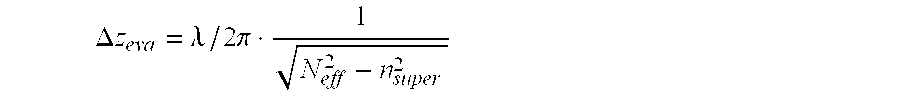Patents
Literature
700 results about "Label free" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
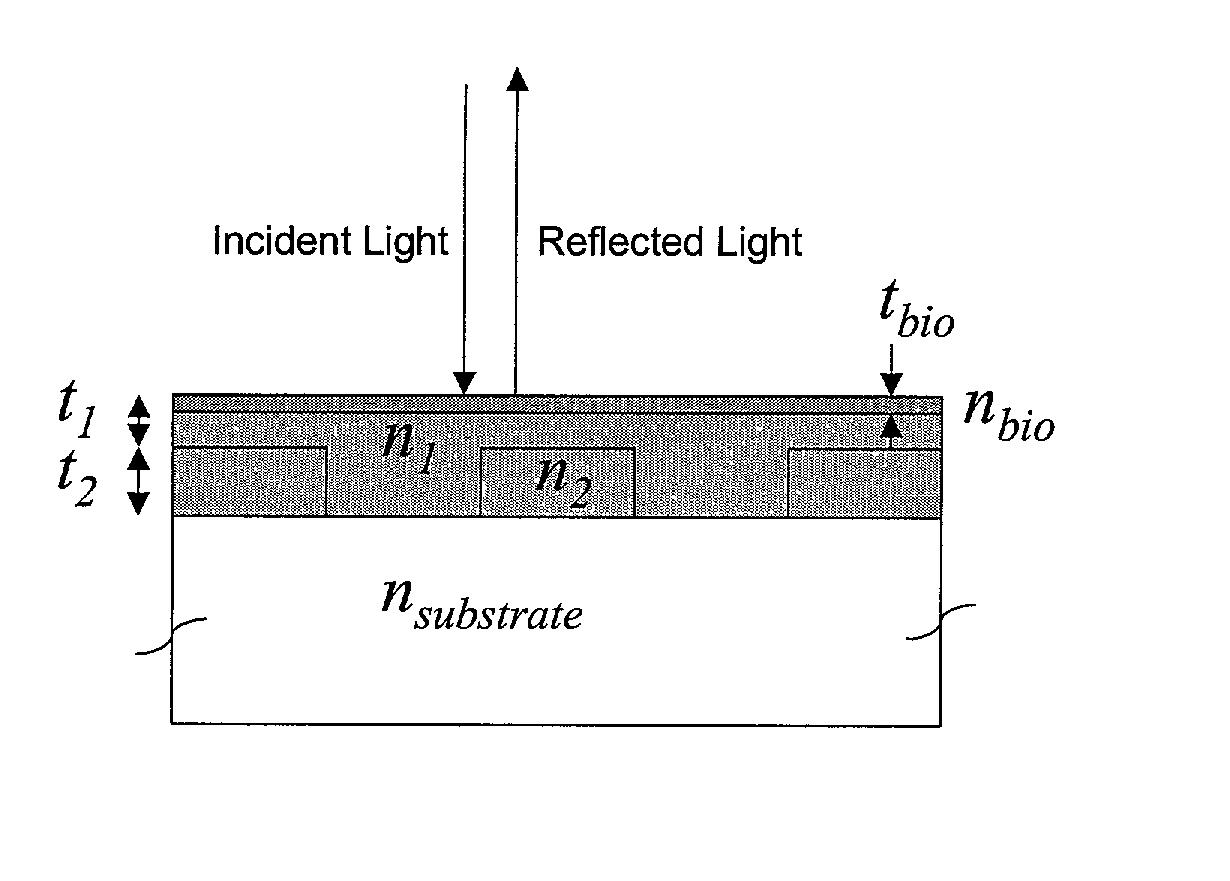
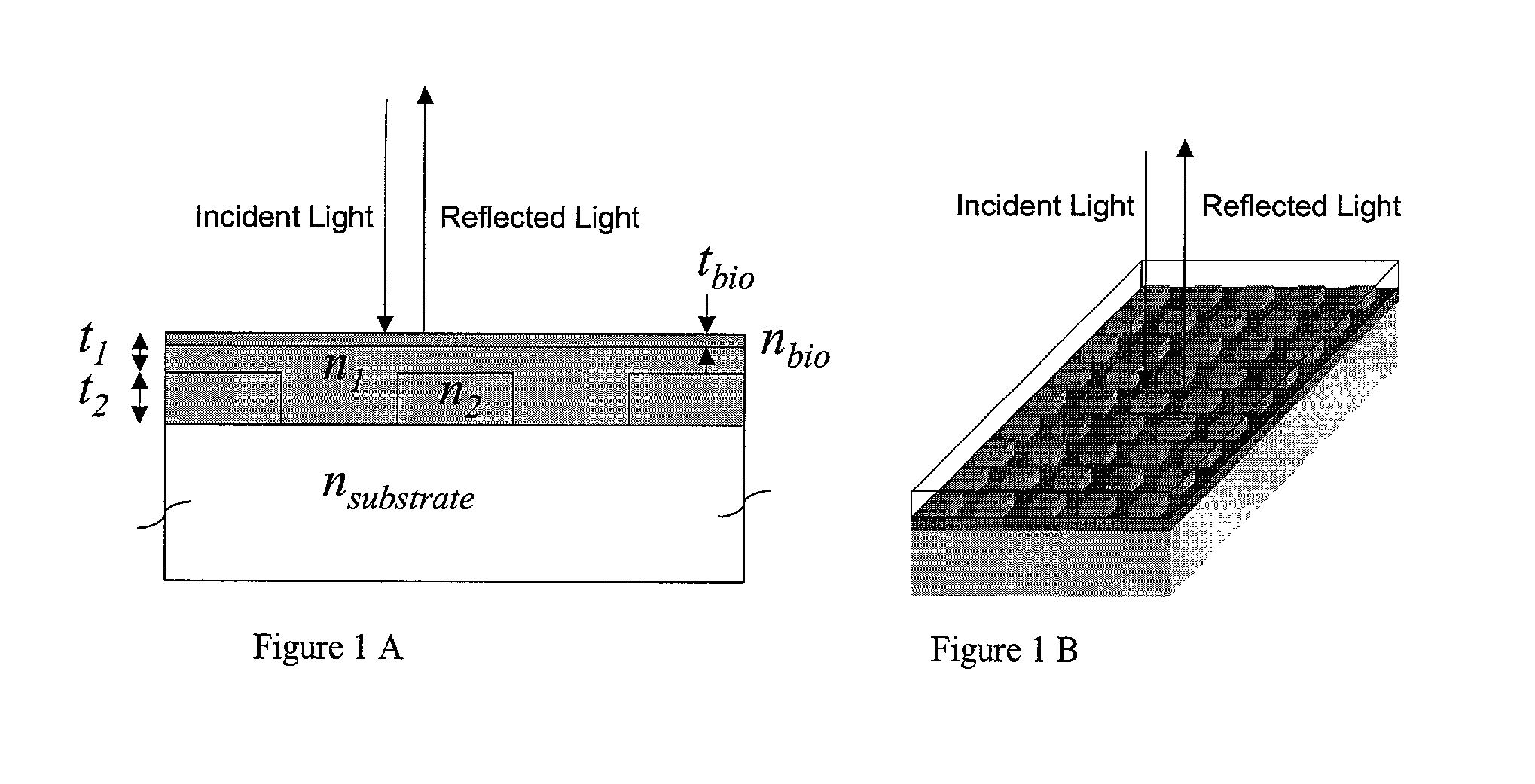
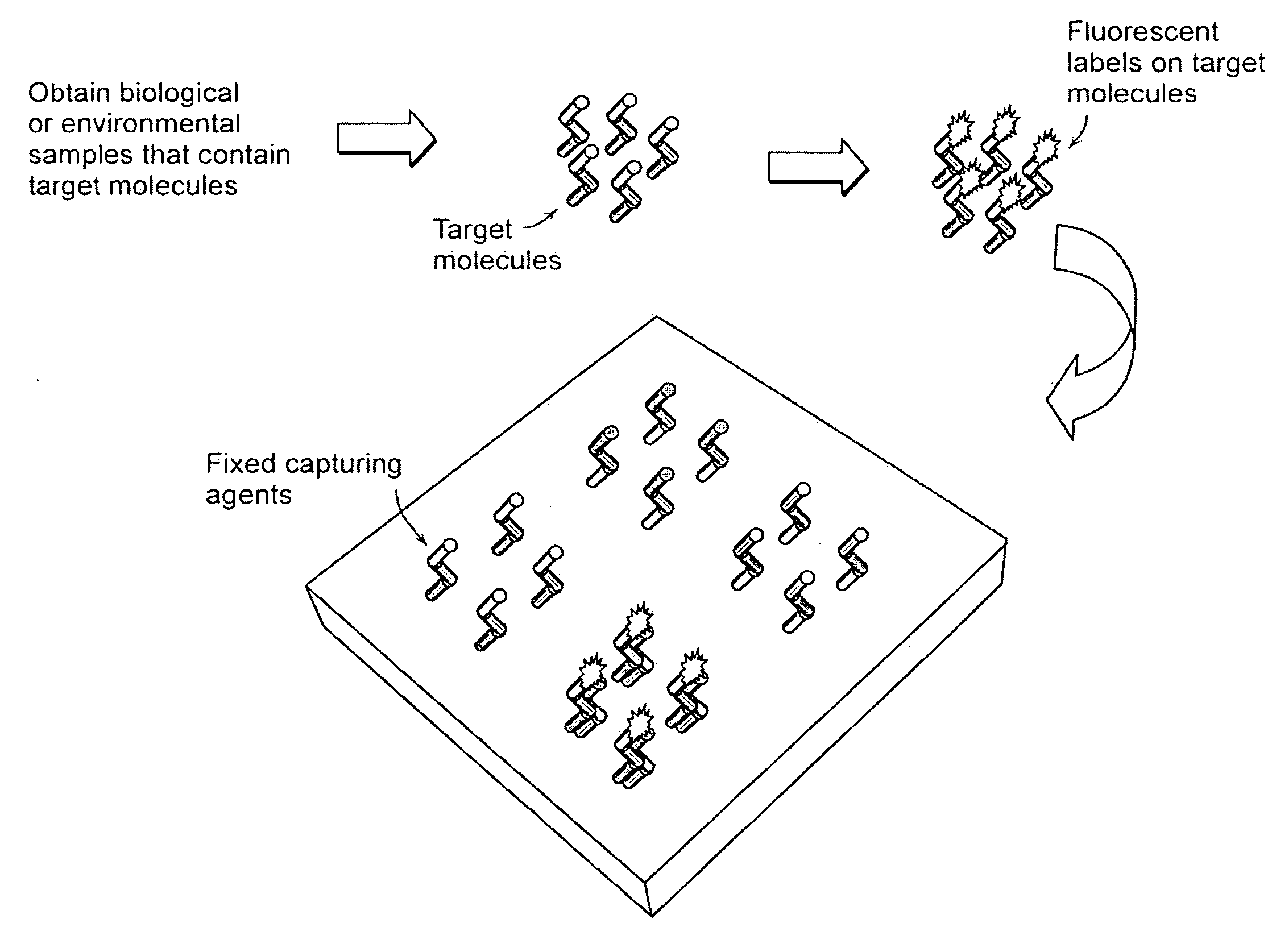
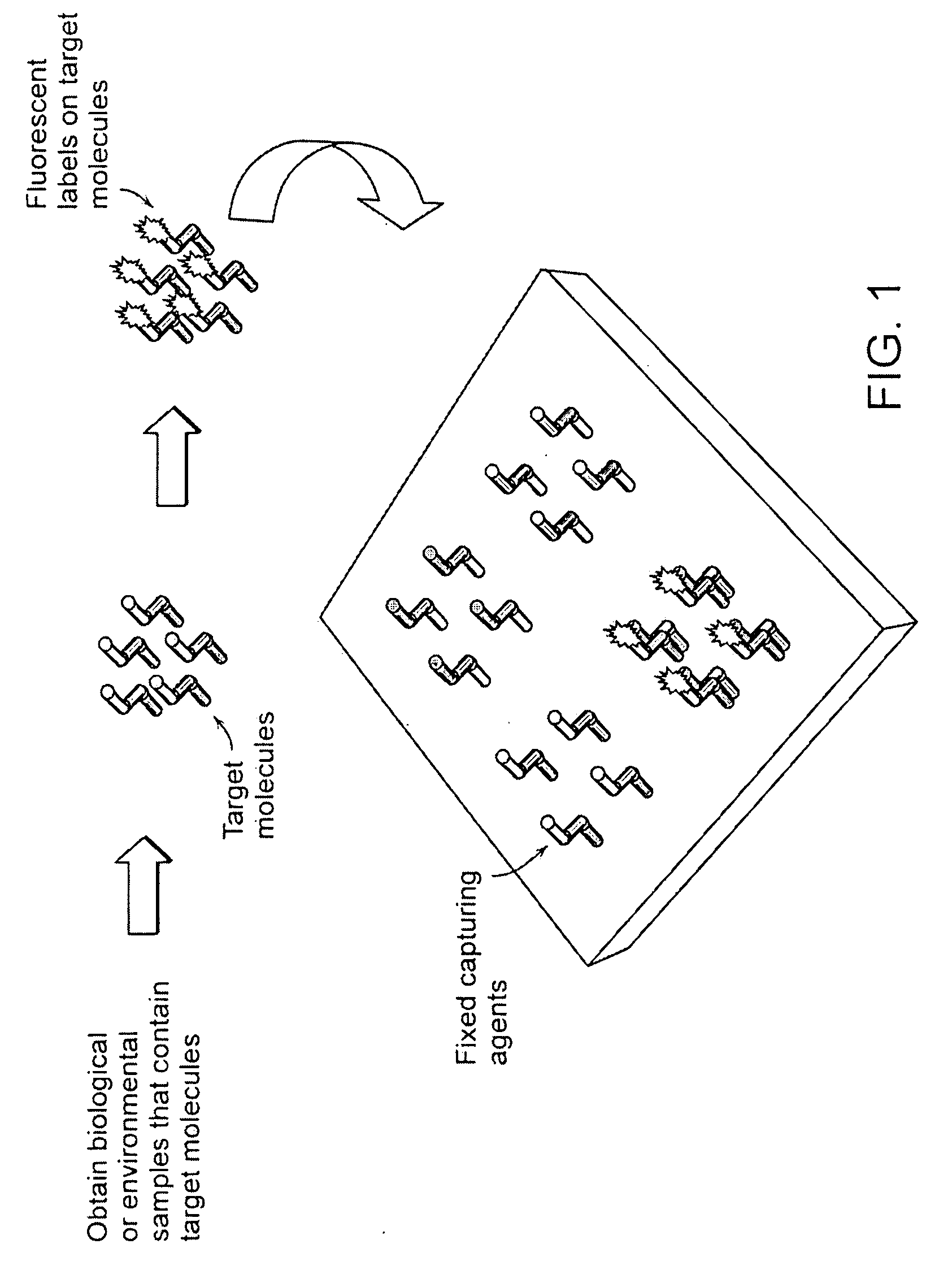
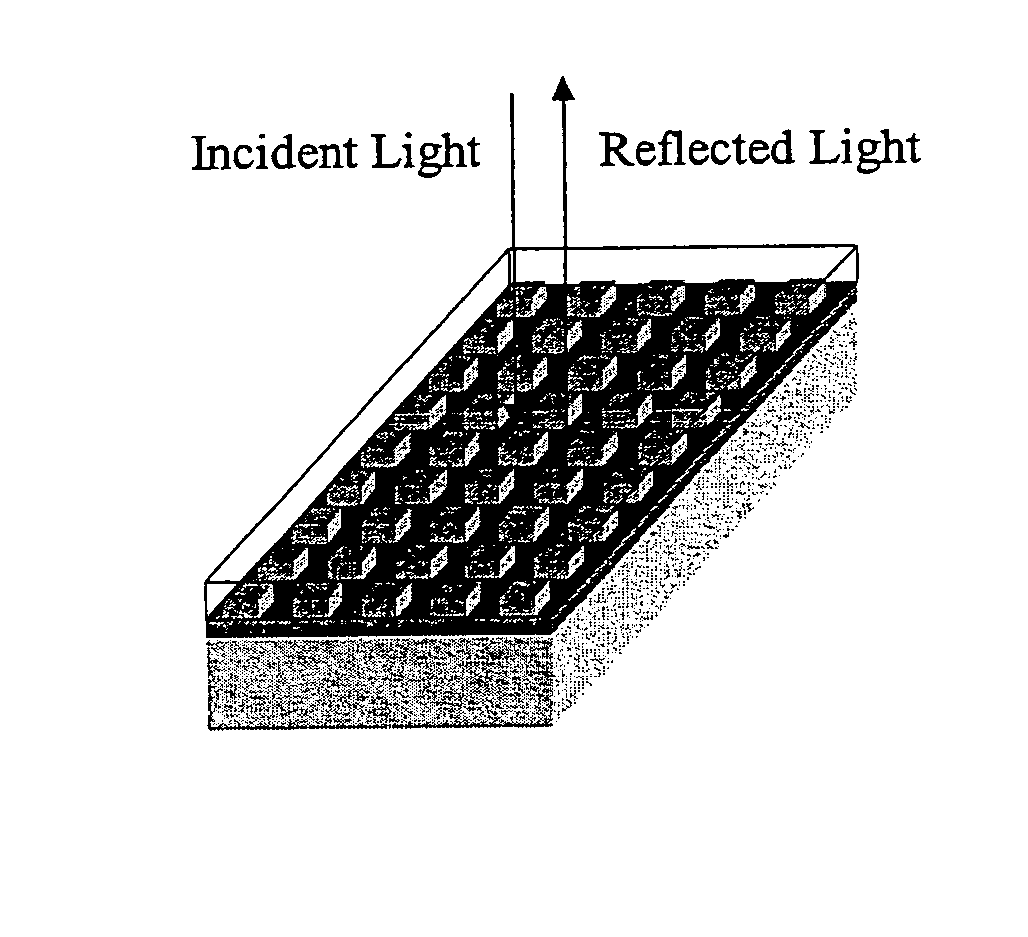
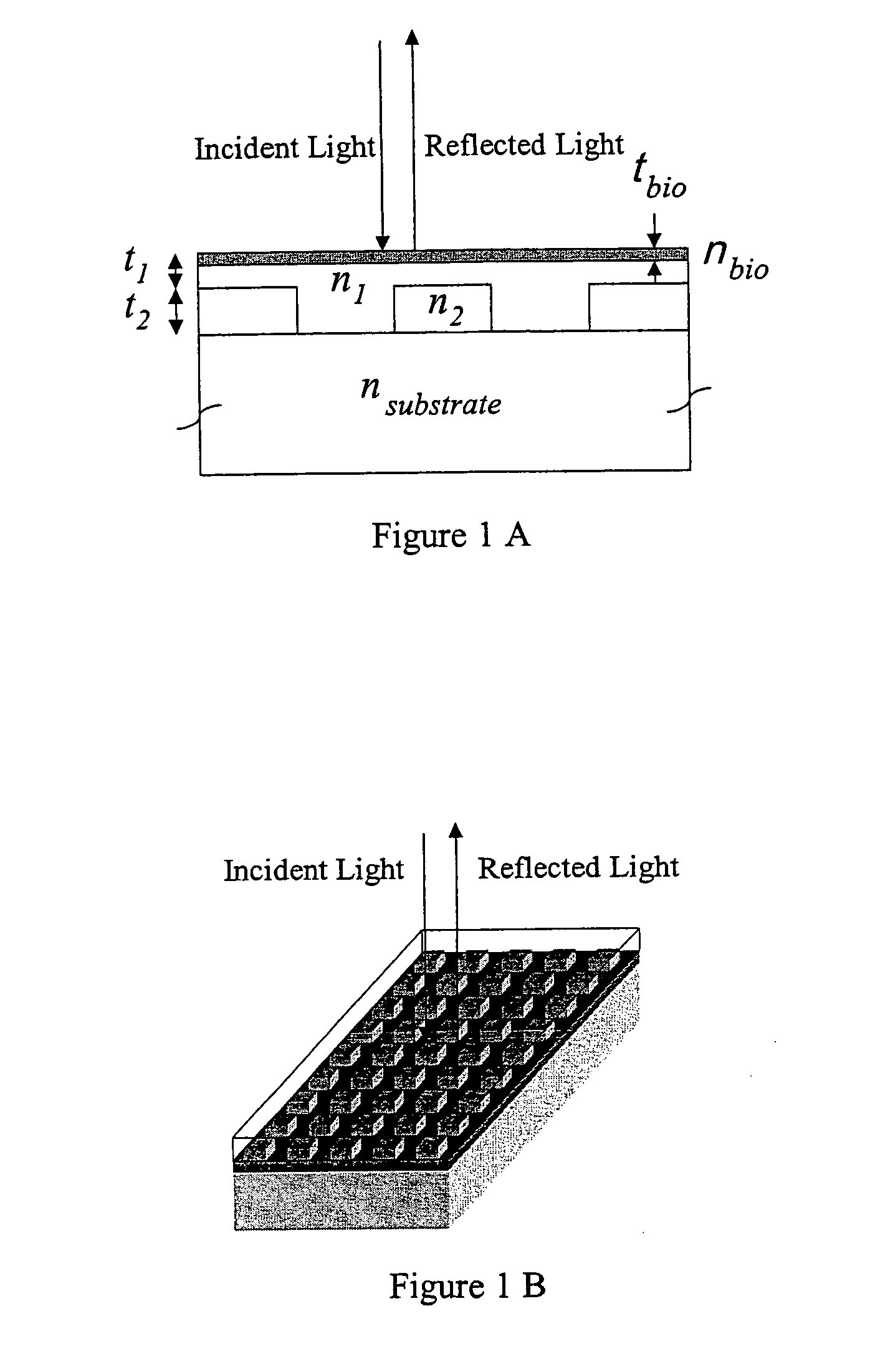
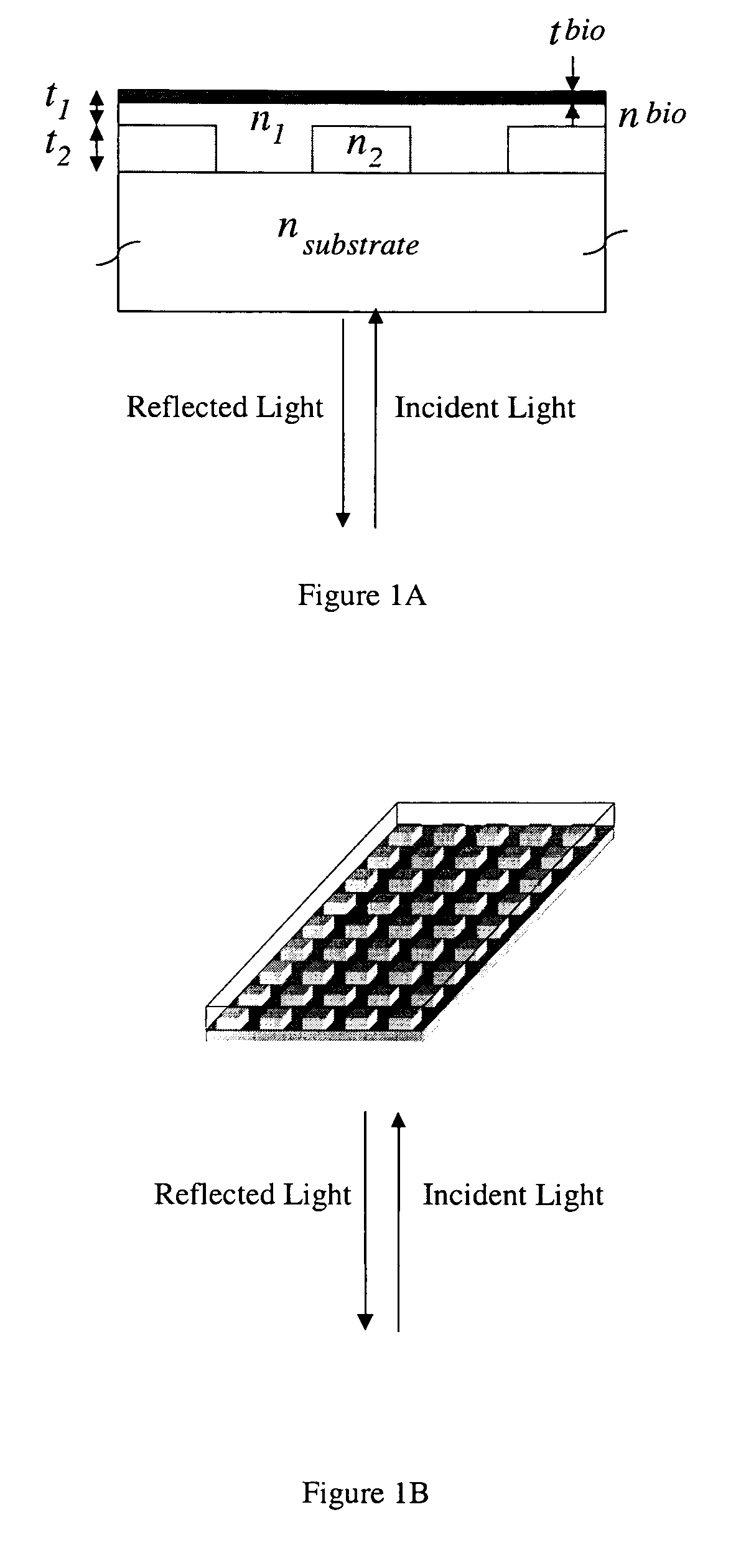
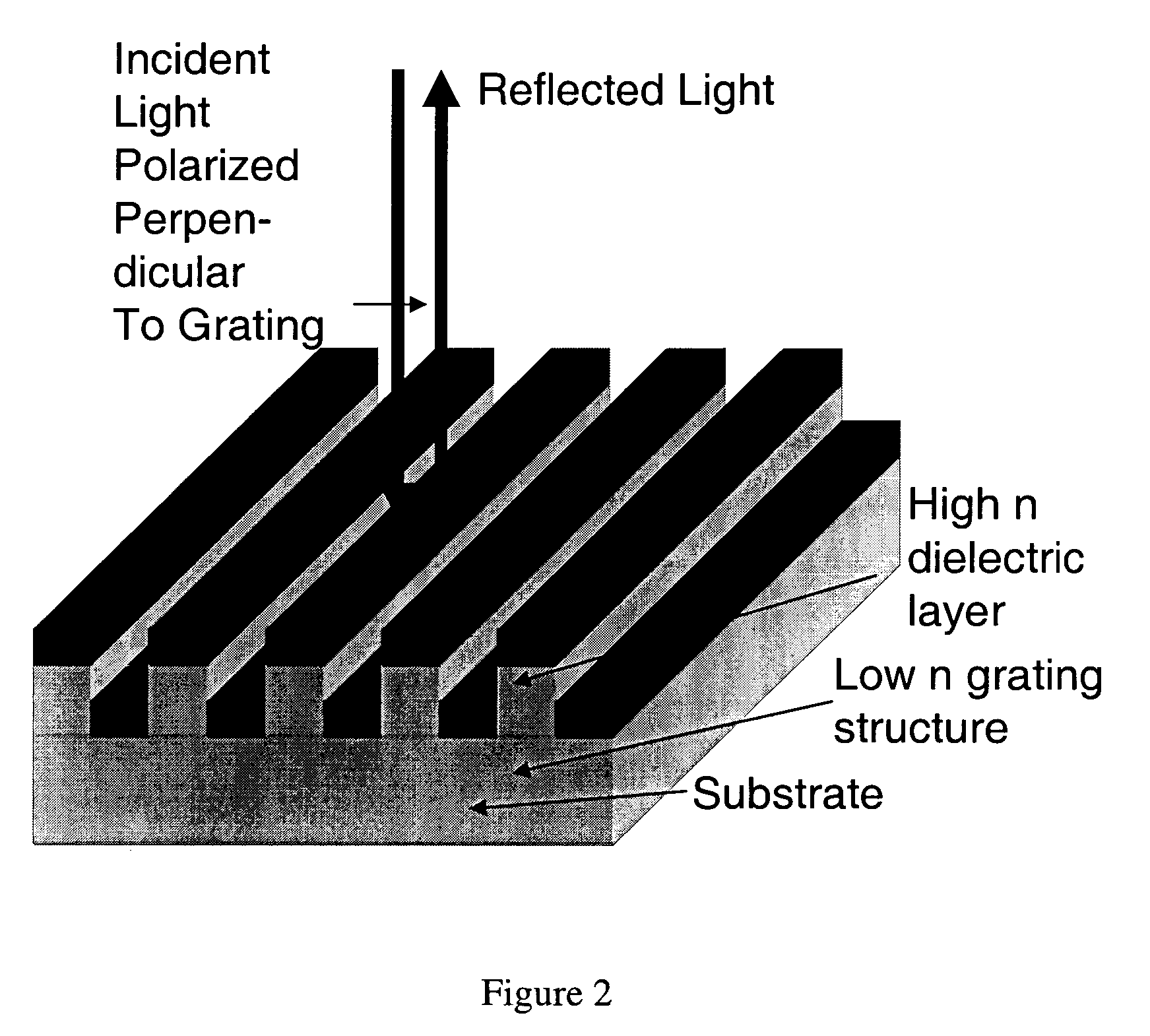
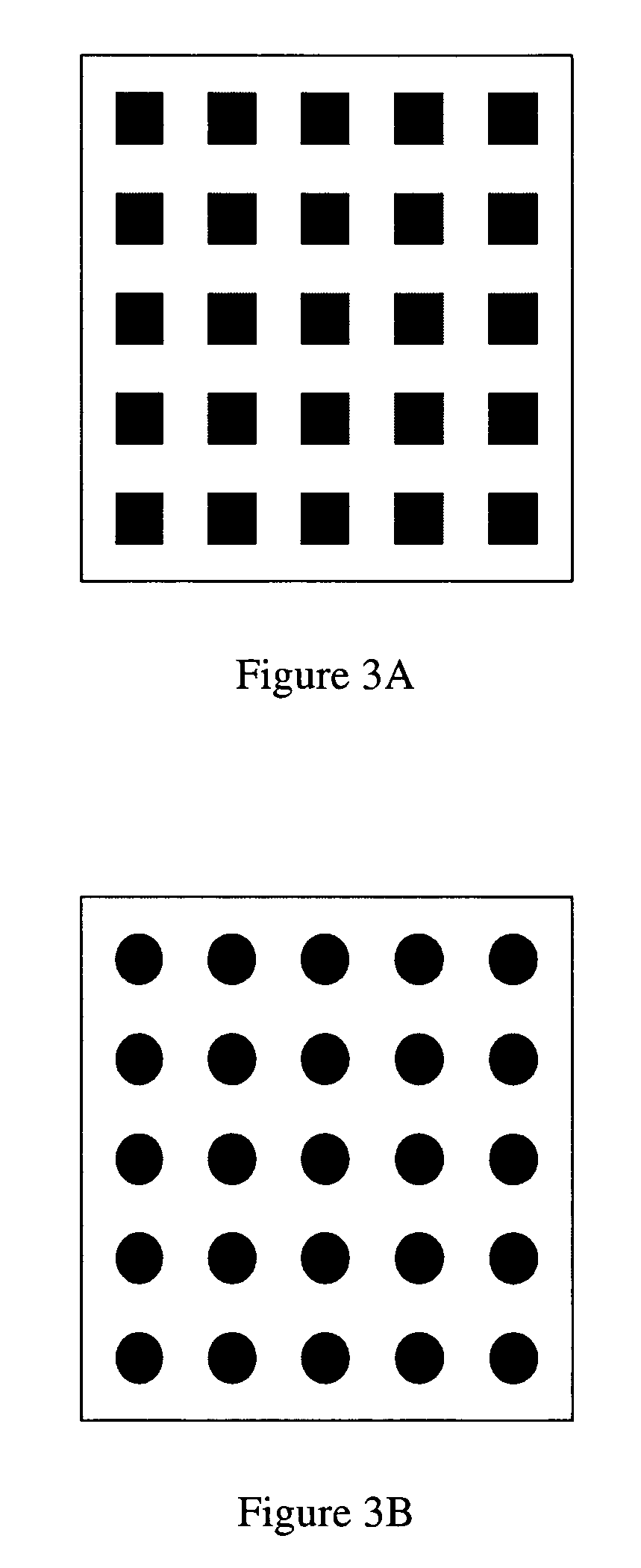
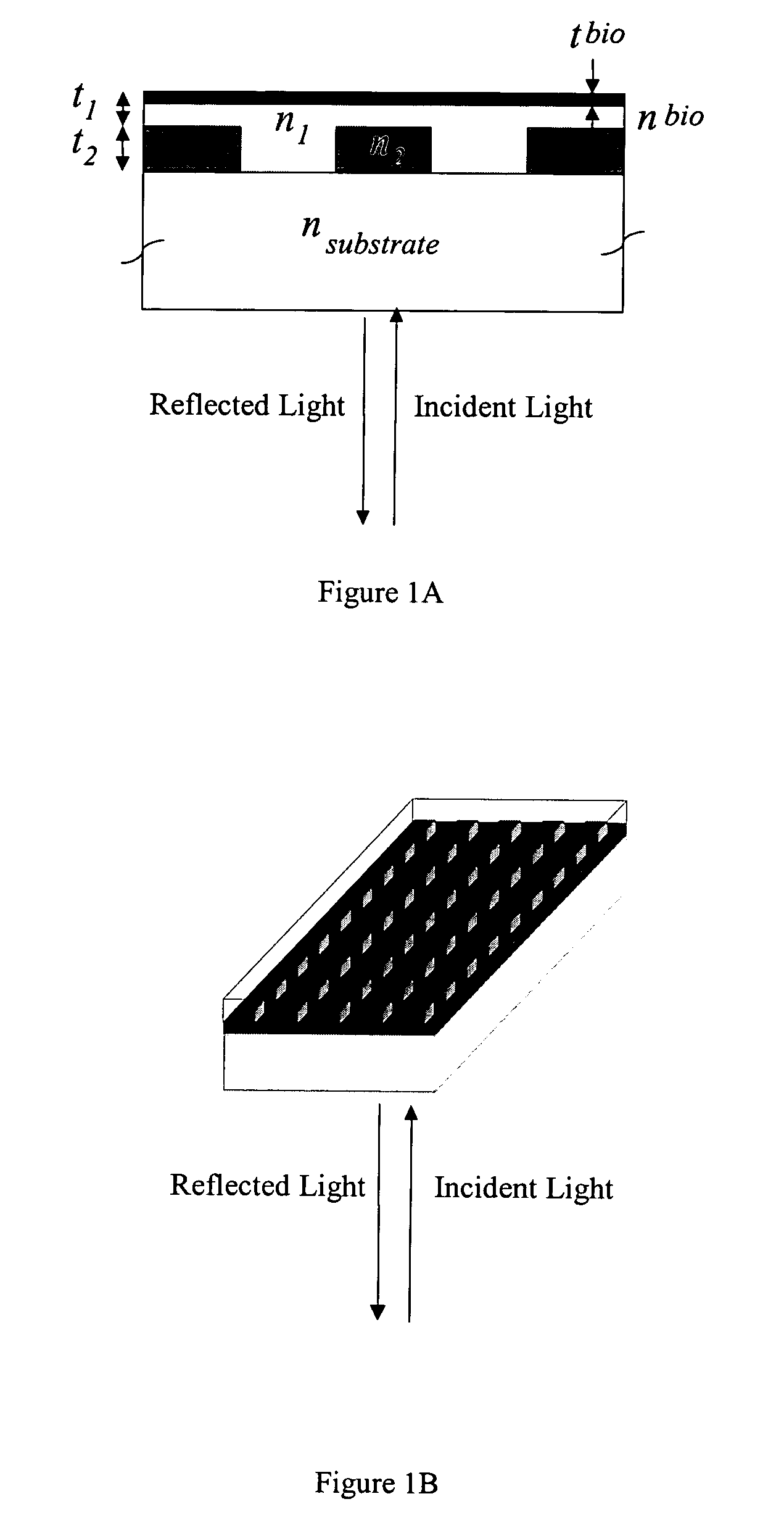
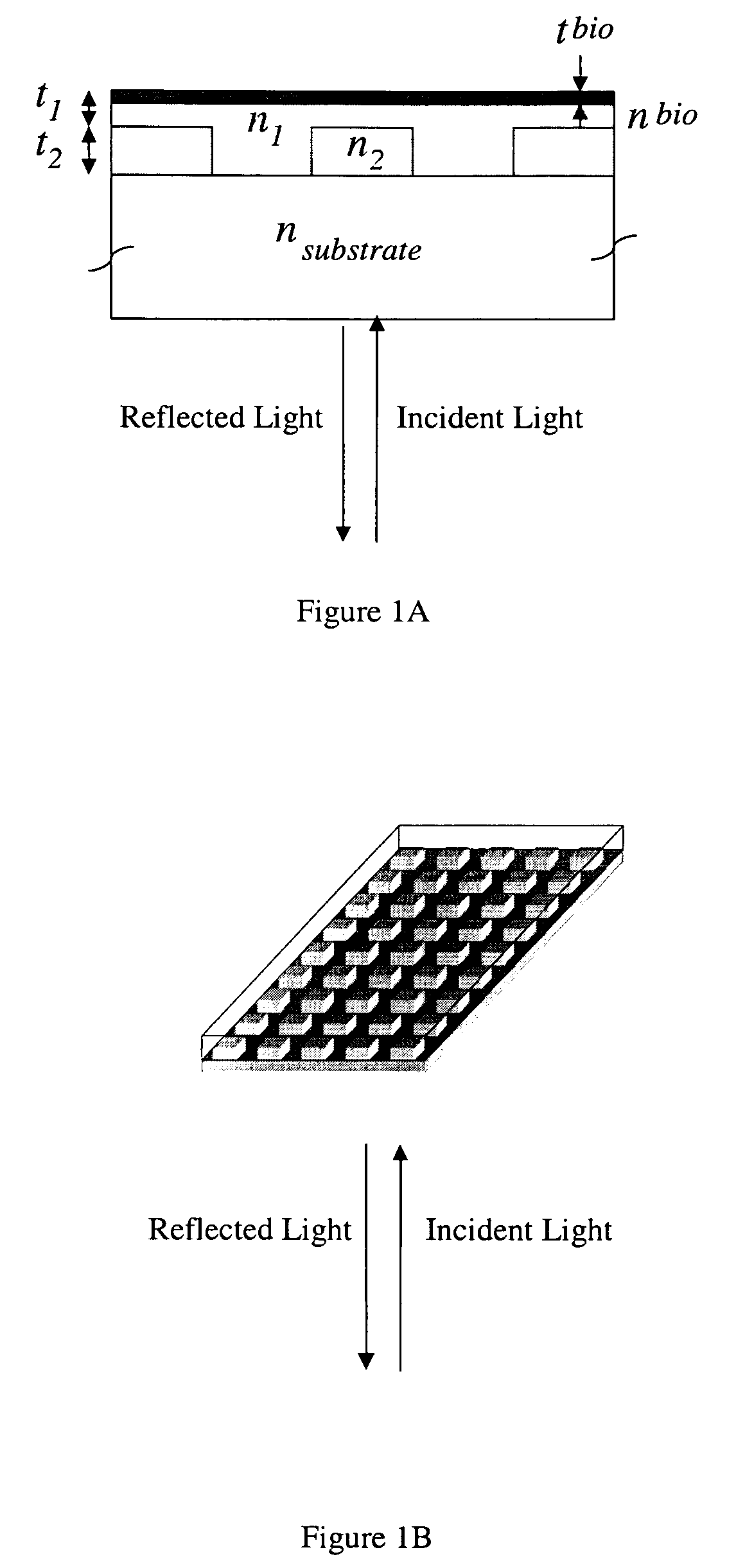
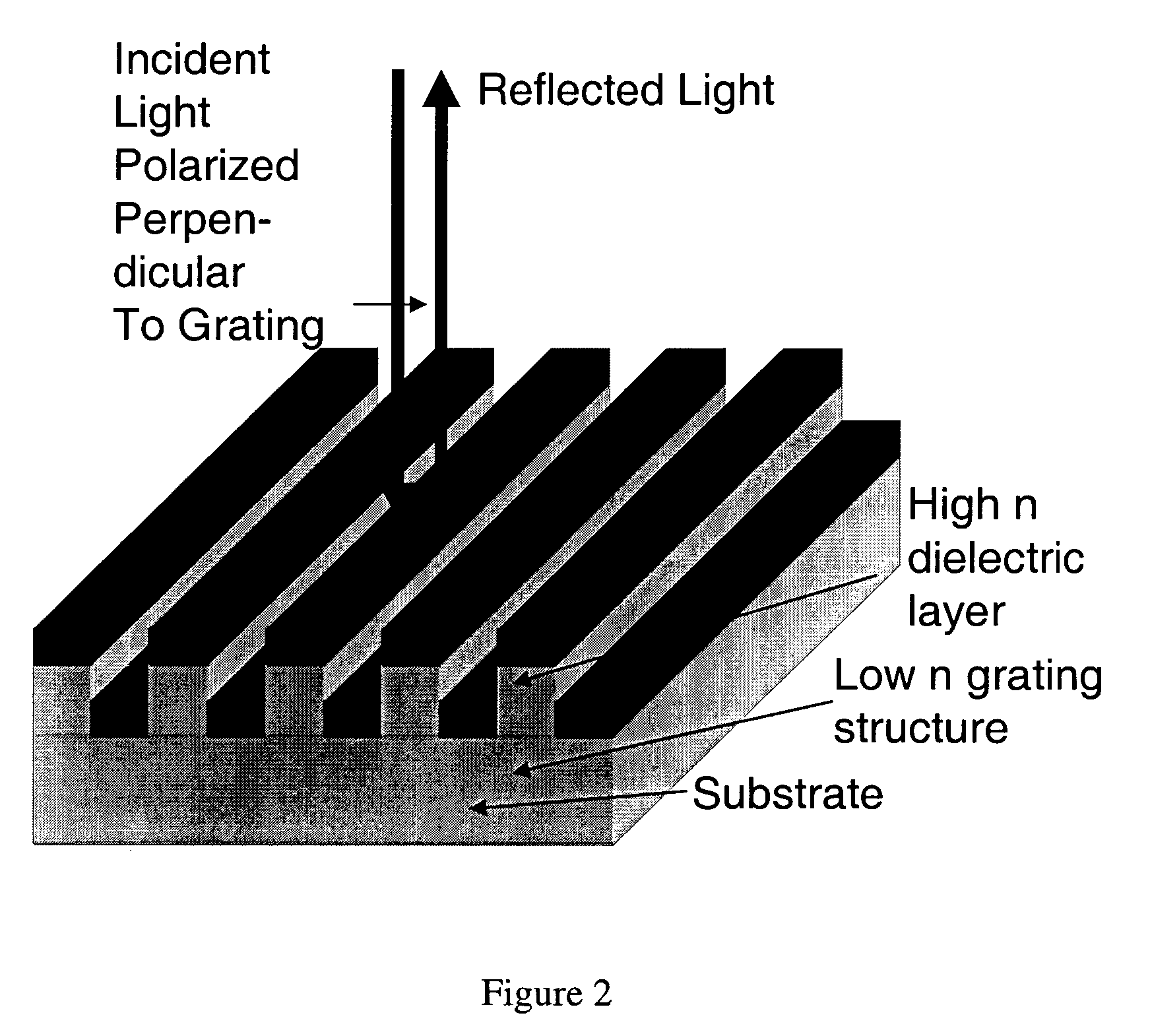
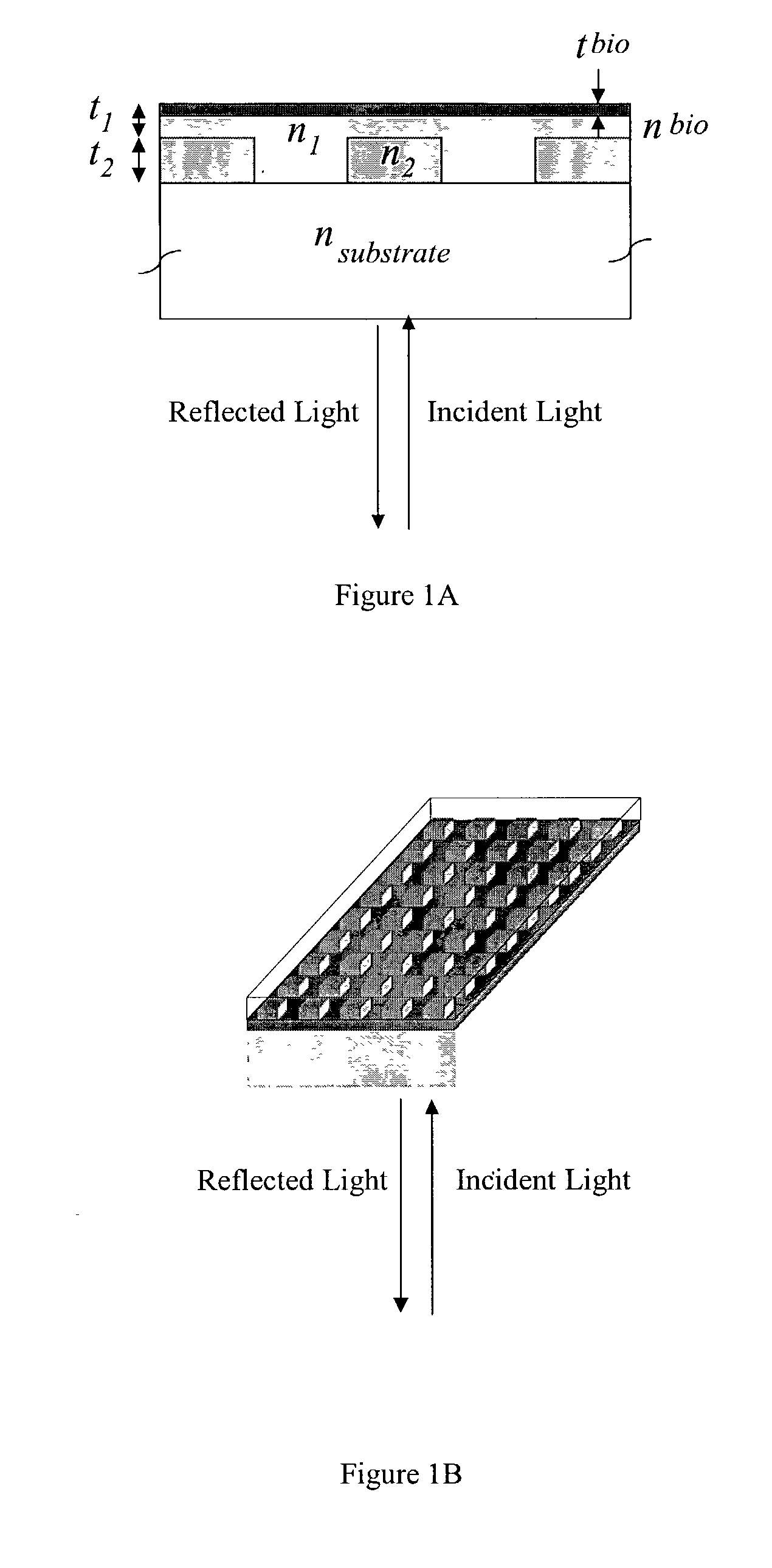
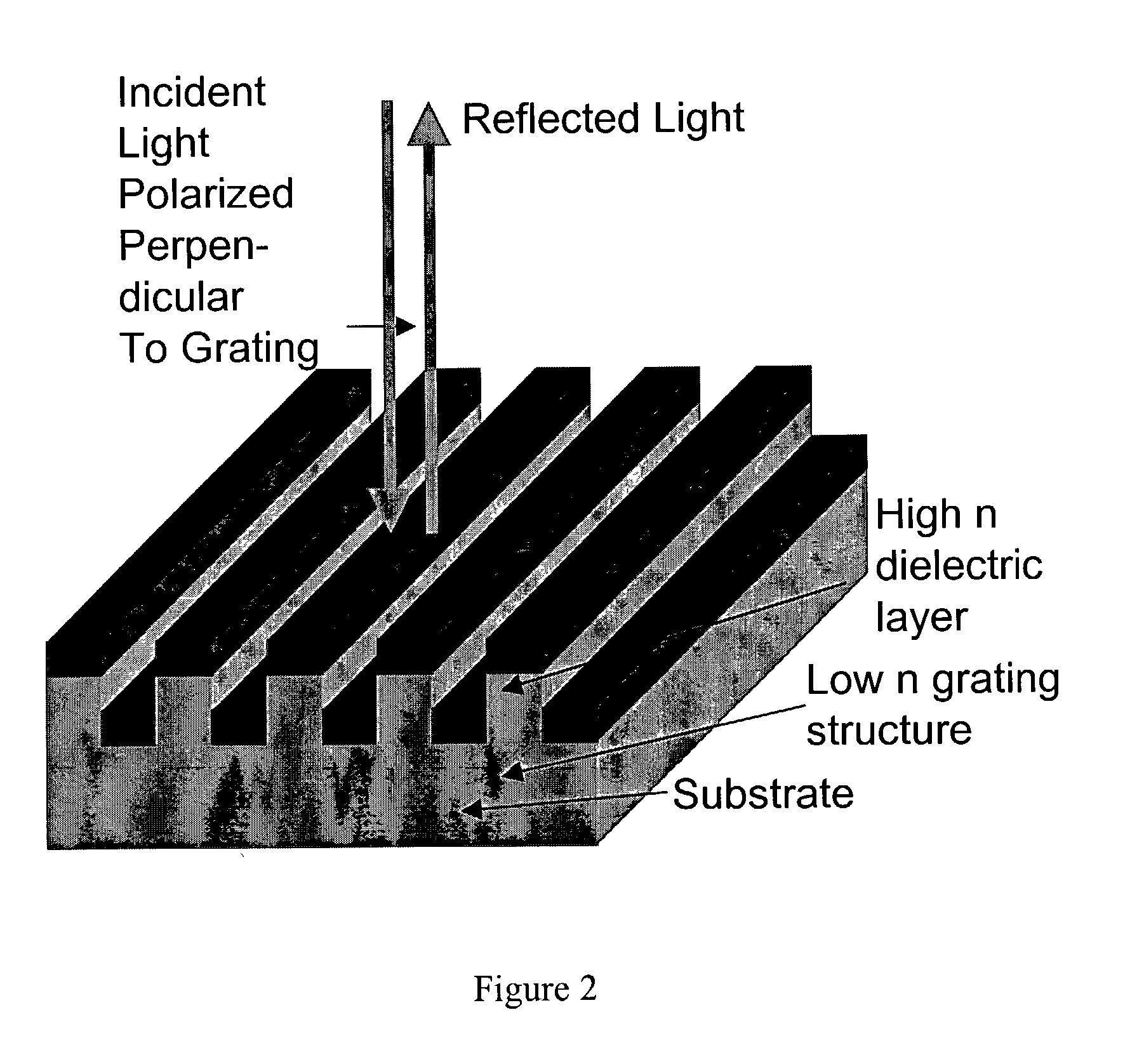
Label-free high-throughput optical technique for detecting biomolecular interactions
InactiveUS20020127565A1Inexpensively incorporatedHigh-throughput screeningBioreactor/fermenter combinationsBiological substance pretreatmentsMolecular interactionsThroughput
Methods and compositions are provided for detecting biomolecular interactions. The use of labels is not required and the methods can be performed in a high-throughput manner. The invention also provides optical devices useful as narrow band filters.
Owner:X BODY
Optical chemical/biochemical sensor
InactiveUS6395558B1Radiation pyrometryMaterial analysis by observing effect on chemical indicatorTransducerLuminescence
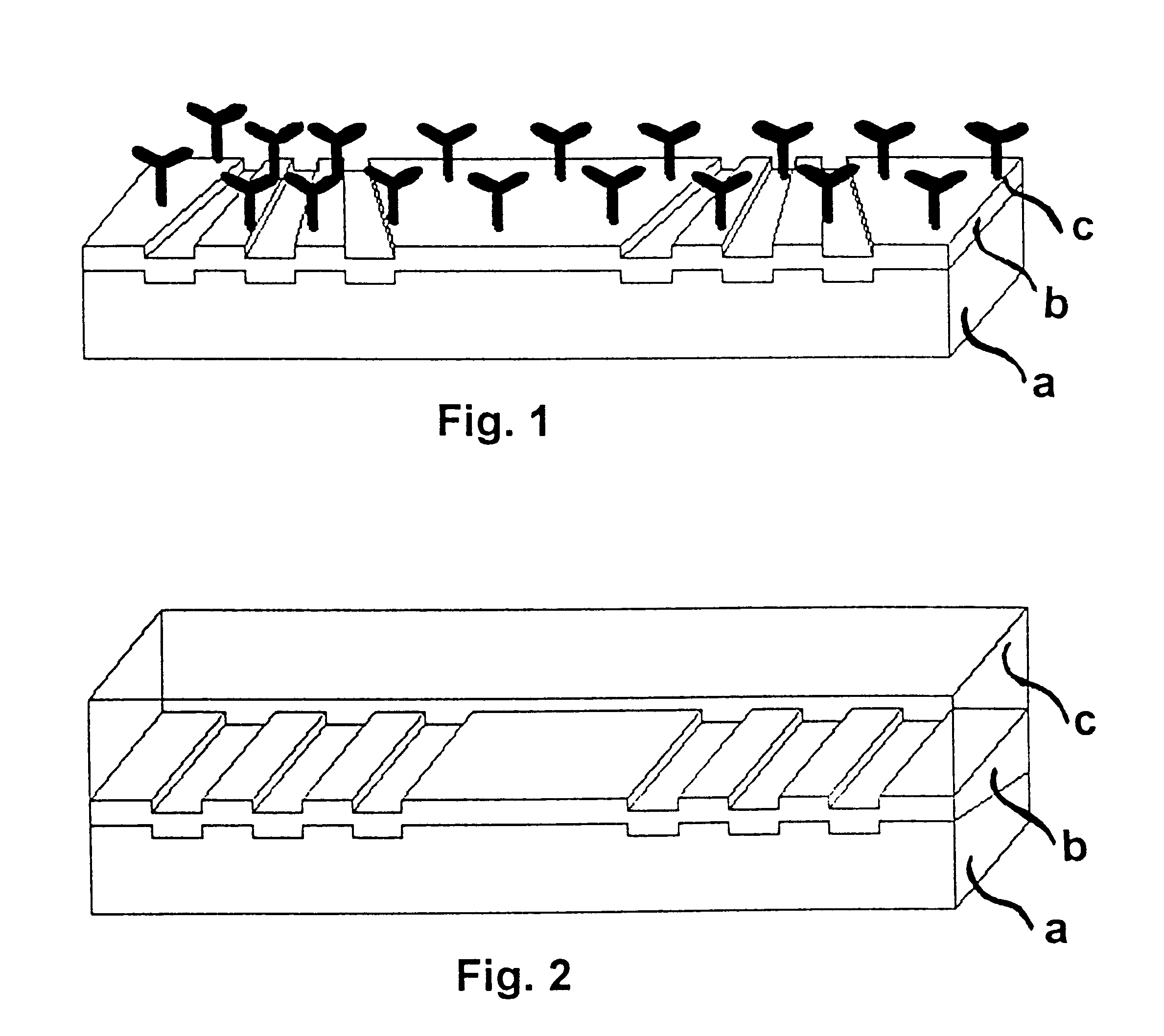
The invention relates to a planar ATR and evanescently exited luminescence optical sensor platform, consisting of a transducer and a recognition layer, wherein changes in the effective refractive index of the recognition layer are converted into a measurable variable in accordance with the integrated-optical light pointer principle. The invention relates also to the use of the method and to the method itself using the sensor platform, for example in label-free biosensory analysis.
Owner:ZEPTOSENS
Microelectronic device and method for label-free detection and quantification of biological and chemical molecules
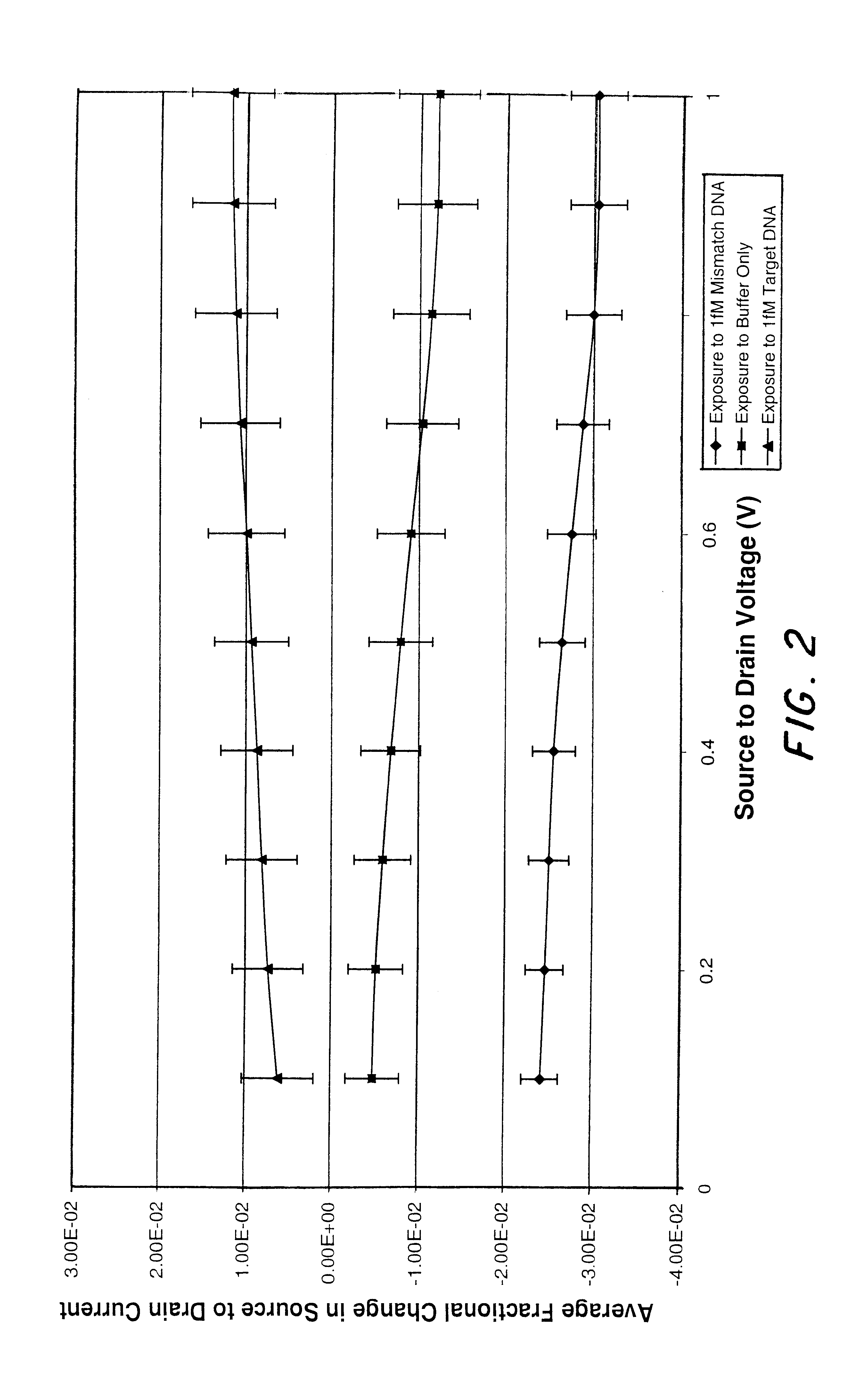
InactiveUS6482639B2Sensitive and accurate detectionWide scope of practical and worthwhile utilizationBioreactor/fermenter combinationsBiological substance pretreatmentsCapacitanceField-effect transistor
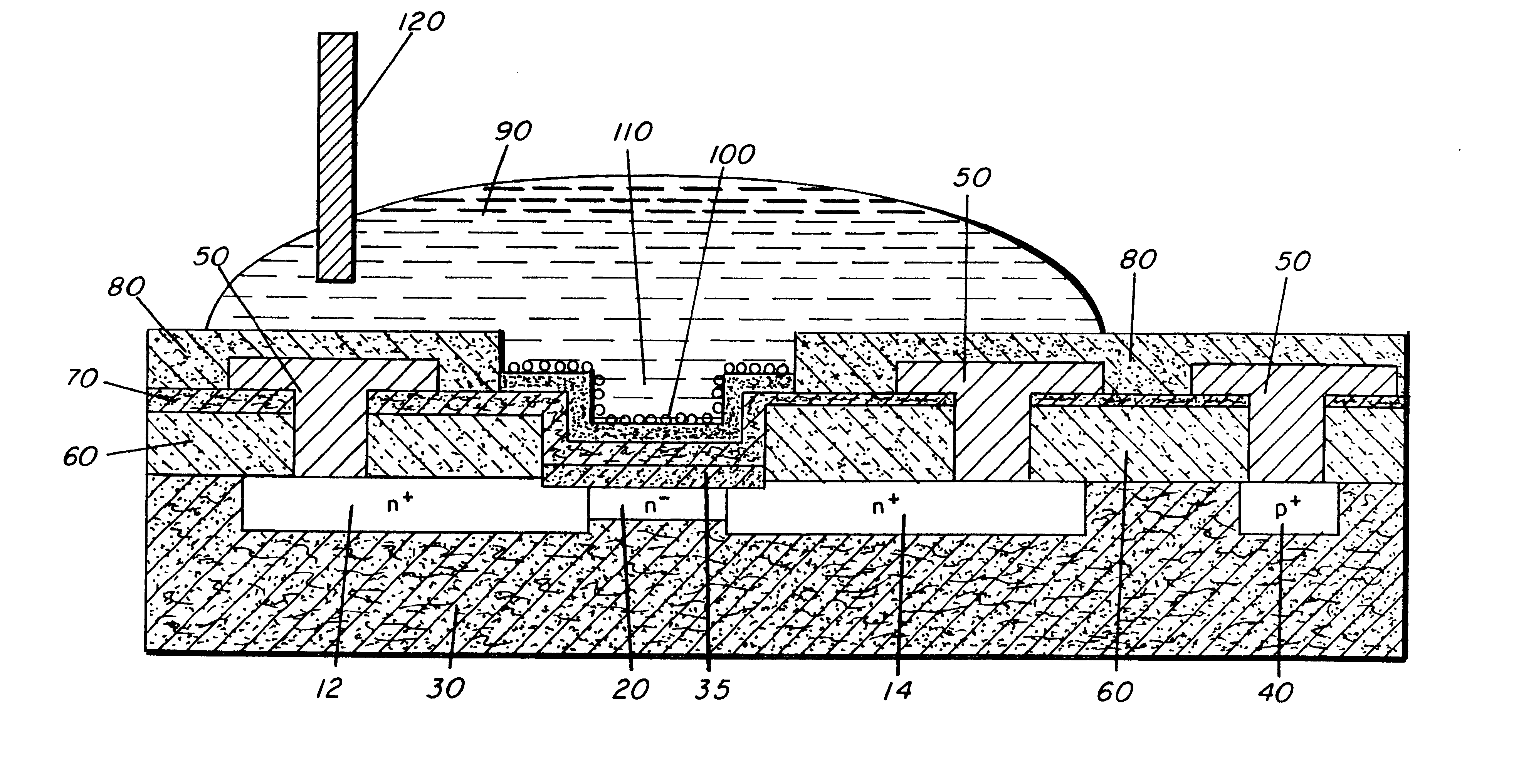
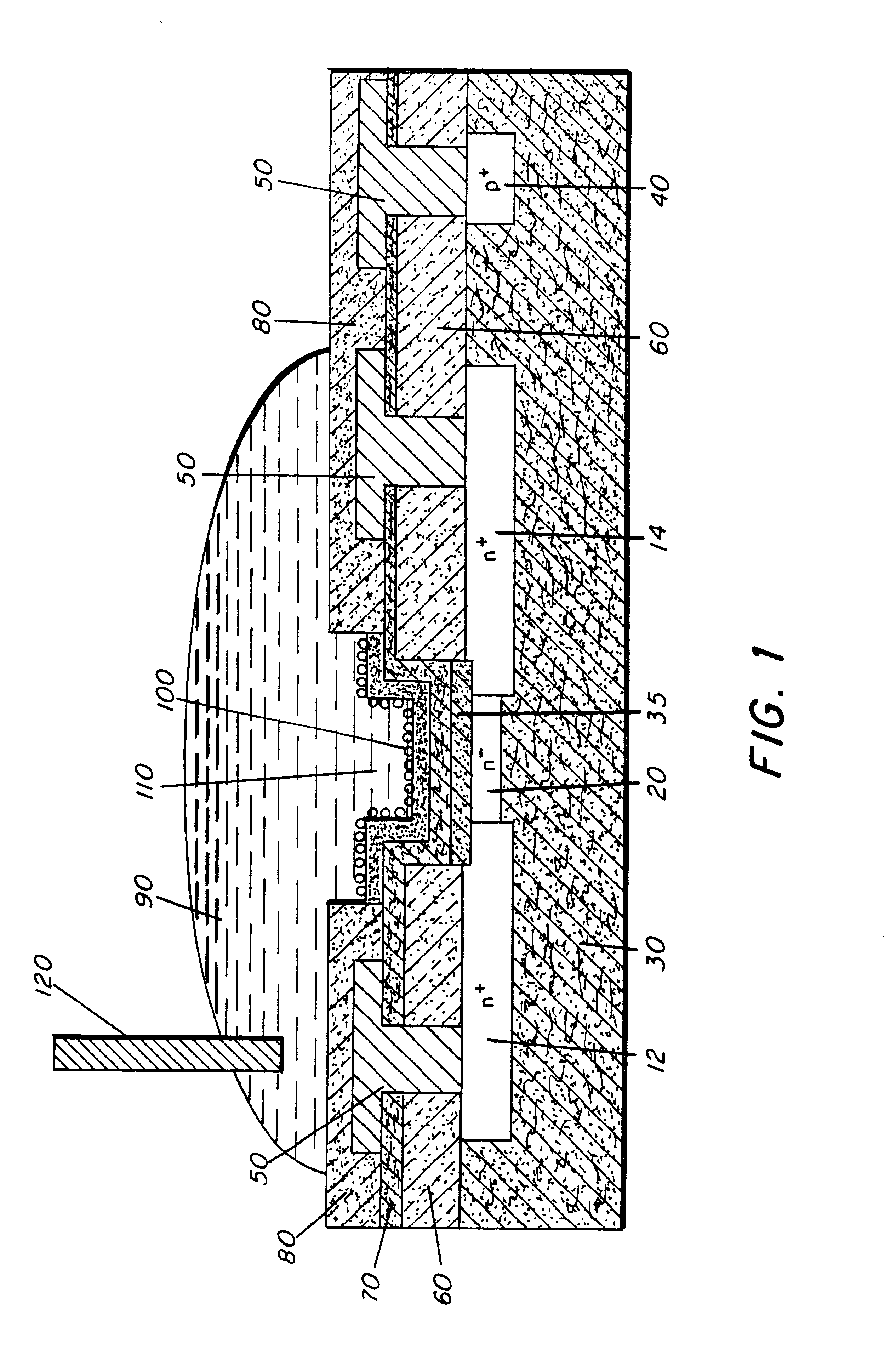
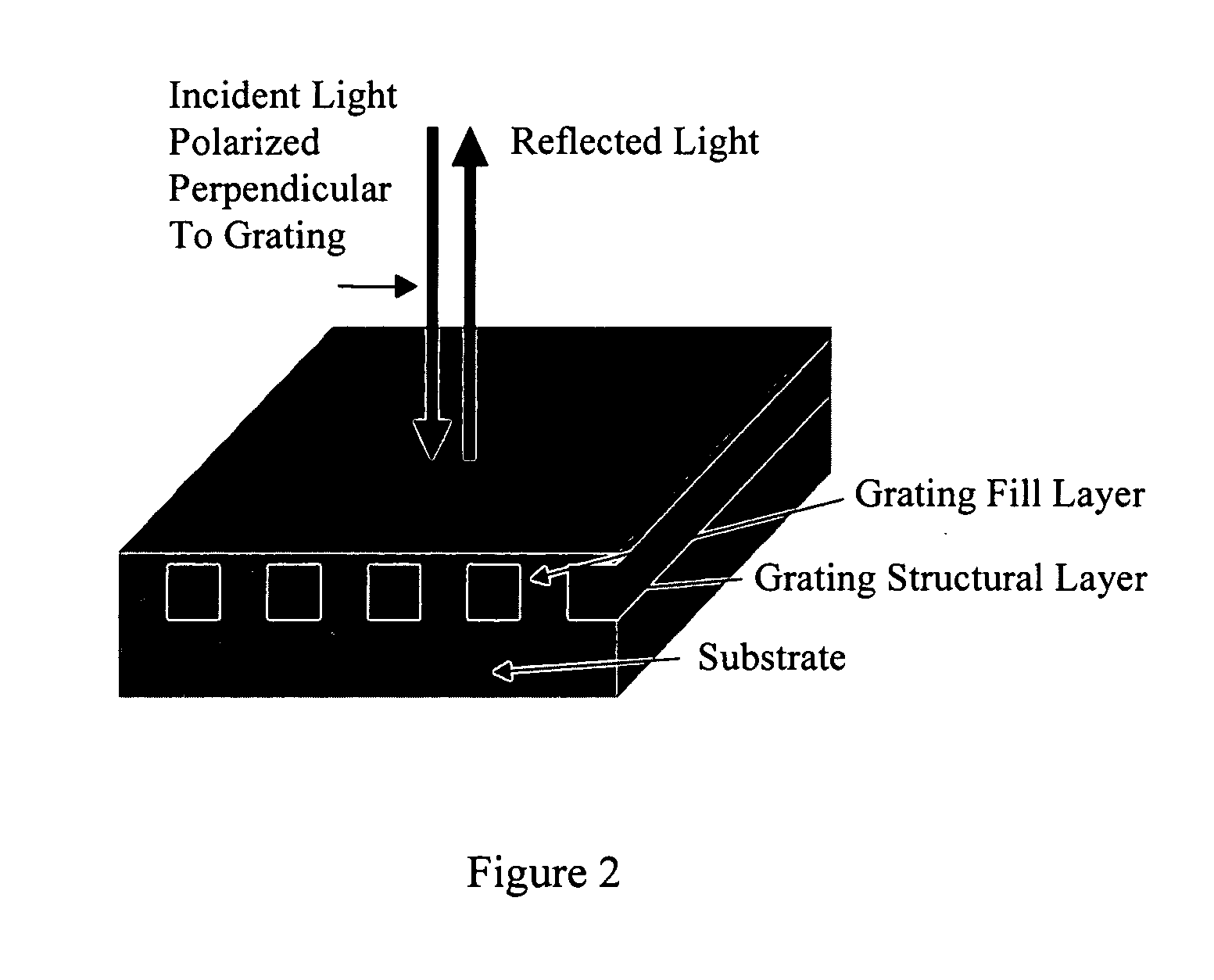
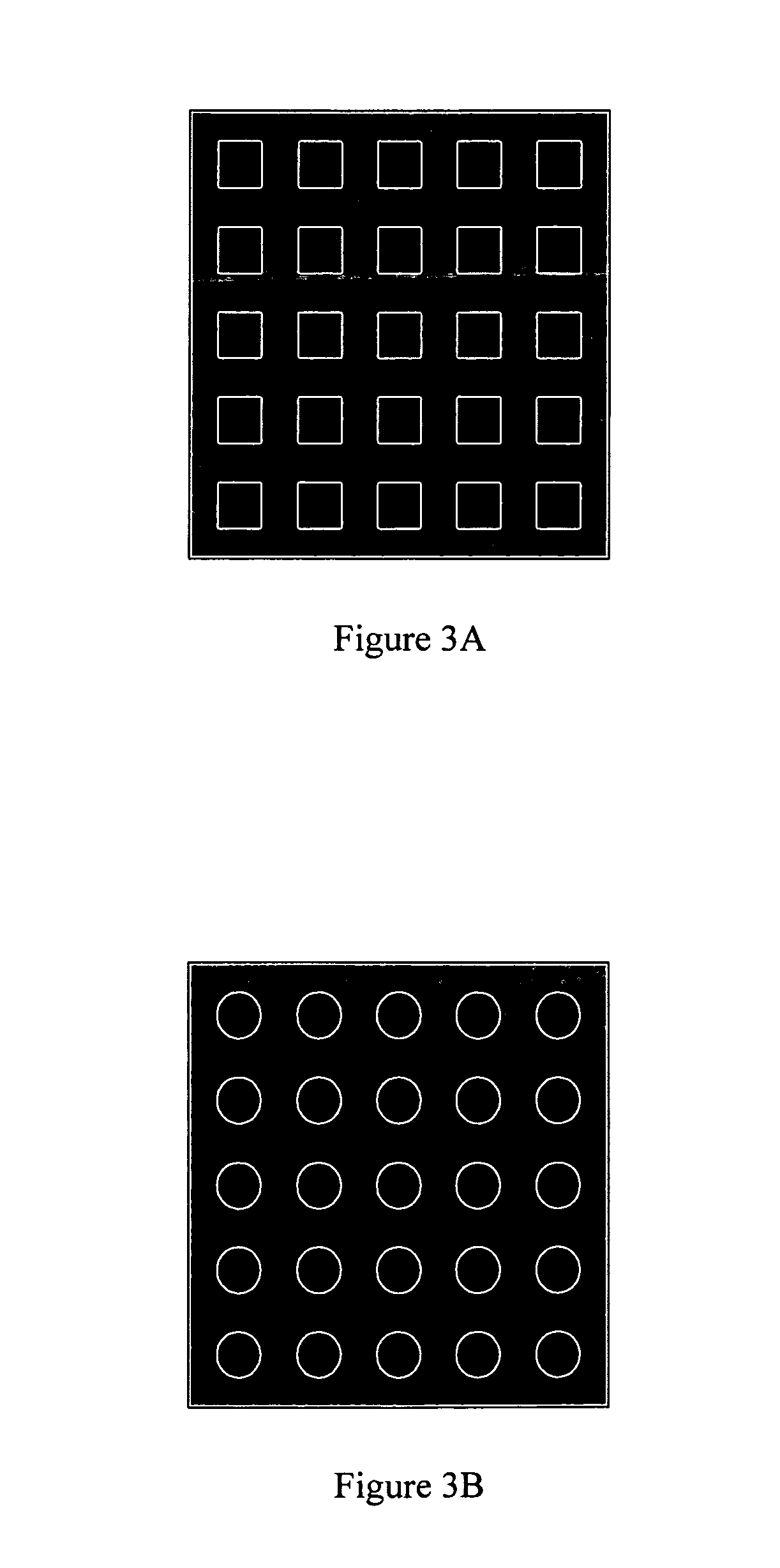
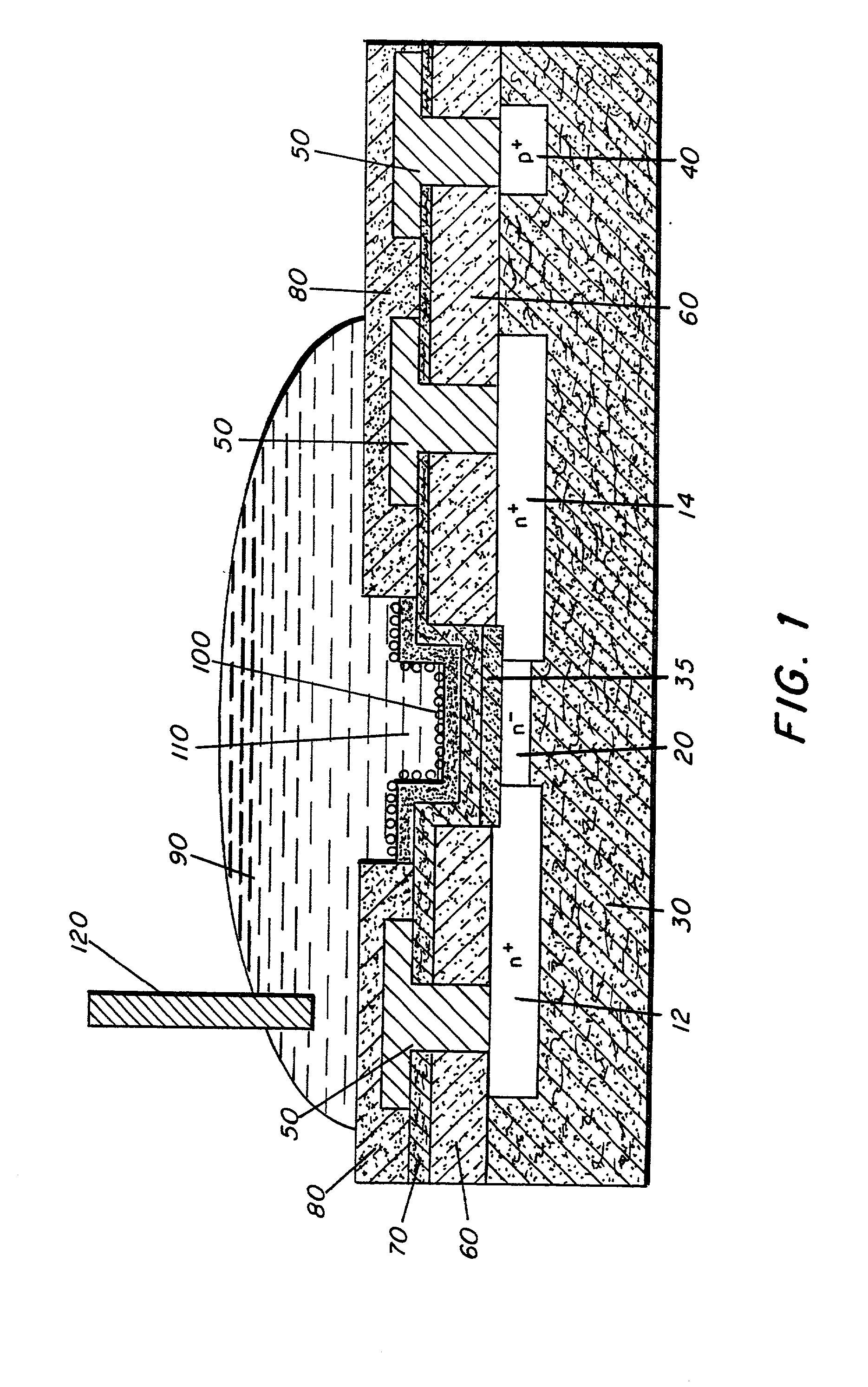
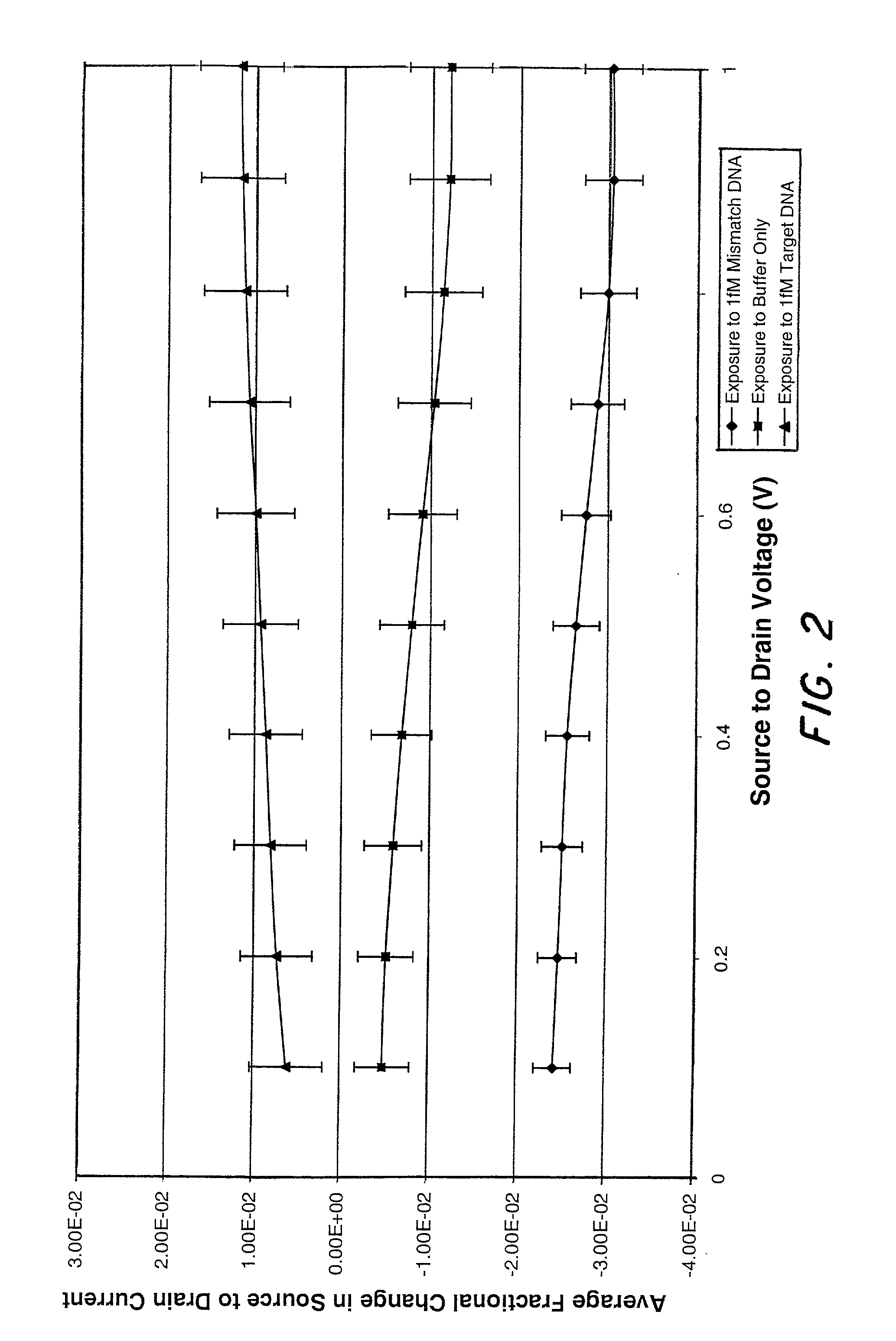
Molecular recognition-based electronic sensor, which is gateless, depletion mode field effect transistor consisting of source and drain diffusions, a depletion-mode implant, and insulating layer chemically modified by immobilized molecular receptors that enables miniaturized label-free molecular detection amenable to high-density array formats. The conductivity of the active channel modulates current flow through the active channel when a voltage is applied between the source and drain diffusions. The conductivity of the active channel is determined by the potential of the sample solution in which the device is immersed and the device-solution interfacial capacitance. The conductivity of the active channel modulates current flow through the active channel when a voltage is applied between the source and drain diffusions. The interfacial capacitance is determined by the extent of occupancy of the immobilized receptor molecules by target molecules. Target molecules can be either charged or uncharged. Change in interfacial capacitance upon target molecule binding results in modulation of an externally supplied current through the channel.
Owner:THE UNITED STATES OF AMERICA AS REPRESENTED BY THE SECRETARY OF THE NAVY
Optical sensor unit and procedure for the ultrasensitive detection of chemical or biochemical analytes
InactiveUS6346376B1High sensitivitySelective retentionBioreactor/fermenter combinationsBiological substance pretreatmentsSpecific detectionNon-covalent interactions
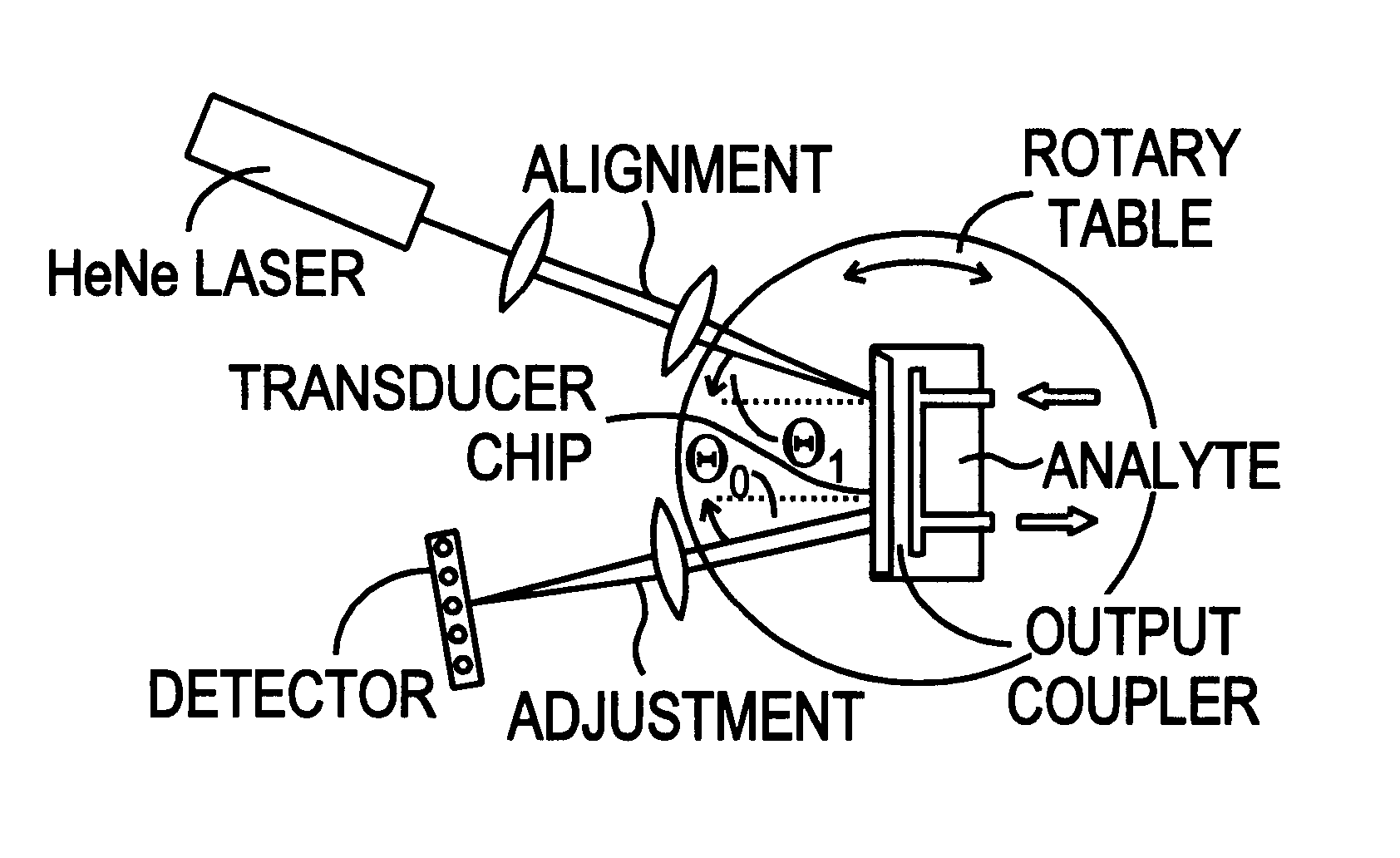
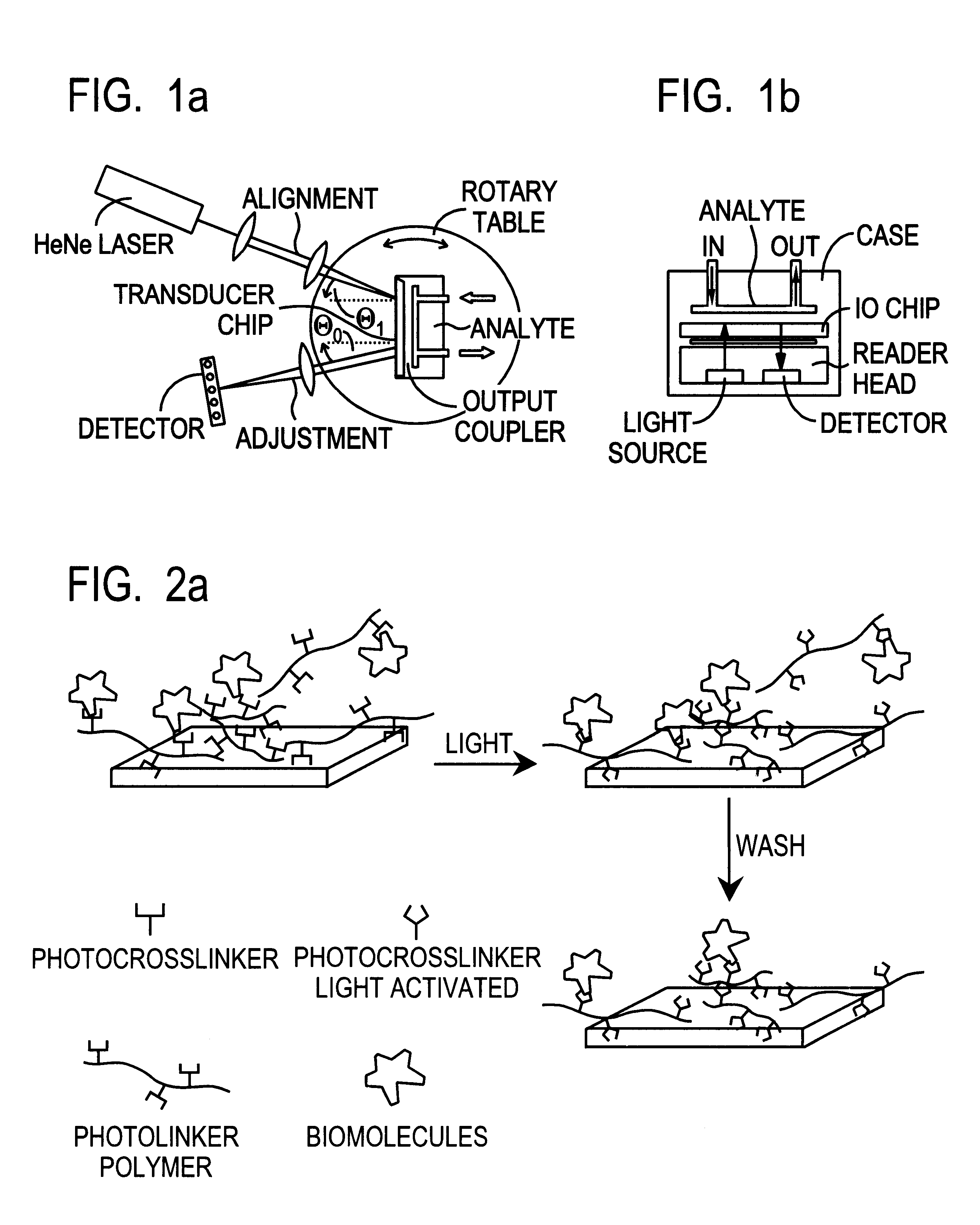
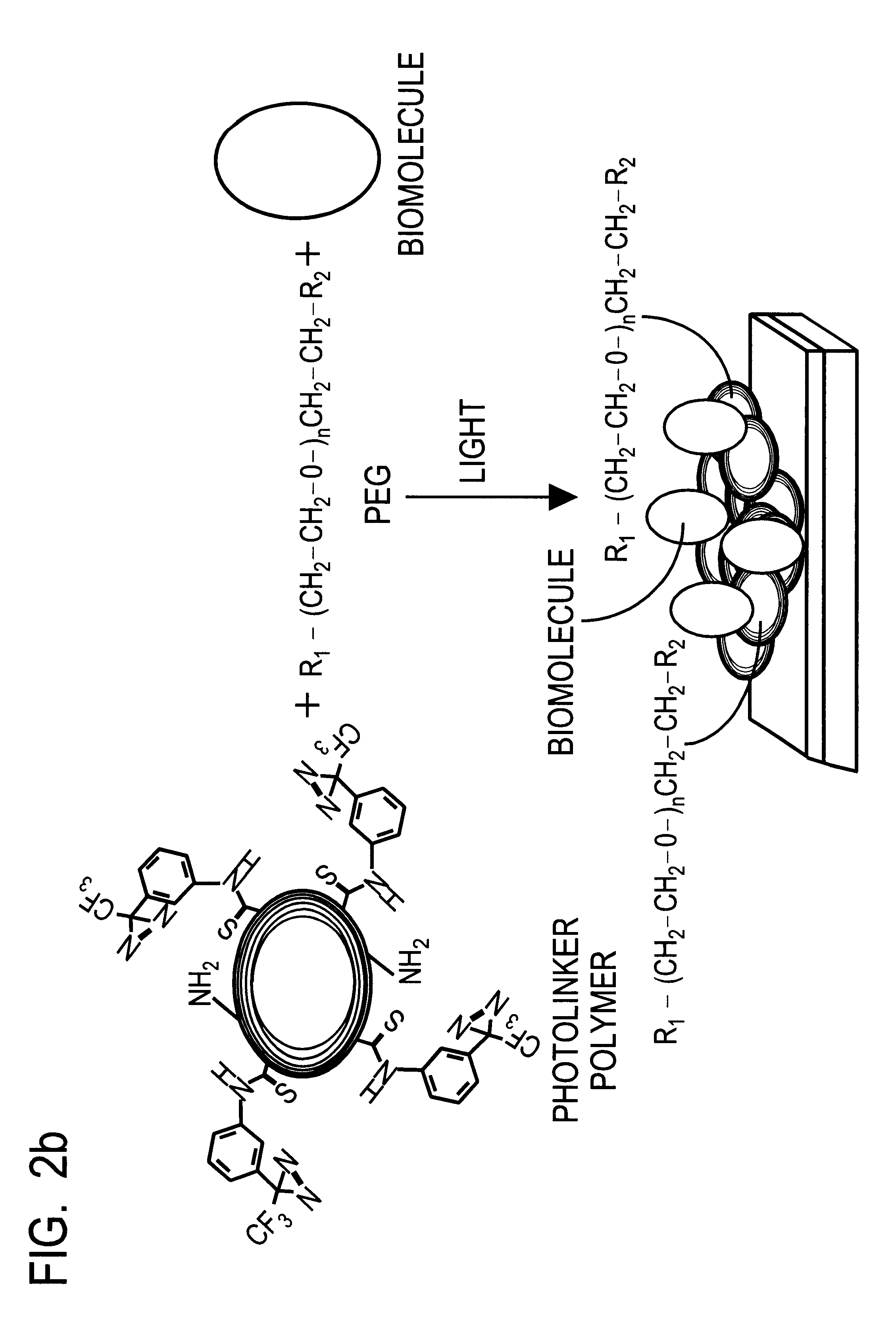
This document describes an optical sensor unit and a procedure for the specific detection and identification of biomolecules at high sensitivity in real fluids and tissue homogenates. High detection limits are reached by the combination of i) label-free integrated optical detection of molecular interactions, ii) the use of specific bioconstituents for sensitive detection and iii) planar optical transducer surfaces appropriately engineered for suppression of non-specific binding, internal referencing and calibration. Applications include the detection of prion proteins and identification of those biomolecules which non-covalently interact with surface immobilized prion proteins and are intrinsically involved in the cause of prion related disease.
Owner:CSEM CENT SUISSE DELECTRONIQUE & DE MICROTECHNIQUE SA RECH & DEV
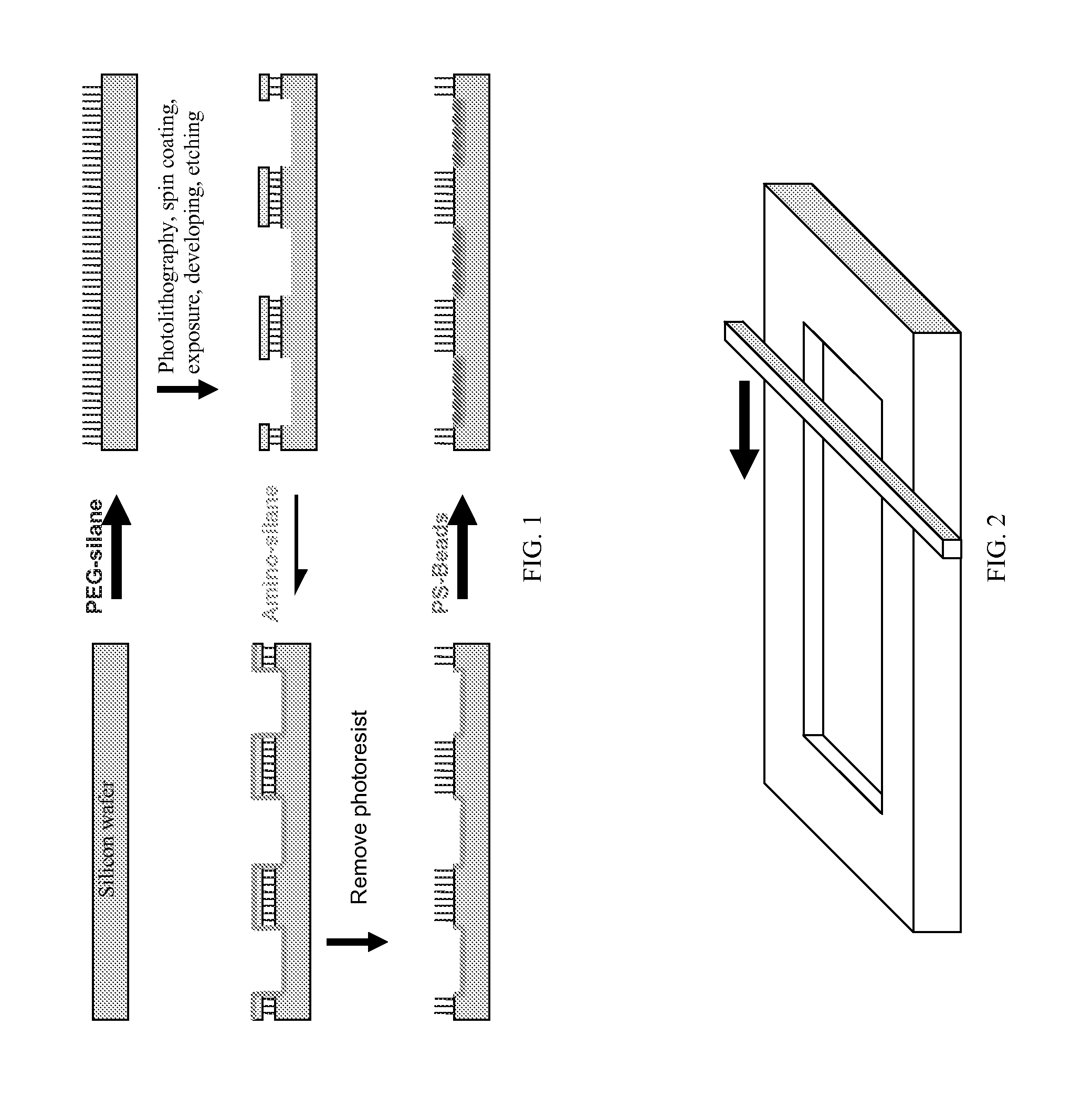
Structured Substrates for Optical Surface Profiling
ActiveUS20090011948A1Increase heightQuick changePeptide librariesNucleotide librariesOptical surfaceBiology
This disclosure provides methods and devices for the label-free detection of target molecules of interest. The principles of the disclosure are particularly applicable to the detection of biological molecules (e.g., DNA, RNA, and protein) using standard SiO2-based microarray technology.
Owner:TRUSTEES OF BOSTON UNIV
Optical detection of label-free biomolecular interactions using microreplicated plastic sensor elements
InactiveUS20060057707A1Inexpensively incorporatedHigh-throughput screeningBioreactor/fermenter combinationsBiological substance pretreatmentsHigh fluxLight filter
Methods and compositions are provided for detecting biomolecular interactions. The use of labels is not required and the methods can be performed in a high-throughput manner. The invention also provides optical devices useful as narrow band filters.
Owner:X BODY
Microelectronic device and method for label-free detection and quantification of biological and chemical molecules
InactiveUS20020012937A1Wide scope of practicalWide scope of worthwhile utilizationBioreactor/fermenter combinationsBiological substance pretreatmentsCapacitanceField-effect transistor
Molecular recognition-based electronic sensor, which is gateless, depletion mode field effect transistor consisting of source and drain diffusions, a depletion-mode implant, and insulating layer chemically modified by immobilized molecular receptors that enables miniaturized label-free molecular detection amenable to high-density array formats. The conductivity of the active channel modulates current flow through the active channel when a voltage is applied between the source and drain diffusions. The conductivity of the active channel is determined by the potential of the sample solution in which the device is immersed and the device-solution interfacial capacitance. The conductivity of the active channel modulates current flow through the active channel when a voltage is applied between the source and drain diffusions. The interfacial capacitance is determined by the extent of occupancy of the immobilized receptor molecules by target molecules. Target molecules can be either charged or uncharged. Change in interfacial capacitance upon target molecule binding results in modulation of an externally supplied current through the channel.
Owner:THE UNITED STATES OF AMERICA AS REPRESENTED BY THE SECRETARY OF THE NAVY
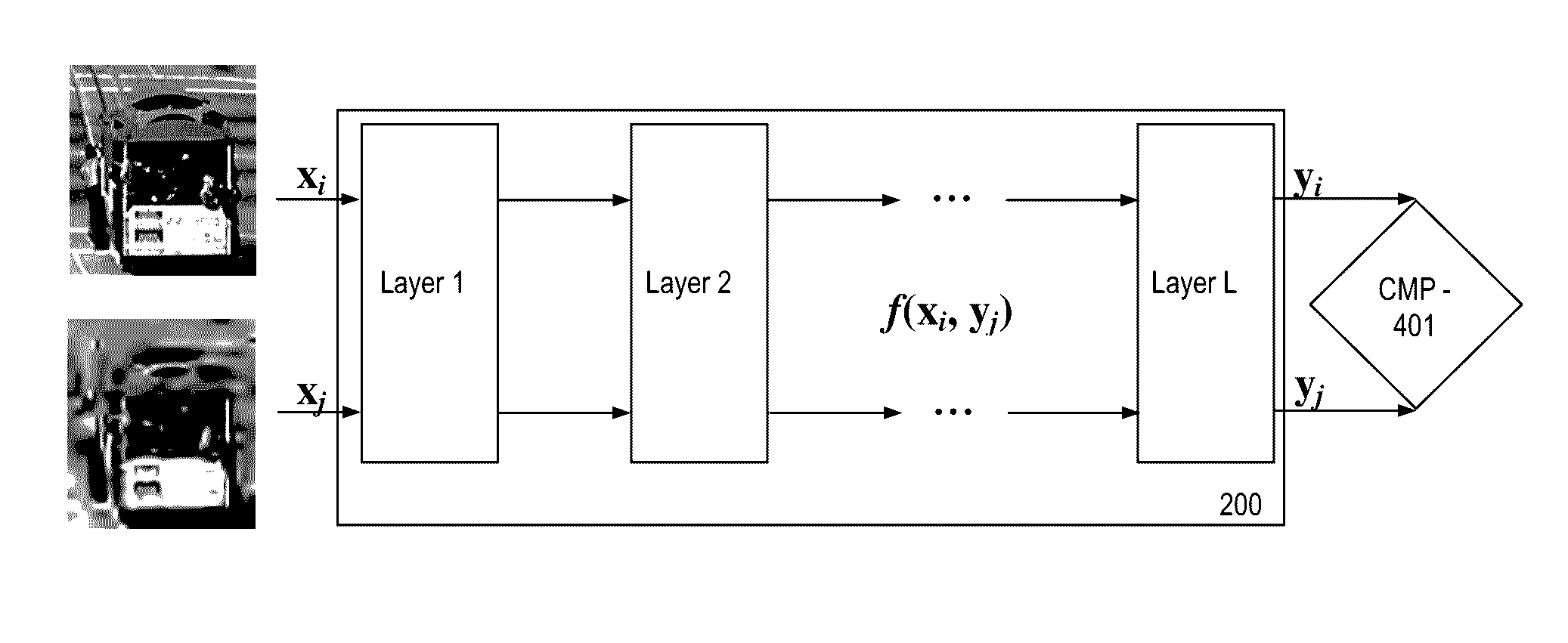
Label-free non-reference image quality assessment via deep neural network
A method for training a neural network to perform assessments of image quality is provided. The method includes: inputting into the neural network at least one set of images, each set including an image and at least one degraded version of the image; performing comparative ranking of each image in the at least one set of images; and training the neural network with the ranking information. A neural network and image signal processing tuning system are disclosed.
Owner:SAMSUNG ELECTRONICS CO LTD
Label-free high-throughput optical technique for detecting biomolecular interactions
InactiveUS20050227374A1Bioreactor/fermenter combinationsBiological substance pretreatmentsHigh fluxLight filter
Methods and compositions are provided for detecting biomolecular interactions. The use of labels is not required and the methods can be performed in a high-throughput manner. The invention also provides optical devices useful as narrow band filters.
Owner:X BODY
Signal amplified biological detection with conjugated polymers
ActiveUS20120252986A1Good water solubilityReduce non-specific assay interactionChemical structureAnalyte
The devices and method are provided for detecting labeled and label-free analytes, such as nucleic acids and proteins, employing conjugated cationic, anionic and neutral polymers. The analytes can be immobilized on a solid support material. The solid support material can be disposed in the pre-fabricated patterns on a substrate. The chemical structures of the polymers employed by the various embodiments of the present invention are described herein.
Owner:NAT UNIV OF SINGAPORE +1
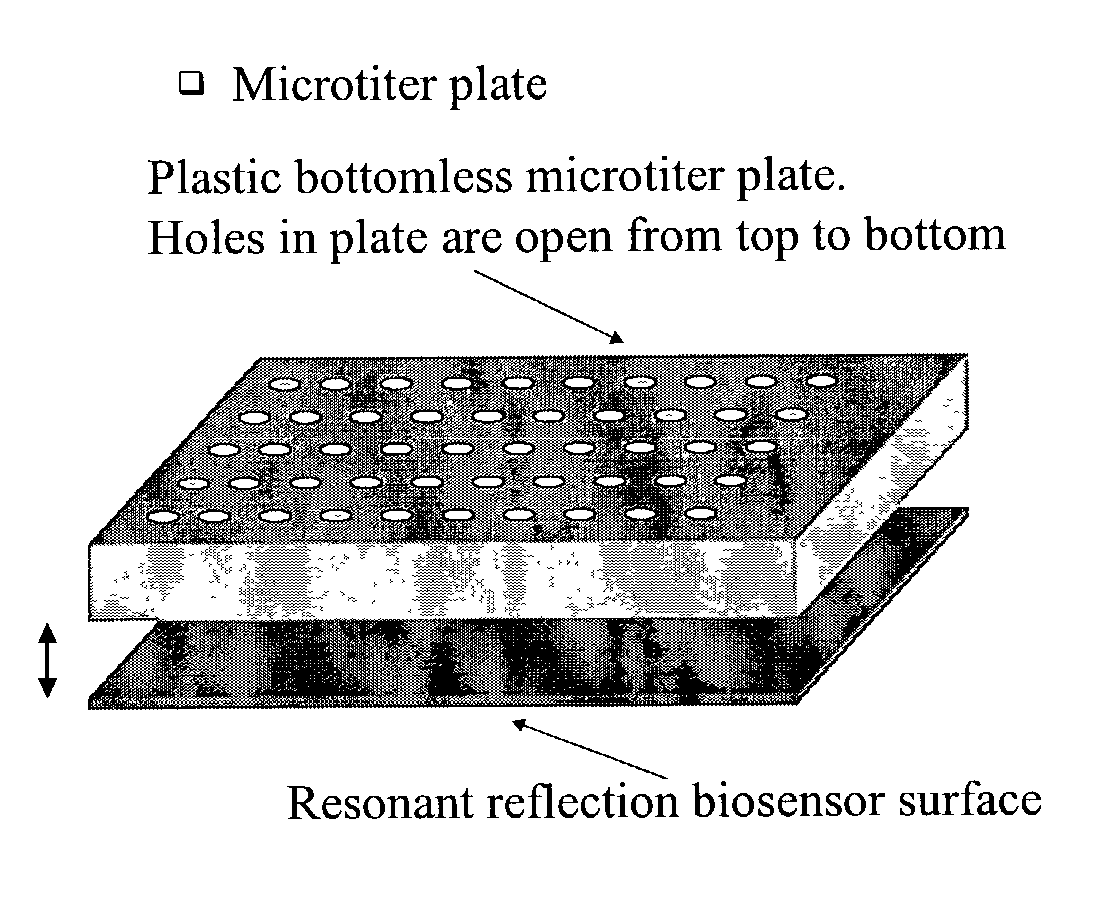
Label-free methods for performing assays using a colorimetric resonant reflectance optical biosensor
InactiveUS20070054339A1Bioreactor/fermenter combinationsBiological substance pretreatmentsAssayAnalytical chemistry
Methods are provided for detecting biomolecular interactions. The use of labels is not required and the methods can be performed in a high-throughput manner. The invention also relates to optical devices.
Owner:X BODY
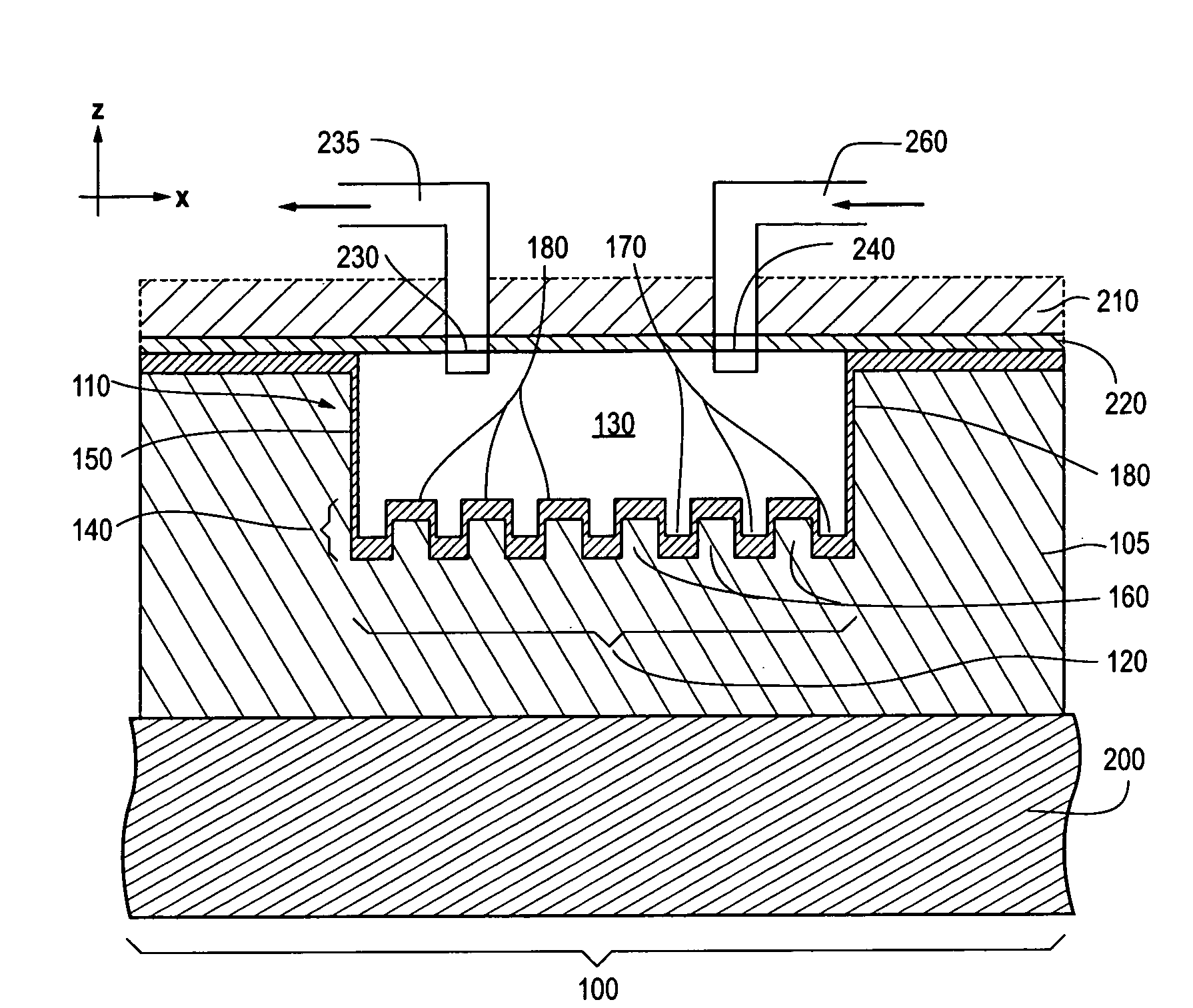
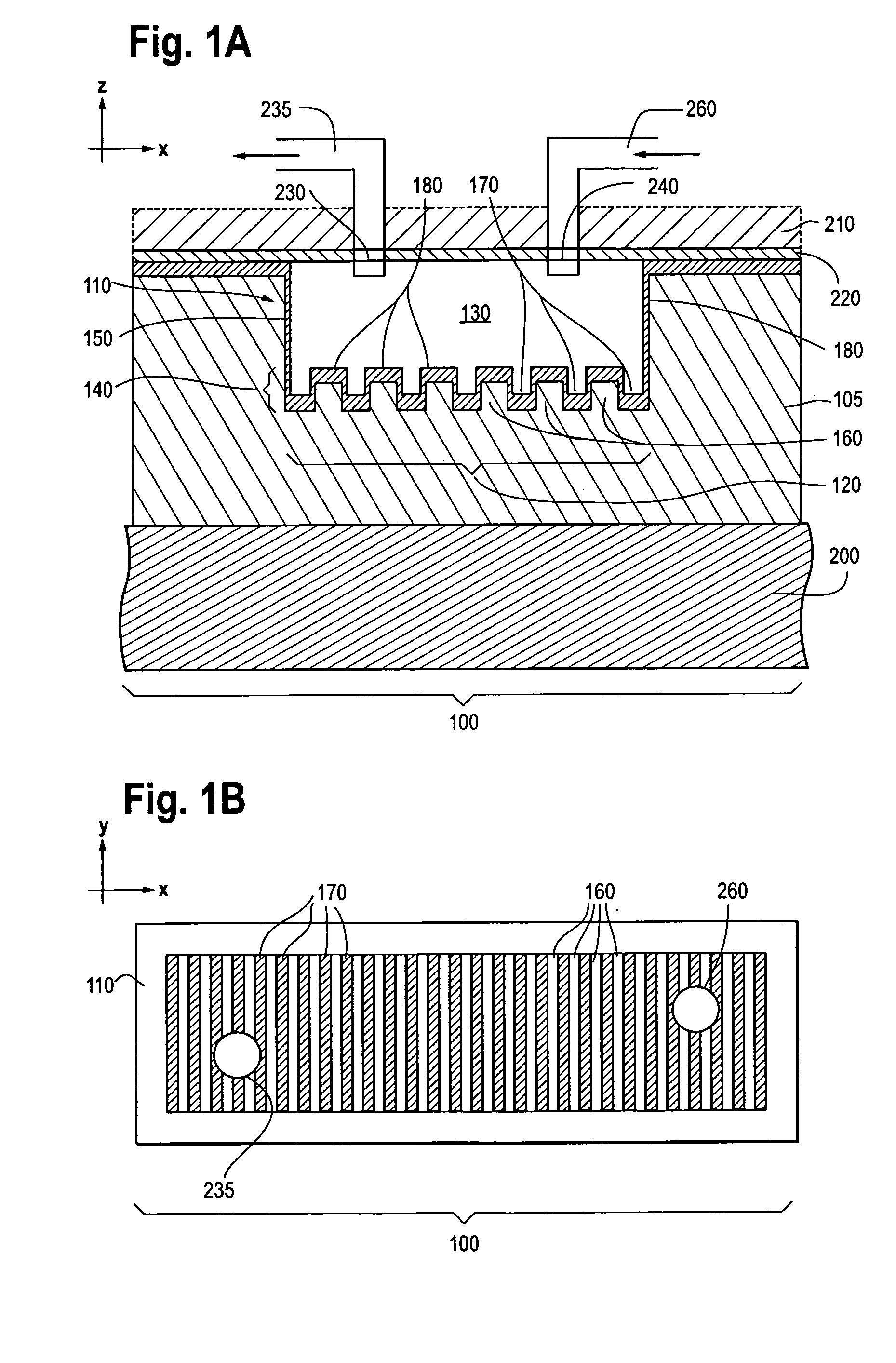
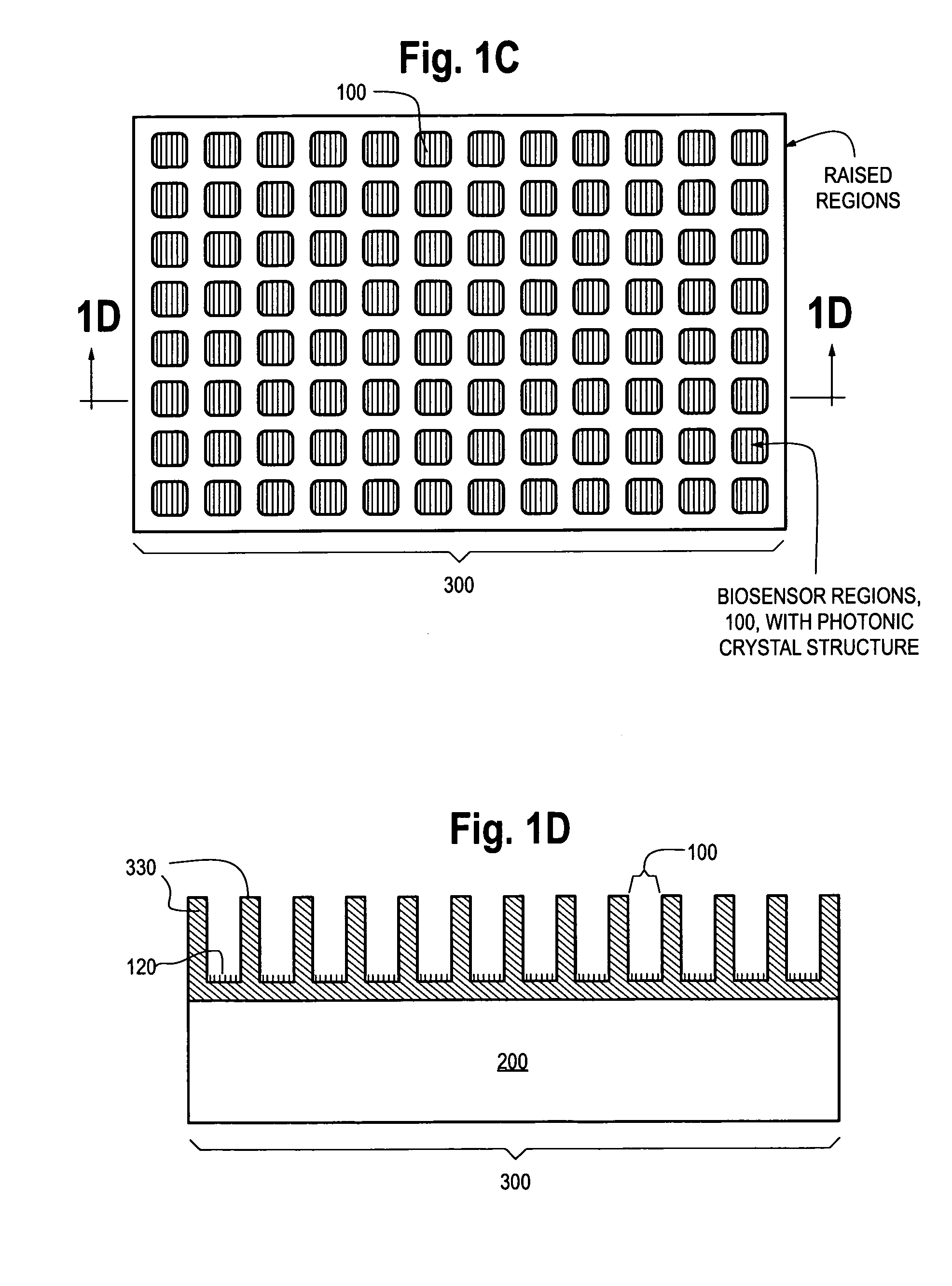
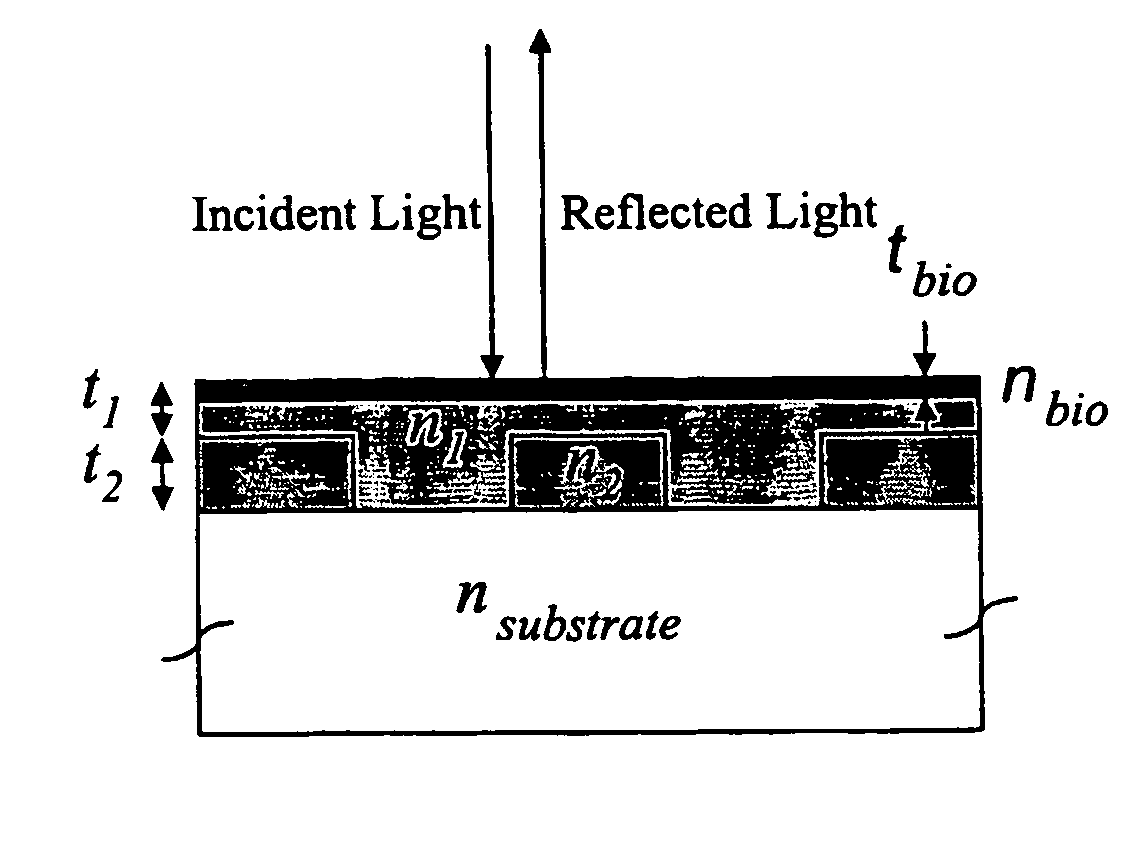
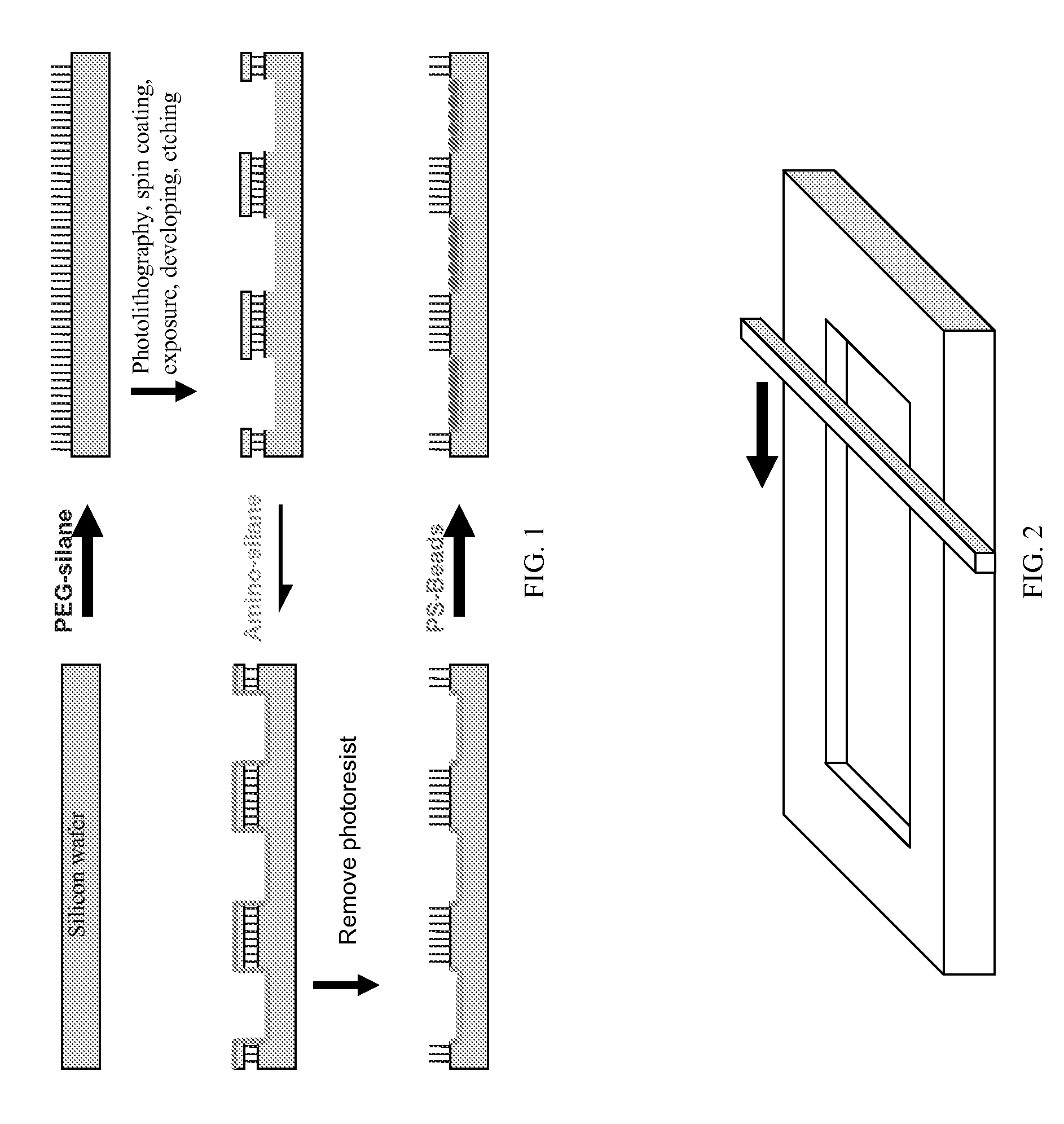
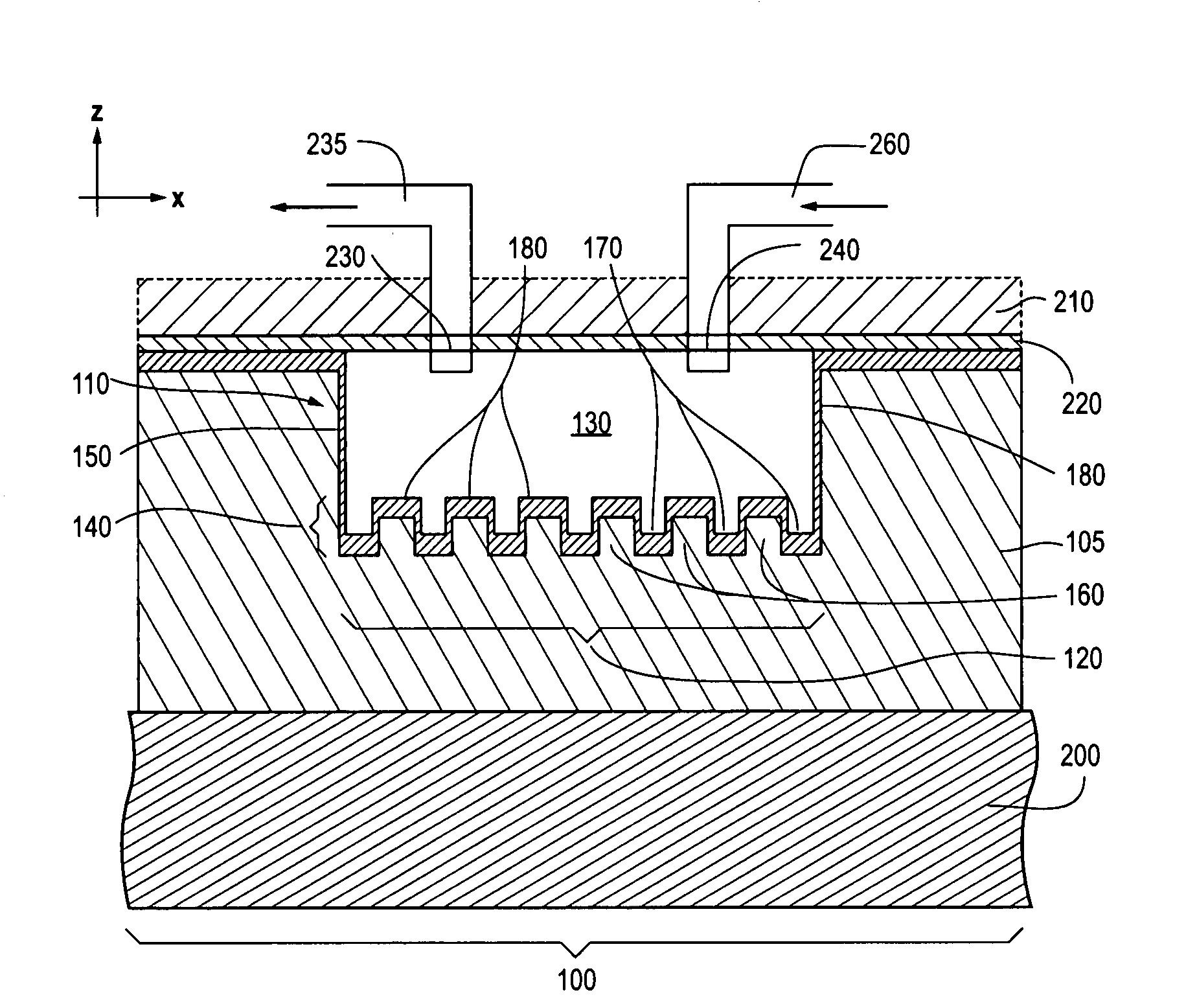
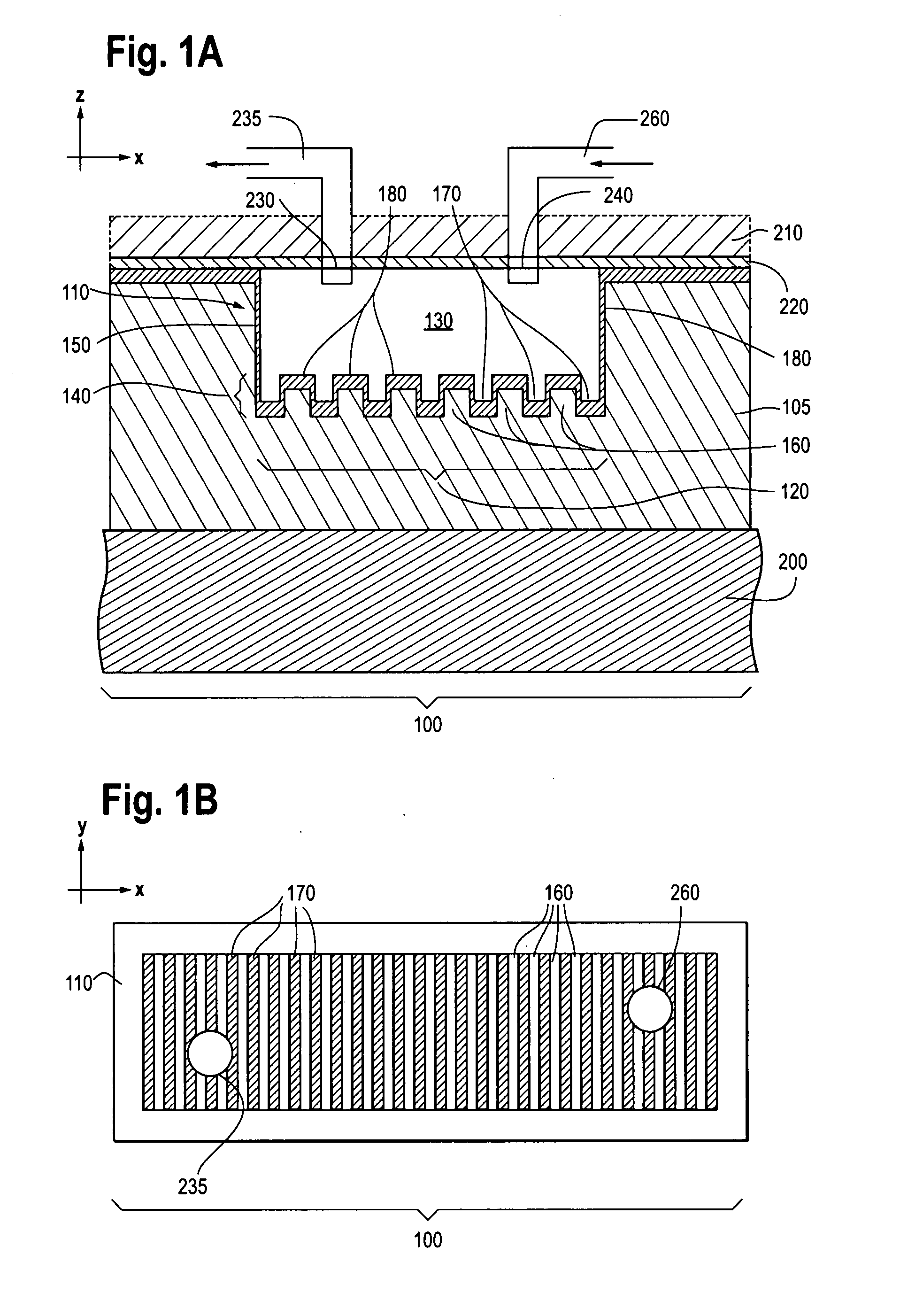
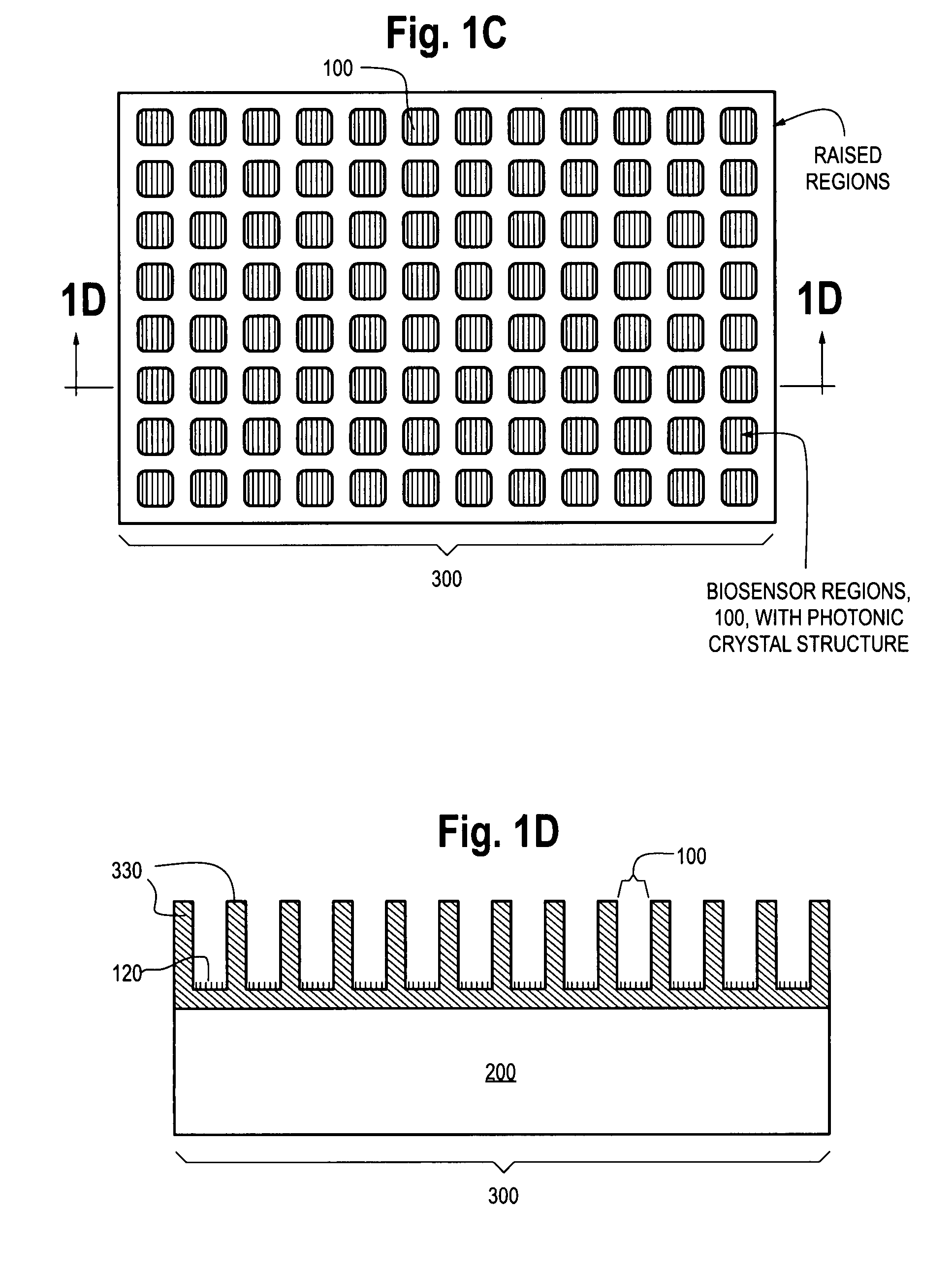
Photonic crystal sensors with integrated fluid containment structure, sample handling devices incorporating same, and uses thereof for biomolecular interaction analysis
ActiveUS20090079976A1Prevent leakageEasy to carryWithdrawing sample devicesInvestigating moving fluids/granular solidsSensor arrayPhotonic crystal structure
Photonic crystal (PC) sensors, and sensor arrays and sensing systems incorporating PC sensors are described which have integrated fluid containment and / or fluid handling structures. The PC sensors are further integrated into a sample handling device such as a microwell plate. Sensors and sensing systems of the present disclosure are capable of high throughput sensing of analytes in fluid samples, bulk refractive index detection, and label-free detection of a range of molecules, including biomolecules and therapeutic candidates. The present disclosure also provides a commercially attractive fabrication platform for making photonic crystal sensors and systems wherein an integrated fluid containment structure and a photonic crystal structure are fabricated in a single molding or imprinting processing step amendable to high throughput processing.
Owner:X BODY
Label-free high-throughput optical technique for detecting biomolecular interactions
InactiveUS7202076B2Inexpensively incorporatedHigh-throughput screeningBioreactor/fermenter combinationsBiological substance pretreatmentsMolecular interactionsThroughput
Methods and compositions are provided for detecting biomolecular interactions. The use of labels is not required and the methods can be performed in a high-throughput manner. The invention also provides optical devices useful as narrow band filters.
Owner:X BODY
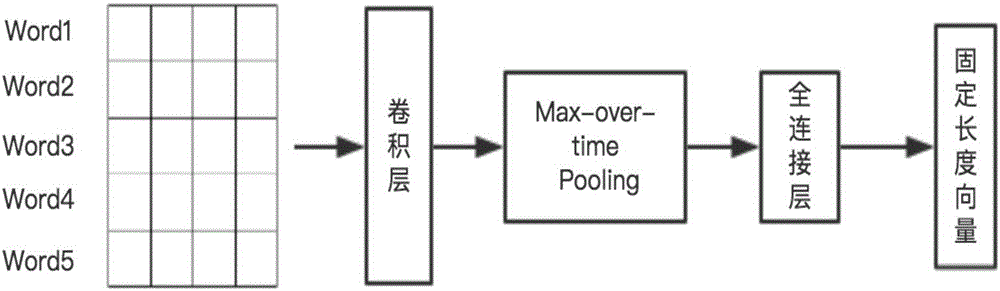
Online traditional Chinese medicine text named entity identifying method based on deep learning
InactiveCN106682220AReduce complexityReduce workloadCharacter and pattern recognitionNatural language data processingFeature extractionNamed-entity recognition
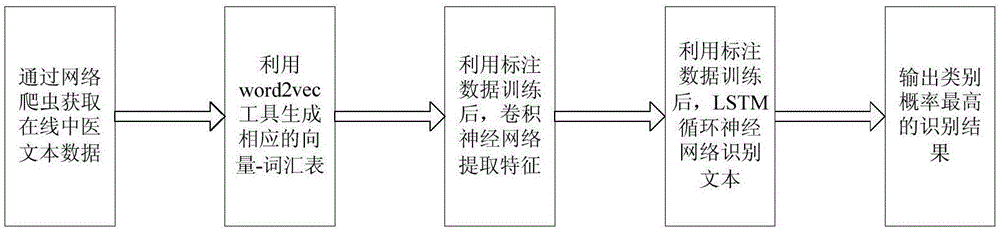
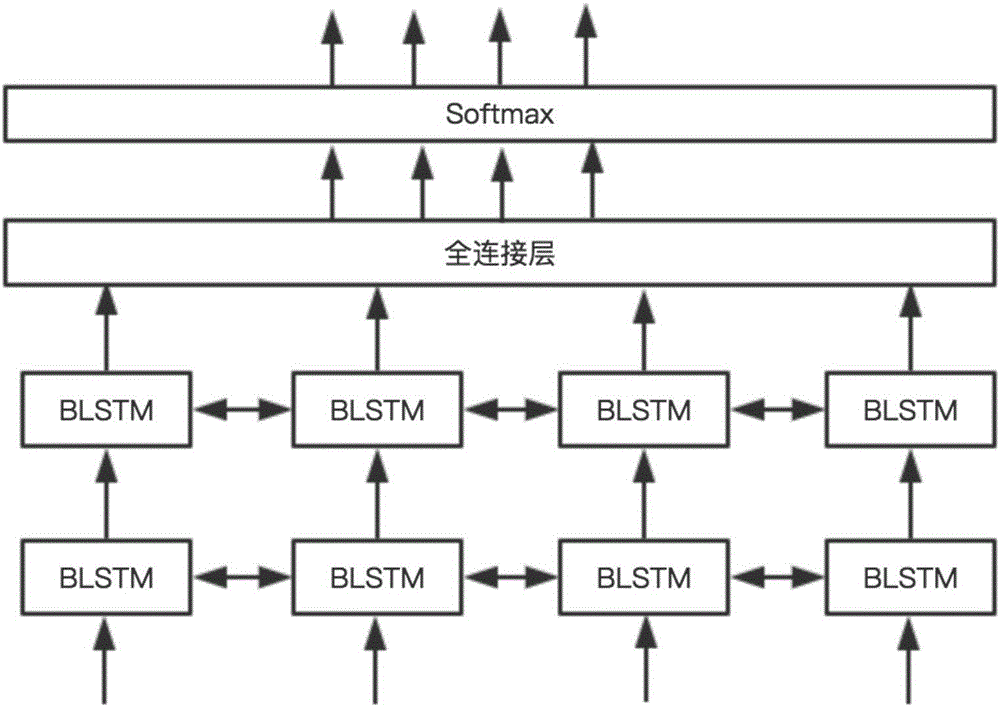
The invention discloses an online traditional Chinese medicine text named entity identifying method based on deep learning. The method includes the steps that online traditional Chinese medicine text data are obtained through a web crawler, and named entities of the obtained online traditional Chinese medicine text data are labeled with existing terminological dictionaries and human assistance; a word2vec tool is used for carrying out learning on large-scale label-free linguistic data, and word vectors with fixed length are obtained and used for forming a corresponding glossary; word segmentation is carried out on the online traditional Chinese medicine text data, words are converted into the word vectors with the fixed length by searching for the glossary, the word vectors serve as input of a convolutional neural network, and a blank character is used for filling when sentence length is insufficient; output of the convolutional neural network serves as input of a bidirectional long-short-time memory recurrent neural network, and an identification result of the online traditional Chinese medicine text data words to be identified is output. Compared with a traditional method for named entity identifying, the method reduces complexity and workload of feature extraction, simplifies the processing process and remarkably improves identification efficiency.
Owner:SOUTH CHINA UNIV OF TECH
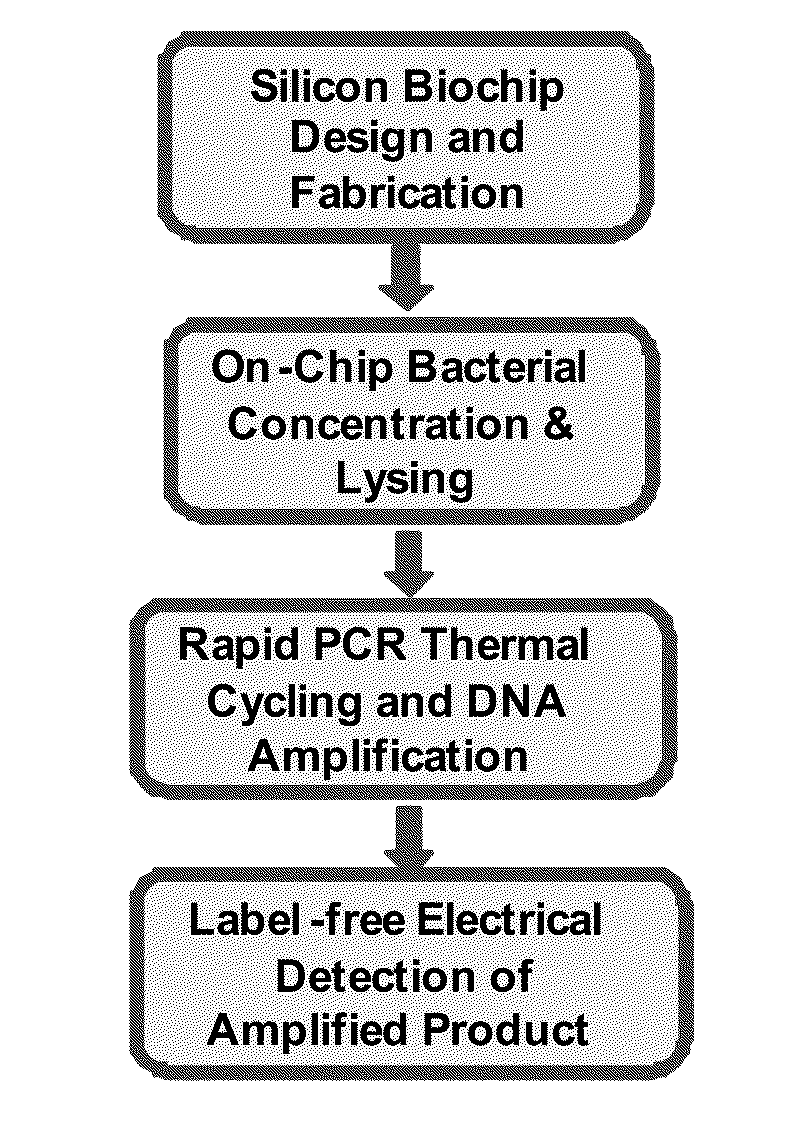
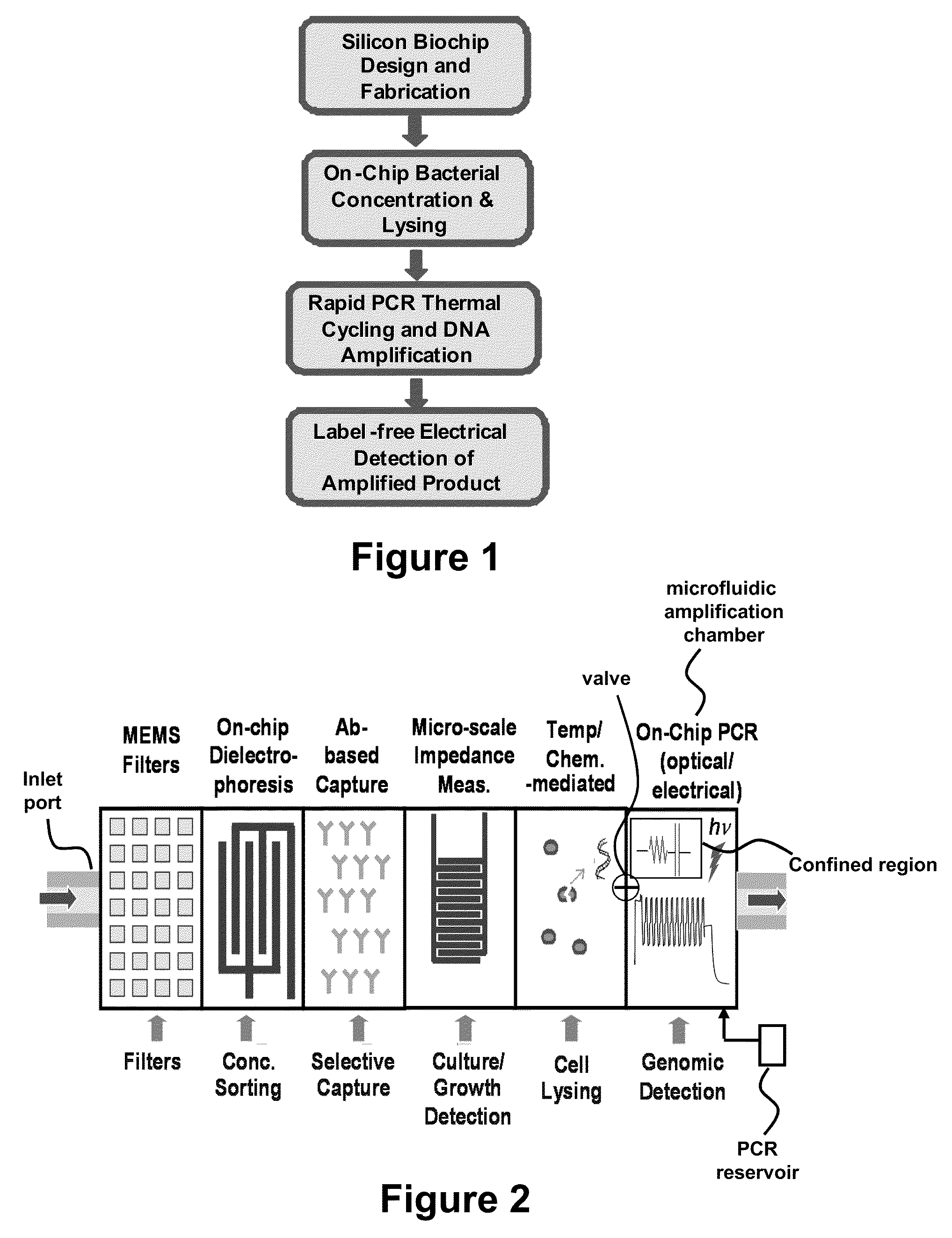
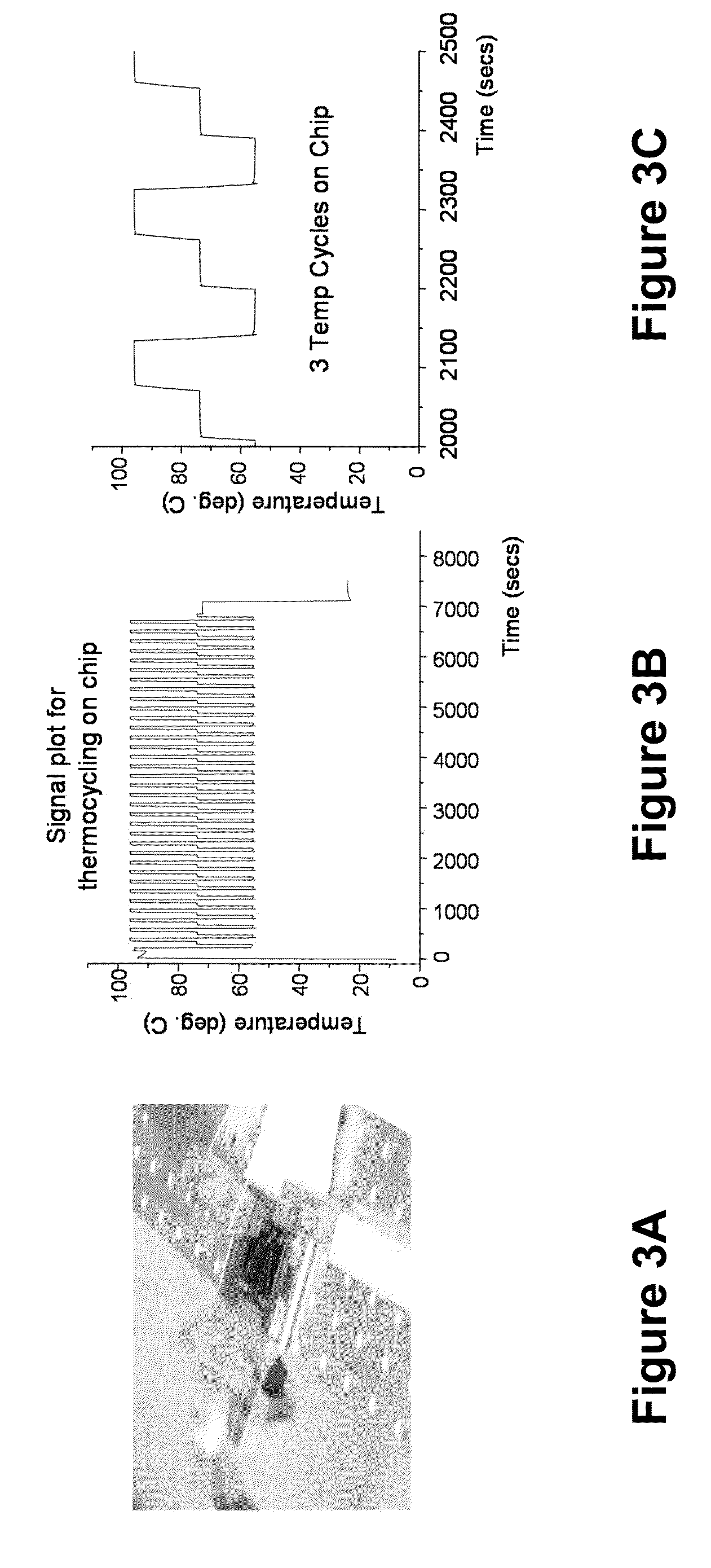
Label Free Detection of Nucleic Acid Amplification
ActiveUS20110086352A1High sensitivityIncrease detection characteristicBioreactor/fermenter combinationsHeating or cooling apparatusPoint of careBiology
Provided are methods and devices for label-free detection of nucleic acids that are amplified by polymerase chain reaction. A solution containing the components necessary for a PCR is introduced to a microfluidic amplification chamber and an electric field applied to a confined region in which PCR occurs. PCR product generated in the confined region is detected by measuring an electrical parameter that is, for example, solution impedance. The devices and methods provided herein are used, for example, in assays to detect one or more pathogens or for point-of-care tests. In an aspect, the PCR product is confined to droplets and the assay relates to detecting an electrical parameter of a flowing droplet, thereby detecting PCR product without a label. In an aspect, the PCR occurs in the droplet.
Owner:THE BOARD OF TRUSTEES OF THE UNIV OF ILLINOIS
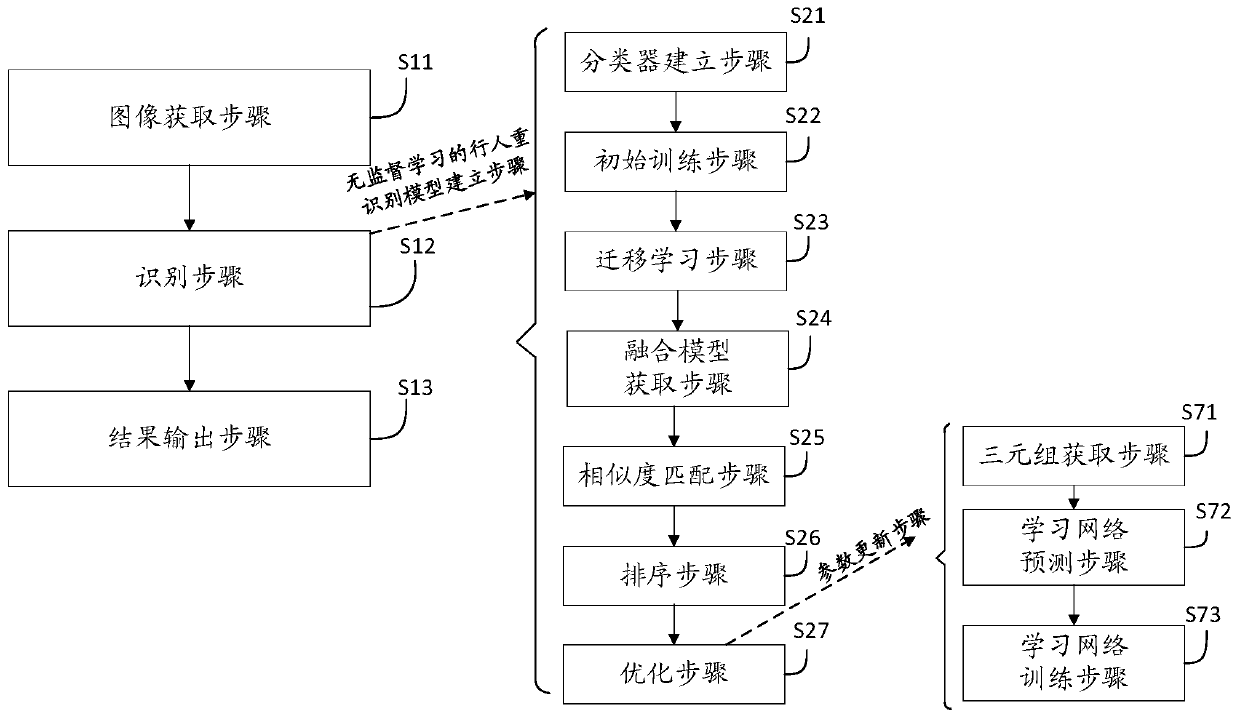
Pedestrian re-identification method and device based on unsupervised learning and medium
InactiveCN110263697AClose to realizationRealization of re-identificationBiometric pattern recognitionNeural architecturesData setSpeed learning
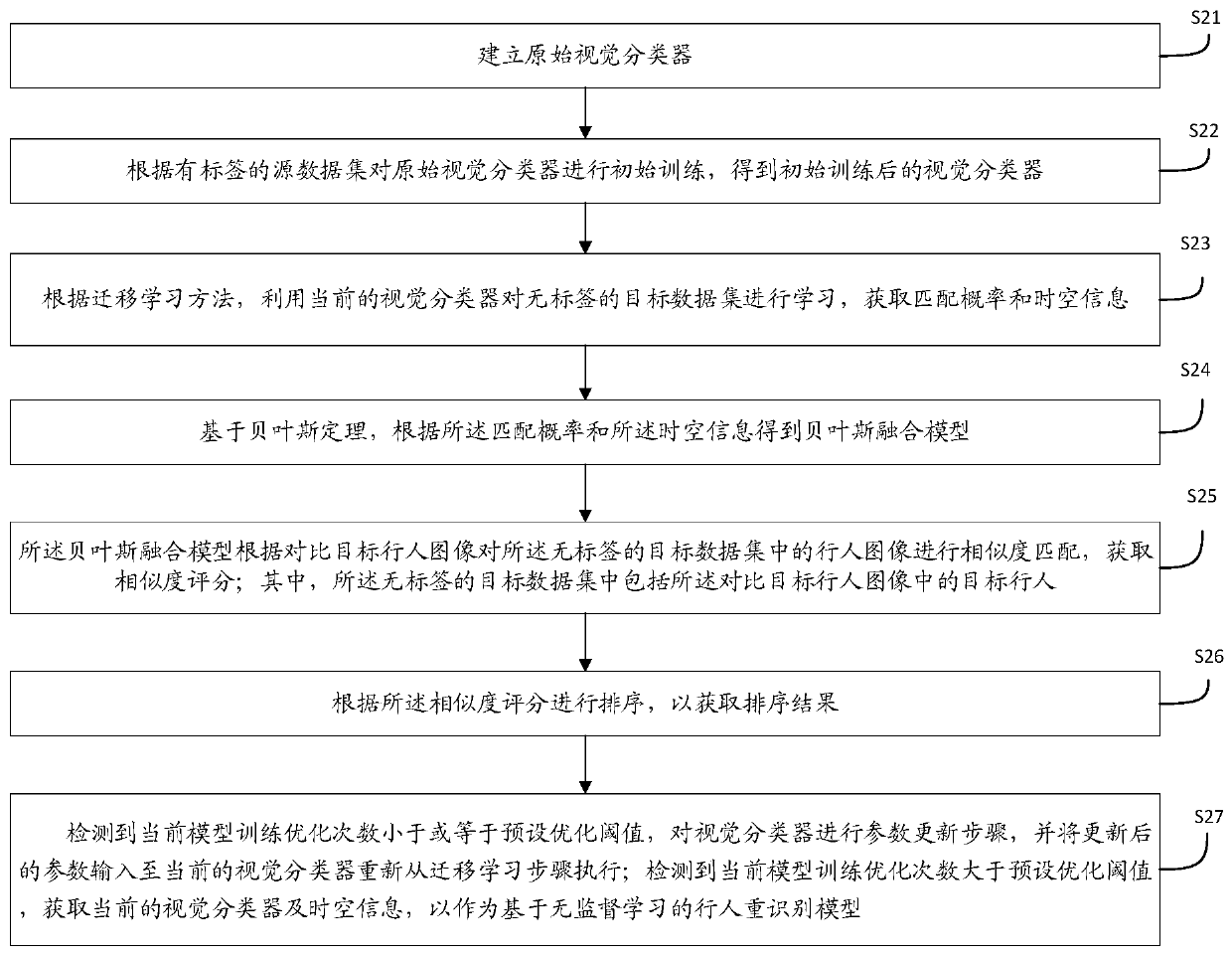
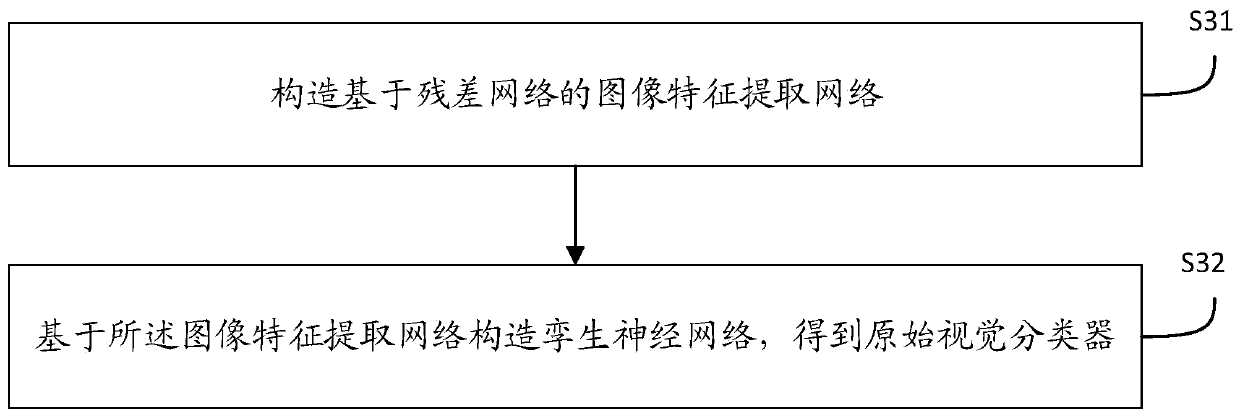
The invention discloses a pedestrian re-identification method and device based on unsupervised learning and a medium, and the method comprises the steps: obtaining a target image and a comparison image, and identifying whether a pedestrian exists in the target image in the comparison image through a pedestrian re-identification model based on unsupervised learning; outputting a recognition result; establishing a pedestrian re-identification model: carrying out initial training on the visual classifier according to the labeled source data set to obtain a visual classifier; learning the label-free target data set by using the vision classifier after initial training to obtain a matching probability and space-time information; obtaining a Bayesian fusion model according to the matching probability and the space-time information; carrying out similarity matching on pedestrian images in the unlabeled target data set by the Bayesian fusion model according to the comparison target pedestrian images to obtain a similarity score; sorting the similarity scores according to a preset threshold value to obtain a sorting result; when it is detected that the current model training optimization frequency is smaller than or equal to a preset optimization threshold value, performing parameter updating on the visual classifier.
Owner:HARBIN INST OF TECH SHENZHEN GRADUATE SCHOOL
Label-free methods for performing assays using a colorimetric resonant optical biosensor
InactiveUS7264973B2Bioreactor/fermenter combinationsBiological substance pretreatmentsReagentLabel free
The instant invention provides compositions and methods for determining cell interactions that are faster than conventional methods and that require the use of fewer reagents than conventional methods.
Owner:X BODY
Firework identification method and firework identification system based on deep learning of image
InactiveCN104408469AImproving the Speed of Unsupervised LearningFew parametersCharacter and pattern recognitionData setFireworks
The invention discloses a firework identification method and a firework identification system based on deep learning of an image. The firework identification method comprises the following steps of step 1, acquiring a label-free sample image set and a label sample image set; step 2, obtaining a label-free training data set and a label training data set; step 3, performing whitening preliminary processing on training data; step 4, based on the label-free training data subjected to the whitening preliminary processing, constructing a deep neutral network based on sparse self coding by adopting unsupervised learning, and extracting a basic image feature set of the label-free training data; step 5, convolving basic image features and pooling image data; step 6, training a Softmax classifier based on the convolved and pooled label training data set; step 7, inputting the convolved and pooled images to be identified into the trained Softmax classifier to obtain the identification result. According to the firework identification method and the firework identification system disclosed by the invention, the visual identification rate of fireworks and a similar object can be effectively improved, and automatic identification with higher precision for the fireworks can be realized.
Owner:WUHAN UNIV
Label-free methods for performing assays using a colorimetric resonant reflectance optical biosensor
InactiveUS7300803B2Bioreactor/fermenter combinationsBiological substance pretreatmentsHigh fluxMolecular interactions
Methods are provided for detecting biomolecular interactions. The use of labels is not required and the methods can be performed in a high-throughput manner. The invention also relates to optical devices.
Owner:X BODY
Label-free methods for performing assays using a colorimetric resonant reflectance optical biosensor
InactiveUS7153702B2EffectivenessBioreactor/fermenter combinationsPeptide librariesMolecular interactionsReflectivity
Methods are provided for detecting biomolecular interactions. The use of labels is not required and the methods can be performed in a high-throughput manner. The invention also relates to optical devices.
Owner:X BODY
Applications of laser-processed substrate for molecular diagnostics
Owner:EBSTEIN STEVEN M
Detection And Quantification Of Biomarkers Via A Piezoelectric Cantilever Sensor
InactiveUS20090078023A1High sensitivityShort timeMaterial analysis using sonic/ultrasonic/infrasonic wavesMicrobiological testing/measurementEscherichia coliMultiple sensor
Quantification of a target analyte is performed using a single sample to which amounts of the target analyte are added. Calibration is performed as part of quantification on the same sample. The target analyte is detectable and quantifiable using label free reagents and requiring no sample preparation. Target analytes include biomarkers such as cancer biomarkers, pathogenic Escherichia coli, single stranded DNA, and staphylococcal enterotoxin. The quantification process includes determining a sensor response of a sensor exposed to the sample and configured to detect the target analyte. Sensor responses are determined after sequential additions of the target analyte to the sample. The amount of target analyte detected by the sensor when first exposed to the sample is determined in accordance with the multiple sensor responses.
Owner:DREXEL UNIV
Packaged chip for multiplexing photonic crystal waveguide and photonic crystal slot waveguide devices for chip-integrated label-free detection and absorption spectroscopy with high throughput, sensitivity, and specificity
ActiveUS20130005606A1Most efficientPrecise positioningRadiation pyrometryColor measuring devicesSlot-waveguidePhotonic crystal
Systems and methods for chip-integrated label-free detection and absorption spectroscopy with high throughput, sensitivity, and specificity are disclosed. The invention comprises packaged chips for multiplexing photonic crystal waveguide and photonic crystal slot waveguide devices. Other embodiments are described and claimed.
Owner:OMEGA OPTICS
Label-free high-throughput optical technique for detecting biomolecular interactions
InactiveUS20060030033A1Inexpensively incorporatedHigh-throughput screeningBioreactor/fermenter combinationsBiological substance pretreatmentsHigh fluxLight filter
Methods and compositions are provided for detecting biomolecular interactions. The use of labels is not required and the methods can be performed in a high-throughput manner. The invention also provides optical devices useful as narrow band filters.
Owner:IGT +1
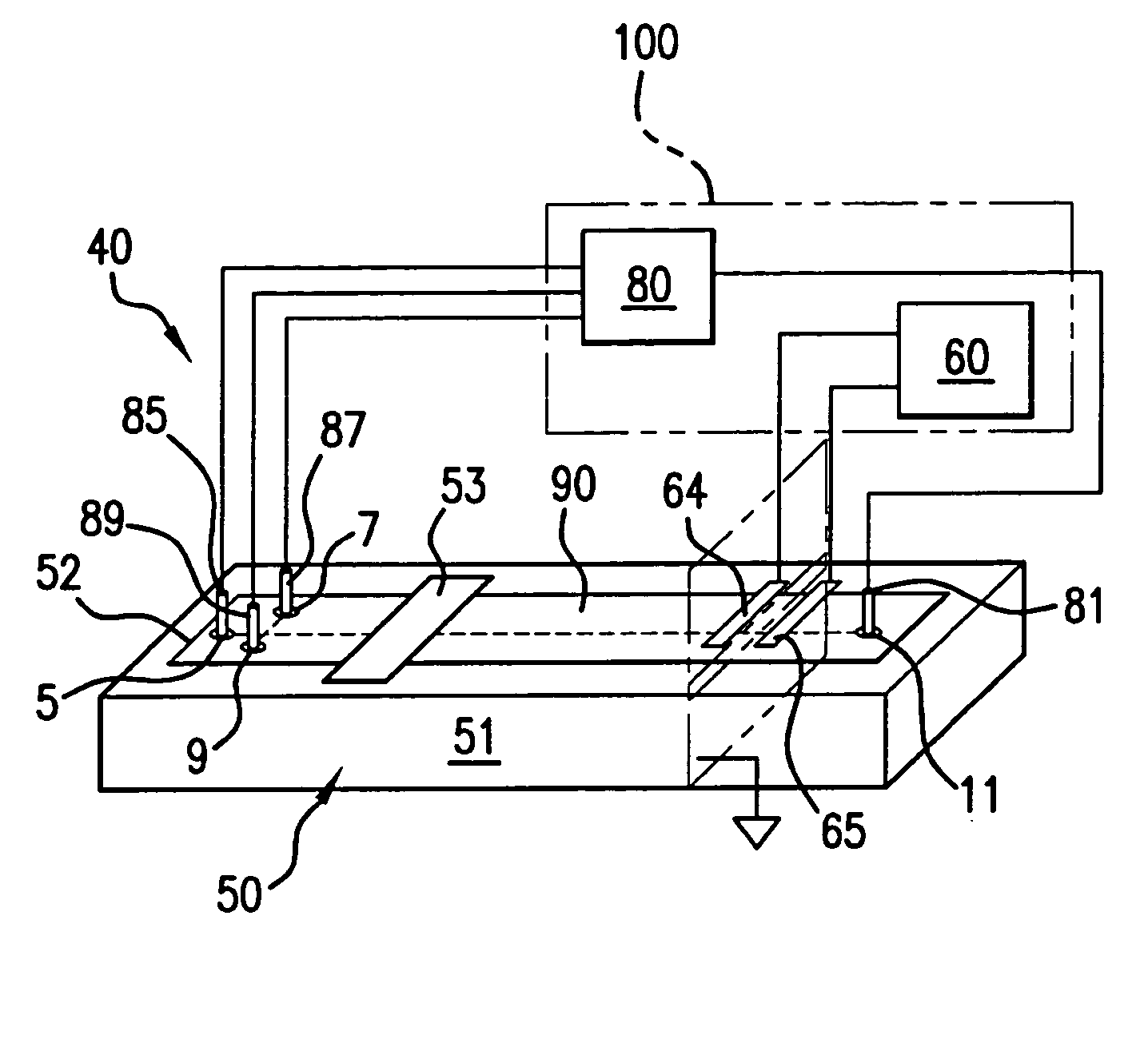
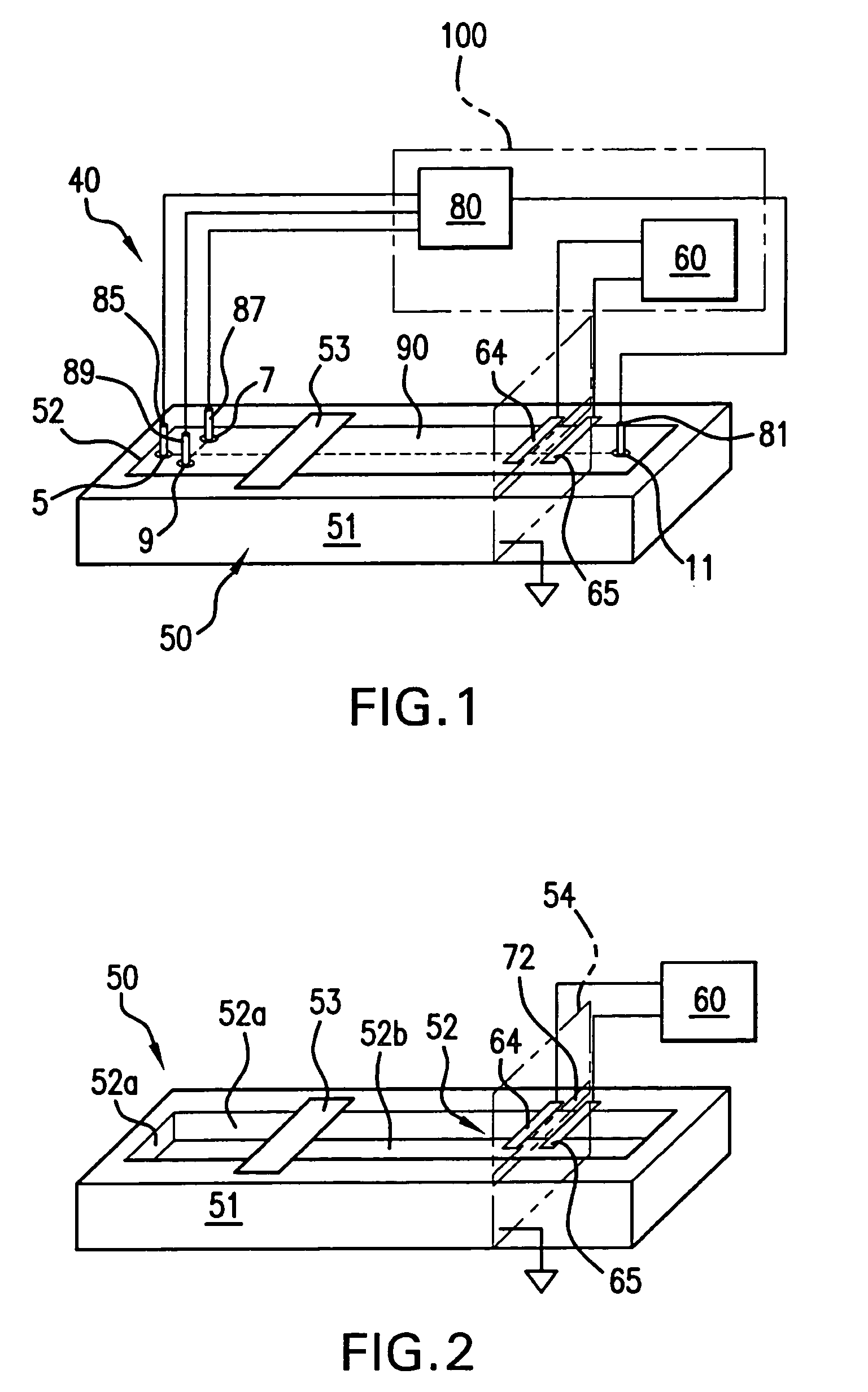
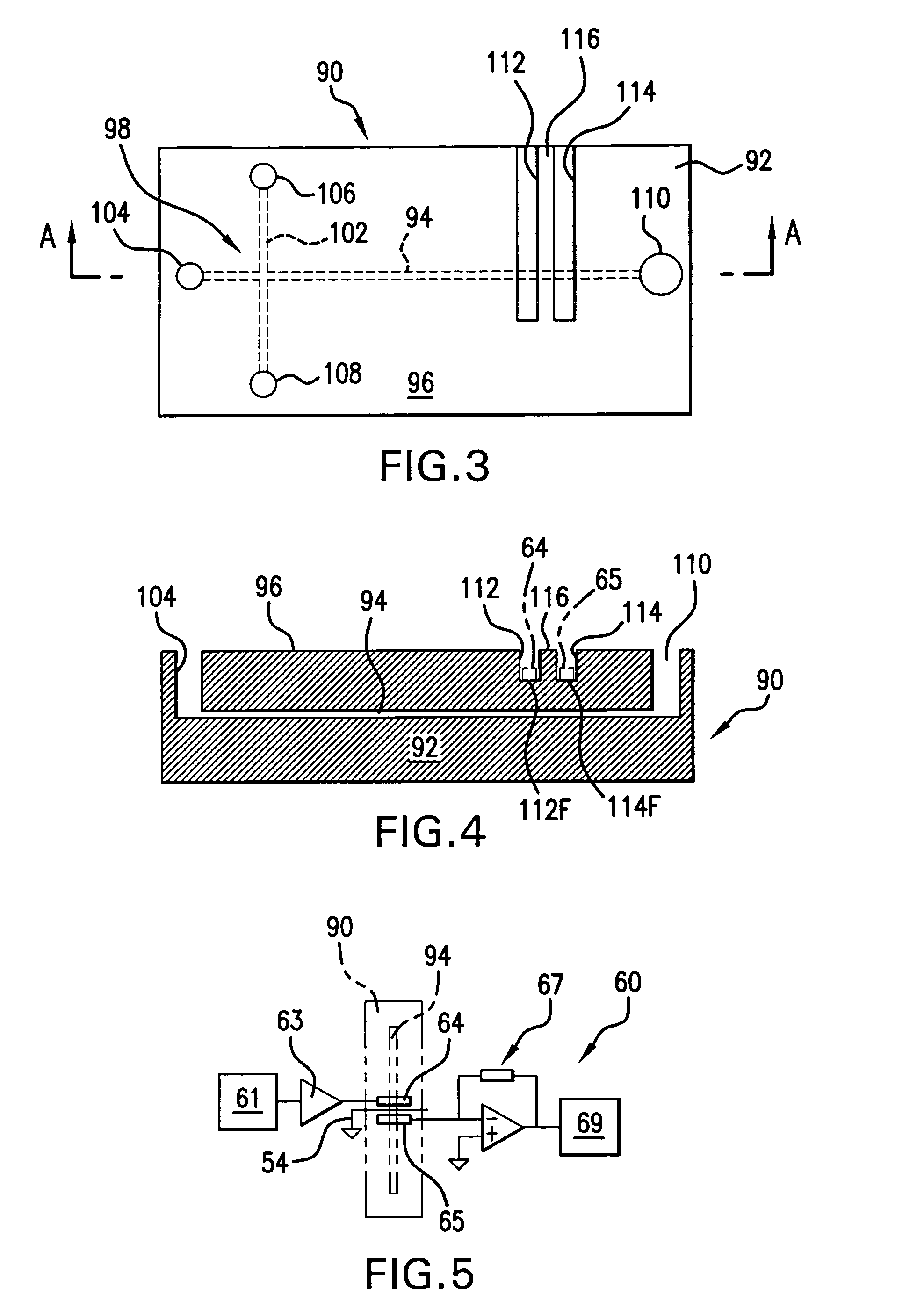
Method and apparatus for performing high-voltage contactless conductivity (HV-CCD) electrophoresis
InactiveUS20050109621A1Reduce adhesionSludge treatmentVolume/mass flow measurementAnalyteCapillary electrophoresis
A chip-based capillary electrophoresis assembly including: a holder including a frame for removably receiving a chip; a capillary electrophoresis microchip dimensioned to fit onto the holder, and comprising a body and a separation channel defined in the body; and a pair of adhesive detection electrodes integrated with an electronic conductometric detection circuit, wherein the assembly is assemblable by disposing the capillary electrophoresis microchip on the holder and removably placing the adhesive electrodes on the microchip body near the separation channel. A label-free analyte conductometric detection method and a label-free capillary electrophoresis immunoassay method are also described.
Owner:HAUSER PETER C
Signal amplified biological detection with conjugated polymers
The devices and method are provided for detecting labeled and label-free analytes, such as nucleic acids and proteins, employing conjugated cationic, anionic and neutral polymers. The analytes can be immobilized on a solid support material. The solid support material can be disposed in the pre-fabricated patterns on a substrate. The chemical structures of the polymers employed by the various embodiments of the present invention are described herein.
Owner:NAT UNIV OF SINGAPORE +1
Photonic crystal sensors with intergrated fluid containment structure
ActiveUS20080219615A1Efficiently conveyedPrevent leakageMaterial analysis by optical meansNanoopticsSensor arrayRefractive index
Photonic crystal (PC) sensors, and sensor arrays and sensing systems incorporating PC sensors are described which have integrated fluid containment and / or fluid handling structures. Sensors and sensing systems of the present disclosure are capable of high throughput sensing of analytes in fluid samples, bulk refractive index detection, and label-free detection of a range of molecules, including biomolecules and therapeutic candidates. The present disclosure also provides a commercially attractive fabrication platform for making photonic crystal sensors and systems wherein an integrated fluid containment structure and a photonic crystal structure are fabricated in a single molding or imprinting processing step amendable to high throughput processing.
Owner:X BODY +1
Packaged Chip For Multiplexing Photonic Crystal Microcavity Coupled Waveguide And Photonic Crystal Slot Waveguide Devices For Chip-Integrated Label-Free Detection And Absorption Spectroscopy With High Throughput, Sensitivity, Specificity, And Wide Dynamic Range
ActiveUS20140140655A1Specificity of bindingHigh sensitivityMaterial analysis by optical meansNanoopticsMicrofluidic channelWaveguide
Systems and methods for chip-integrated label-free detection and absorption spectroscopy with high throughput, sensitivity, and specificity are disclosed. The invention comprises packaged chips for multiplexing photonic crystal microcavity waveguide and photonic crystal slot waveguide devices. The packaged chips comprise crossing waveguides to prevent leakage of fluids from the microfluidic channels from the trenches or voids around the light guiding waveguides. Other embodiments are described and claimed.
Owner:OMEGA OPTICS
Applications of laser-processed substrate for molecular diagnostics
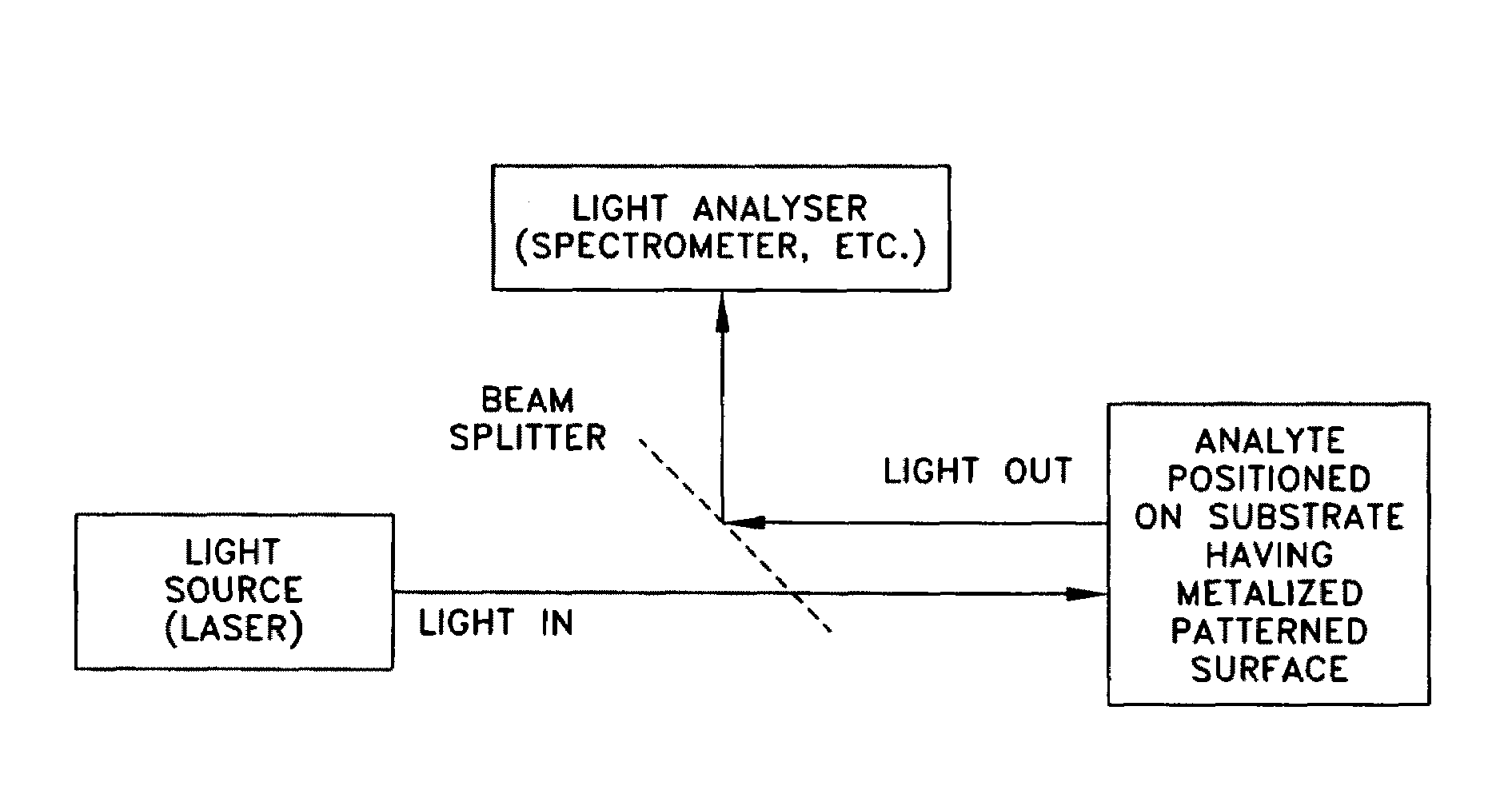
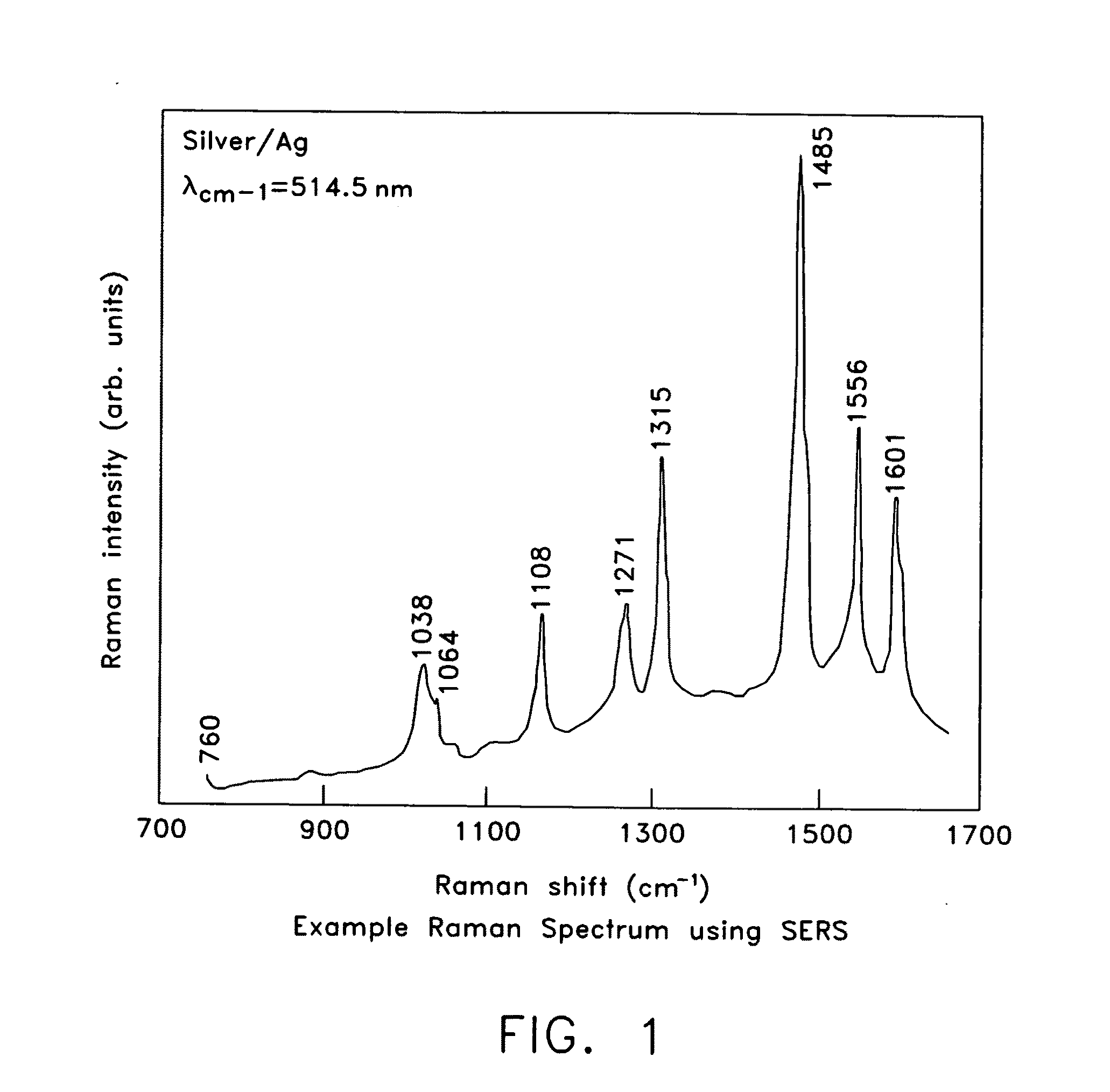
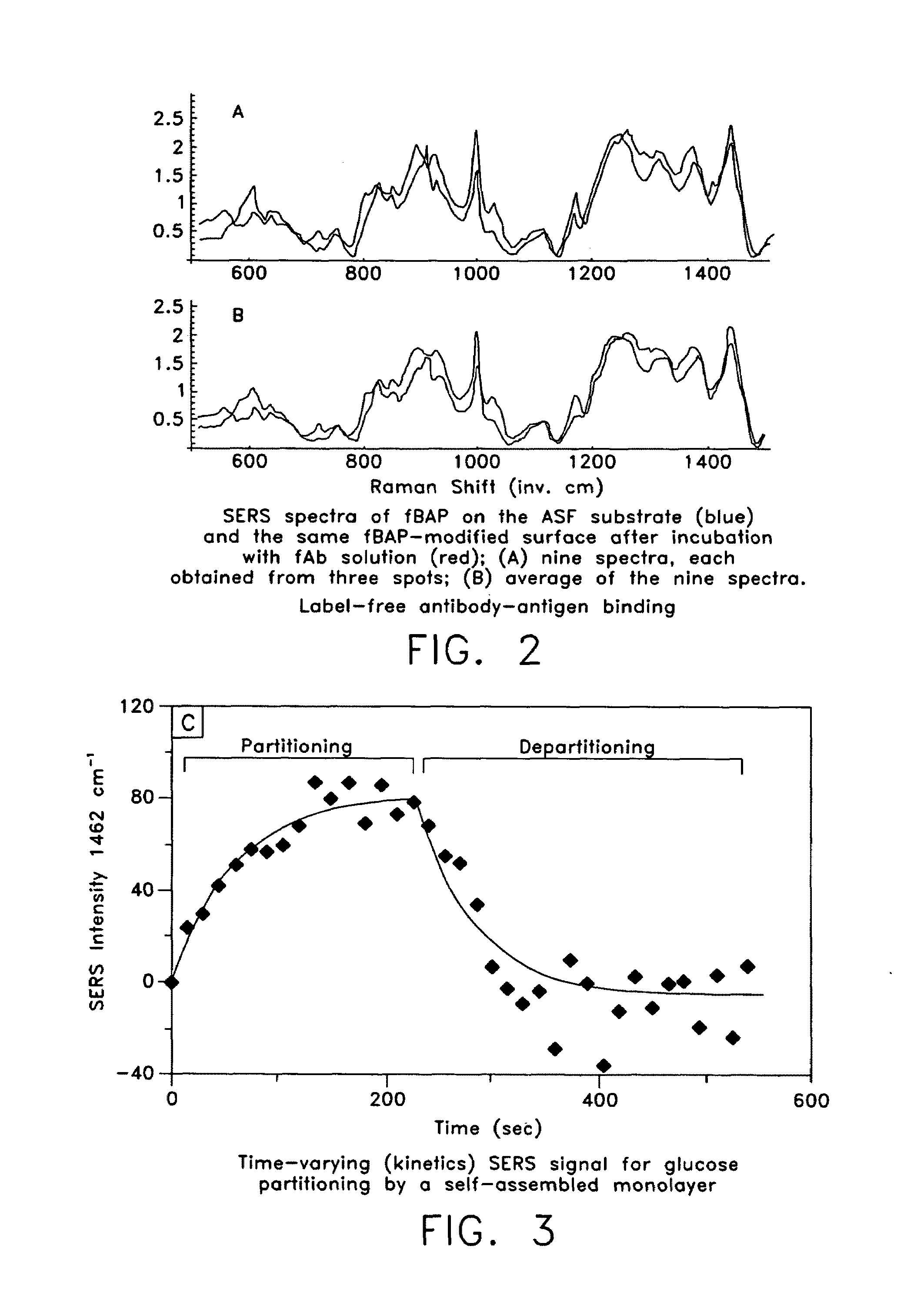
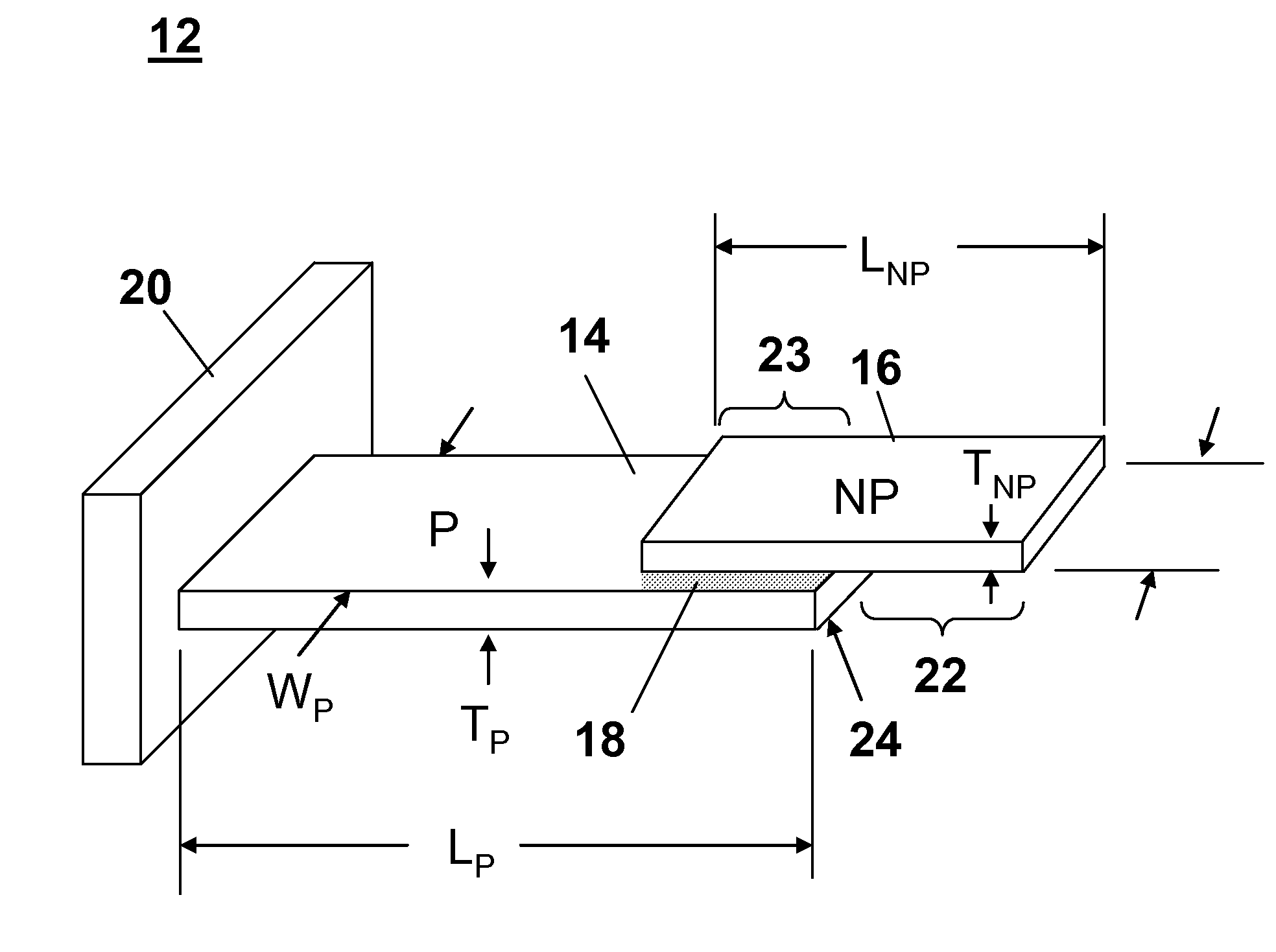
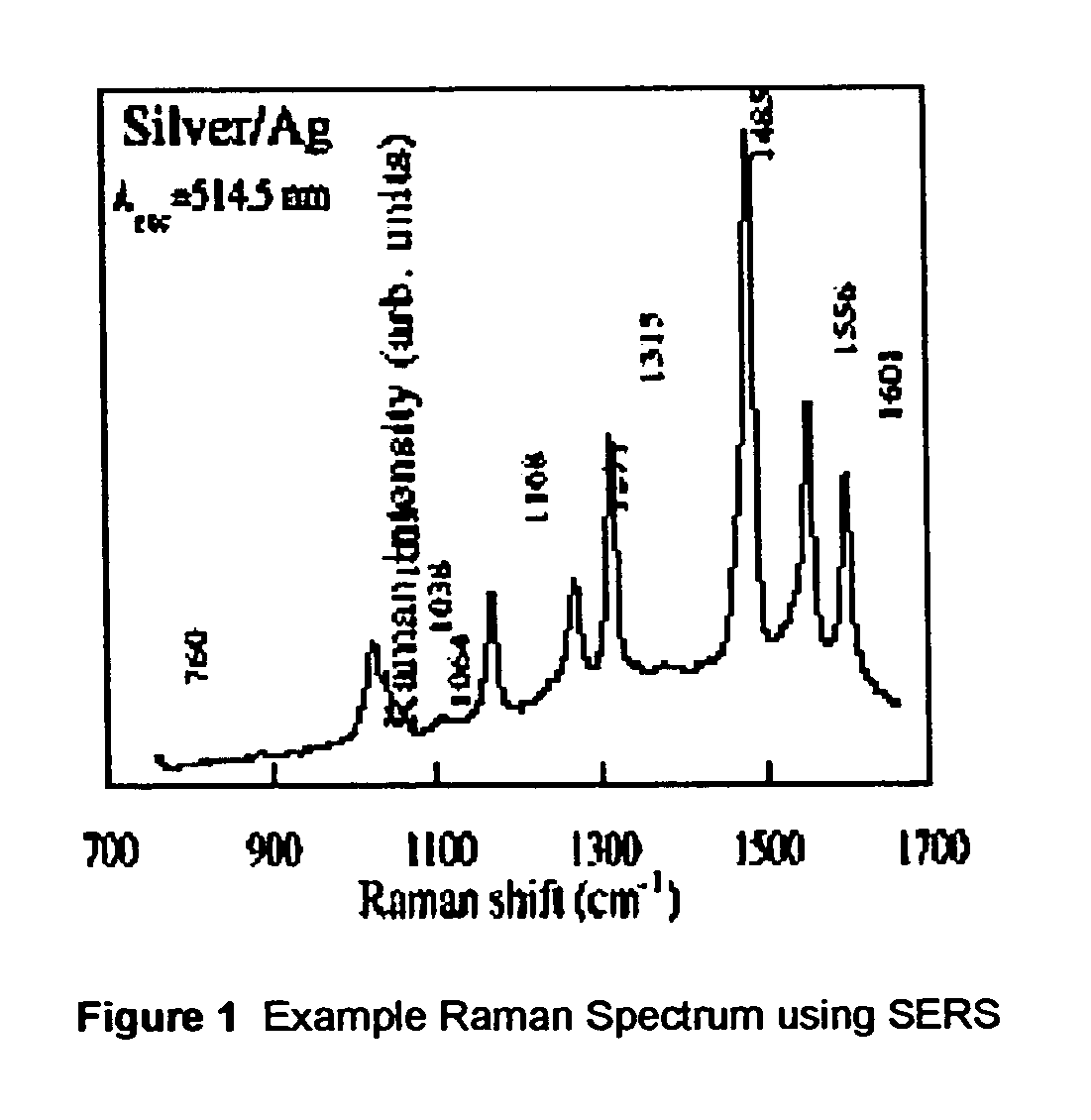
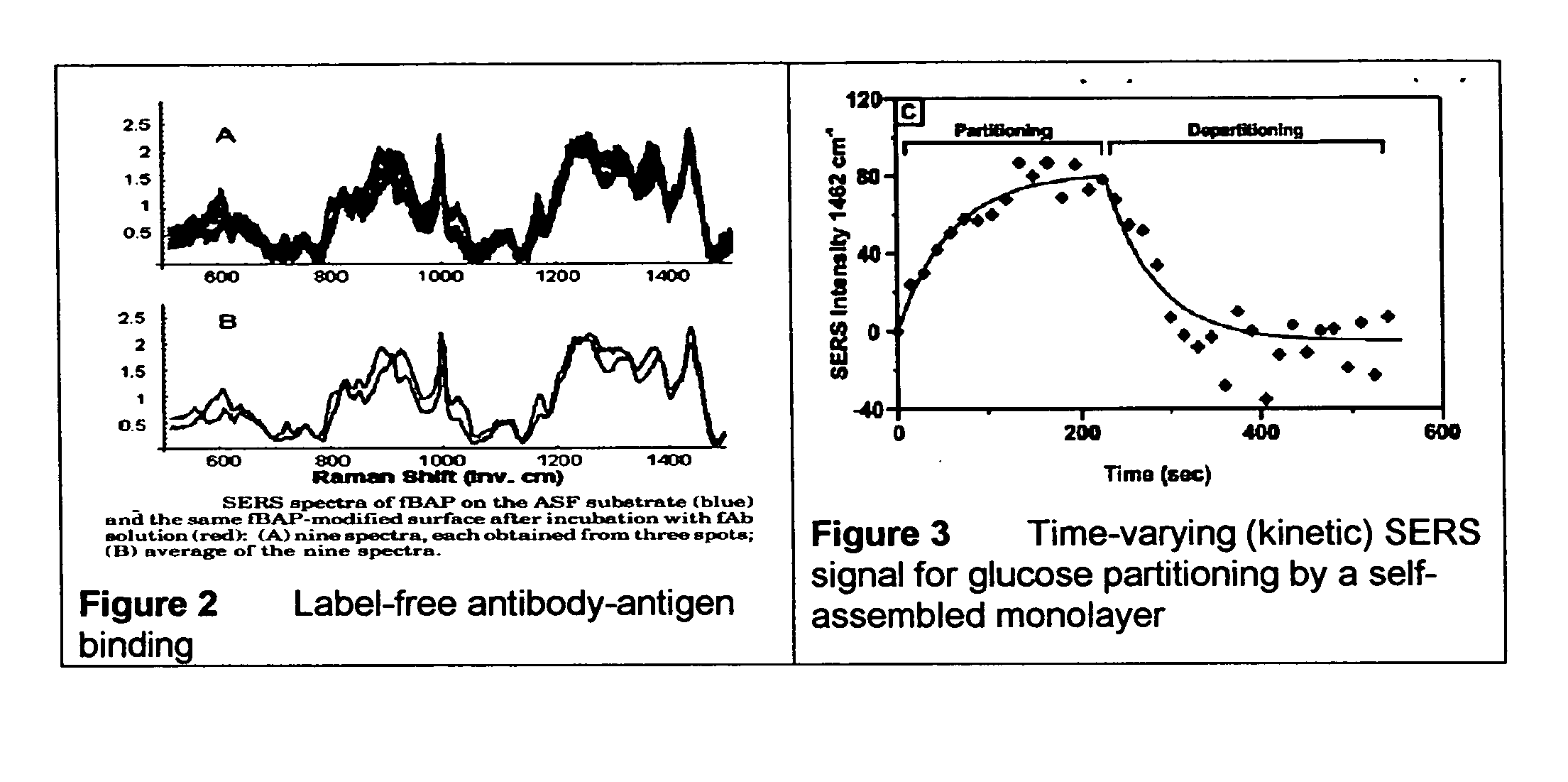
ActiveUS20070115469A1Facilitate SERS analysisMaterial nanotechnologyRadiation pyrometryChemical reactionPhotonics
Surface enhanced Raman Scattering (SERS) and related modalities offer greatly enhanced sensitivity and selectivity for detection of molecular species through the excitation of plasmon modes and their coupling to molecular vibrational modes. One of the chief obstacles to widespread application is the availability of suitable nanostructured materials that exhibit strong enhancement of Raman scattering, are inexpensive to fabricate, and are reproducible. I describe nanostructured surfaces for SERS and other photonic sensing that use semiconductor and metal surfaces fabricated using femtosecond laser processing. A noble metal film (e.g., silver or gold) is evaporated onto the resulting nanostructured surfaces for use as a substrate for SERS. These surfaces are inexpensive to produce and can have their statistical properties precisely tailored by varying the laser processing. Surfaces can be readily micropatterned and both stochastic and self-organized structures can be fabricated. This material has application to a variety of genomic, proteomic, and biosensing applications including label free applications including binding detection. Using this material, monolithic or arrayed substrates can be designed. Substrates for cell culture and microlabs incorporating microfluidics and electrochemical processing can be fabricated as well. Laser processing can be used to form channels in the substrate or a material sandwiched onto it in order to introduce reagents and drive chemical reactions. The substrate can be fabricated so application of an electric potential enables separation of materials by electrophoresis or electro-osmosis.
Owner:EBSTEIN STEVEN M
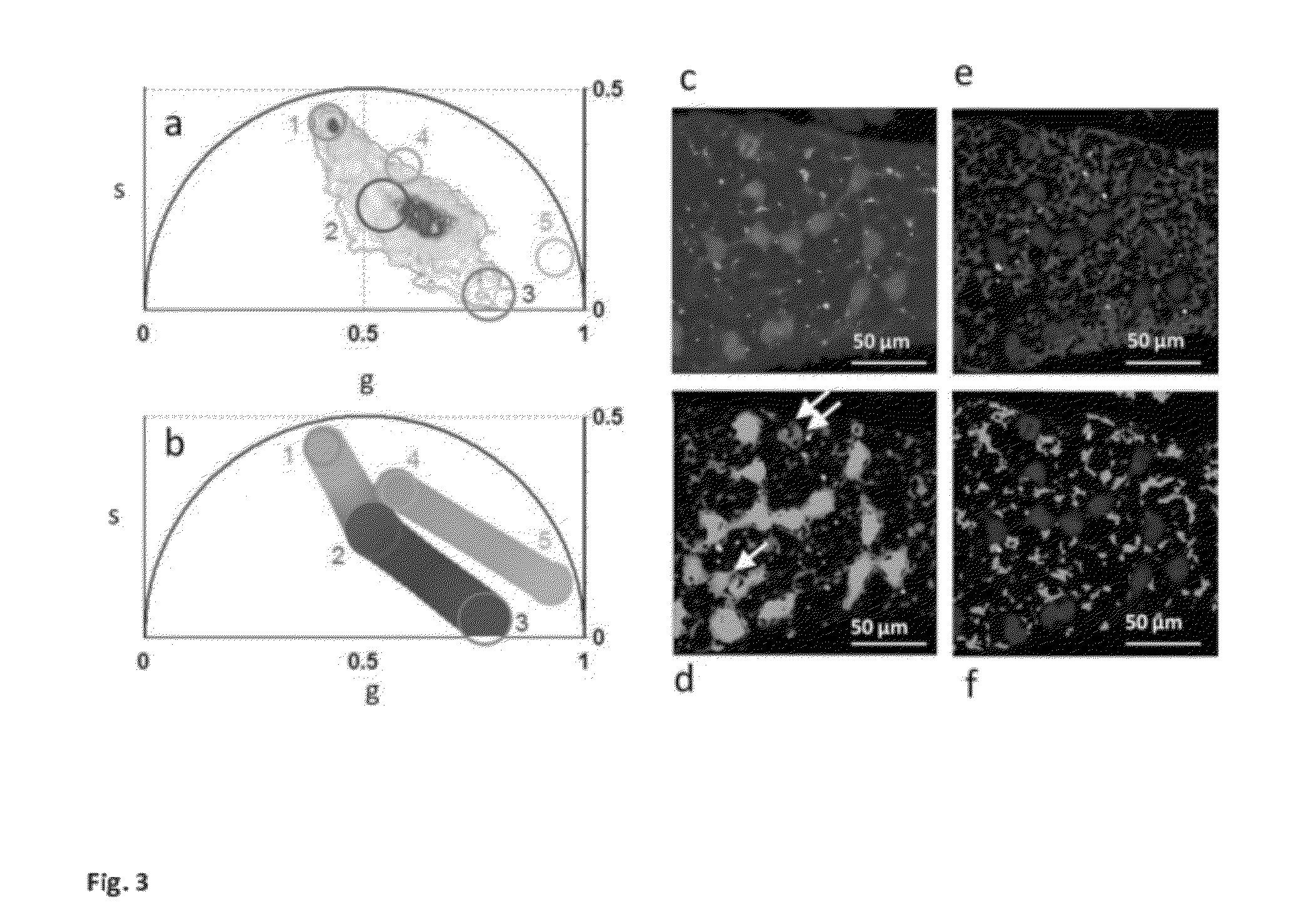
Phasor Method to Fluorescence Lifetime Microscopy to Discriminate Metabolic State of Cells in Living Tissue
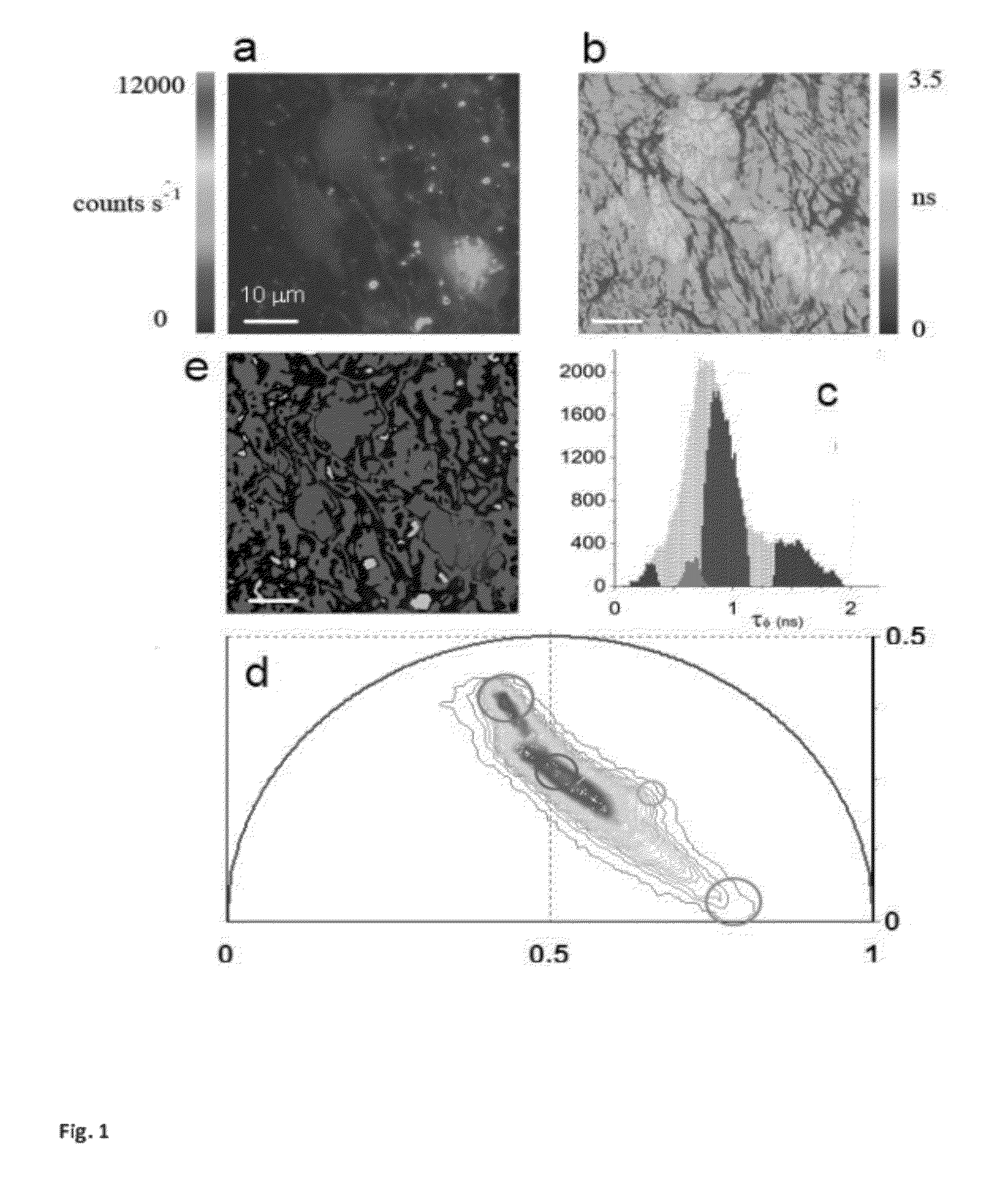
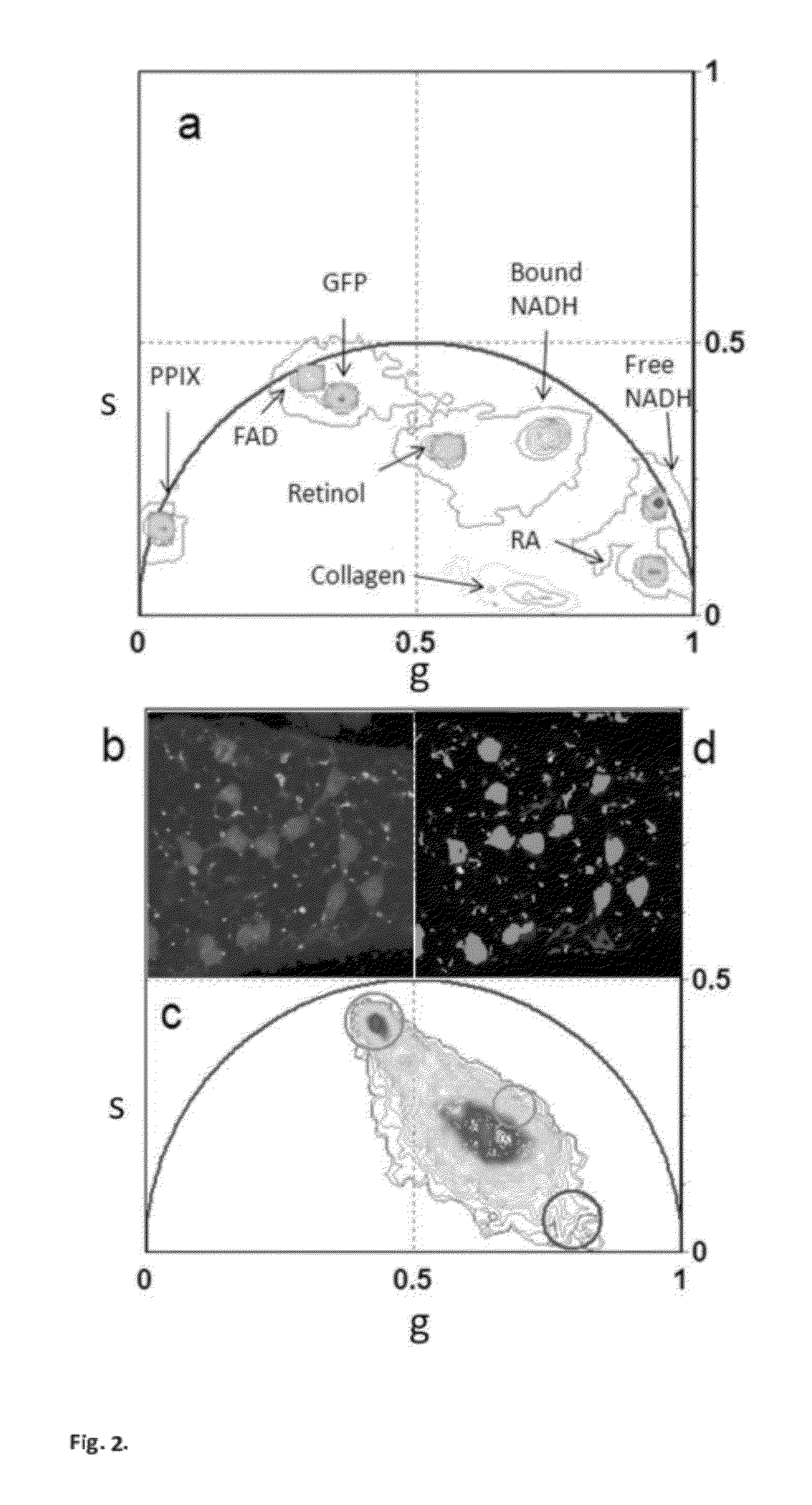
ActiveUS20120276578A1Bioreactor/fermenter combinationsBiological substance pretreatmentsPlant Germ CellsPorphyrin
“A label-free imaging method to monitor stem cell metabolism discriminates different states of stem cell as they differentiate in a living tissues. We use intrinsic fluorescence biomarkers and the phasor approach to Fluorescence Lifetime Imaging Microscopy (FLIM). We identify and map intrinsic fluorophores such as collagen, retinol, retinoic acid, flavins, nicotinamide adenine dinucleotide (NADH) and porphyrin. We measure the phasor values of germ cells in C. Elegans germ line. Their metabolic fingerprint cluster according to their differentiation state, reflecting changes in FAD concentration and NADH binding during the differentiation pathway. The phasor approach to lifetime imaging provides a label-free, fit-free and sensitive method to identify different metabolic state of cells during differentiation, to sense small changes in the redox state of cells and may identify symmetric and asymmetric divisions and predict cell fate.”
Owner:RGT UNIV OF CALIFORNIA