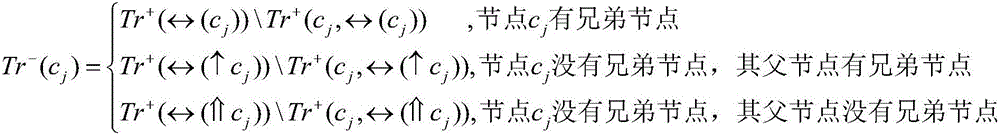Patents
Literature
572 results about "Function proteins" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
Proteins also have structural or mechanical functions, such as actin and myosin in muscle and the proteins in the cytoskeleton, which form a system of scaffolding that maintains cell shape. Other proteins are important in cell signaling, immune responses, cell adhesion, and the cell cycle.
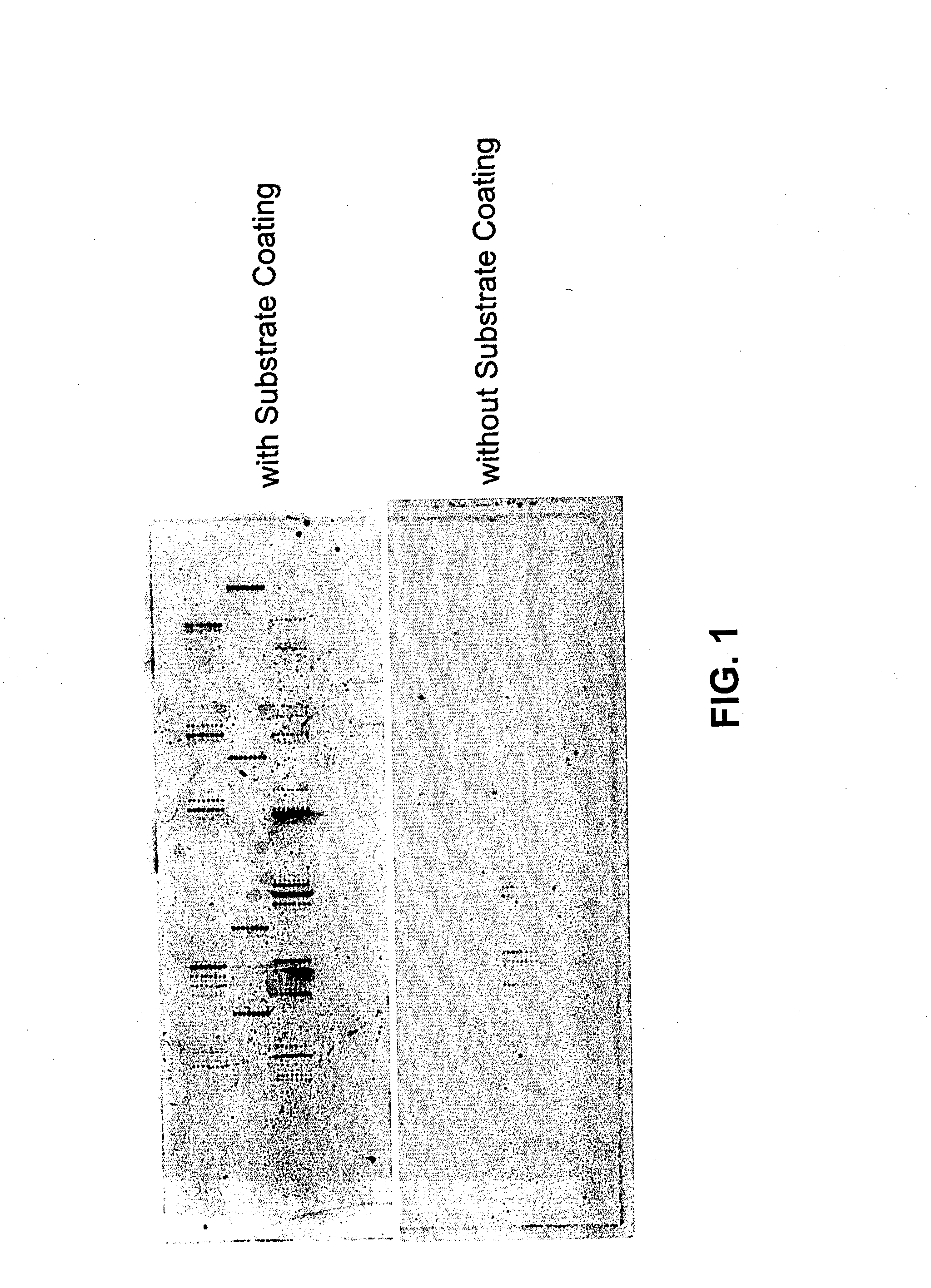
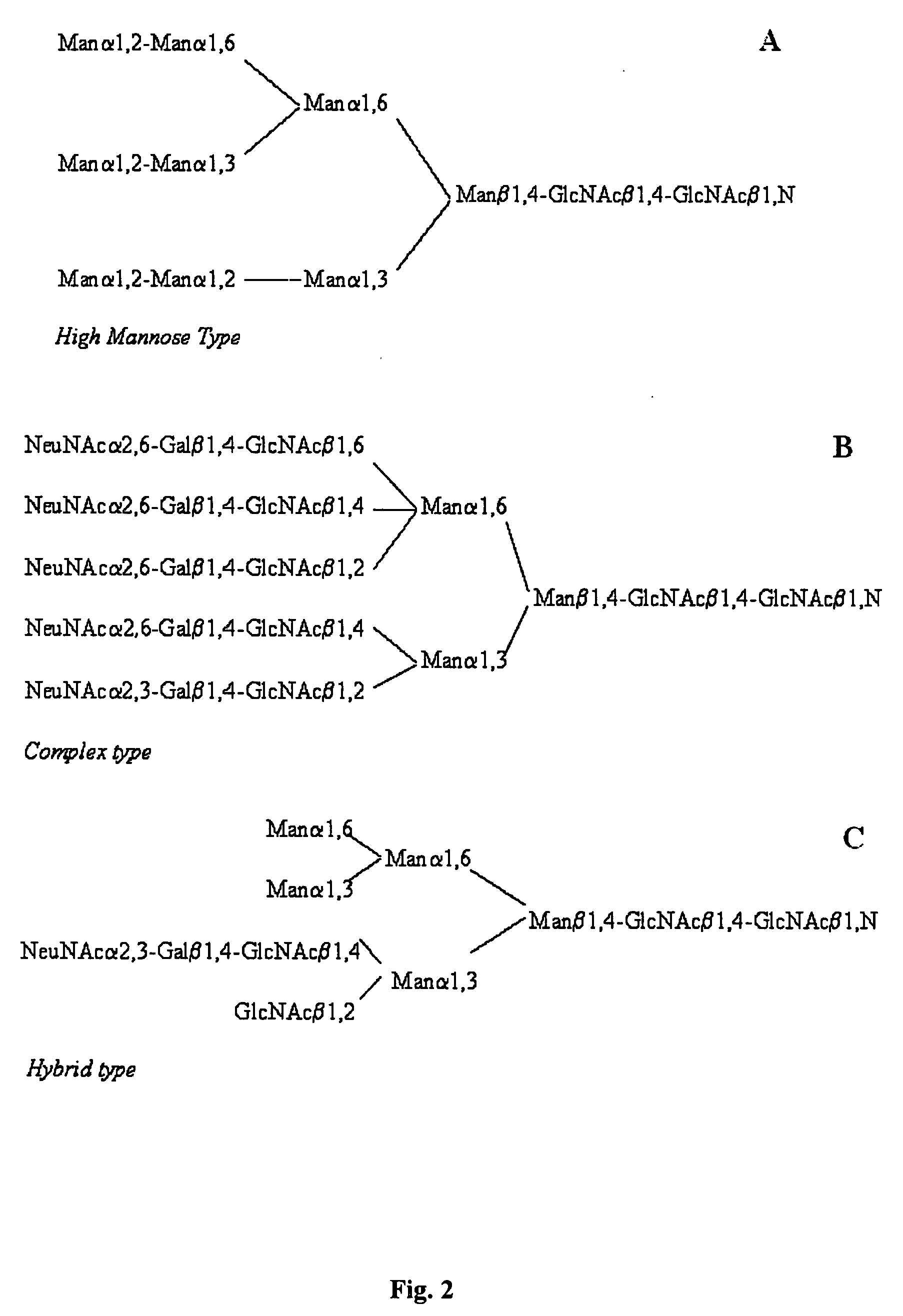
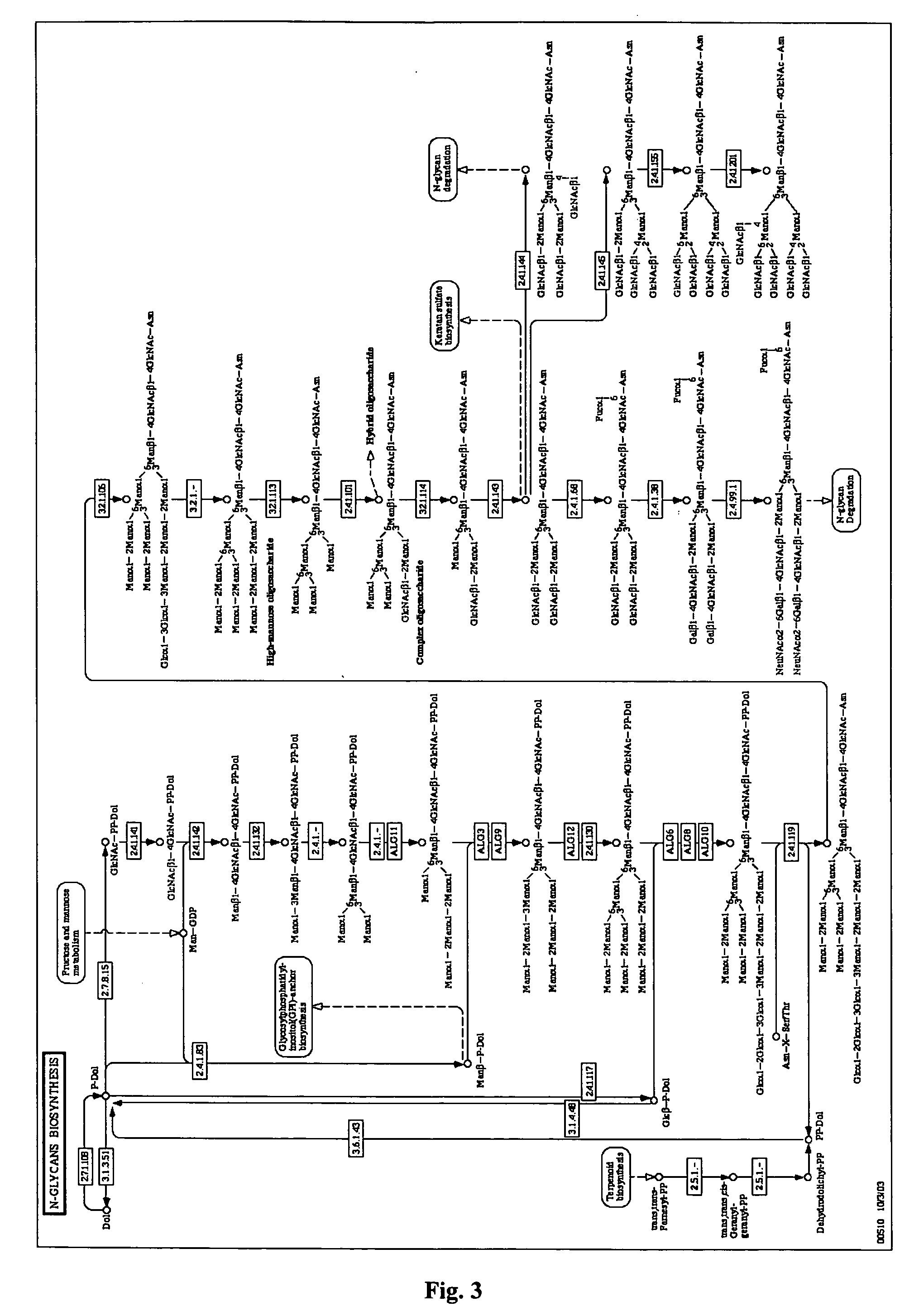
Methods and products related to the improved analysis of carbohydrates
The invention relates, in part, to the improved analysis of carbohydrates. In particular, the invention relates to the analysis of carbohydrates, such as N-glycans and O-glycans found on proteins. Improved methods, therefore, for the study of glycosylation patterns on cells, tissue and body fluids are also provided. Information regarding the analysis of glycans, such as the glycosylation patterns on cells, tissues and in body fluids, can be used in diagnostic and treatment methods as well as for facilitating the study of the effects of glycosylation / altered glycosylation on protein function. Such methods are also provided. Methods are also provided to assess protein production processes, to assess the purity of proteins produced, and to select proteins with the desired glycosylation.
Owner:MOMENTA PHARMA +2
Method for in vitro molecular evolution of protein function
InactiveUS6989250B2Altered propertyIncreased complexitySugar derivativesMicrobiological testing/measurementNucleotideIn vivo
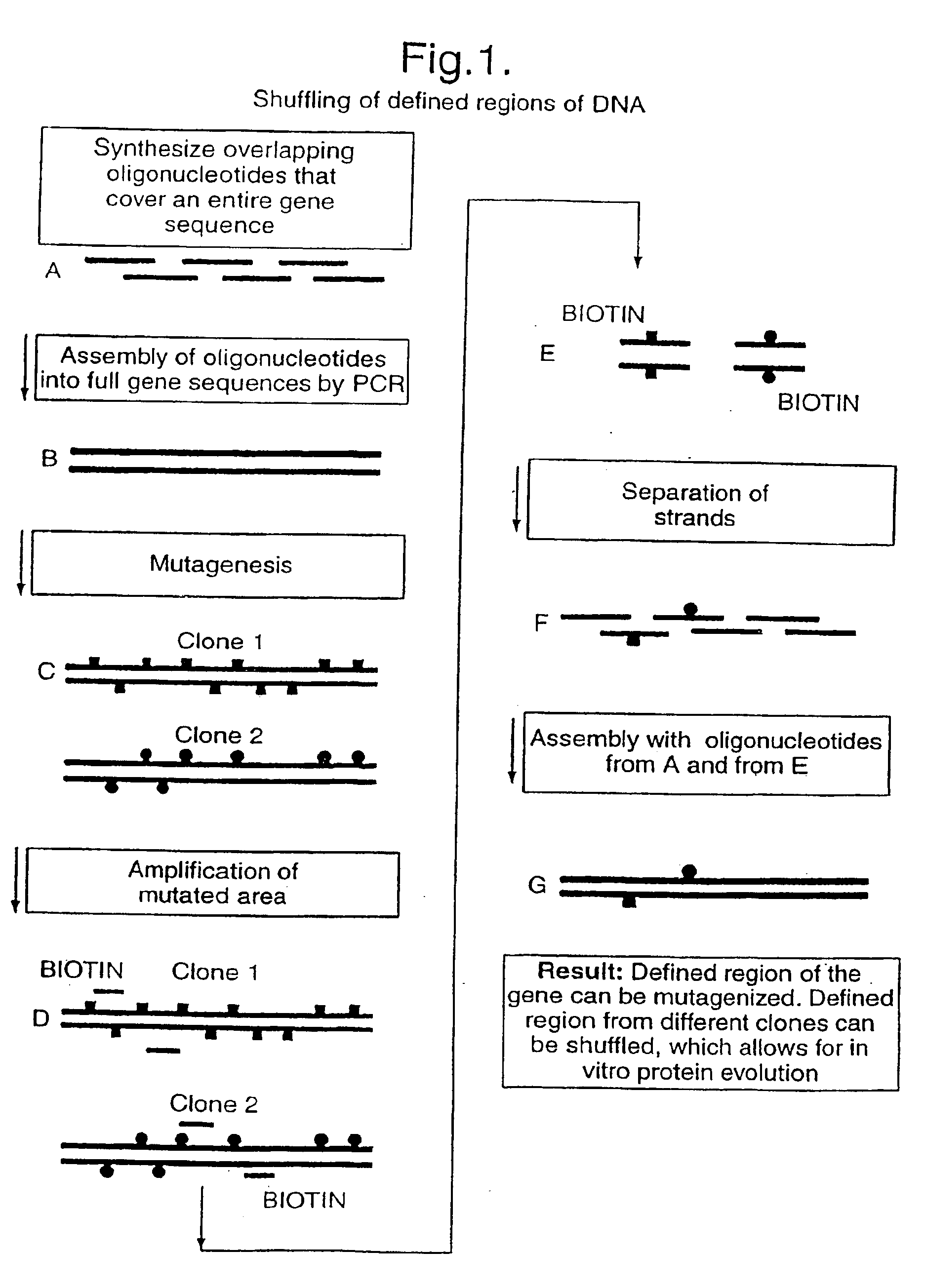
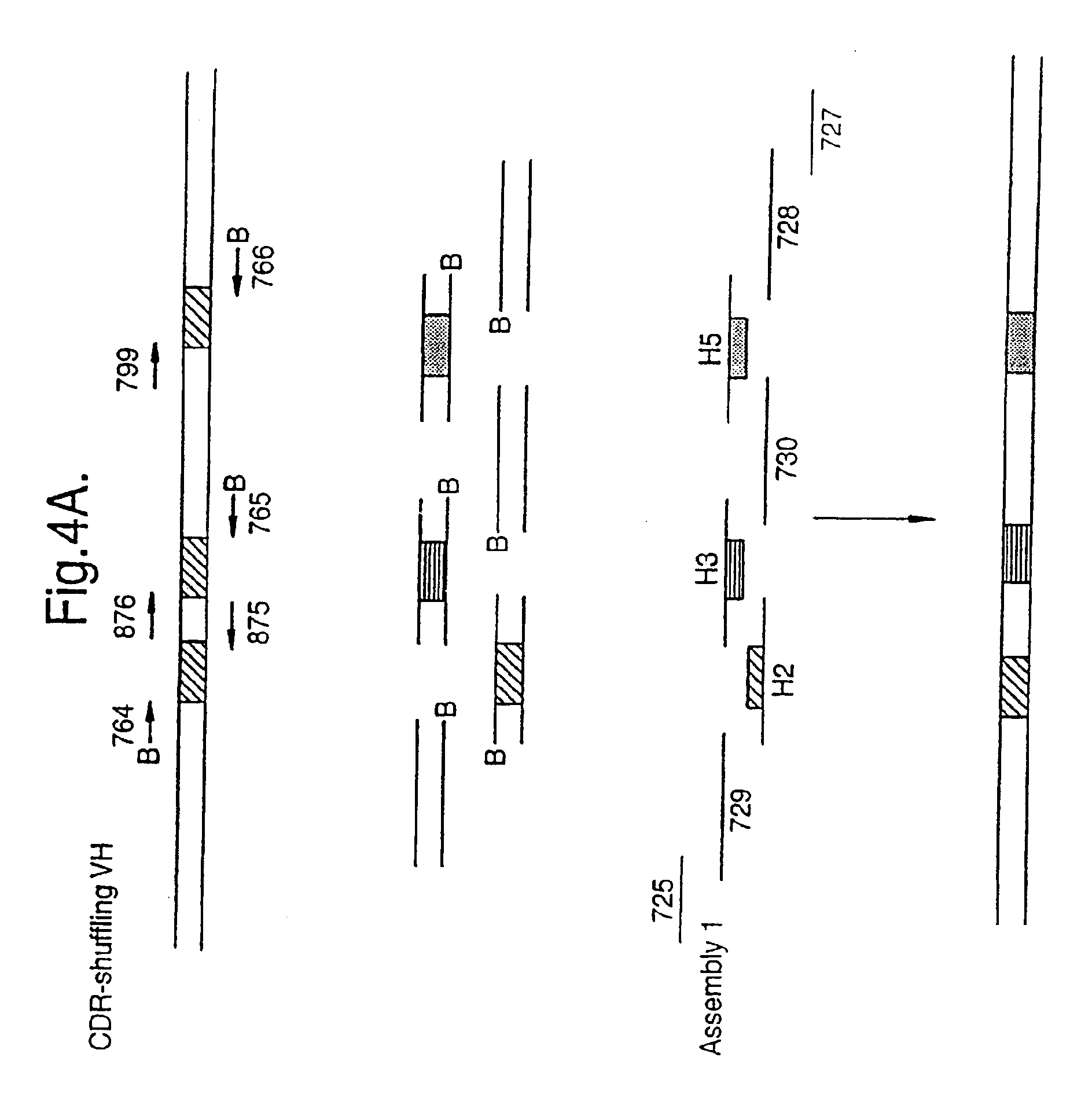
The present invention relates to a method for in vitro creation of molecular libraries evolution of protein function. Particularly, it relates to variability and modification of protein function by shuffling polynucleotide sequence segments. A protein of desired characteristics can be obtained by incorporating variant peptide regions (variant motifs) into defined peptide regions (scaffold sequence). The variant motifs can be obtained from parent DNA which has been subjected to mutagenesis to create a plurality of differently mutated derivatives thereof or they can be obtained from in vivo sequences. These variant motifs can then be incorporated into a scaffold sequence and the resulting coded protein screened for desired characteristics. This method is ideally used for obtaining antibodies with desired characteristics by isolating individual CDR DNA sequences and incorporating them into a scaffold which may, for example, be from a totally different antibody.
Owner:BIOINVENT INT AB
Bispecific antibody substituting for functional proteins
ActiveUS20070041978A1Enhances enzymatic reactionImmunoglobulins against blood coagulation factorsFactor VIIBlood coagulation factor VIIIBlood Coagulation Factor X
The present inventors succeeded in constructing bispecific antibodies, which bind to both the blood coagulation factor IX / activated blood coagulation factor IX and blood coagulation factor X, and functionally substitute for blood coagulation factor VIII / activated blood coagulation factor VIII which enhances the enzymatic reaction.
Owner:CHUGAI PHARMA CO LTD
Double Specific Antibodies Substituting For Functional Proteins
InactiveUS20080075712A1Reduced activityOutstanding sustainabilityImmunoglobulins against blood coagulation factorsNervous disorderBlood coagulation factor VIIIBlood Coagulation Factor X
The present inventors succeeded in separating bispecific antibodies that functionally substitute for ligands of type I interferon receptors comprising two types of molecules: AR1 chain and AR2 chain. Furthermore, the present inventors succeeded in producing bispecific antibodies that substitute for the enzyme reaction-accelerating function of blood coagulation factor VIII / activated blood coagulation factor VIII, which bind to both blood coagulation factor IX / activated blood coagulation factor IX and blood coagulation factor X.
Owner:CHUGAI PHARMA CO LTD
Methods, reagents, kits and apparatus for protein function analysis
InactiveUS20030207290A1Bioreactor/fermenter combinationsBiological substance pretreatmentsProtein functionEnzyme
Owner:MESO SCALE TECH LLC
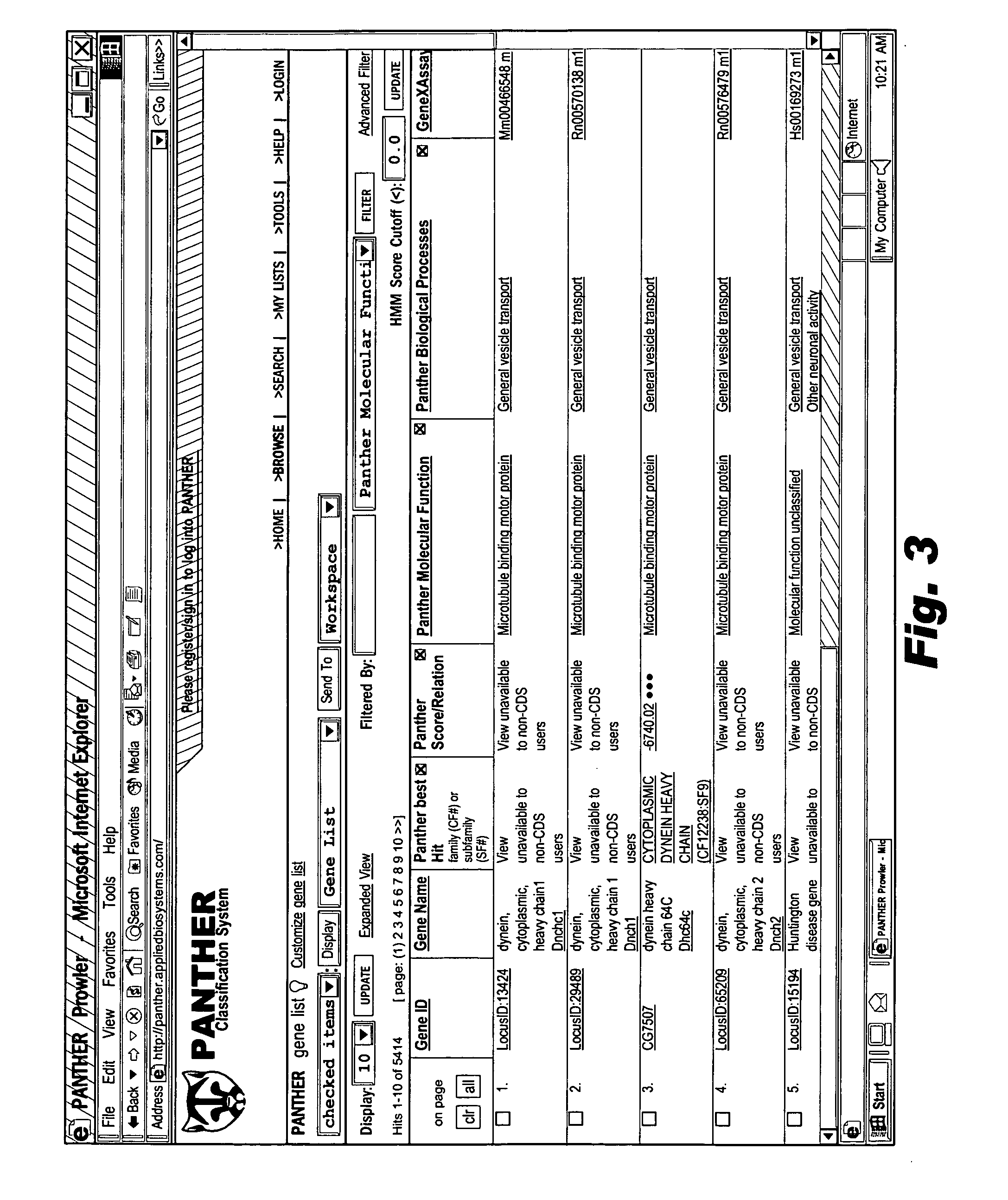
Browsable database for biological use
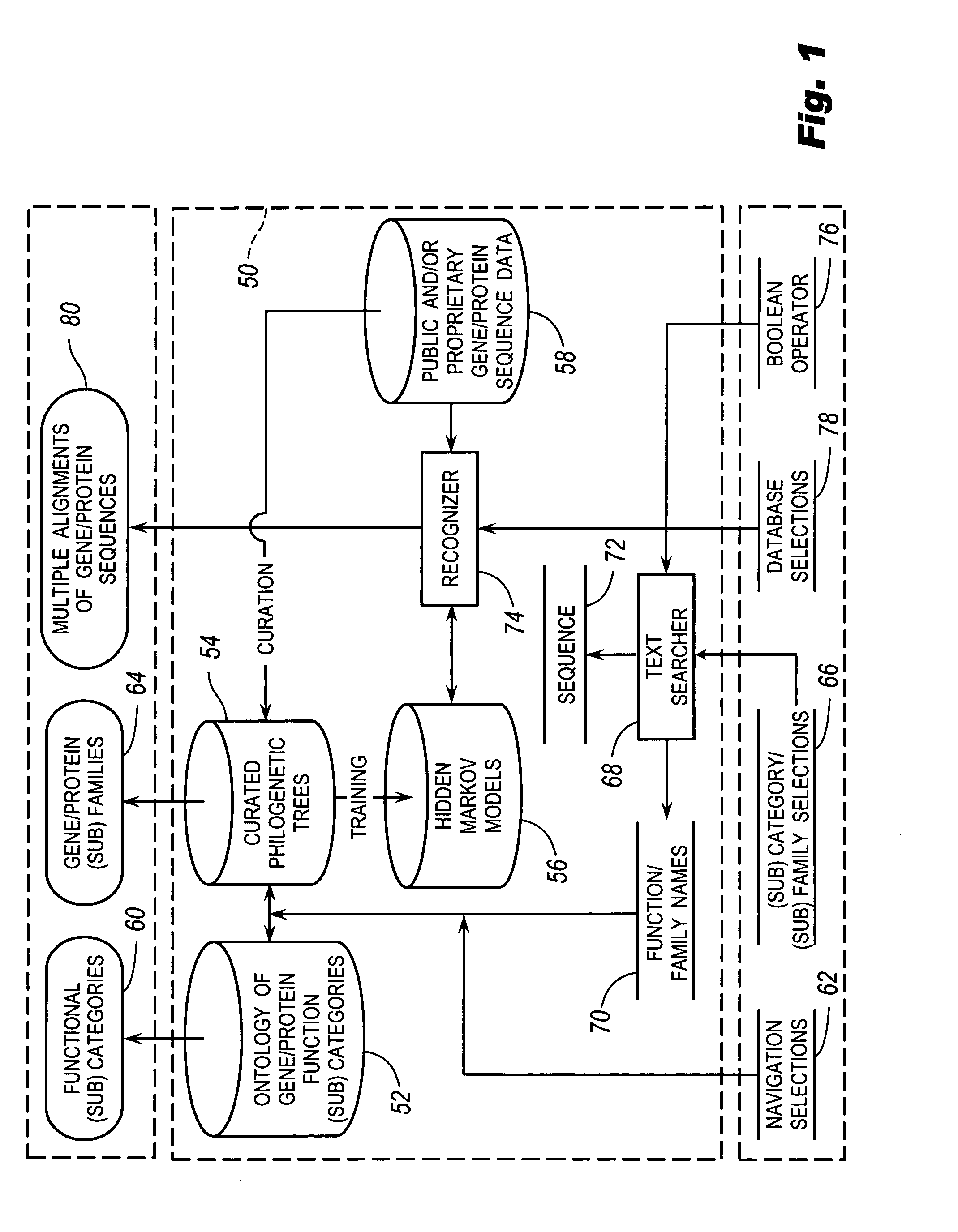
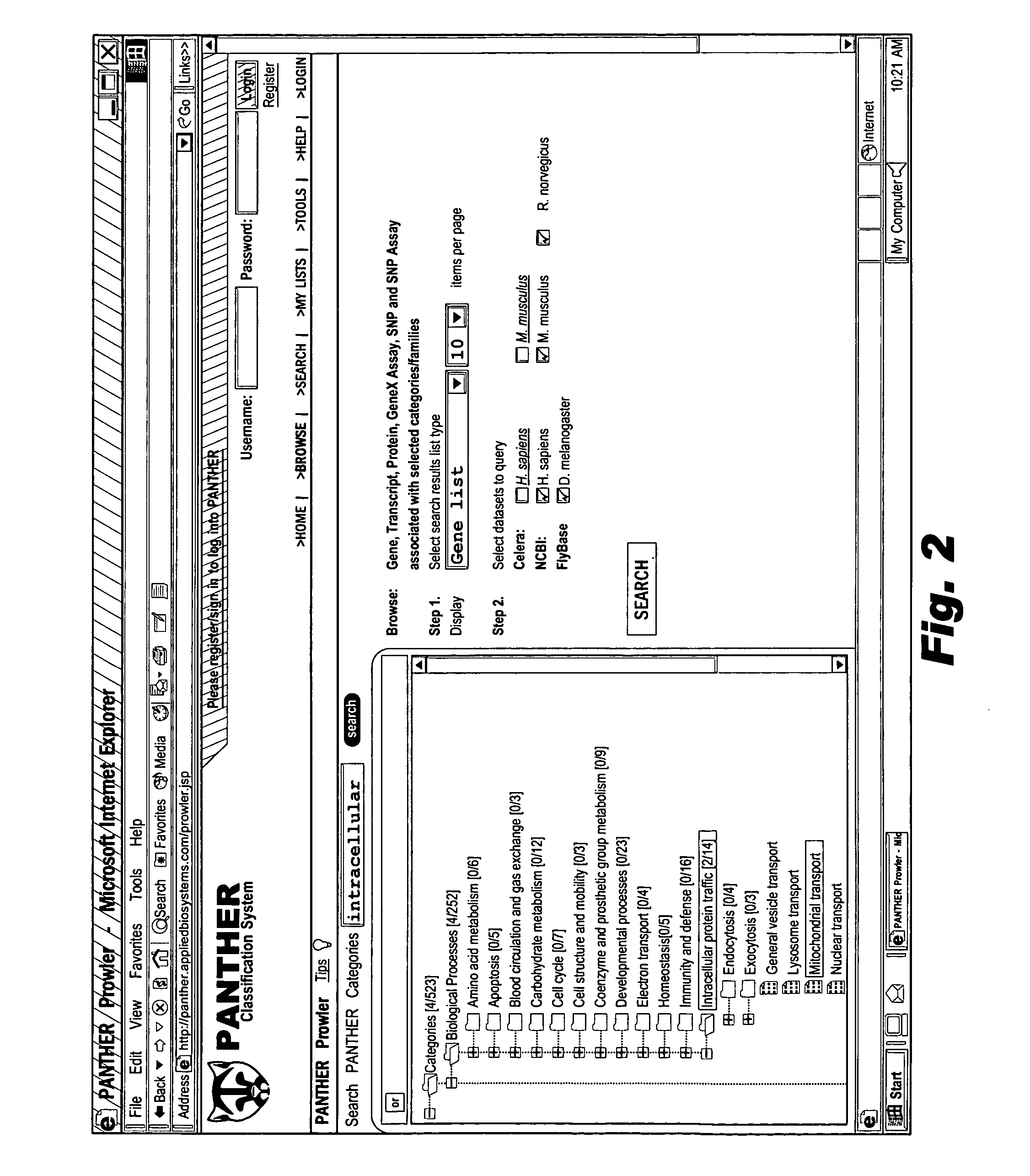
The browsable database can allow for high-throughput analysis of protein sequences. One helpful feature may be a simplified ontology of protein function, which allows browsing of the database by biological functions. Biologist curators may have associated the ontology terms with Hidden Markov Models (HMMs), rather than individual sequences, so that they can be applied to additional sequences. To ensure accurate functional classification, HMMs may be constructed not only for families, but for curator-defined subfamilies, whenever family members have divergent functions or nomenclature. Multiple sequence alignments and phylogenetic trees, including curator-assigned information, can be available for each family. Various versions of the browsable database may include training sequences from all organisms in the GenBank non-redundant protein database, and the HMMs can be used to classify gene products across the entire genomes of human, and Drosophila melanogaster.
Owner:APPL BIOSYSTEMS INC
Hormone receptor functional dimers and methods of their use
InactiveUS7057015B1Enhance possibility of producingIncrease flexibilityFusion with DNA-binding domainSugar derivativesADAMTS ProteinsProtein Unit
The invention provides chimeric proteins having at least two functional protein units, each containing the dimerization domain of a member of the steroid / thyroid hormone nuclear receptor superfamily. The chimeric proteins can fold under crystallization conditions to form functional entities. The functional entities optionally contain a novel flexible peptide linker of variable lengths between at least two of the protein units. In a preferred embodiment, the linker is designed to be increased in increments of 12 amino acids each to aid in preparation of variant chimeric proteins. The DNA binding characteristics of the invention functional entities differ from those of wild-type complexes formed between “monomeric” receptors and their binding partners. Some functional entities, e.g. dimers expressed as fusion proteins, transactivate responsive promoters in a manner similar to wild-type complexes, while others do not promote transactivation and function instead essentially as constitutive repressors. The invention further provides nucleotide sequences encoding the invention chimeric proteins, cells containing such nucleotide sequences, and methods for using the invention chimeric proteins to modulate expression of one or more exogenous genes in a subject organism. In addition, isolated protein crystals suitable for x-ray diffraction analysis and methods for obtaining putative ligands for the invention chimeric proteins are provided.
Owner:SALK INST FOR BIOLOGICAL STUDIES
Bispecific antibody substituting for functional proteins
ActiveUS8062635B2Enhances enzymatic reactionImmunoglobulins against blood coagulation factorsFactor VIIBlood coagulation factor VIIIBlood Coagulation Factor X
The present inventors succeeded in constructing bispecific antibodies, which bind to both the blood coagulation factor IX / activated blood coagulation factor IX and blood coagulation factor X, and functionally substitute for blood coagulation factor VIII / activated blood coagulation factor VIII which enhances the enzymatic reaction.
Owner:CHUGAI PHARMA CO LTD
Small molecule-dependent inteins and uses thereof
ActiveUS20140065711A1Low splicing efficiencySlow splicingHydrolasesFusion with post-translational modification motifInteinNatural state
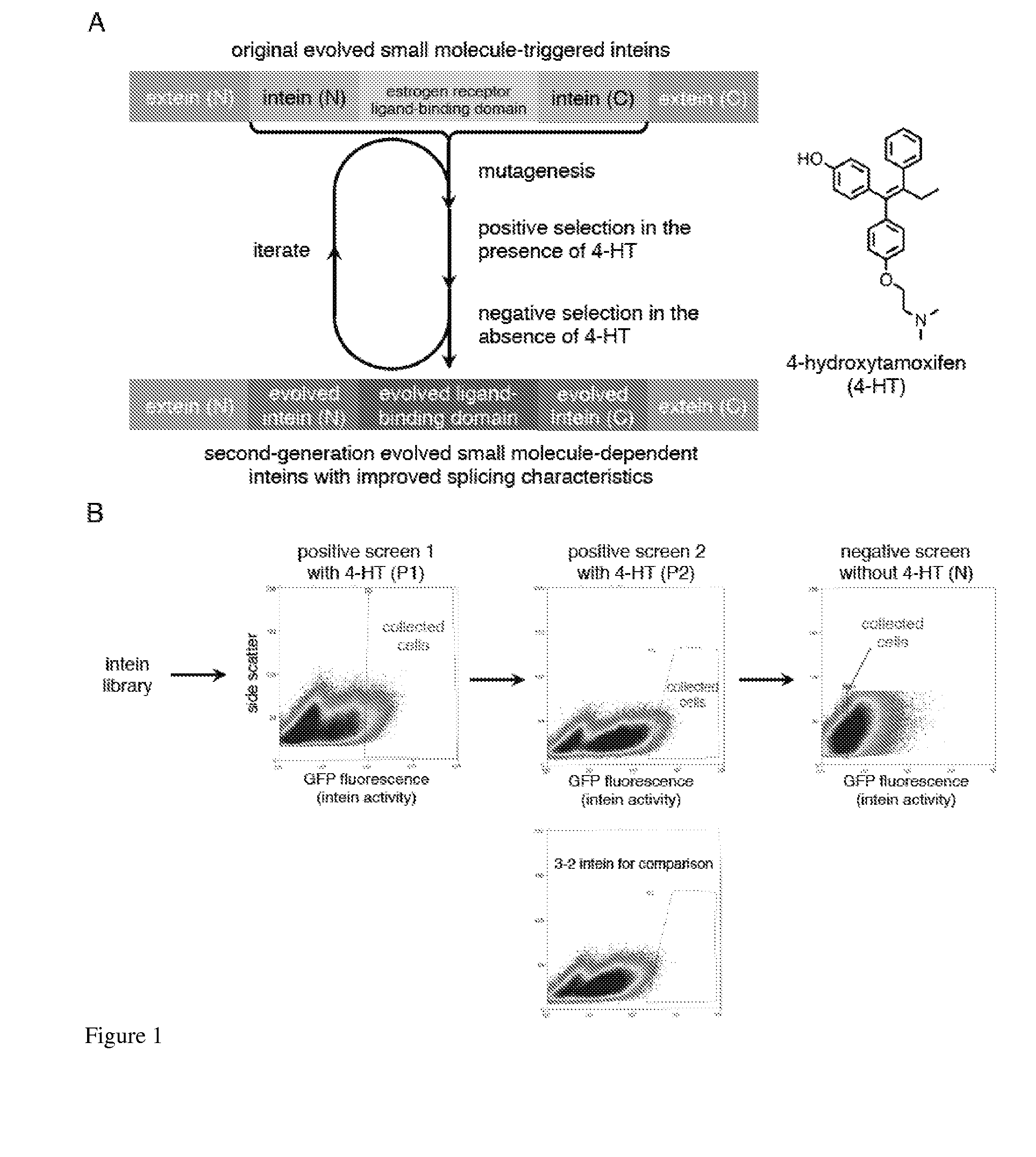
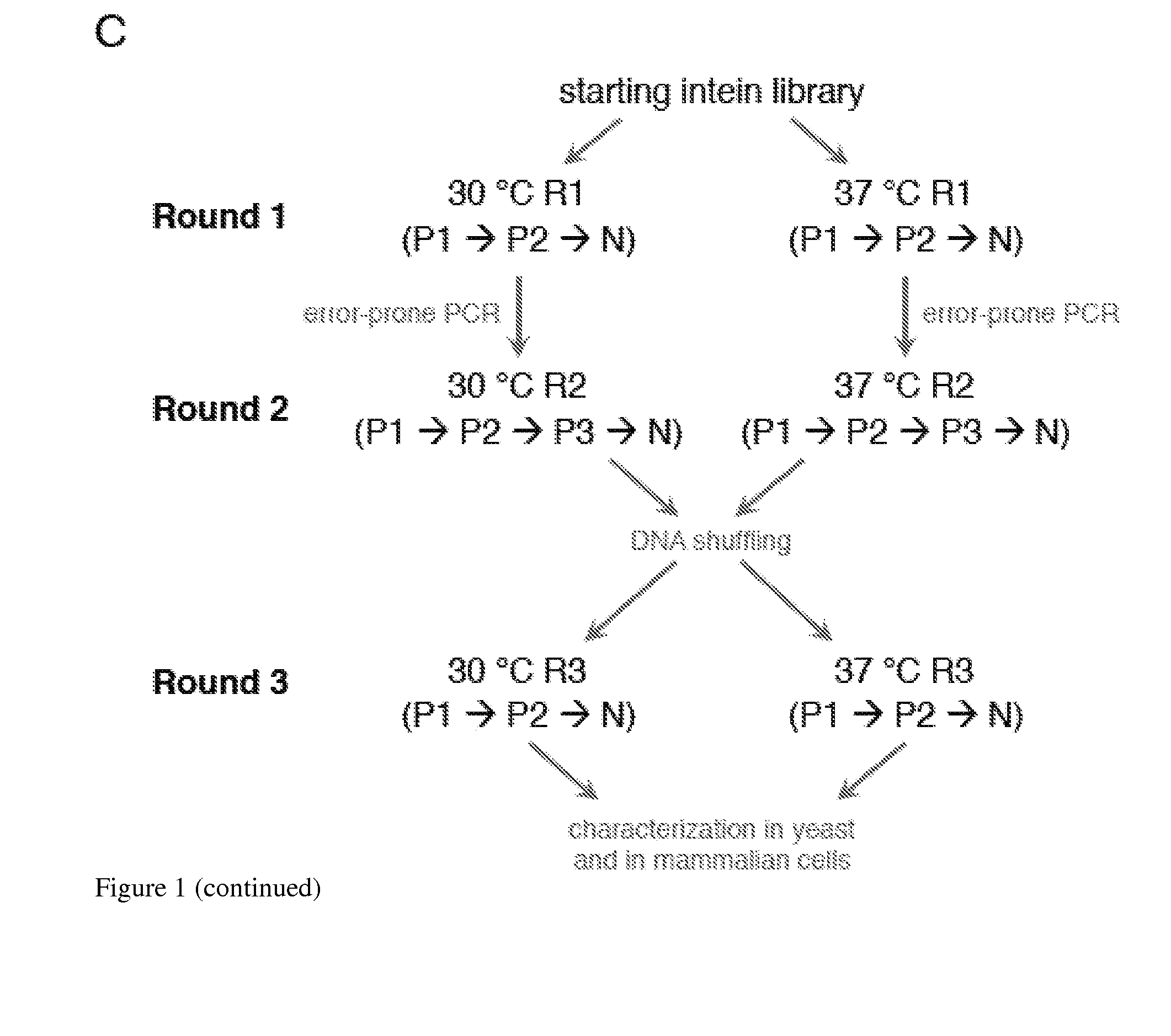
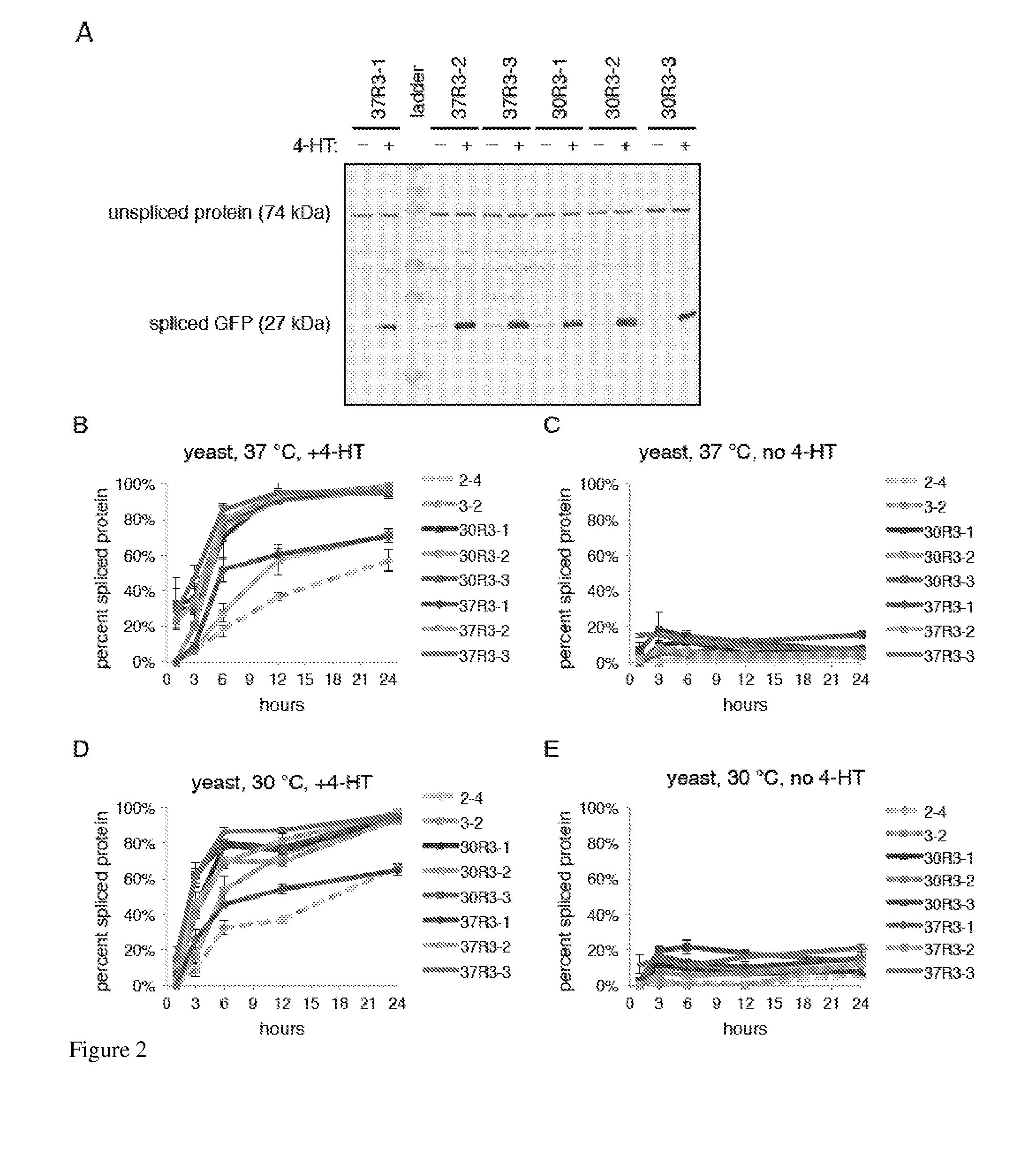
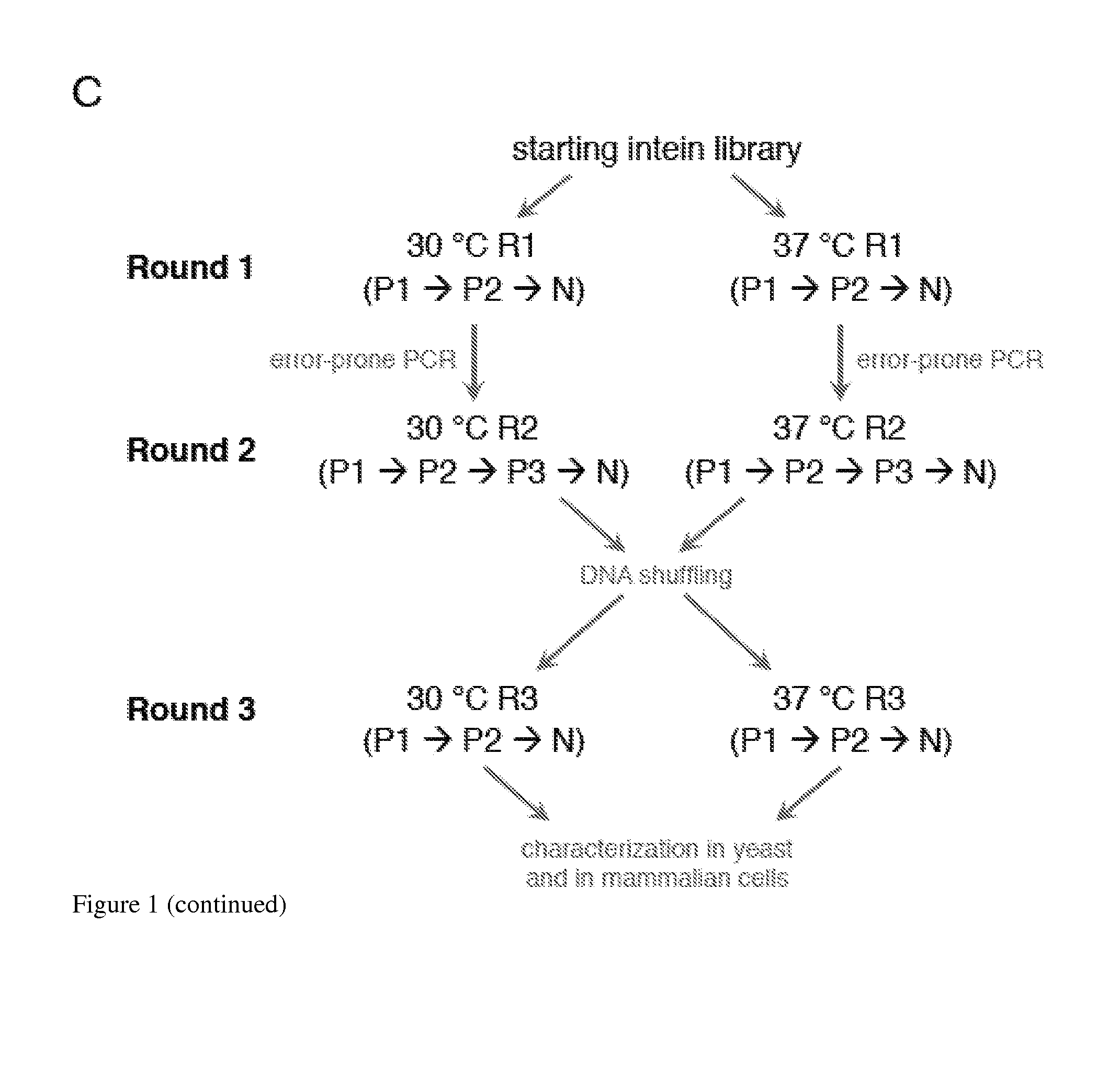
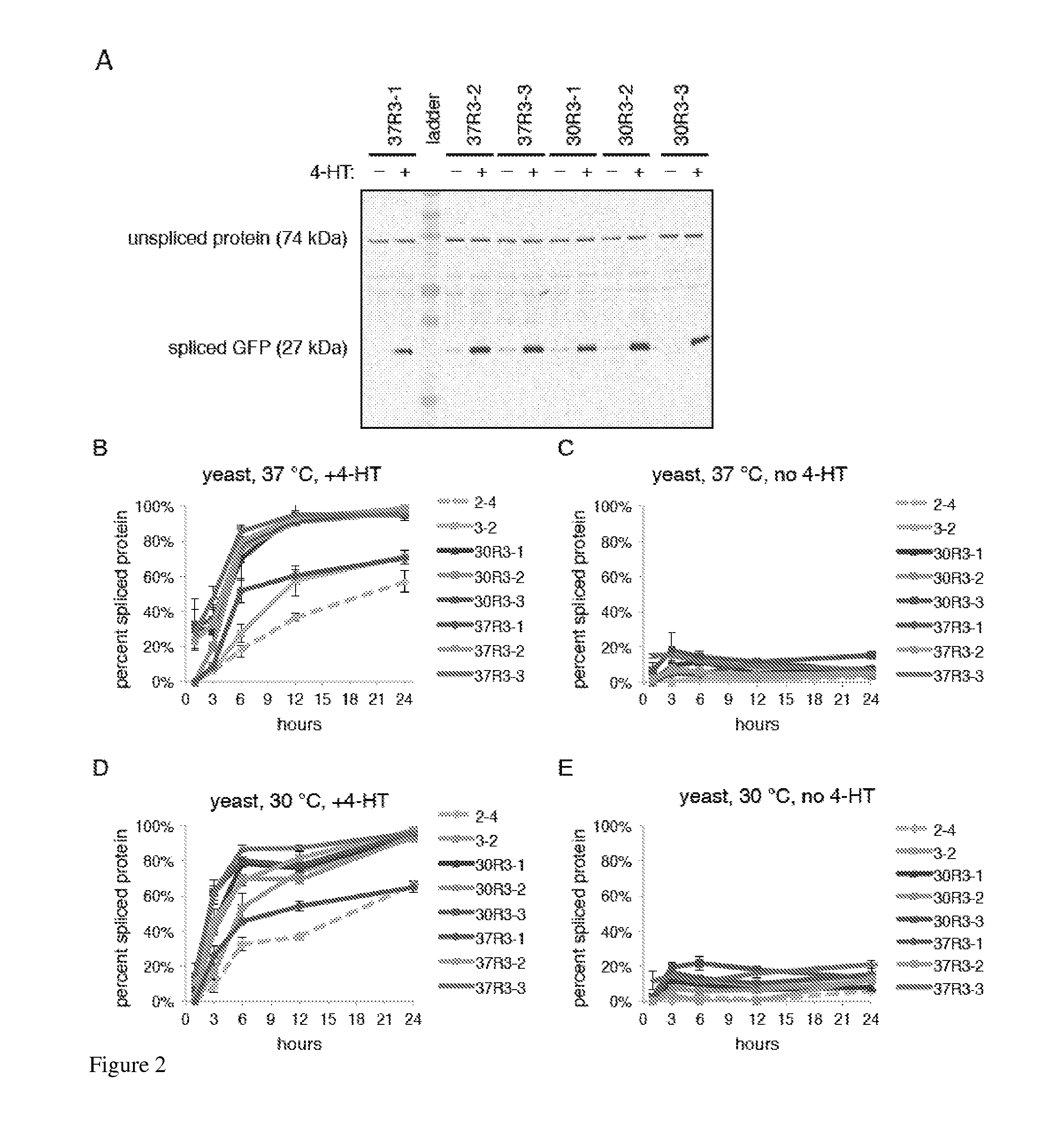
Elucidating the function of proteins in mammalian cells is particularly challenging due to the inherent complexity of these systems. Methods to study protein function in living cells ideally perturb the activity of only the protein of interest but otherwise maintain the natural state of the host cell or organism. Ligand-dependent inteins offer single-protein specificity and other desirable features as an approach to control protein function in cells post-translationally. Some aspects of this invention provide second-generation ligand-dependent inteins that splice to substantially higher yields and with faster kinetics in the presence of the cell-permeable small molecule 4-HT, especially at 37° C., while exhibiting comparable or improved low levels of background splicing in the absence of 4-HT, as compared to the parental inteins. These improvements were observed in four protein contexts tested in mammalian cells at 37° C., as well as in yeast cells assayed at 30° C. or 37° C. The newly evolved inteins described herein are therefore promising tools as conditional modulators of protein structure and function in yeast and mammalian cells.
Owner:PRESIDENT & FELLOWS OF HARVARD COLLEGE
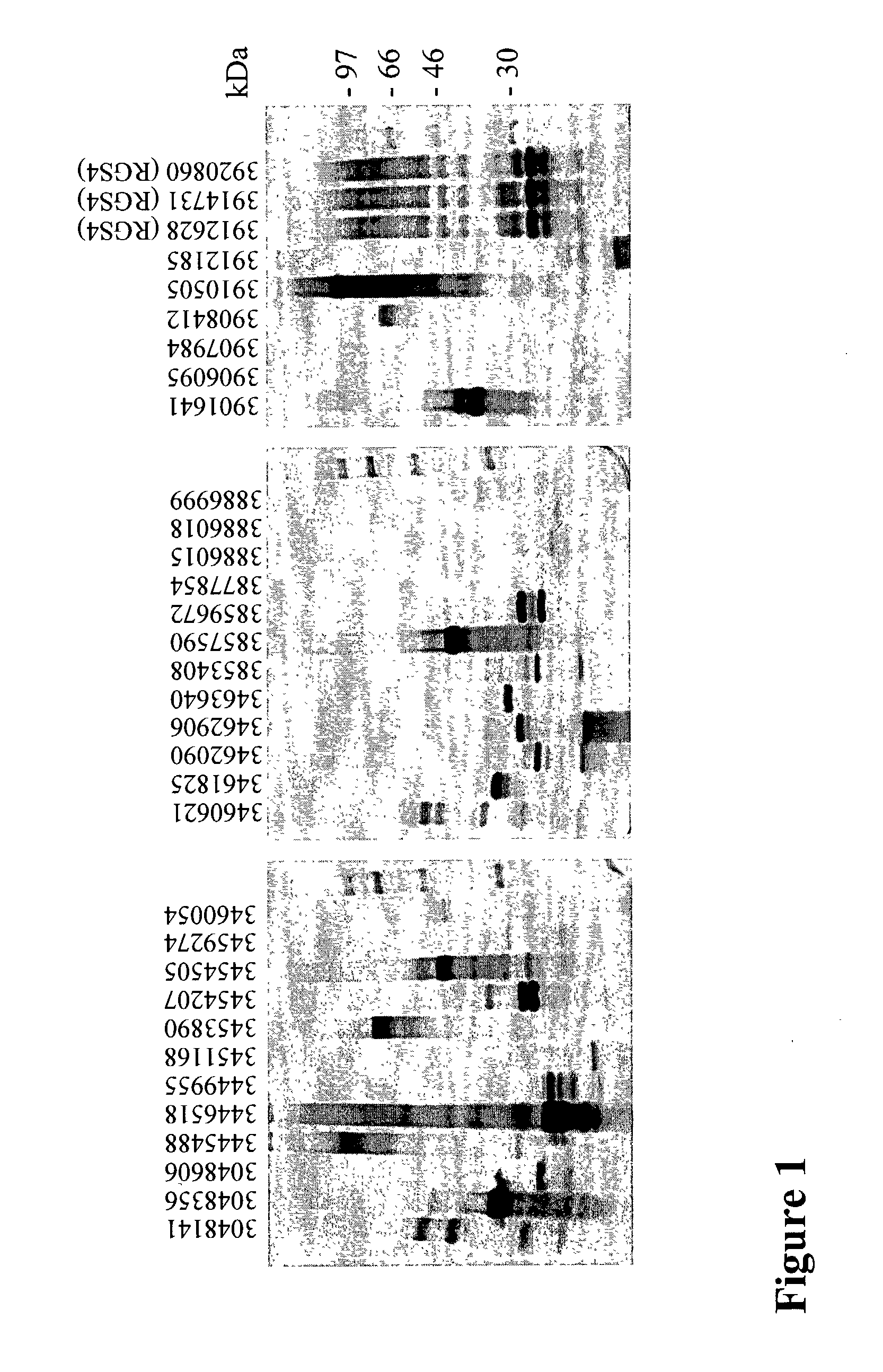
Process for analyzing protein samples
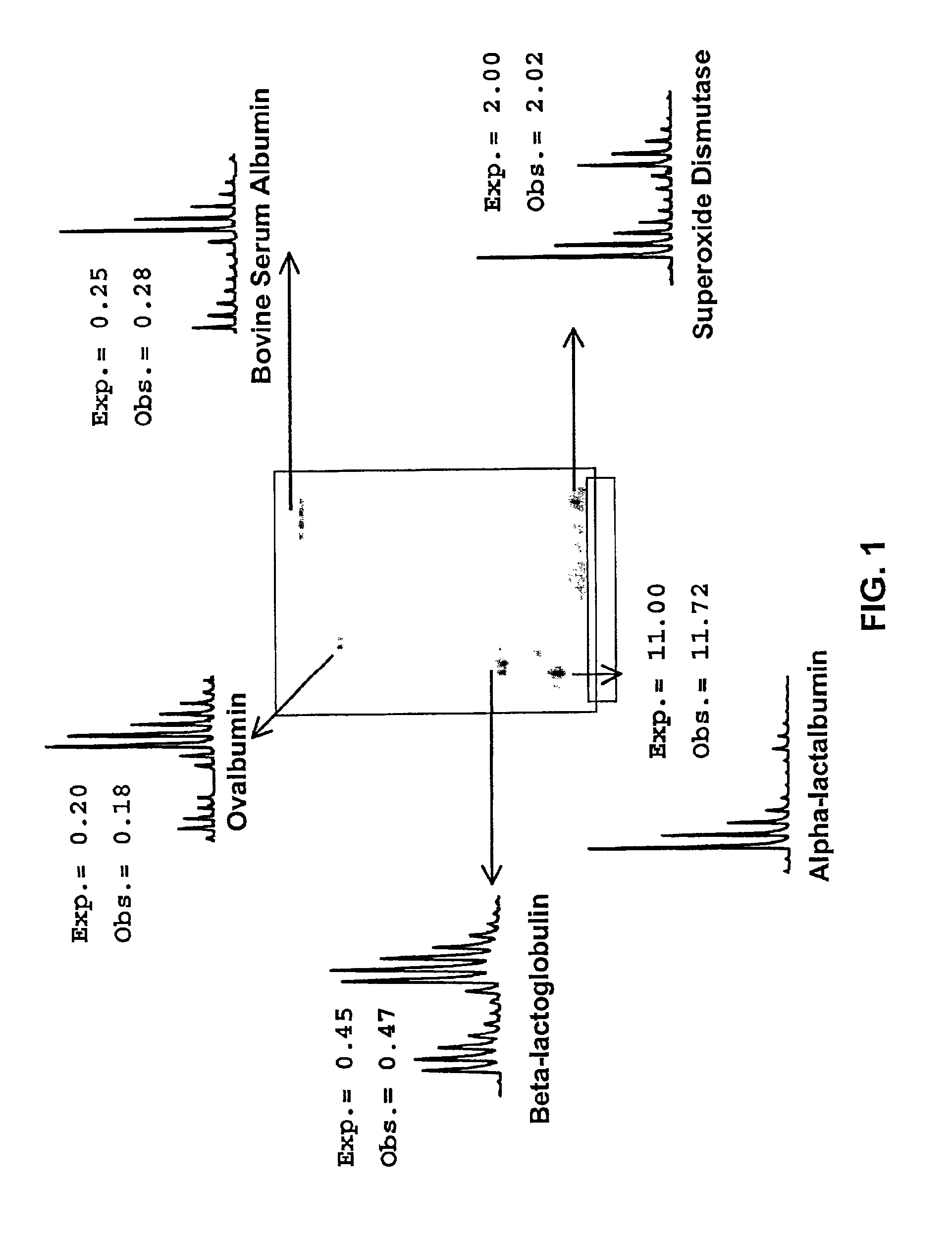
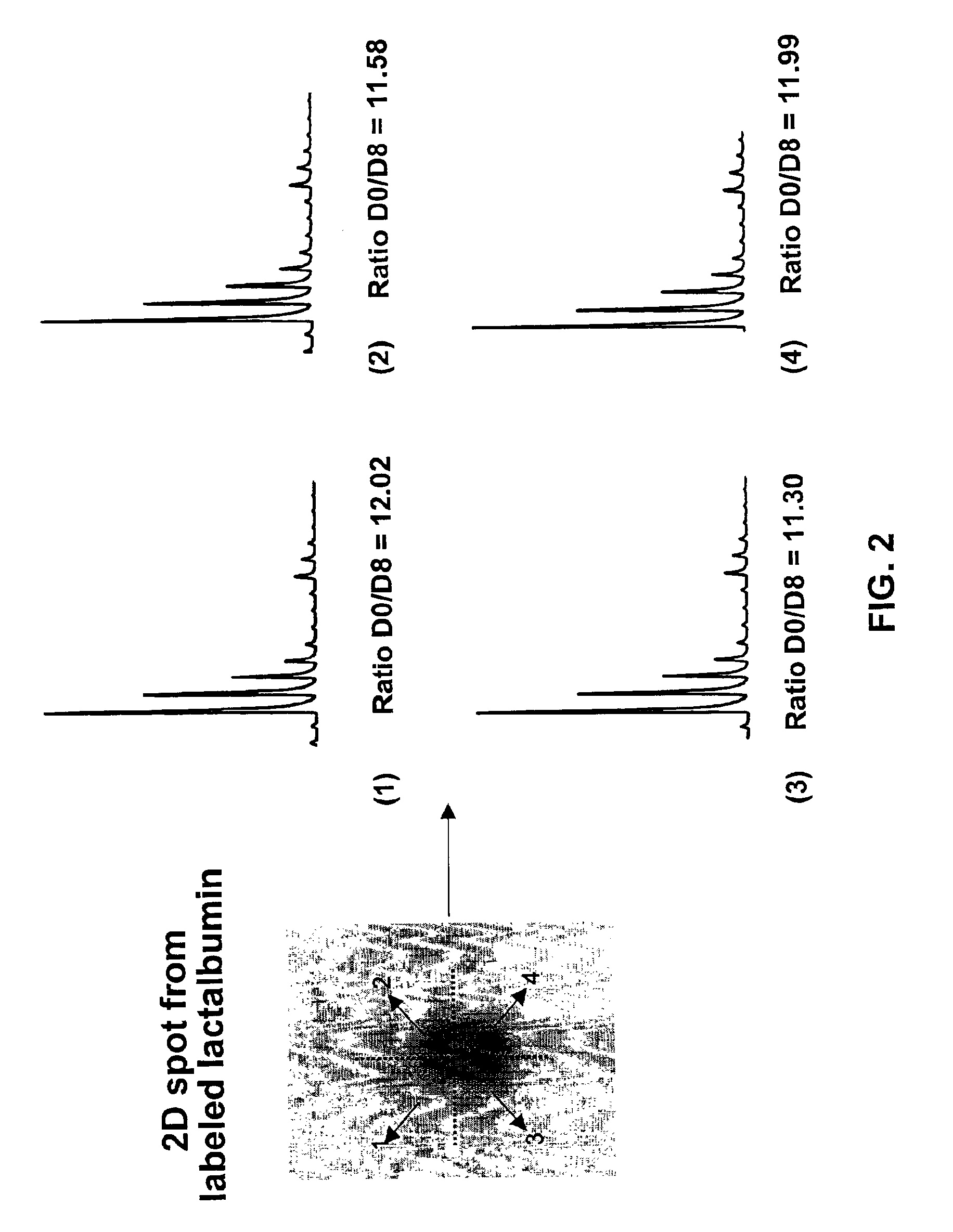
InactiveUS7045296B2Reduce disulfide bondEasy labelingSamplingMicrobiological testing/measurementProtein expression profileUnit operation
Methods using gel electrophoresis and mass spectrometry for the rapid, quantitative analysis of proteins or protein function in mixtures of proteins derived from two or more samples in one unit operation are disclosed. In one embodiment the method includes (a) preparing an extract of proteins from each of at least two different samples; (b) providing a set of substantially chemically identical and differentially isotopically labeled protein reagents; (c) reacting the extract of proteins from different samples of step (a) with a different isotopically labeled reagent from the set of step (b) to provide two or more sets of isotopically differentially labeled proteins; (d) mixing each of the two or more sets of isotopically labeled proteins to form a single mixture of isotopically differentially labeled proteins; (e) electrophoresing the mixture of step (d) by an electrophoresing method capable of separating proteins within the mixture; (f) digesting at least a portion of one or more separated proteins of step (e) and (g) detecting the difference in the expression levels of the proteins in the two samples by mass spectrometry based on one or more peptides in the sample of labeled peptides. The analytical method can be used for qualitative and particularly for quantitative analysis of global protein expression profiles in cells and tissues, i.e. the quantitative analysis of proteomes.
Owner:DH TECH DEVMENT PTE +1
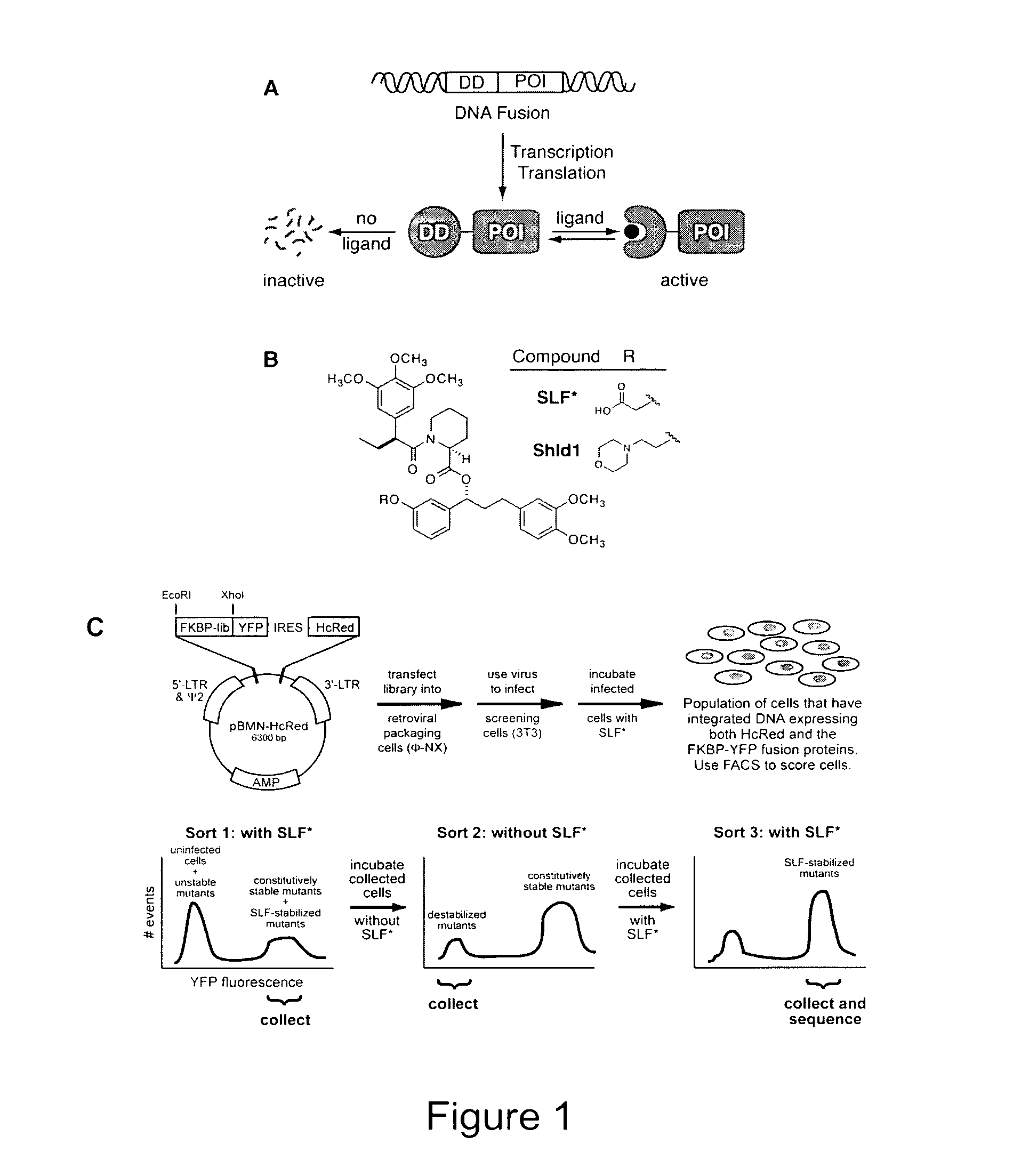
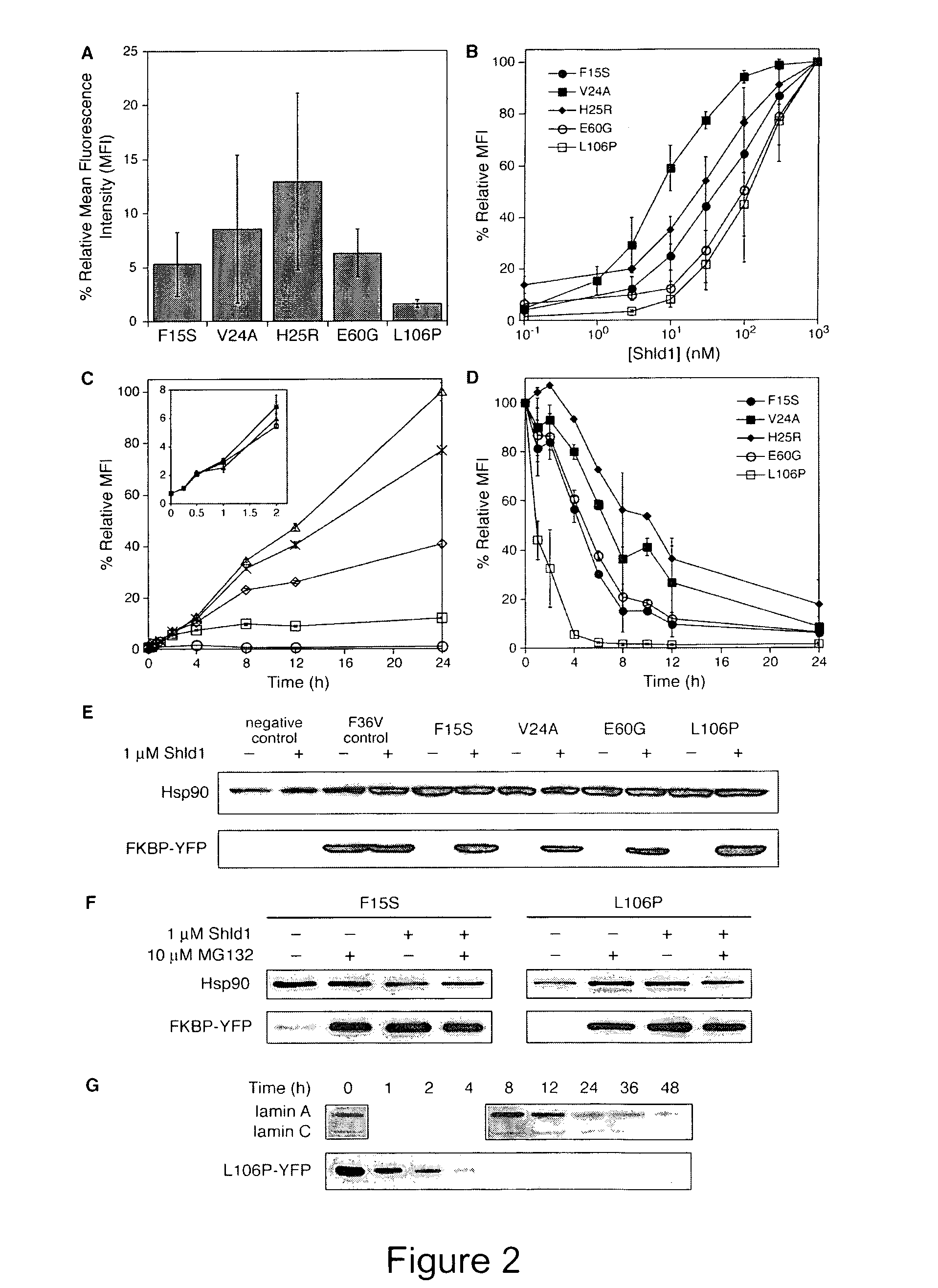
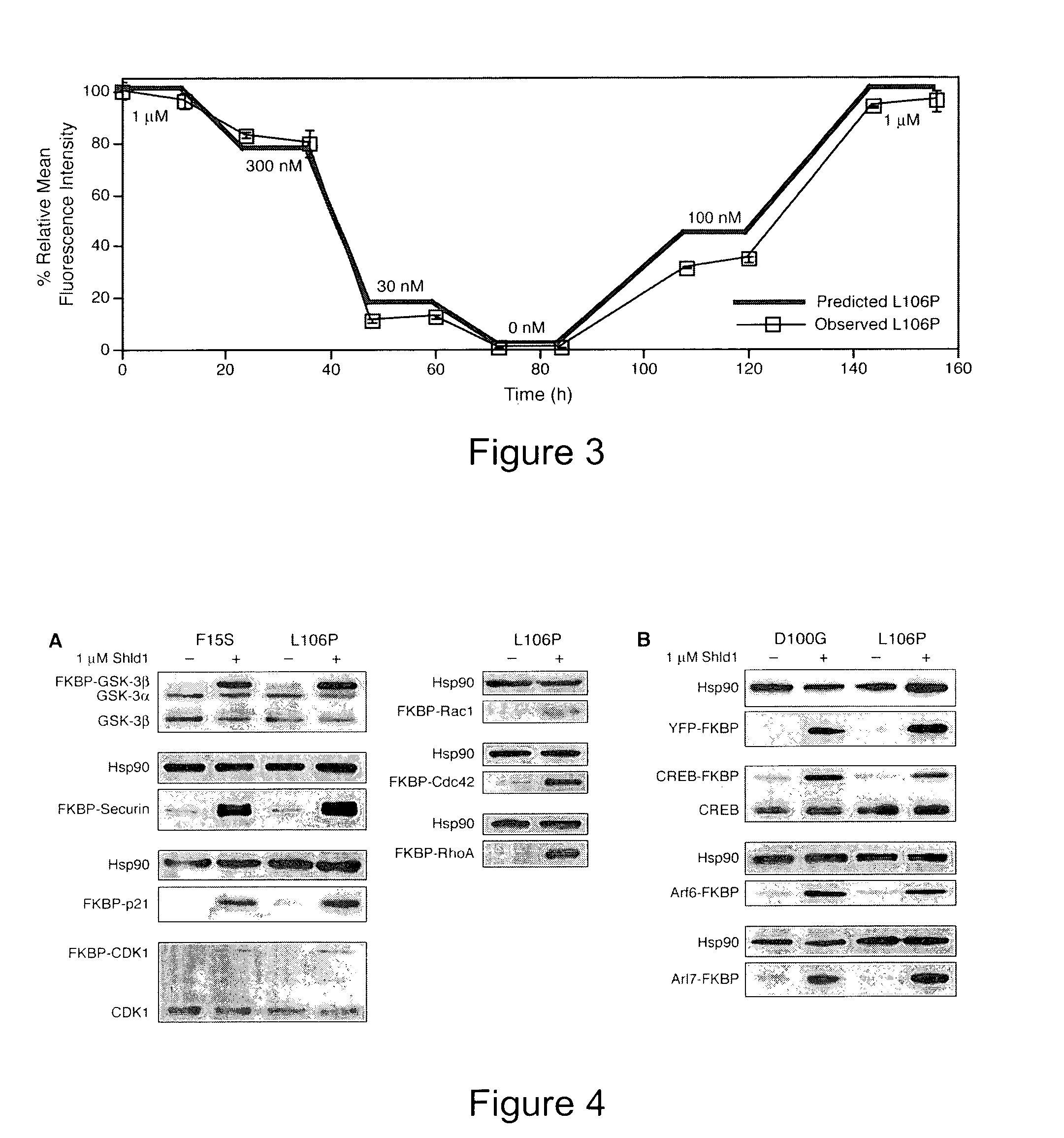
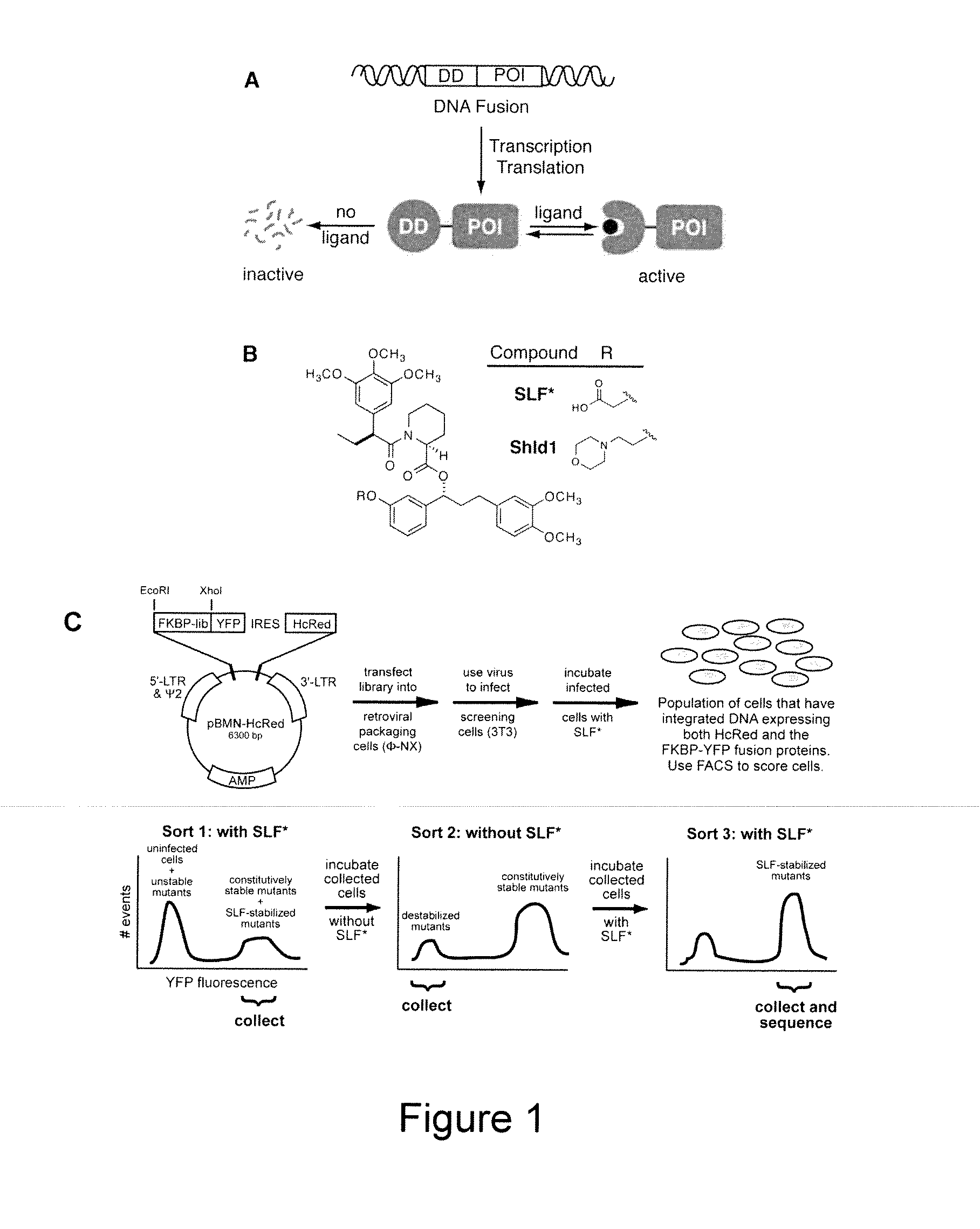
Method for regulating protein function in cells using synthetic small molecules
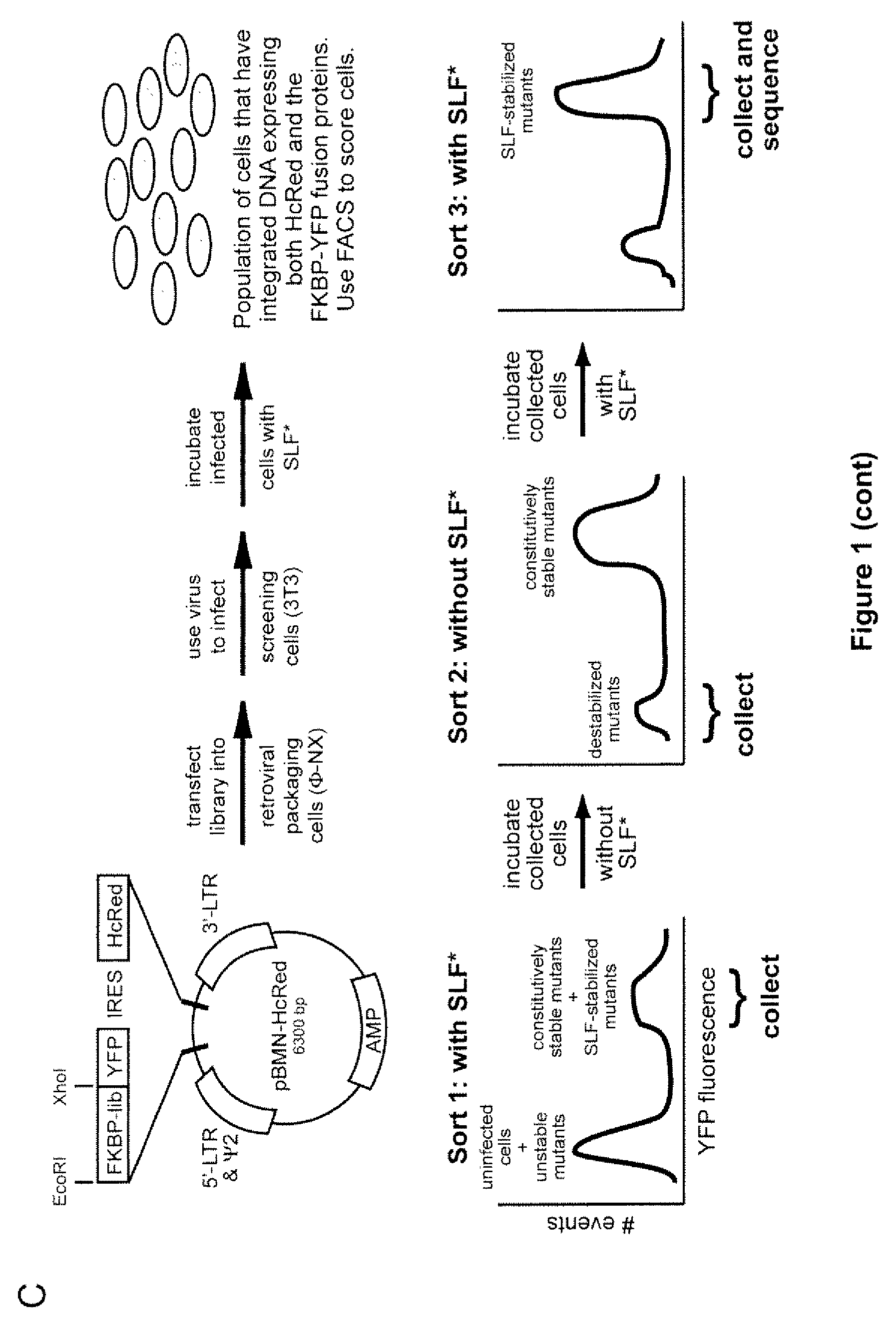
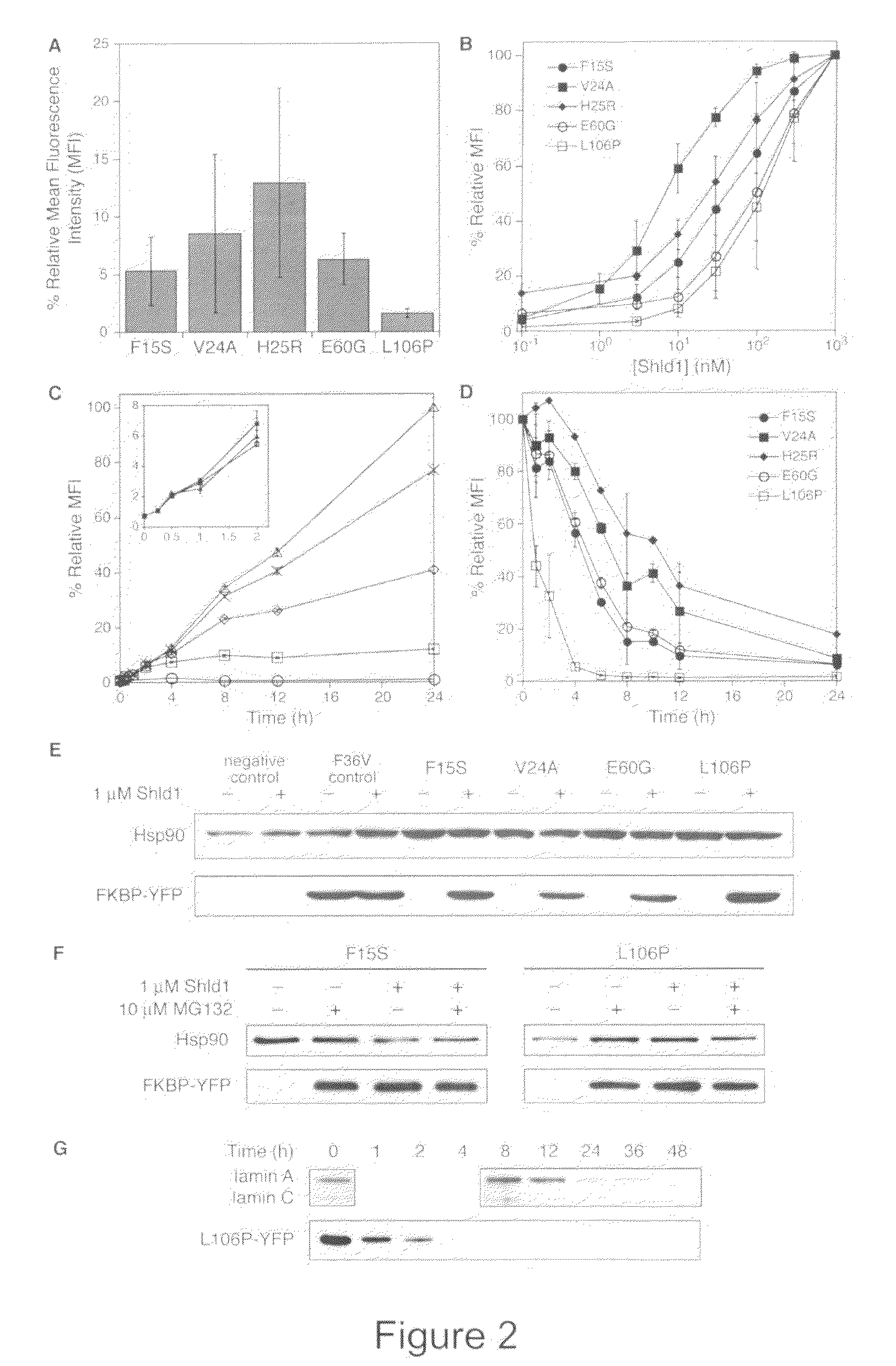
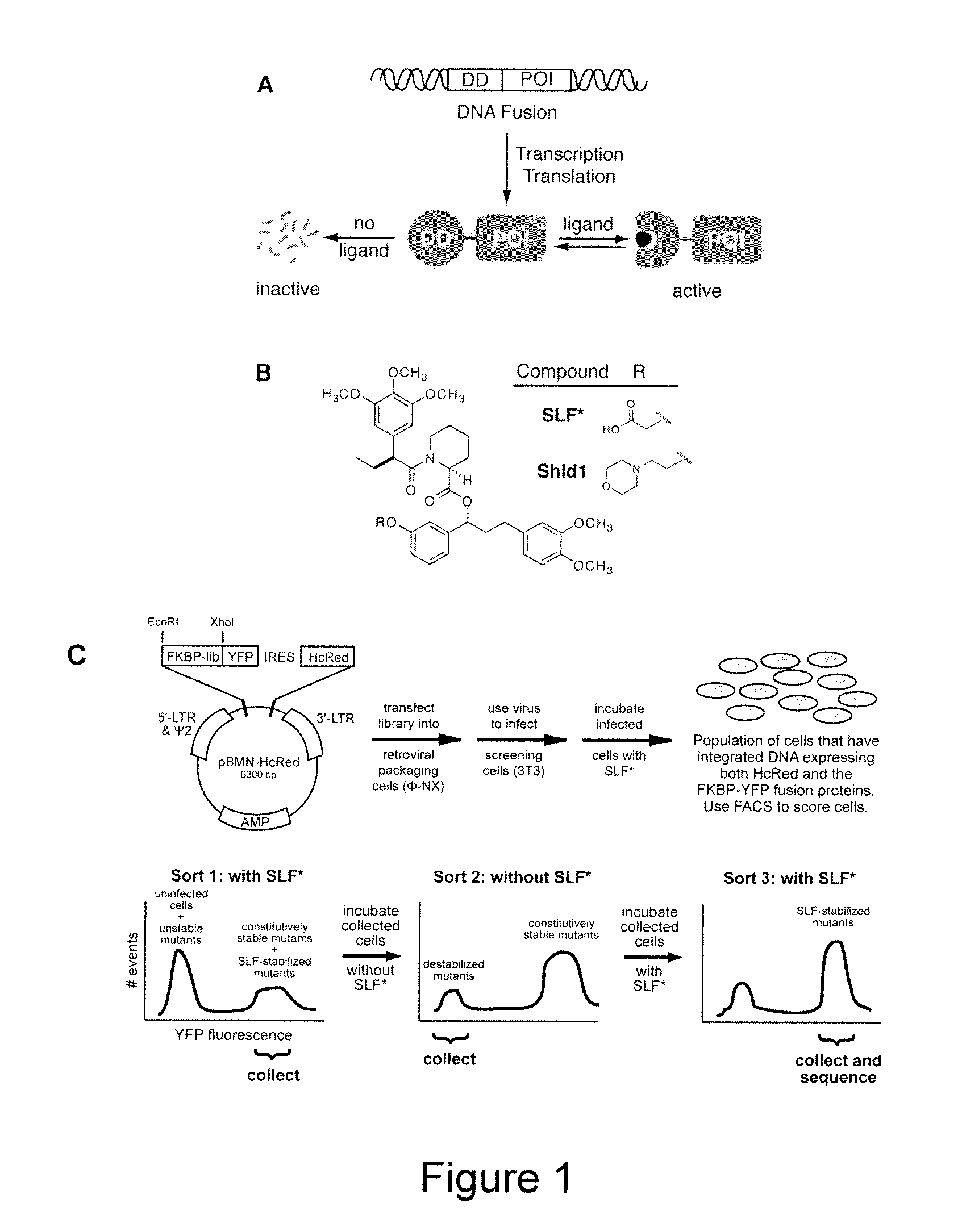
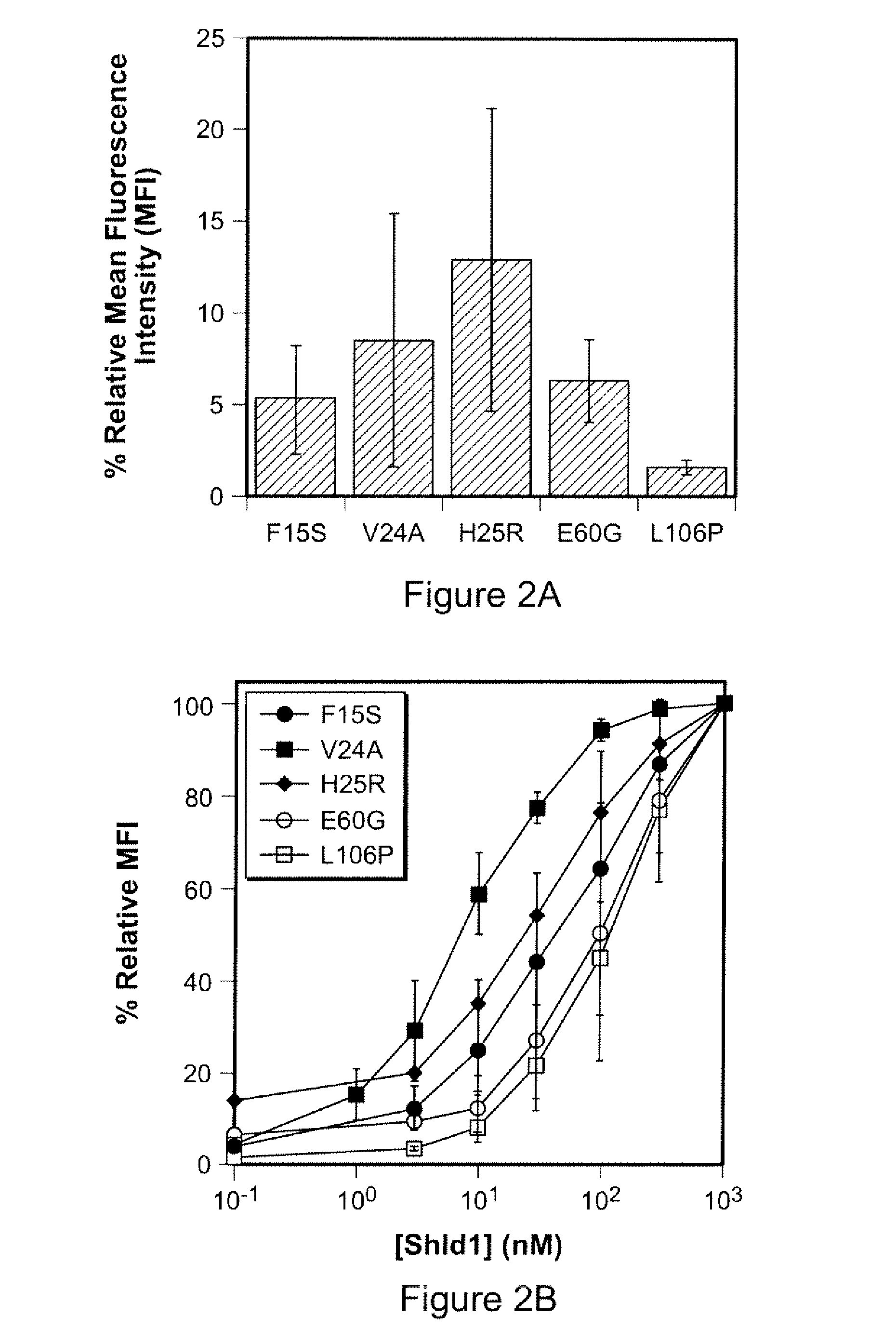
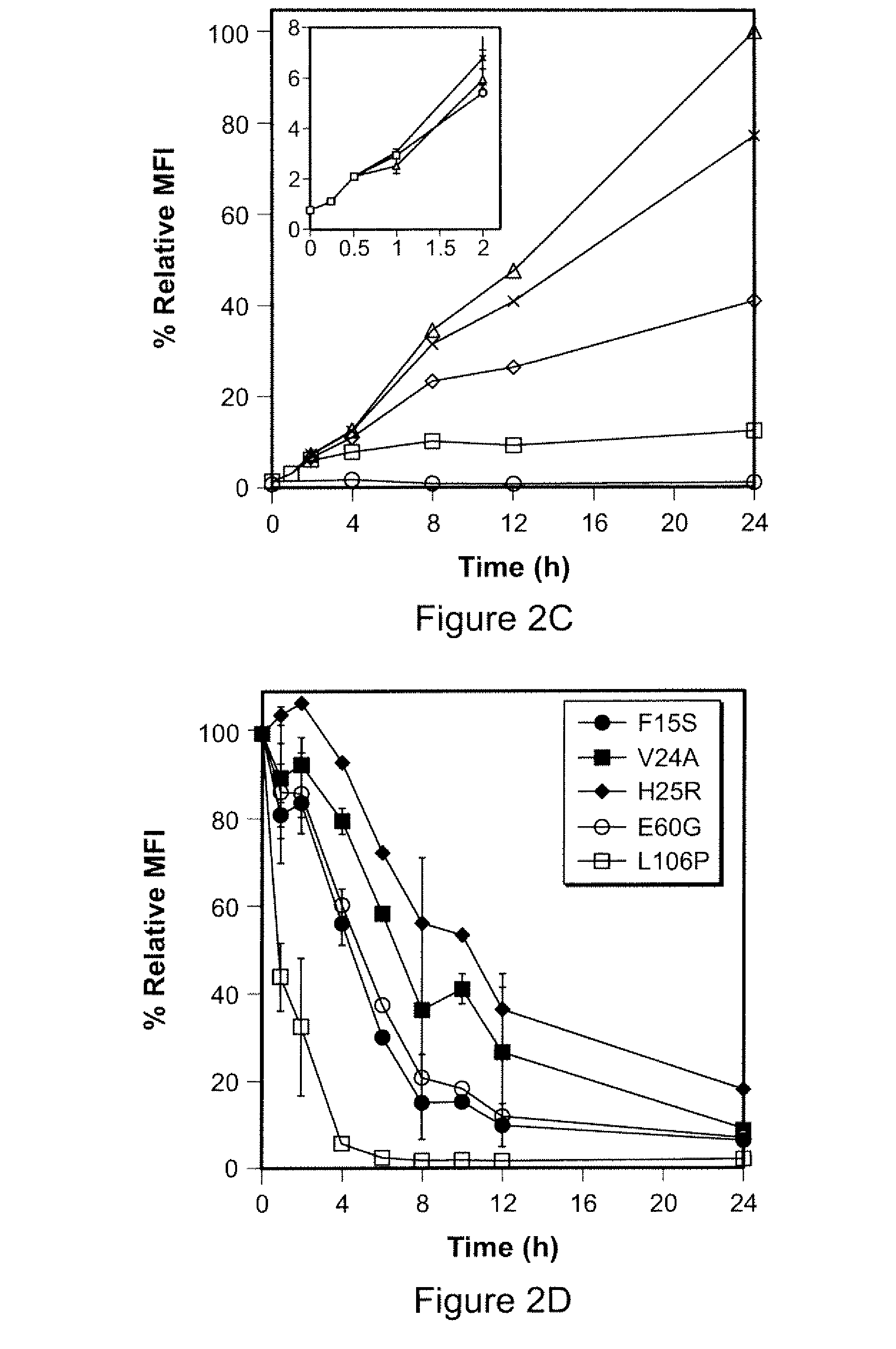
Methods and compositions for the rapid and reversible destabilizing of specific proteins using cell-permeable, synthetic molecules are described. Stability-affecting proteins, e.g., derived from FKBP and DHFR proteins are fused to a protein of interest and the presence or absence of the ligand is used to modulate the stability of the fusion protein.
Owner:THE BOARD OF TRUSTEES OF THE LELAND STANFORD JUNIOR UNIV
Methods for conducting assays for enzyme activity on protein microarrays
The present invention relates to methods of conducting assays for enzymatic activity on microarrays useful for the large-scale study of protein function, screening assays, and high-throughput analysis of enzymatic reactions. The invention relates to methods of using protein chips to assay the presence, amount, activity and / or function of enzymes present in a protein sample on a protein chip. In particular, the methods of the invention relate to conducting enzymatic assays using a microarray wherein a protein and a substance are immobilized on the surface of a solid support and wherein the protein and the substance are in proximity to each other sufficient for the occurrence of an enzymatic reaction between the substance and the protein. The invention also relates to microarrays that have an enzyme and a substrate immobilized on their surface wherein the enzyme and the substrate are in proximity to each other sufficient for the occurrence of an enzymatic reaction between the enzyme and the substrate.
Owner:LIFE TECH CORP
Four-gene-deletion weak-toxin strain for African swine fever viruses and application of four-gene-deletion weak-toxin strain
InactiveCN110551695AEasy to solveViral antigen ingredientsMicrobiological testing/measurementAfrican swine feverToxin
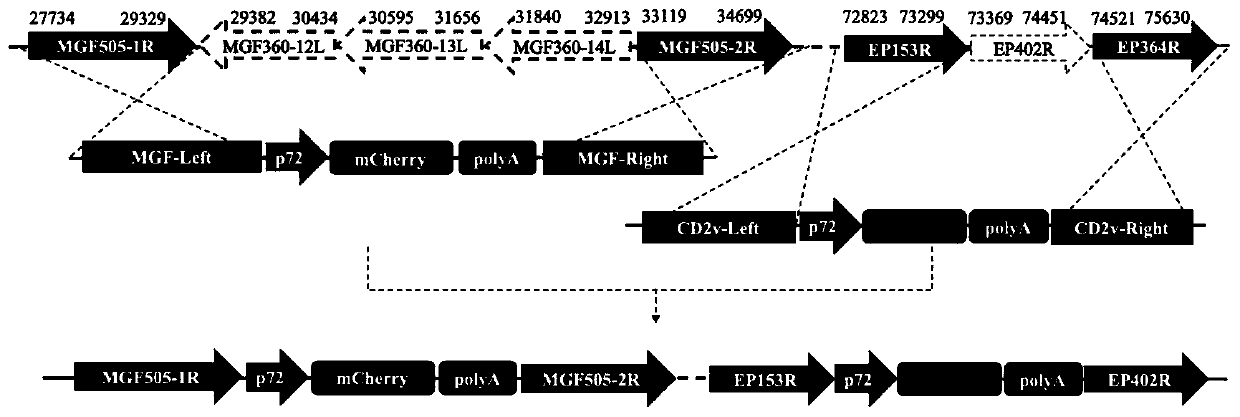
The invention discloses a four-gene-deletion weak-toxin strain for African swine fever viruses. The weak-toxin strain is the four-gene-deletion weak-toxin strain for an African swine fever virus SY18separation strain, and the following gene function protein is deleted: CD2v gene coding products and three multigene family genes( MGF360-12L, MGF360-13L and MGF360-14L ) coding products. The invention further discloses an application of the weak-toxin strain of the African swine fever viruses to preparation of vaccines for preventing or treating African swine fever. The weak-toxin strain of the African swine fever viruses can provide complete immunoprotection effect on attack of ASFV parent toxin strains, is high in safety, and is suitable for being used as vaccine candidate strains for preventing the African swine fever.
Owner:SHANGHAI VETERINARY RES INST CHINESE ACAD OF AGRI SCI
Protein chips for high throughput screening of protein activity
InactiveUS20030207467A1Material nanotechnologyBioreactor/fermenter combinationsHigh-Throughput Screening MethodsChemical reaction
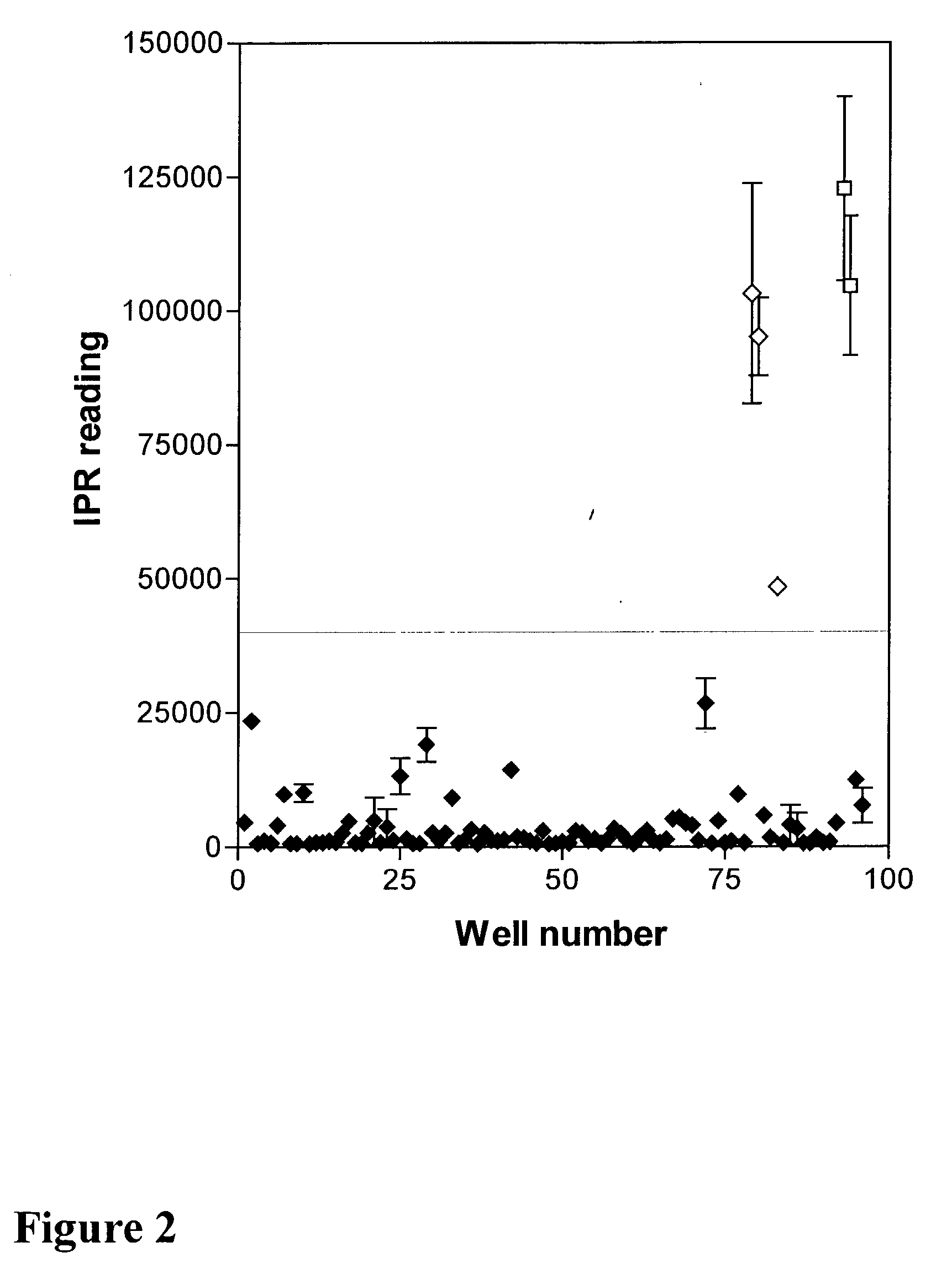
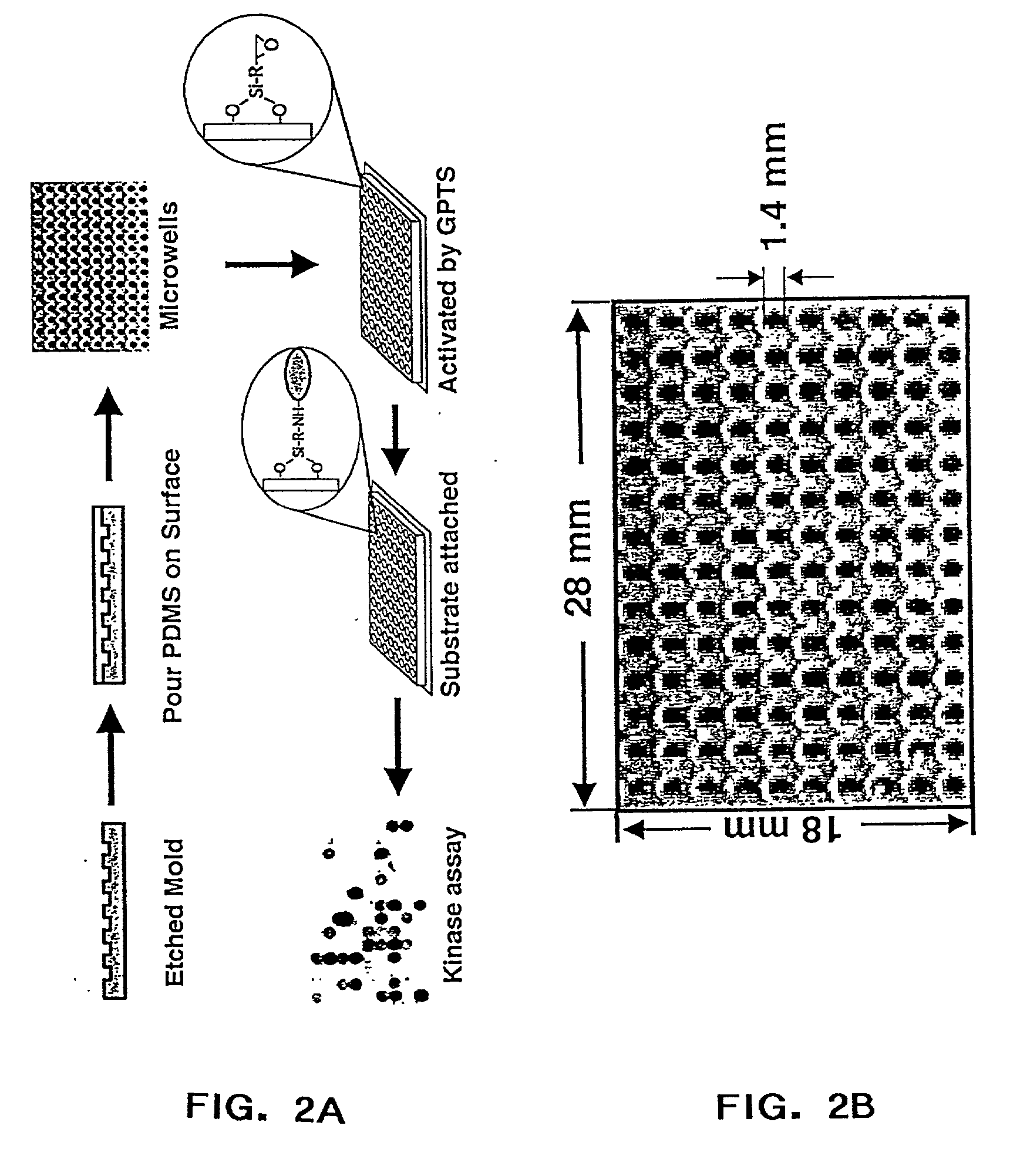
The present invention relates to protein chips useful for the large-scale study of protein function where the chip contains densely packed reaction wells. The invention also relates to methods of using protein chips to assay simultaneously the presence, amount, and / or function of proteins present in a protein sample or on one protein chip, or to assay the presence, relative specificity, and binding affinity of each probe in a mixture of probes for each of the proteins on the chip. The invention also relates to methods of using the protein chips for high density and small volume chemical reactions. Also, the invention relates to polymers useful as protein chip substrates and methods of making protein chips. The invention further relates to compounds useful for the derivatization of protein chip substrates.
Owner:YALE UNIV
Systems and methods for silencing expression of a gene in a cell and uses thereof
InactiveUS20060178297A1Minimal disturbanceFast and nearly-complete uptakePeptide/protein ingredientsMicroencapsulation basedPrimary cellNeuron
The present invention provides RNA-based systems that are capable of silencing expression of a gene in a cell. Also provided are pharmaceutical compositions, cells, and kits that include the systems; cultures of primary cells that have been contacted with the systems; use of the systems in methods of studying protein function in one or more cells; and use of the systems in methods of studying interactions between neurons in culture; and use of the systems in a method of studying the function of mRNA. The present invention further provides methods for silencing expression of a gene in a cell, and methods for determining the function of a gene in a cell. Additionally, the present invention provides systems for use in genetic screening, and methods for performing genetic screening.
Owner:THE TRUSTEES OF COLUMBIA UNIV IN THE CITY OF NEW YORK
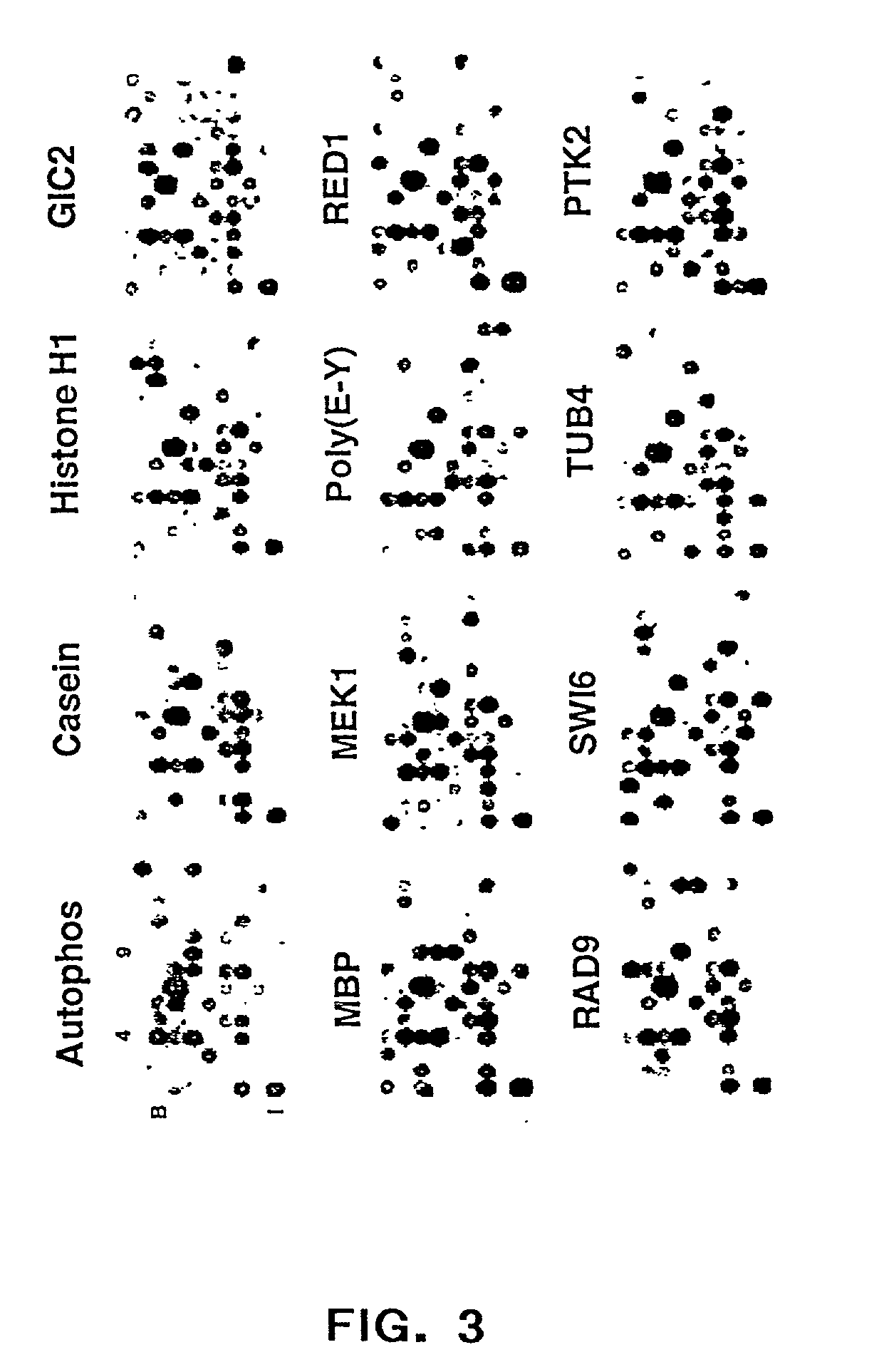
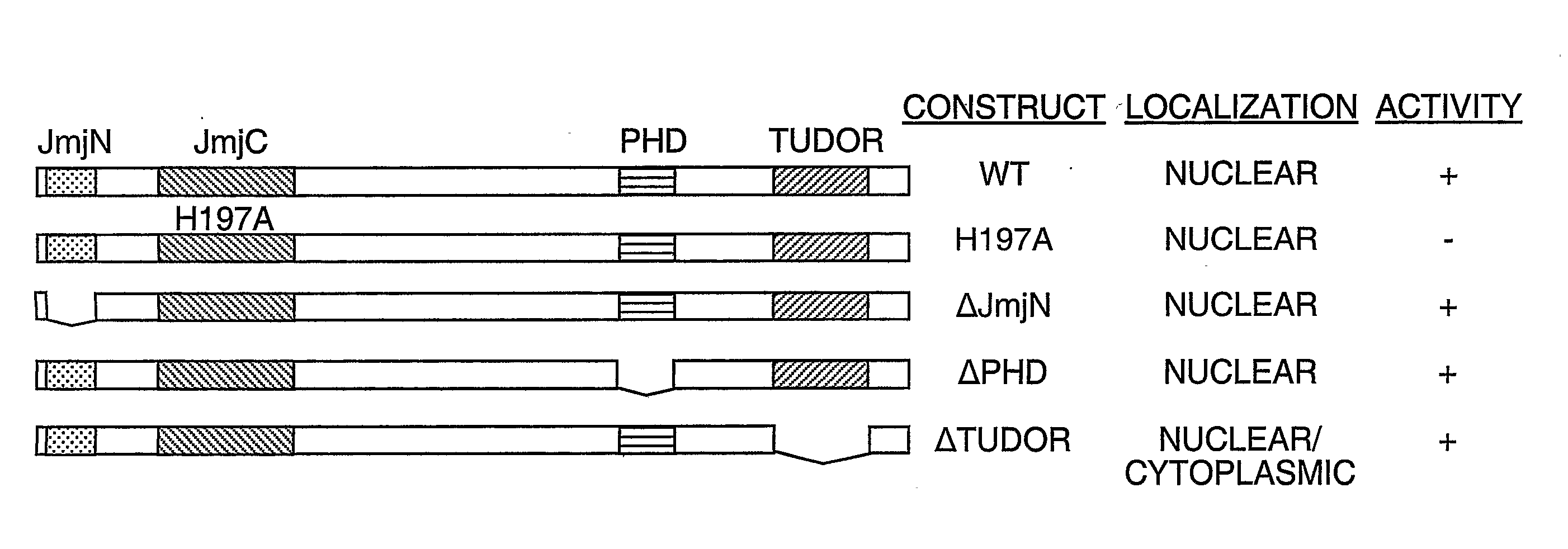
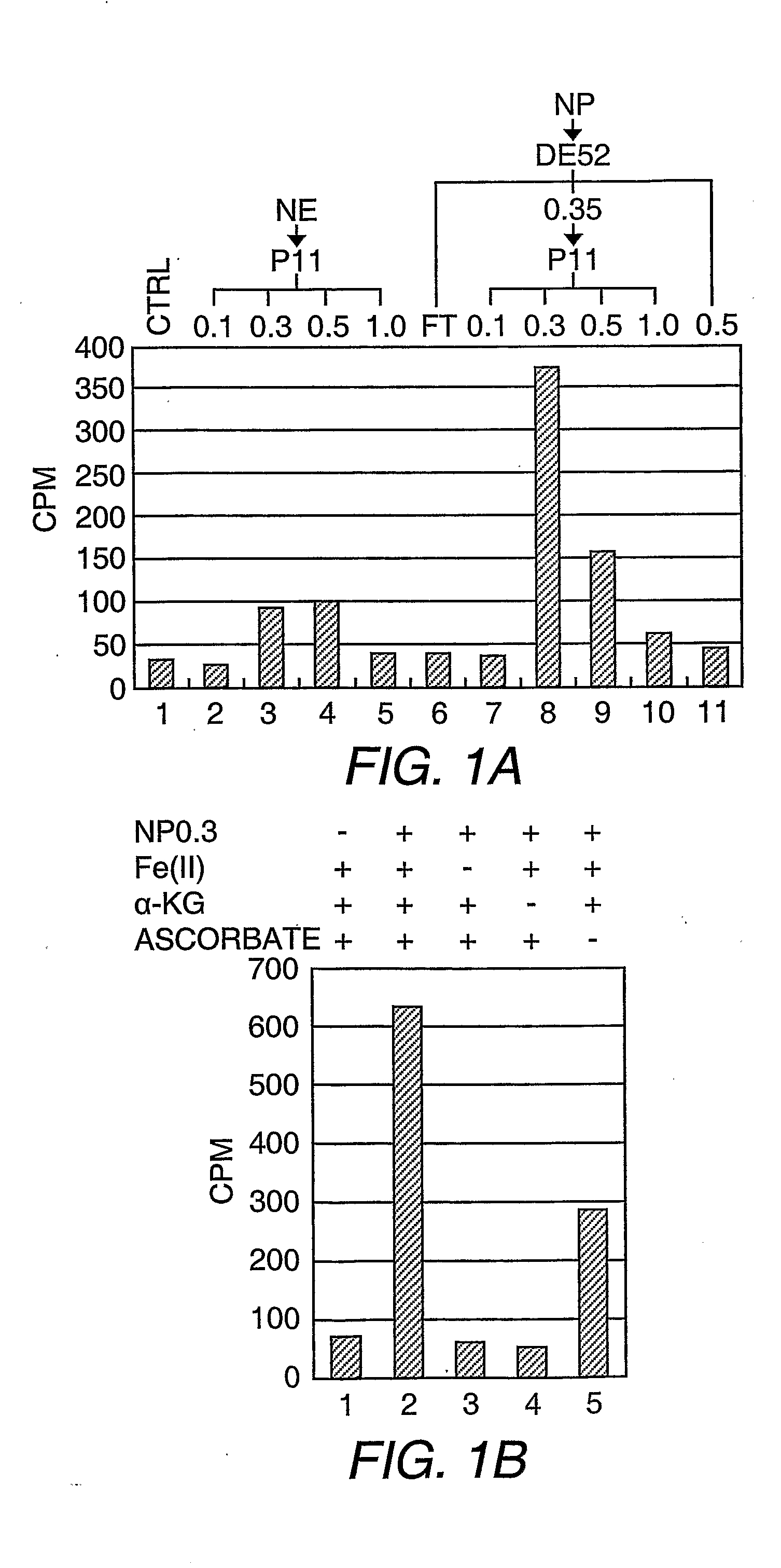
Protein demethylases comprising a jmjc domain
Post-translational modification, including protein methylation, plays an important role in regulating protein function. The present invention provides a novel assay for evaluating demethylase activity and the discovery of a family of protein demethylases comprising a novel demethylase motif.
Owner:THE UNIV OF NORTH CAROLINA AT CHAPEL HILL
Method for regulating protein function in cells using synthetic small molecules
Methods and compositions for the rapid and reversible destabilizing of specific proteins using cell-permeable, synthetic molecules are described. Stability-affecting proteins, e.g., derived from FKBP and DHFR proteins are fused to a protein of interest and the presence or absence of the ligand is used to modulate the stability of the fusion protein.
Owner:THE BOARD OF TRUSTEES OF THE LELAND STANFORD JUNIOR UNIV
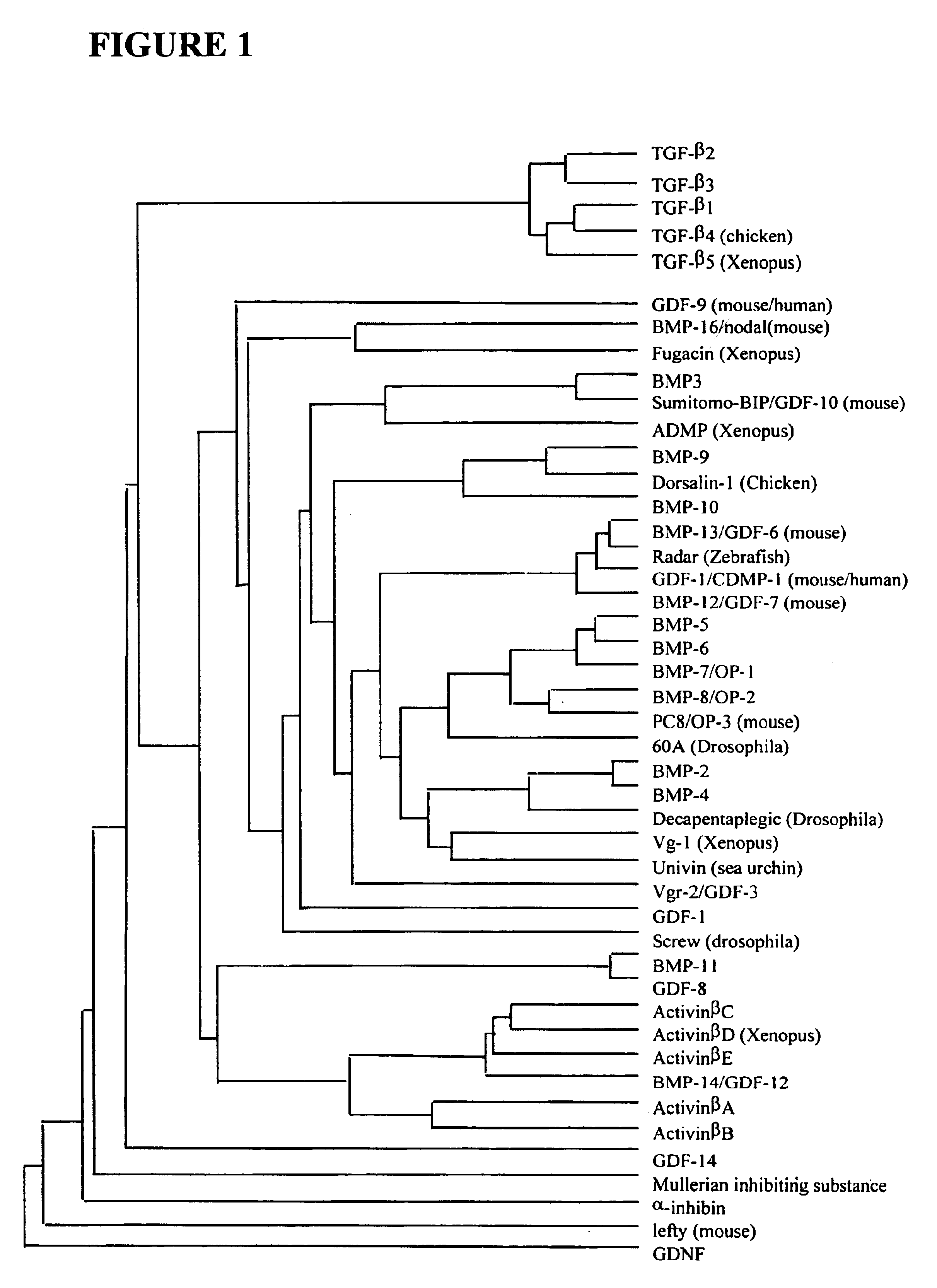
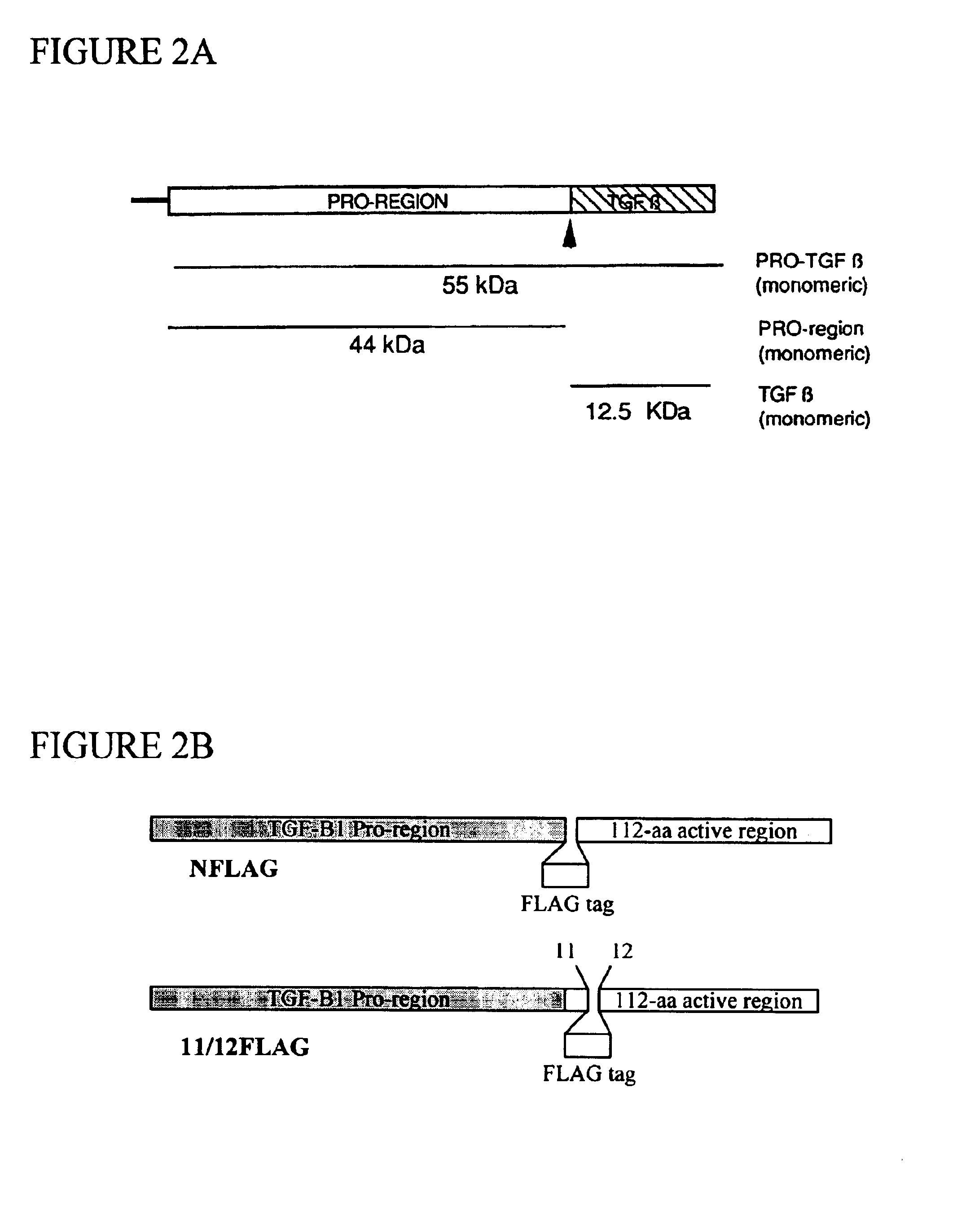
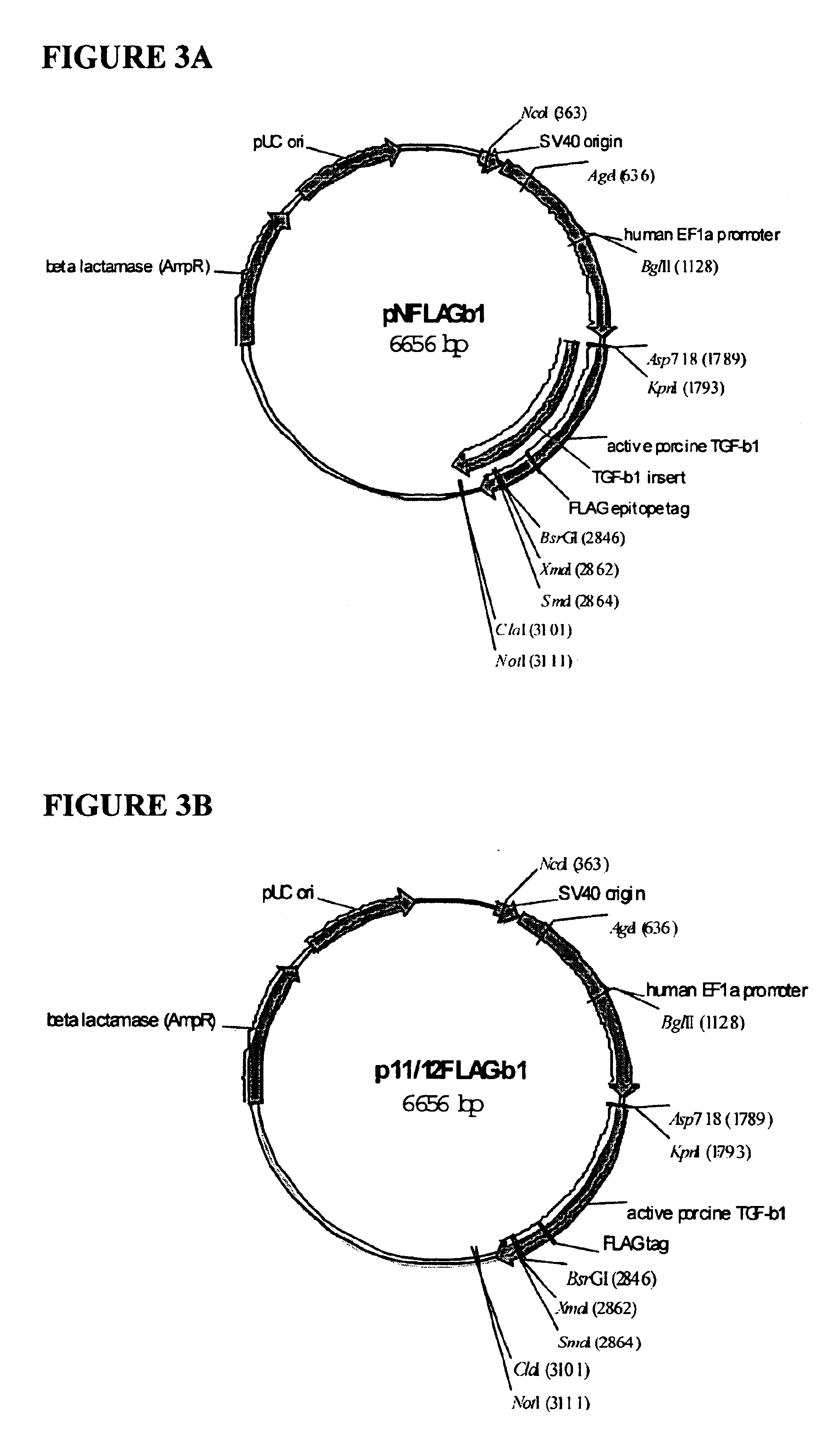
Functionalized TGF-beta fusion proteins
This disclosure relates to TGF-beta family protein fusions that display substantial native TGF-beta family protein function while also having an additional functionality conveyed by the addition of a functionalizing peptide domain. Such functionalizing peptide domain can be a tag peptide (e.g., an epitope tag, a purification tag, a molecular size differentiation tag, etc.) or a passenger or targeting protein. Also provided are methods of making these fusions, as well as methods of using them for diagnosis and diagnosis of various conditions, in measuring and monitoring levels of the fusion molecule in experimental systems and subjects, and in measuring and detecting receptor proteins.
Owner:GOVERNMENT OF THE UNITED STATE OF AMERICA AS REPRESENTED BY THE SEC OF THE DEPT OF HEALTH & HUMAN SERVICES THE
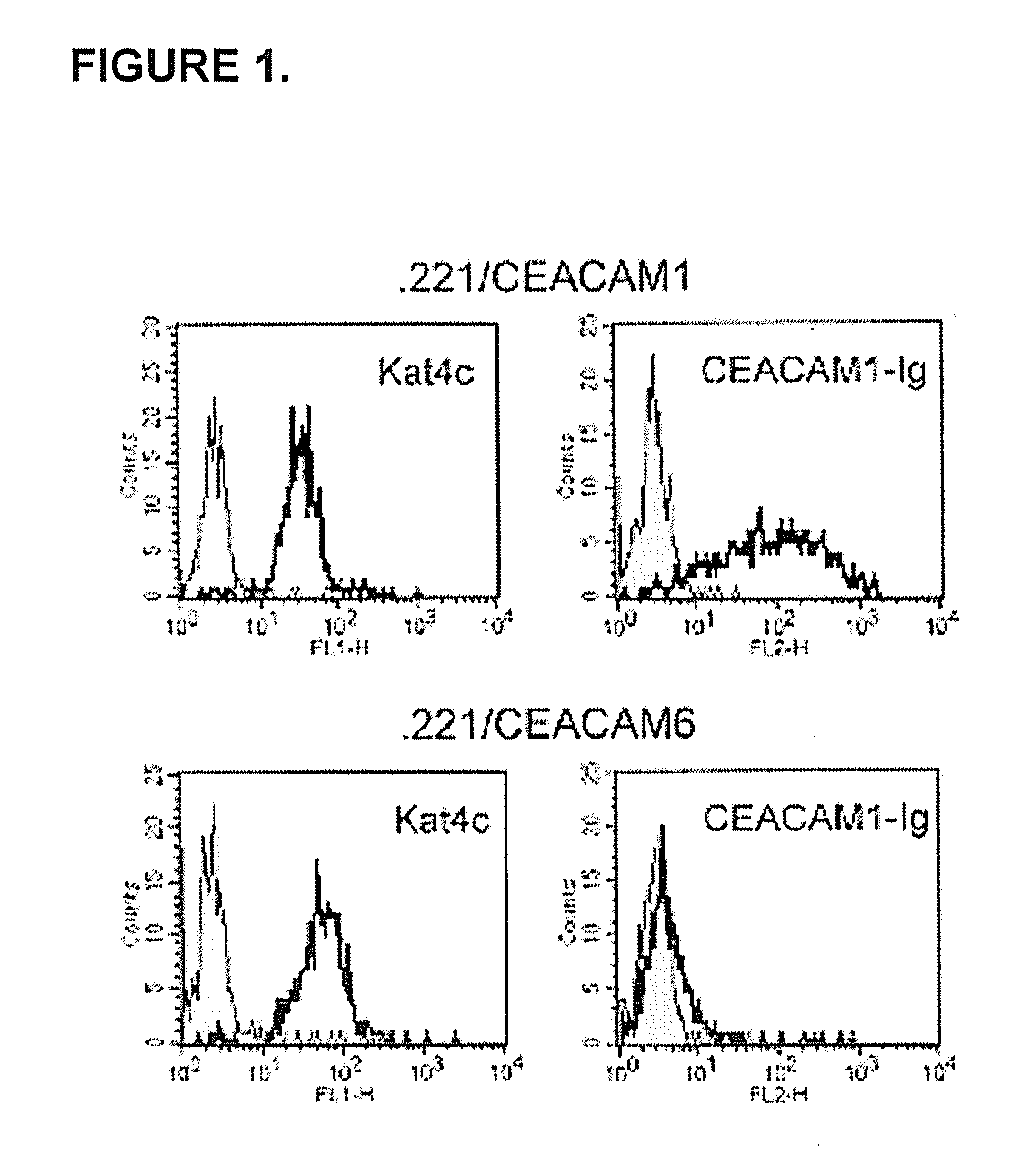
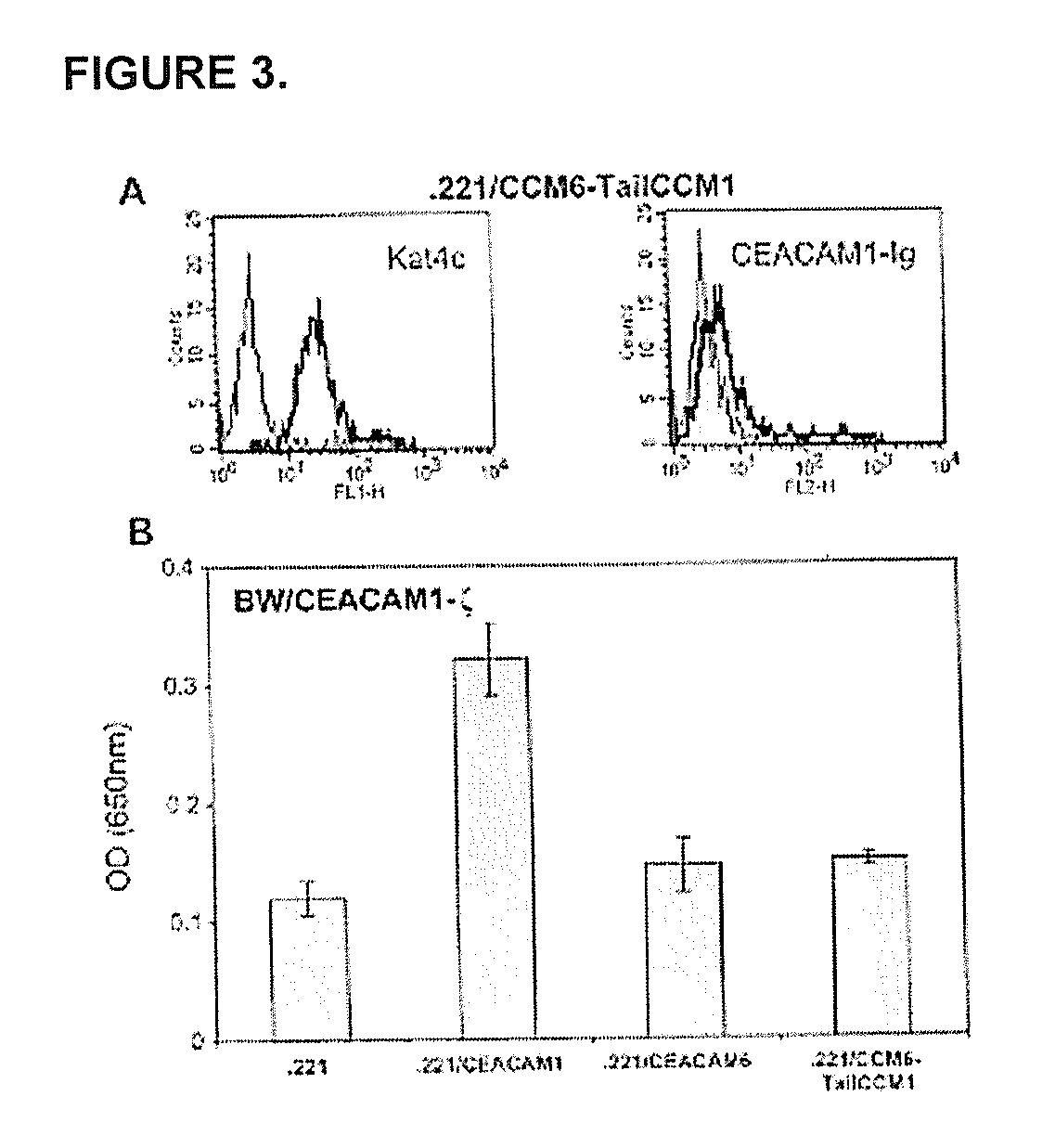
Modulation of Immunity and Ceacam1 Activity
ActiveUS20070110668A1Good curative effectCompounds screening/testingBiocideLymphocyteSpecific immunity
The present technology comprises methods for regulating an the immune system, and in particular methods for the regulation of a specific immune response, including the regulation of lymphocyte activity. Methods of the present technology comprise both the negative and positive modulation of CEACAM1 protein function.
Owner:MARKEL GAL
Rapid quantitative analysis of proteins or protein function in complex mixtures
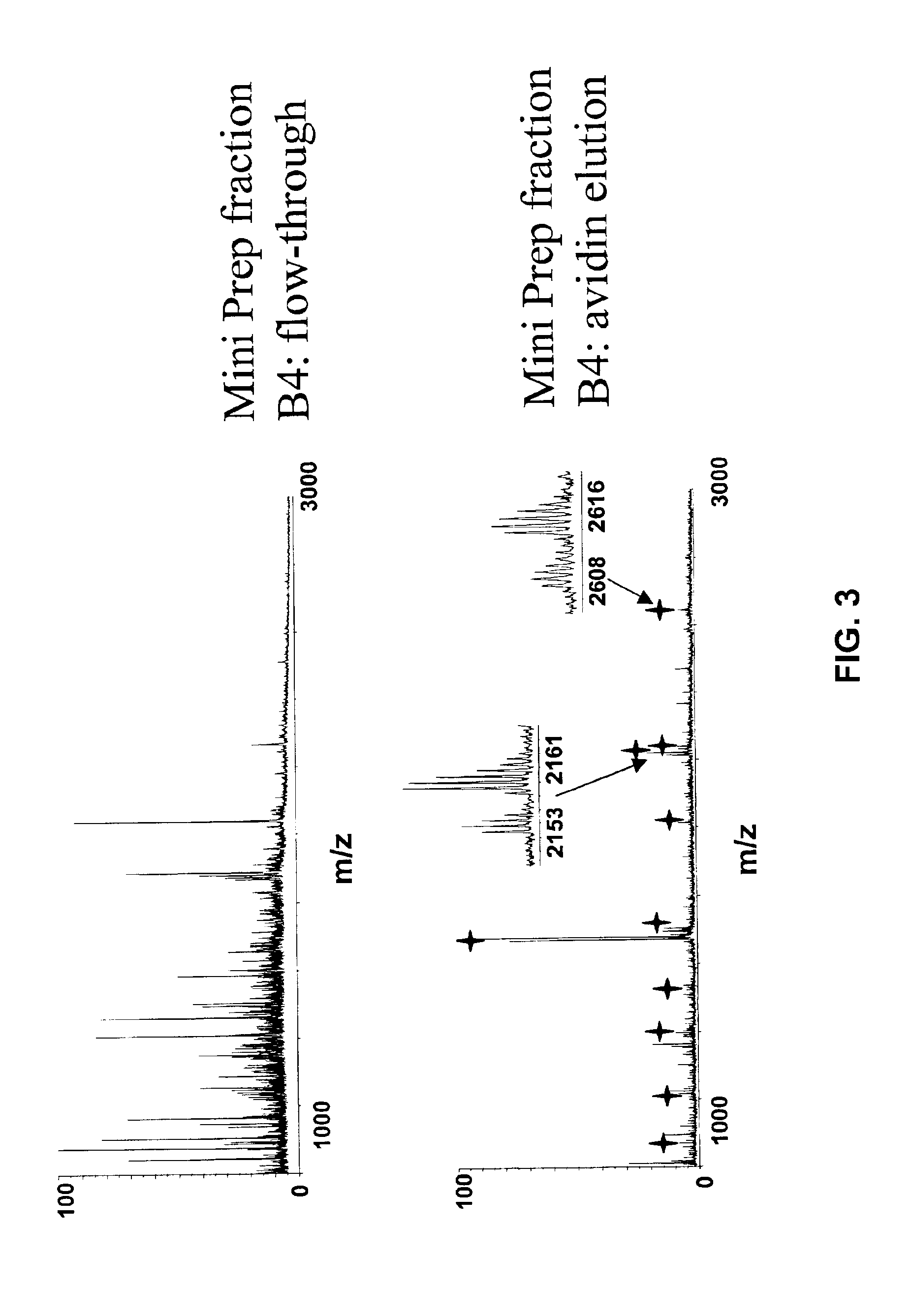
InactiveUS7544518B2Facilitates quantitative determinationFacilitates quantitative determination of the absolute amountsComponent separationMaterial analysis by electric/magnetic meansIsotopic labelingProtein expression profile
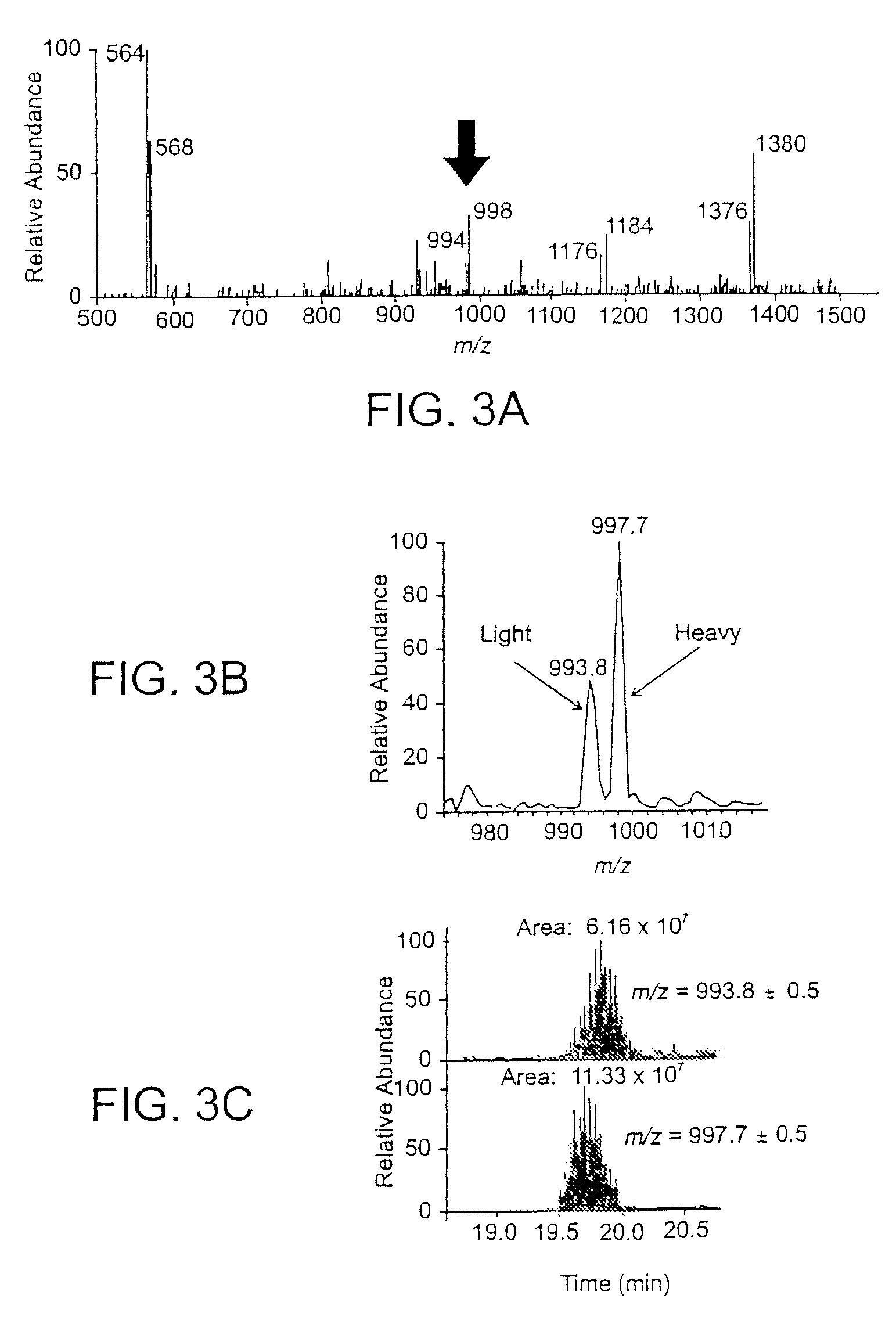
Analytical reagents and mass spectrometry-based methods using these reagents for the rapid, and quantitative analysis of proteins or protein function in mixtures of proteins. The methods employ affinity labeled protein reactive reagents having three portions: an affinity label (A) covalently linked to a protein reactive group (PRG) through a linker group (L). The linker may be differentially isotopically labeled, e.g., by substitution of one or more atoms in the linker with a stable isotope thereof. These reagents allow for the selective isolation of peptide fragments or the products of reaction with a given protein (e.g., products of enzymatic reaction) from complex mixtures. The isolated peptide fragments or reaction products are characteristic of the presence of a protein or the presence of a protein function in those mixtures. Isolated peptides or reaction products are characterized by mass spectrometric (MS) techniques. The reagents also provide for differential isotopic labeling of the isolated peptides or reaction products which facilitates quantitative determination by mass spectrometry of the relative amounts of proteins in different samples. The methods of this invention can be used for qualitative and quantitative analysis of global protein expression profiles in cells and tissues, to screen for and identify proteins whose expression level in cells, tissue or biological fluids is affected by a stimulus or by a change in condition or state of the cell, tissue or organism from which the sample originated.
Owner:UNIV OF WASHINGTON
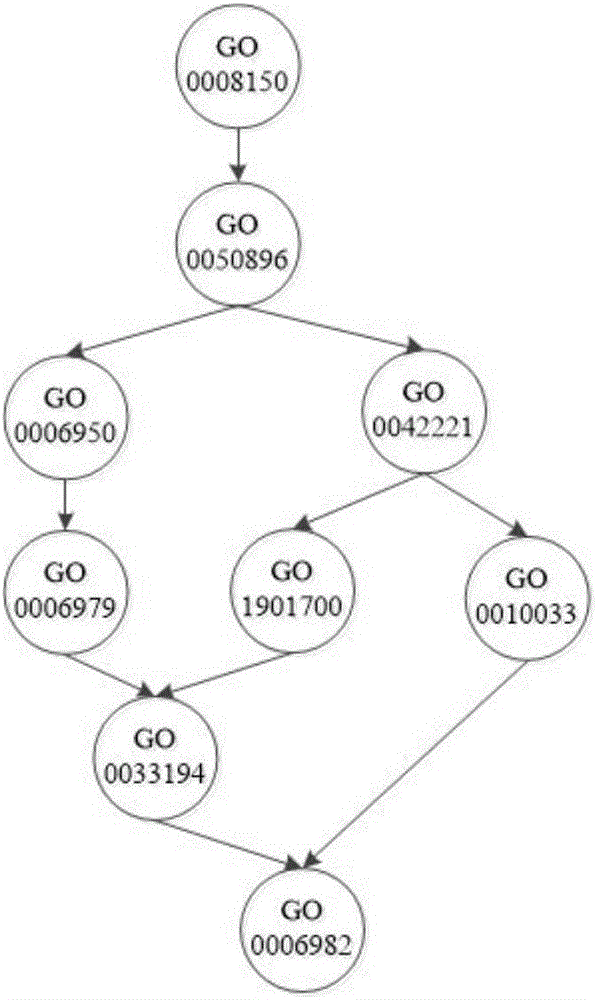
Method for identifying protein functions based on protein-protein interaction network and network topological structure features
InactiveCN105138866ARobustSignificant predictive advantageSpecial data processing applicationsNODALData set
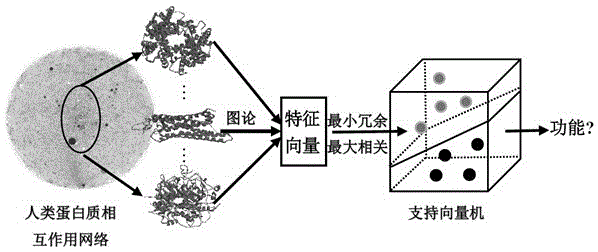
The invention discloses a method for identifying protein functions based on a protein-protein interaction network and network topological structure features. Firstly, a node and side-weighted protein-protection interaction network is established, wherein the node represents protein while the edge represents the interaction; then the nodes and the sides in the network are weighted by protein first-grade structural description and protein-protein interaction trust scoring; protection functional annotation data is collected to establish a data set, and a new protein with overall and local information network topological structure features is provided based on a graph theory; and finally, the protein functions are predicated by choosing features through adopting a minimum-redundancy maximum-correlation method and by modeling through a support vector machine. The protein function predication method is greatly better than the prior art, and has robustness on sequence similarity and sampling; and meanwhile, information of three-dimensional structure and the like of protein is not required, so that the method is simple, rapid, accurate and efficient, and the method is expected to be applied in the research fields of proteomics and the like.
Owner:SYSU CMU SHUNDE INT JOINT RES INST +2
Hierarchical multi-label classification method for protein function prediction
ActiveCN106126972ASolve the multi-label problemAchieve forecastBiostatisticsProteomicsData setProtein function prediction
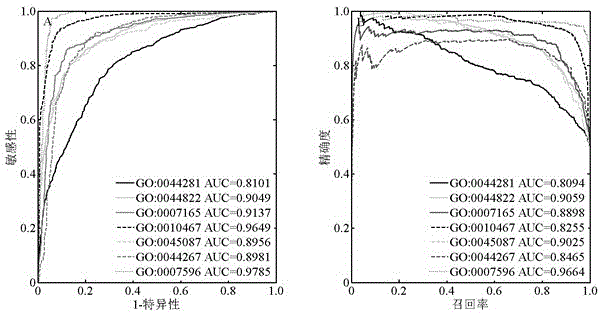
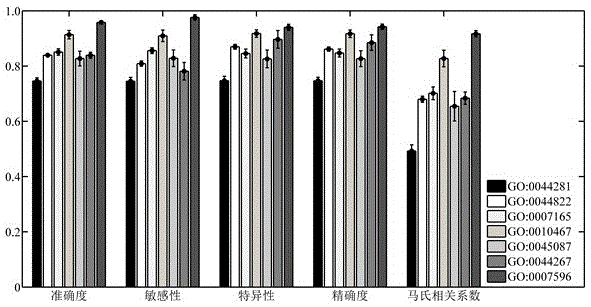
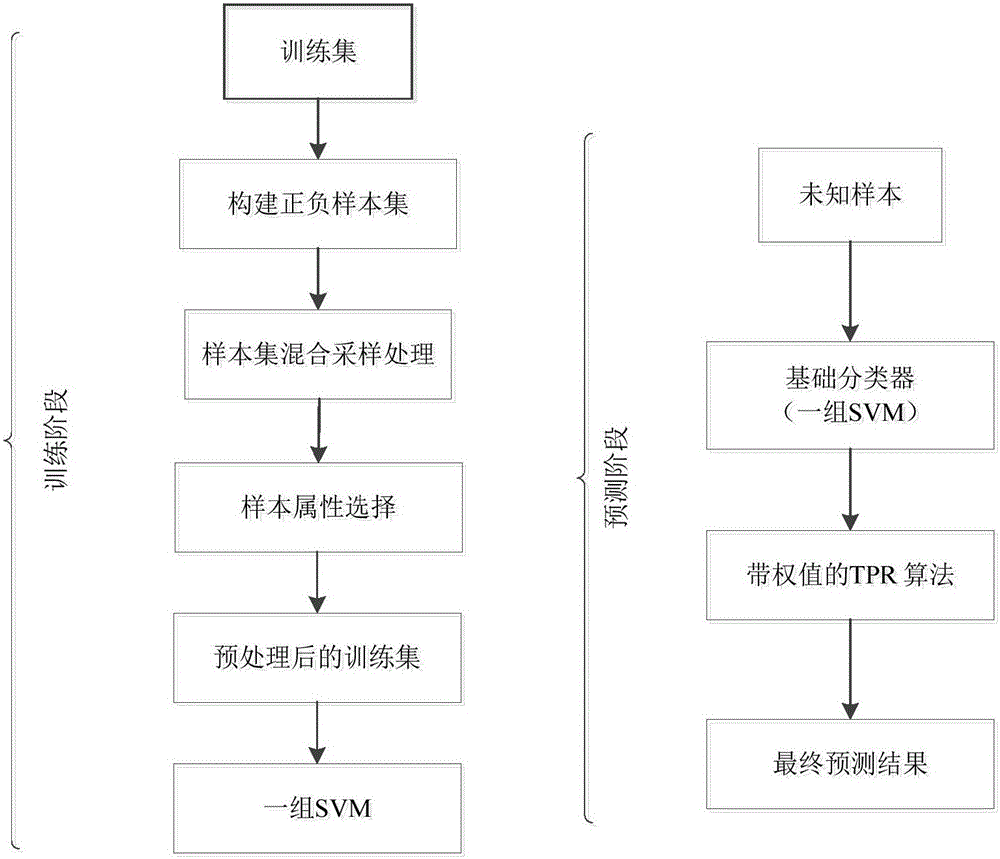
The invention relates to the field of bioinformatics and data mining, in particular to a hierarchical multi-label classification method for protein function prediction, and aims at solving the data set imbalance problem, multi-label problem and hierarchical constraint problem when the conventional classification methods are used for predicting protein functions. The method comprises the following steps of: 1, a training stage: training a data set of each node in a class label hierarchical structure by adopting an SVM classifier in the training stage so as to obtain a group of basic classifiers; and 2, a prediction stage: firstly obtaining preliminary results of unknown samples by using the group of basic classifiers obtained in the training stage in the prediction stage, and processing the results by adopting a TPR algorithm with a weight so as to obtain a final result which satisfies a hierarchical constraint condition and realize the prediction of the protein functions. The hierarchical multi-label classification method for protein function prediction is applied to the field of bioinformatics and data mining.
Owner:NAT INST OF ADVANCED MEDICAL DEVICES SHENZHEN
Small molecule-dependent inteins and uses thereof
ActiveUS9200045B2Low efficiencySlow splicingHydrolasesFusion with post-translational modification motifInteinNatural state
Owner:PRESIDENT & FELLOWS OF HARVARD COLLEGE
Method for regulating protein function in cells in vivo using synthetic small molecules
ActiveUS8530636B2Improve survivalInhibit tumor growthOrganic active ingredientsBiocideIn vivoProtein function
Owner:THE BOARD OF TRUSTEES OF THE LELAND STANFORD JUNIOR UNIV
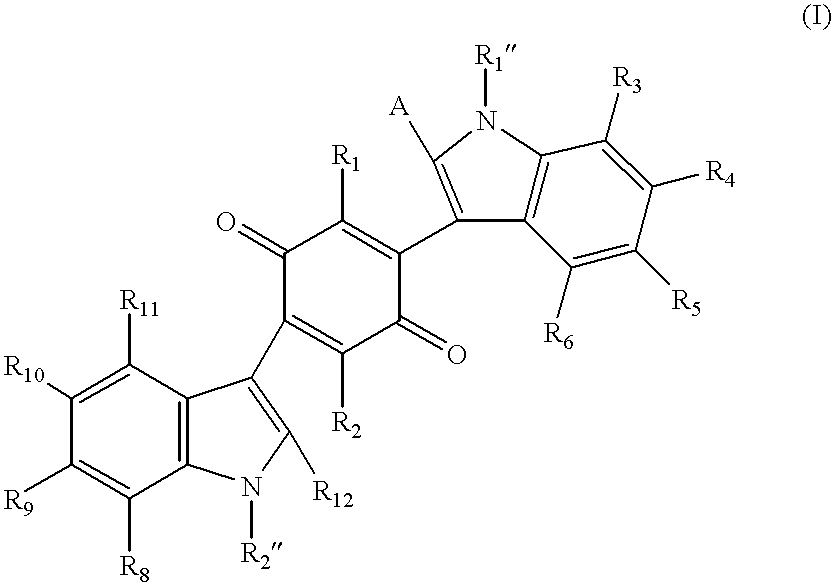
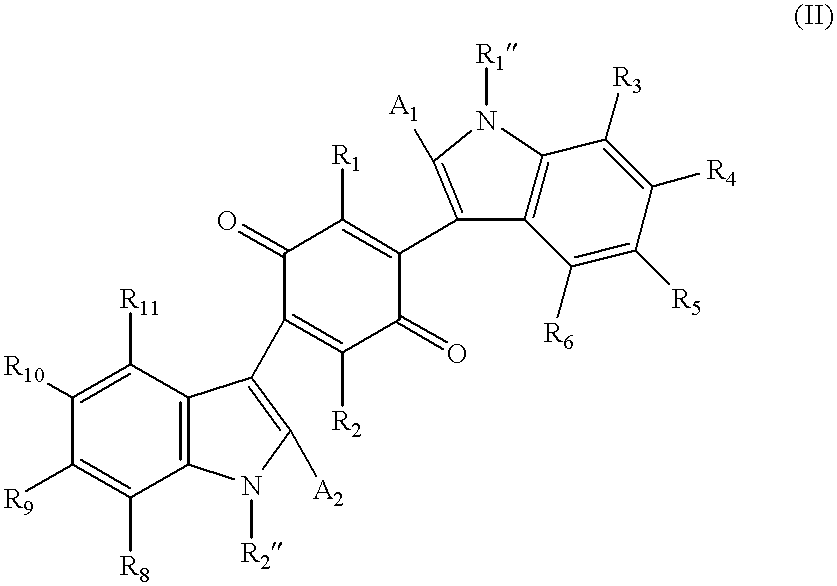
Mono- and bis-indolylquinones and prophylactic and therapeutic uses thereof
InactiveUS6376529B1Effective treatmentLower blood sugar levelsBiocideOrganic chemistryMedicineInsulin deficiency
The present invention relates to a class of indolylquinone compounds that inhibit GRB-2 adaptor protein function, pharmaceutical compositions comprising these compounds, and methods for ameliorating the symptoms of cell proliferative disorders associated with GRB-2 adaptor protein function using these compounds. The present invention further relates to methods for treating insulin-related disorders, such as diabetes, insulin resistance, insulin deficiency and insulin allergy, and for ameliorating the symptoms of insulin-related disorders, using certain indolylquinone compounds and pharmaceutical compositions thereof. The present invention also relates to novel synthetic methods for the preparation of mono- and bis-indolylquinone compounds.
Owner:SUGEN INC
Nitrosated and nitrosylated heme proteins
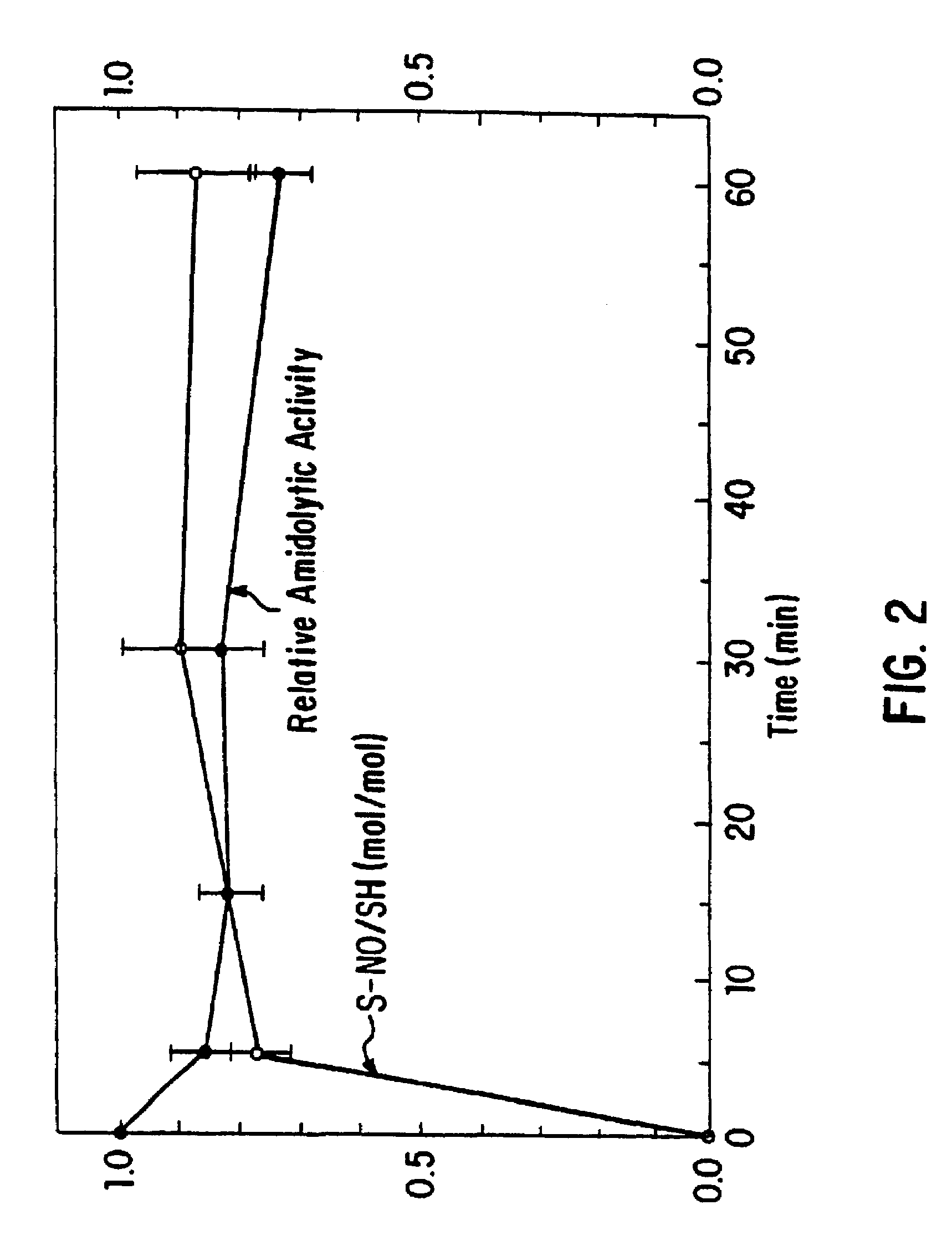
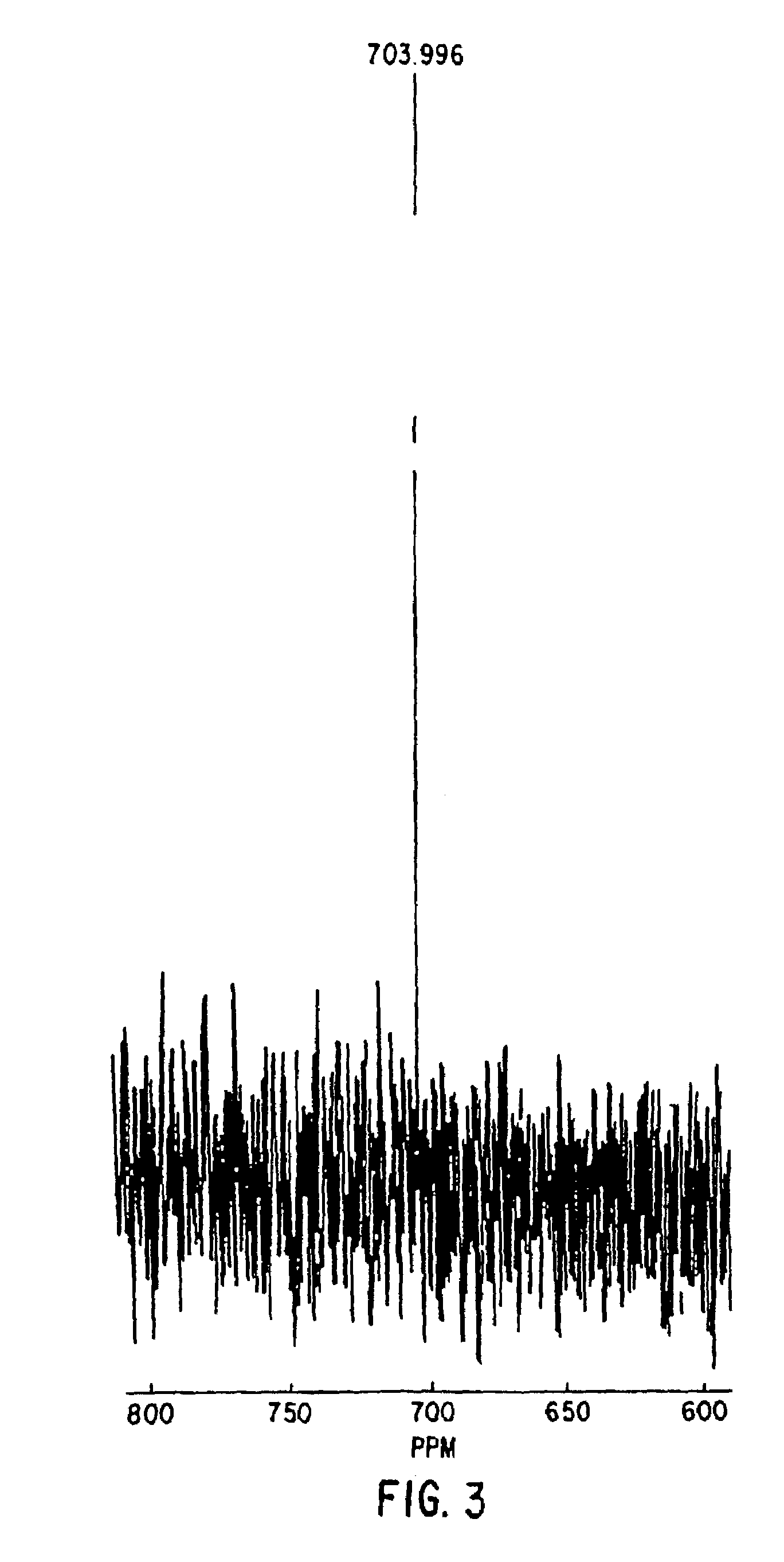
Nitrosylation of proteins and amino acid groups enables selective regulation of protein function, and also endows the proteins and amino acids with additional smooth muscle relaxant and platelet inhibitory capabilities. Thus, the invention relates to novel compounds achieved by nitrosylation of protein thiols. Such compounds include: S-nitroso-heme proteins, S-nitroso-t-PA, S-nitroso-cathepsin; S-nitroso-lipoproteins; and S-nitroso-immunoglobulins. The invention also relates to therapeutic use if S-nitroso-protein compounds for regulating protein function, cellular metabolism and effecting vasodilation, platelet inhibition, relaxation of non-vascular smooth muscle, and increasing blood oxygen transport by hemoglobin and myoglobin. The compounds are also used to deliver nitric oxide in its most bioactive form in order to achieve the effects described above, or for in vitro nitrosylation of molecules present in the body. The invention also relates to the nitrosylation of oxygen, carbon and nitrogen moieties present on proteins and amino acids, and the use thereof to achieve the above physiological effects.
Owner:THE BRIGHAM & WOMEN S HOSPITAL INC
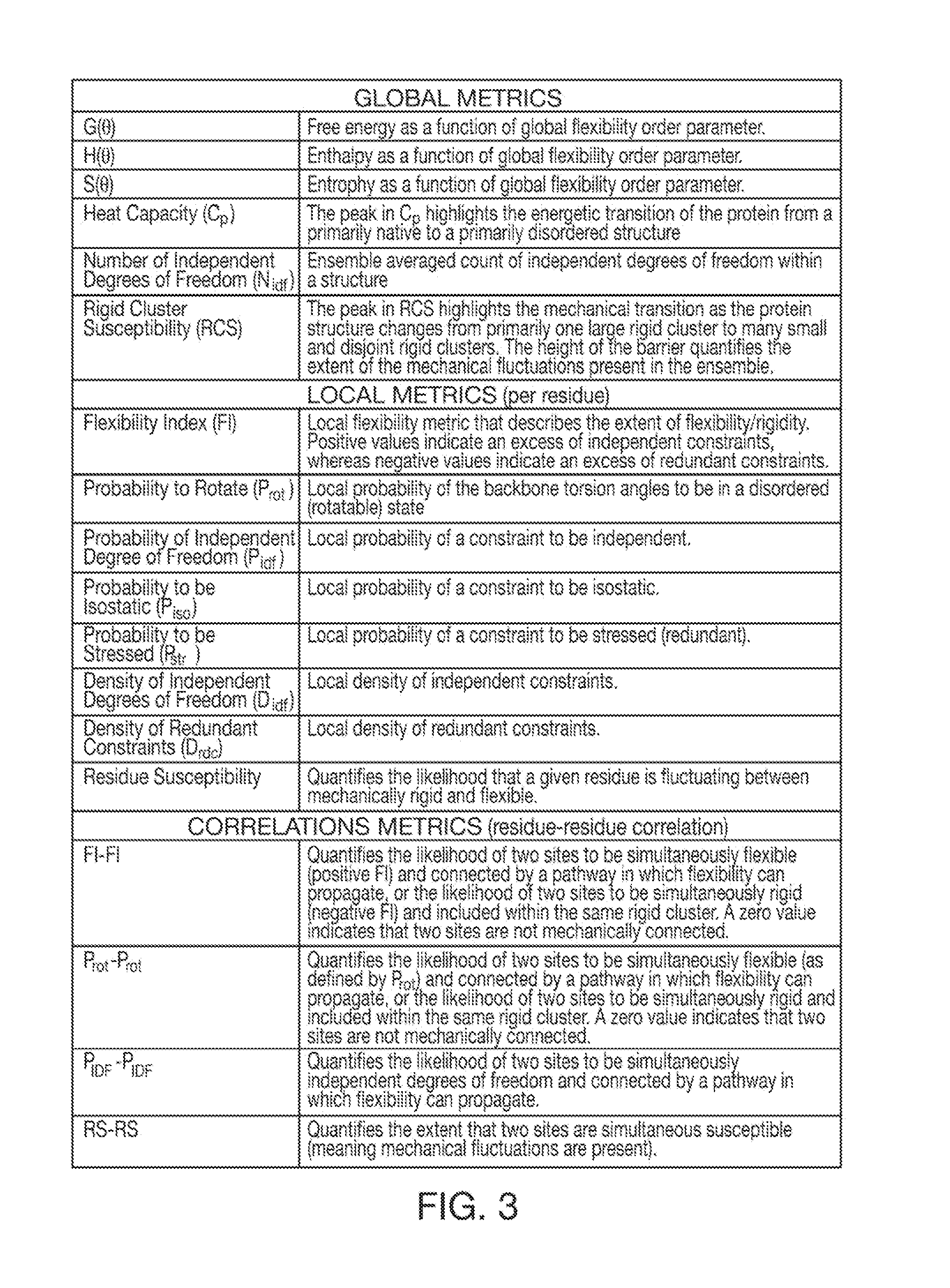
Computer implemented system for protein and drug target design utilizing quantified stability and flexibility relationships to control function
InactiveUS8374828B1High catalytic efficiencyFast computerMolecular designMicrobiological testing/measurementHigh dimensionalDrug target
Owner:THE UNIV OF NORTH CAROLINA AT CHAPEL HILL
Method for Regulating Protein Function in Cells In Vivo Using Synthetic Small Molecules
ActiveUS20100034777A1Improve survivalInhibit tumor growthBiocideOrganic active ingredientsIn vivoProtein function
Owner:THE BOARD OF TRUSTEES OF THE LELAND STANFORD JUNIOR UNIV
Methods for conducting assays for enzyme activity on protein microarrays
The present invention relates to methods of conducting assays for enzymatic activity on microarrays useful for the large-scale study of protein function, screening assays, and high-throughput analysis of enzymatic reactions. The invention relates to methods of using protein chips to assay the presence, amount, activity and / or function of enzymes present in a protein sample on a protein chip. In particular, the methods of the invention relate to conducting enzymatic assays using a microarray wherein a protein and a substance are immobilized on the surface of a solid support and wherein the protein and the substance are in proximity to each other sufficient for the occurrence of an enzymatic reaction between the substance and the protein. The invention also relates to microarrays that have an enzyme and a substrate immobilized on their surface wherein the enzyme and the substrate are in proximity to each other sufficient for the occurrence of an enzymatic reaction between the enzyme and the substrate.
Owner:PROTOMETRIX
Novel Genes Encoding Novel Proteolytic Enzymes
The invention relates to newly identified gene sequences that encode novel proteases obtainable from Aspergillus niger. The invention features the full length gene sequence of the novel genes, their cDNA sequences as well as the full-length functional protein and fragments thereof. The invention also relates to methods of using these enzymes in industrial processes and methods of diagnosing fungal infections. Also included in the invention are cells transformed with DNA according to the invention and cells wherein a protease according to the invention is genetically modified to enhance or reduce its activity and / or level of expression.
Owner:DSM IP ASSETS BV