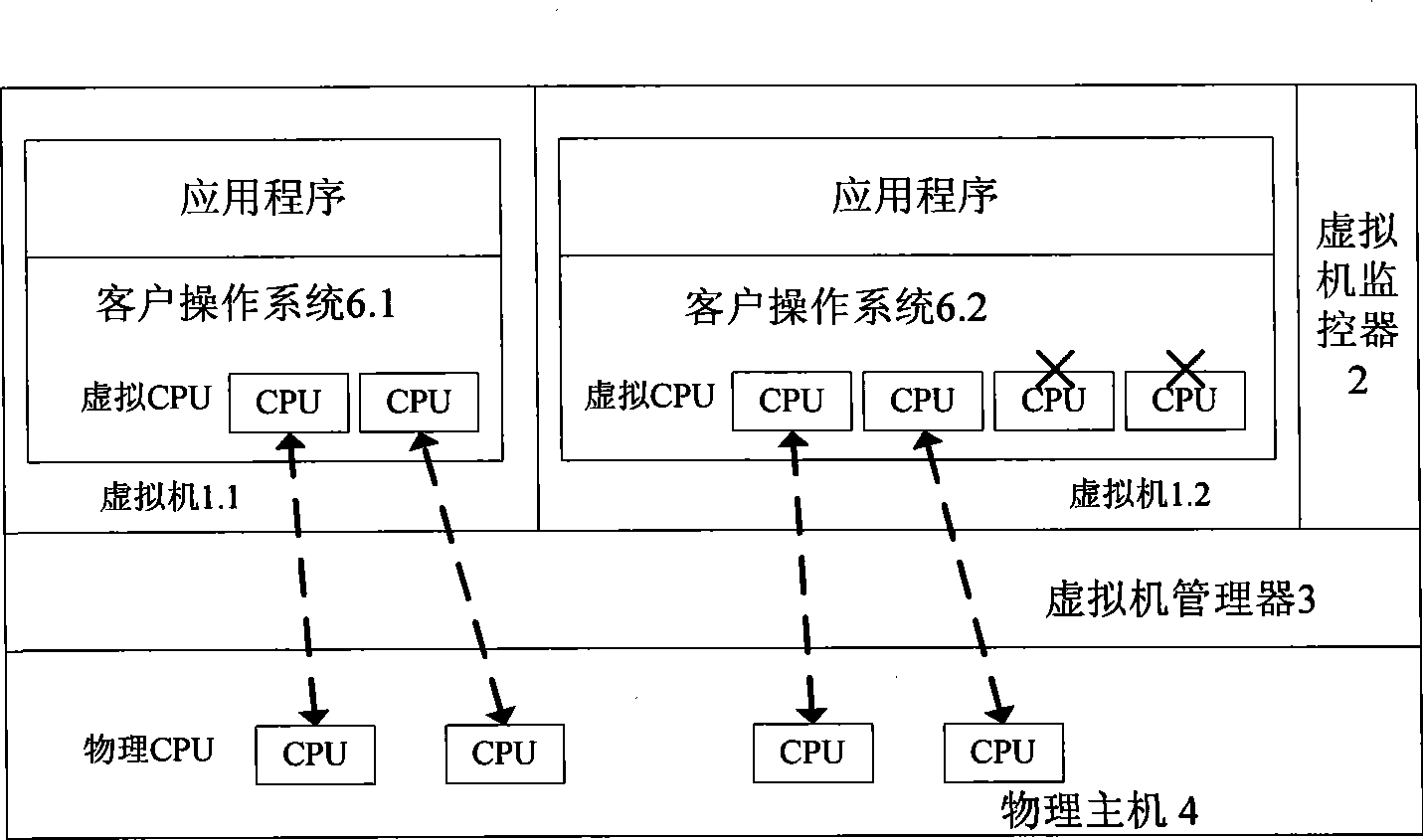
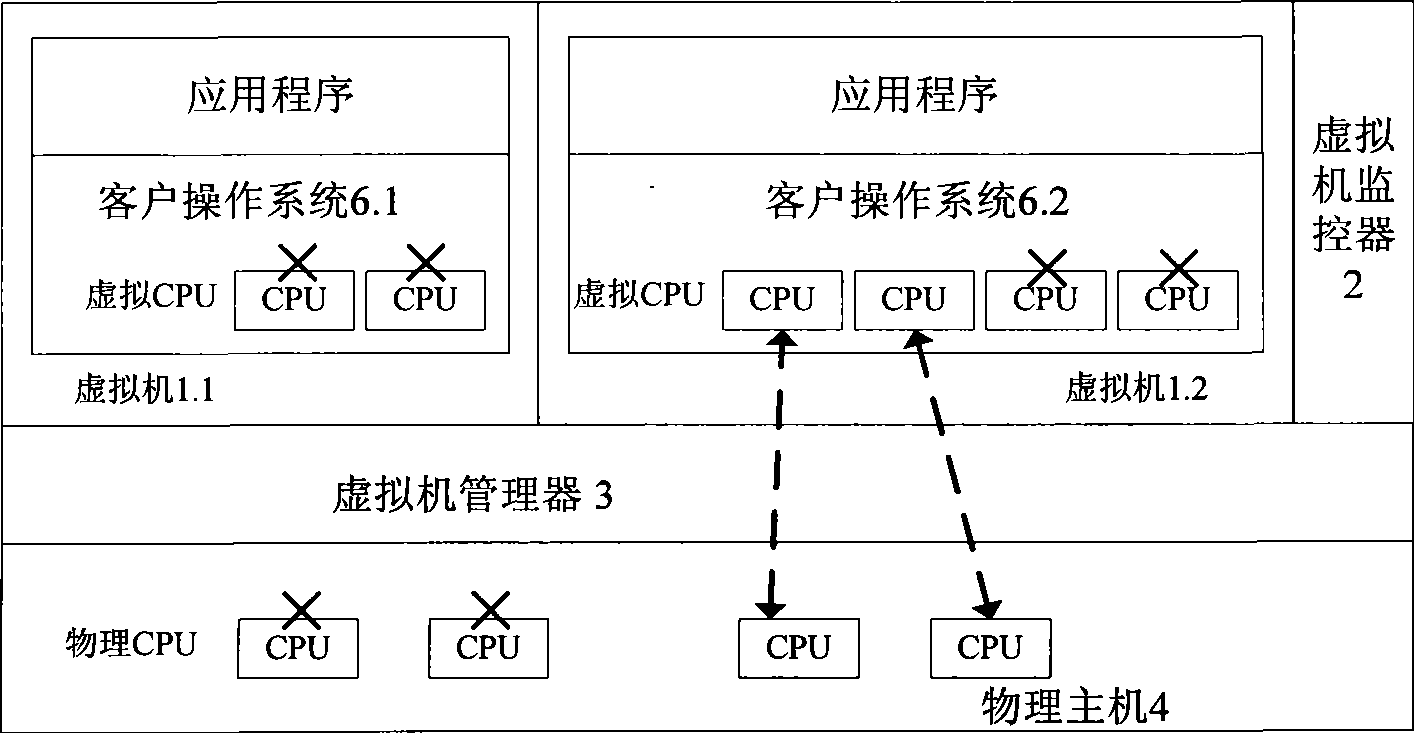
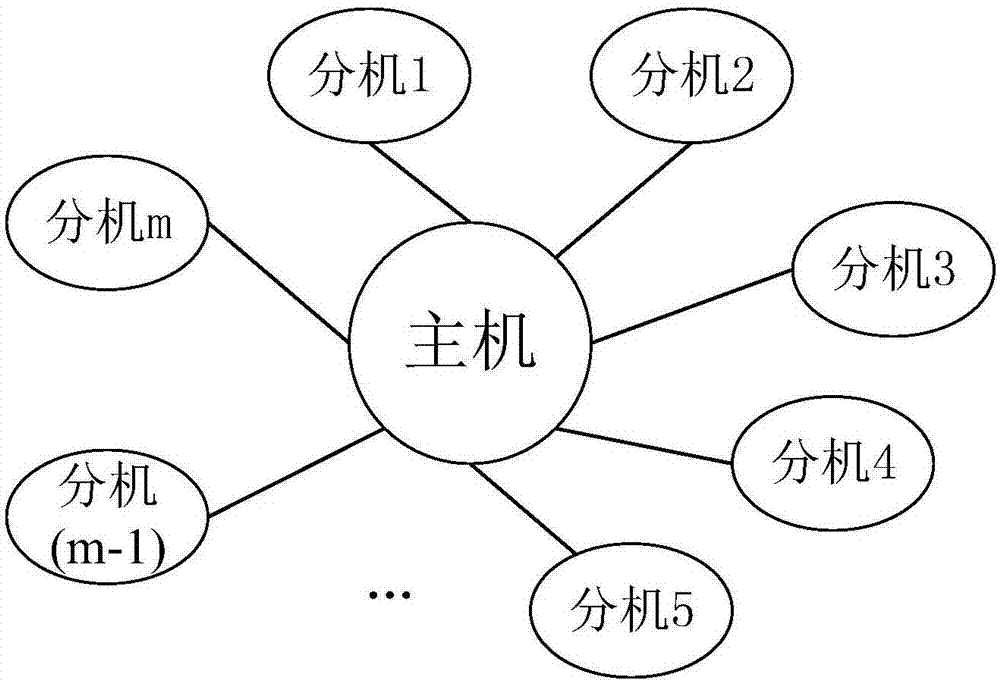
Multi-core computing resource management system based on virtual computing technology
InactiveCN101488098AReasonable distributionImprove management abilityEnergy efficient ICTResource allocationMulti core computingResource Management System
The invention relates to a multi-core system computer resource management system based on a virtual computer technology. The system comprises a plurality of virtual machines, a virtual machine monitor and a virtual machine manager. The virtual machine monitor monitors the load condition and the operation state of the virtual machine at real time. The virtual machine manager is a bond for communication between a virtual machine and a physical host. The virtual machine operates on the virtual machine manager and provides the user with a virtual platform. At the same time, the invention divides the virtual machine into three general categories, and different resource adjusting strategies are adopted for each category of the virtual machine respectively. The invention provides a practical and feasible way for the dynamic adjusting and distributing problem of multi-core computer resource and realizes the maximization of energy saving and resource utilization.
Owner:HUAZHONG UNIV OF SCI & TECH
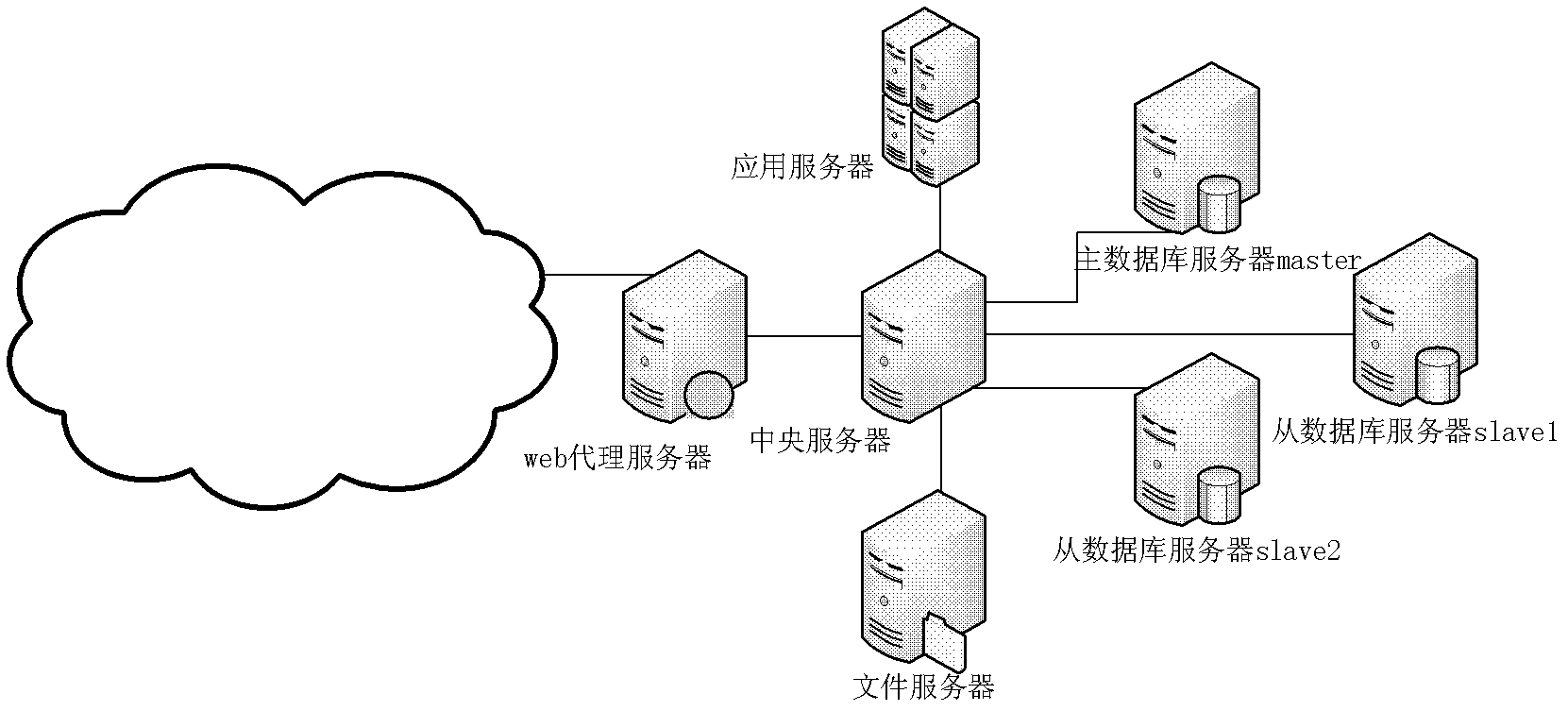
Scheduling method of cloud computing open platform
ActiveCN102681889AHigh speedImprove responsivenessResource allocationMulti core computingOpen platform
The invention discloses a scheduling method of a cloud computing open platform, which particularly characterized in that call requests of mass users are monitored through maintaining a central server; business service and data service of a cluster server are dynamically scheduled; and meanwhile, detachable service components are called and a multithread ability of a multi-core processor is scheduled and distributed according to use levels of the users. Through using the method, the defects that the traditional open platform is short of expansibility and flexibility can be effectively corrected, two targets of rapidly constructing and deploying an application and operation environments and dynamically adjusting the application and operation environments are established, the existing equipment is maximally utilized and the service is maximally constructed; and meanwhile, through adopting a multi-core computing manner and a multithread technology, the scheduling speed and the response ability of the system service are improved.
Owner:UNIV OF ELECTRONICS SCI & TECH OF CHINA
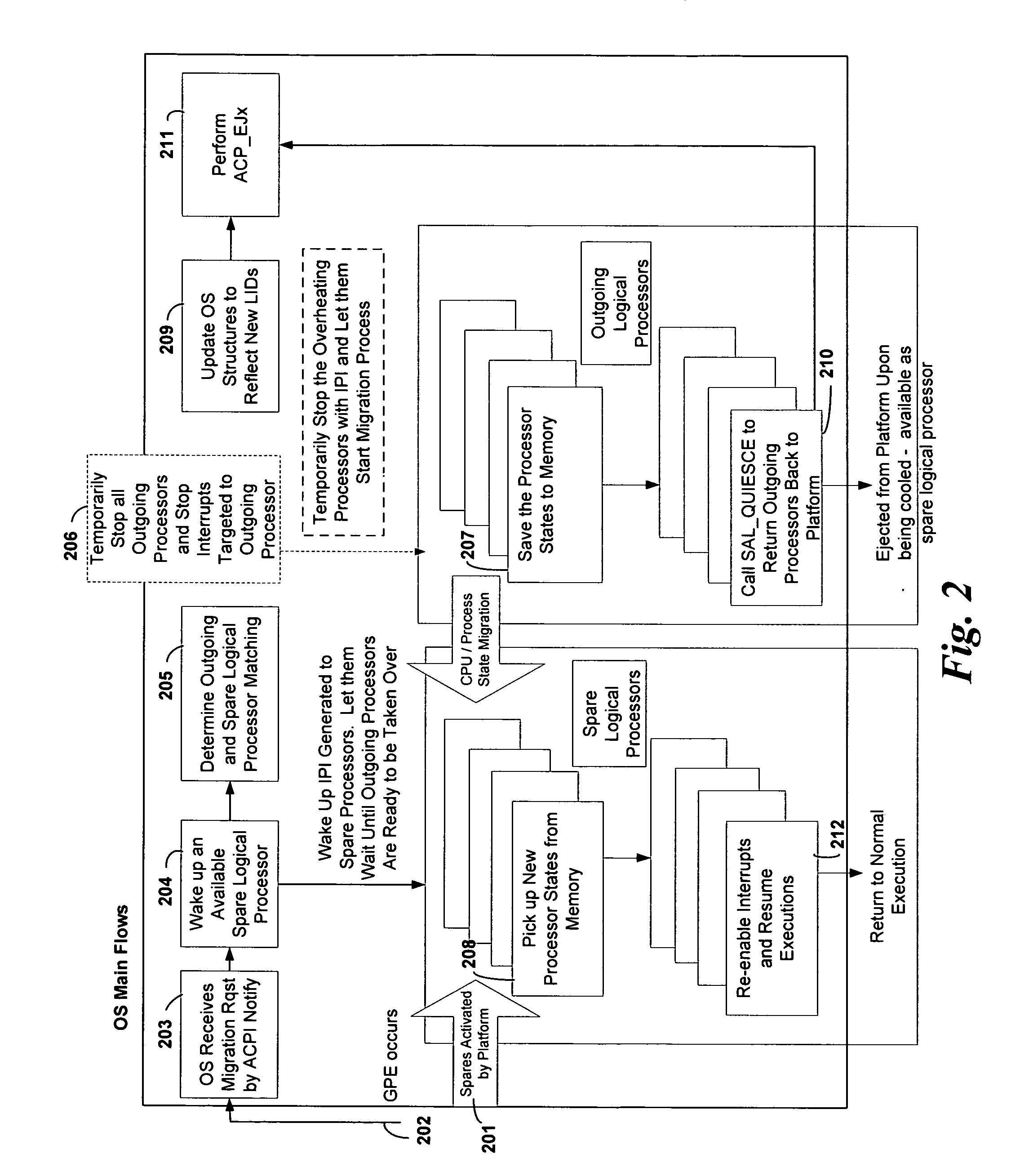
System and method to establish fine-grained platform control
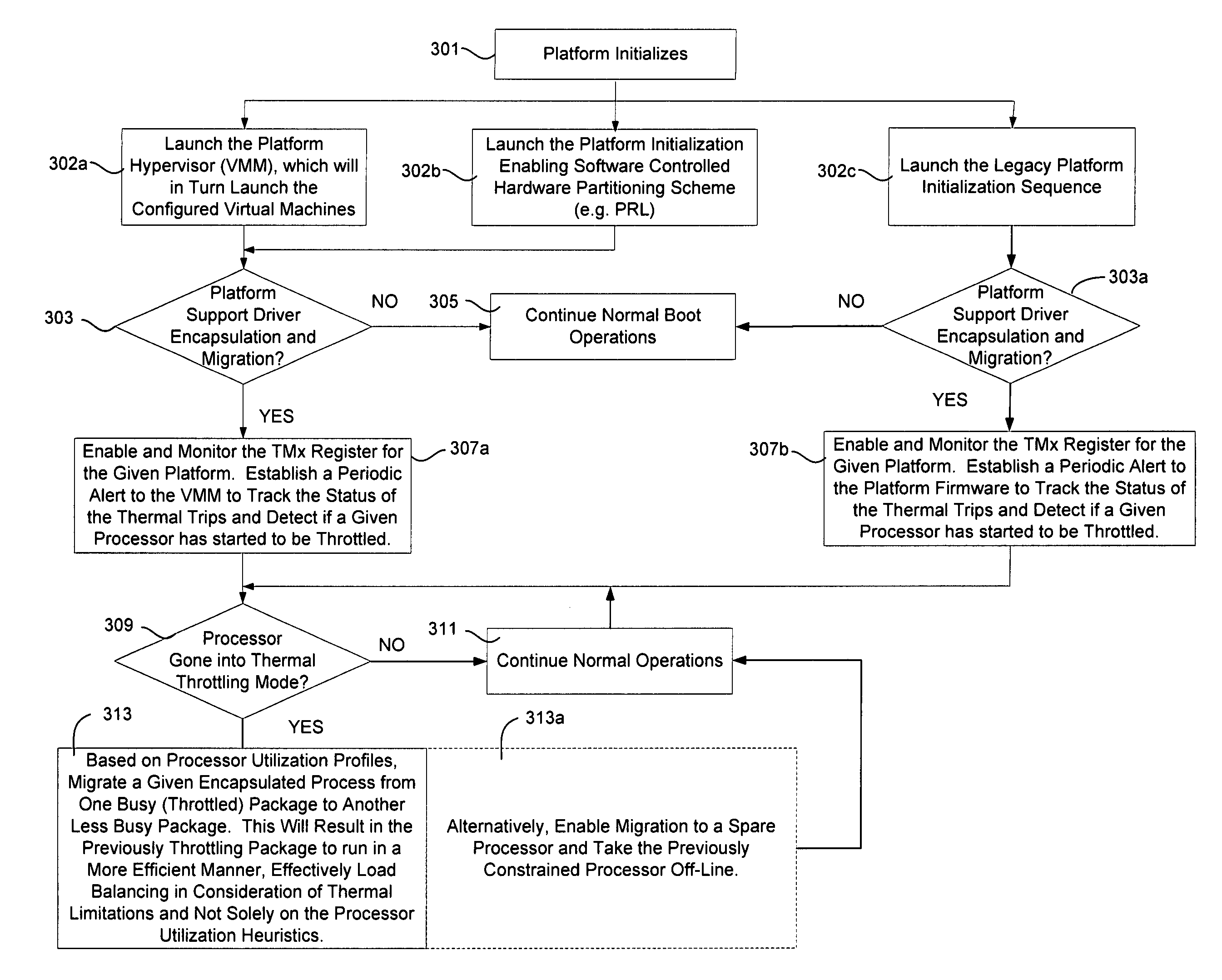
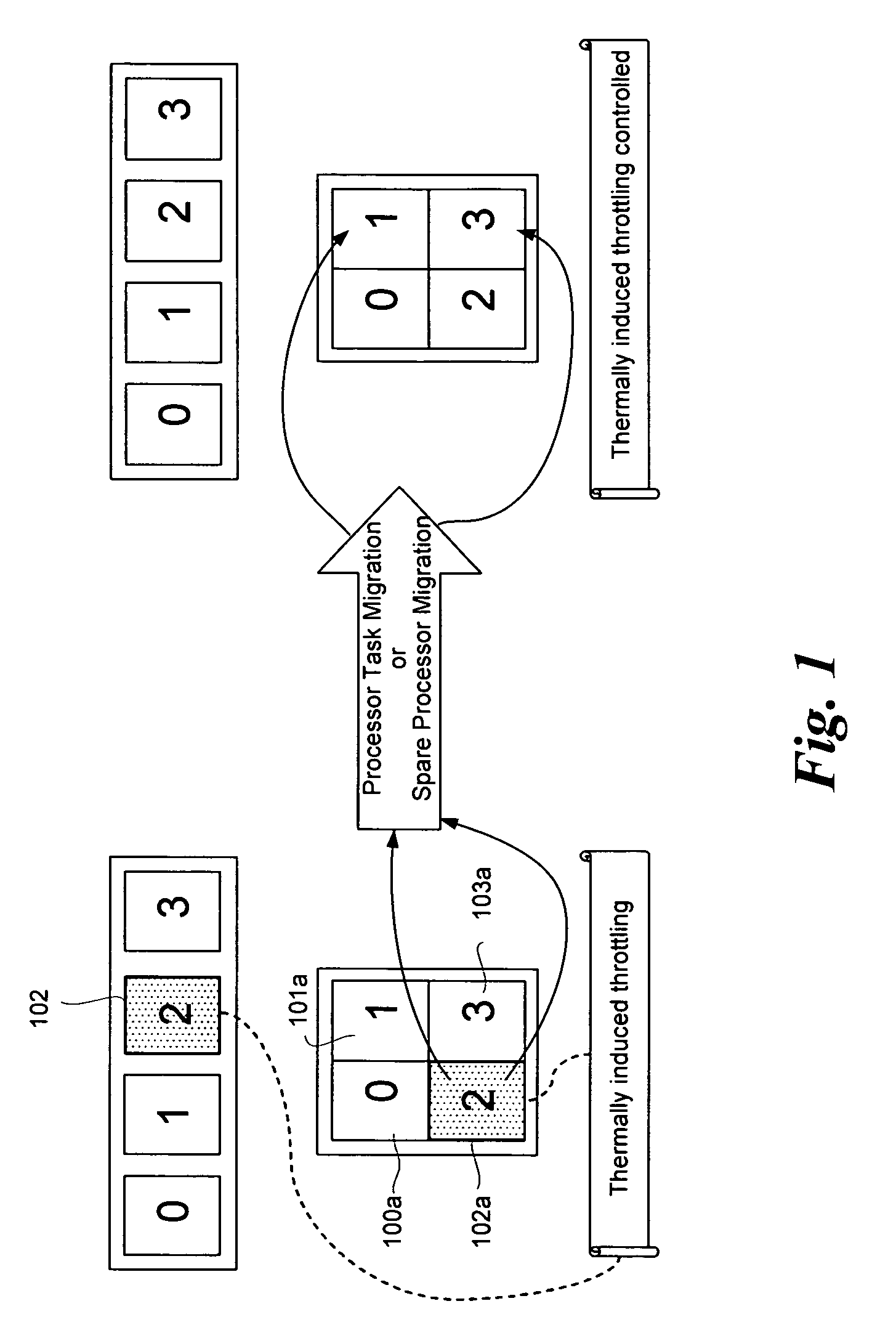
InactiveUS20080115010A1Avoid overall overheatingMinimize throttlingEnergy efficient ICTFault responseMulti core computingFine grain
In an embodiment, processes are to be migrated in a multi-core computing system. Task migration is performed between and among cores to prevent over tasking or overheating of individual cores. In a platform with multi-core processors, each core is thermally isolated and has individual thermal sensors to indicate overheating. Processes are migrated among cores, and possibly among cores on more than one processor, to efficiently load balance the platform to avoid undue throttling or ultimate shutdown of an overheated processor. Utilization profiles may be used to determine which core(s) are to be used for task migration. Other embodiments are described and claimed.
Owner:INTEL CORP
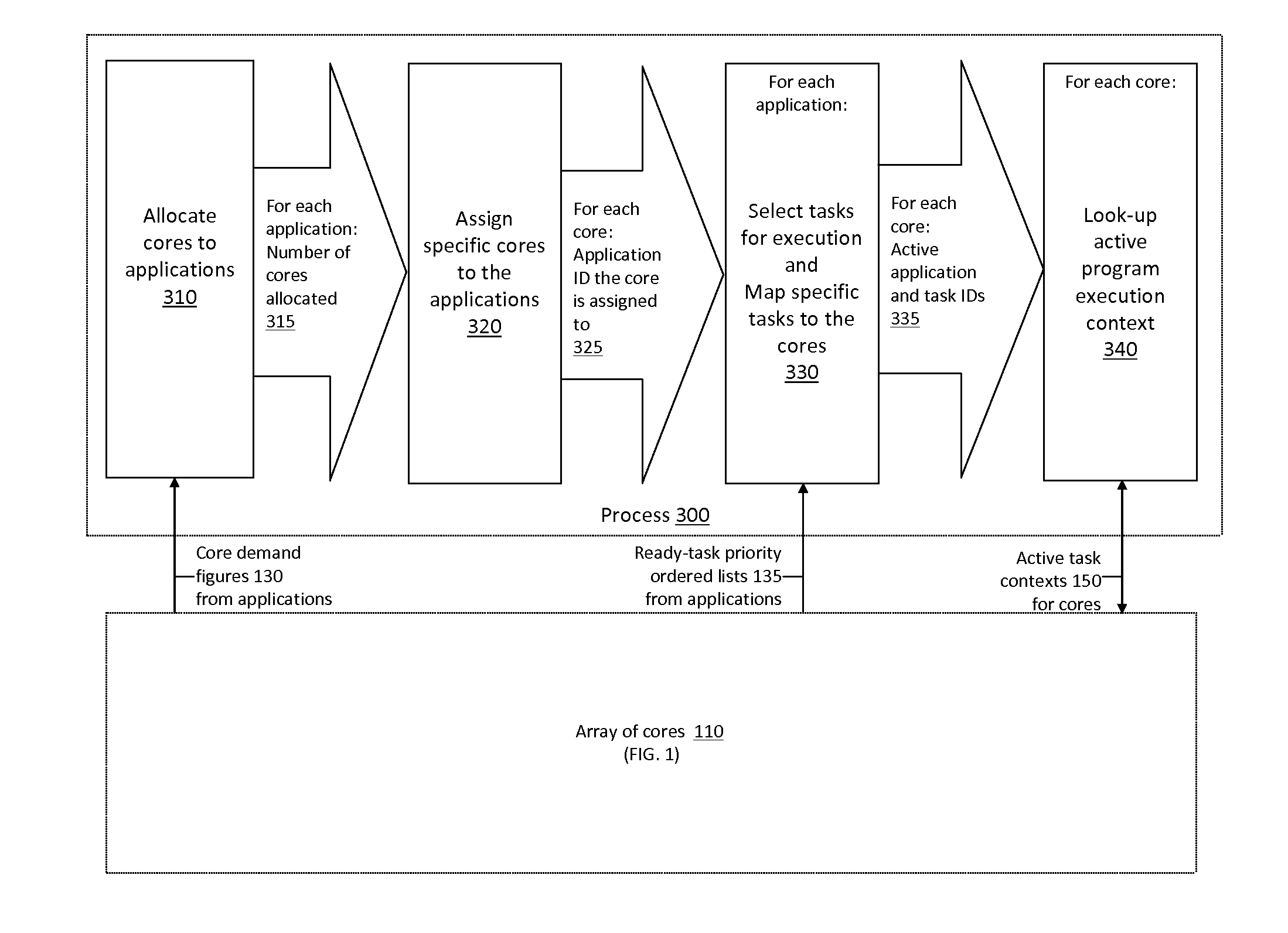
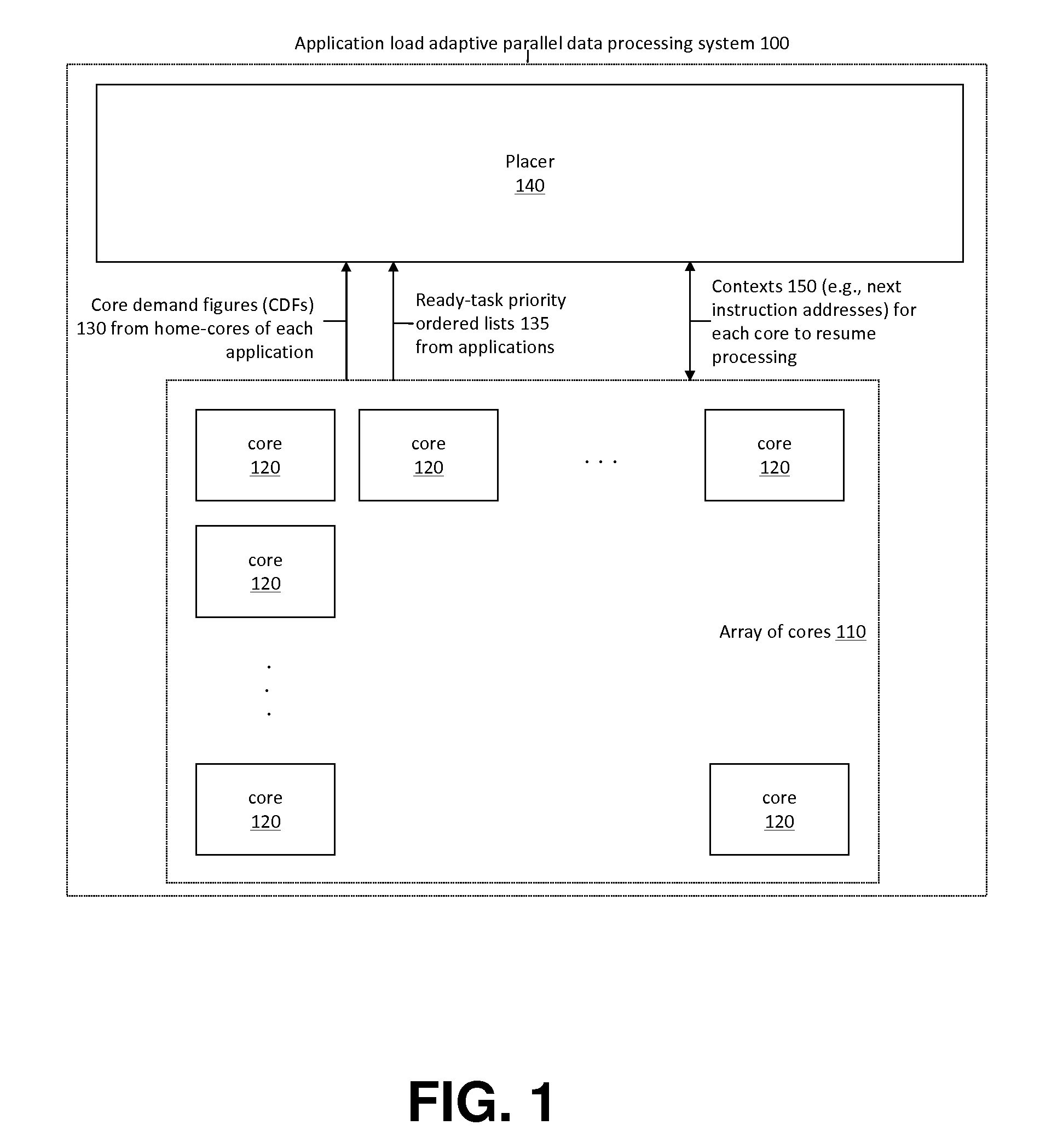
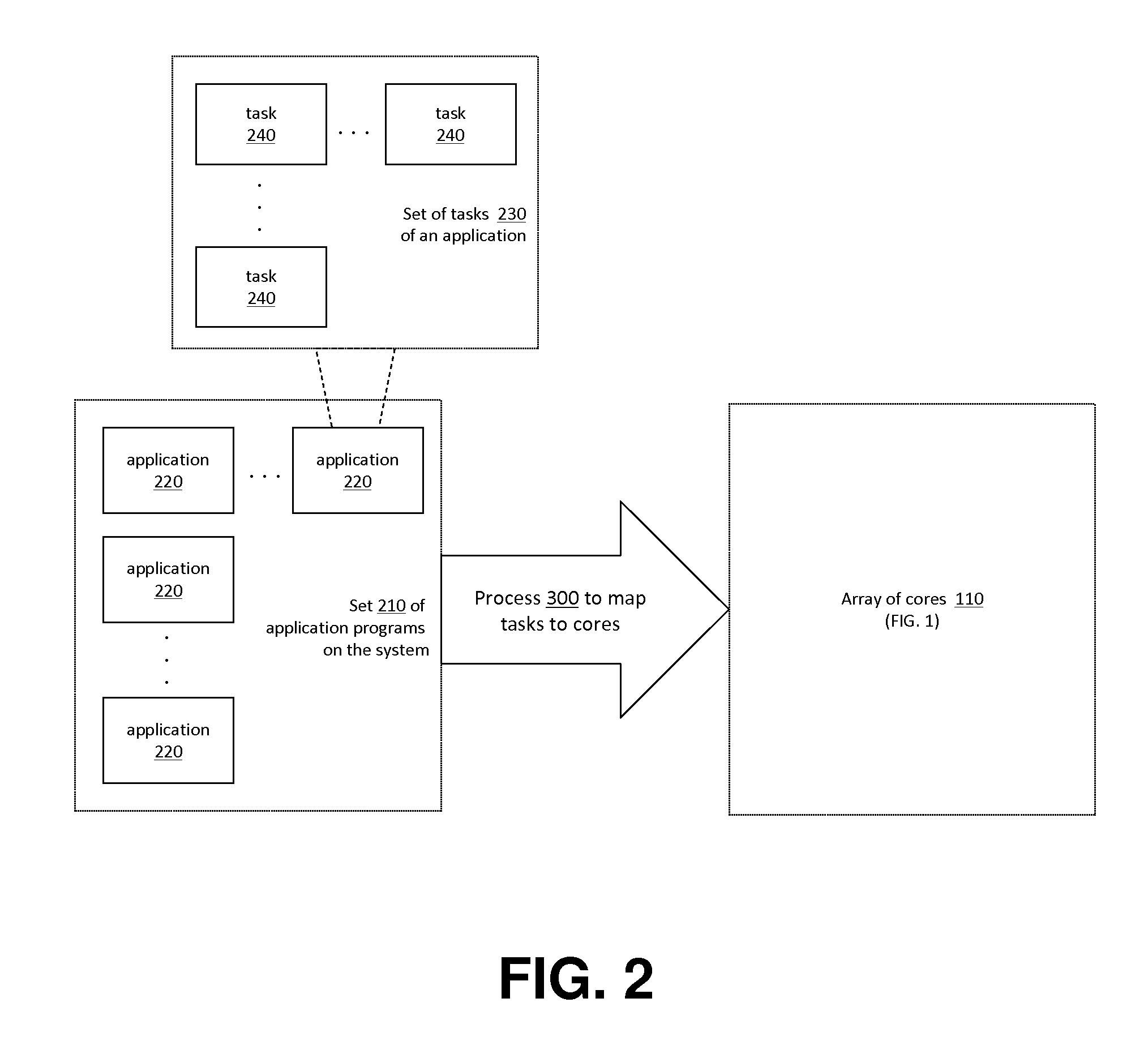
Application Load Adaptive Processing Resource Allocation
InactiveUS20120079501A1Maximized number of processing coreMaximizing whole system data processing throughputMultiprogramming arrangementsMemory systemsData processing systemMulti core computing
The invention provides hardware-automated systems and methods for efficiently sharing a multi-core data processing system among a number of application software programs, by dynamically reallocating processing cores of the system among the application programs in an application processing load adaptive manner. The invention enables maximizing the whole system data processing throughput, while providing deterministic minimum system access levels for each of the applications. With invented techniques, each application on a shared multi-core computing system dynamically gets a maximized number of cores that it can utilize in parallel, so long as all applications on the system still get at least up to their entitled number of cores whenever their actual processing load so demands. The invention provides inherent security and isolation between applications, as each application resides in its dedicated system memory segments, and can safely use the shared processing system as if it was the sole application running on it.
Owner:THROUGHPUTER
Control method for multiple operating systems of multi-core computer and multi-core computer
ActiveCN102929719AGuaranteed uptimeTake advantage of fault toleranceProgram initiation/switchingResource allocationMulti core computingOperational system
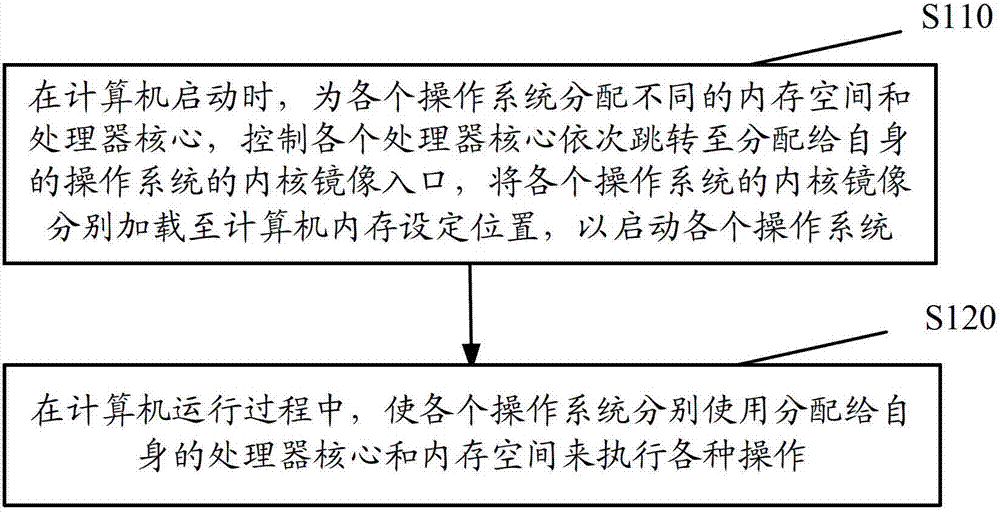
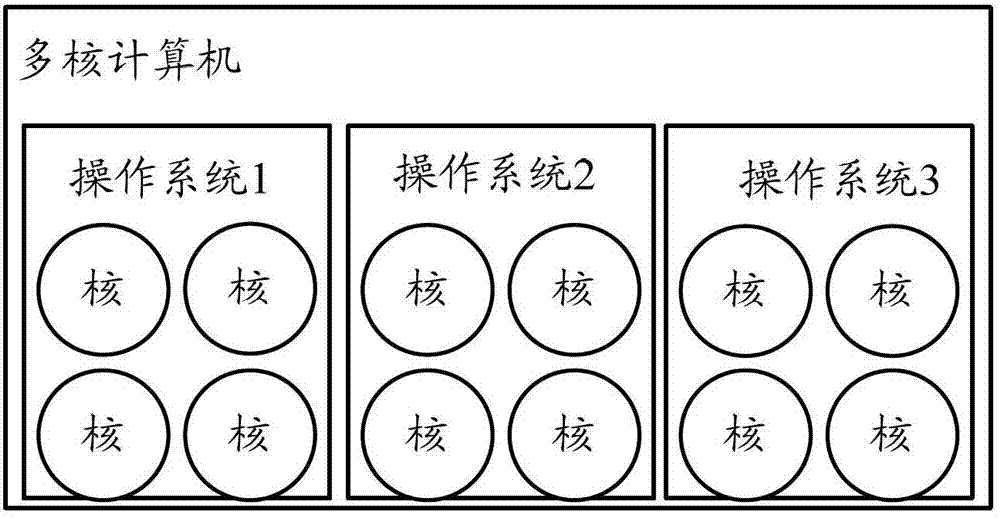
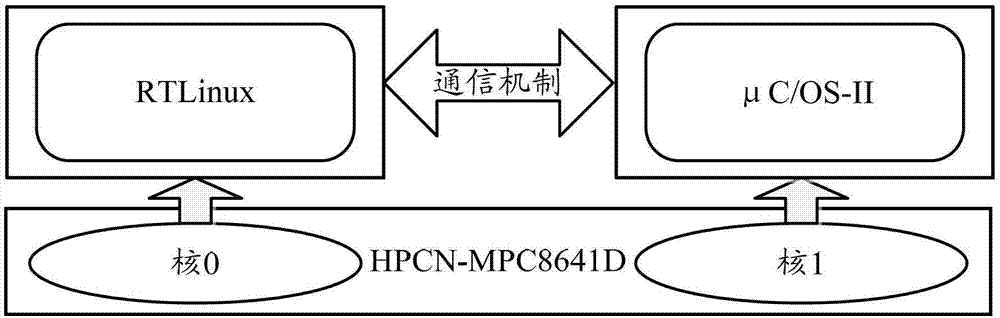
The invention discloses a control method for multiple operating systems of a multi-core computer, comprising the following steps: when the computer is started, different memory spaces and processor cores are allocated for the operating systems, the processor cores are controlled to skip to the respectively-allocated kernel image entrances of the operating systems in sequence, kernel images of the operating systems are respectively loaded to the preset address of the memory of the computer, so that the operating systems can be started; in the running process of the computer, the operating systems respectively use the respectively-allocated processor cores and memory spaces to execute various operations. According to the control method, different memory spaces and processor cores are allocated for multiple operating systems, the multiple operating system installed in the multi-core processor are controlled to manage the multiple processor cores, each operating system uses the allocated privately-owned memory, so that the error tolerance of the multi-core processor is sufficiently utilized while the computing capability of the multi-core processor is utilized.
Owner:CHINA STANDARD SOFTWARE
Devices and Methods for Optimizing Data-Parallel Processing in Multi-Core Computing Systems
InactiveUS20120131584A1Digital computer detailsMultiprogramming arrangementsMulti core computingMulti processor
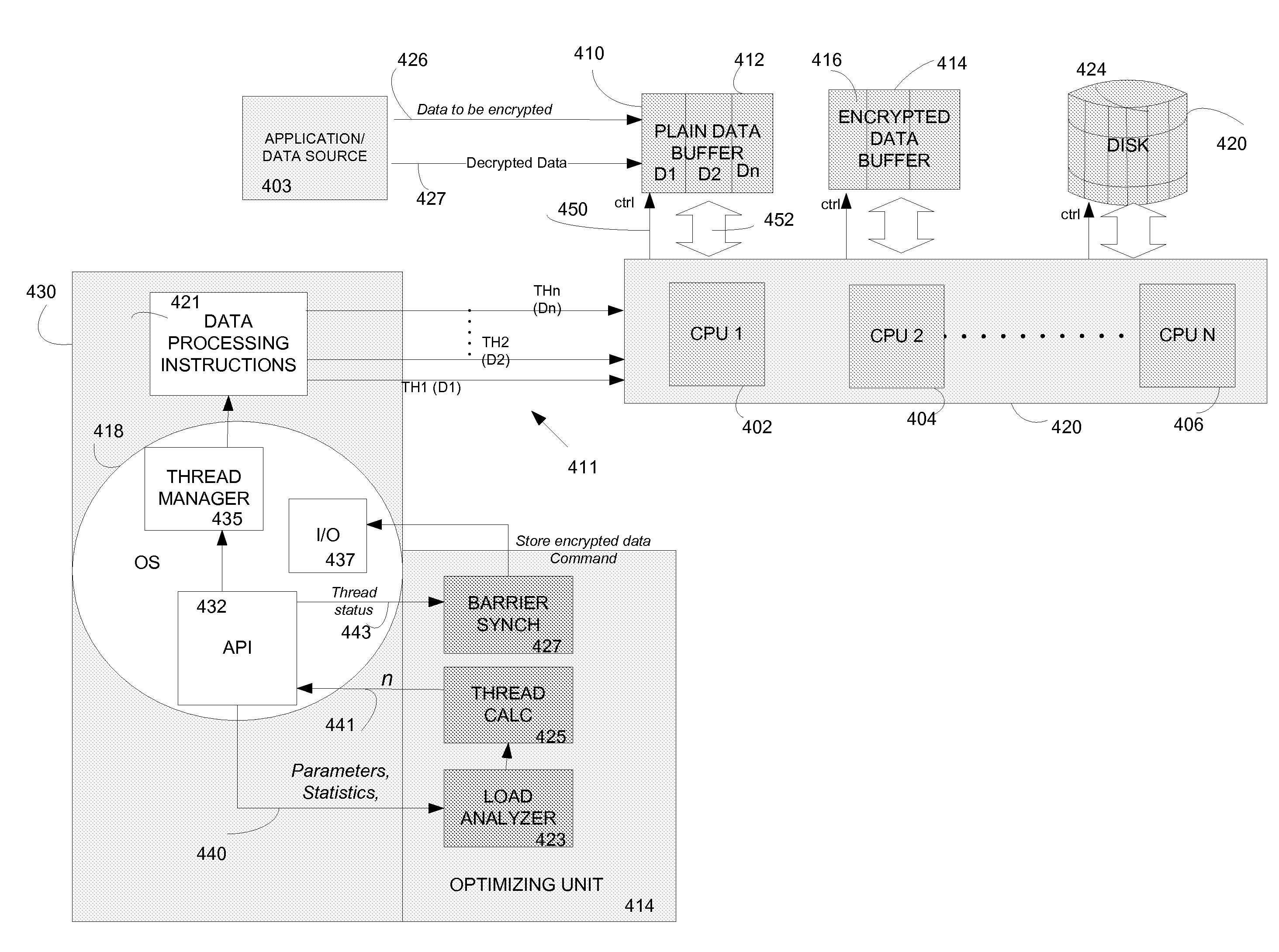
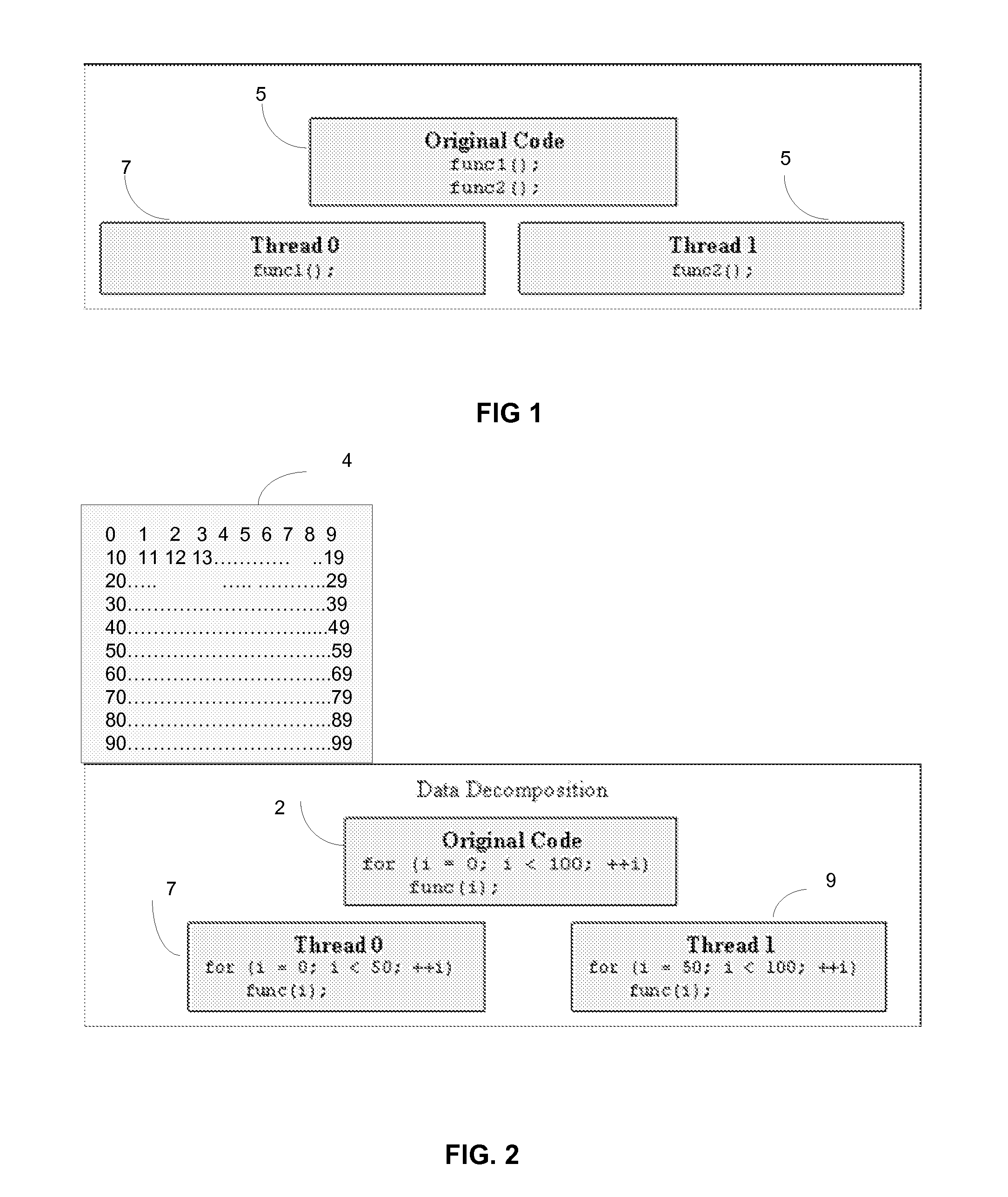
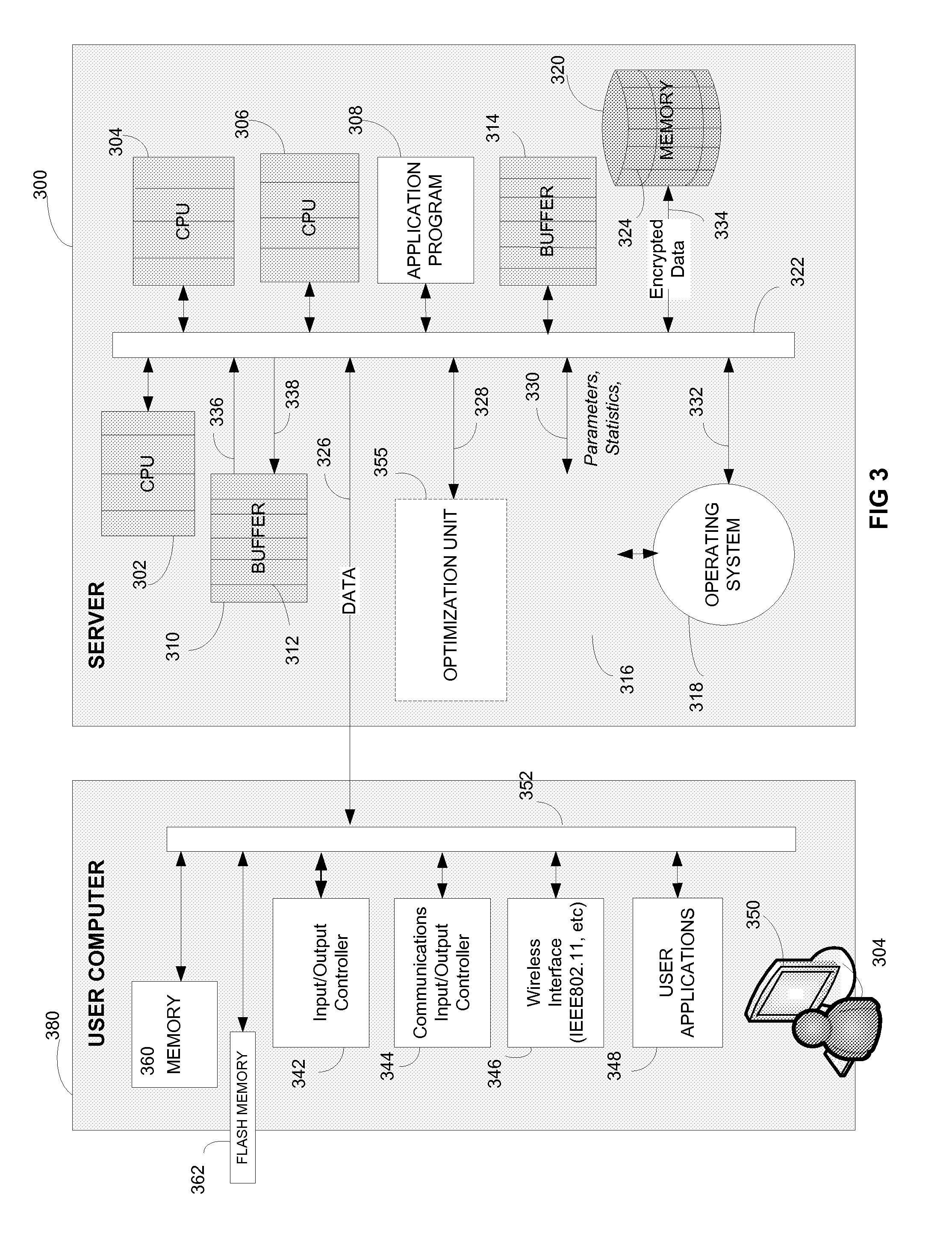
According to an embodiment of a method of the invention, at least a portion of data to be processed is loaded to a buffer memory of capacity (B). The buffer memory is accessible to N processing units of a computing system. The processing task is divided into processing threads. An optimal number (n) of processing threads is determined by an optimizing unit of the computing system. The n processing threads are allocated to the processing task and executed by at least one of the N processing units. After processing by at least one of N processing units, the processed data is stored on a disk defined by disk sectors, each disk sector having storage capacity (S). The storage capacity (B) of the buffer memory is optimized to be a multiple X of sector storage capacity (S). The optimal number (n) is determined based, at least in part on N, B and S. The system and method are implementable in a multithreaded, multi-processor computing system. The stored encrypted data may be later recalled and decrypting using the same system and method.
Owner:RAEVSKY ALEXEY
Task scheduling system of on-chip multi-core computing platform and method for task parallelization
ActiveCN102129390AJudging relevanceImprove parallelismMultiprogramming arrangementsMultiple digital computer combinationsMulti core computingService module
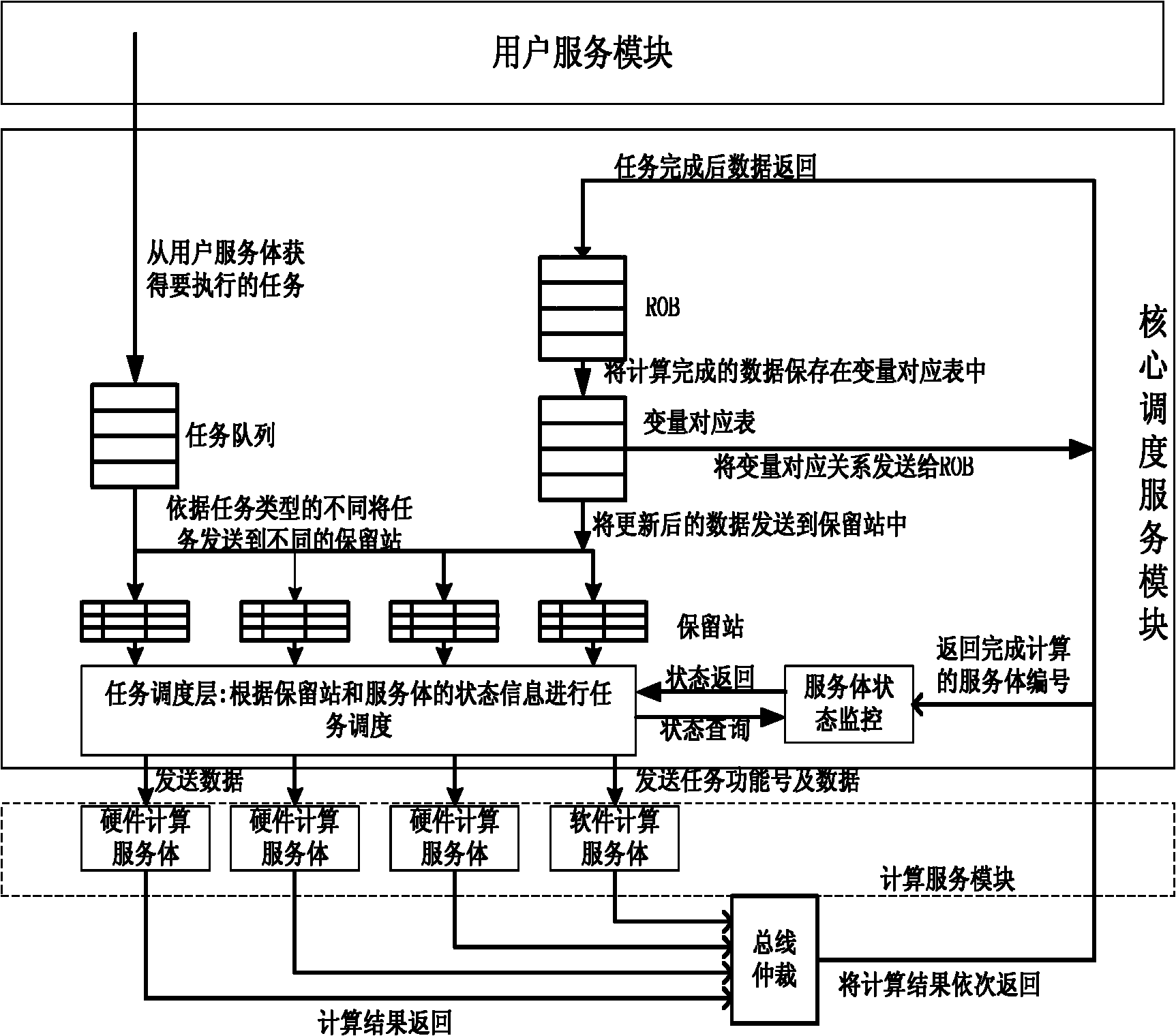
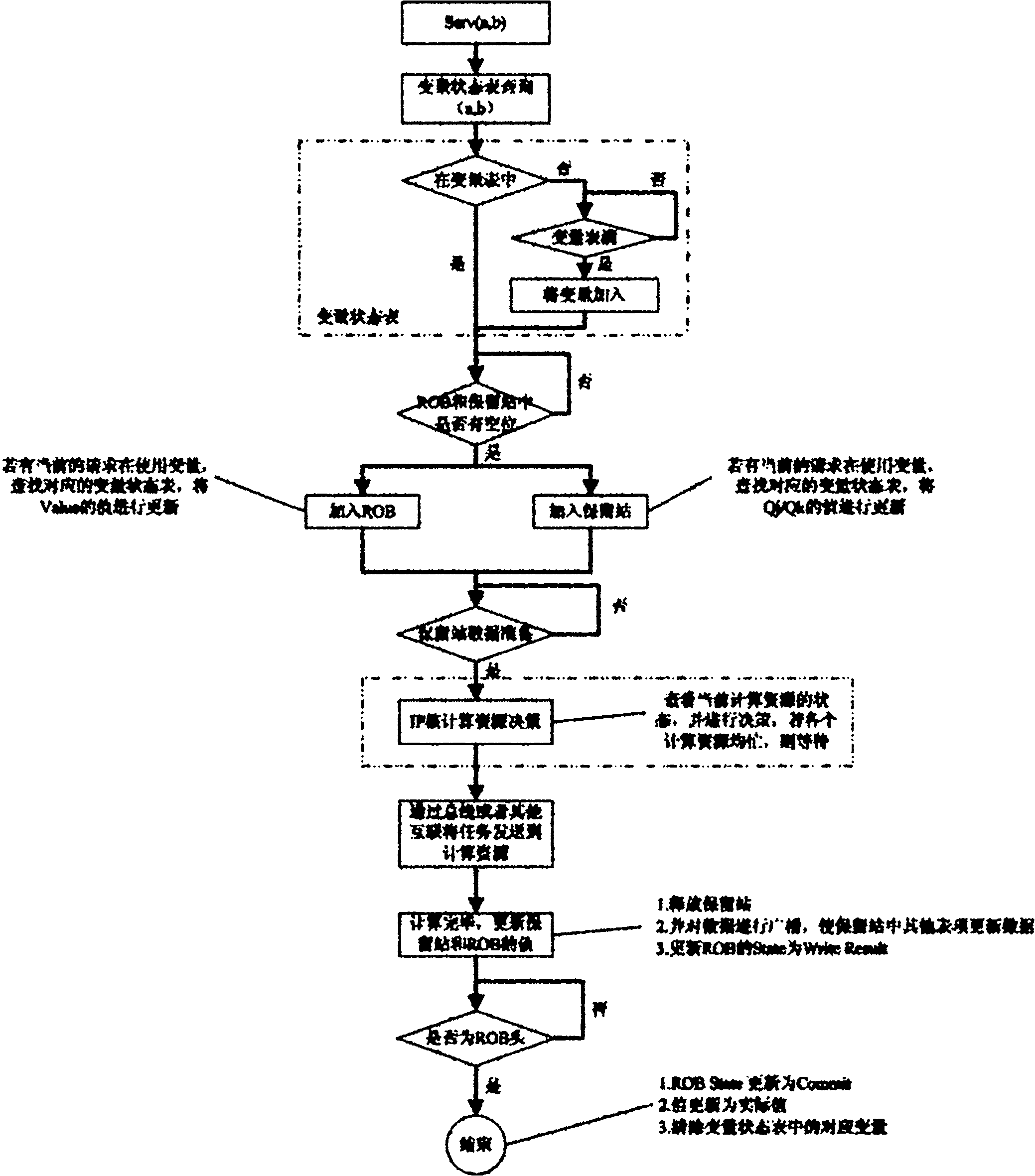
The invention discloses a task scheduling system of an on-chip multi-core computing platform and a method for task parallelization, wherein the system comprises user service modules for providing tasks which are needed to be executed, and computation service modules for executing a plurality of tasks on the on-chip multi-core computing platform, and the system is characterized in that core scheduling service modules are arranged between the user service modules and the computation service modules, the core scheduling service modules receive task requests of the user service modules as input, judge the data dependency relations among different tasks through records, and schedule the task requests in parallel to different computation service modules for being executed. The system enhances platform throughput and system performance by performing correlation monitoring and automatic parallelization on the tasks during running.
Owner:SUZHOU INST FOR ADVANCED STUDY USTC
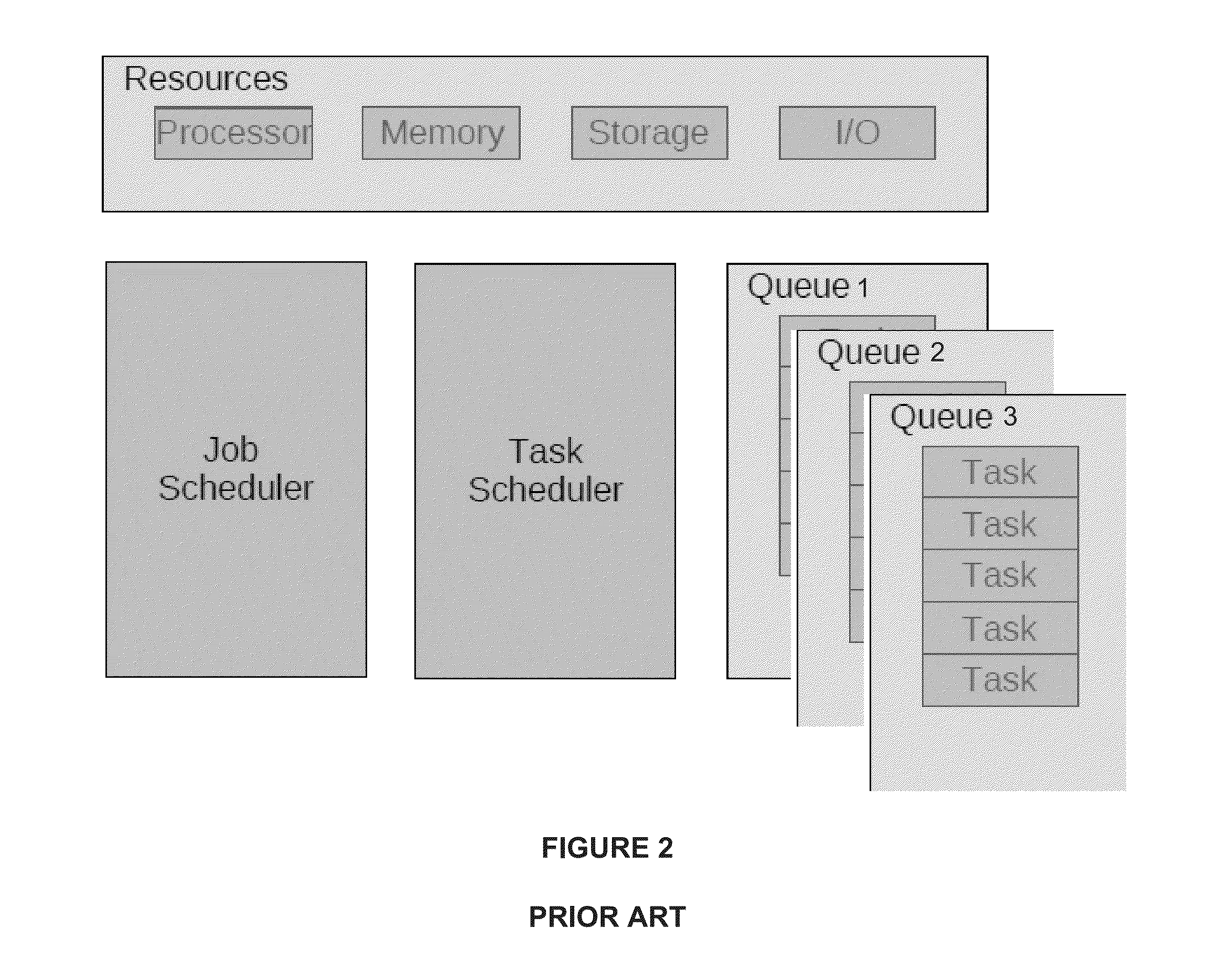
Efficient method for the scheduling of work loads in a multi-core computing environment
InactiveUS20130061233A1Maximize useReduce executionResource allocationMemory systemsComputer resourcesMulti core computing
A computer in which a single queue is used to implement all of the scheduling functionalities of shared computer resources in a multi-core computing environment. The length of the queue is determined uniquely by the relationship between the number of available work units and the number of available processing cores. Each work unit in the queue is assigned an execution token. The value of the execution token represents an amount of computing resources allocated for the work unit. Work units having non-zero execution tokens are processed using the computing resources allocate to each one of them. When a running work unit is finished, suspended or blocked, the value of the execution token of at least one other work unit in the queue is adjusted based on the amount of computing resources released by the running work unit.
Owner:EXLUDUS
System and Method for Massively Multi-Core Computing Systems
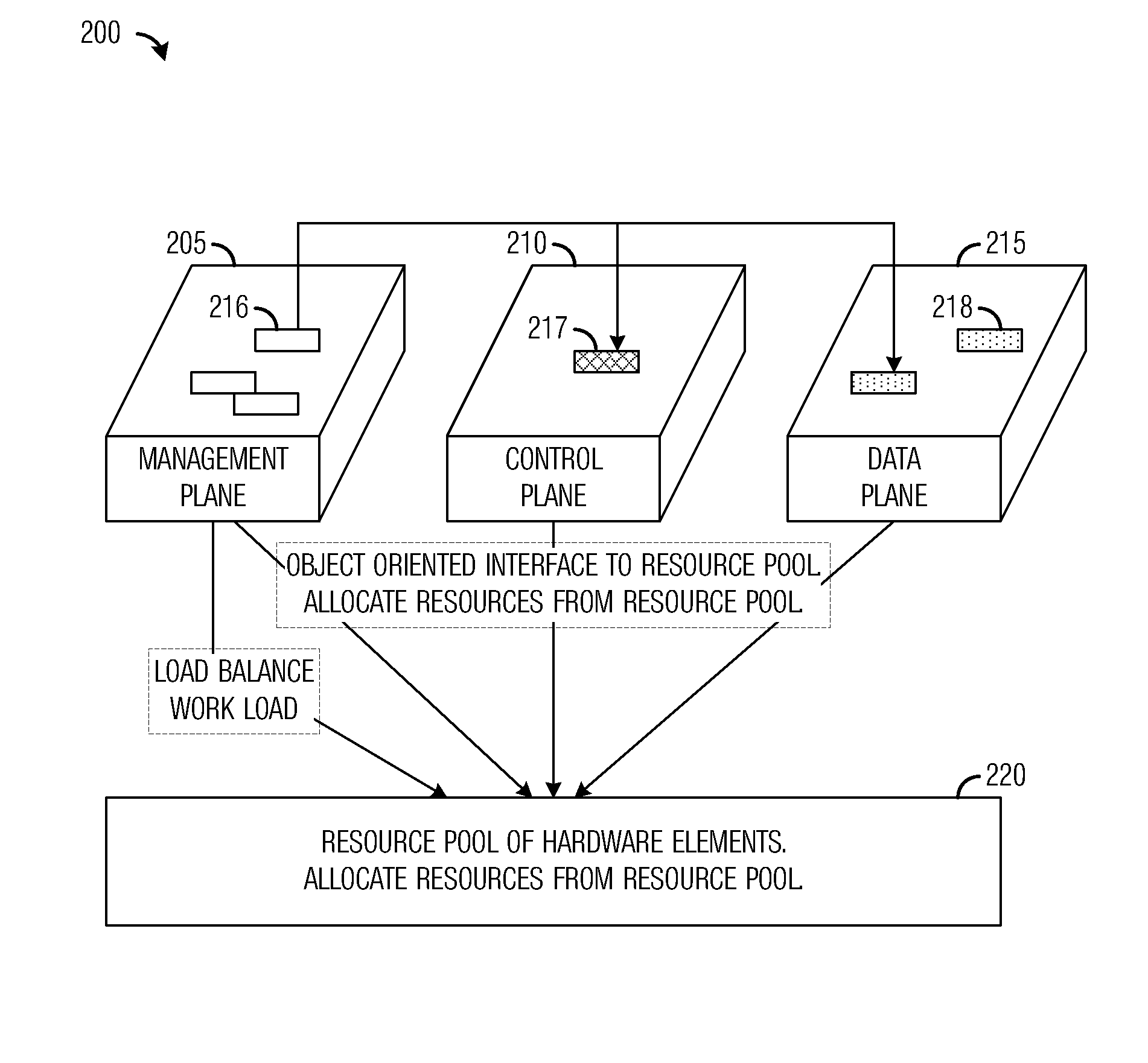
ActiveUS20120198465A1Reduce power consumptionAdjust power consumptionEnergy efficient ICTDigital computer detailsResource poolMulti core computing
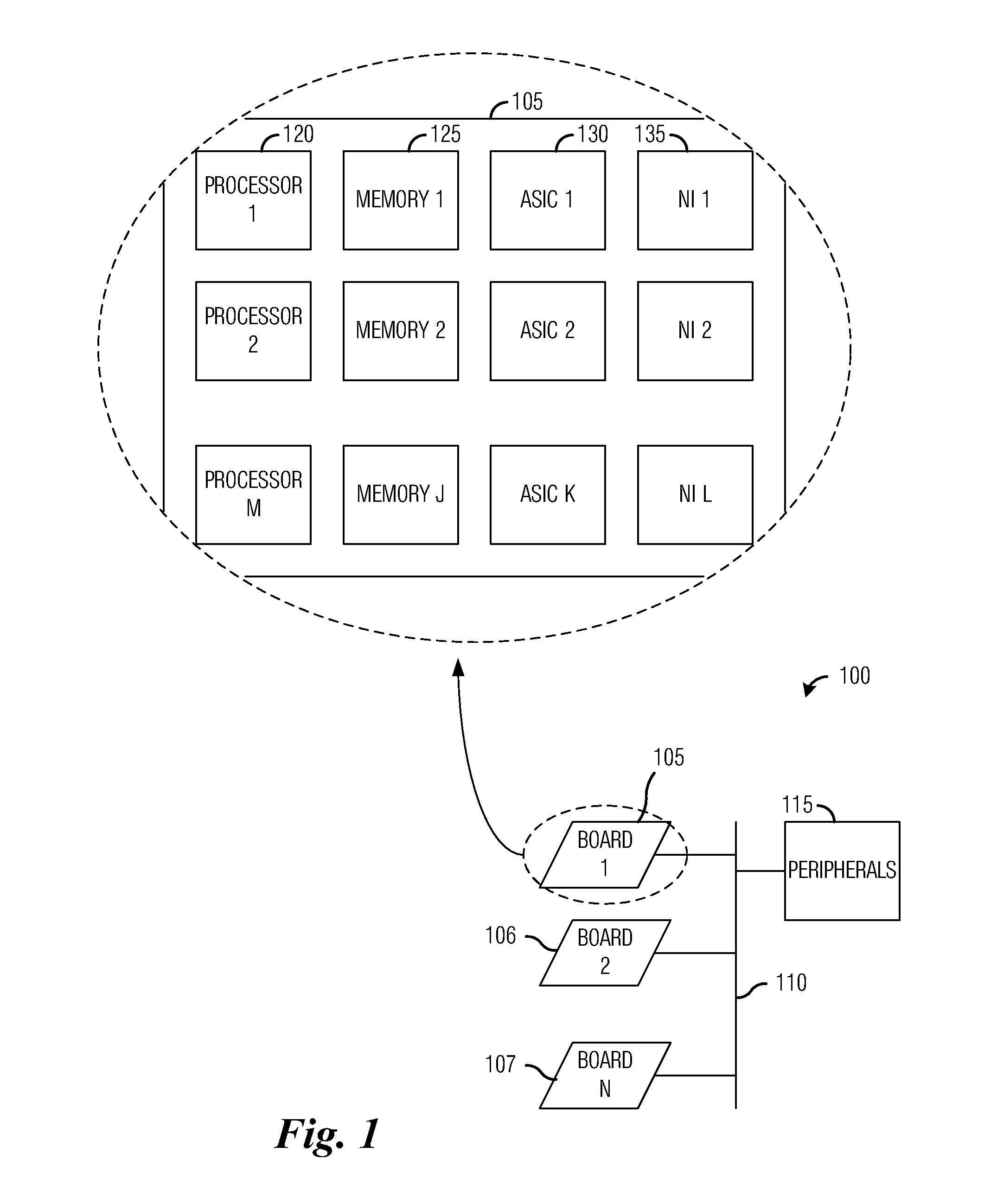
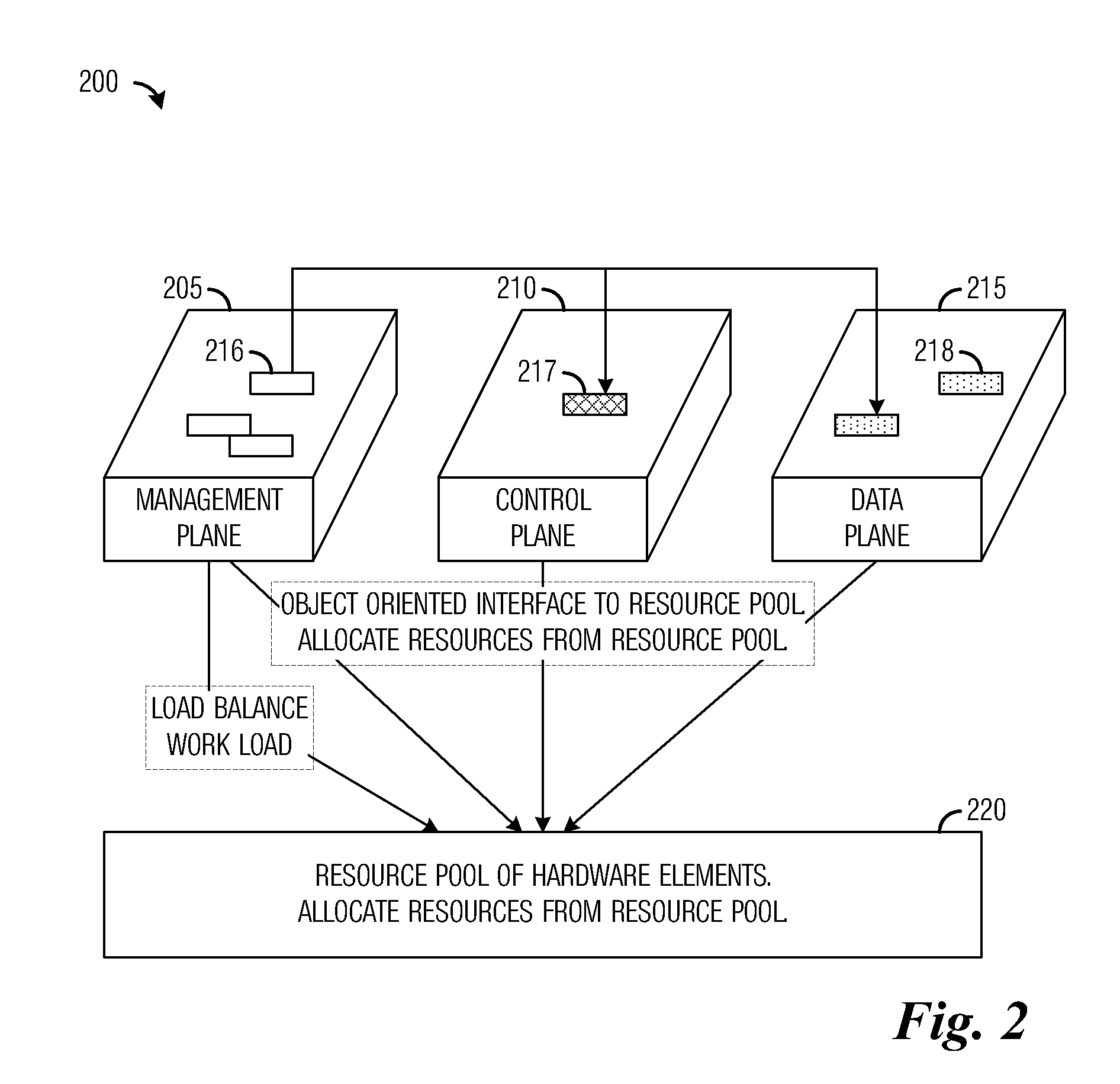
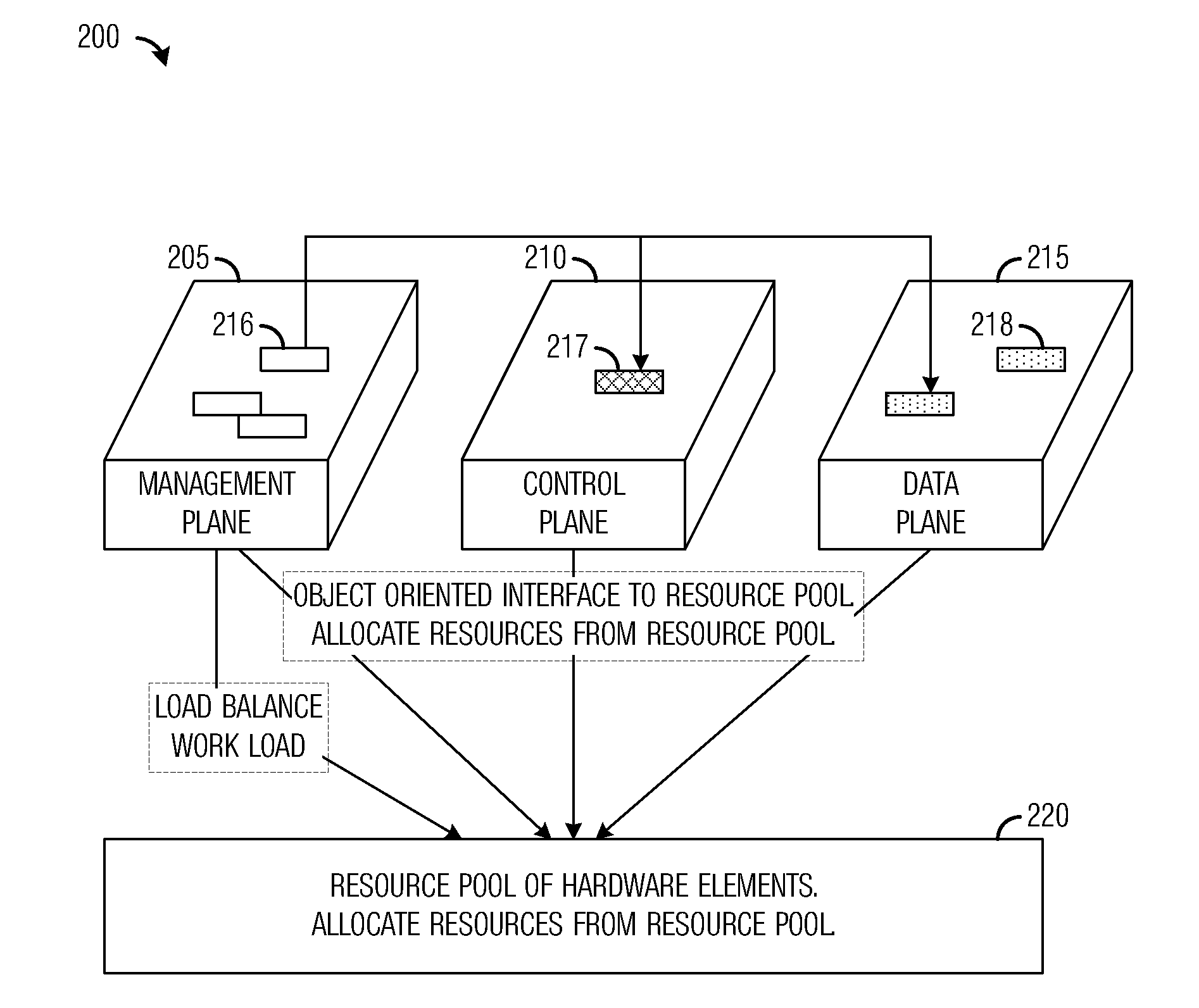
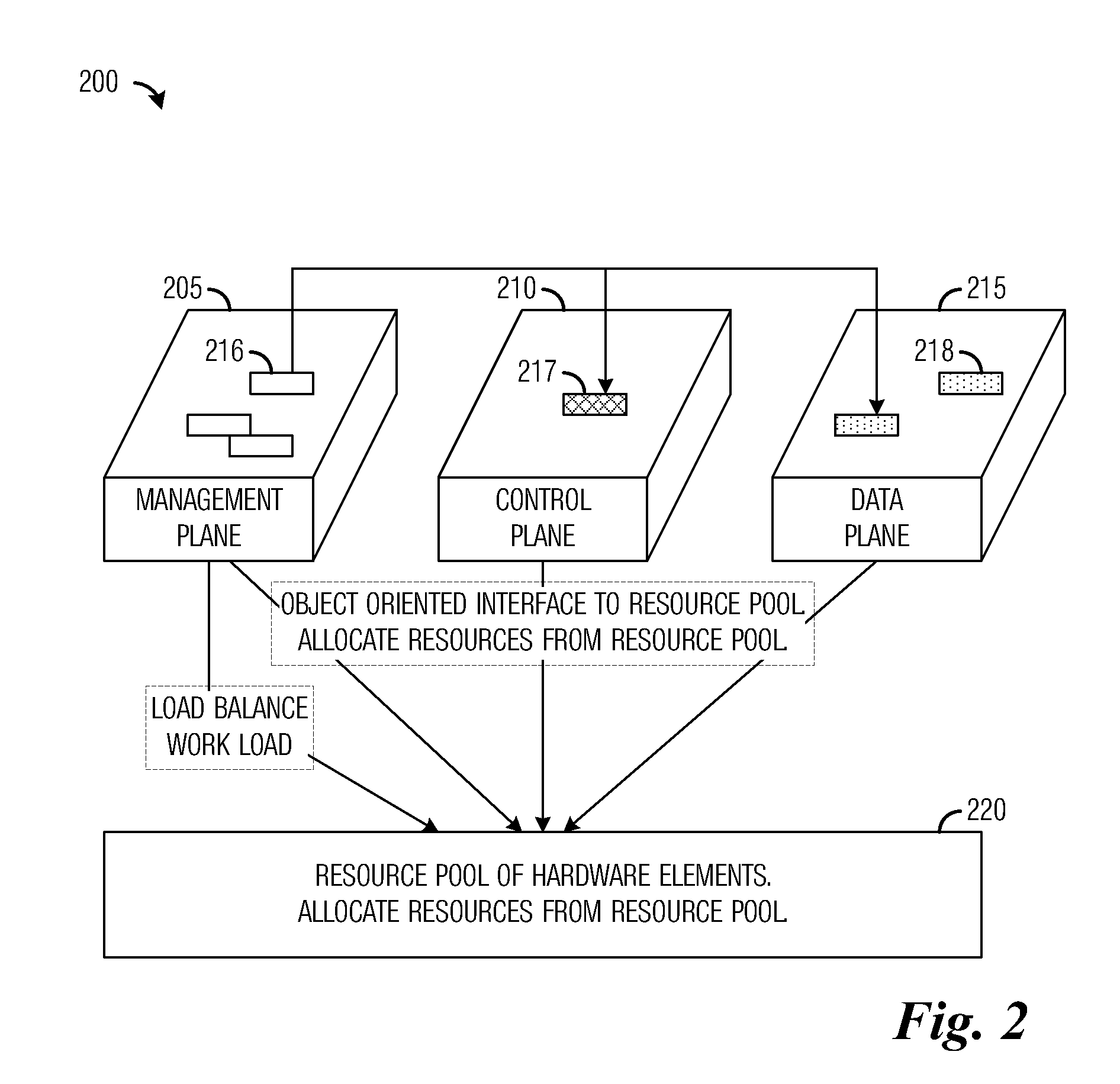
A system and method for massively multi-core computing are provided. A method for computer management includes determining if there is a need to allocate at least one first resource to a first plane. If there is a need to allocate at least one first resource, the at least one first resource is selected from a resource pool based on a set of rules and allocated to the first plane. If there is not a need to allocate at least one first resource, it is determined if there is a need to de-allocate at least one second resource from a second plane. If there is a need to de-allocate at least one second resource, the at least one second resource is de-allocated. The first plane includes a control plane and / or a data plane and the second plane includes the control plane and / or the data plane. The resources are unchanged if there is not a need to allocate at least one first resource and if there is not a need to de-allocate at least one second resource.
Owner:FUTUREWEI TECH INC
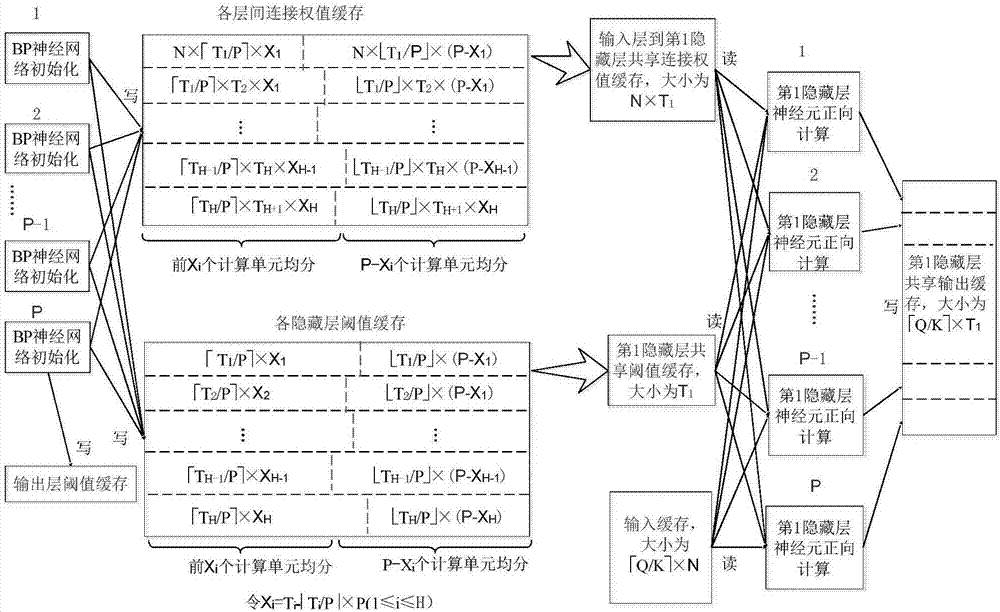
BP neural network parallelization method for multi-core computing environment
InactiveCN106909971AReduce replacementImprove parallelismResource allocationNeural learning methodsHidden layerMulti core computing
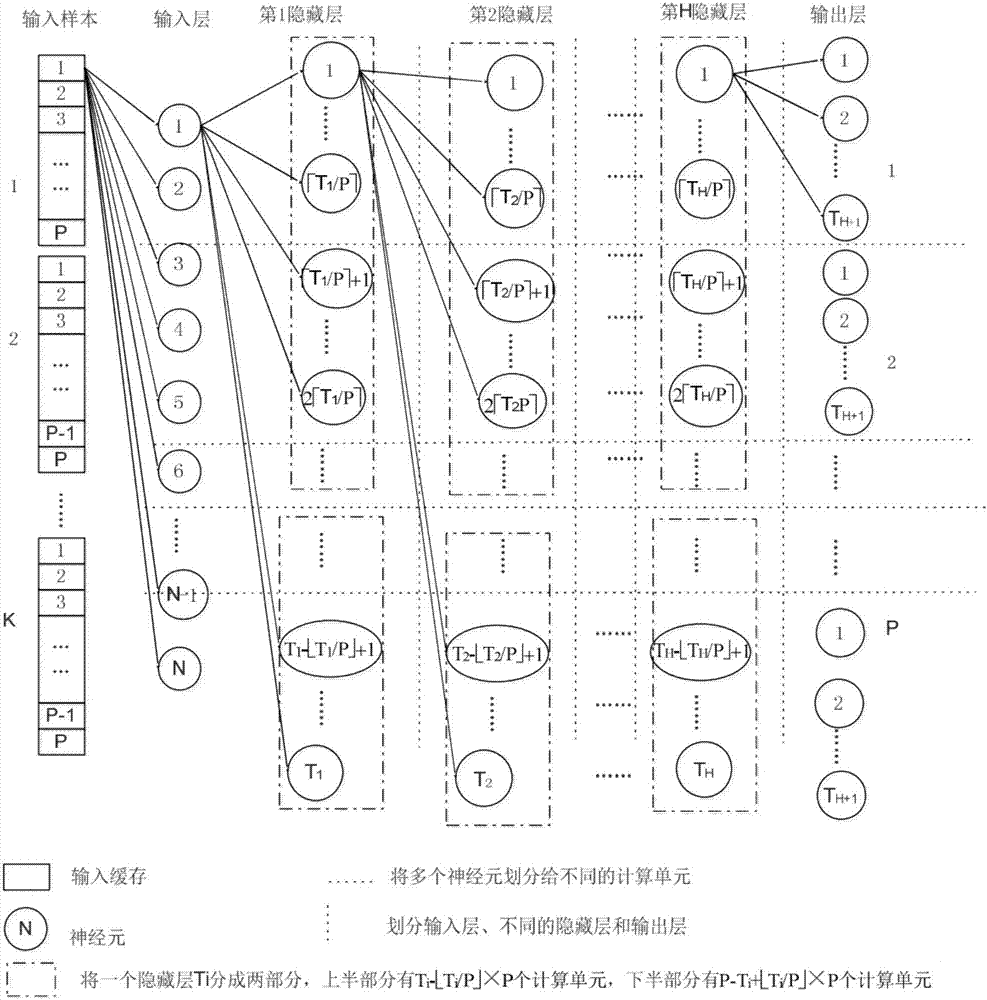
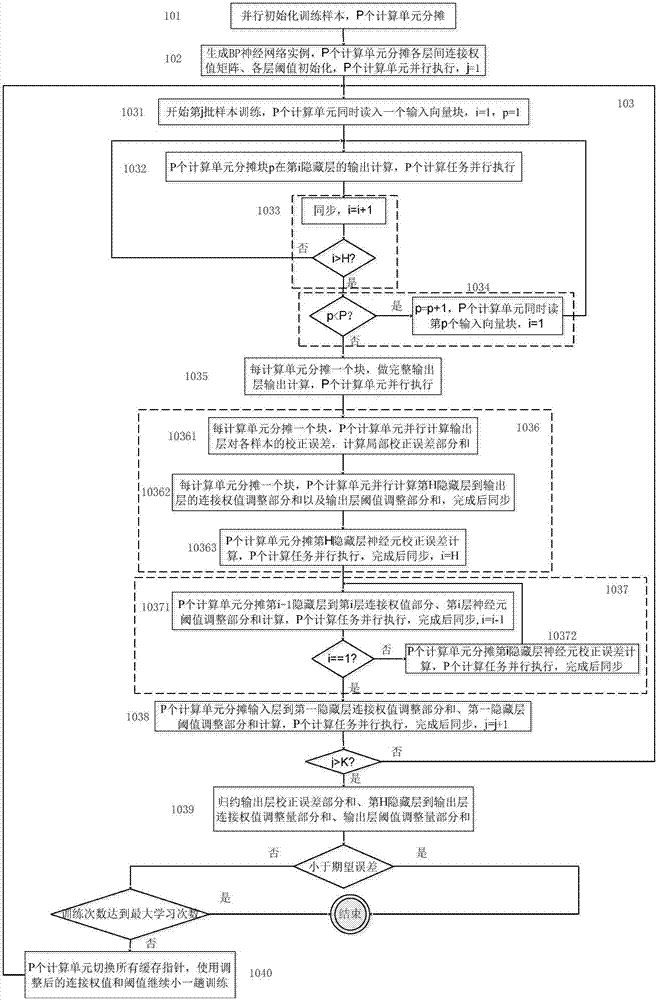
The invention provides a BP neural network parallelization method oriented to a multi-core computing environment. Including: parallel calculation task division and mapping method; cache setting method for storing intermediate calculation results of each layer; parallel training method of BP neural network. For the hardware platform containing P computing units, the task division method combines the hidden layer and output layer computing tasks into a larger granularity task as a whole, so as to improve parallelism; the cache setting method, in a training process, The same variable can be used in the next subtask after being accessed by the previous subtask, and the next subtask will not cause Cache loss; in the BP neural network training method, the samples are divided into K batches to enter the network training, and the K value is designed on a comprehensive computing platform The size of the secondary cache, combined with the cache settings, maximizes hardware performance and is suitable for BP neural network applications under multi-core computing platforms.
Owner:SOUTH CHINA UNIV OF TECH +1
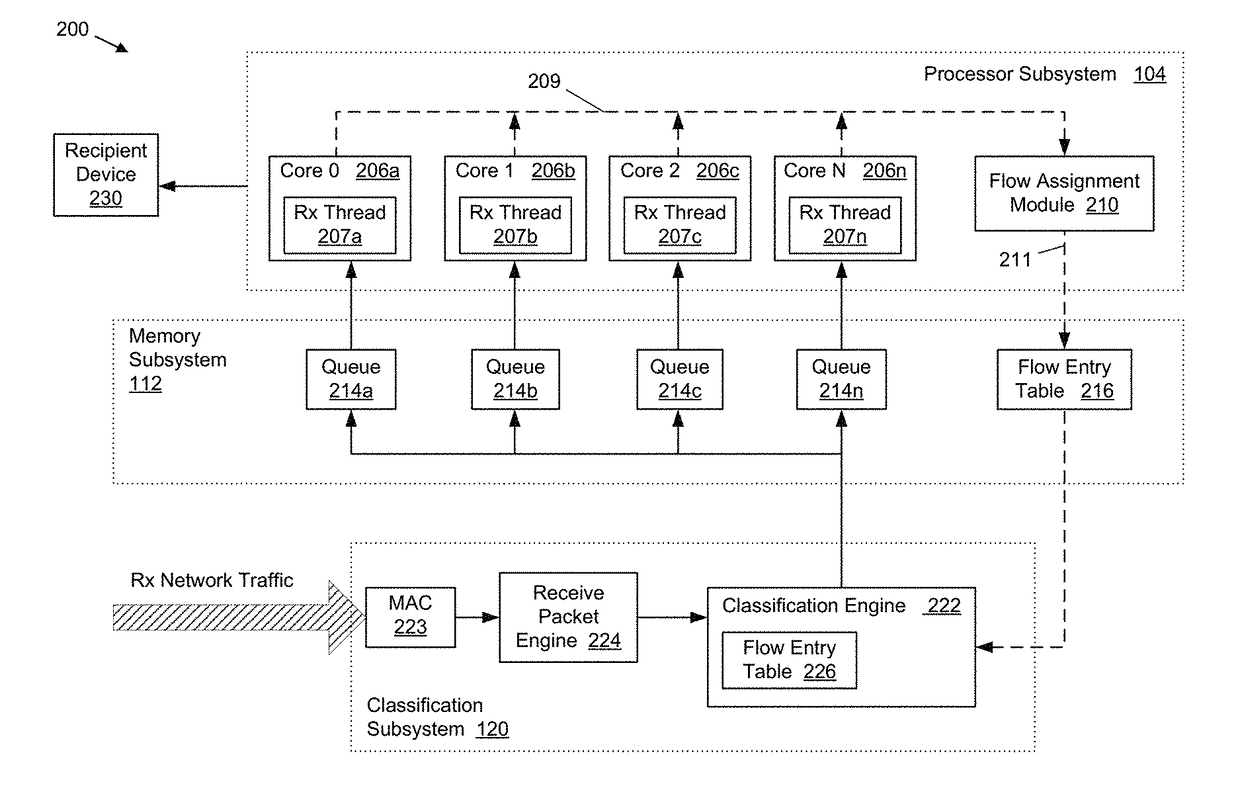
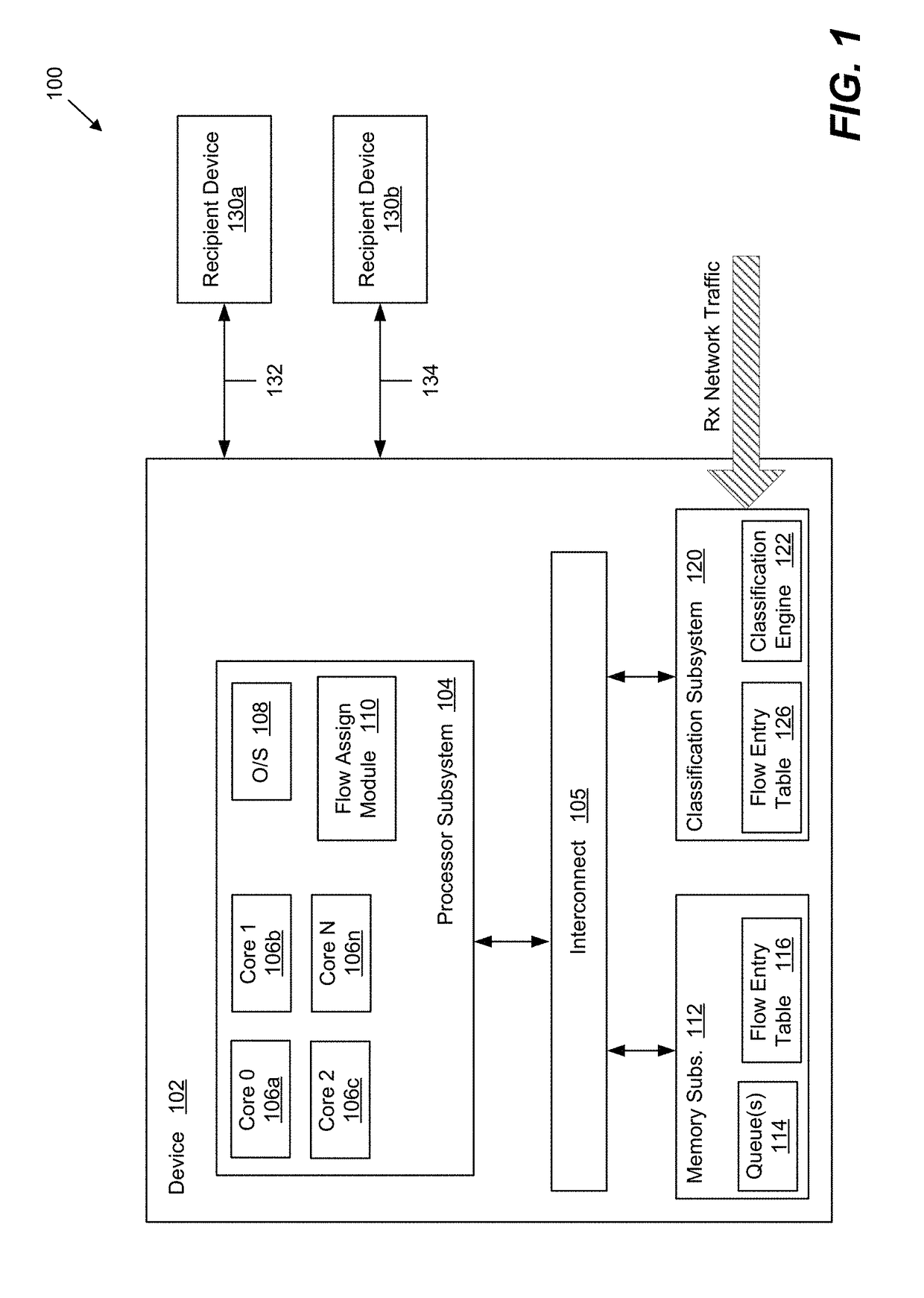
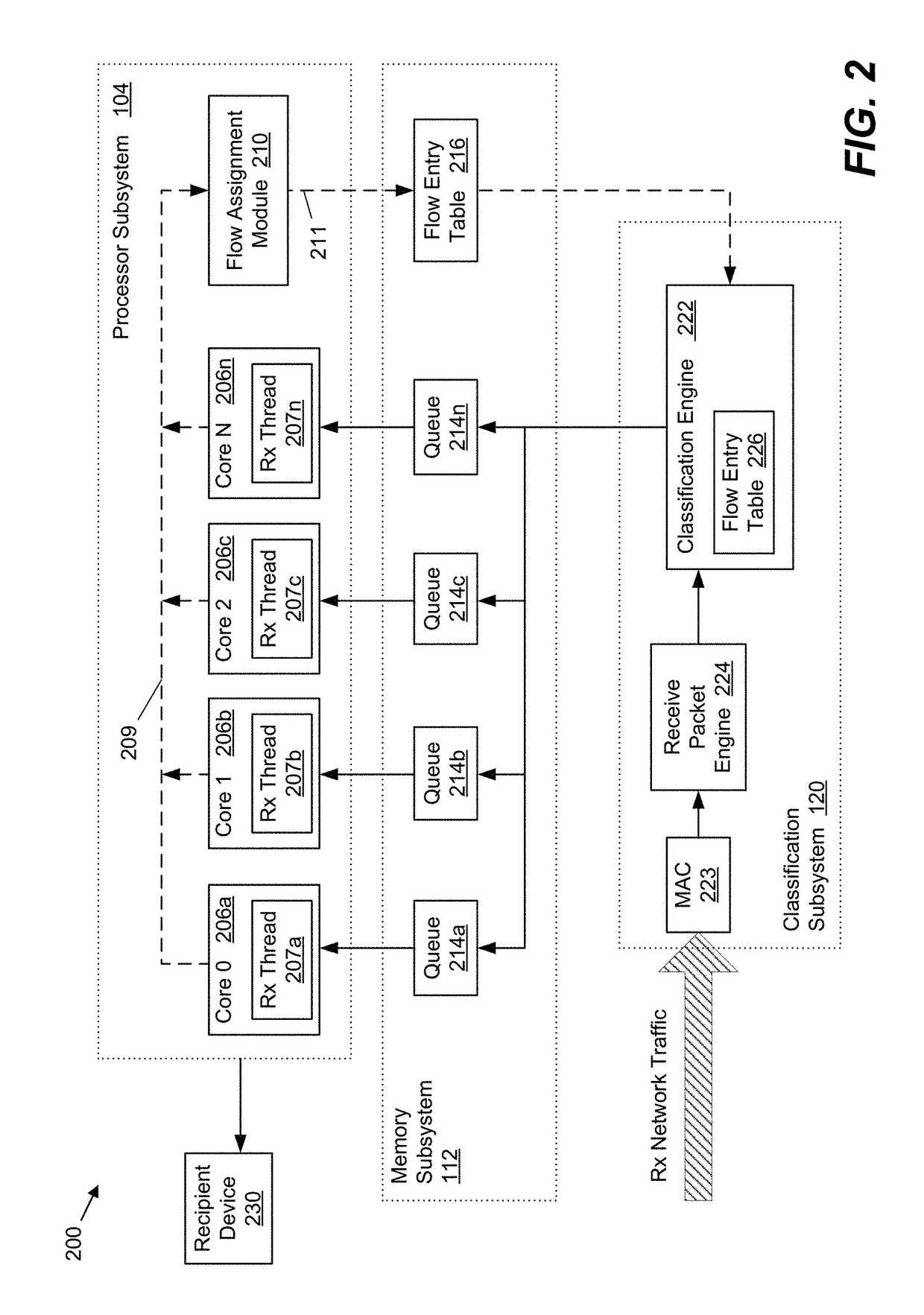
Method and system for providing efficient receive network traffic distribution that balances the load in multi-core processor systems
InactiveUS20170318082A1Efficient mappingAvoid congestionProgram controlData switching networksTraffic capacityMulti core computing
Systems and methods for improved received network traffic distribution in a multi-core computing device are presented. A hardware classification engine of the computing device receives a data packet comprising a portion of a received network traffic data flow. Packet information from the data packet is identified. Based in part on the packet information, the classification engine determines whether a core of a multi-core processor subsystem is assigned to the data flow of which the packet is a part. In embodiments, this determination may be made based on one or more criteria, such as a work load of the core(s) of the processor subsystem, a priority level of the data flow, etc. Responsive to the determination that a core is not assigned to the data flow, a core of the multi-core processor is assigned to the data flow and the data packet is sent to the first core for processing.
Owner:QUALCOMM INC
Performance acceleration method of heterogeneous multi-core computing platform on chip
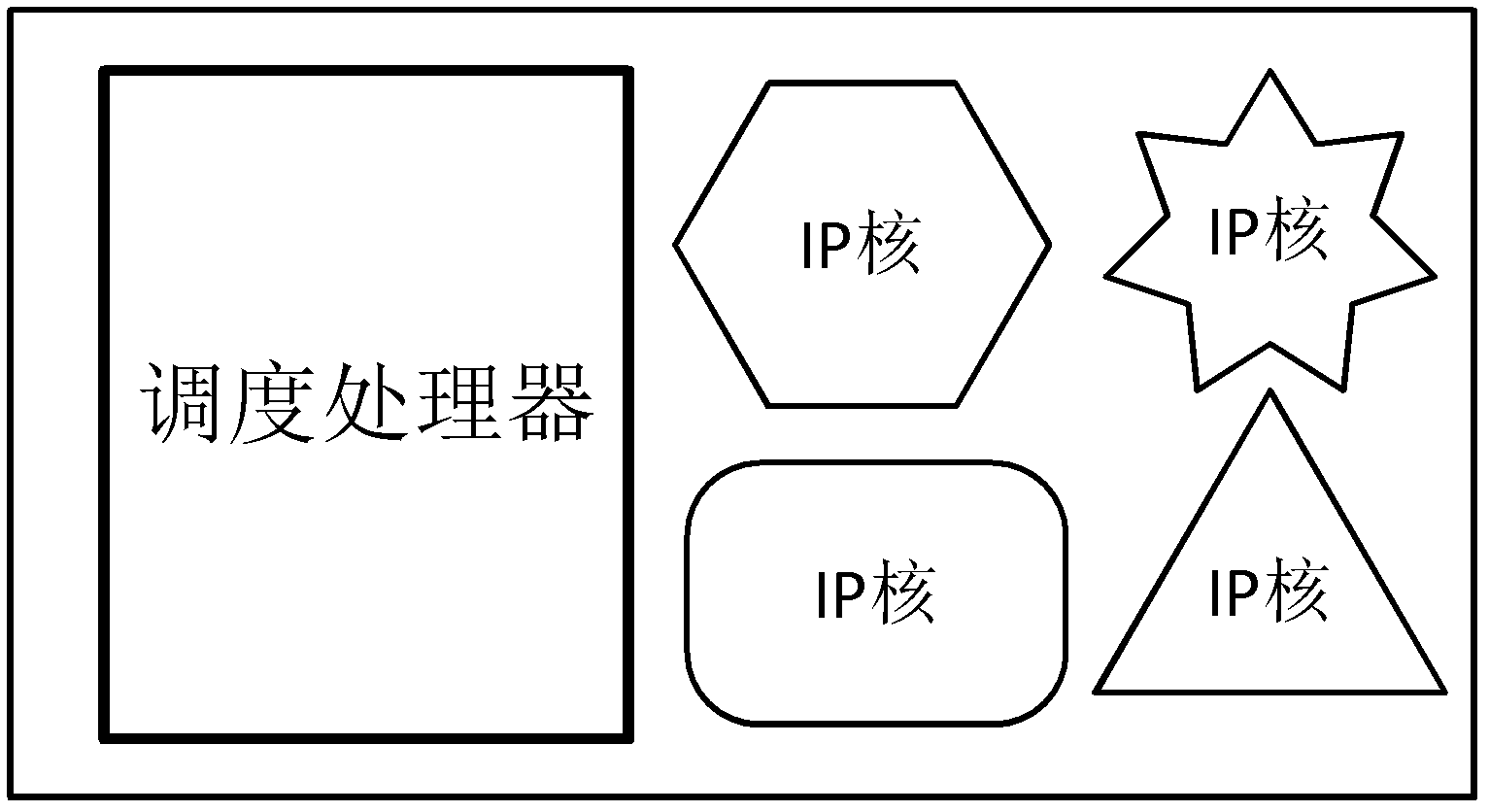
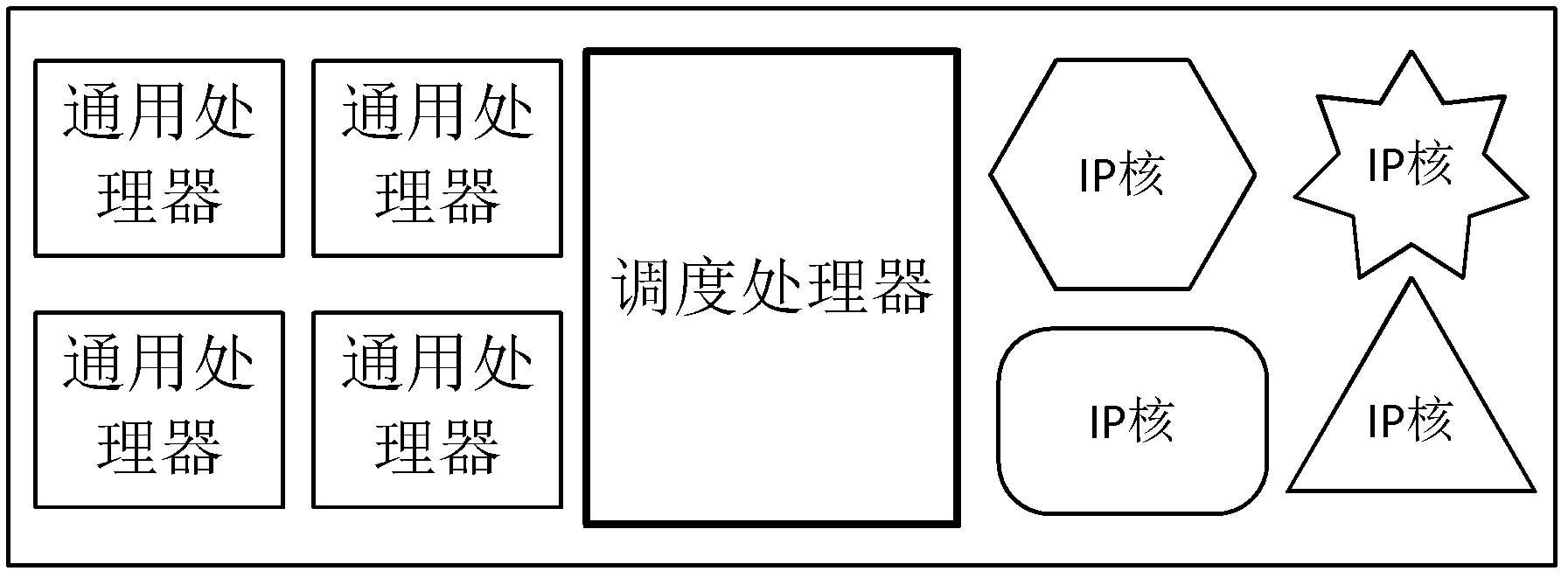
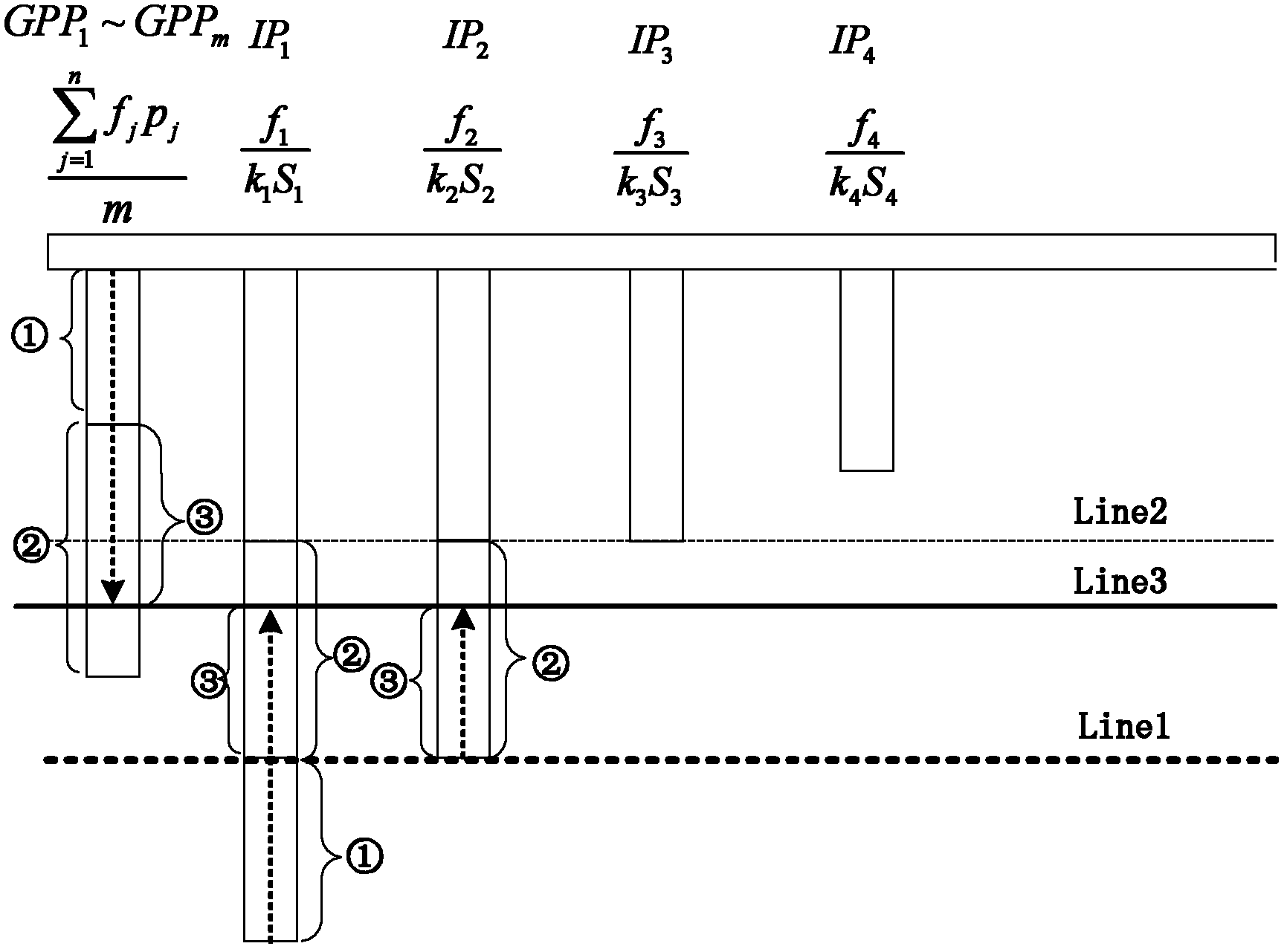
ActiveCN102360313ATake advantage ofImprove acceleration performanceResource allocationArchitecture with multiple processing unitsMulti core computingSingle-core
The invention discloses a performance acceleration method of a heterogeneous multi-core computing platform on chip, wherein the heterogeneous multi-core computing platform on chip comprises a plurality of general processors for executing tasks, a plurality of hardware IP (Internet Protocol) cores and a core scheduling module; and the core scheduling module is responsible for task partitioning and scheduling, so as to allocate the tasks to different computing units for execution. The performance acceleration method is characterized by comprising the following steps of: (1) taking a performance acceleration ratio of a single-core processor as an evaluation index, evaluating the influence of a software and hardware task partitioning scheme of the core scheduling module on the acceleration ratio under the premise of fixed hardware platform, and obtaining the task type, the number of the general processors, the number of hardware acceleration parts and an acceleration ratio parameter of single hardware acceleration part under the optimal performance condition; and (2) reconfiguring the hardware platform according to the task type, the number of the general processors, the number of hardware acceleration parts and the acceleration ratio parameter of single hardware acceleration part under the optimal performance condition. The method can obviously improve the accelerated running efficiency of the system, so that all the resource of the system can be fully used.
Owner:SUZHOU INST FOR ADVANCED STUDY USTC
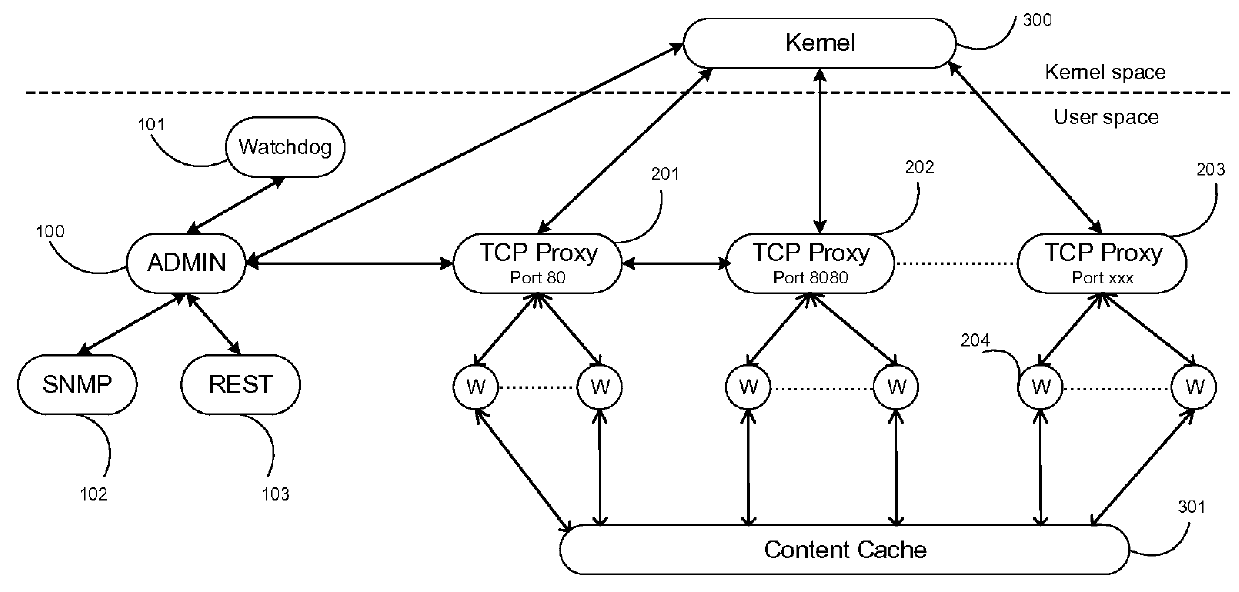
Optimizing stream-mode content cache
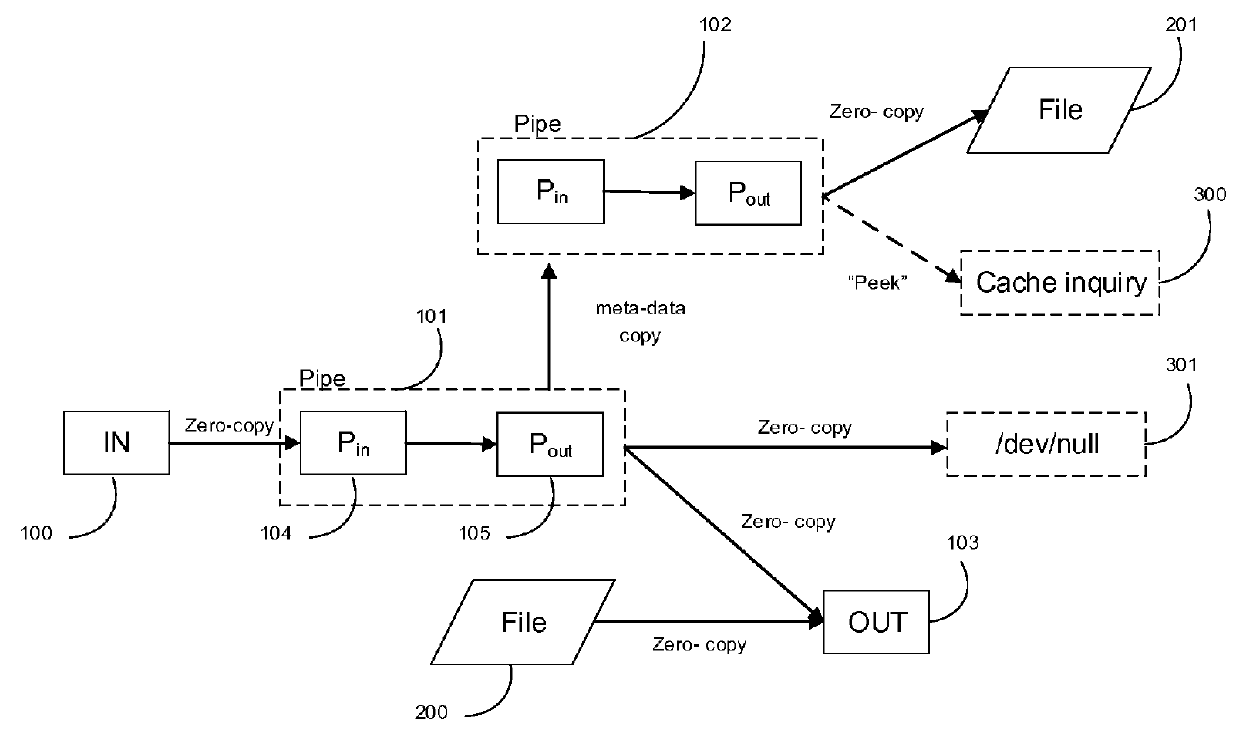
ActiveUS20160036879A1Delay minimizationZero copyingProgram initiation/switchingMultiple digital computer combinationsCache accessMulti core computing
A system and method, called ADD (adaptive data descriptor) and FP (functional proximity), optimizes data transfer between inbound and outbound streams for TCP or other data transfer mechanisms. ADD minimizes data copying in the steam mode, while allowing simultaneous reading from the inbound stream, and writing into the outbound stream from a stored file. ADD and FP jointly minimize total latency in stream-mode data transfer, with or without caching, over multi-core computing systems.FP assigns processes in a multi-core computing system to minimize cache misses and cache access in shared on-chip caches. FP also assigns threads in a TCP splicing box that provides TCP splicing between a plurality of senders and a plurality of receivers, by assigning one thread or 2 threads per TCP connection. The threads are assigned to thread groups so that each thread group is assigned to a single CPU or core. This assignment maximizes cache hits in shared on-chip caches in a multi-core computing system.
Owner:BADU NETWORKS
Parallel universal sequence alignment method running on multi-core computer platform
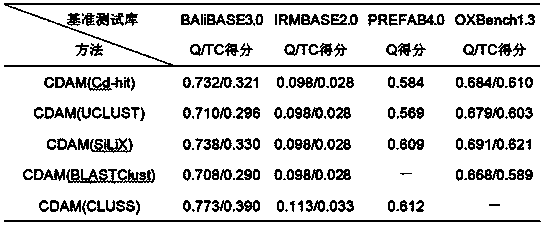
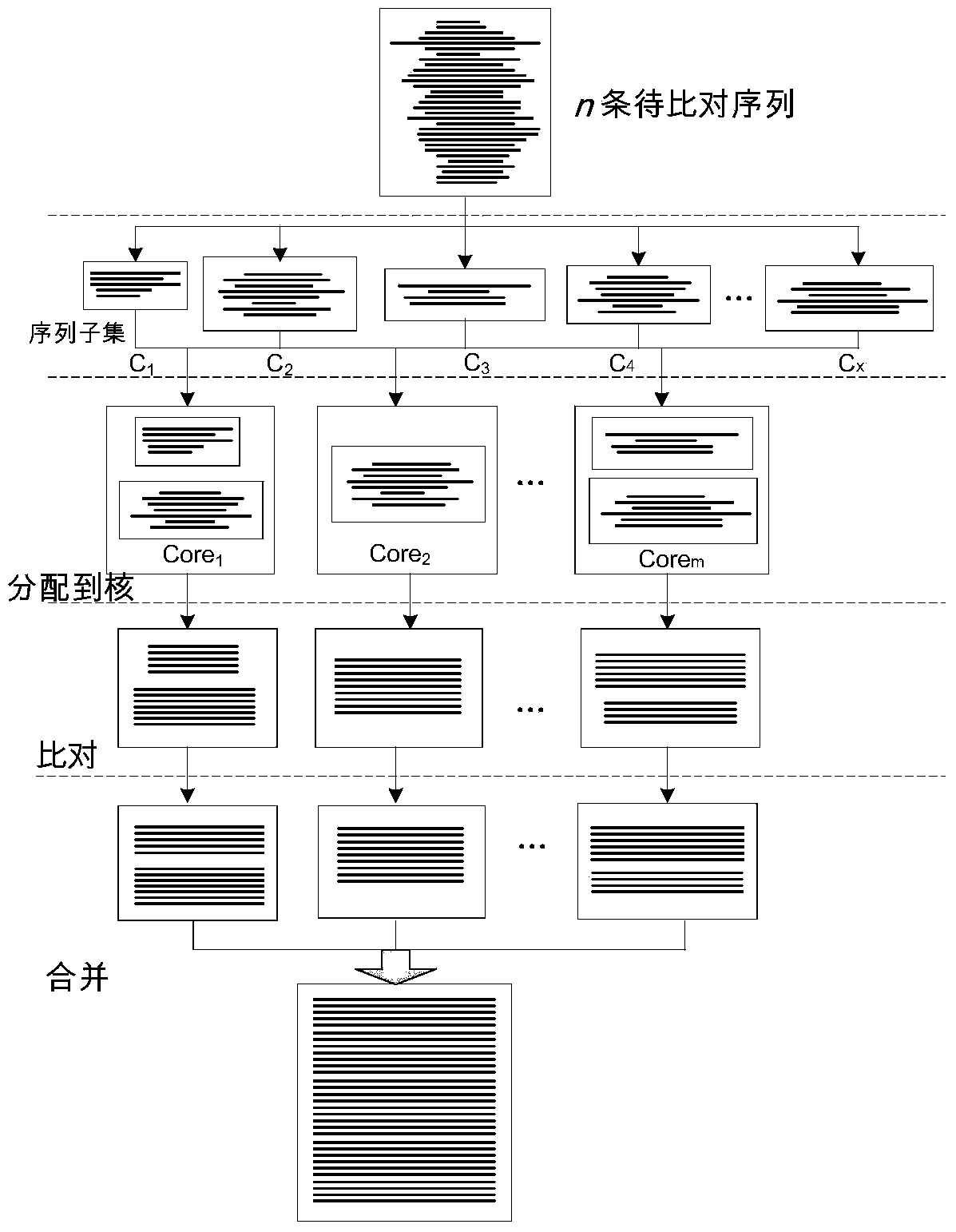
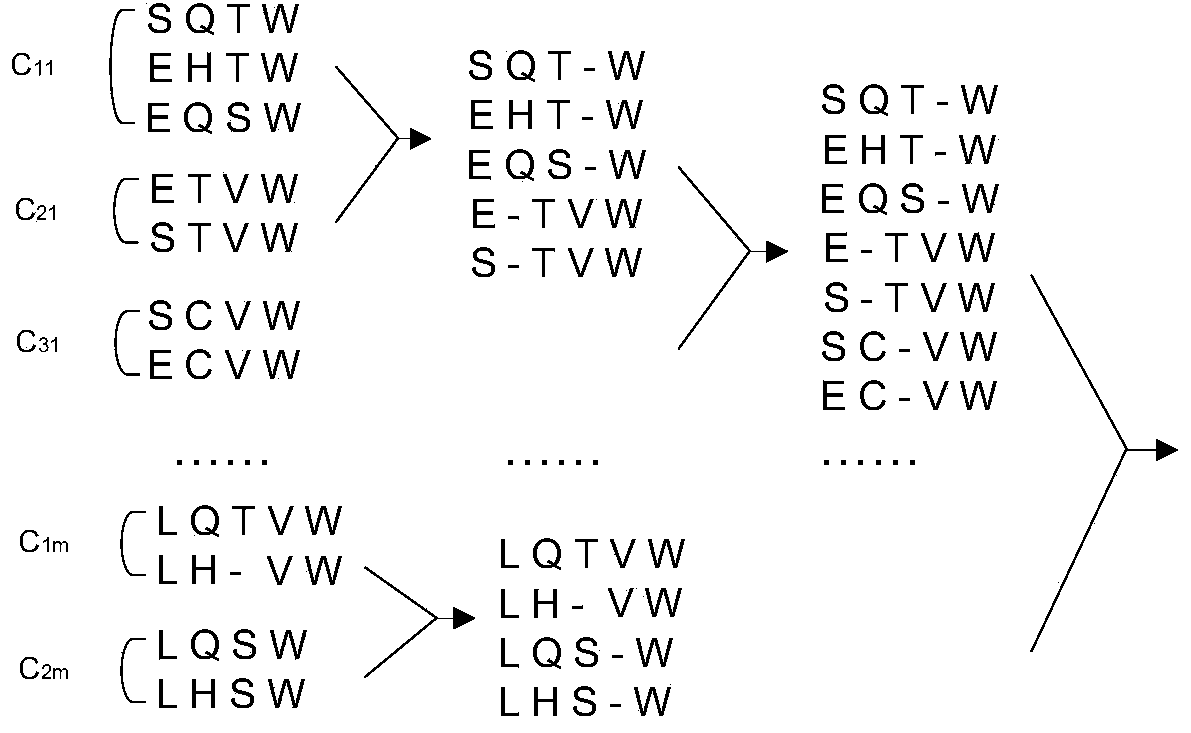
InactiveCN104239732AImprove comparison processing efficiencySpecial data processing applicationsMulti core computingDistribution method
The invention discloses a parallel universal sequence alignment method running on a multi-core computer platform. The parallel universal sequence alignment method comprises the following steps: firstly performing classification on to-be-aligned sequence sets by utilizing a clustering method (Cluster) to obtain subsequence sets (C1, C2, ... , Cm) unequal in size; then, distributing to-be-aligned subsequence sets to all computing cores (Core1, Core2, ... , Coren) by applying a distribution method (Distribute), wherein load balance on each core is taken as the final goal of distribution; subsequently, respectively aligning (Align) all the subsequence sets by applying the traditional sequence alignment method; finally, merging aligned subsequence sets by applying a merging method (Merge) to obtain final alignment results of the to-be-aligned subsequence sets. According to the parallel universal sequence alignment method disclosed by the invention, upon the multi-core computer platform, by fully utilizing data parallel computing strategy, the processing efficiency of biological sequence alignment is obviously improved.
Owner:HUNAN UNIV
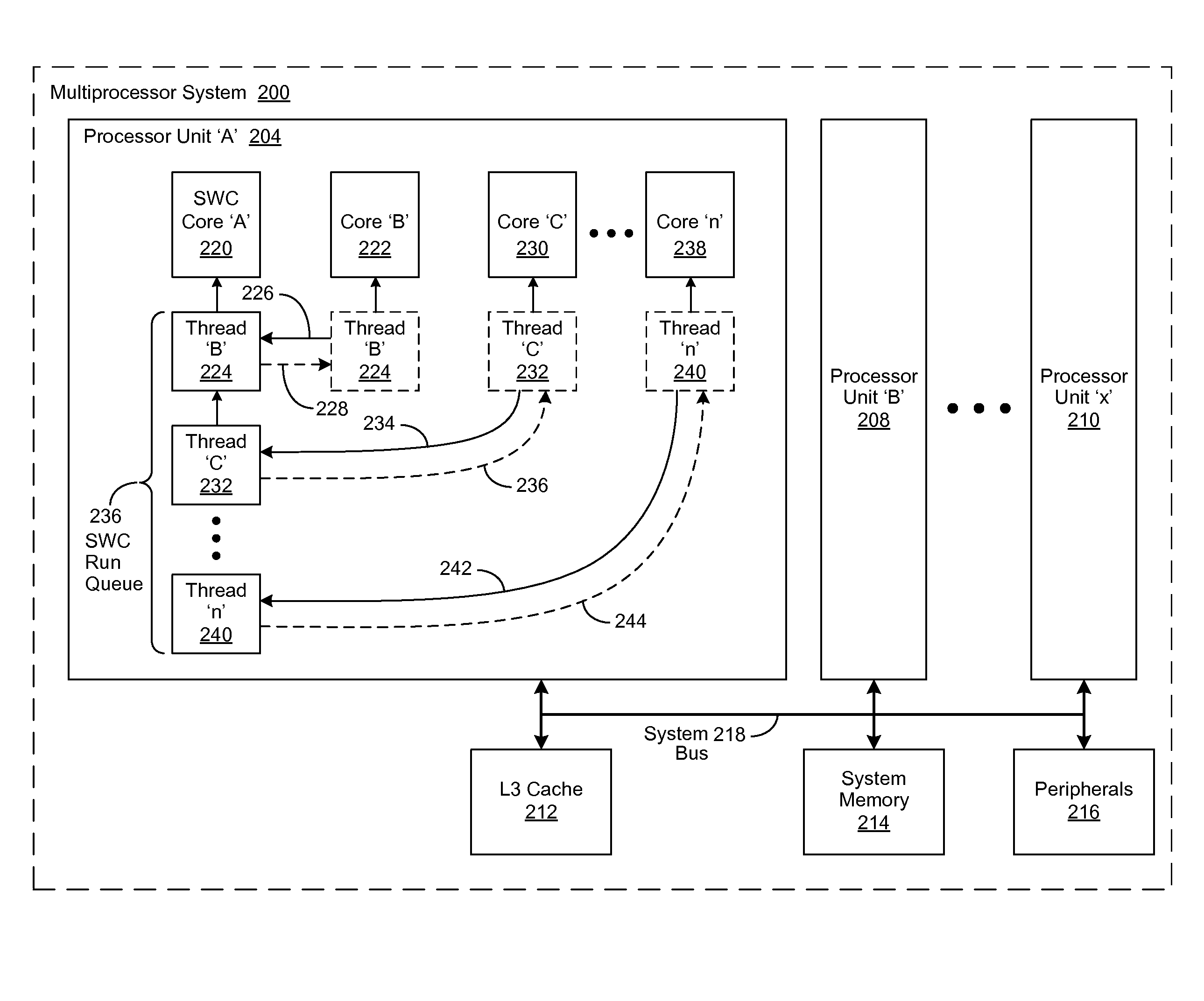
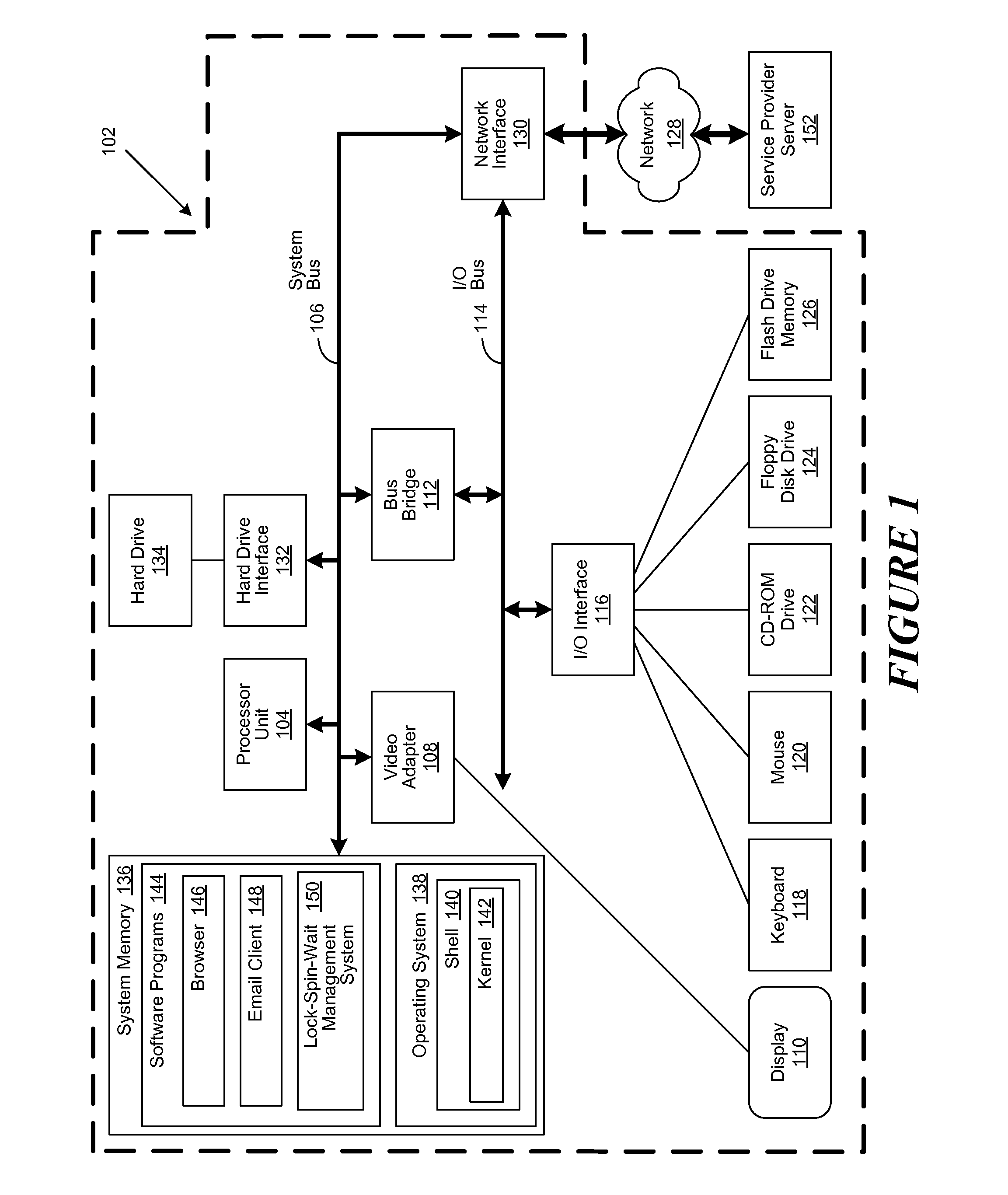
Lock spin wait operation for multi-threaded applications in a multi-core computing environment
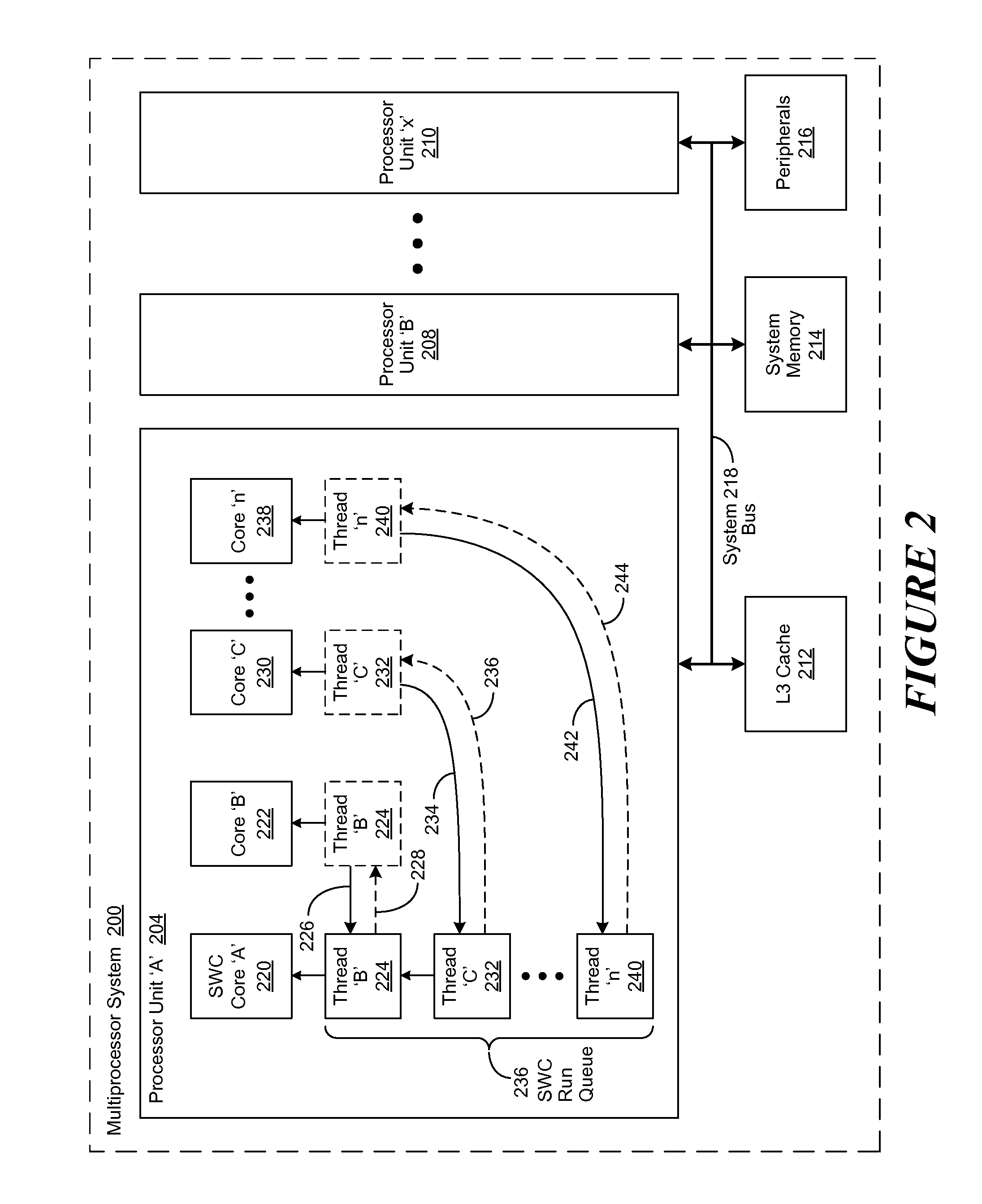
A method, system and computer-usable medium are disclosed for a lock-spin-wait operation for managing multi-threaded applications in a multi-core computing environment. A target processor core, referred to as a “spin-wait core” (SWC), is assigned (or reserved) for primarily running spin-waiting threads. Threads operating in the multi-core computing environment that are identified as spin-waiting are then moved to a run queue associated with the SWC to acquire a lock. The spin-waiting threads are then allocated a lock response time that is less than the default lock response time of the operating system (OS) associated with the SWC. If a spin-waiting fails to acquire a lock within the allocated lock response time, the SWC is relinquished, ceding its availability for other spin-waiting threads in the run queue to acquire a lock. Once a spin-waiting thread acquires a lock, it is migrated to its original, or an available, processor core.
Owner:INT BUSINESS MASCH CORP
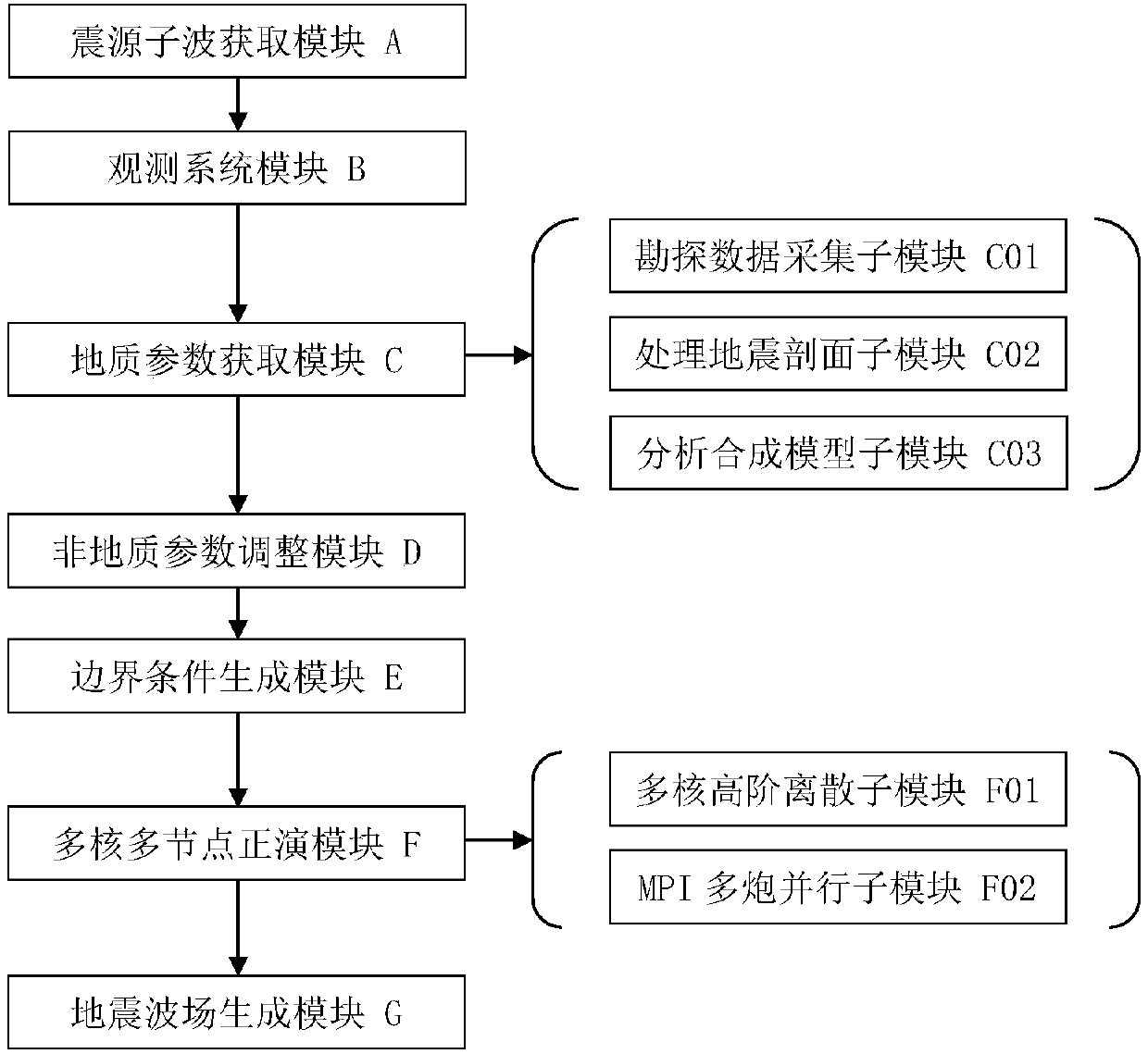
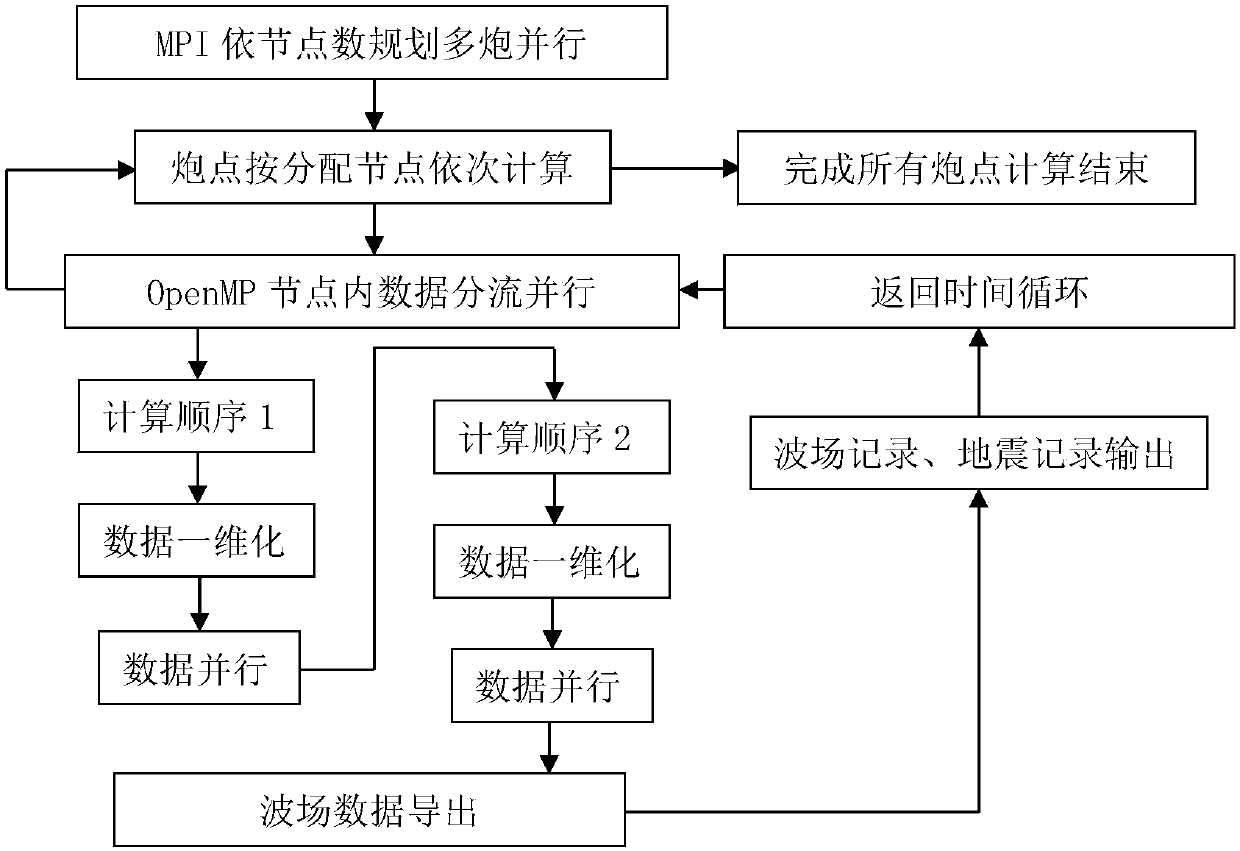
Multi-core multi-node parallel three-dimensional seismic wave field generation method and system
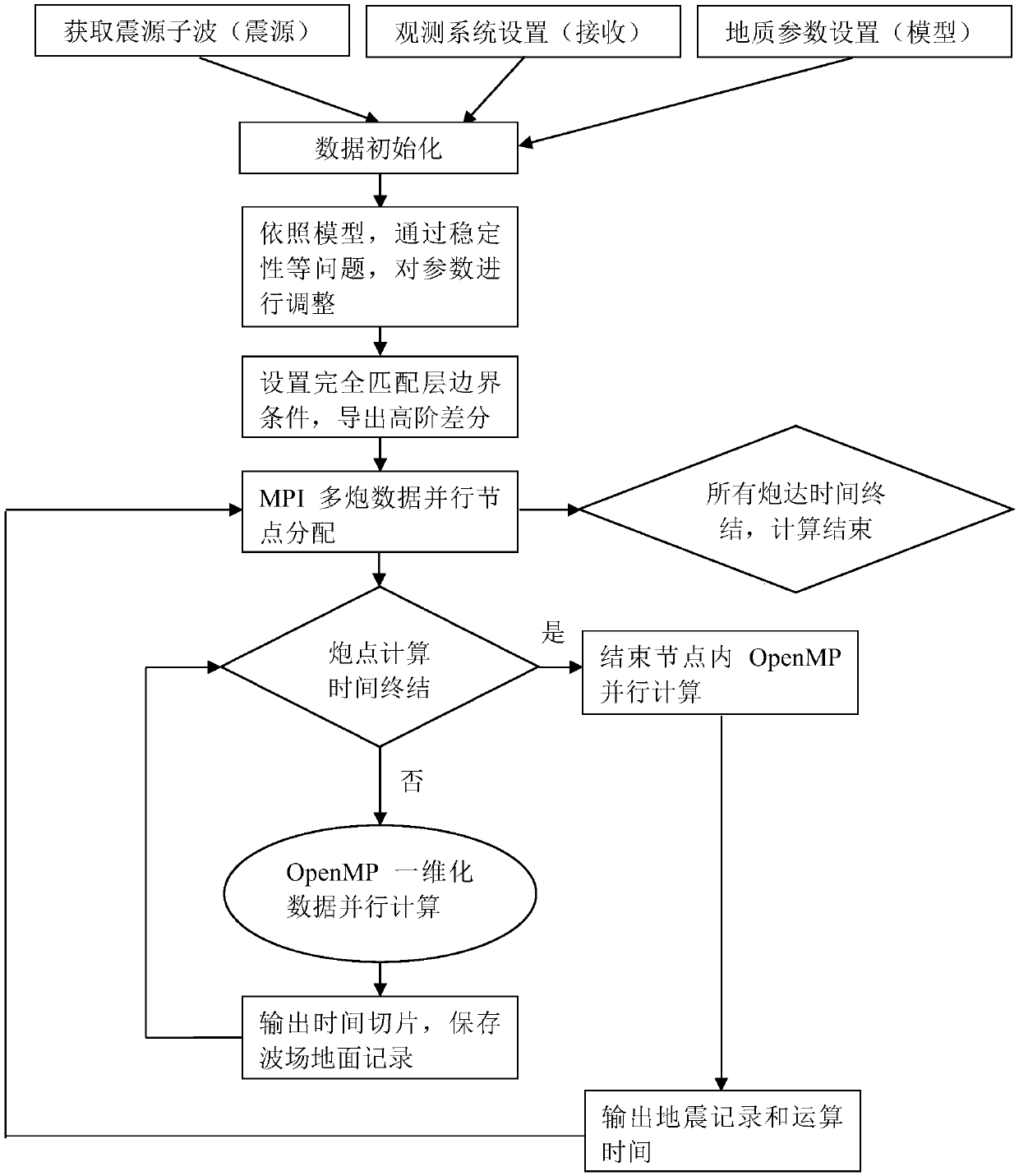
InactiveCN107561585AHigh precisionImprove numerical stabilitySeismic signal processingMulti core computingWave equation
The invention discloses a multi-core multi-node parallel three-dimensional seismic wave field generation method and system. High-precision and high-efficiency solving of a three-dimensional seismic wave equation can be realized based on high-order finite difference and multi-core multi-node parallel acceleration. The method comprises the steps that data information is acquired; the initial condition, the generation boundary condition and the algorithm stability condition of a seismic wave propagation three-dimensional forward simulation model are determined, and high-order finite difference and numerical simulation are performed on the seismic wave propagation equation; a hybrid multi-core multi-node parallel forward simulation algorithm structure based on MPI / OpenMP is designed through combination of the processing capacity of the MPI for multi-node parallel and OpenMP for multi-core computing; and multi-gun data cyclic parallel computing is controlled by the MPI, and the multi-gun wave field slice result and the ground seismic record data are outputted to simulate and generate the exploration seismic wave field. The computing efficiency can be greatly enhanced and the computing accuracy can be effectively enhanced so that the requirements of three-dimensional seismic wave field generation for the computing efficiency and accuracy can be met.
Owner:PEKING UNIV
Lock Spin Wait Operation for Multi-Threaded Applications in a Multi-Core Computing Environment
InactiveUS20150254113A1Response requirements are lowLow benefitProgram synchronisationMemory systemsMulti core computingOperational system
A method, system and computer-usable medium are disclosed for a lock-spin-wait operation for managing multi-threaded applications in a multi-core computing environment. A target processor core, referred to as a “spin-wait core” (SAC), is assigned (or reserved) for primarily running spin-waiting threads. Threads operating in the multi-core computing environment that are identified as spin-waiting are then moved to a run queue associated with the SAC to acquire a lock. The spin-waiting threads are then allocated a lock response time that is less than the default lock response time of the operating system (OS) associated with the SAC. If a spin-waiting fails to acquire a lock within the allocated lock response time, the SAC is relinquished, ceding its availability for other spin-waiting threads in the run queue to acquire a lock. Once a spin-waiting thread acquires a lock, it is migrated to its original, or an available, processor core.
Owner:IBM CORP
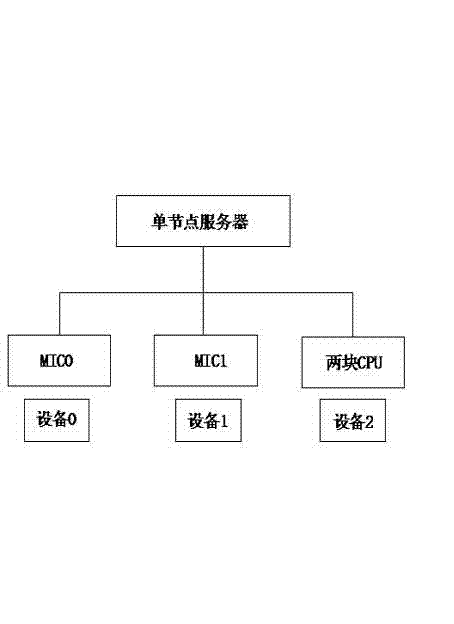
High-efficiency system based on central processing unit (CPU)/many integrated core (MIC) heterogeneous system structure
InactiveCN103049329ALower build costsReduce managementResource allocationConcurrent instruction executionMulti core computingPerformance computing
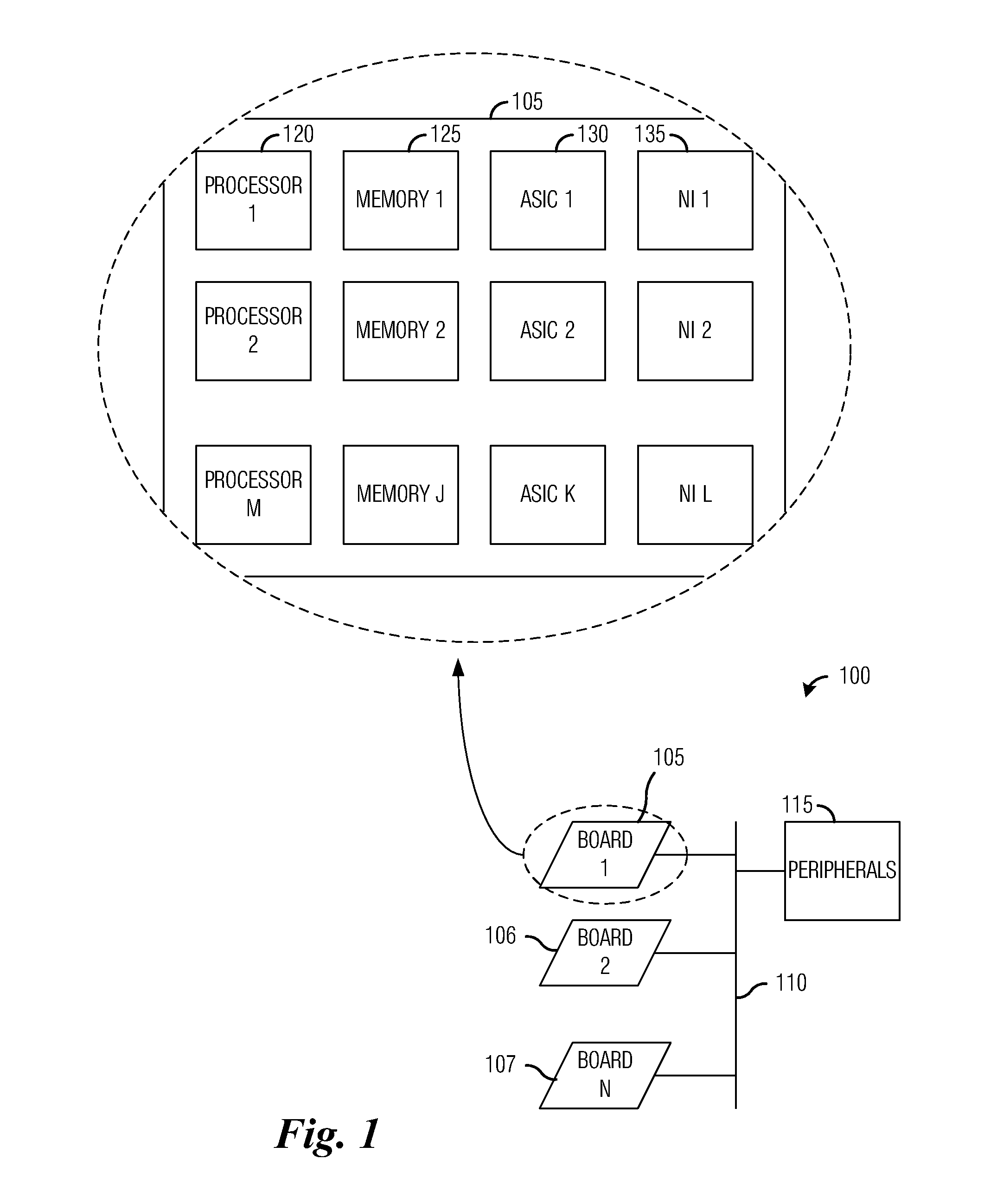
The invention provides a high-efficiency system based on a CPU / MIC heterogeneous system structure, and relates to the field of high-performance computing of computers. The whole system design comprises hardware portion design, system environment configuration and software portion design. The system achieves software and hardware integrated design, the CPU / MIC heterogeneous system structure is utilized, the system integrates the multi-core computing capacity of the CPU platform with the many-core computing capacity of the MIC, wherein the CPU participates in logic computing and intensive core computing, the MIC only participates in intensive core computing, and the performance is maximized through common computing of the CPU and the MIC. The high-efficiency system has the advantages that the system solves the problems of the performance bottleneck and power consumption of high-performance computing applications through the cooperative computing of the CPU and the MIC and has the advantages of being high in performance and low in power consumption, and the computer room construction cost and management, operation and maintenance costs are reduced.
Owner:LANGCHAO ELECTRONIC INFORMATION IND CO LTD
Mass data parallel processing method based on big data
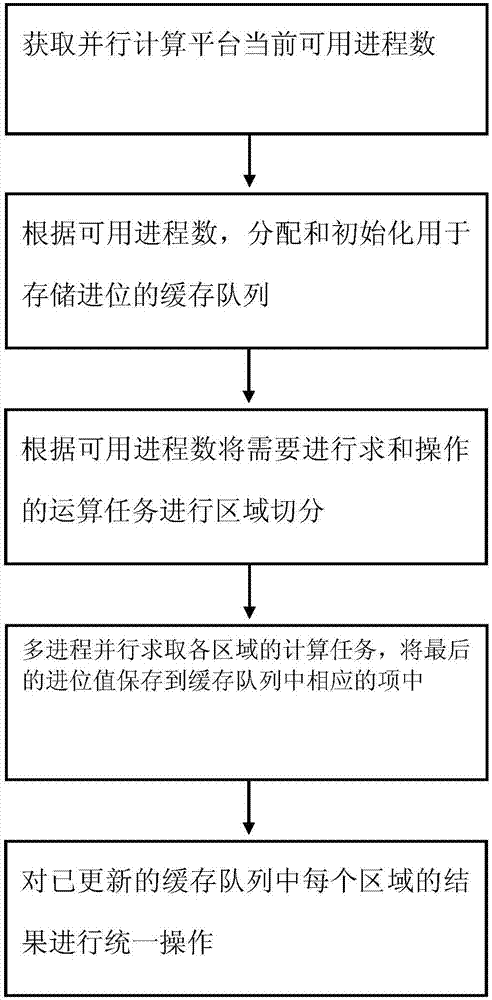
InactiveCN107977444AFast operationSpecial data processing applicationsMulti core computingMultiple modes
The invention provides a mass data parallel processing method based on big data. The method comprises the steps that (1) a current available process number of a parallel computing platform is acquired; (2) a cache queue used for storing carry is allocated and initialized according to the available process number; (3) region segmentation is performed on operation tasks needing summation operation according to the available process number; (4) computing tasks in all regions are solved parallelly in multiple processes, and a last carry value is saved into an corresponding item in the cache queue;and (5) unified operation is performed on a result of each region in the updated cache queue. Through the mass data parallel processing method based on the big data, on the basis of the multi-core computing platform, a distributed parallel environment is fully utilized to increase operation speed.
Owner:成都博睿德科技有限公司
System and method for massively multi-core computing systems
ActiveUS8516493B2Reduce power consumptionAdjust power consumptionEnergy efficient ICTDigital computer detailsResource poolMulti core computing
A system and method for massively multi-core computing are provided. A method for computer management includes determining if there is a need to allocate at least one first resource to a first plane. If there is a need to allocate at least one first resource, the at least one first resource is selected from a resource pool based on a set of rules and allocated to the first plane. If there is not a need to allocate at least one first resource, it is determined if there is a need to de-allocate at least one second resource from a second plane. If there is a need to de-allocate at least one second resource, the at least one second resource is de-allocated. The first plane includes a control plane and / or a data plane and the second plane includes the control plane and / or the data plane. The resources are unchanged if there is not a need to allocate at least one first resource and if there is not a need to de-allocate at least one second resource.
Owner:FUTUREWEI TECH INC
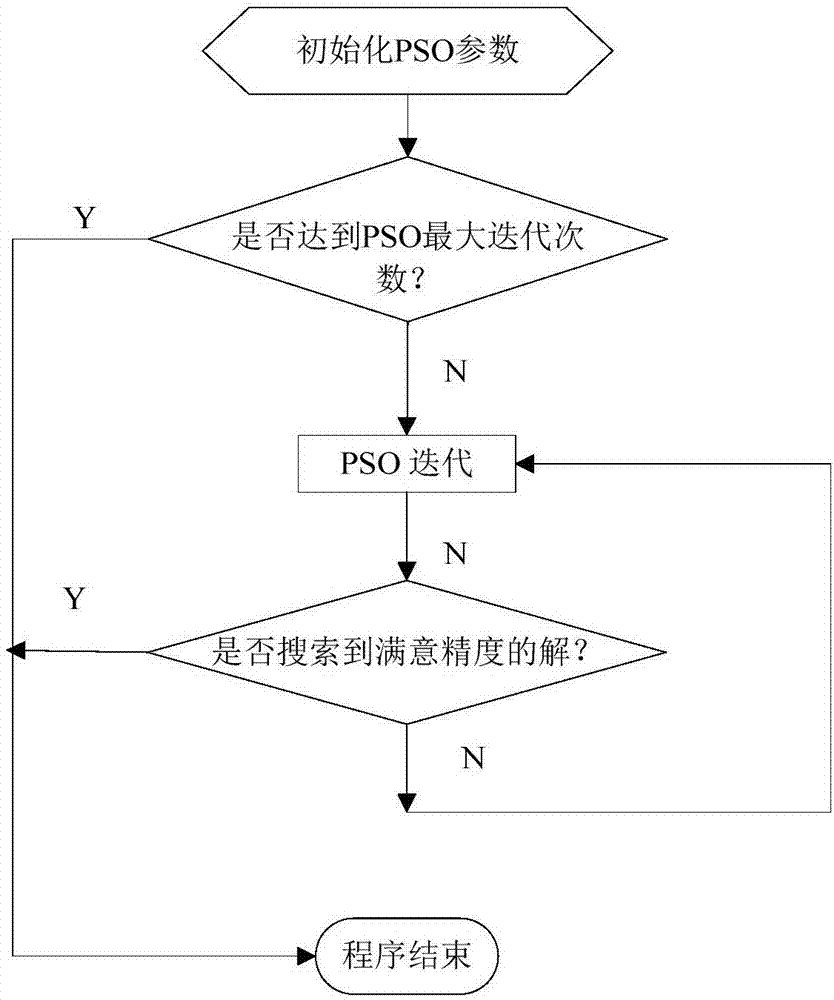
Particle swarm optimization algorithm, multi-computer parallel processing method and system
InactiveCN106951957AExpand coverageSolve the problem that the optimization algorithm is easy to fall into local optimumBiological modelsMultiple digital computer combinationsLocal optimumMulti core computing
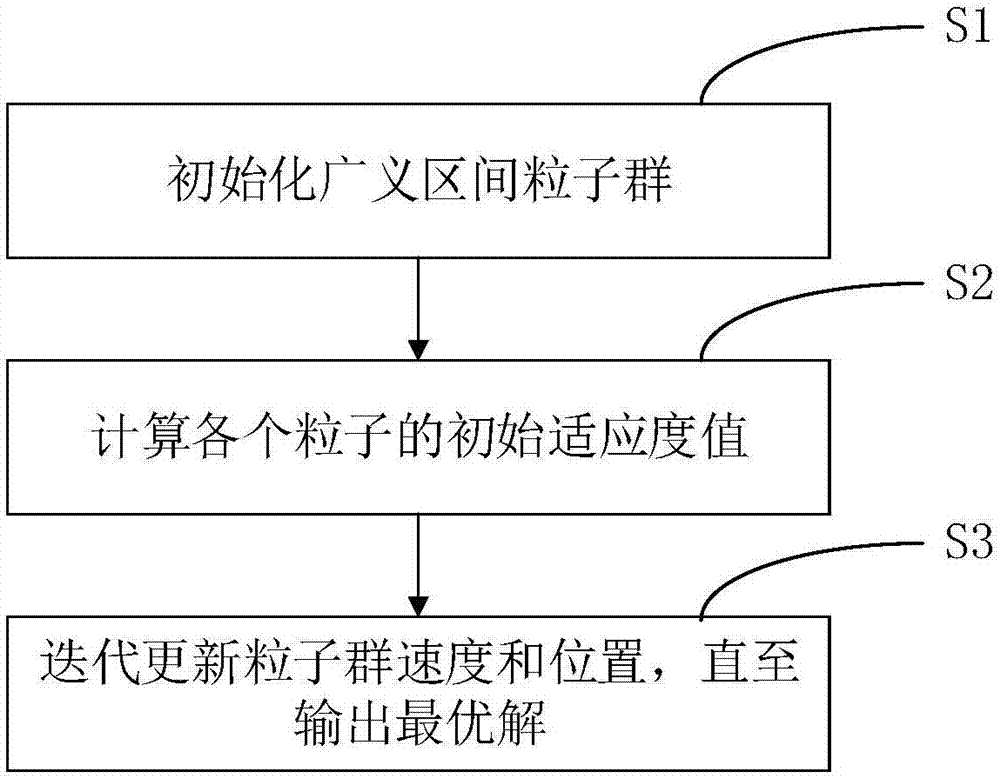
The invention relates to a particle swarm optimization algorithm, a multi-computer parallel processing method and a system. The algorithm comprises steps of step S1, initializing a particle swarm in a generalized section; S2, calculating an initial fitness value of each particle; and step S3, updating speed and positions of the particle swarm in an iteration manner until the overall optimal solution is output. According to the invention, in a way of replacing precise particles through the generalized section, the coverage range of the particle swarm is greatly expanded; a problem that the initial value of the precise particles easily allow the optimization algorithm to suffer from local optimum is effectively solved; and the parallel processing ability of the current multi-core computer can be combined, so precision and efficiency of calculation results are remarkably improved.
Owner:CHANGZHOU COLLEGE OF INFORMATION TECH
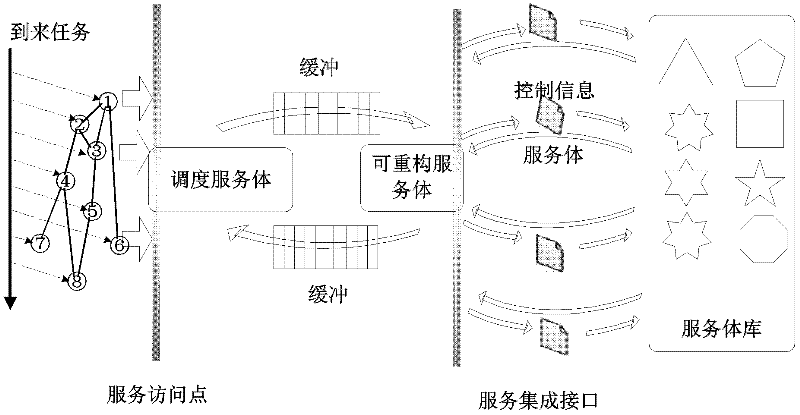
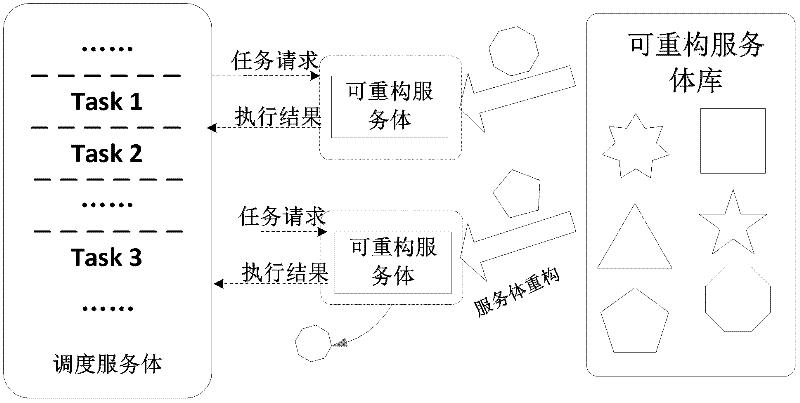
Service-oriented multi-core computing platform on reconfigurable chip and reconfiguration method thereof
InactiveCN102508711AReduce overheadImprove parallelismProgram initiation/switchingMulti core computingService-orientation
The invention discloses a service-oriented multi-core computing platform on a reconfigurable chip and a reconfiguration method thereof. The service-oriented multi-core computing platform is characterized by comprising a scheduling servant, a reconfigurable servant, an interconnection module based on first-in first-out buffer and a servant library containing a plurality of servants, wherein the scheduling servant is interconnected with the reconfigurable servant by virtue of the interconnection module; and the reconfigurable servant is connected with the servant library through an integrated port. According to the platform system provided by the invention, a plurality of types of computing services can be rapidly provided and the reconfiguration time of the reconfigurable servant is greatly reduced.
Owner:SUZHOU INST FOR ADVANCED STUDY USTC
Neural network operation optimization and data processing method and device and storage medium
ActiveCN110187965AImprove computing efficiencyResource allocationNeural architecturesMulti core computingComputation process
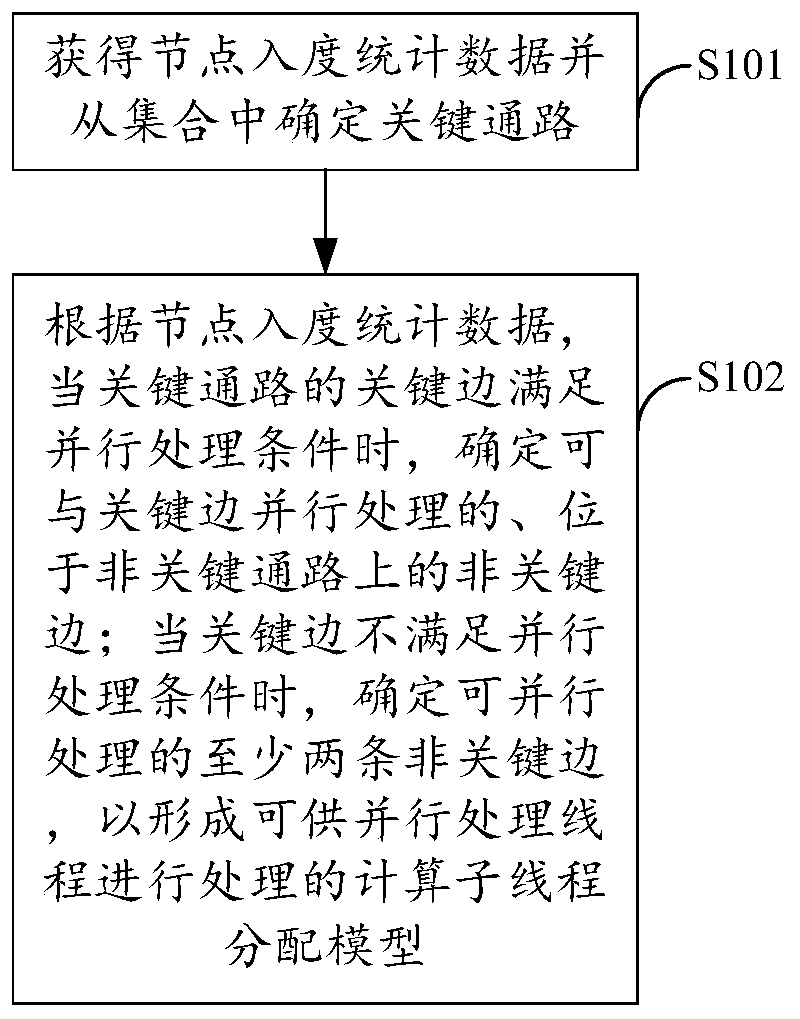
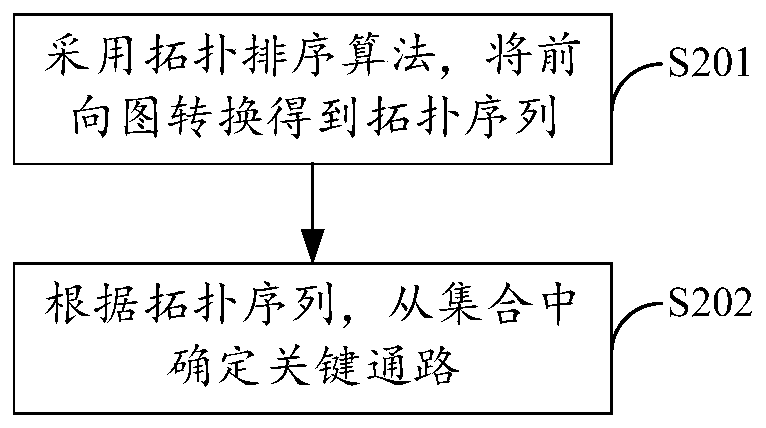
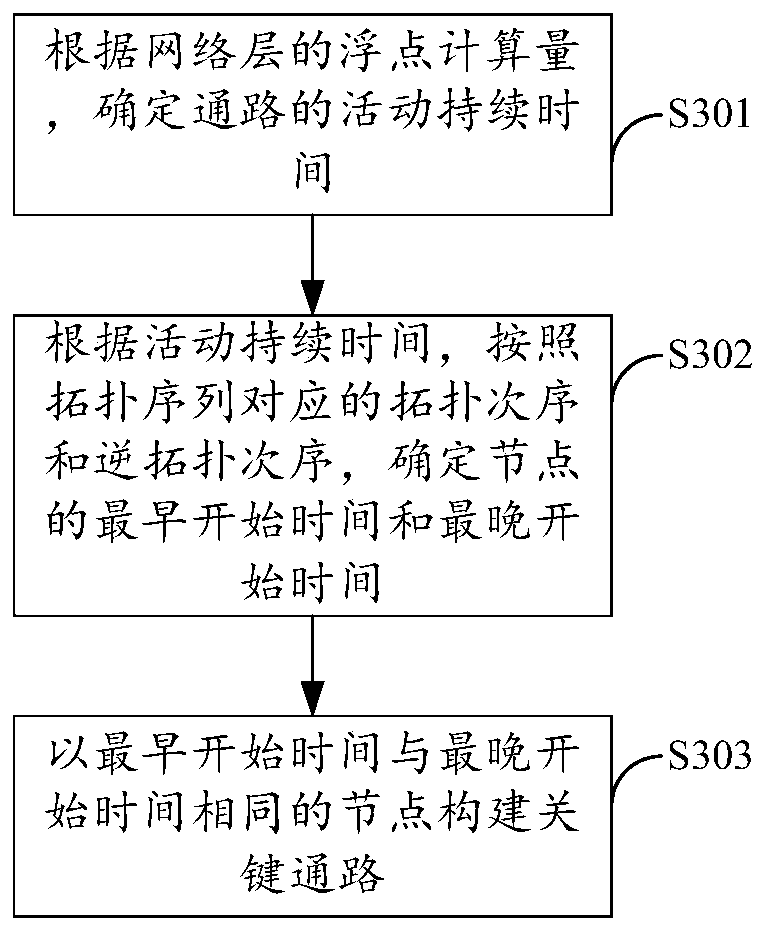
The method is suitable for the technical field of computers, and provides a neural network operation optimization and data processing method and device and a storage medium. The method comprises the steps of during a forward calculation process of a neural network, obtaining the node in-degree statistical data and determining a key path from a path set; if the key edges of the key paths meet the parallel processing conditions, determining the non-key edges which can be processed with the key edges in parallel and are located on the non-key paths, and if the key edges do not meet the parallel processing conditions, determining at least two non-key edges which can be processed in parallel to form a calculation sub-thread distribution model which can be processed by the parallel processing threads. Therefore, the multi-core parallel accelerated optimization on the hierarchical structure of the neural network can be realized, the computing efficiency of the neural network is effectively improved, and the popularization and application of the large-scale neural retention on the computing equipment using the multi-core computing resources are facilitated.
Owner:SHENZHEN UNIV
Method to efficiently implement synchronization using software managed address translation
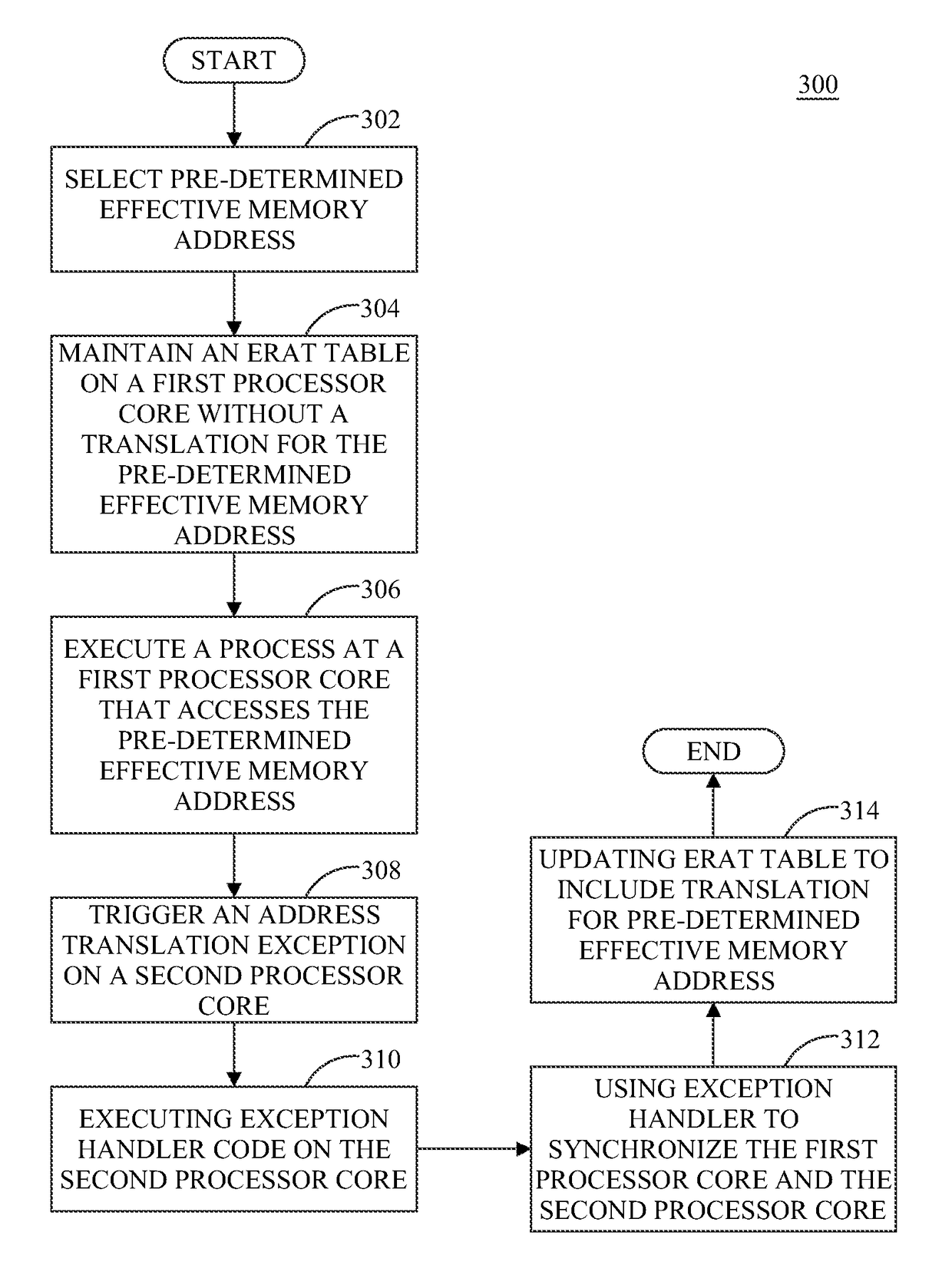
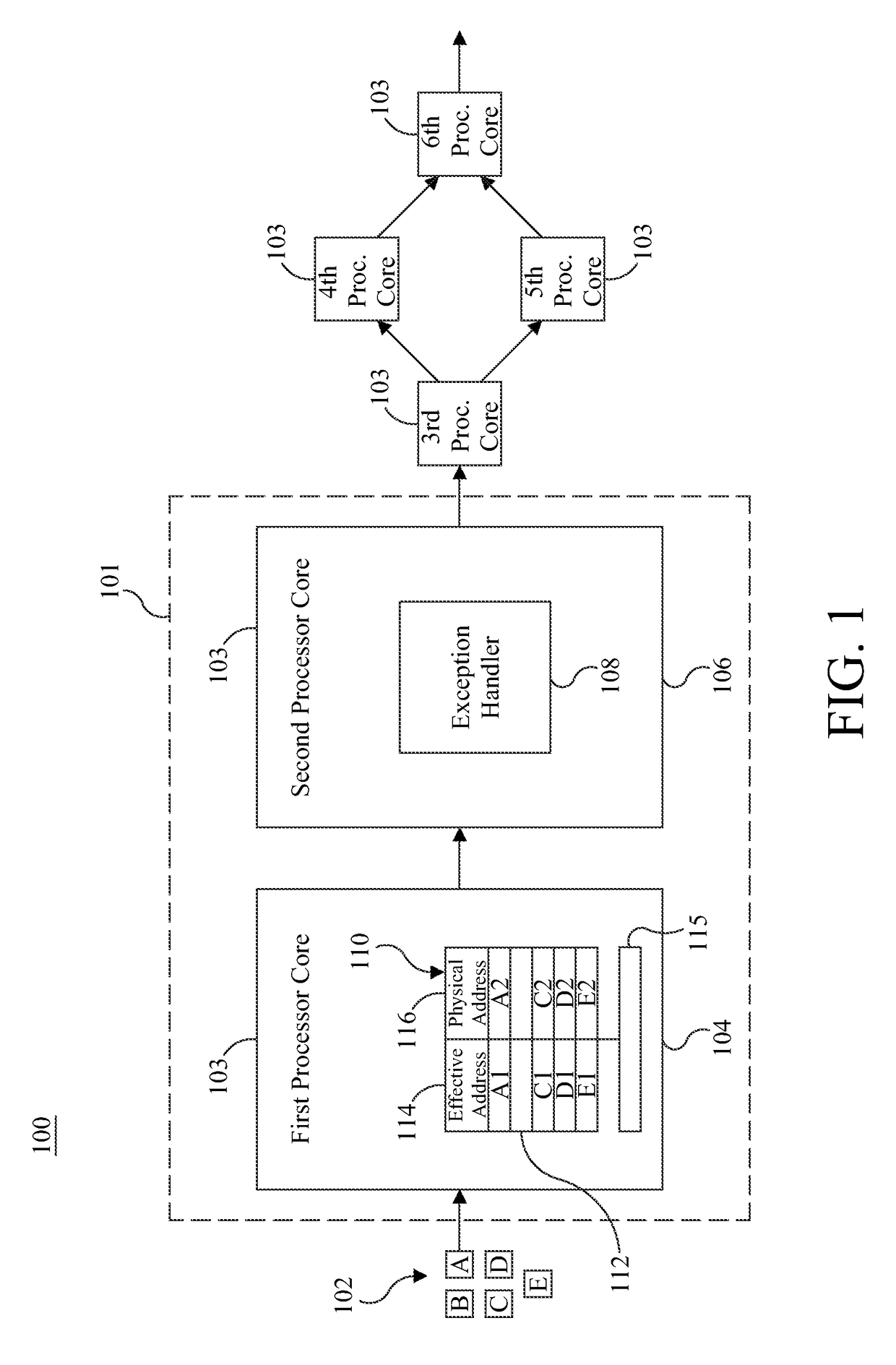
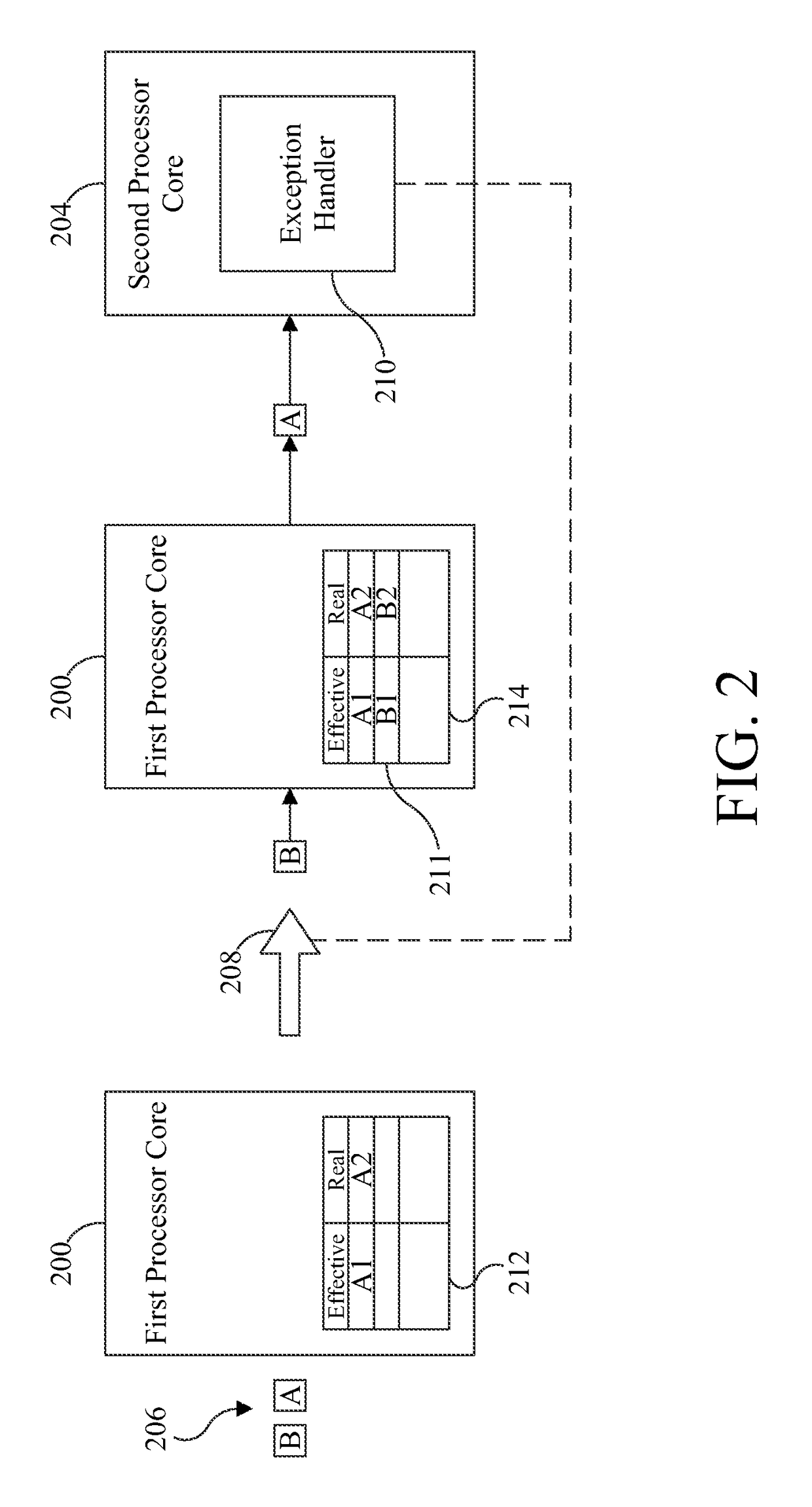
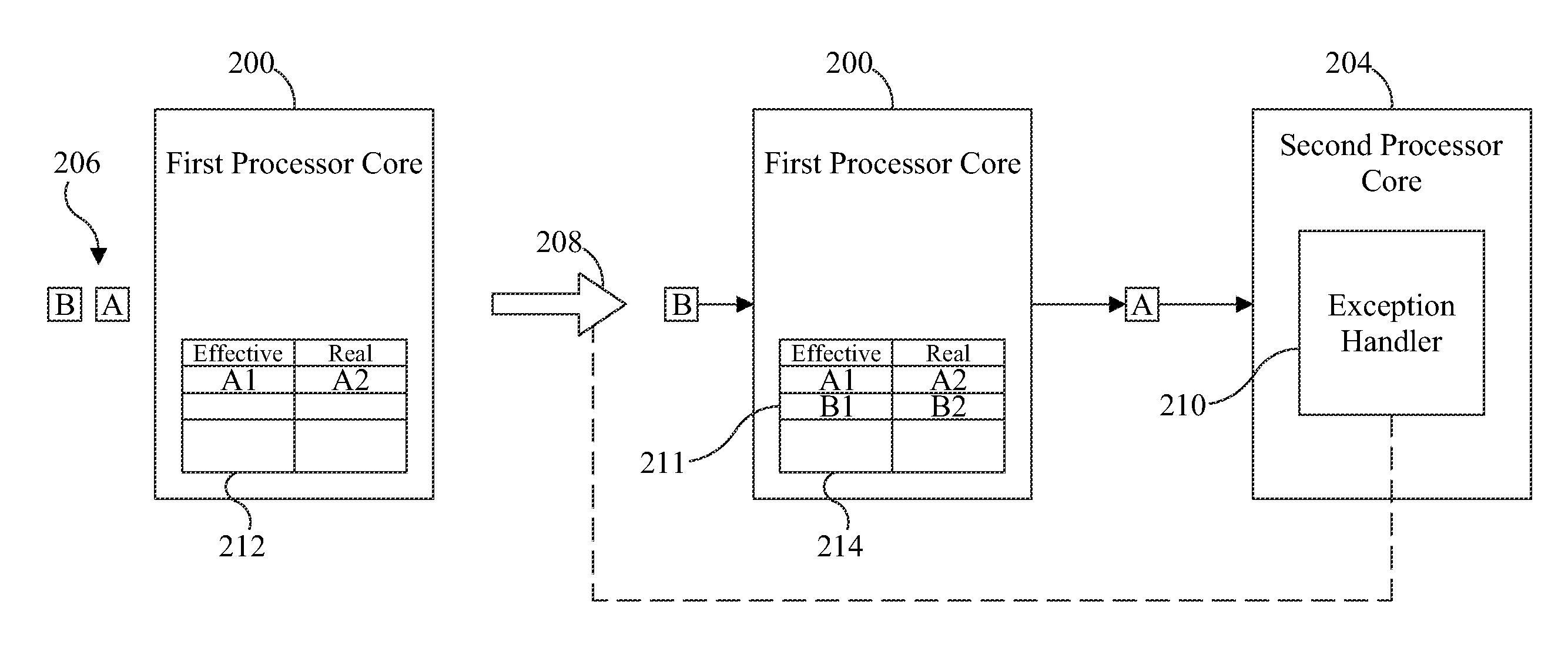
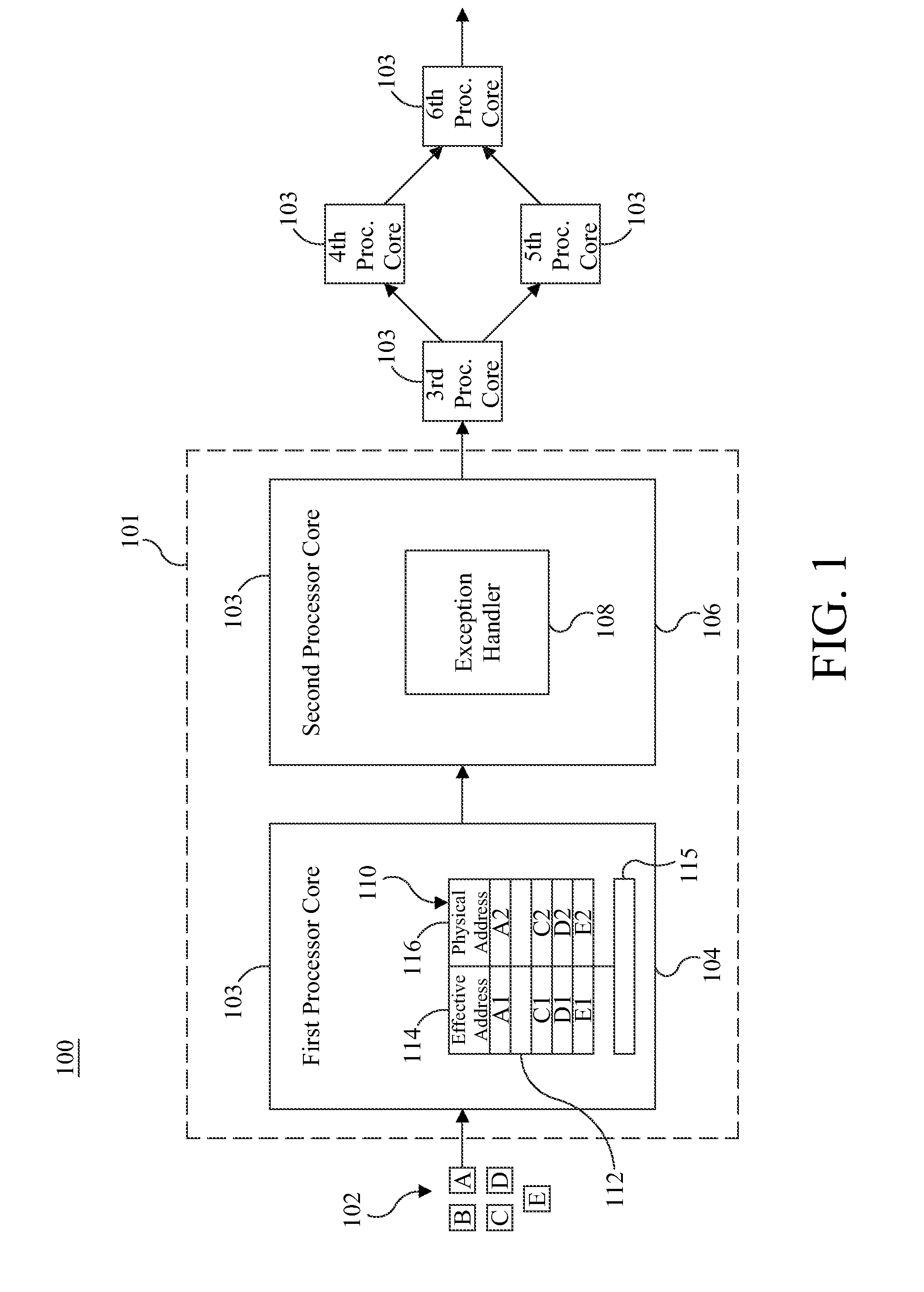
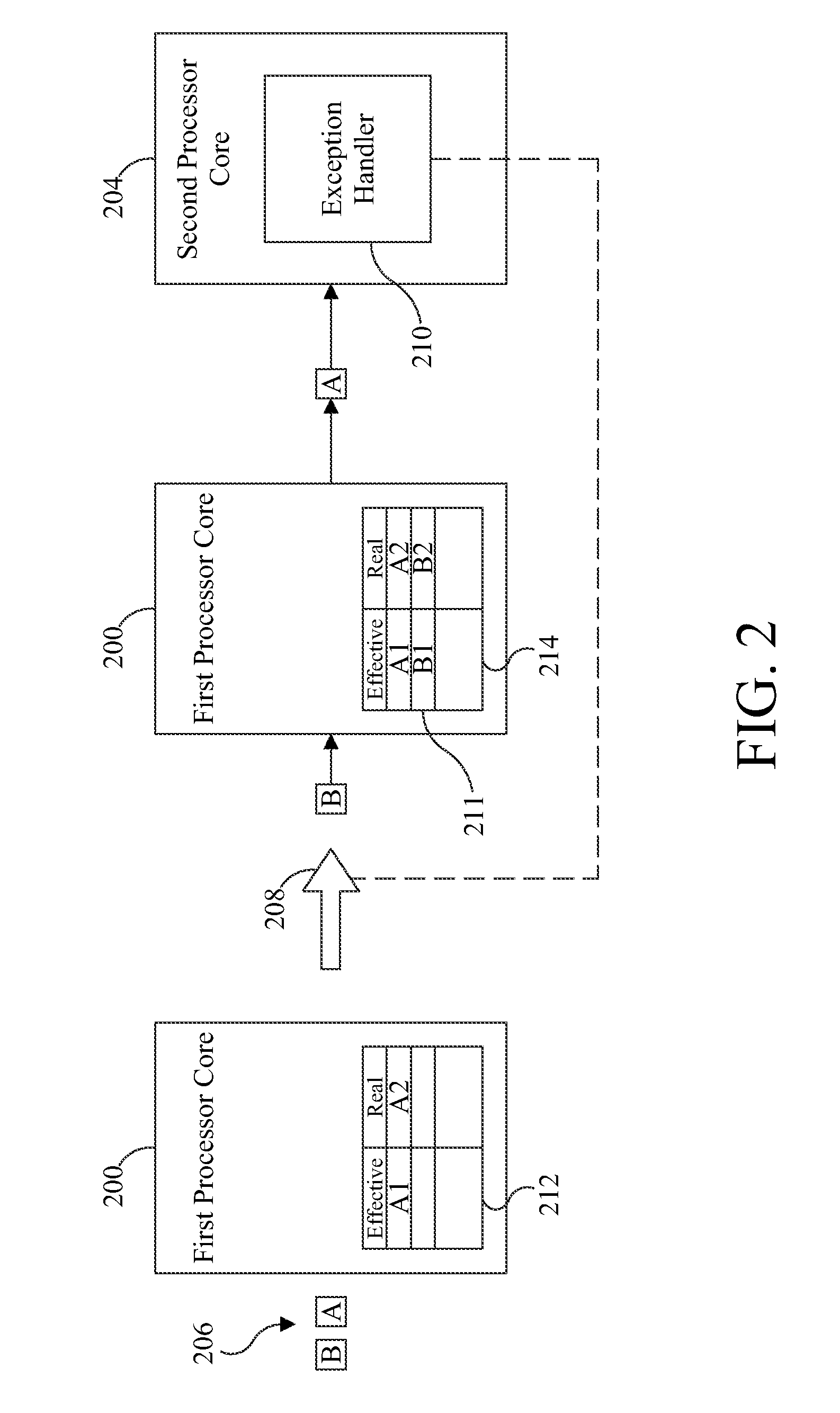
ActiveUS9658940B2Minimal impactReduce overheadMemory architecture accessing/allocationMemory adressing/allocation/relocationMemory addressMulti core computing
Software-managed resources are used to utilize effective-to-real memory address translation for synchronization among processes executing on processor cores in a multi-core computing system. A failure to find a pre-determined effective memory address translation in an effective-to-real memory address translation table on a first processor core triggers an address translation exception in a second processor core and causes an exception handler on the second processor core to start a new process, thereby acting as a means to achieve synchronization among processes on the first processor core and the second processor core. The specific functionality is implemented in the exception handler, which is tailored to respond to the exception based on the address that generated it.
Owner:INT BUSINESS MASCH CORP
Method to efficiently implement synchronization using software managed address translation
ActiveUS20160274996A1Reduce overheadReducing performance degradationMemory architecture accessing/allocationMemory adressing/allocation/relocationMemory addressMulti core computing
Software-managed resources are used to utilize effective-to-real memory address translation for synchronization among processes executing on processor cores in a multi-core computing system. A failure to find a pre-determined effective memory address translation in an effective-to-real memory address translation table on a first processor core triggers an address translation exception in a second processor core and causes an exception handler on the second processor core to start a new process, thereby acting as a means to achieve synchronization among processes on the first processor core and the second processor core. The specific functionality is implemented in the exception handler, which is tailored to respond to the exception based on the address that generated it.
Owner:IBM CORP
Efficient detection method for massive malicious codes
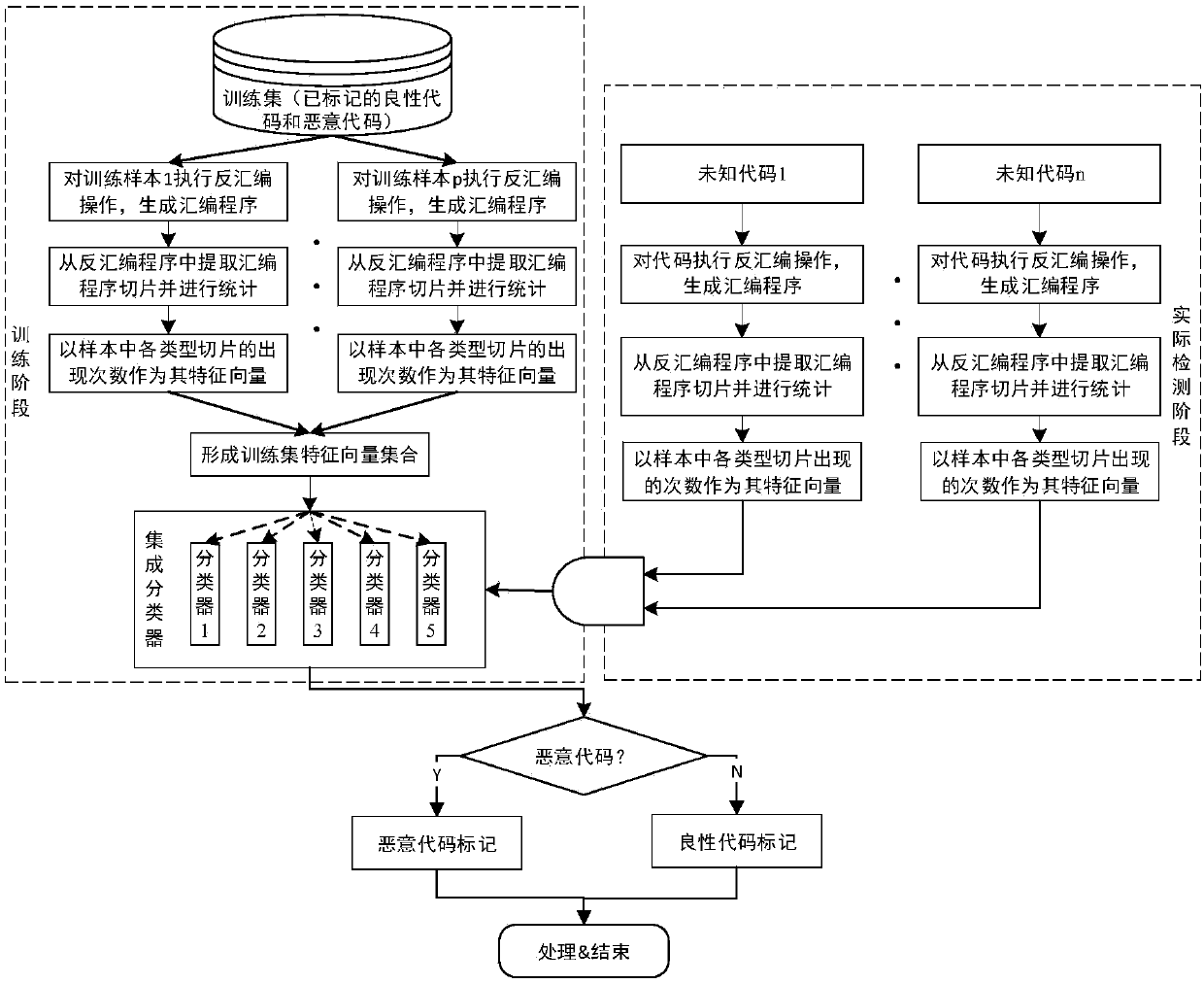
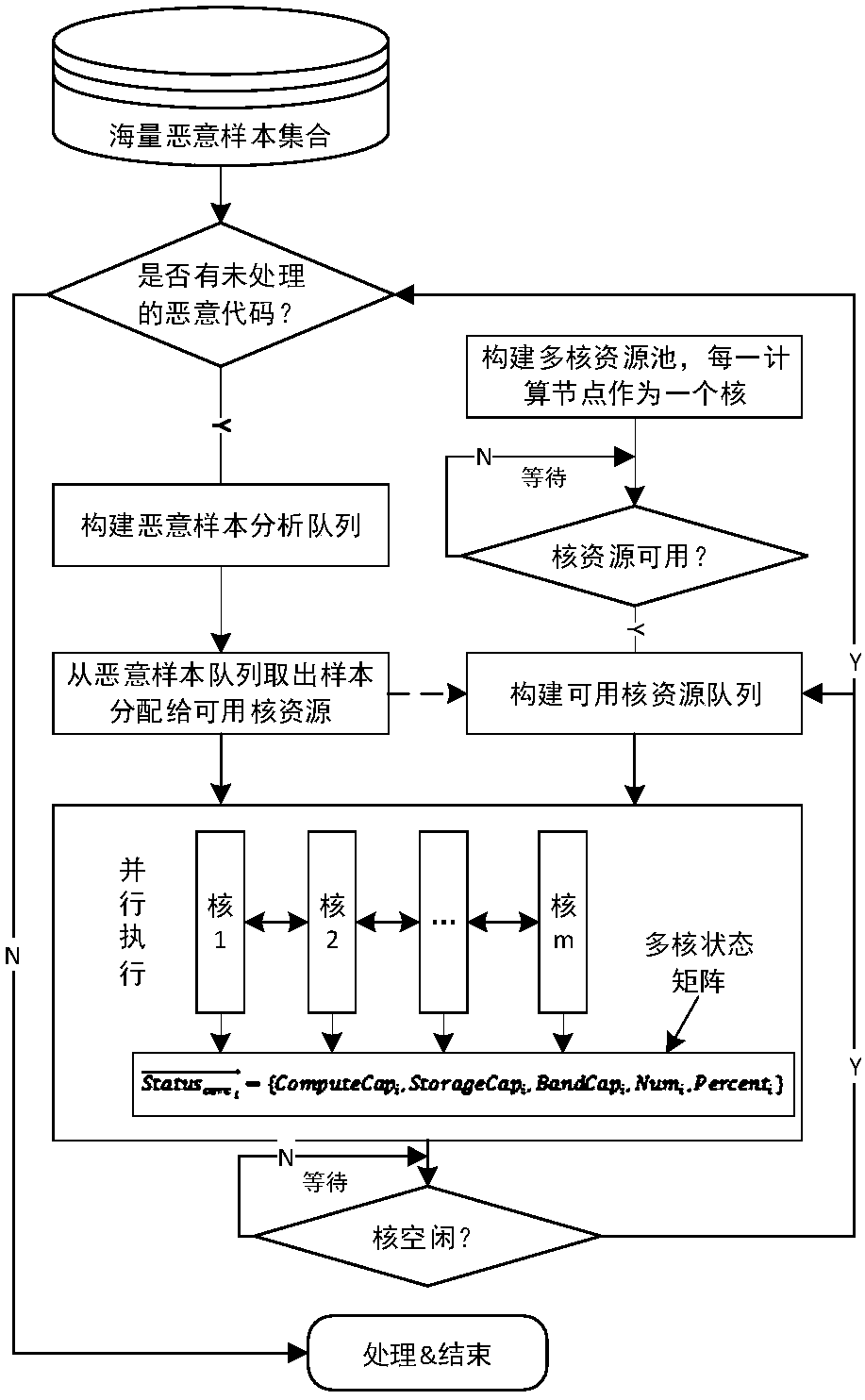
ActiveCN108021810AShorten the timeImprove detection efficiencyPlatform integrity maintainanceFeature vectorProgramming language
The invention discloses an efficient detection method for massive malicious codes, can realize efficient detection of massive malicious codes. According to assembler samples, malicious code detectionis conducted, the method adopts multi-core computing resources to execute steps of sample identification of assembler in parallel, the steps of sample identification of the assembler are as follows: assembler slices are extracted, the assembler slice is a statement or an expression that affects a specified variable in a assembler sample. Based on the preset assembler slice types, the number of occurrences of each type of assembler slice extracted from the assembler sample is counted, and is taken as a feature vector of the assembler sample. According to the feature vectors of the assembler samples, a classifier is obtained by pre-training, the malicious codes are identified by adopting the classifier.
Owner:BEIJING INSTITUTE OF TECHNOLOGYGY +1
Transformational architecture for multi-layer systems
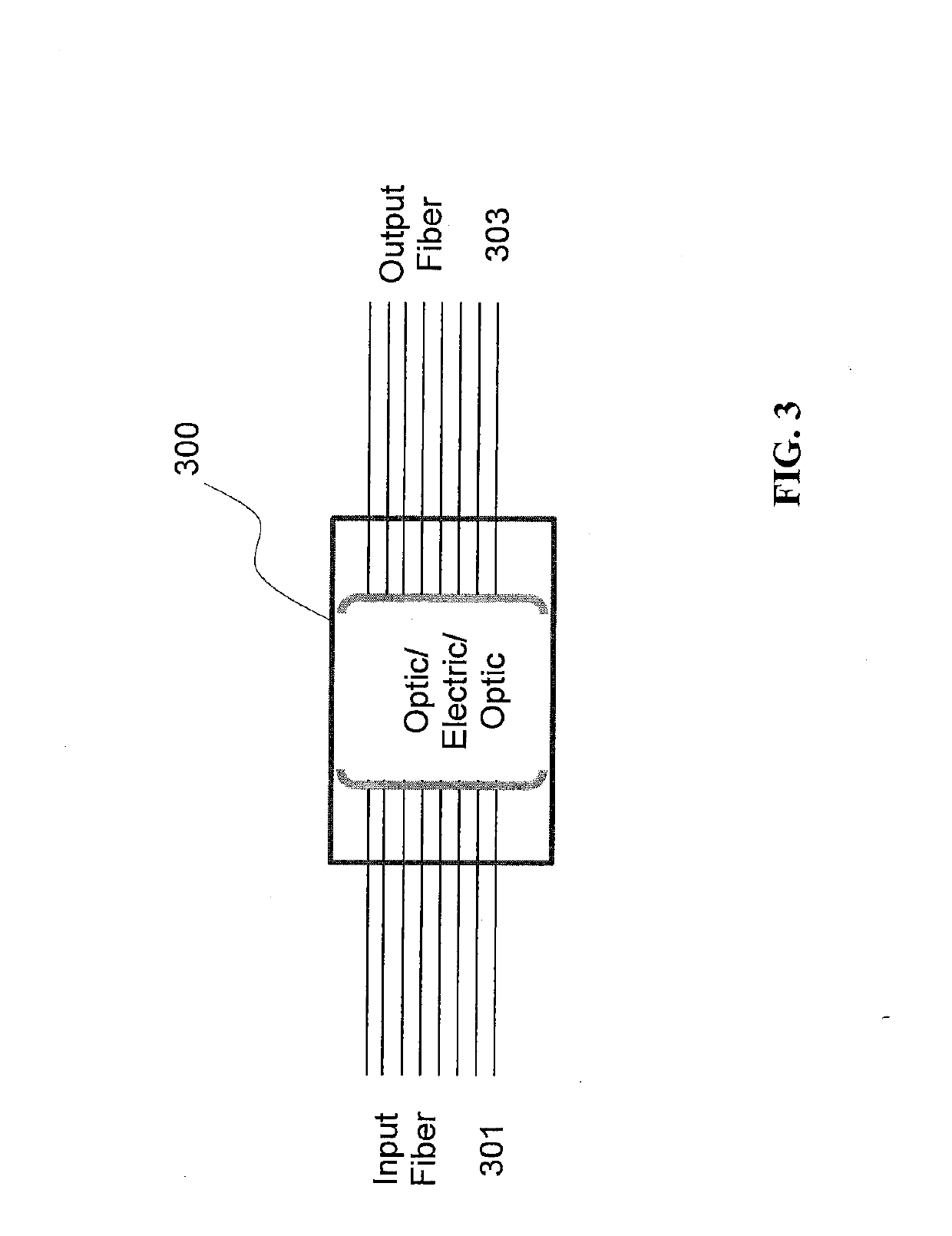
ActiveUS10108573B1Easy to handleEnergy efficient computingElectric digital data processingScalable systemPhotonics
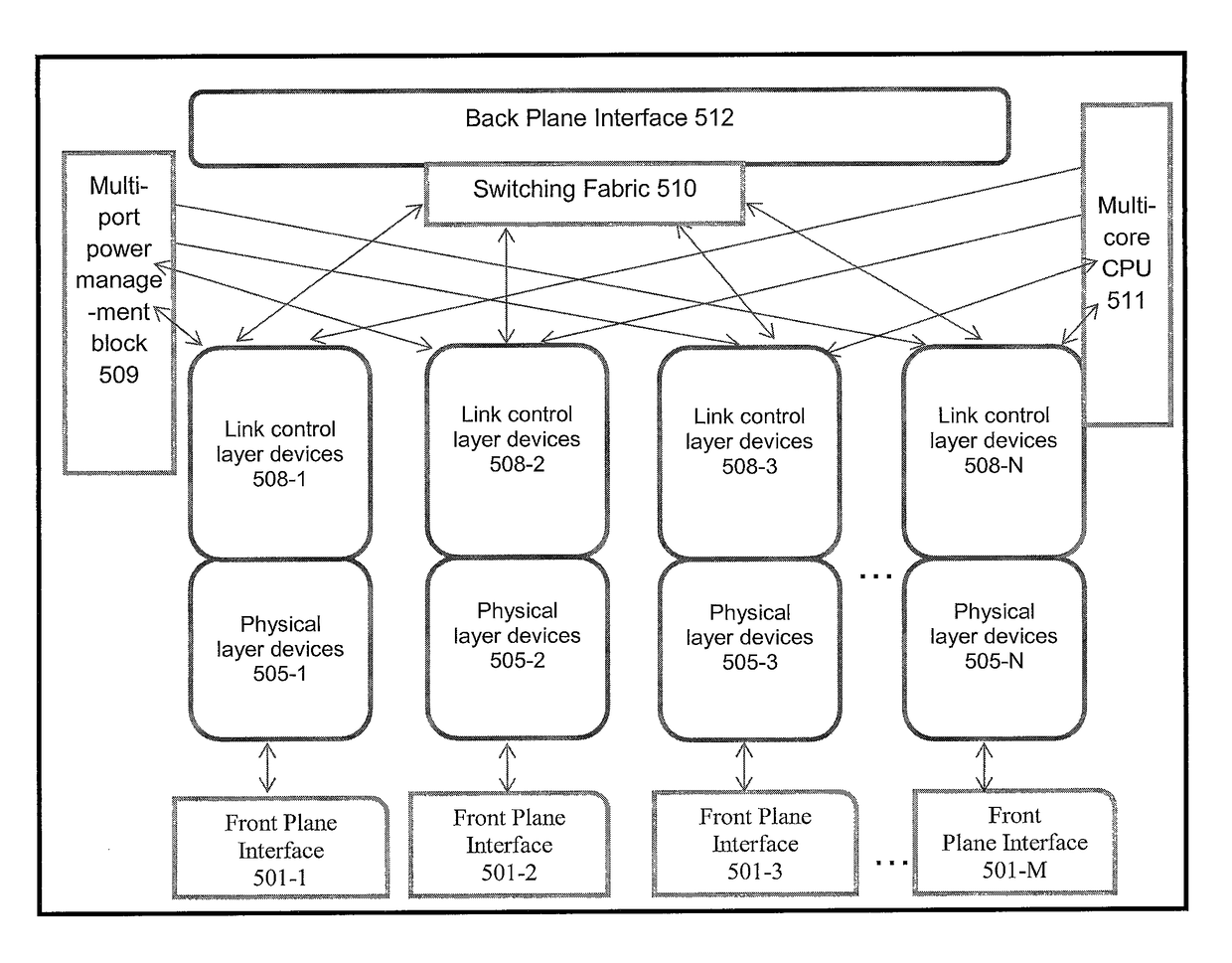
The new architecture disclosed herein exploits advances in system and chip technologies to implement a scalable multi-port open network. Using System-on-a-Chip (SOCs) and / or Multi-Chip-Module (MCM) technology, the architecture is implemented to efficiently handle multi-port switching. The novelty lies in using multi-core computing model in the data, control and management planes of multi-port networking cards implemented as an elemental scalable system (ESS) comprising N number of Elemental Units (EUs). EUs comprise device arrays on an integrated circuit (IC) platform using integrated silicon photonics or discrete electro-optics. TX4M™ system architecture therefore includes multiple EUs, switch fabric, multi-core central processing unit (CPU), multi-port power management module with embedded programmable logic, a back plane interface (BPI) as well as selectable functions for front plane interface (FPI) implemented in FPGAs for integration of front plane interface optics on host or on pluggable modules.
Owner:OPTEL NETWORKS INC
Method for enhanced security of computational device with multiple cores
ActiveUS20170155502A1Improve security levelEnhanced resistance against security threatsProgram initiation/switchingResource allocationMulti core computingProcessing core
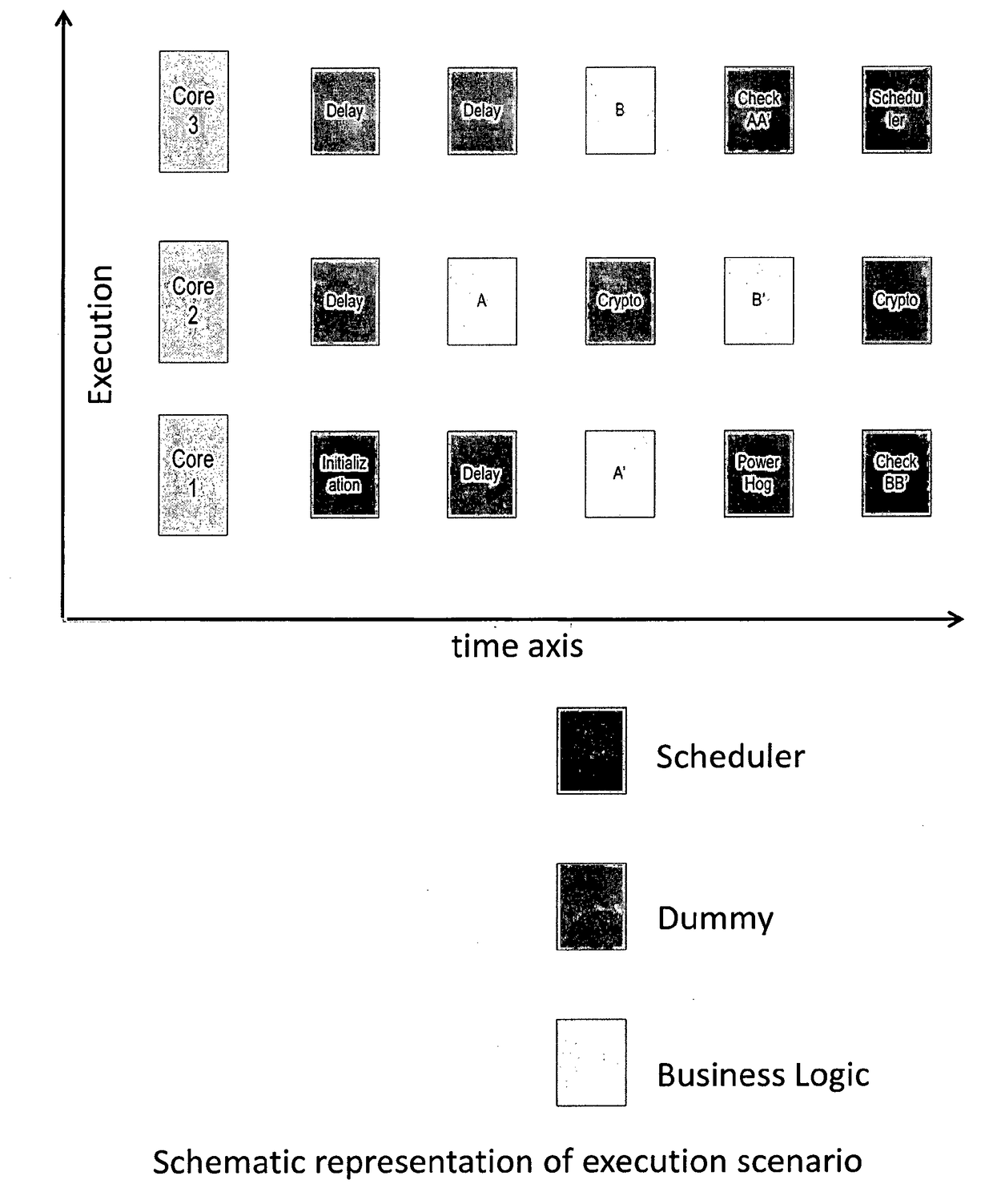
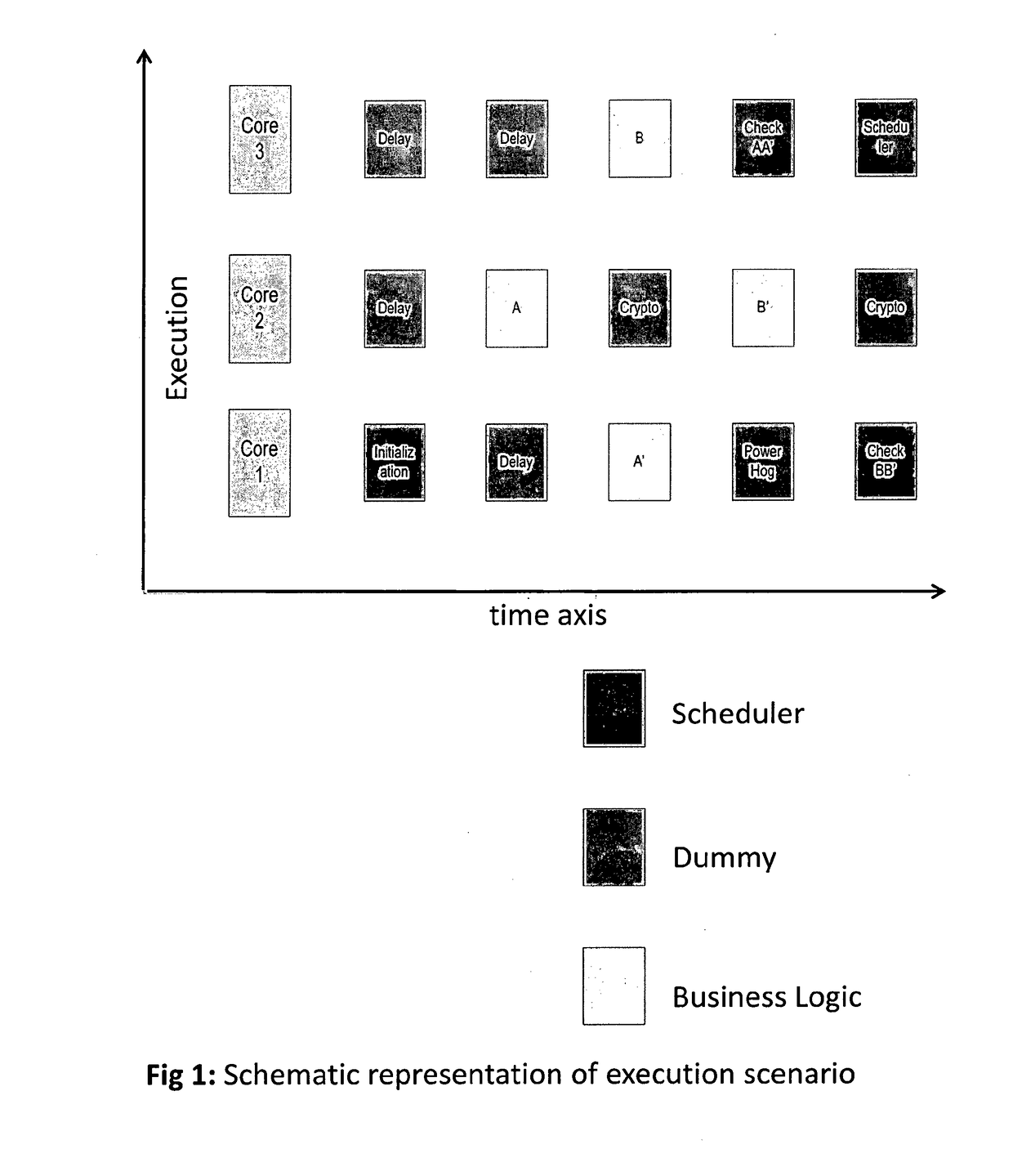
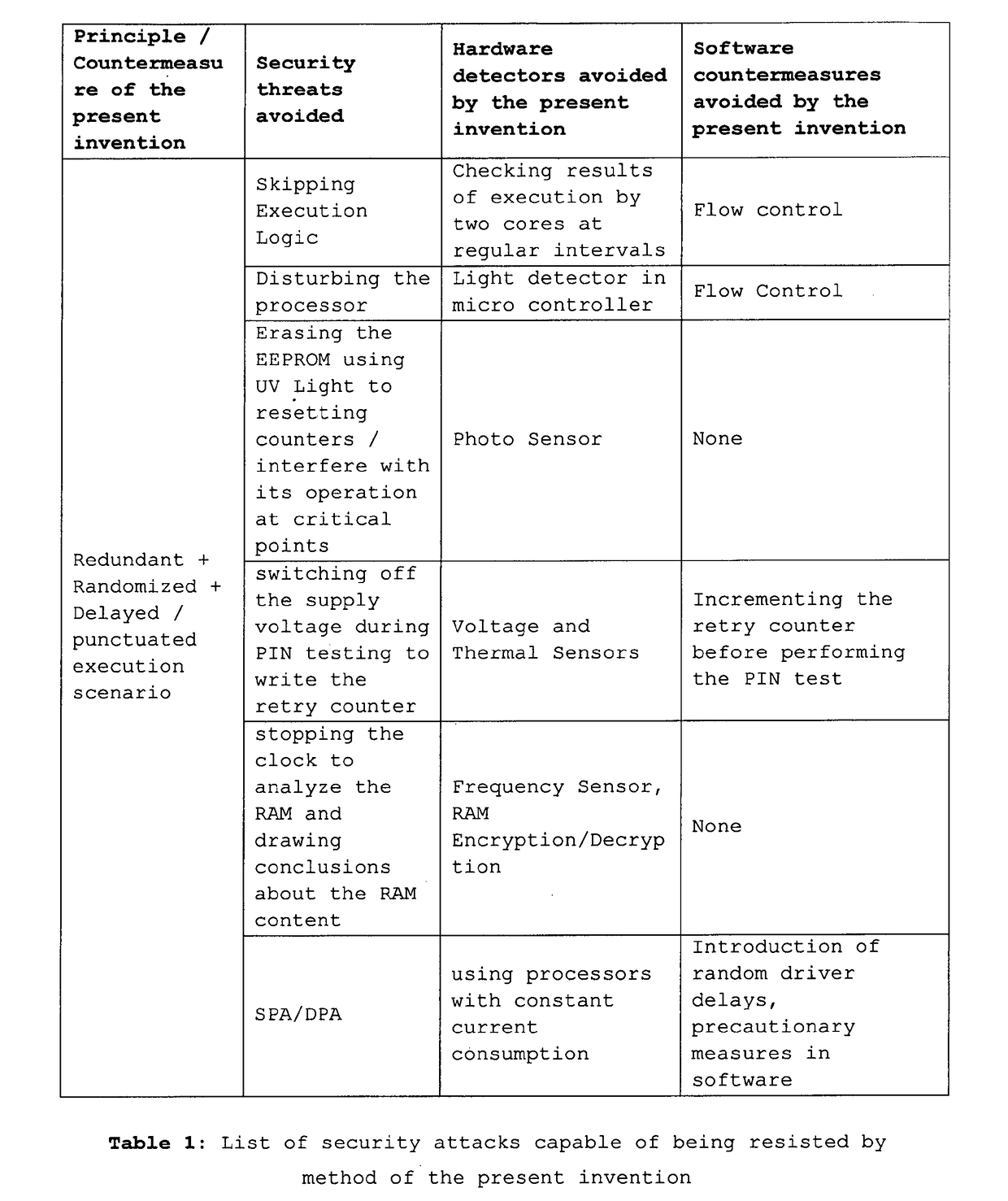
A method to protect computational, in particular cryptographic, devices having multi-core processors from DPA and DFA attacks is disclosed herein. The method implies: Defining a library of execution units functionally grouped into business function related units, security function related units and scheduler function related units; Designating at random one among the plurality of processing cores on the computational device to as a master core for execution of the scheduler function related execution units; and Causing, under control of the scheduler, execution of the library of execution units, so as to result in a randomized execution flow capable of resisting security threats initiated on the computational device.
Owner:GIESECKE & DEVRIENT MOBILE SECURITY GMBH
Multi-robot rapid task scheduling method based on multi-core computing
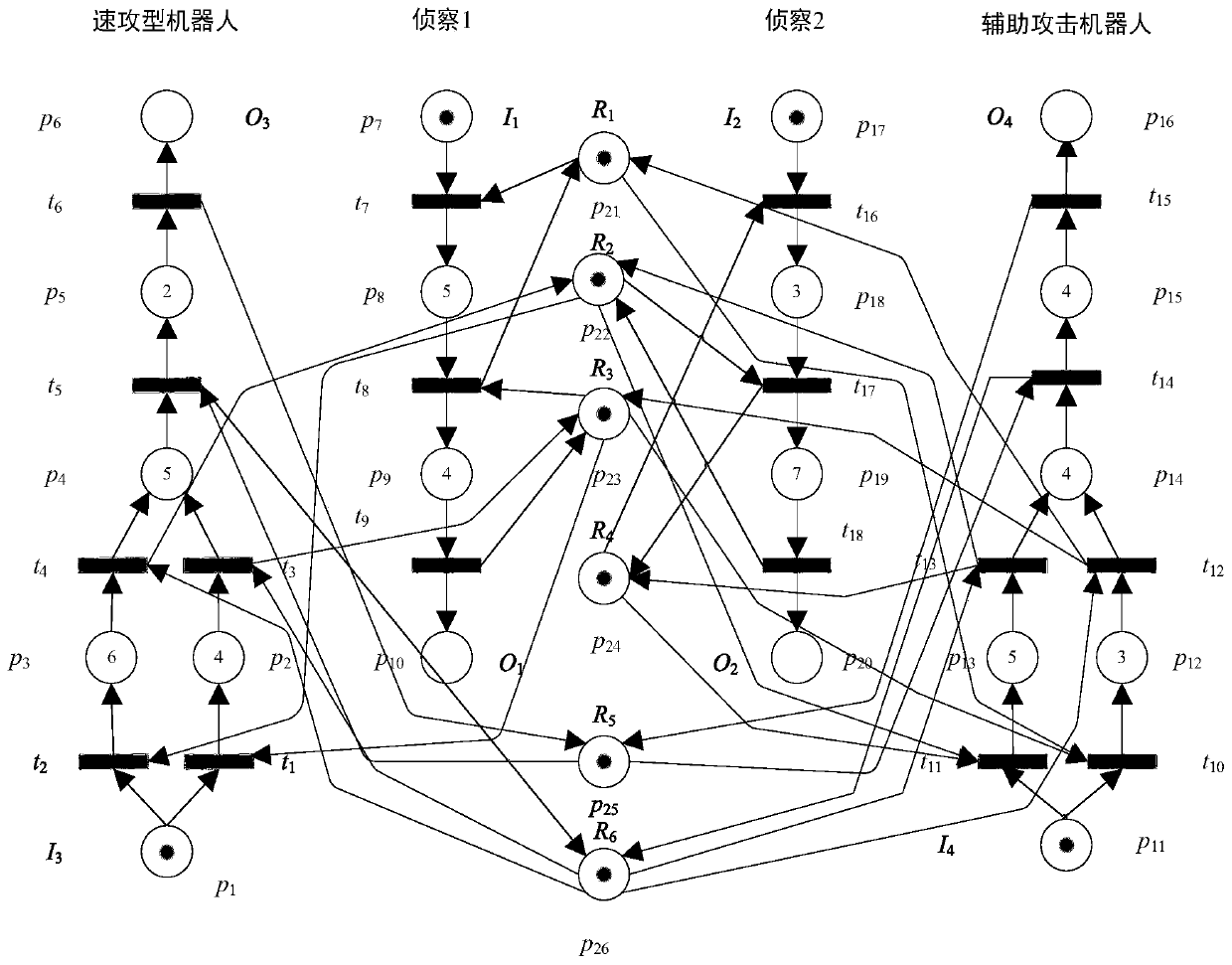
ActiveCN110348693AReduce time complexityFast searchForecastingResourcesTheoretical computer sciencePetri net
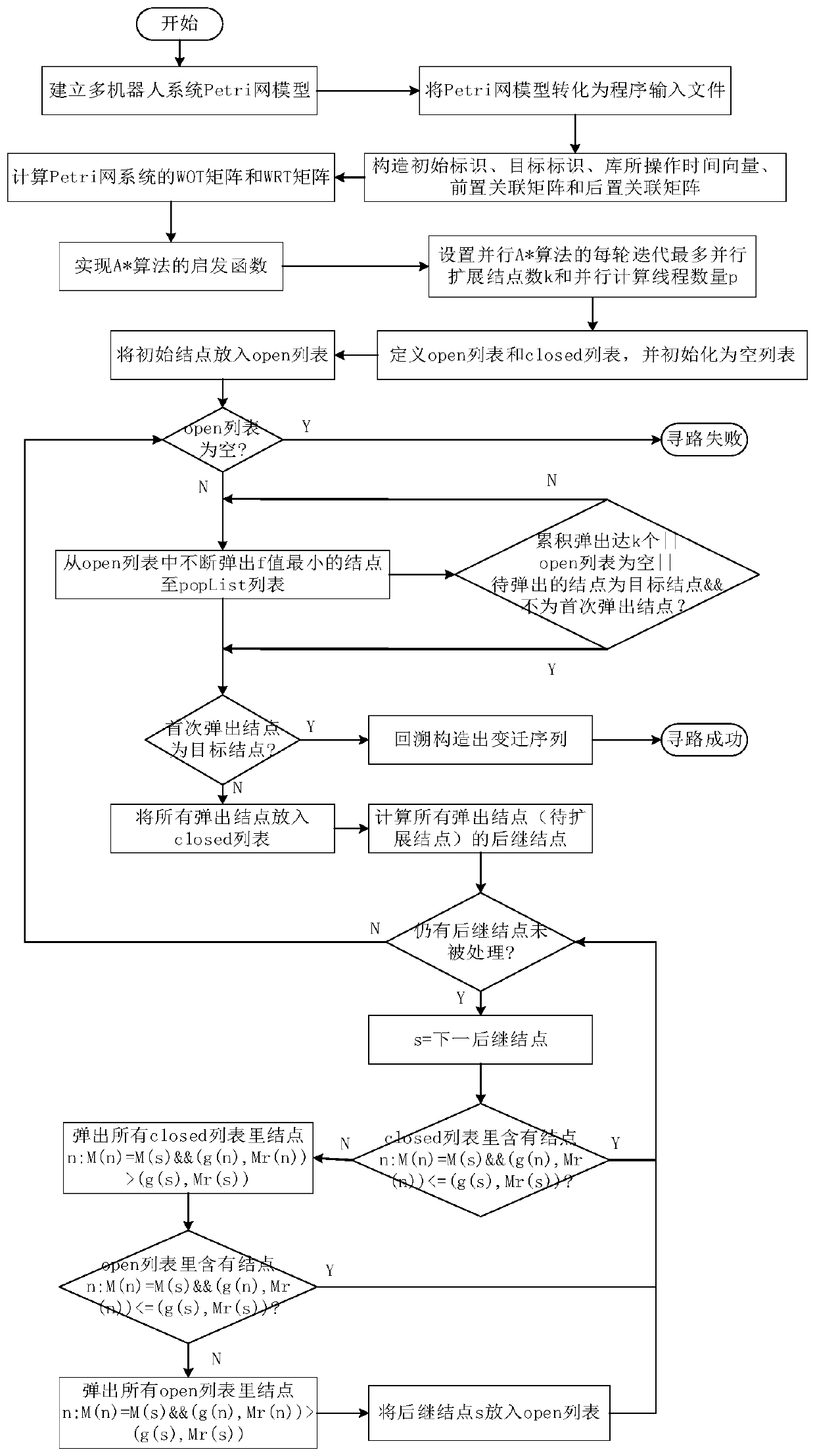
The invention discloses a multi-robot rapid task scheduling method based on multi-core computing. The method comprises the following steps: modeling a multi-robot system by adopting a Petri network modeling method; converting the Petri net model into a program input file according to a self-defined file format; solving a WOT matrix and a WRT matrix of the Petri network to realize a heuristic function required by the A * search algorithm; constructing an initial identifier, a target identifier, a library operation time vector, a front incidence matrix and a rear incidence matrix of a Petri network system according to the input file; constructing a Treap data structure representation open list supporting concurrency and a binary balance tree data structure representation closed list supporting concurrency; setting the maximum number k of parallel expansion nodes and the number p of parallel computing threads of each round of iteration of the parallel A * algorithm; and running a parallelA * algorithm to search a transmission transition sequence from the initial node to the target node in parallel to obtain a task scheduling scheme. The method has the advantages of strong algorithm parallelism, high parallelism efficiency, high path search efficiency, high quality of the obtained task scheduling scheme and the like.
Owner:NANJING UNIV OF SCI & TECH
Transformational architecture for multi-layer systems
InactiveUS20190095373A1Easy to handleEnergy efficient computingElectric digital data processingScalable systemPhotonics
The new architecture disclosed herein exploits advances in system and chip technologies to implement a scalable multi-port open network. Using System-on-a-Chip (SOCs) and / or Multi-Chip-Module (MCM) technology, the architecture is implemented to efficiently handle multi-port switching. The novelty lies in using multi-core computing model in the data, control and management planes of multi-port networking cards implemented as an elemental scalable system (ESS) comprising N number of Elemental Units (EUs). EUs comprise device arrays on an integrated circuit (IC) platform using integrated silicon photonics or discrete electro-optics. TX4M™ system architecture therefore includes multiple EUs, switch fabric, multi-core central processing unit (CPU), multi-port power management module with embedded programmable logic, a back plane interface (BPI) as well as selectable functions for front plane interface (FPI) implemented in FPGAs for integration of front plane interface optics on host or on pluggable modules.
Owner:OPTEL NETWORKS INC