Patents
Literature
381results about How to "Accurate content" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
Apparatus for providing voice dialogue service and method of operating the same
InactiveUS7734461B2Accurate analysisAccurate contentSemantic analysisSpeech recognitionPart of speechSyntax
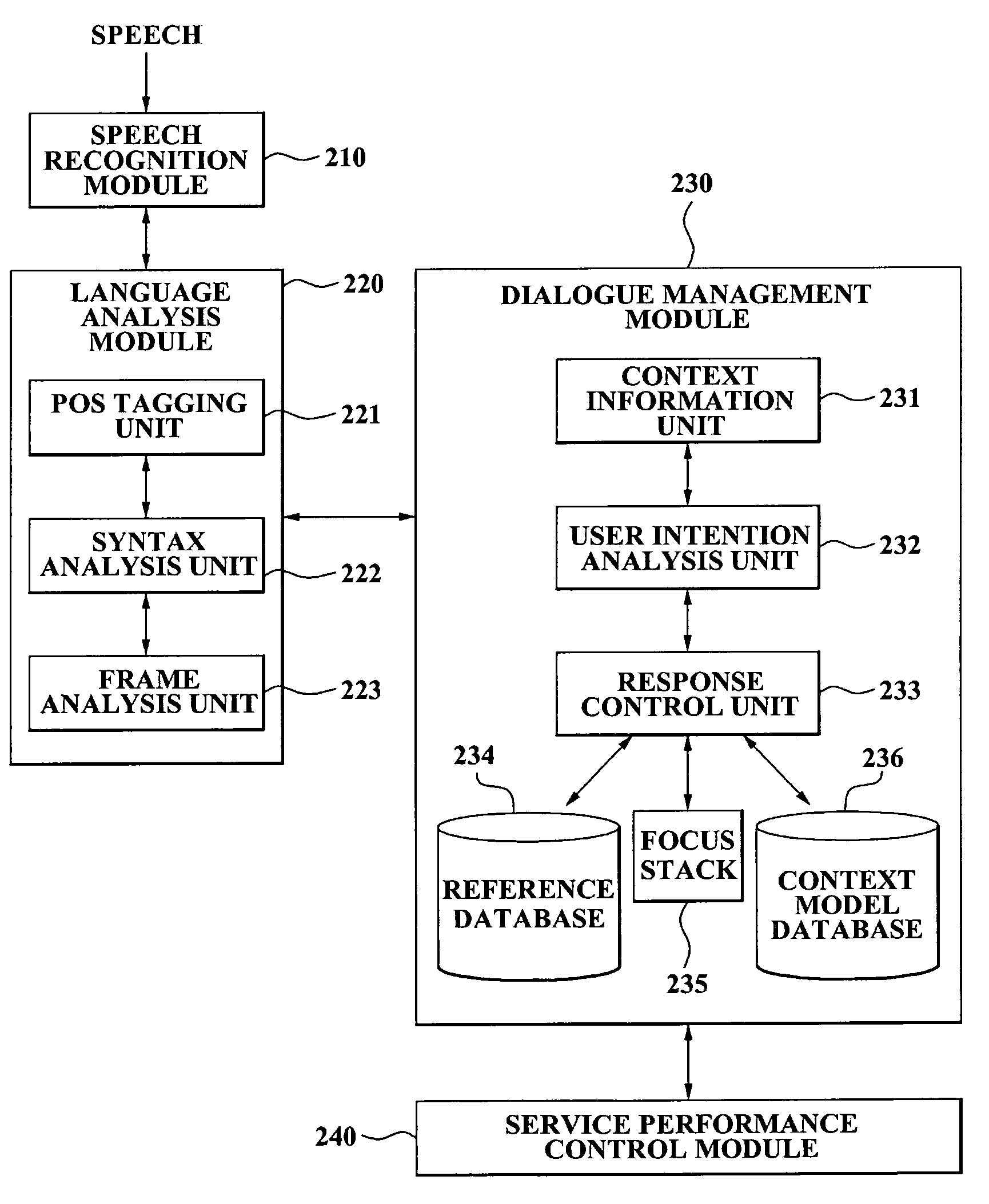
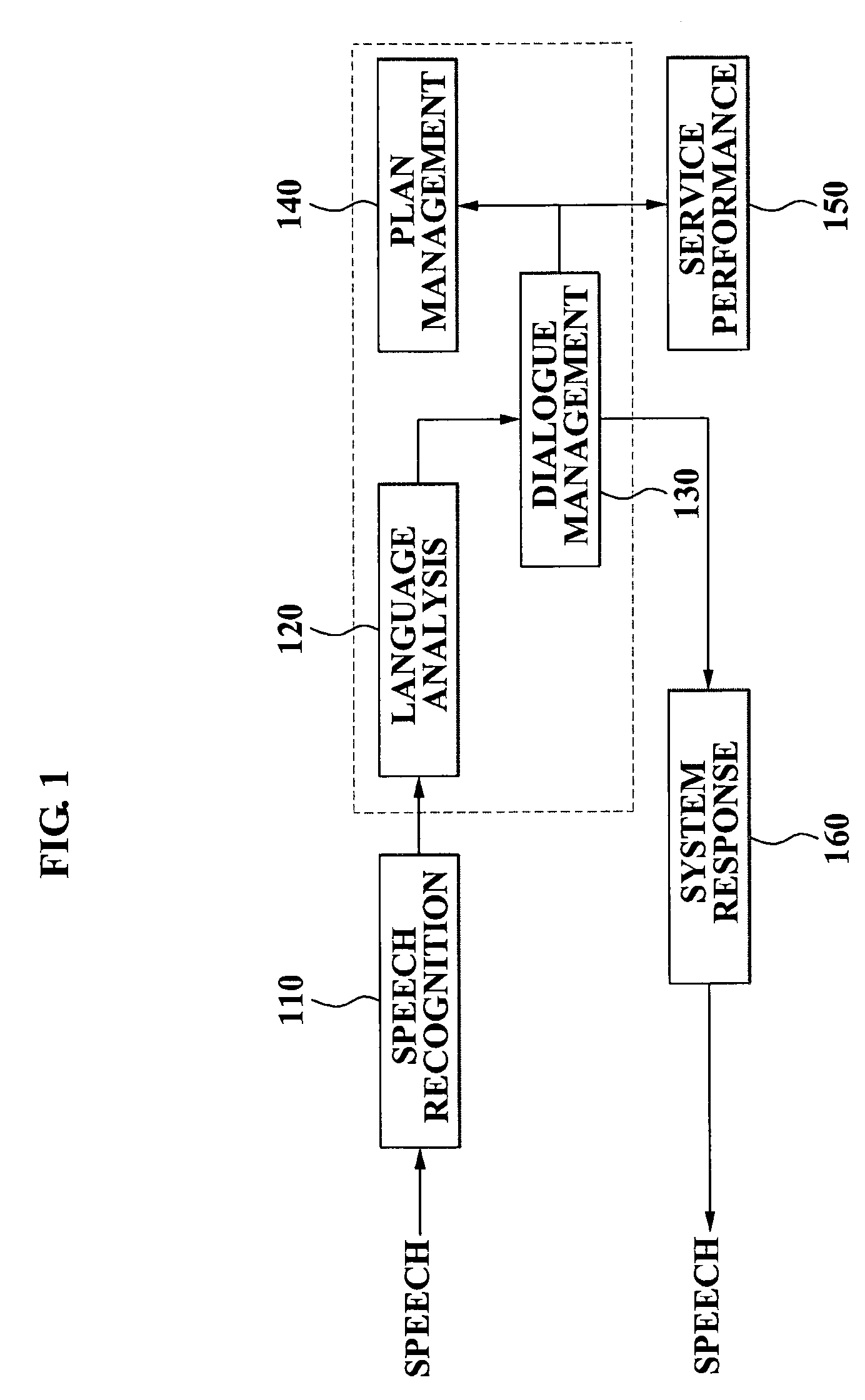
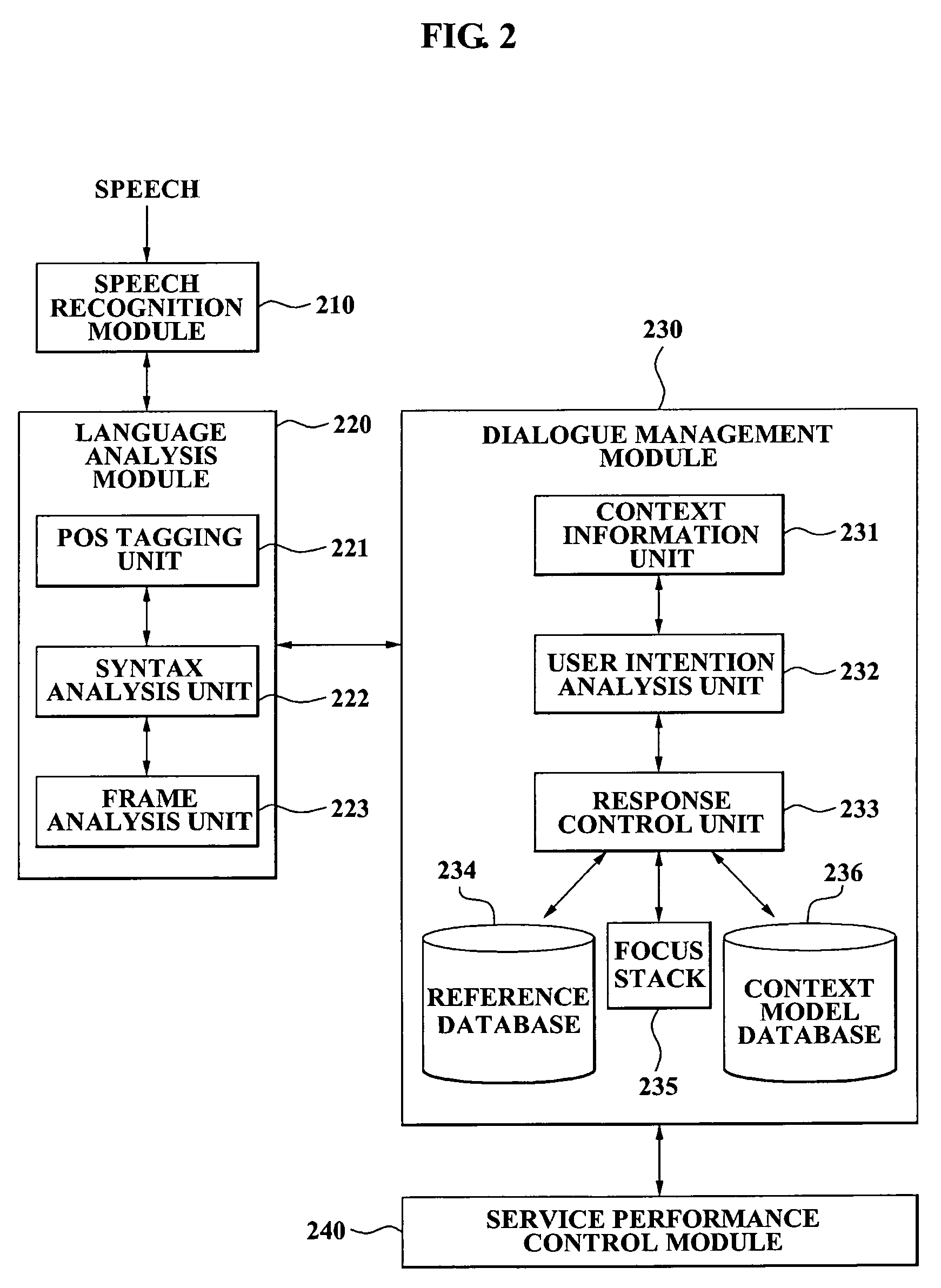
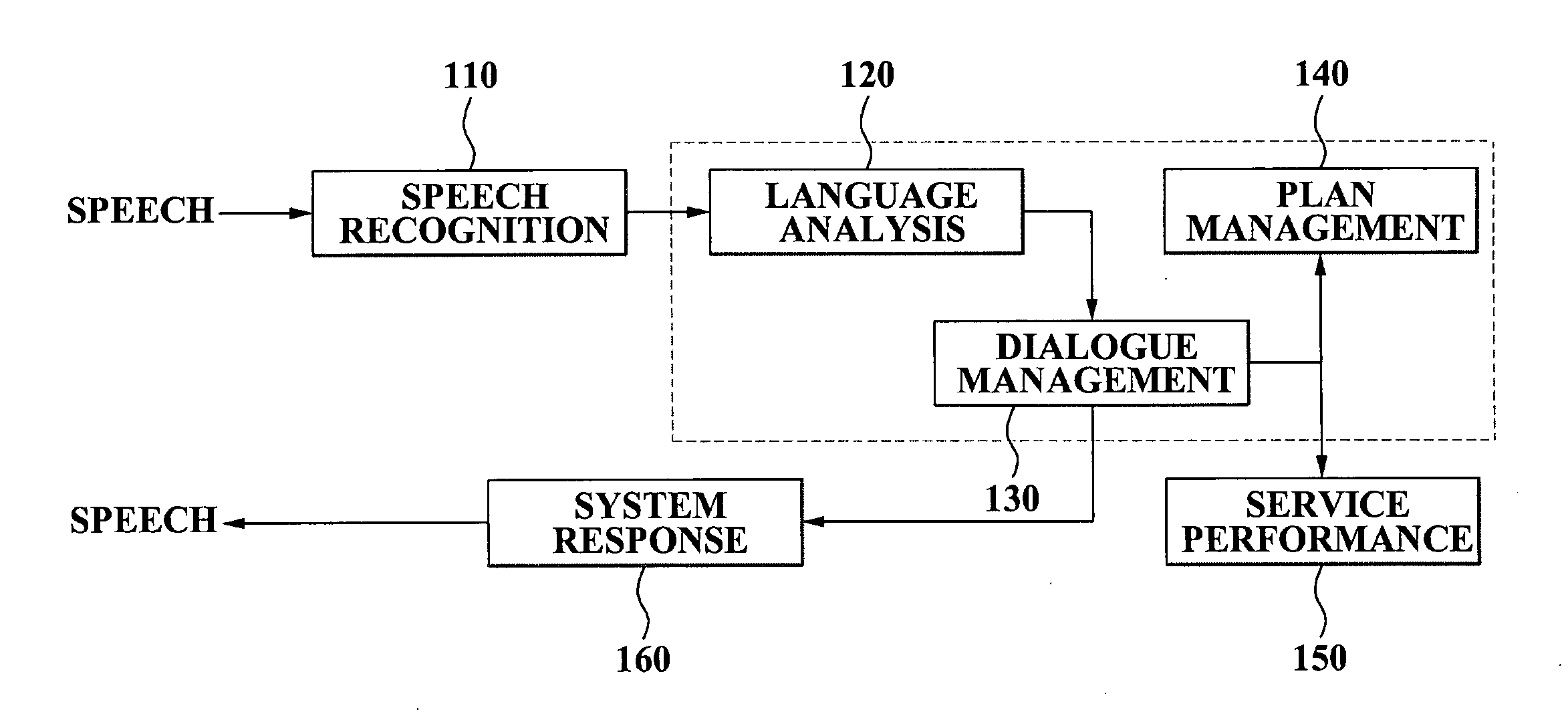
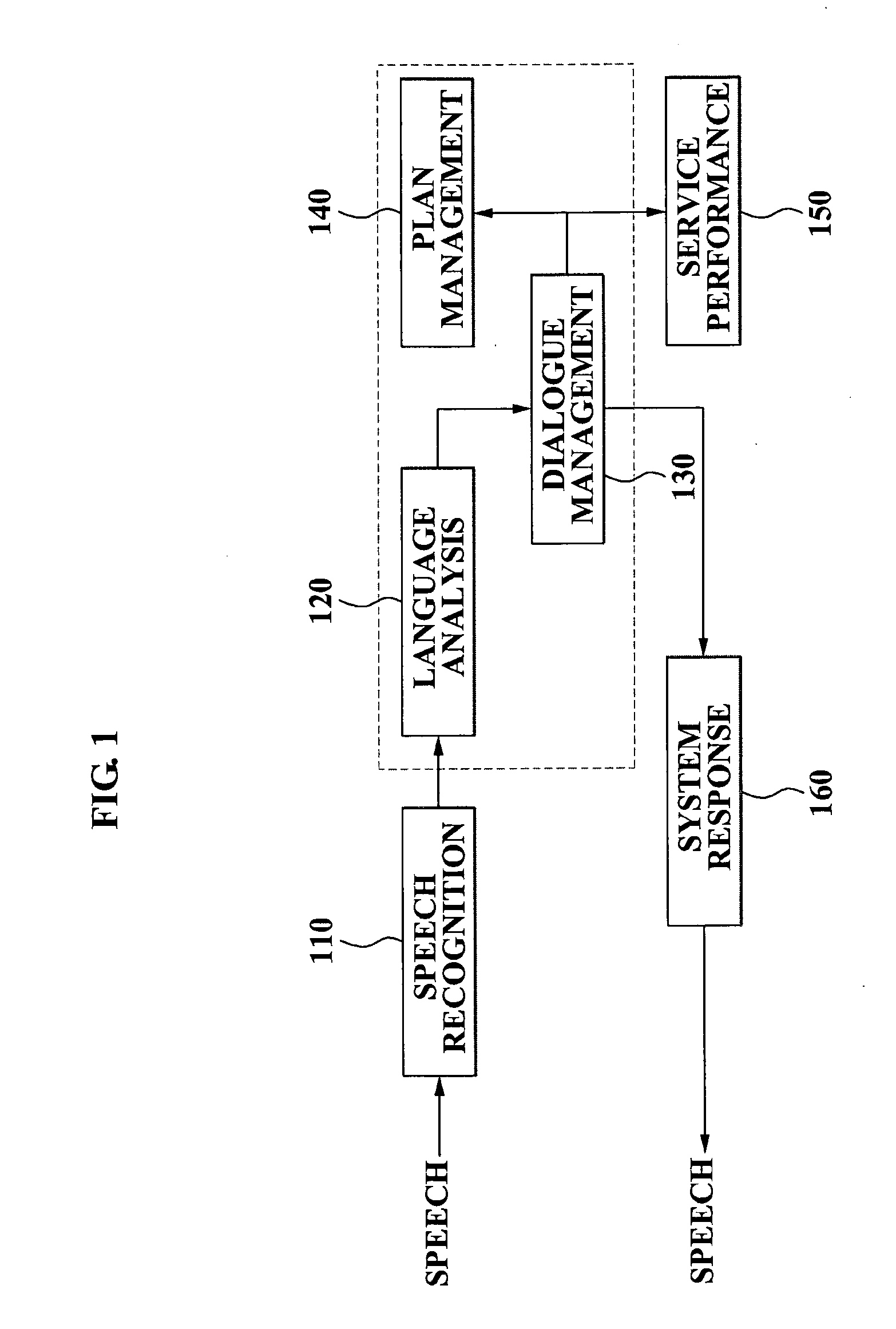
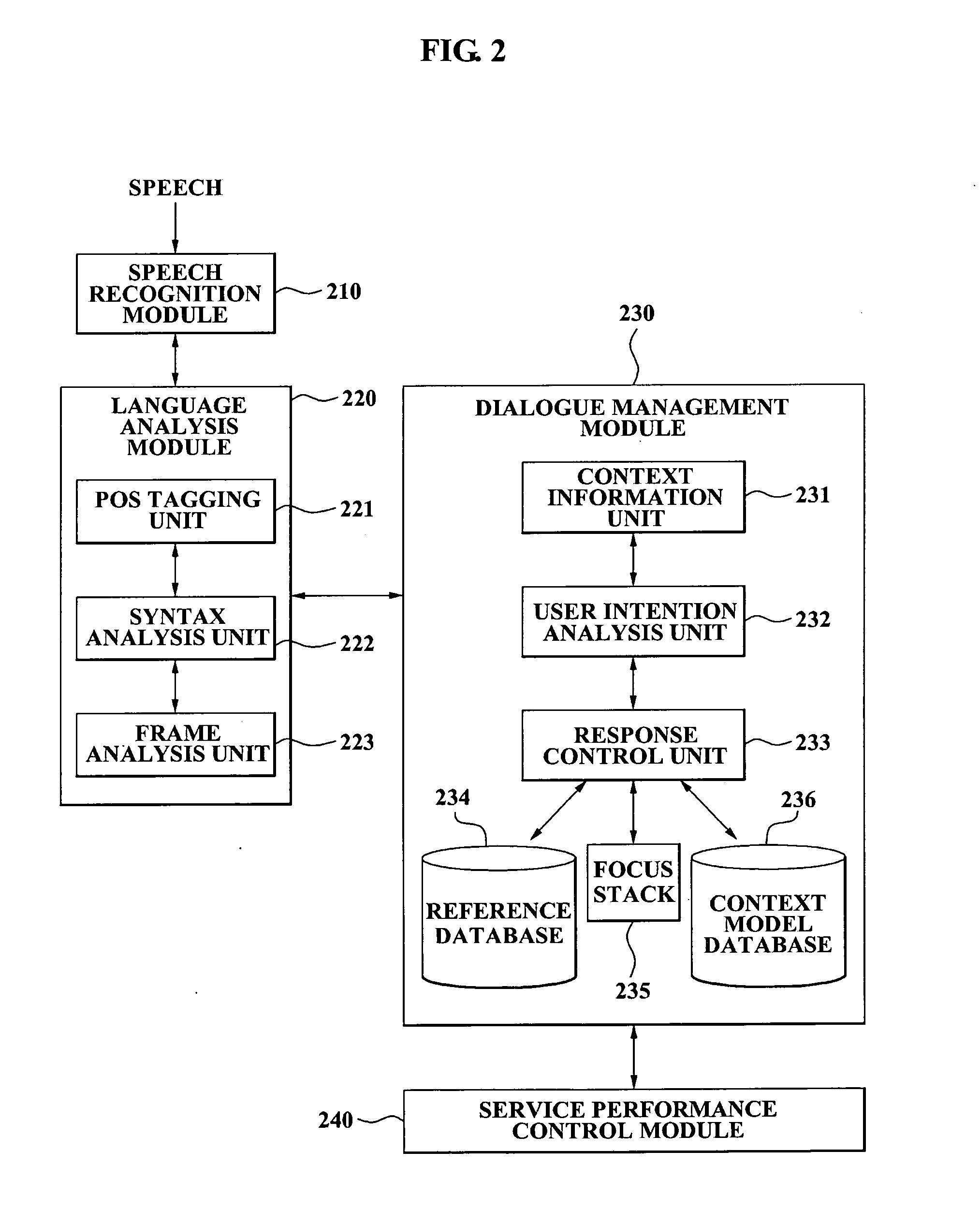
A speech dialogue service apparatus including: a language analysis module tagging a part of speech (POS) of each respective word included in a sentence recorded in a predetermined text, syntactically analyzing the sentence by classifying a meaning of each respective word, and generating at least one semantic frame corresponding to the sentence according to a result of the syntactical analysis; and a dialogue management module analyzing an intention of the sentence corresponding to the at least one respective semantic frame, and generating a system response corresponding to the sentence intention by selecting a predetermined sentence intention according to whether an action corresponding to the intention of the respective sentence can be performed.
Owner:SAMSUNG ELECTRONICS CO LTD
Method and System of Web Page Content Filtering
InactiveUS20120131438A1Low efficiencyAccurate contentComputer security arrangementsTransmissionE-commerceWeb page
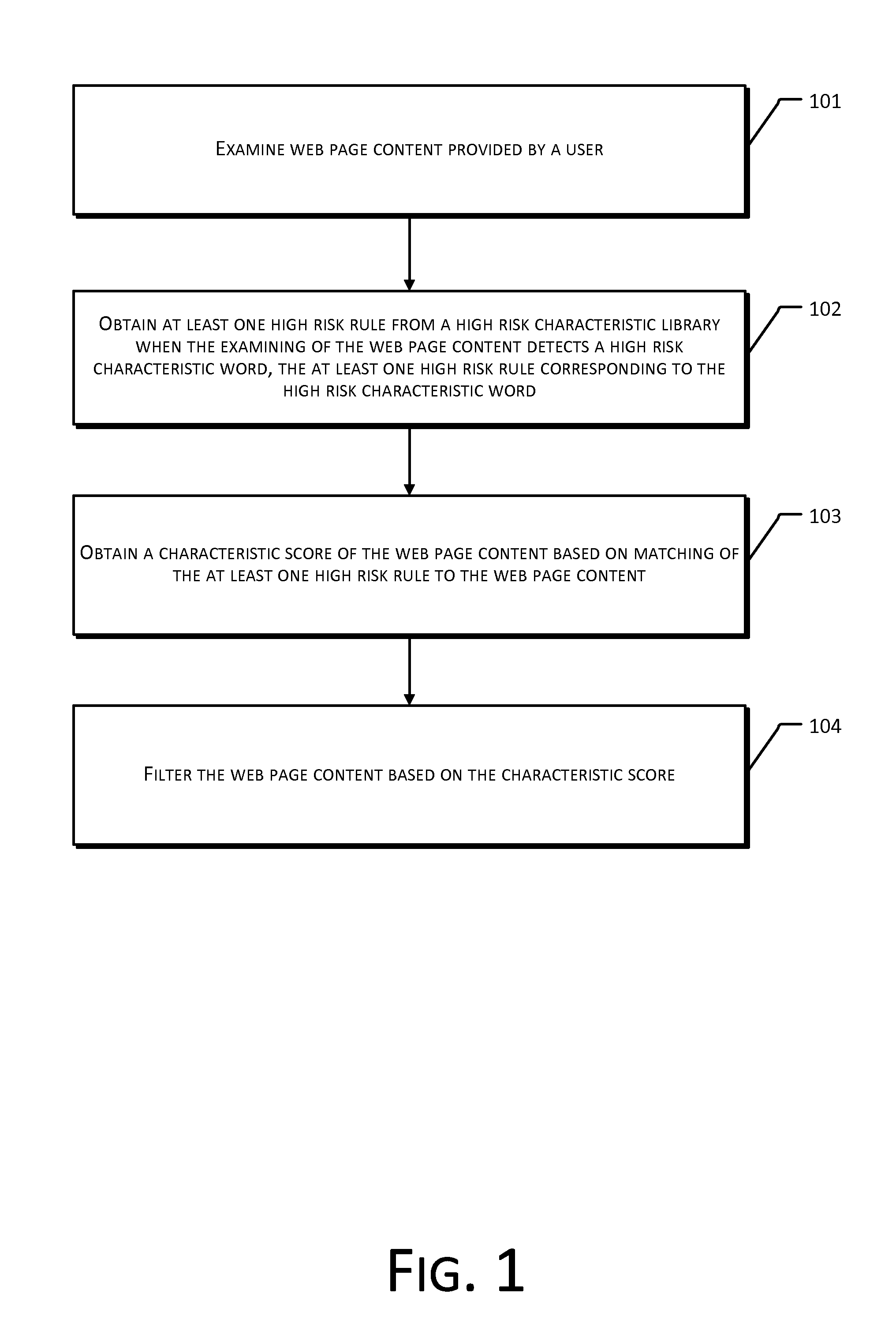
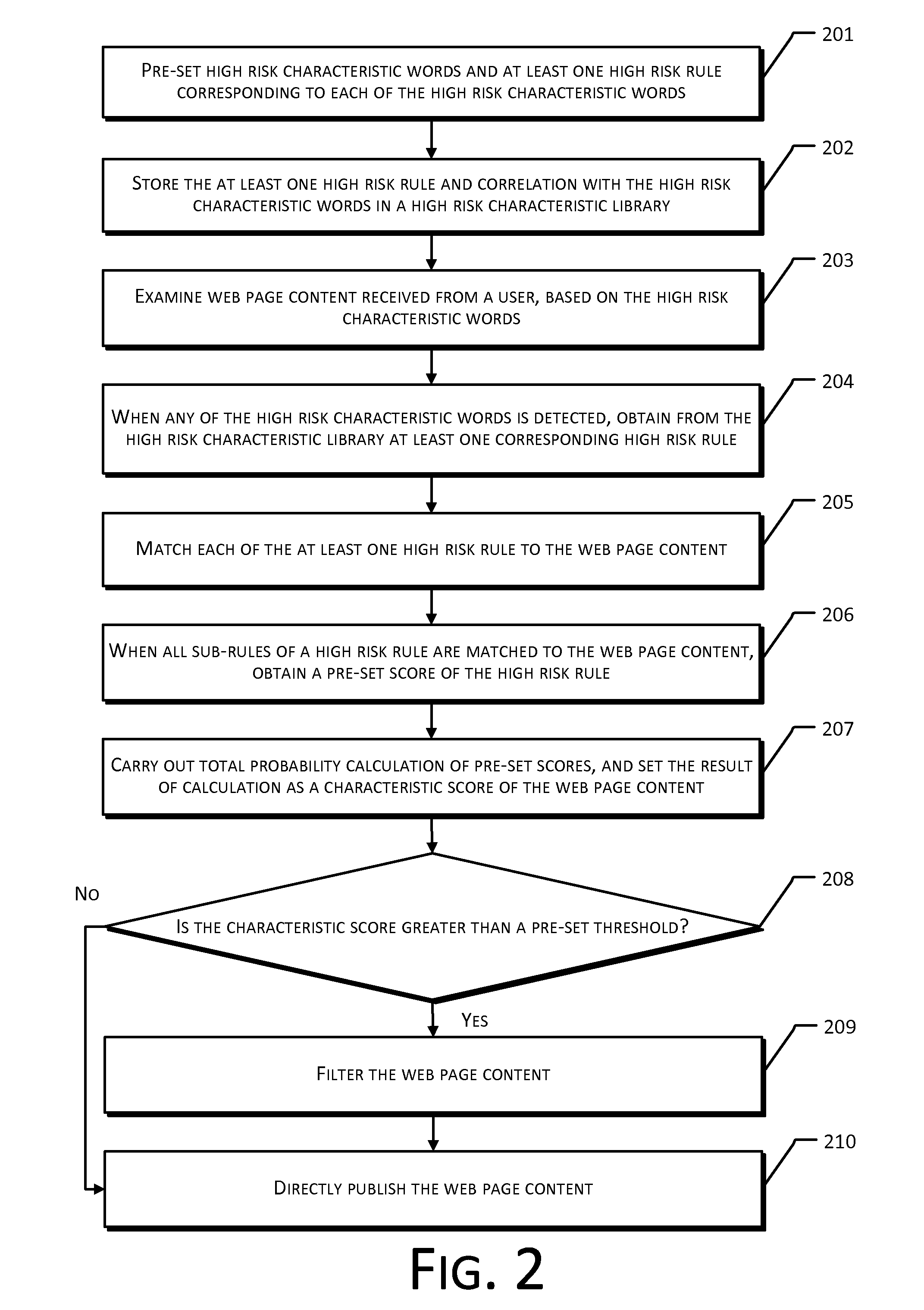
The present disclosure provides a method and system for web page content filtering. A method comprises: examining the web page content provided by a user; obtaining at least one high risk rule from a high risk characteristic library when the examining of the web page content detects a high risk characteristic word, the at least one high risk rule corresponding to the high risk characteristic word; obtaining a characteristic score of the web page content based on matching of the at least one high risk rule to the web page content; and filtering the web page content based on the characteristic score. The difference between the present disclosure and prior art techniques is that the disclosed embodiments can more precisely carry out web page content filtering to achieve better real-time safety and reliability of an e-commerce transaction.
Owner:ALIBABA GRP HLDG LTD
Mission critical NAND flash
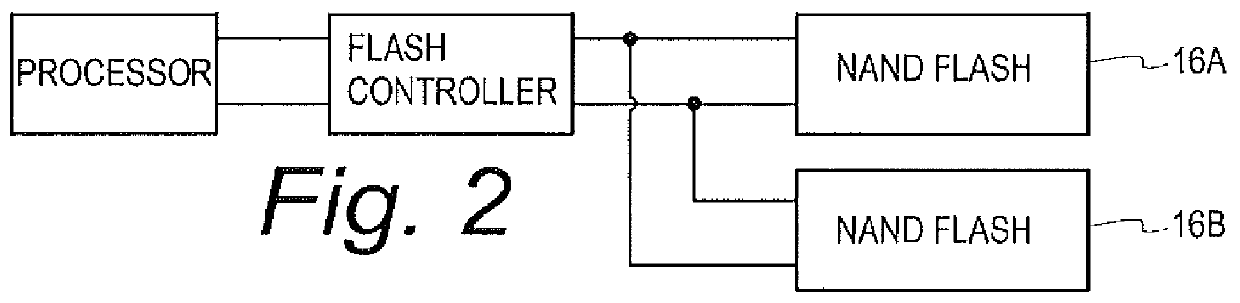
InactiveUS20120226934A1Data storage is reliableAccurate contentMemory systemsRedundant hardware error correctionComputer hardwareComputer architecture
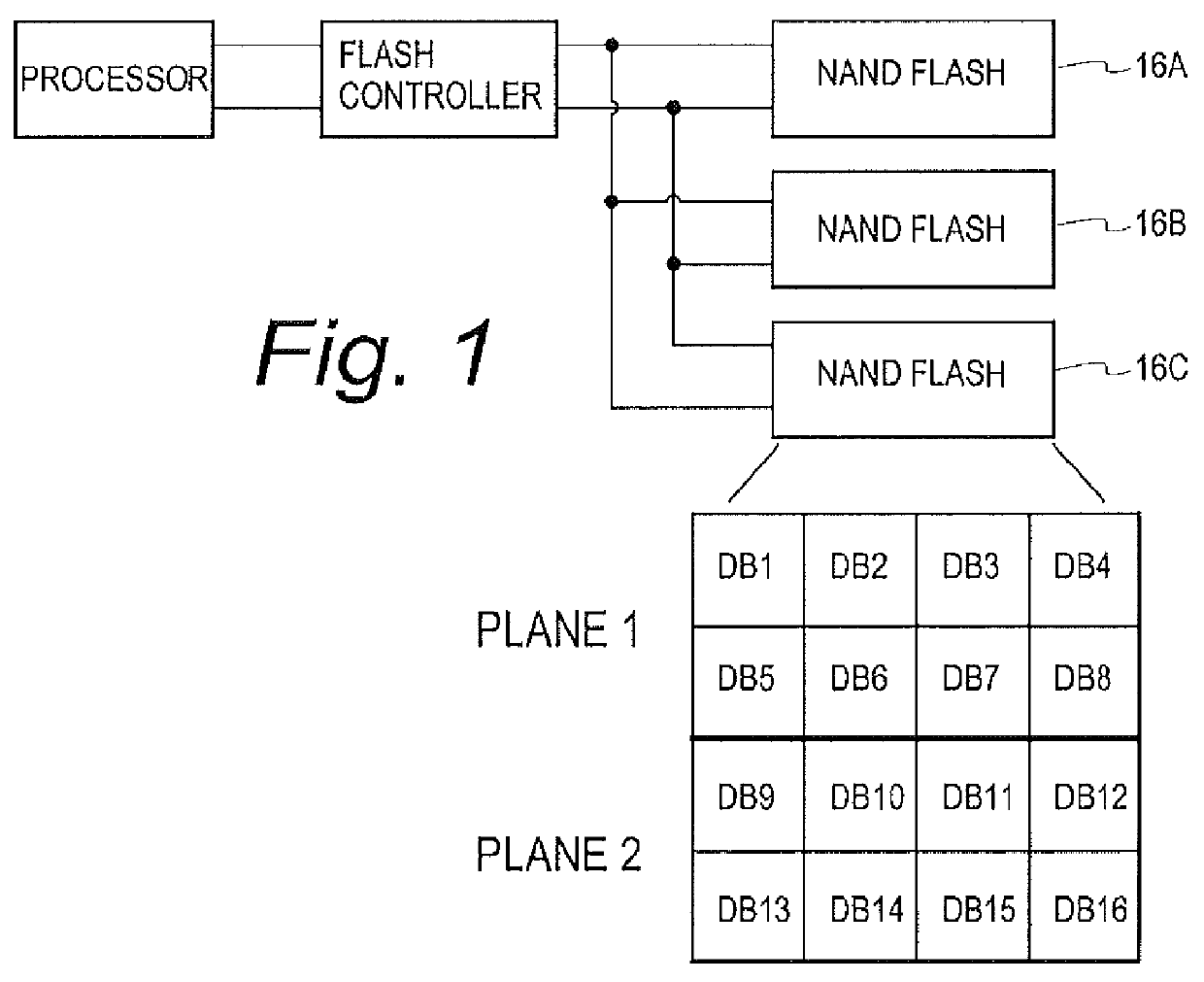
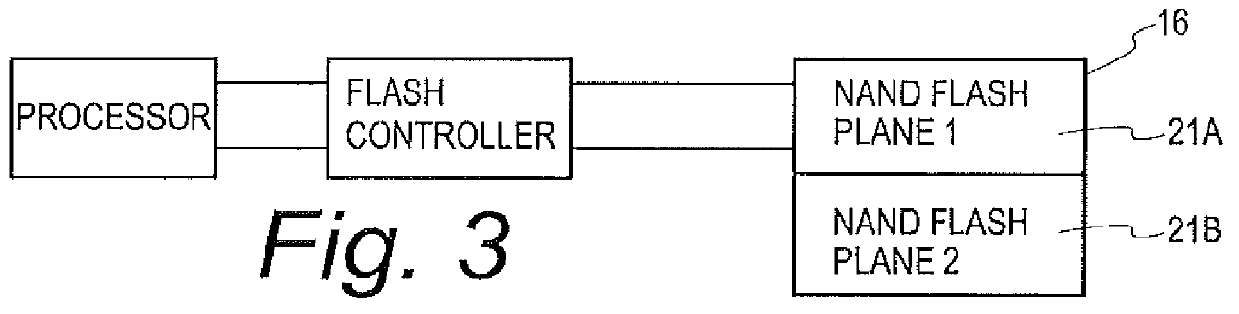
A flash controller reliably stores data in NAND FLASH by encoding data using an encoding algorithm, and storing that data across multiple pages of the memory. In one embodiment, true data is accepted by the controller, and the controller in turn creates coded data that is the bit-for-bit complement of the true data. The true data and the coded data are then written to the NAND FLASH on a page by page basis. A property of the coding techniques used is that, in at least some cases, detected errors can be corrected.
Owner:GREENTHREAD
Message reconstruction from partial detection
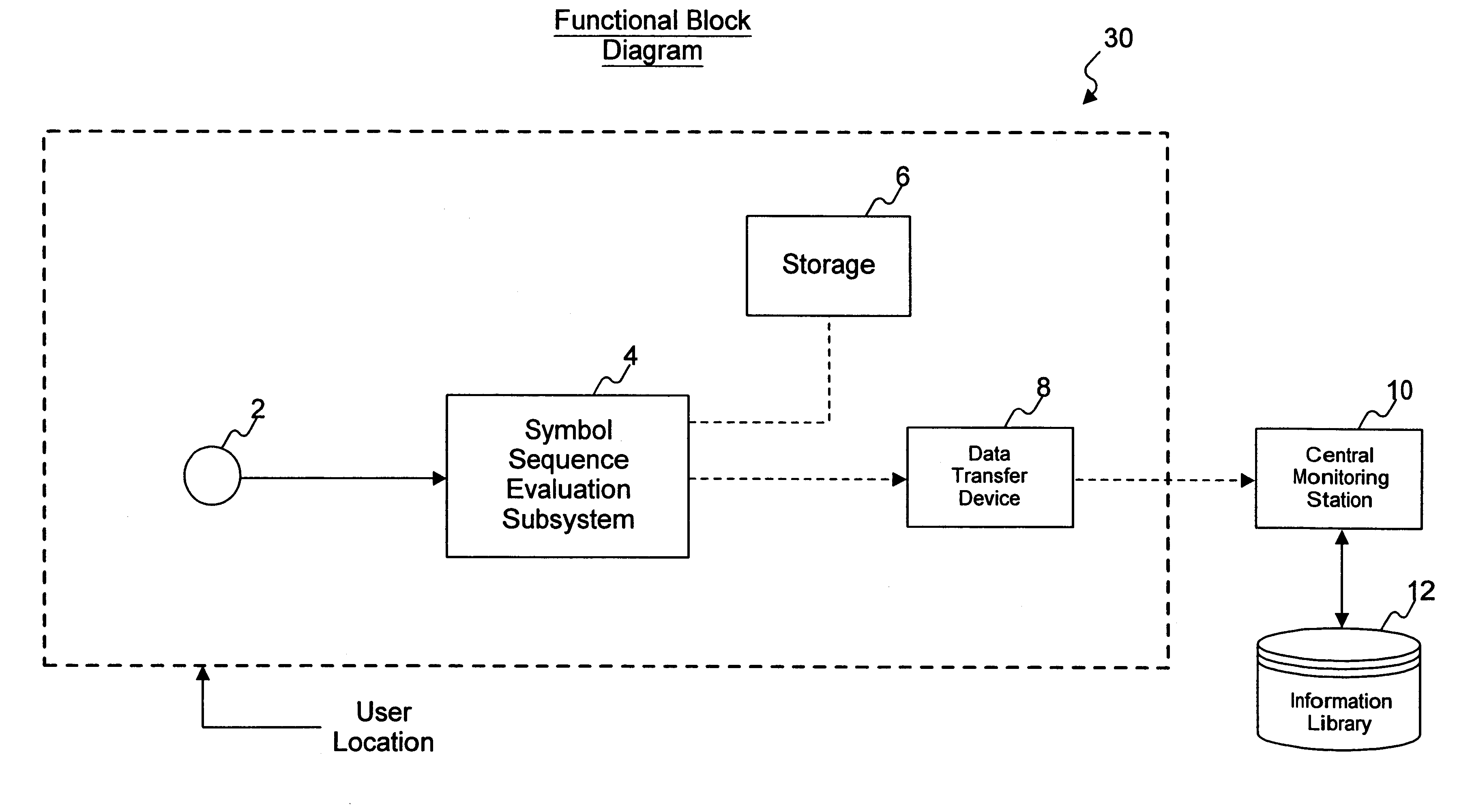
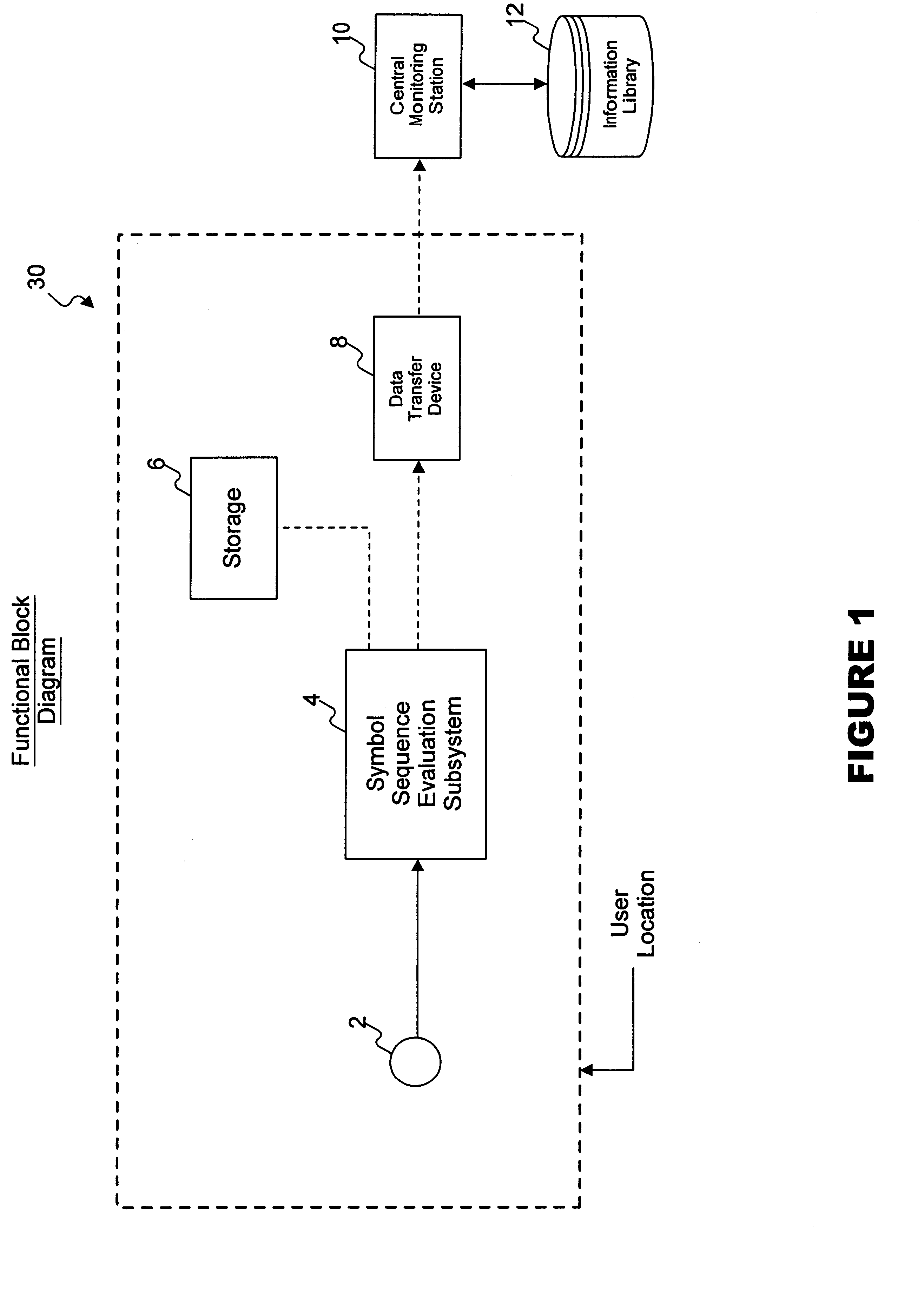
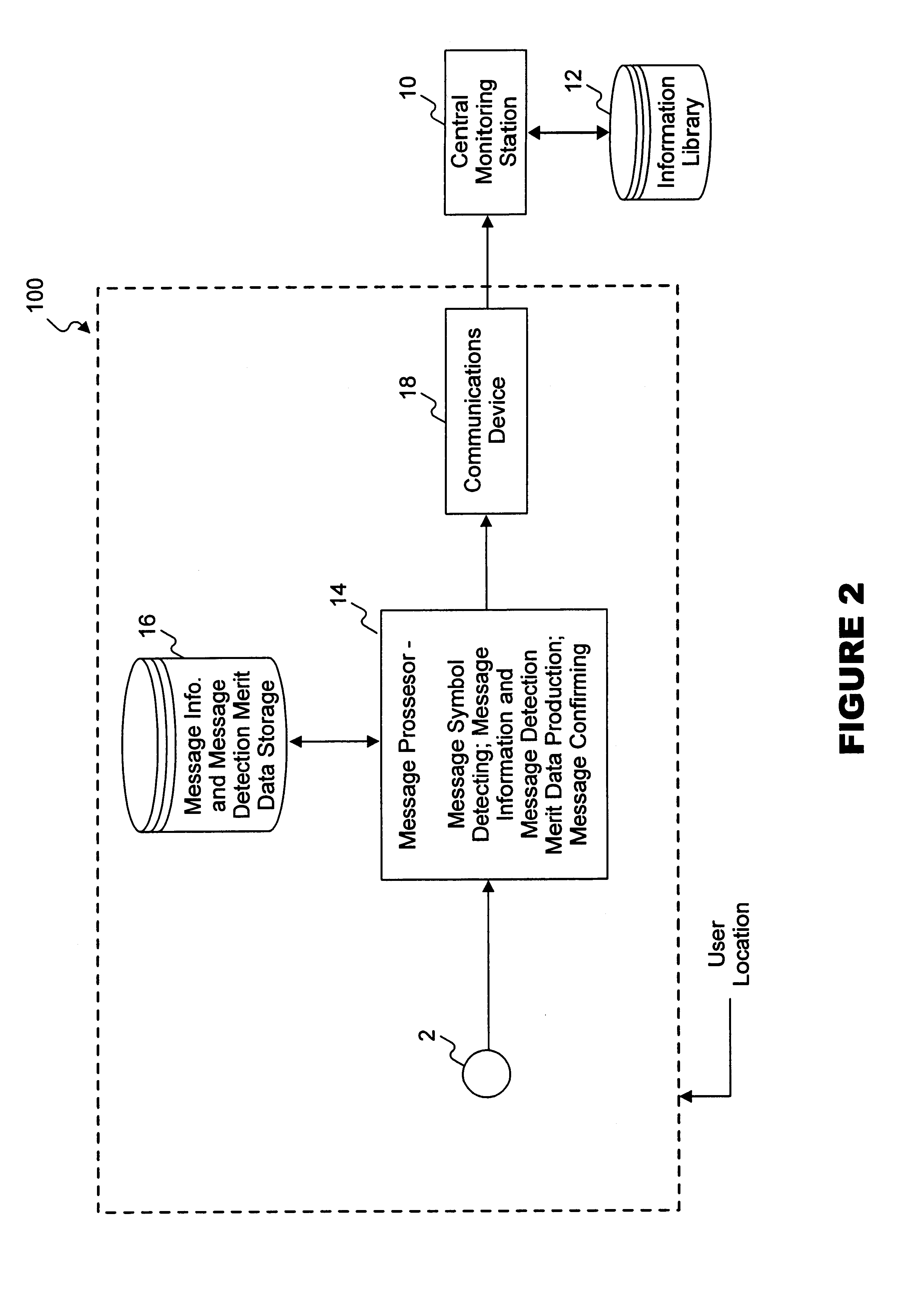
InactiveUS6862355B2Accurate representationAccurate contentError preventionDigital data processing detailsAudio MediaComputer science
A method and system for reliably detecting encoded messages included in audio media data in varying acoustic environments, where only a portion of the predetermined message may have been received or detected.
Owner:NIELSEN HLDG NV +1
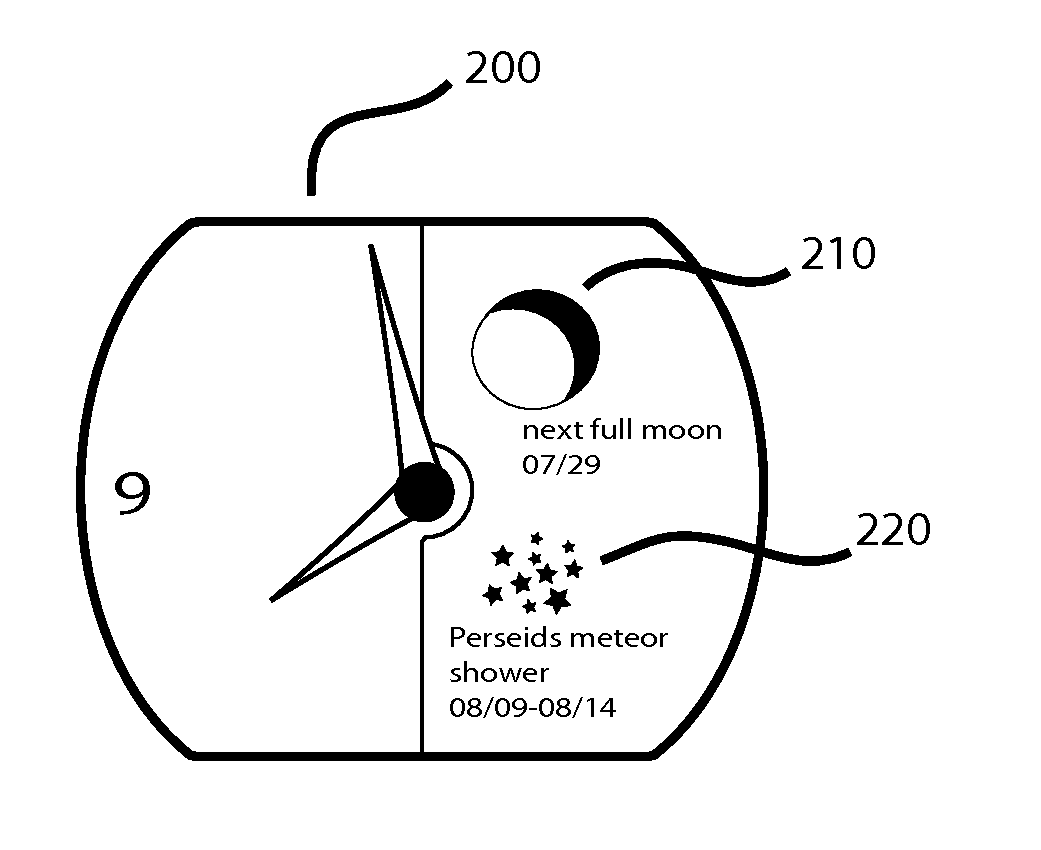
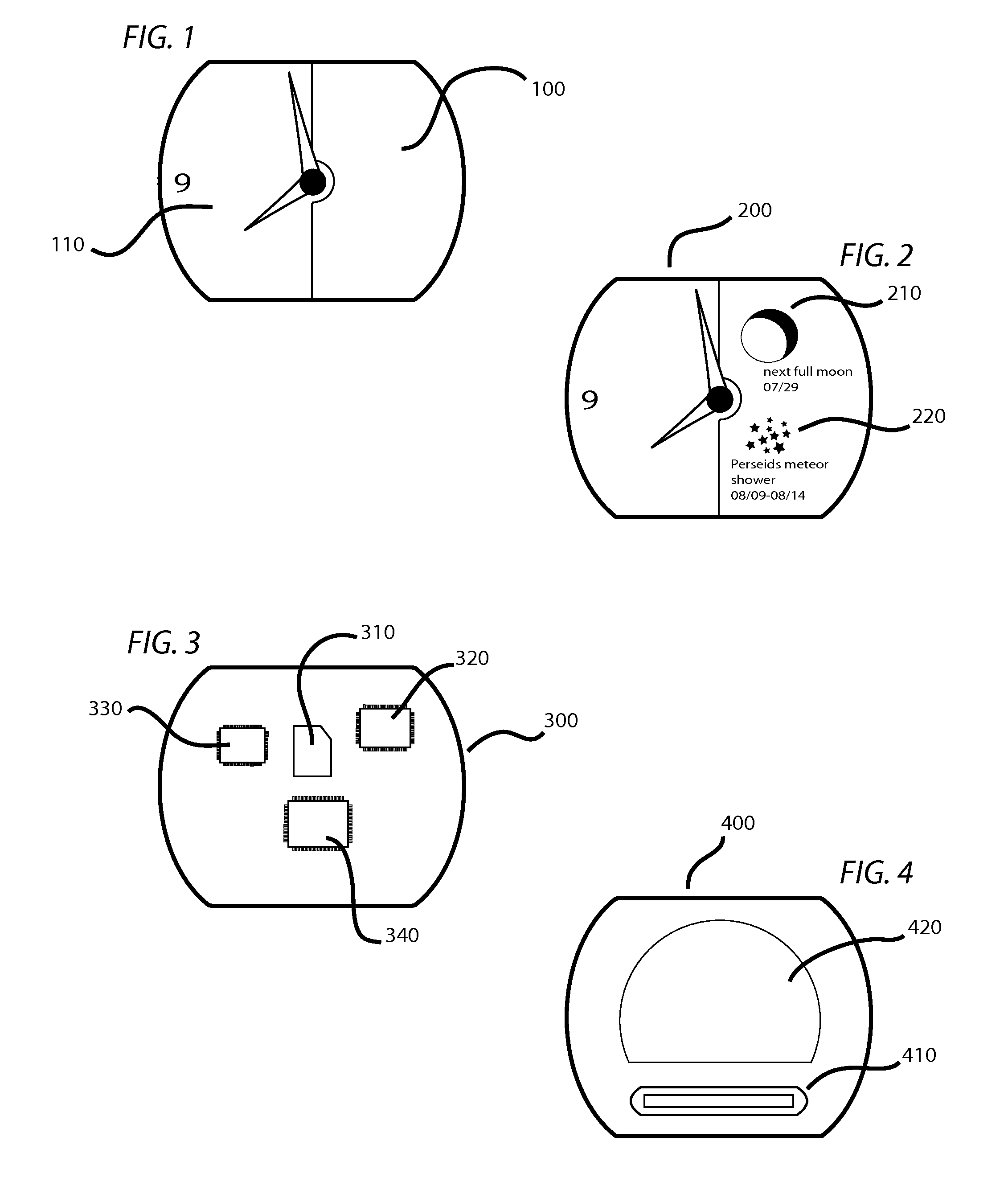
User Customizable Timepiece
InactiveUS20100226213A1Overcome deficienciesPrecise processVisual indicationVisual indicationsGraphicsGeolocation
A timepiece or wristwatch displays celestial “complications” and meteorological events based upon calculations and conditions relevant to the geographic location of the timepiece. A memory storage device, microprocessor, mechanical and software controlled graphical display systems and network connectivity facilitate the input of user selected complications, display options, geographic arguments and other variables. The timepiece is not limited to a preprogramed geographic area or to predefined complications.
Owner:DRUGGE BRIAN ROBERT
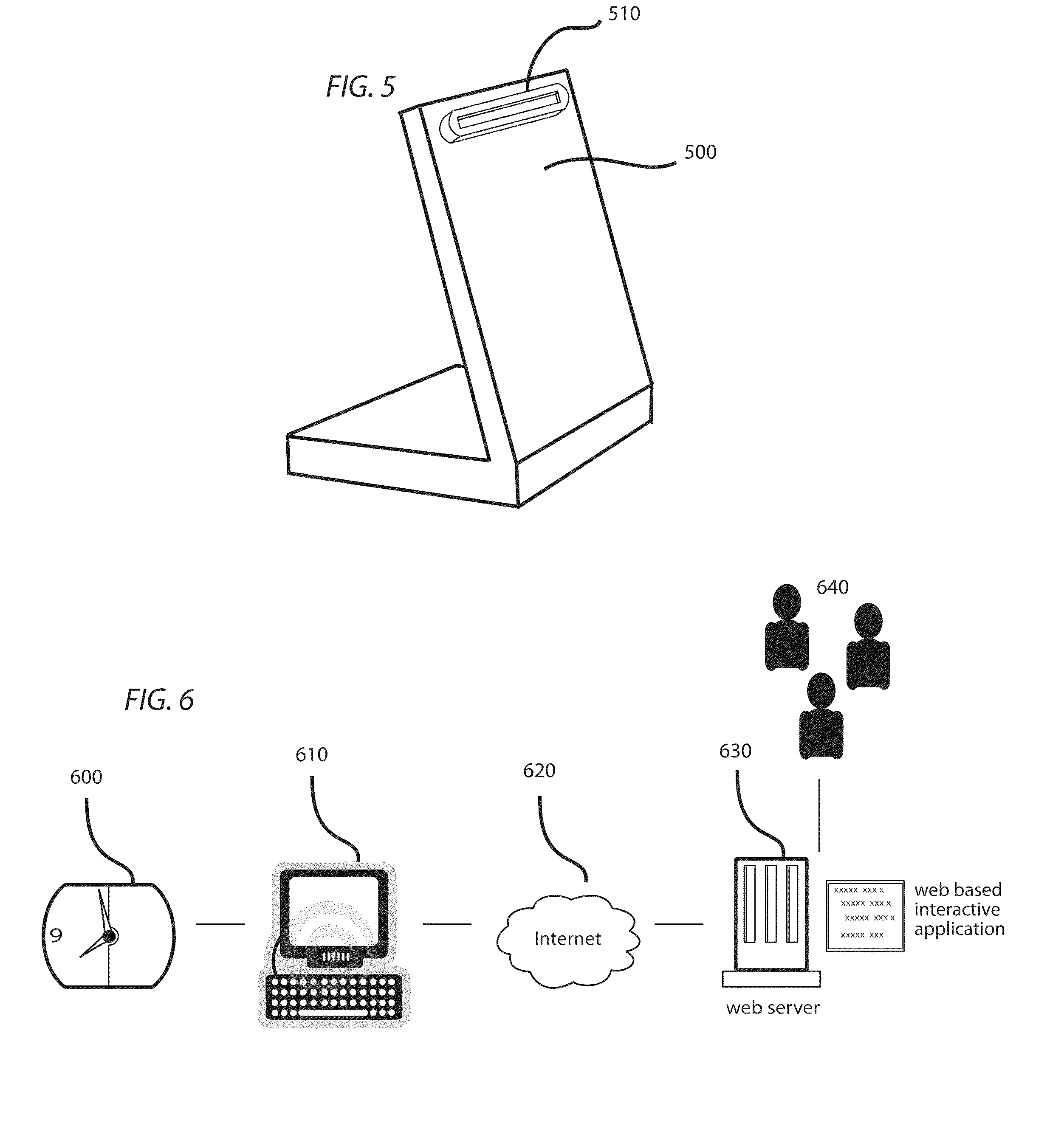
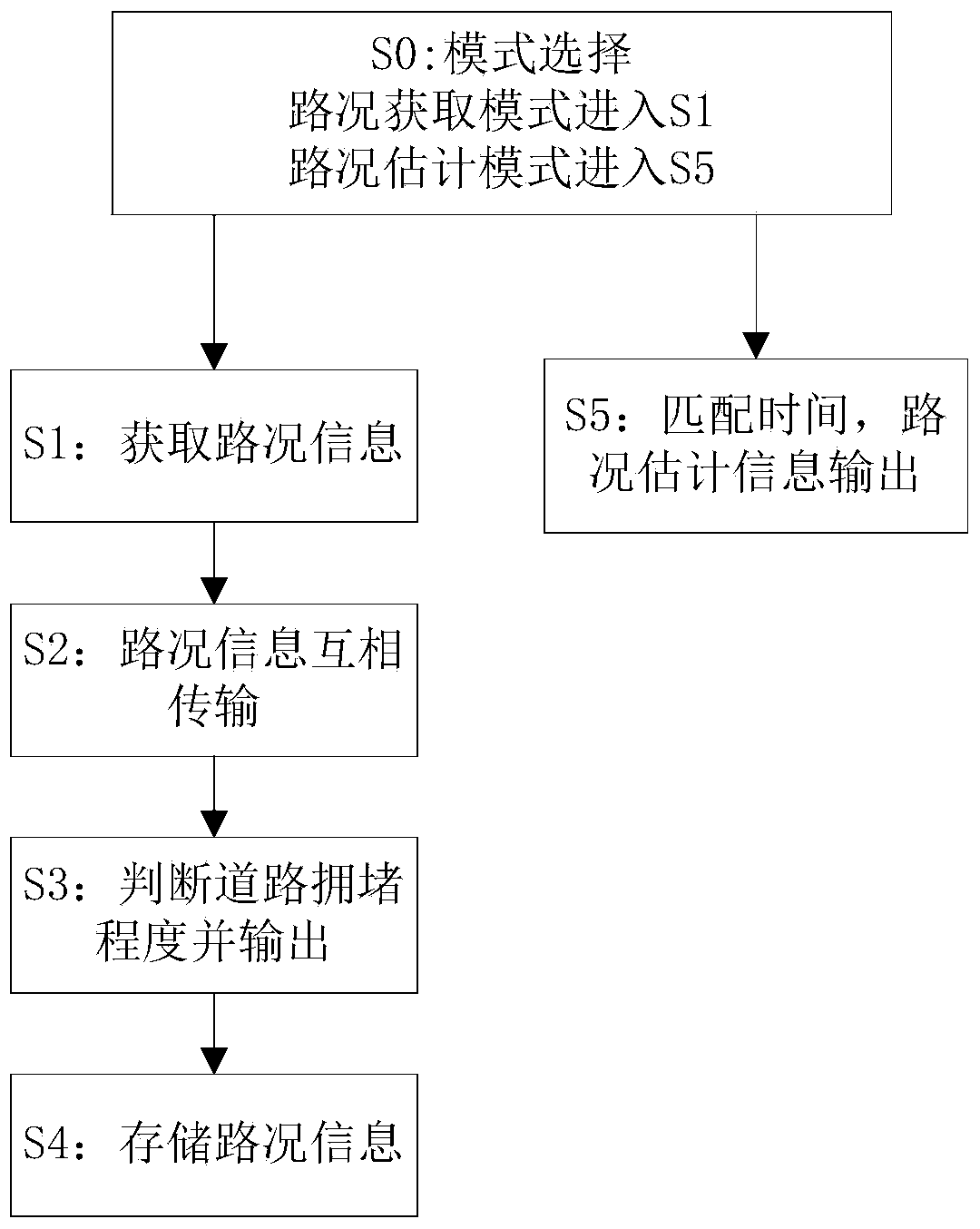
Method and system for acquiring and estimating road conditions based on vehicle-to-vehicle communication
InactiveCN103971529ALarge amount of informationQuickly achieve mutual sharingRoad vehicles traffic controlInformation processingInformation sharing
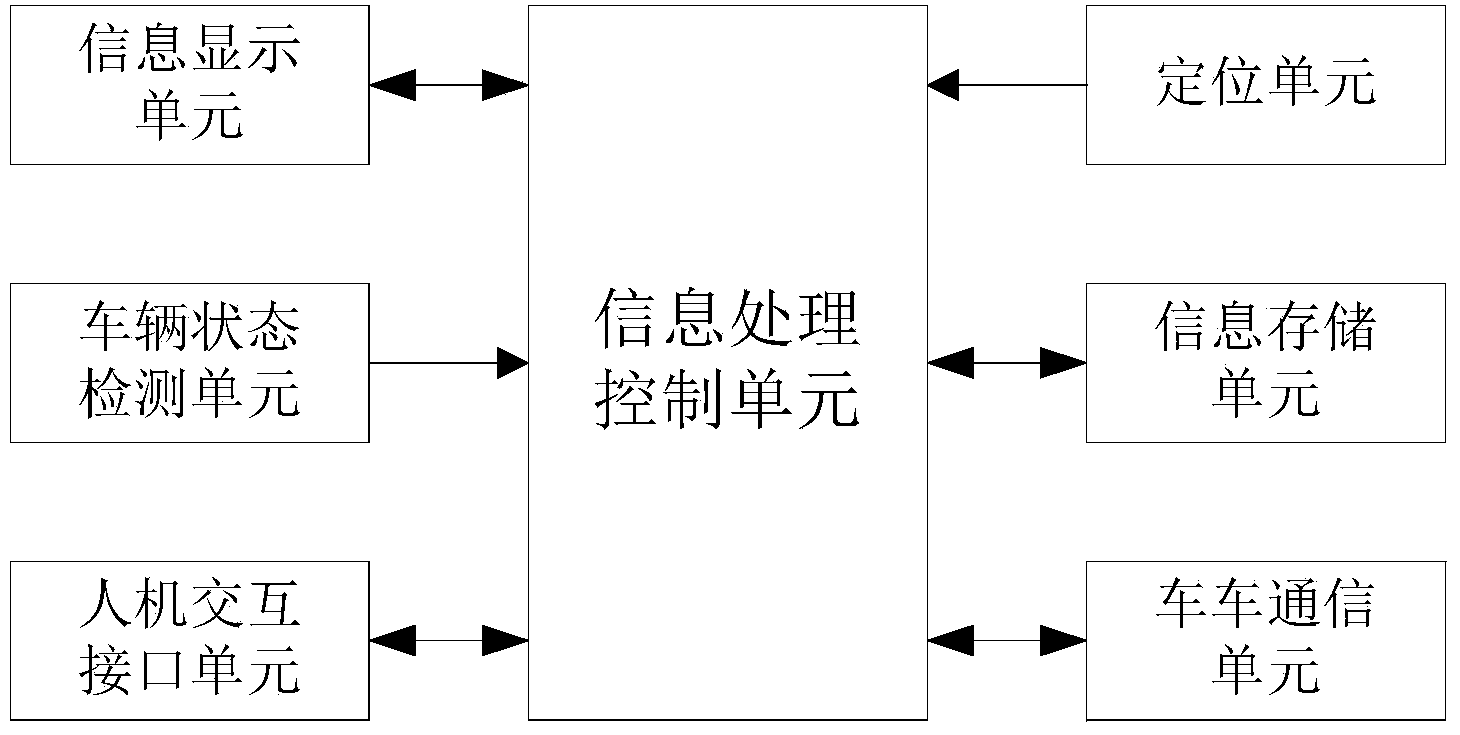
The invention provides a method and system for acquiring and estimating road conditions based on vehicle-to-vehicle communication. Road condition information of vehicles is acquired through a positioning unit, the road condition information is transmitted to other vehicles through a vehicle-to-vehicle communication unit, road condition information of other vehicles is received, the road condition information is processed through an information processing control unit, road condition congestion degrees are acquired and displayed on an electronic map through different colors, and information sharing between the vehicles can be quickly achieved. The quantity of the acquired road condition information is large, the detailed road congestion condition in a certain region surrounding the vehicles can be acquired, all roads in the region can be covered, consumed time in the whole process is short, road conditions can be updated in real time, and road condition services can be provided for specific persons and positions. Positions and time needing road condition estimation are determined, matched road condition information is acquired from stored road condition information and output, the congestion conditions of the roads at a certain moment can be predicted in advance, and users can conveniently conduct path planning in advance.
Owner:SHENZHEN INST OF ADVANCED TECH CHINESE ACAD OF SCI +1
Media player operable to decode content data
InactiveUS20060195909A1Simple authorisation processFunction increaseTelevision system detailsPulse modulation television signal transmissionObfuscationData content
Content is provided on a medium or is transmitted, for example streamed, with in-built protection against unauthorised playback or copying. Video content data is provided in content blocks, each of which relates to a video frame and has a corresponding header. Some content clocks have an excess data item, such as a digital fingerprint, interposed somewhere in the data comprising that content block. Thus, the amount of the data in the content block is increased by the addition of the digital fingerprint. Information identifying the length of the content block in the corresponding header is not altered. Some or all content blocks are obfuscated following the additional of the digital fingerprints. In a media player, de-obfuscation is performed, and digital fingerprints then removed before the resulting content blocks are decoded. Without de-obfuscation and removal of the digital fingerprints, correct decoding would not occur. Thus, playback of the content is limited to a media player that knows the de-obfuscation scheme used and knows the locations of the digital fingerprints.
Owner:ROK PROD LTD
Thermal spray material and process for preparing same
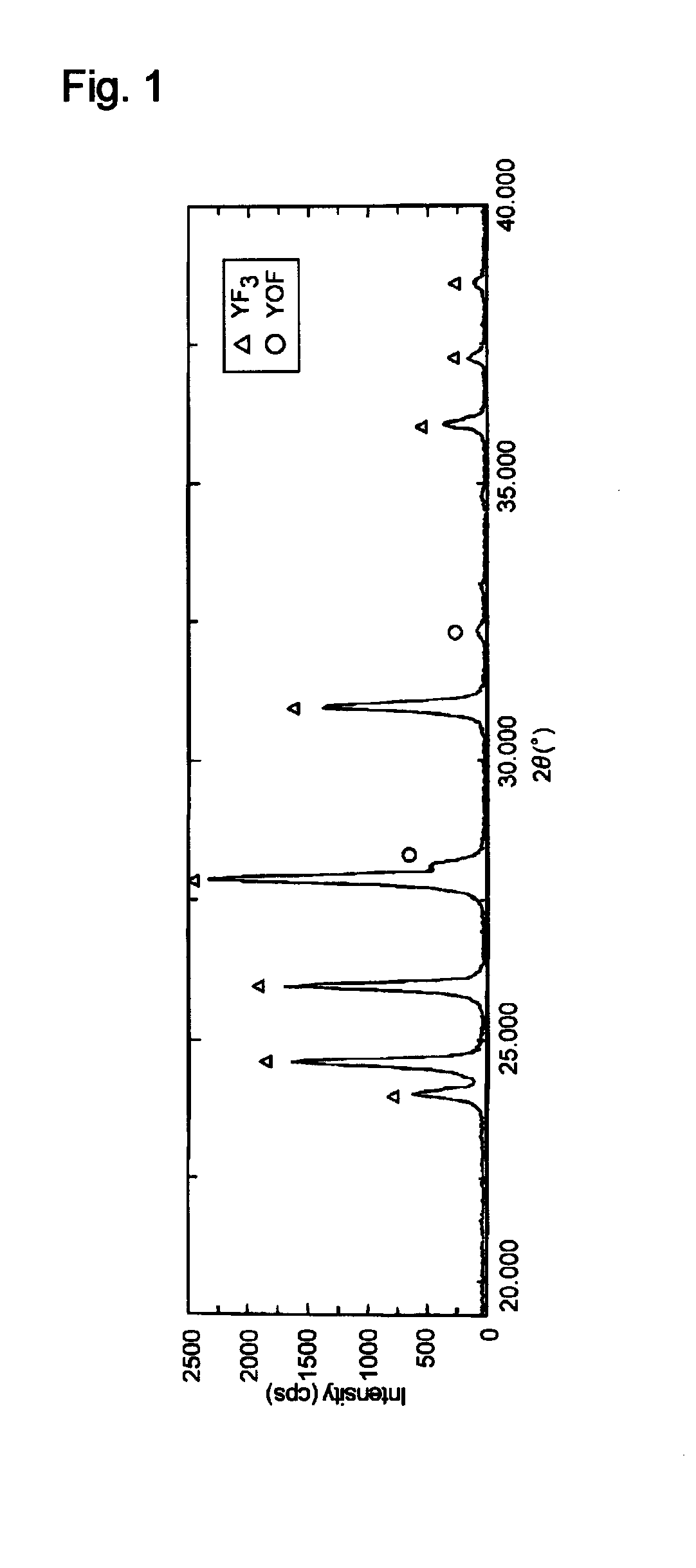
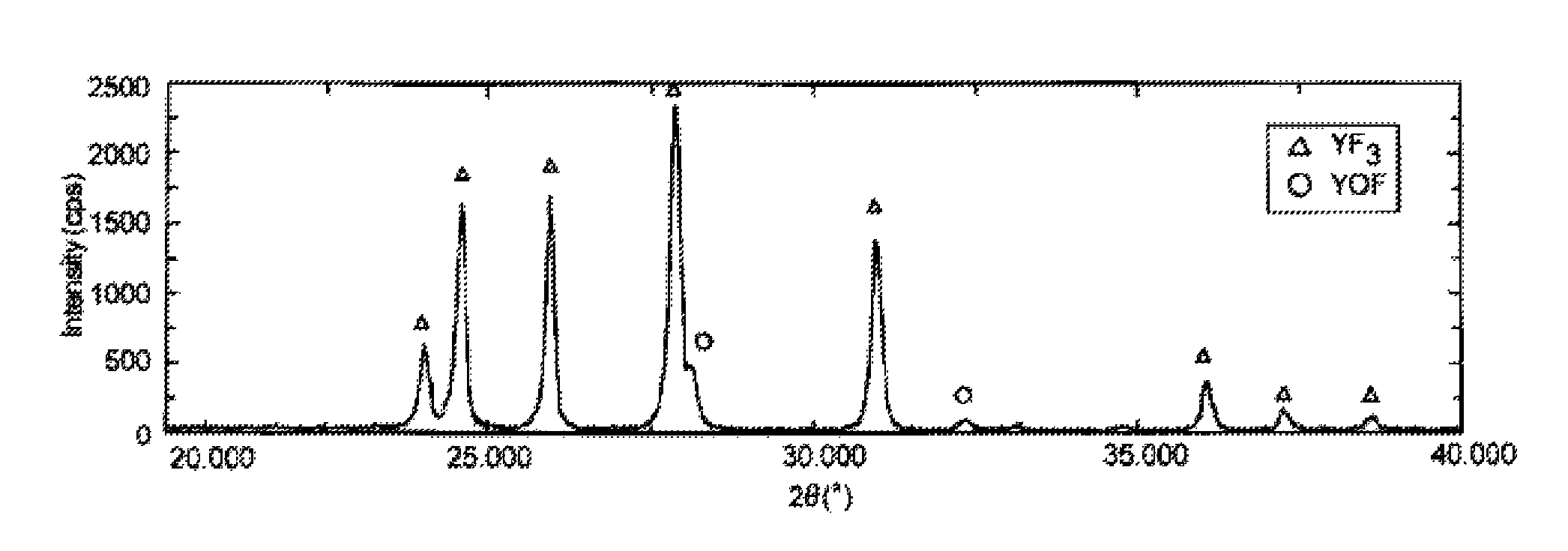
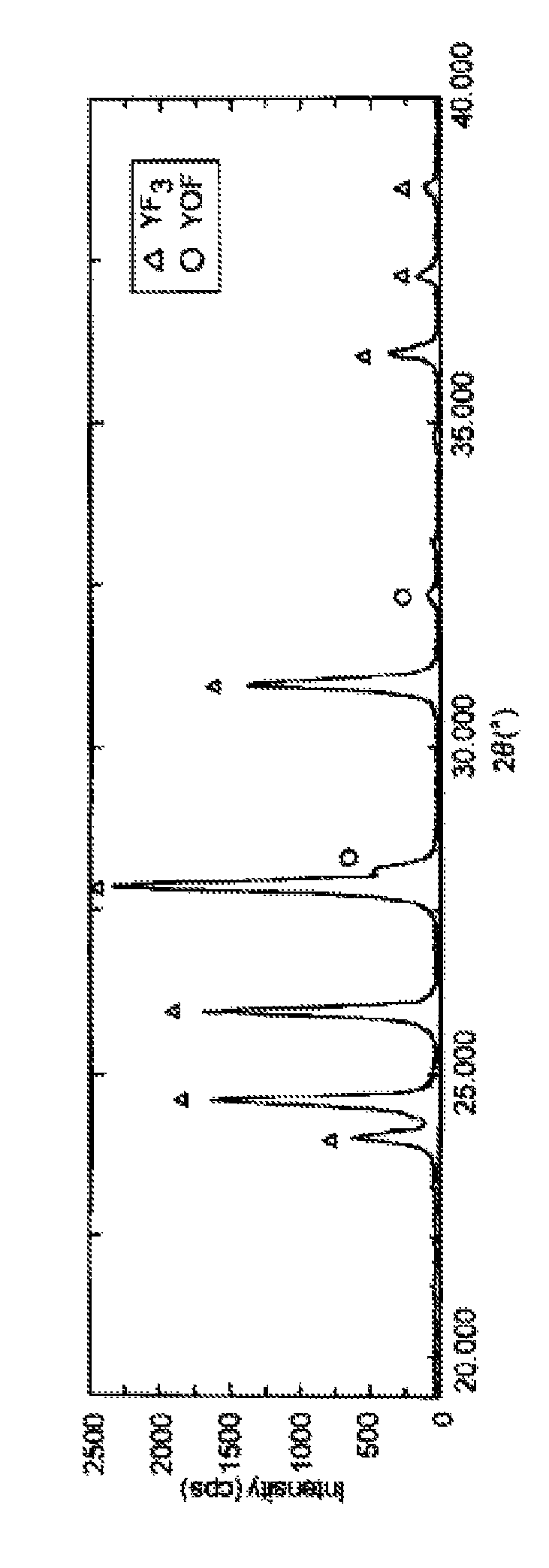
ActiveUS20150096462A1Smooth thermal spray coatingFormed with easeMolten spray coatingRare earth metal compoundsRare-earth elementThermal spraying
A thermal spray material includes granules of an oxyfluoride of yttrium (YOF). The granules may contain a fluoride of yttrium (YF3). The granules preferably have an oxygen content of 0.3 to 13.1 mass %. The granules preferably have a fracture strength of 0.3 MPa or more and less than 10 MPa. Part of yttrium (Y) of the granules may be displaced with at least one rare earth element (Ln) except yttrium, the molar fraction of Ln relative to the sum of Y and Ln being preferably 0.2 or less.
Owner:NIPPON YTTRIUM
Secure access and copy protection management system
InactiveUS20050078822A1Inhibitory contentAccurate contentRecord information storageDigital signal formattingFingerprintCopy protection
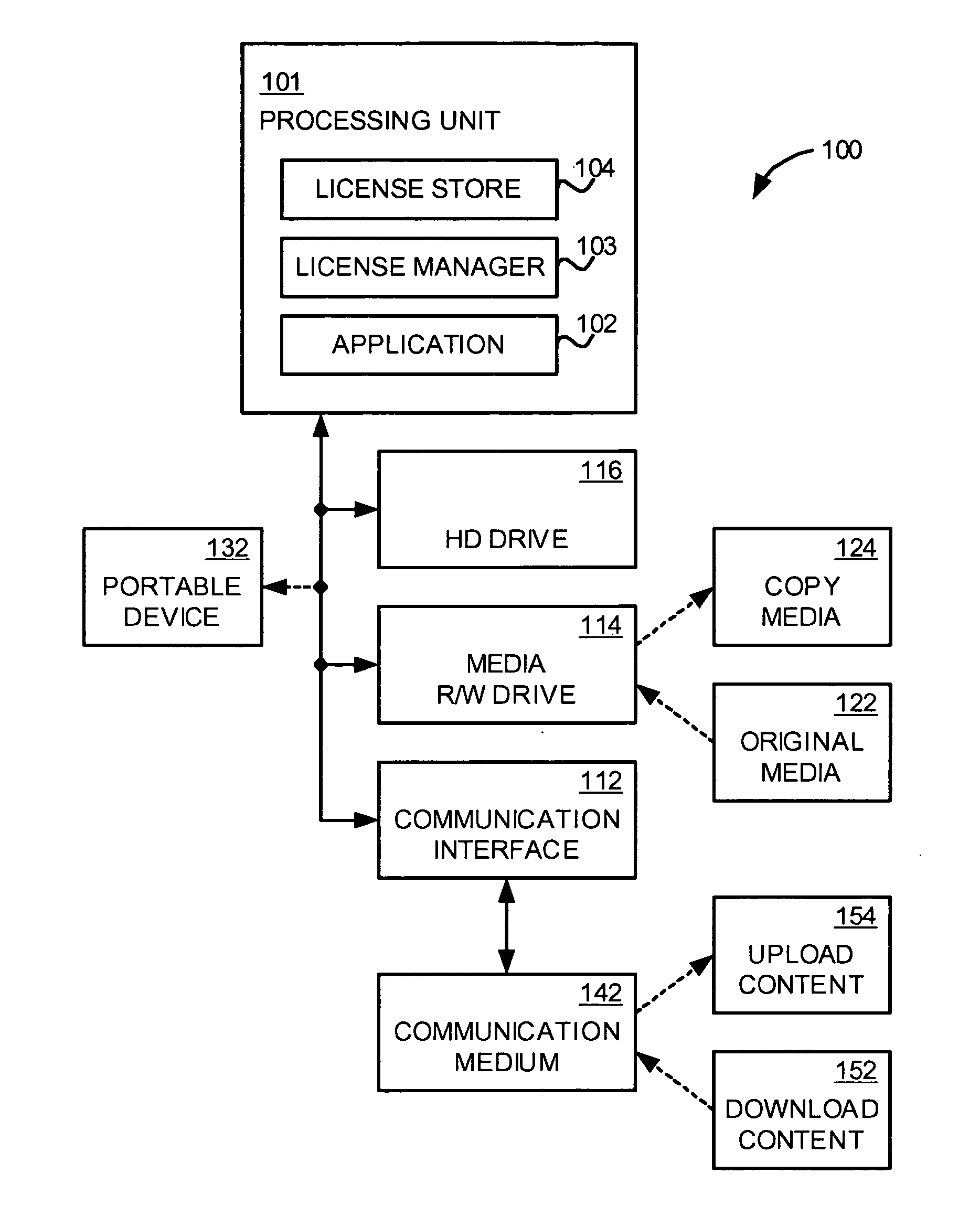
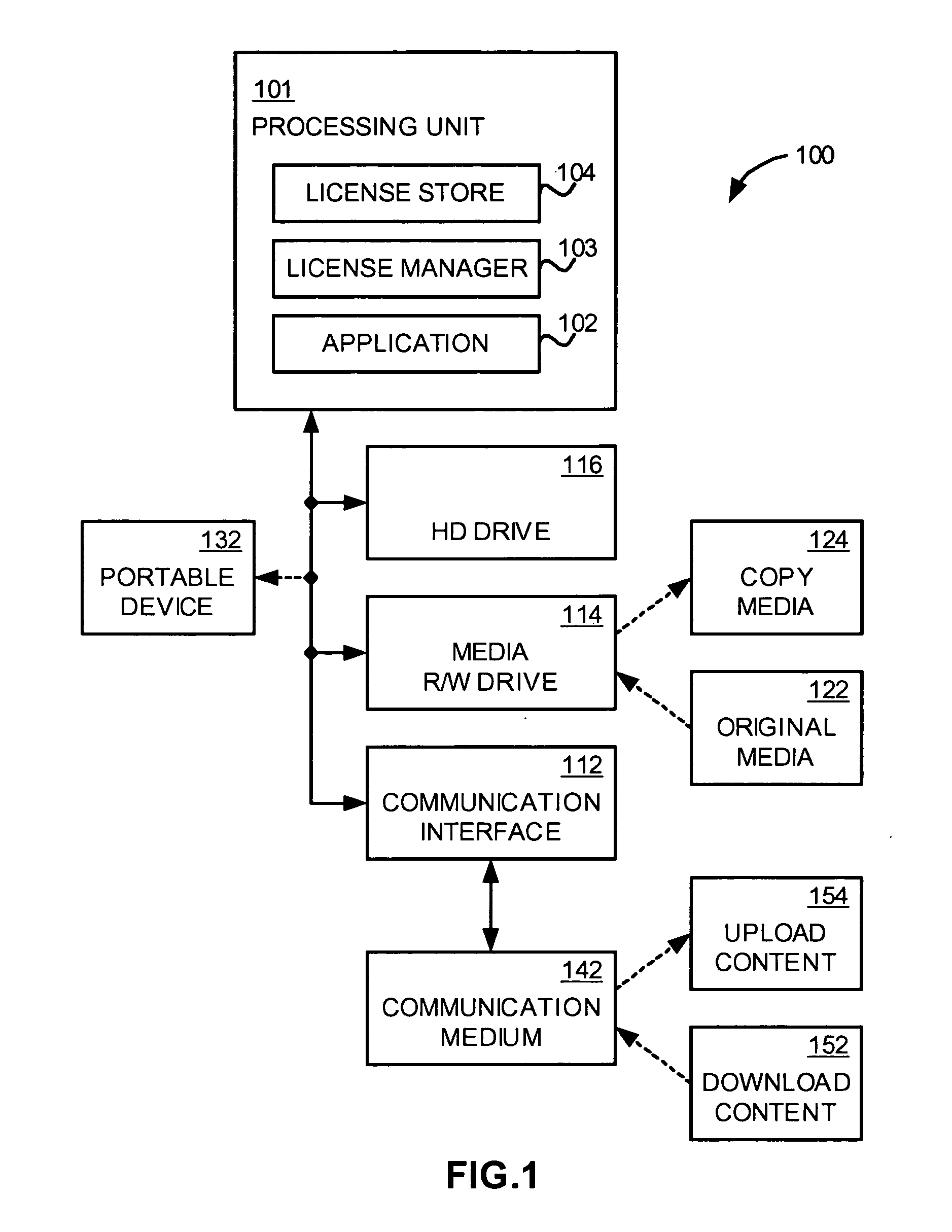
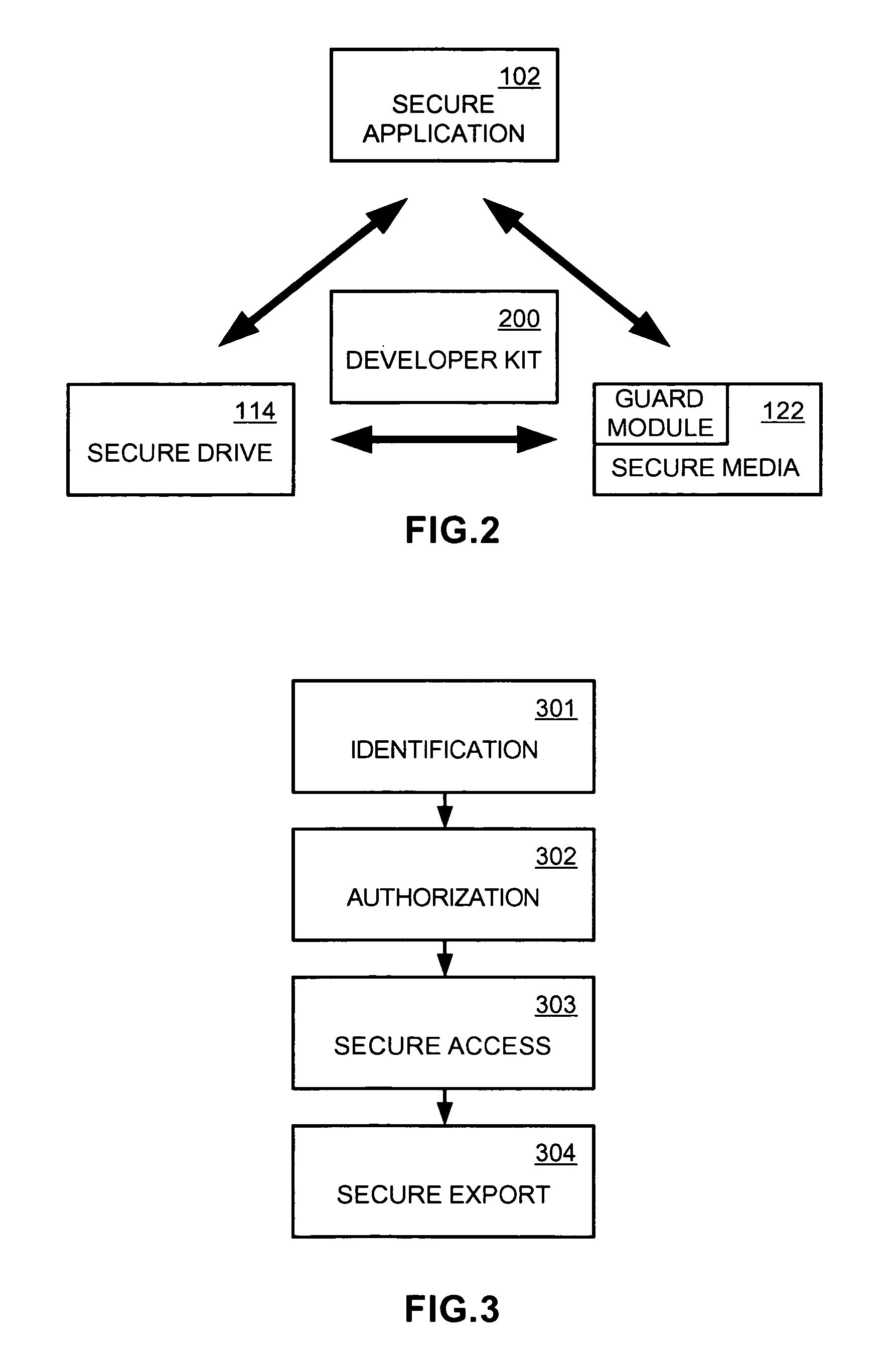
An application, media drive, and media are configured so as to cooperate with one another to provide secure access and copying of protected content on the media. The application cooperates with the media drive to identify the media as a cooperating component, and in the process, also identifies the media drive as a cooperating component. The media includes information facilitating such identifying activity, such as a fingerprint indicating the copy protection method used to protect the content. Also included on the media is a guard module that authenticates the application as a cooperating component, establishes secure channels respectively with the application and the media drive for communicating secret information, installs licenses included on the media, and only allows access to the protected content if the media is an original copy. The application then manages usage and / or copying of the protected content according to the installed licenses.
Owner:ROVI SOLUTIONS CORP
Method and device for combining entities in knowledge map
InactiveCN104484459AAccurate contentCompact structureSpecial data processing applicationsFeature vectorKnowledge graph
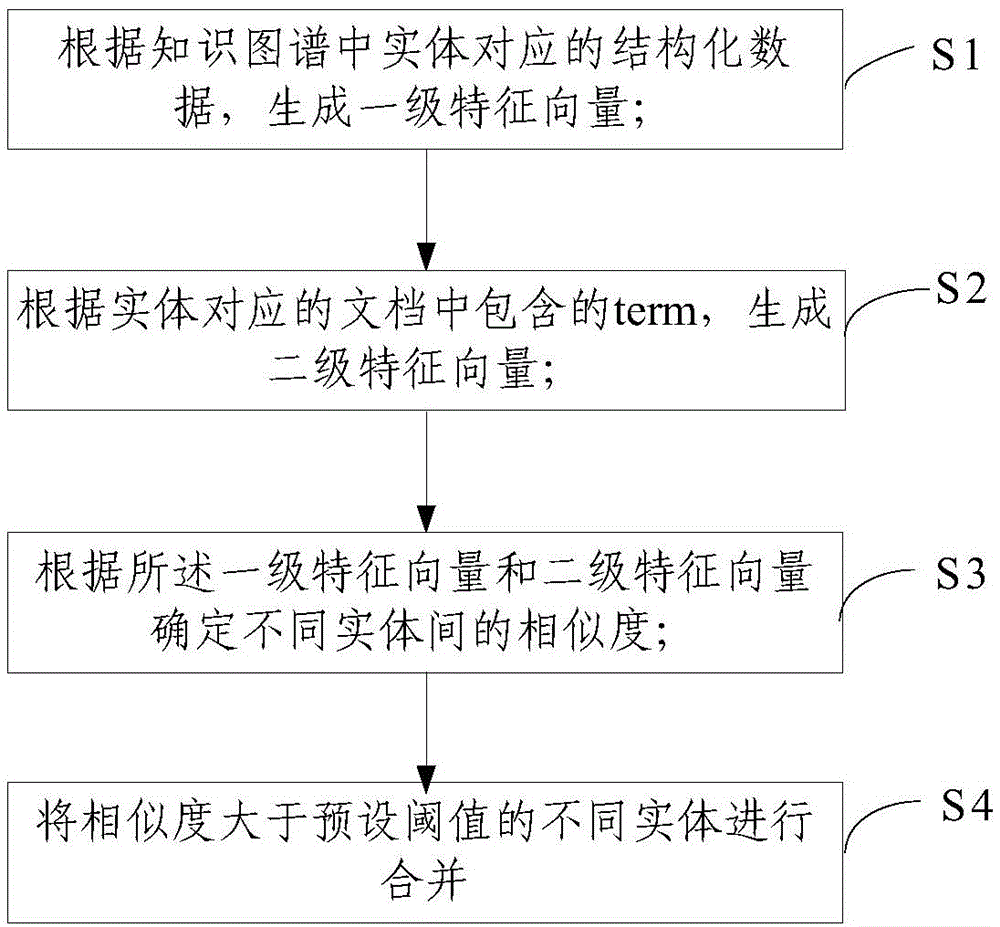
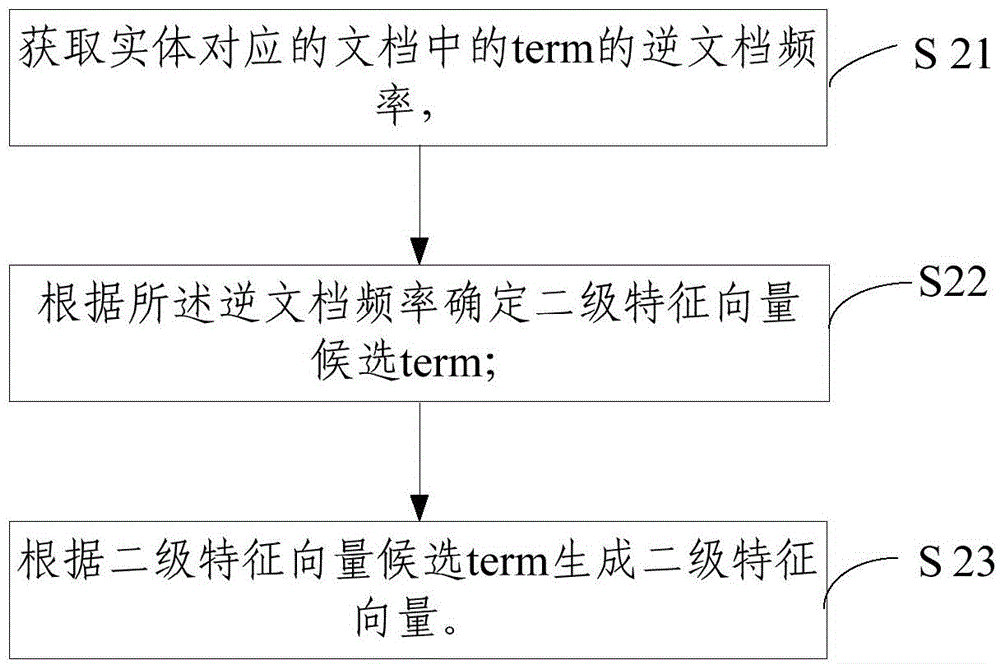
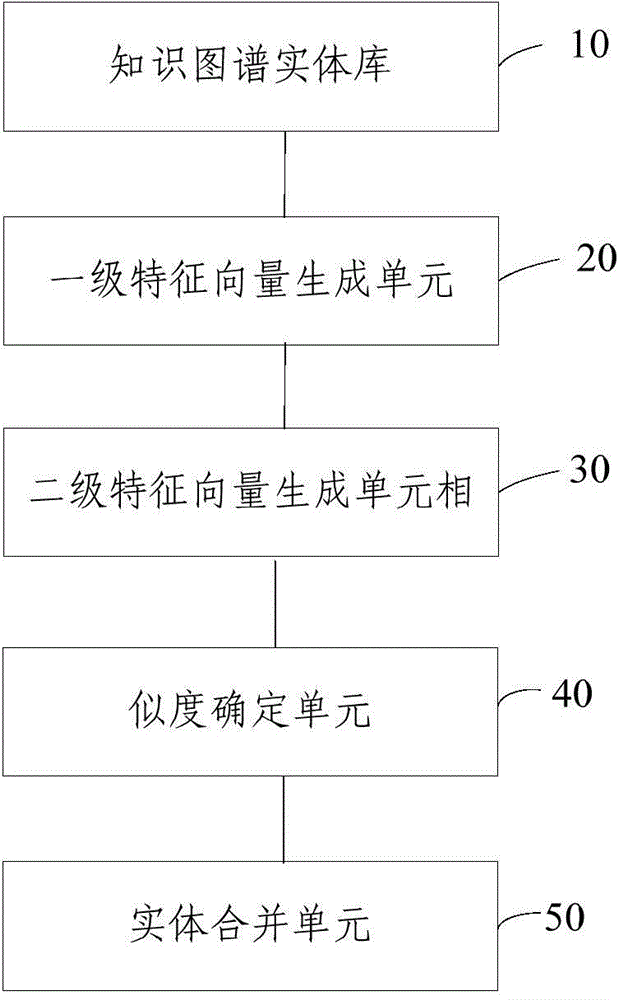
The invention relates to a method and a device for combining entities in a knowledge map. The method comprises the following steps of generating a first-grade feature vector according to structural data corresponding to the entities in the knowledge map; generating a second-grade feature vector according to terms in documents corresponding to the entities; determining the similarity of the different entities according to the first-grade feature vector and the second-grade feature vector. According to the method and the device, the first-grade feature vector and the second-grade feature vector are constructed by entity IDs respectively, and the similarity of the entity IDs with the same name is calculated, and whether the entity IDs with the same name are the same object or not can be accurately judged, so that a condition that one object in the knowledge map has a plurality of entity IDs can be reduced, the content of the knowledge map is more accurate and the structure is more compact.
Owner:BEIJING QIHOO TECH CO LTD +1
Internet streaming and the presentation of dynamic content
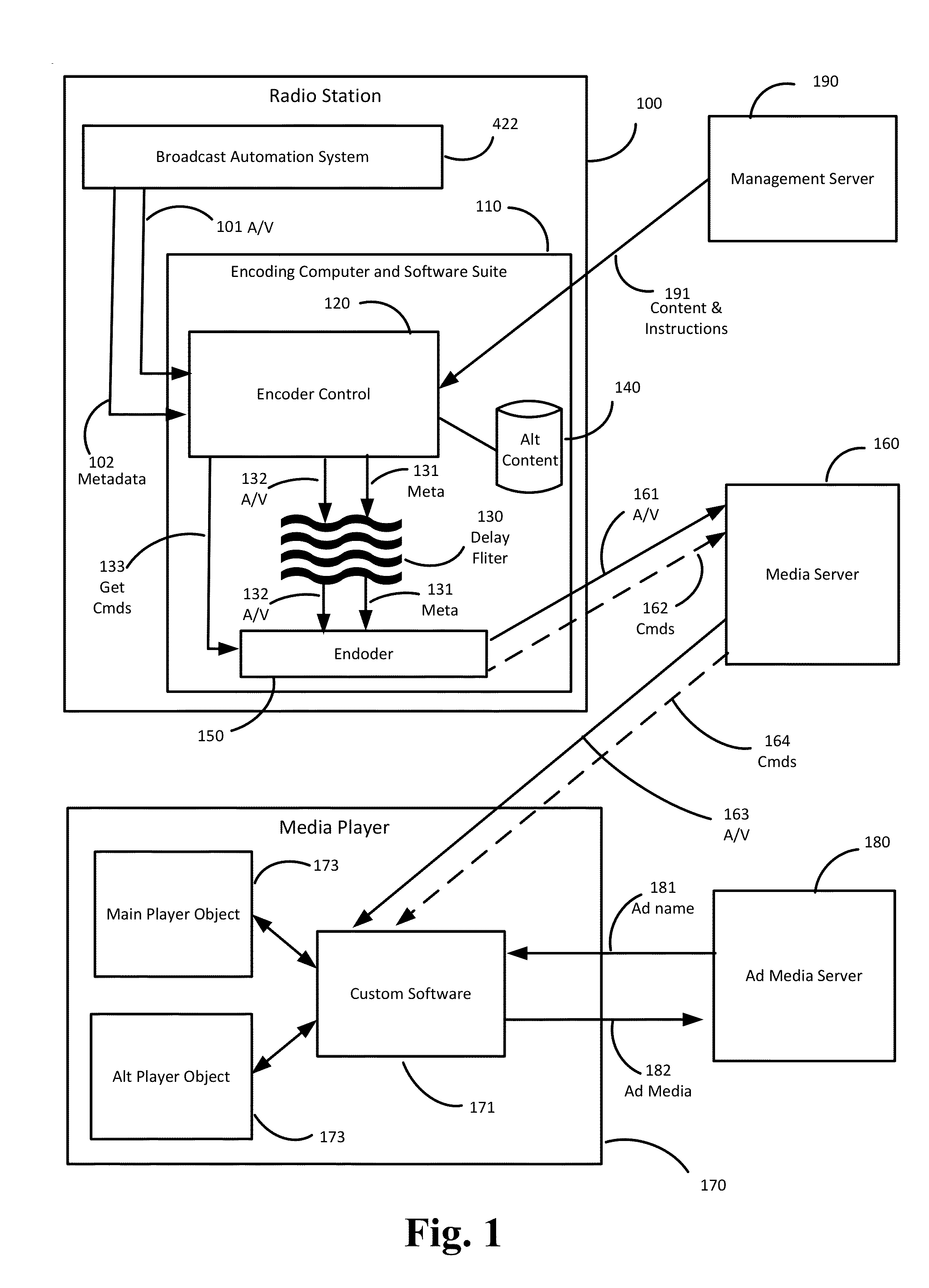
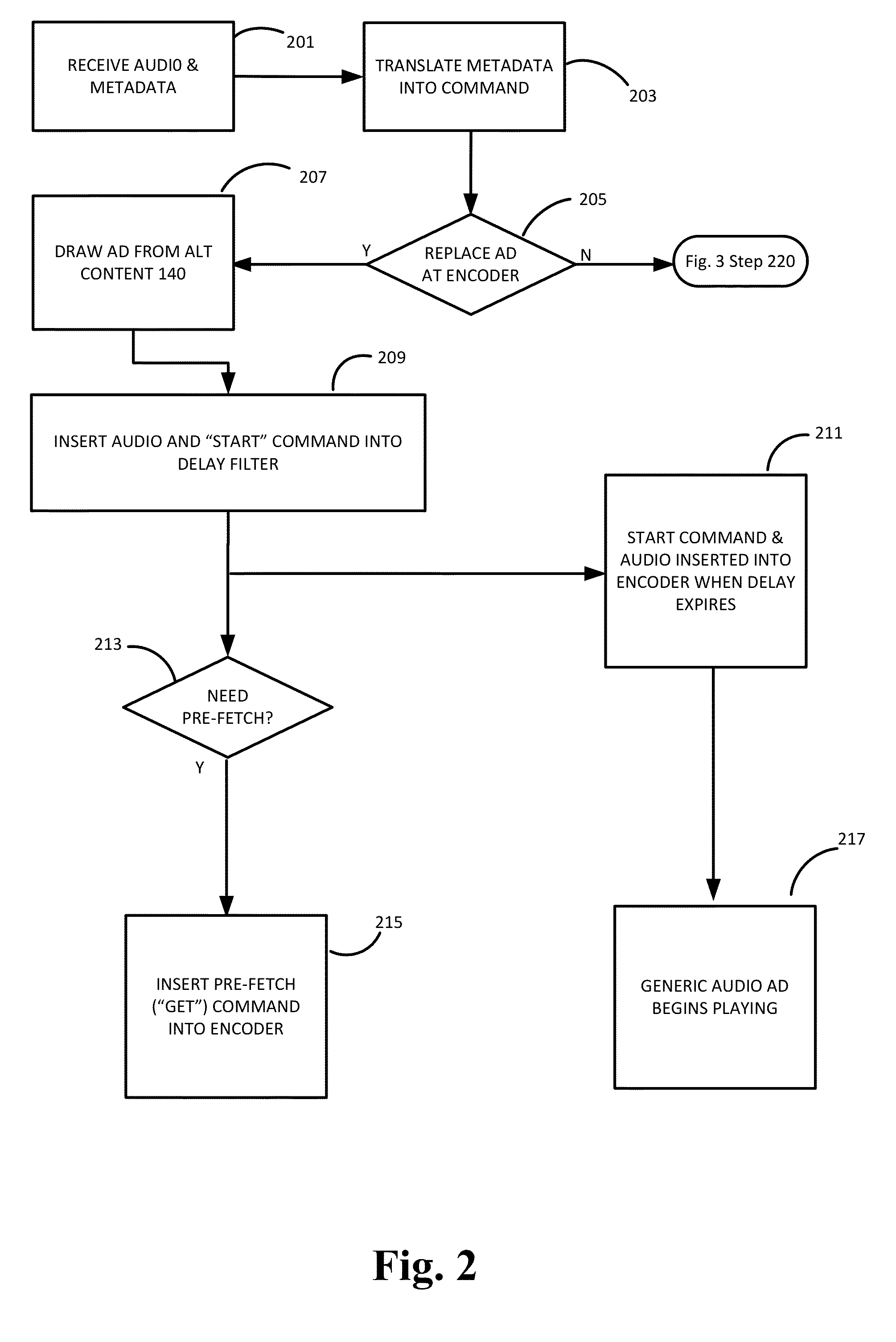
ActiveUS20140149596A1Accurate contentAccurate timingMultiple digital computer combinationsProgram controlThe InternetDistribution system
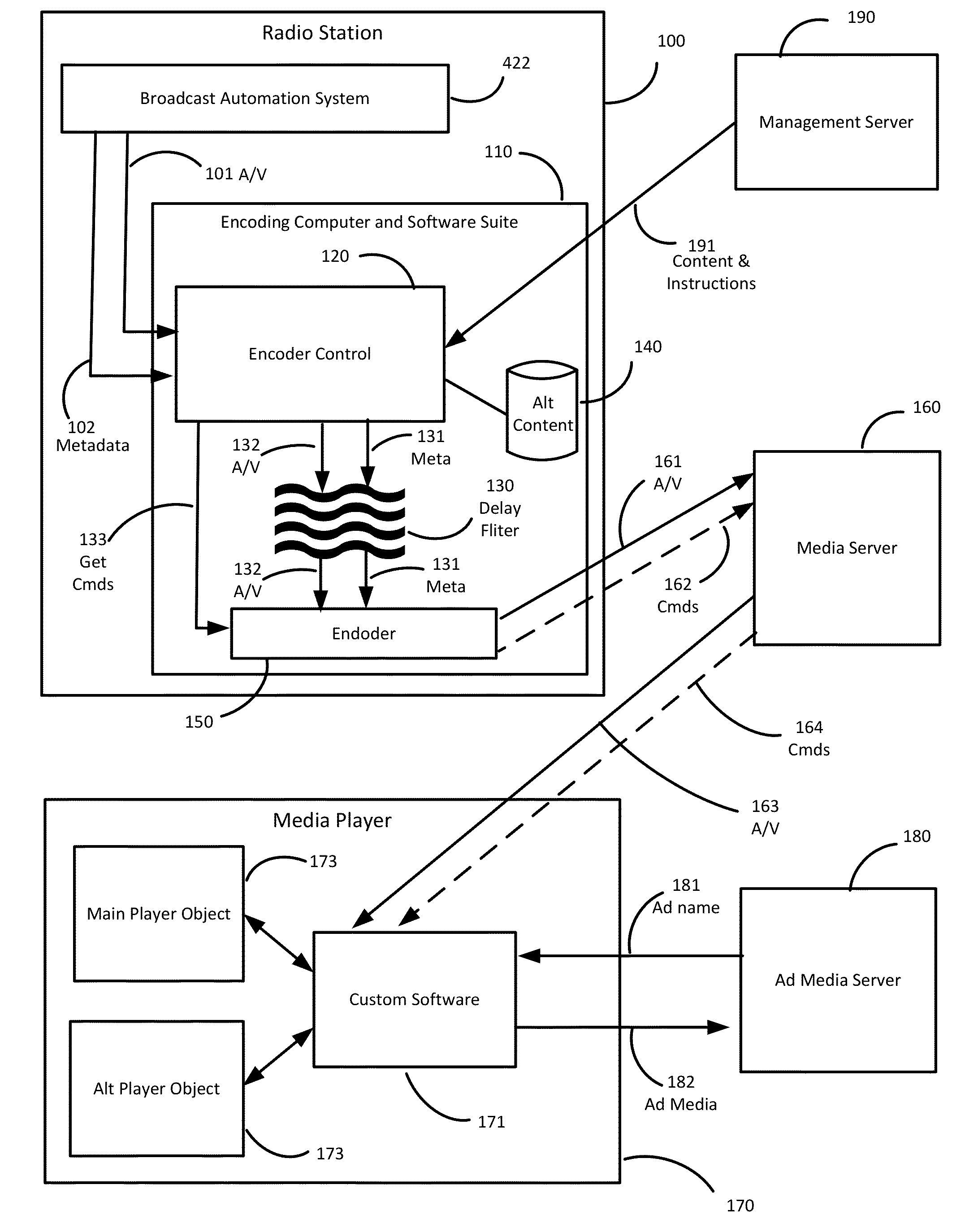
Internet streaming from broadcast radio or television stations is described wherein triggers for dynamic content from internal or external systems cause an encoder system to generate command messages, and to synchronize those command messages with any delays associated with the triggering events. Command messages are delivered through a streaming media distribution system to client media players which obtain or present the dynamic content, in association with any desired configuration changes to the appearance of the media player or the method or manner in which the dynamic content is presented.
Owner:EMERSON III HARRY E
Imaging device, image processing method and program
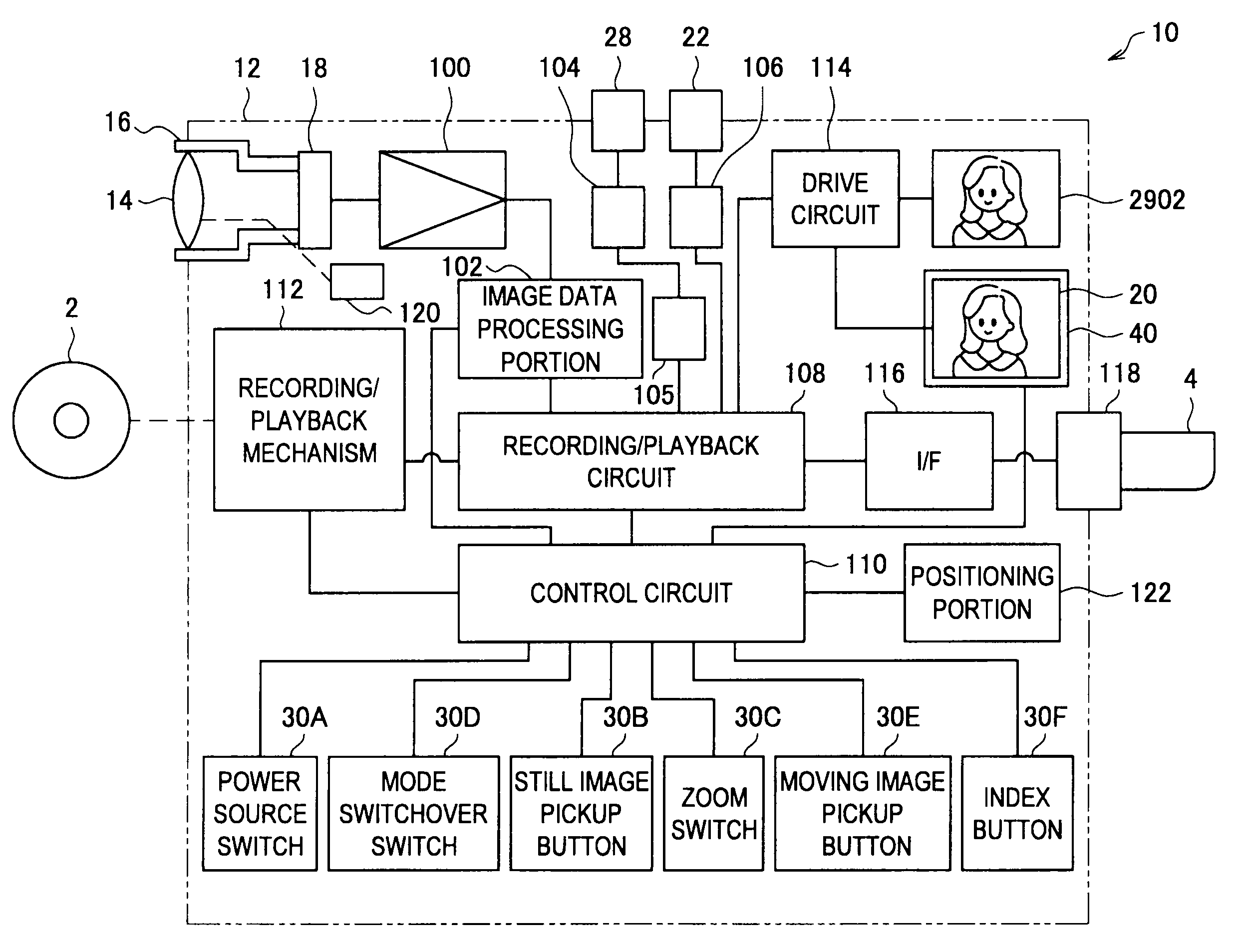
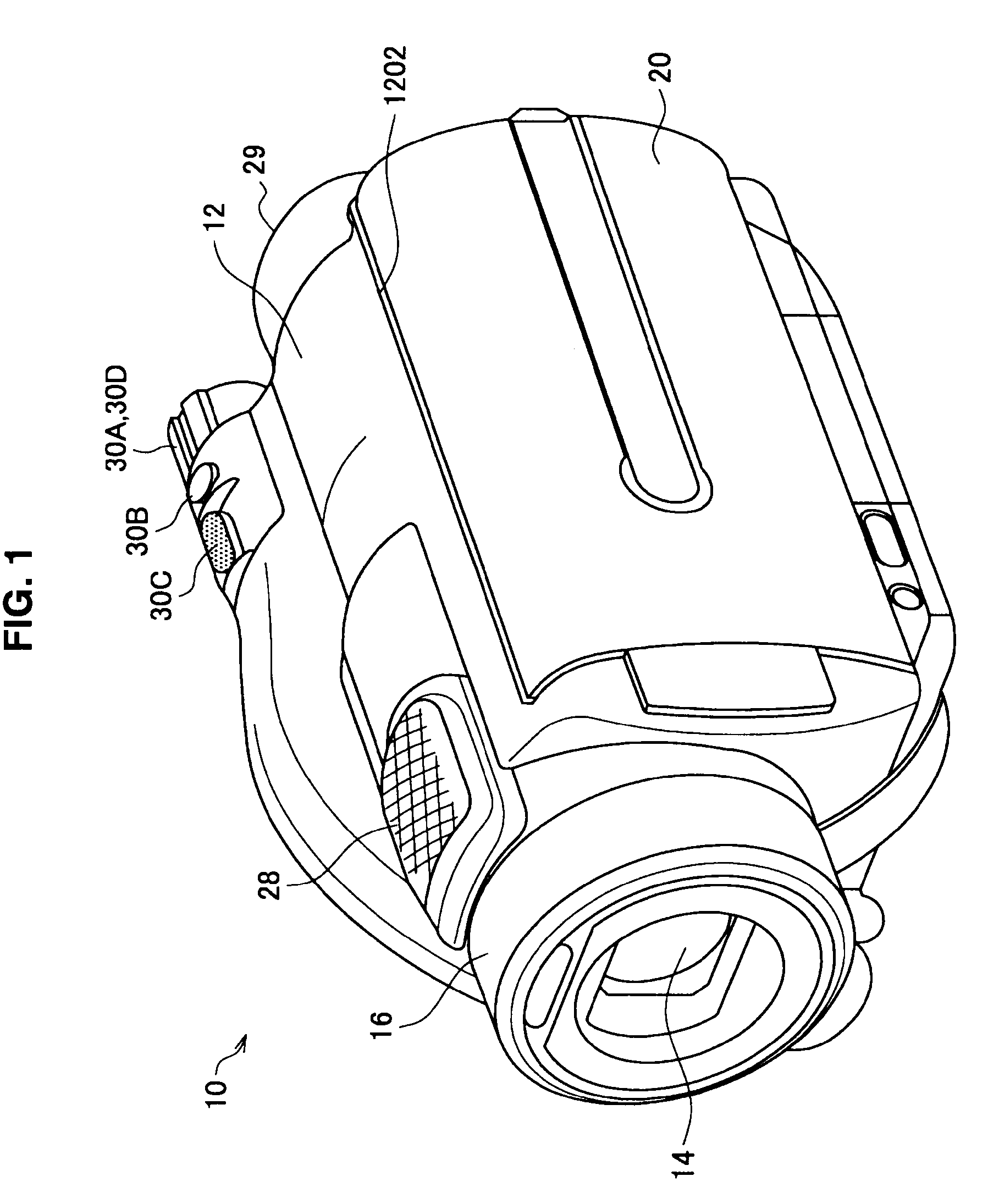
ActiveUS20100310232A1Accurate displayAccurate contentTelevision system detailsColor television signals processingComputer hardwareImaging processing
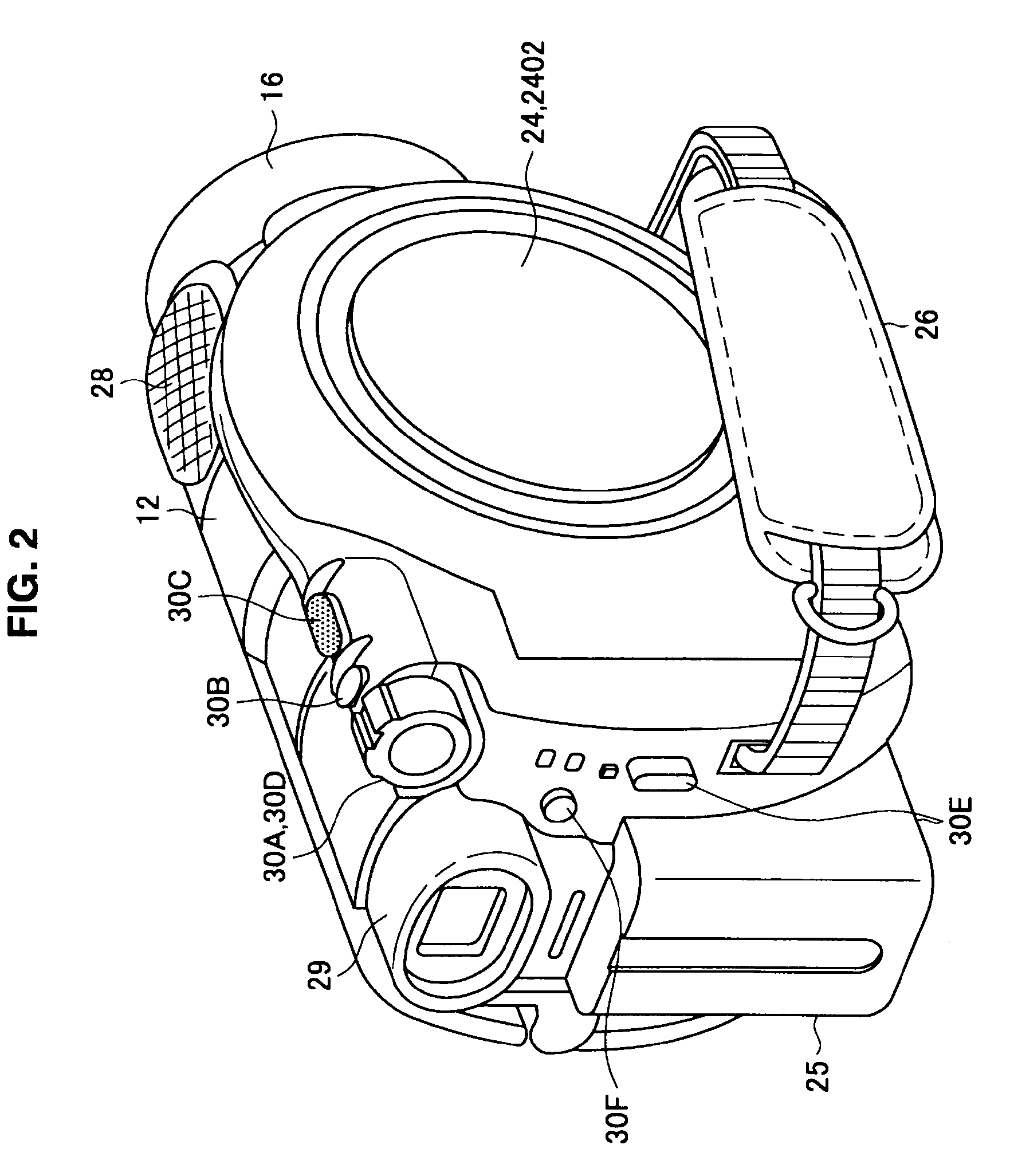
There is provided an imaging device including, an operating portion on which are input a recording start instruction and a recording stop instruction, a recording portion that records onto a recording medium moving images that are picked up during a recording period from the recording start instruction to the recording stop instruction, a thumbnail generation portion that generates a plurality of thumbnail images that represent each of sections obtained by time division of the moving images, and a display control portion that displays a recorded image verification screen on a display portion immediately after the recording stop instruction, the plurality of thumbnail images being arranged in chronological order on the recorded image verification screen.
Owner:SONY CORP
System and method for drm content management
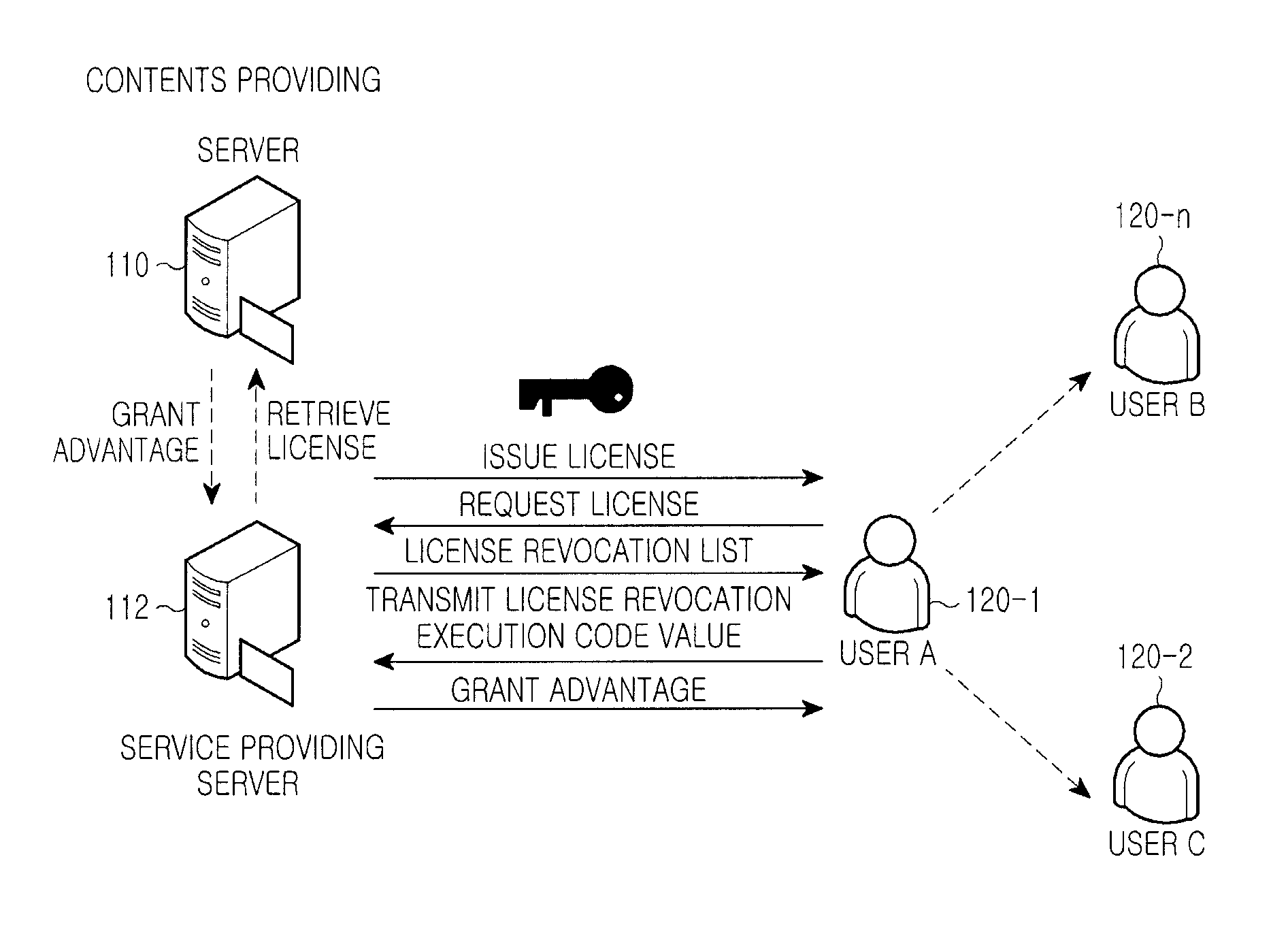
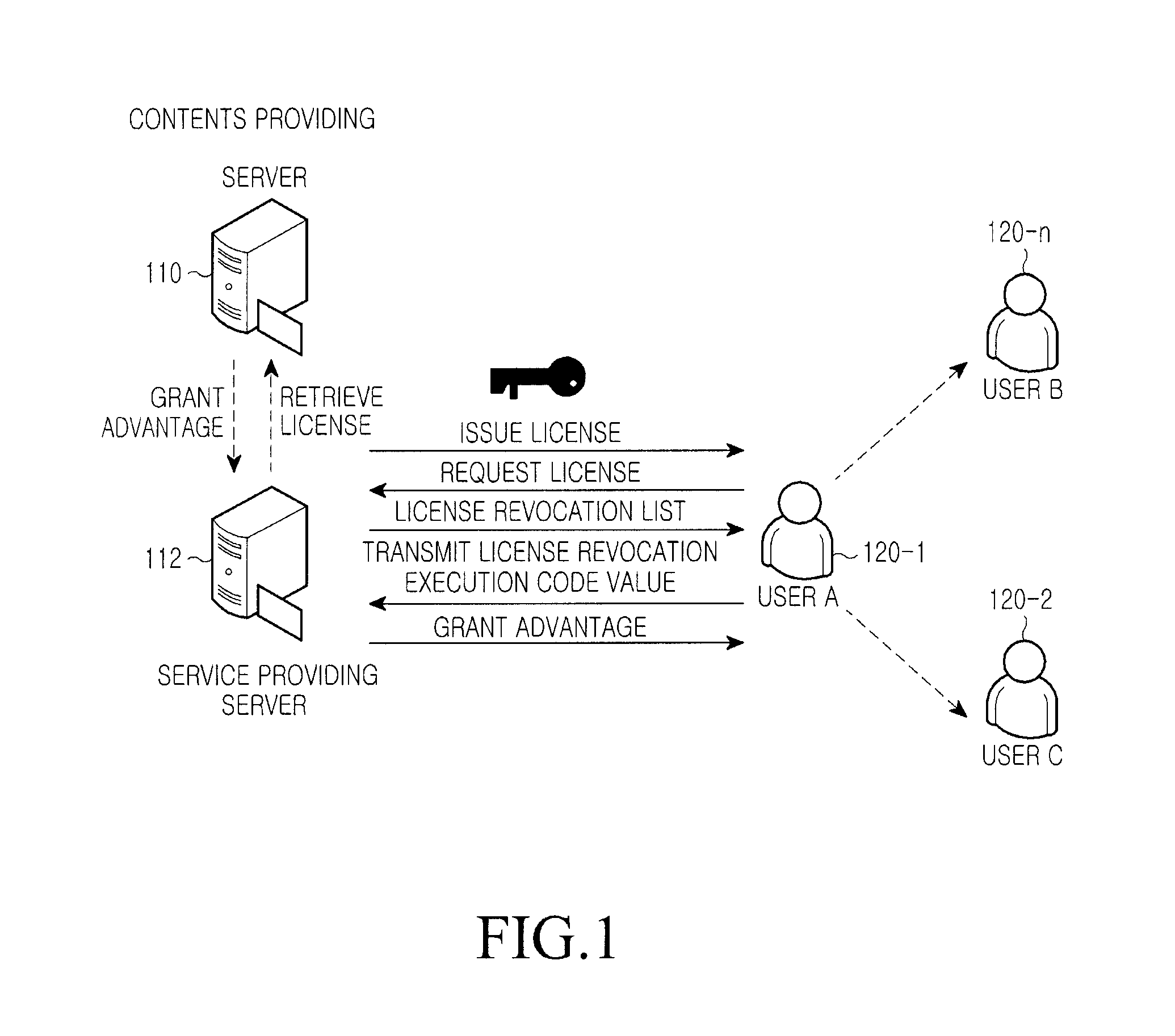
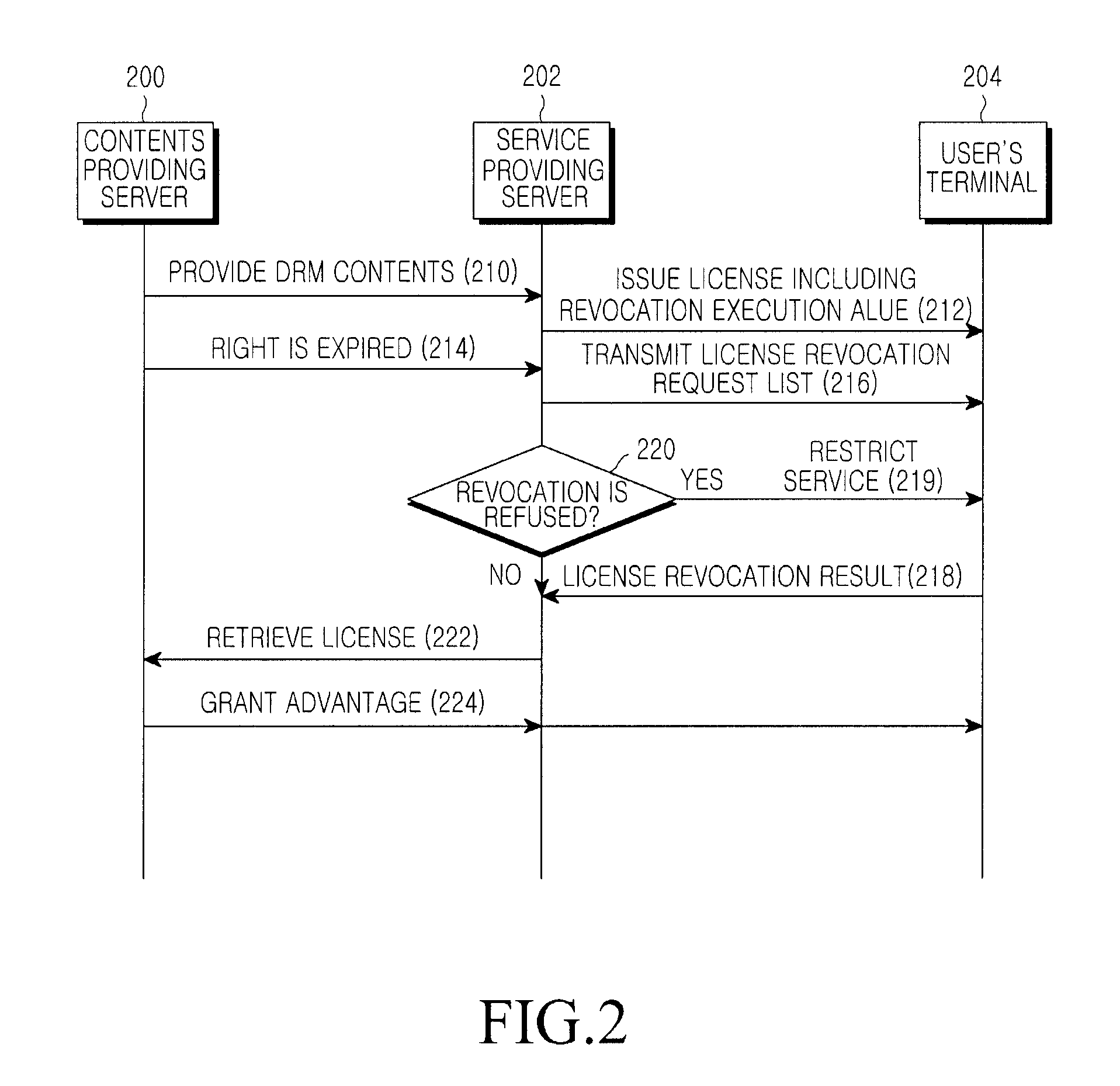
InactiveUS20110047080A1Accurate contentDigital data processing detailsAnalogue secracy/subscription systemsEngineeringDigital rights management
A system for managing a Digital Rights Management (DRM) content includes a content providing server for providing an environment, through which a DRM content and a digital right of the DRM content can be registered, a service providing server for issuing a license serving as a usage authority for each of DRM content files provided from the content providing server, generating a license revocation execution value in a specific field of the issued license, encrypting the generated license revocation execution value for transmission to a user terminal, and the user terminal for inspecting the license of the DRM content file transmitted from the service providing server, and extracting and storing the license and the license revocation execution value to use a corresponding DRM content file according to the license, and upon receiving a revocation request, transmitting a revocation result of the corresponding license to the service providing server.
Owner:SAMSUNG ELECTRONICS CO LTD
Apparatus for providing voice dialogue service and method of operating the same
InactiveUS20070208556A1Accurate analysisAccurate contentSemantic analysisSpeech recognitionPart of speechSpeech sound
A speech dialogue service apparatus including: a language analysis module tagging a part of speech (POS) of each respective word included in a sentence recorded in a predetermined text, syntactically analyzing the sentence by classifying a meaning of each respective word, and generating at least one semantic frame corresponding to the sentence according to a result of the syntactical analysis; and a dialogue management module analyzing an intention of the sentence corresponding to the at least one respective semantic frame, and generating a system response corresponding to the sentence intention by selecting a predetermined sentence intention according to whether an action corresponding to the intention of the respective sentence can be performed.
Owner:SAMSUNG ELECTRONICS CO LTD
System and Method for Combining Media Data
InactiveUS20080189735A1Limited abilityIncrease awarenessElectronic editing digitised analogue information signalsElectrical cable transmission adaptationData setData system
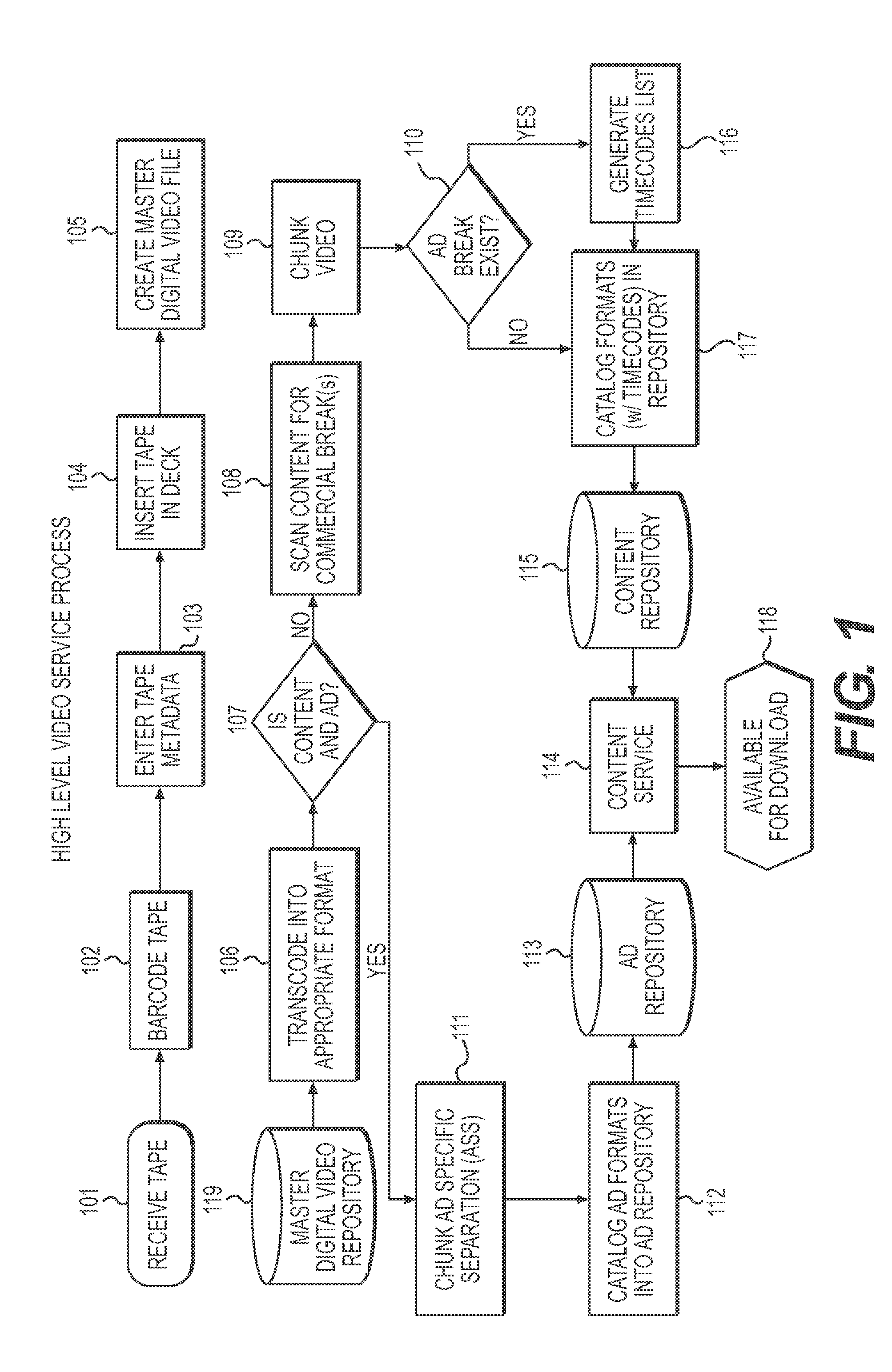
System and method for processing media data set and transmission of the same in a data communication medium. A first set of data is scanned for specific time-indicating markers. A second set of data comprising one or more sub-sets of data are selected according to a predetermined set of criteria, and the second one or more sub-sets of data are inserted into the first set of data at the identified time-indicating markers located in the first data set. The method further comprises converting the data sets to formats suitable for viewing by a consumer. The new, combined data set is transmitted to the consumer.
Owner:ITT MFG ENTERPRISES LLC
Systems and methods for partitioning search indexes for improved efficiency in identifying media segments
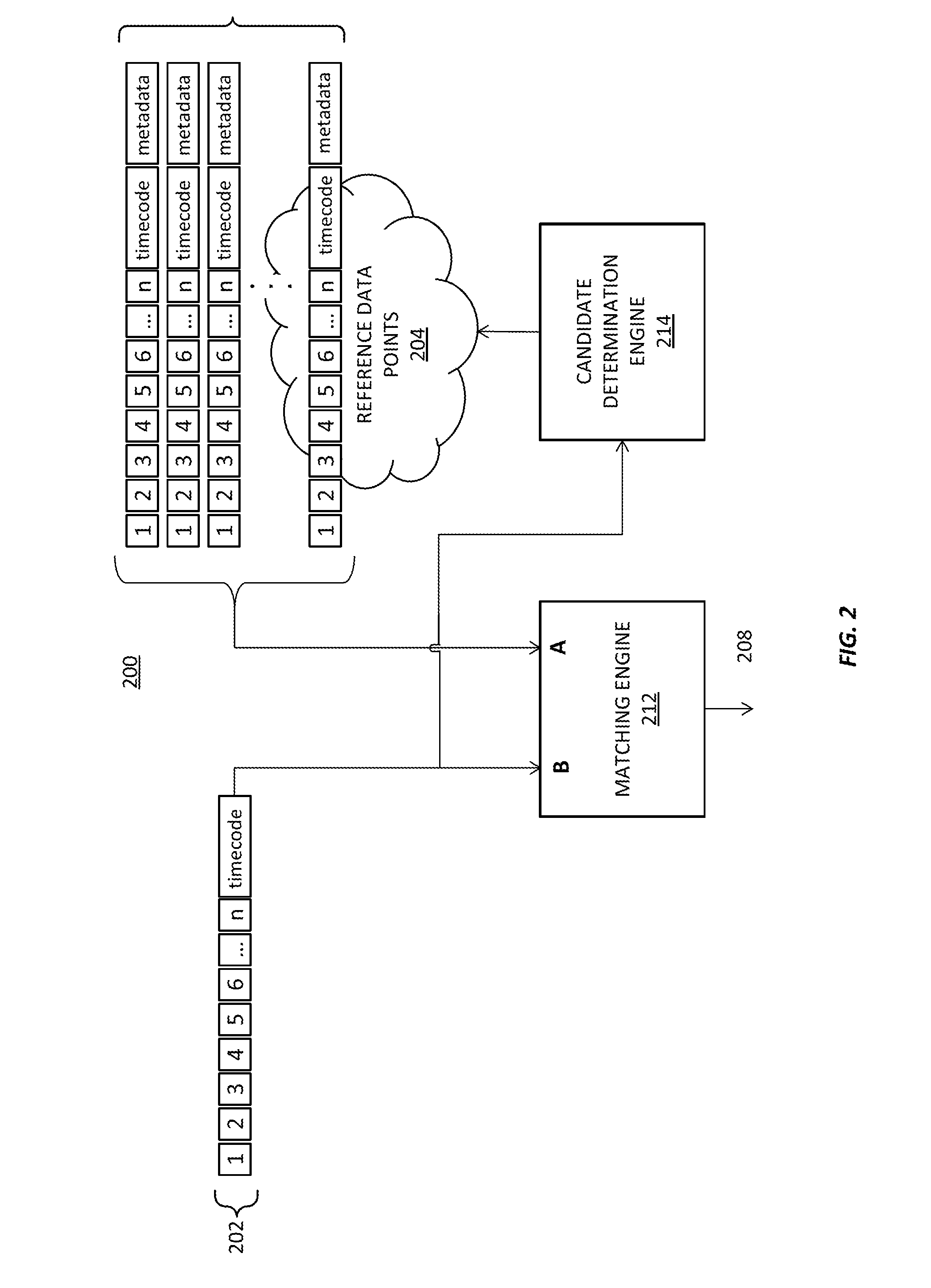
ActiveUS20170017645A1Enhance accuracyImprove accuracyVideo data indexingMultimedia data indexingReference databaseMedia content
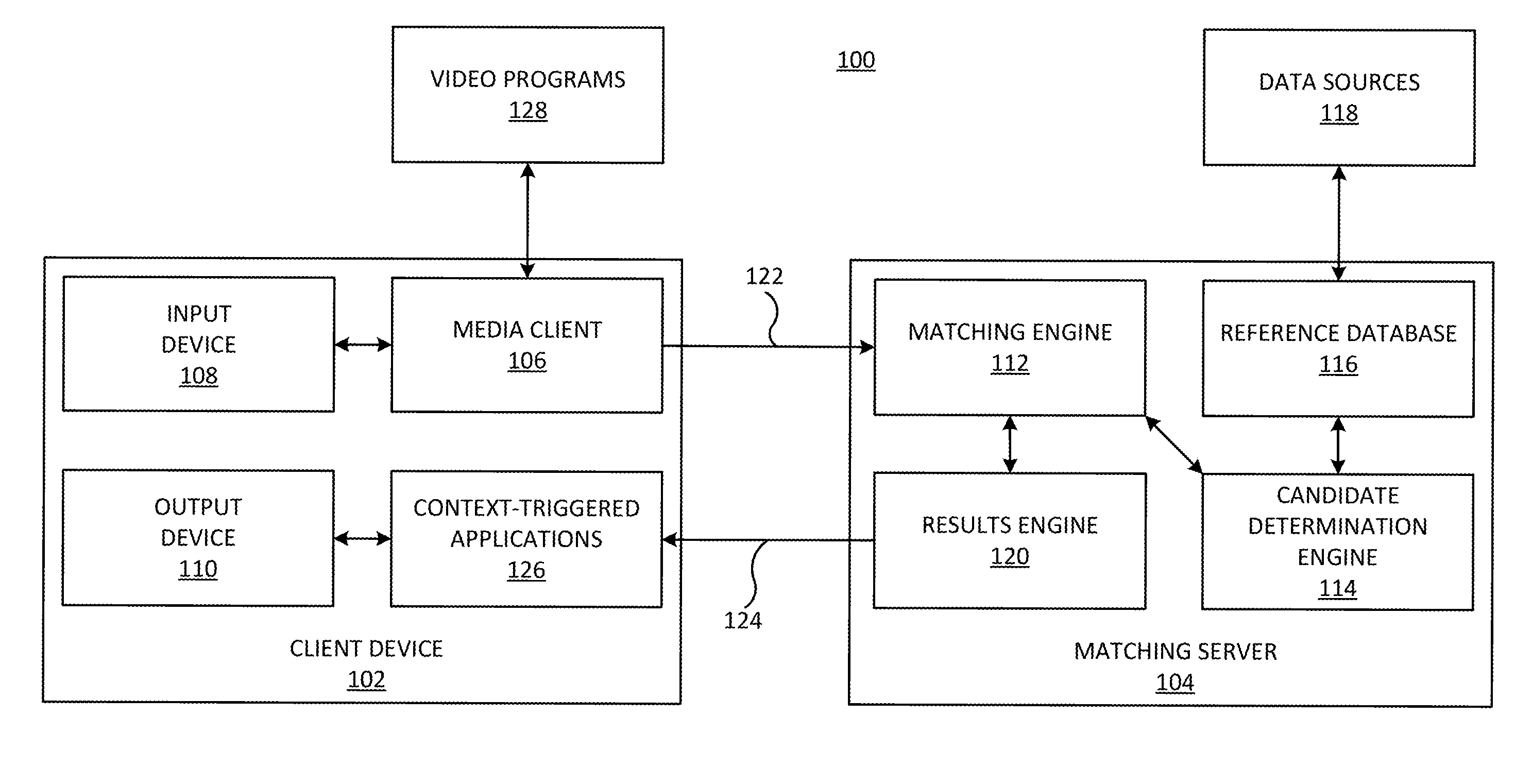
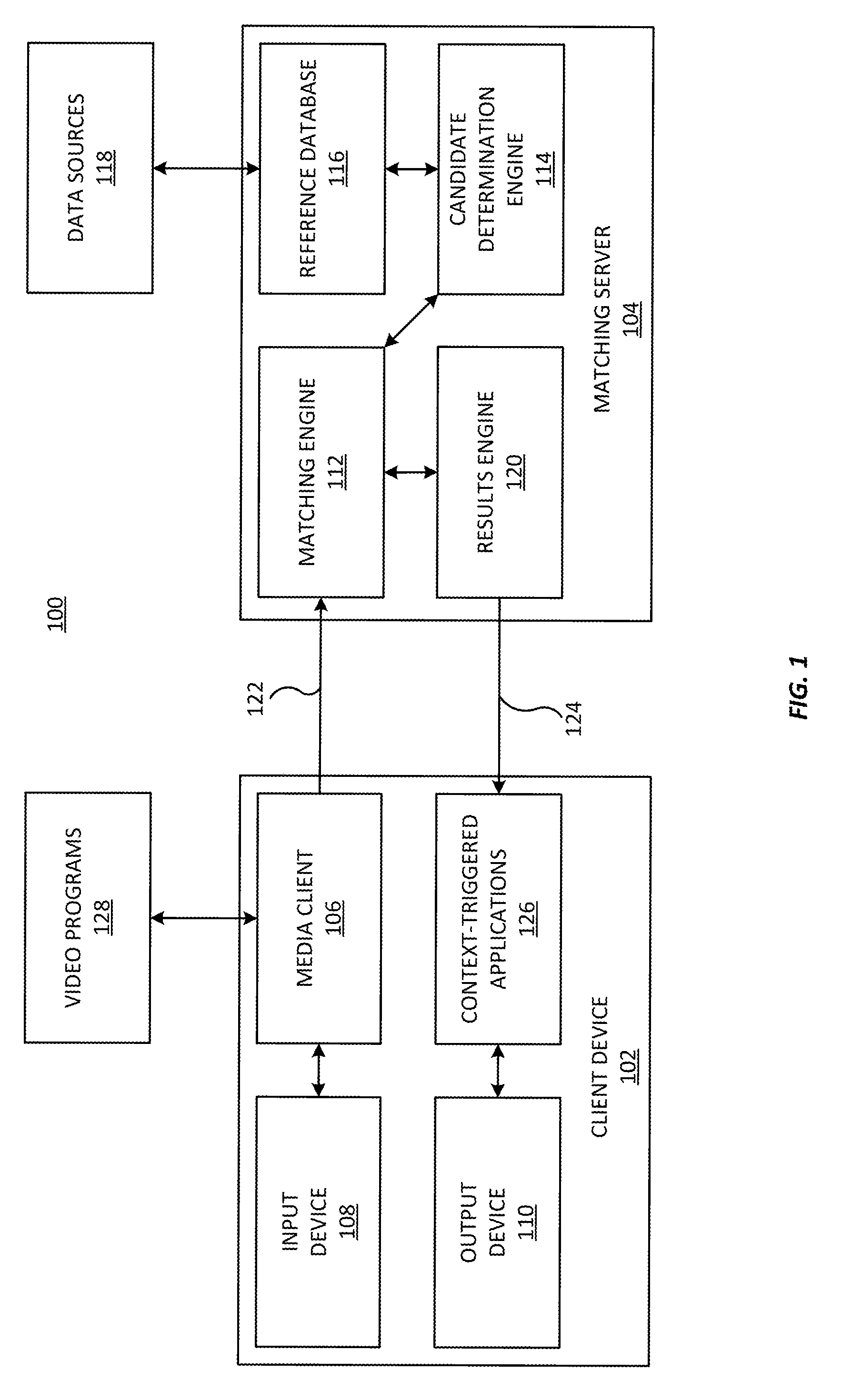
Systems and methods for identifying a media segment of audio or video content are disclosed. The video segment is identified by deriving data from media content and comparing said data to a reference database in order to identify said video segment. Embodiments of the invention improve the speed and accuracy of the media identification process by advantageously partitioning the indexes in subdivisions where high value reference information is separated from the bulk information, for example.
Owner:INSCAPE DATA INC
Method, system, and apparatus for acquiring information concerning broadcast information
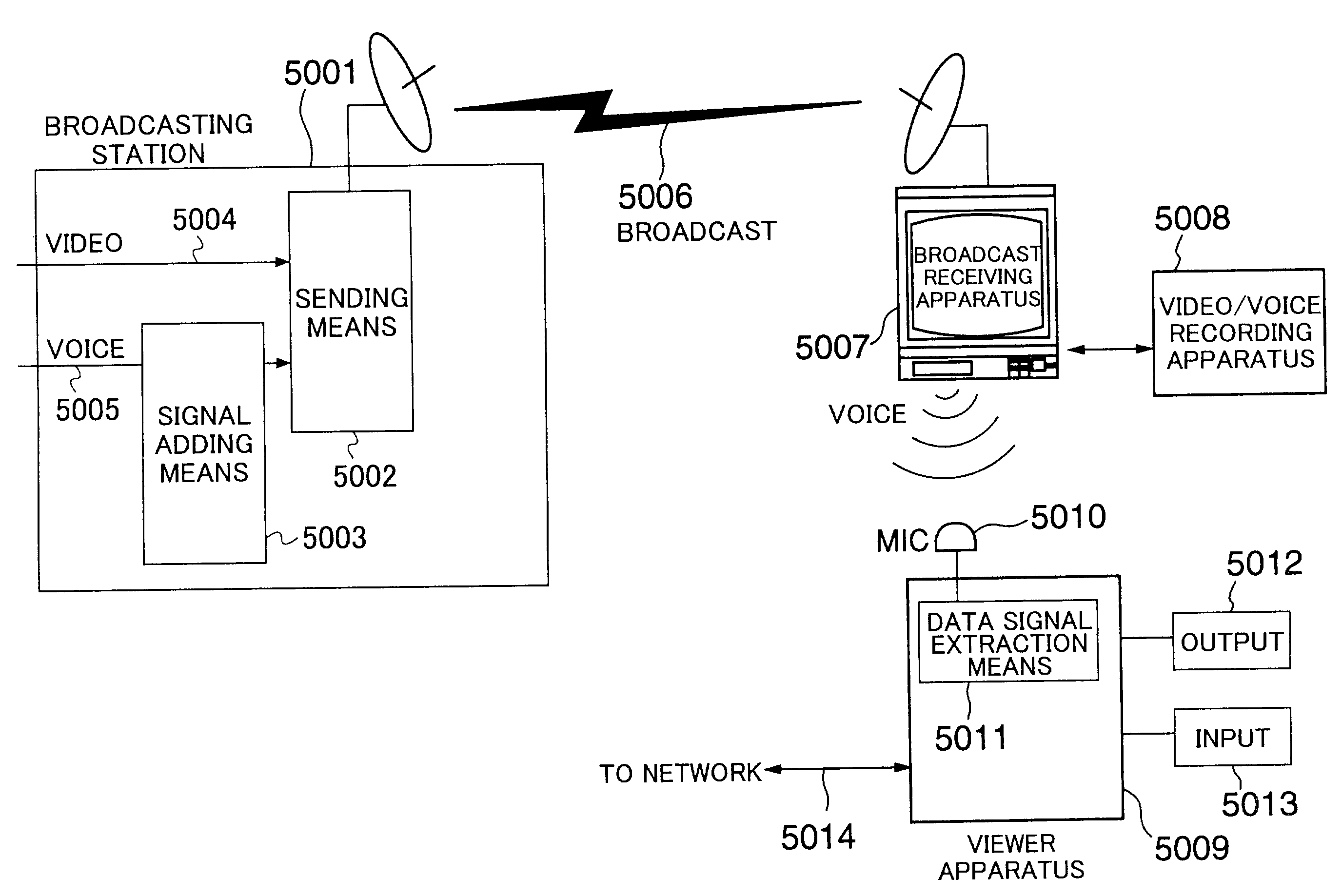
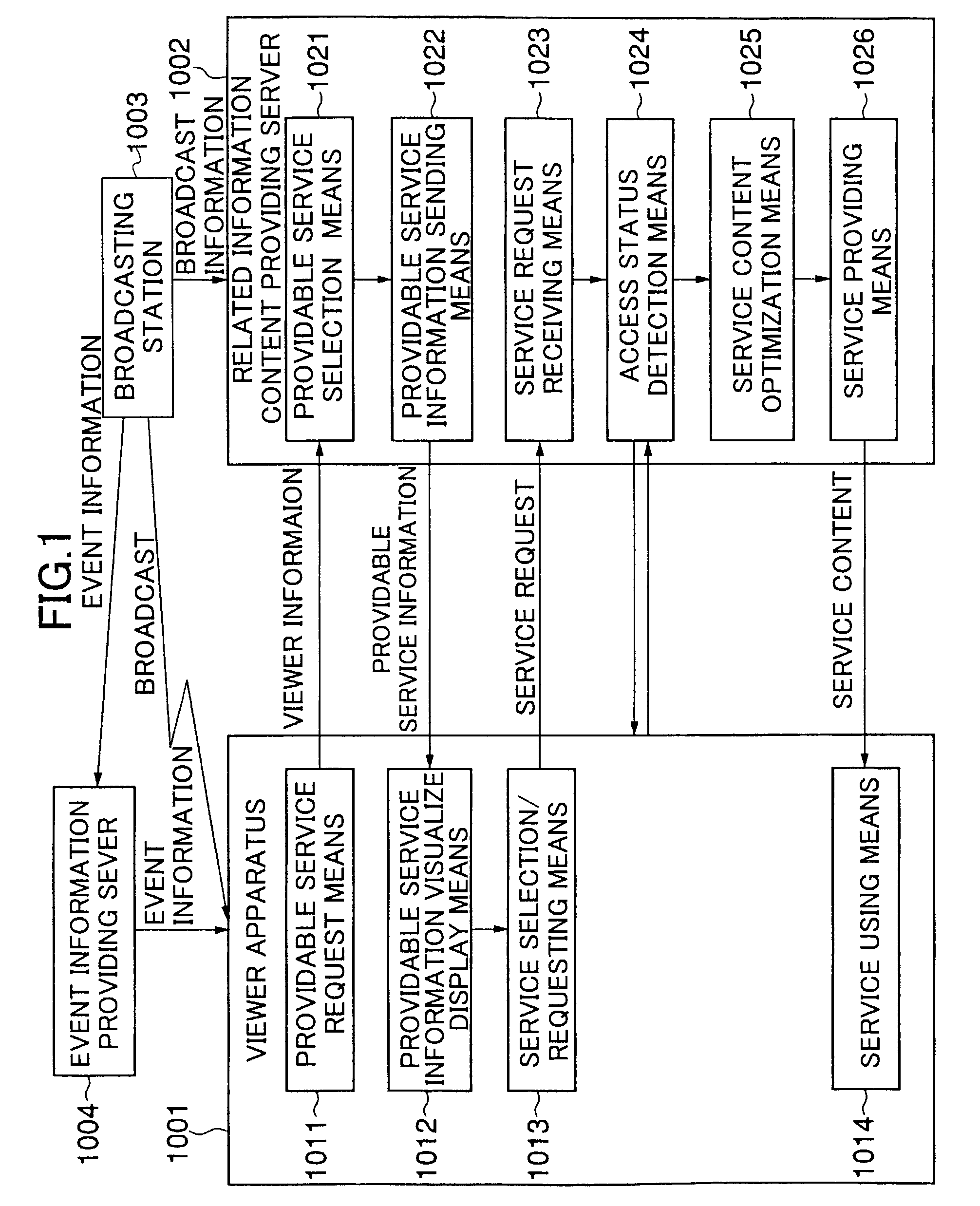
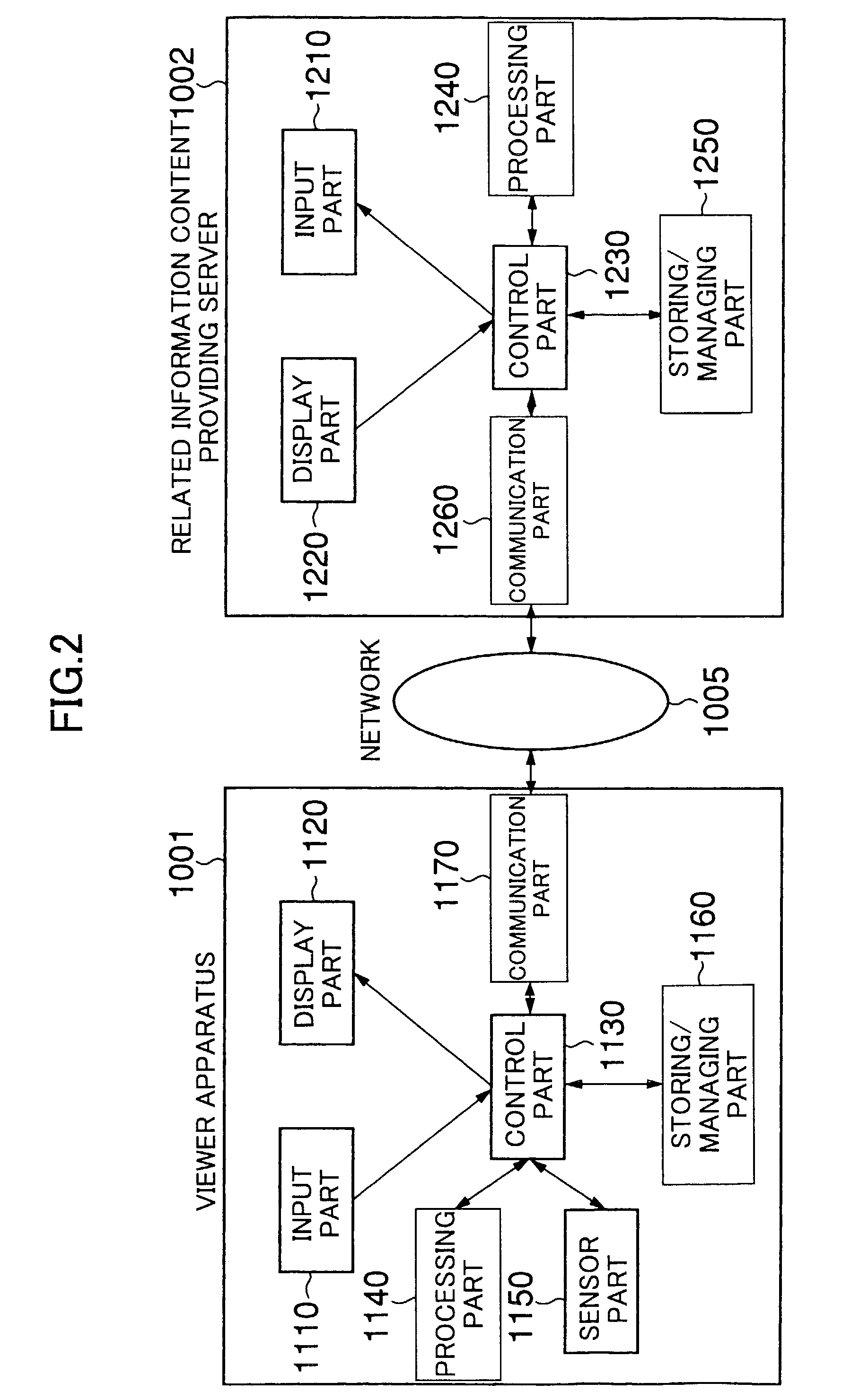
ActiveUS7712123B2Easy accessEasily referredAnalogue secracy/subscription systemsTwo-way working systemsRelevant informationNetwork connection
A technology is provided for a viewer of broadcast information or recorded information to easily obtain information related to the broadcast information or recorded information from a server, in which, in a system in which a viewer apparatus and a content providing server for providing content information related to video or voice are connected via a network, the viewer apparatus obtains related information necessary for obtaining the content information; sends the related information to the content providing server so as to request the content providing server to send the content information; and obtains the content information sent from the content providing server.
Owner:NIPPON TELEGRAPH & TELEPHONE CORP
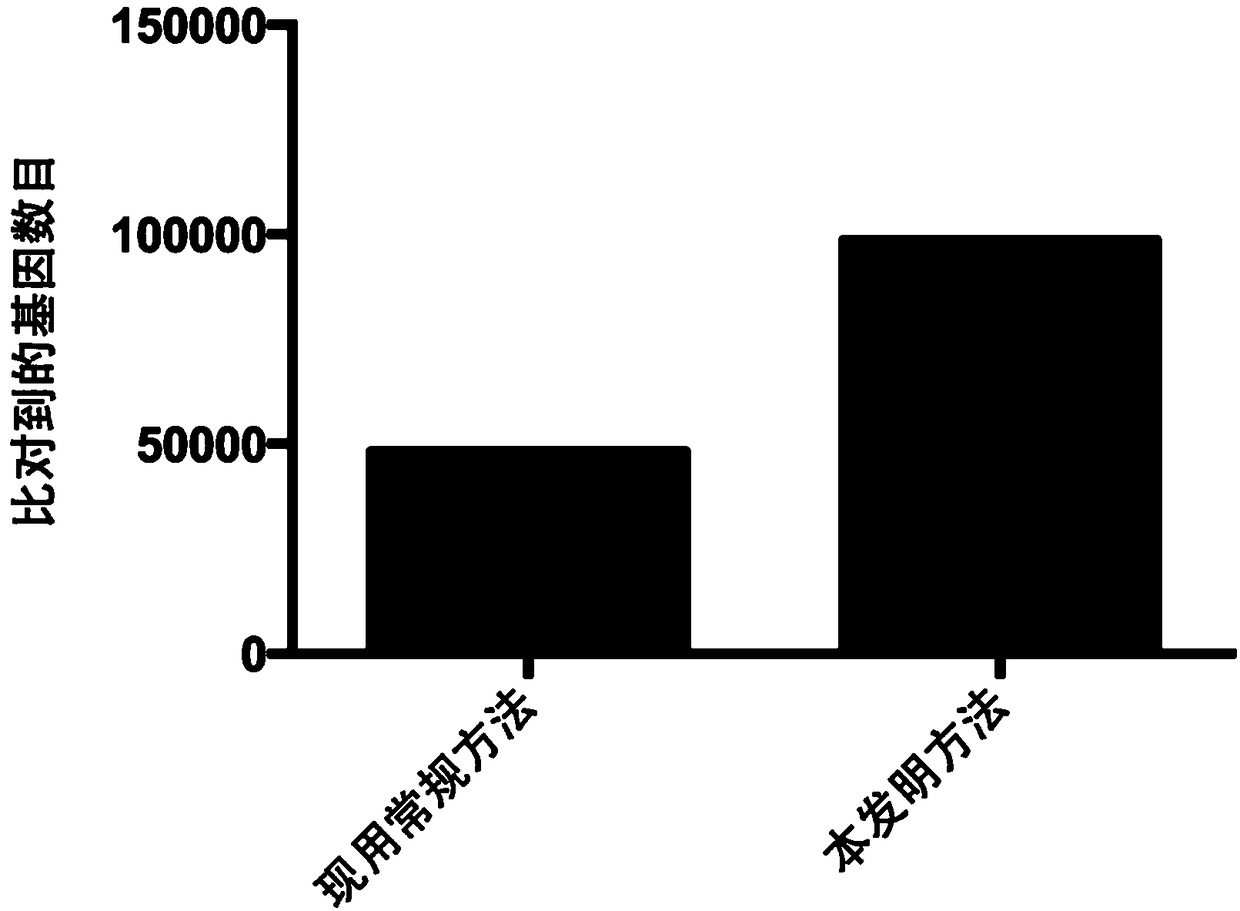
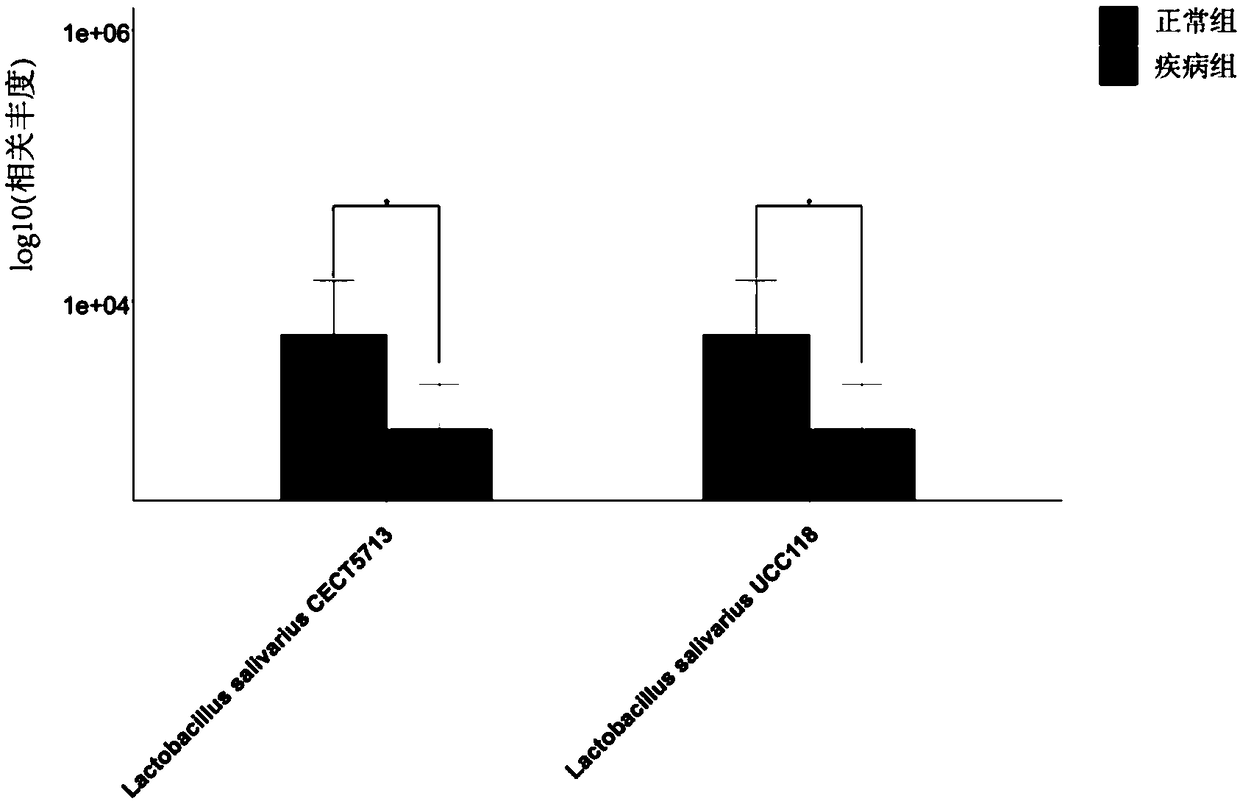
Method for analyzing microbial population function by using metagenome data
ActiveCN108804875AIncrease profitExcellent universality and comprehensiveness of informationMicrobiological testing/measurementSpecial data processing applicationsPopulationTest sample
The invention provides a method for analyzing a microbial population function by using metagenome data. The method comprises the following steps: collecting all known microbial species, gene and function information, and integrating the information as a reference database; sequencing the metagenome of the to-be-tested microbial specifies, controlling the sequencing data quality, computing speciesabundance and gene abundance, analyzing the composition difference of microbes among different samples and gene level difference; annotating the gene function, clustering the genes with the same function to obtain a function module, performing adduction computation on related abundance of all non-redundancy genes in various function modules to obtain abundance values of all function modules, performing difference comparison analysis or overall evaluation on the functions of the to-be-tested sample microbes. Through the method provided by the invention, the step of respectively comparing the splicing data, the assembling data, the predicting data and the sequencing data with the single function database is saved, the time is saved, the utilization efficiency of the sequencing data is improved, and the method can be used for analyzing high-throughput microbe whole-genome sequencing data and screening the function microbe.
Owner:BEIJING INST OF GENOMICS CHINESE ACAD OF SCI CHINA NAT CENT FOR BIOINFORMATION
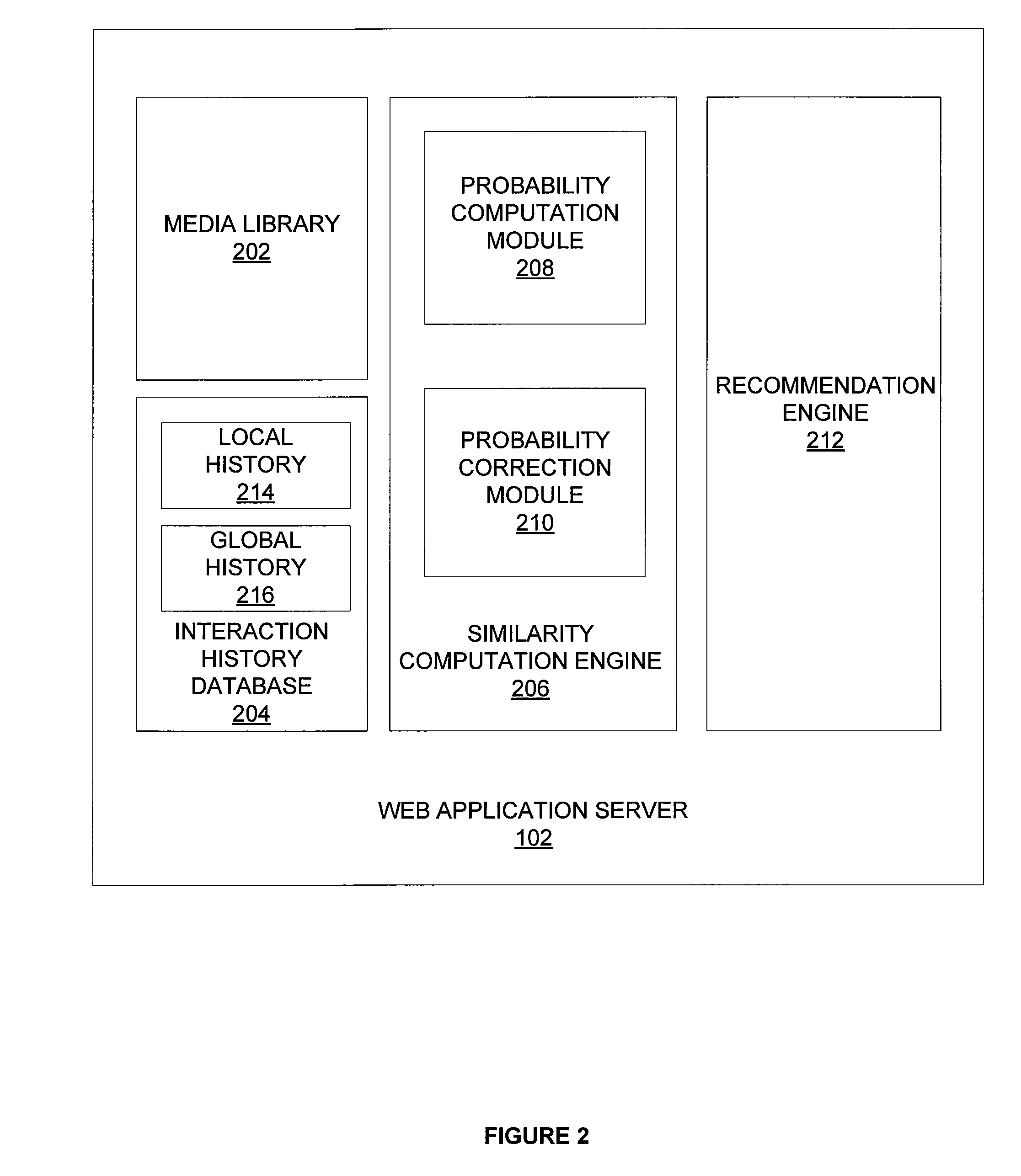
Identifying similar items based on global interaction history
ActiveUS20130013458A1Precise processAccurate contentMarket data gatheringTotal countInteraction history
One embodiment sets forth technique for computing a similarity score between two digital items is computed based on interaction histories associated with global users and interaction histories associated with local users. Global counts indicating the number of interactions associated with each unique pair of digital items are weighted based on a mixing rate. The weighted global counts are then combined with local counts to compute total counts. An effective interaction probability indicating the likelihood of a user interacting with one digital item in the pair of digital items after interacting with the other digital item in the pair is computed based on the total counts. The effective interaction probability is then corrected for noise, resulting in a similarity score indicating the similarity between the pair of digital items.
Owner:NETFLIX
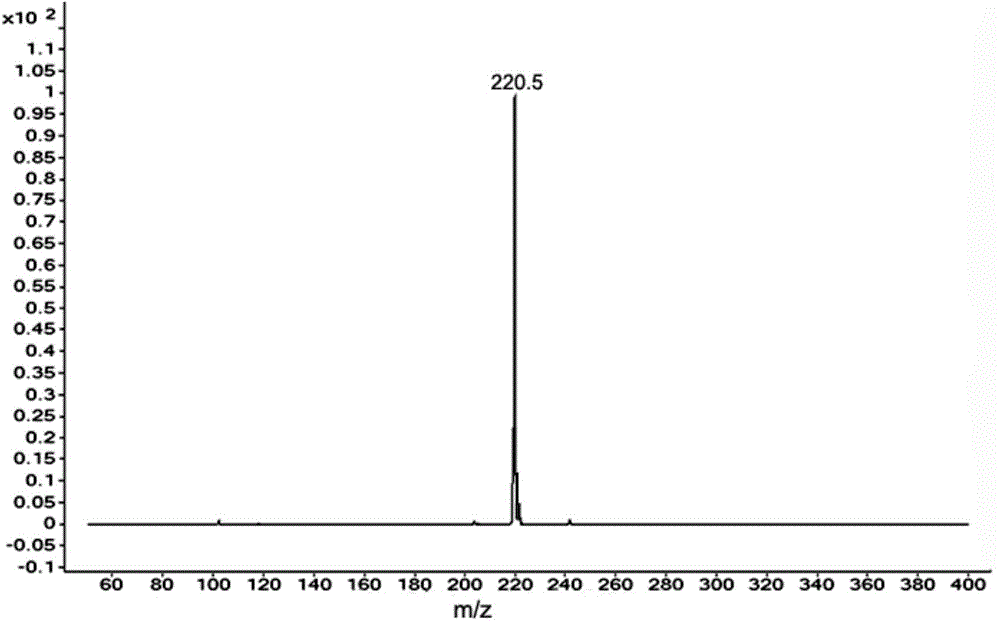
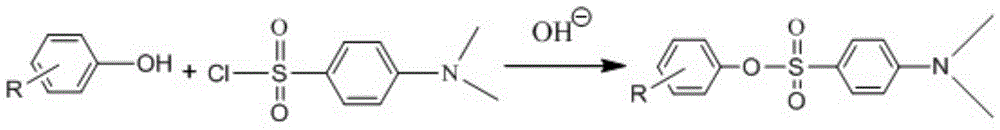
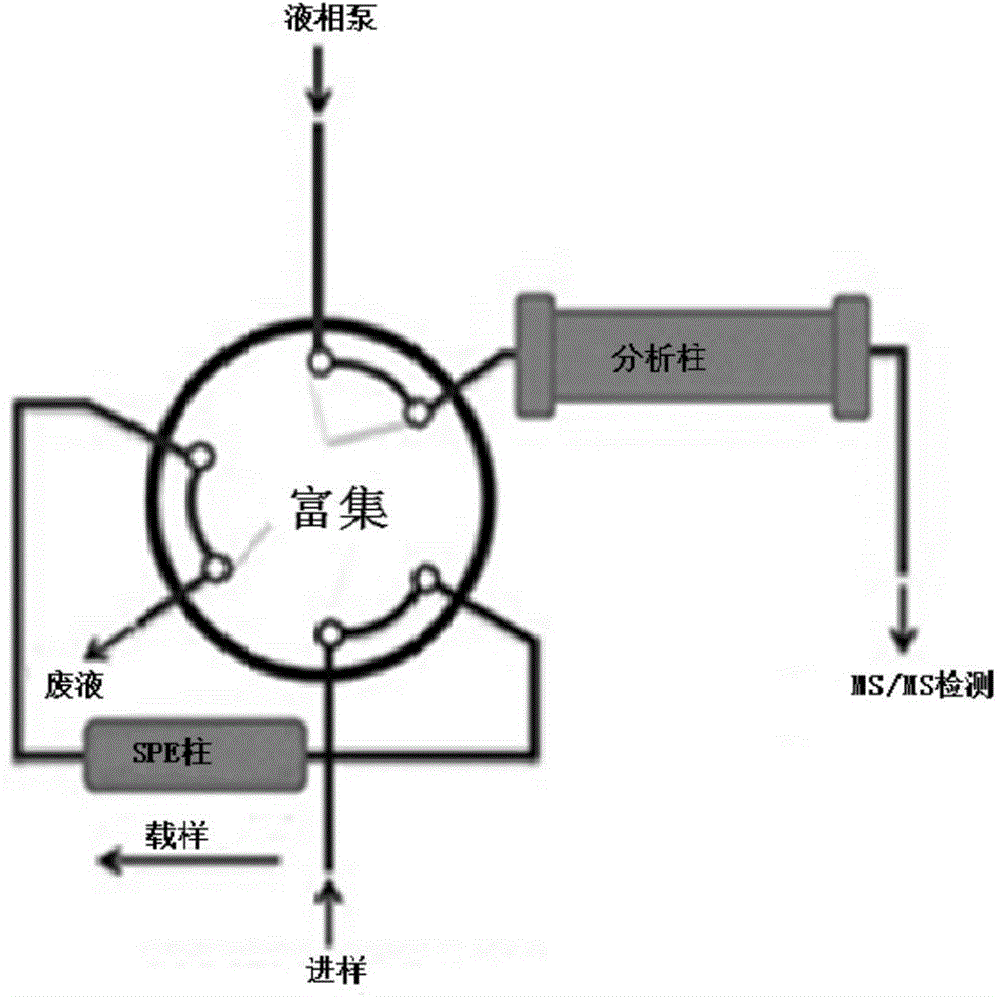
Pretreatment and detection method for biogen amine neurotransmitter and detection kit
ActiveCN104678028AHigh sensitivityAvoid instabilityComponent separationMetabolitePretreatment method
The invention belongs to the technical field of detection and relates to a pretreatment method for biogen amine neurotransmitter and / or metabolites thereof. The method comprises the following steps of: performing derivatization on a sample containing the biogen amine neurotransmitter and / or the metabolites thereof, and carrying out online solid-phase extraction on derivatization products. On the basis, the invention also relates to a detection method for the biogen amine neurotransmitter and / or the metabolites thereof, a detection kit and application thereof. The pretreatment method has the advantages that impurities in the derivatization products can be eliminated, and after being enriched, the derivatization products with proper concentration are transferred into the following detection steps. The detection method has the advantages that the content of the biogen amine neurotransmitter and / or the metabolites thereof in a biological sample can be fast and accurately measured, and the detection sensitivity is high.
Owner:INST OF PHARMACOLOGY & TOXICOLOGY ACAD OF MILITARY MEDICAL SCI P L A
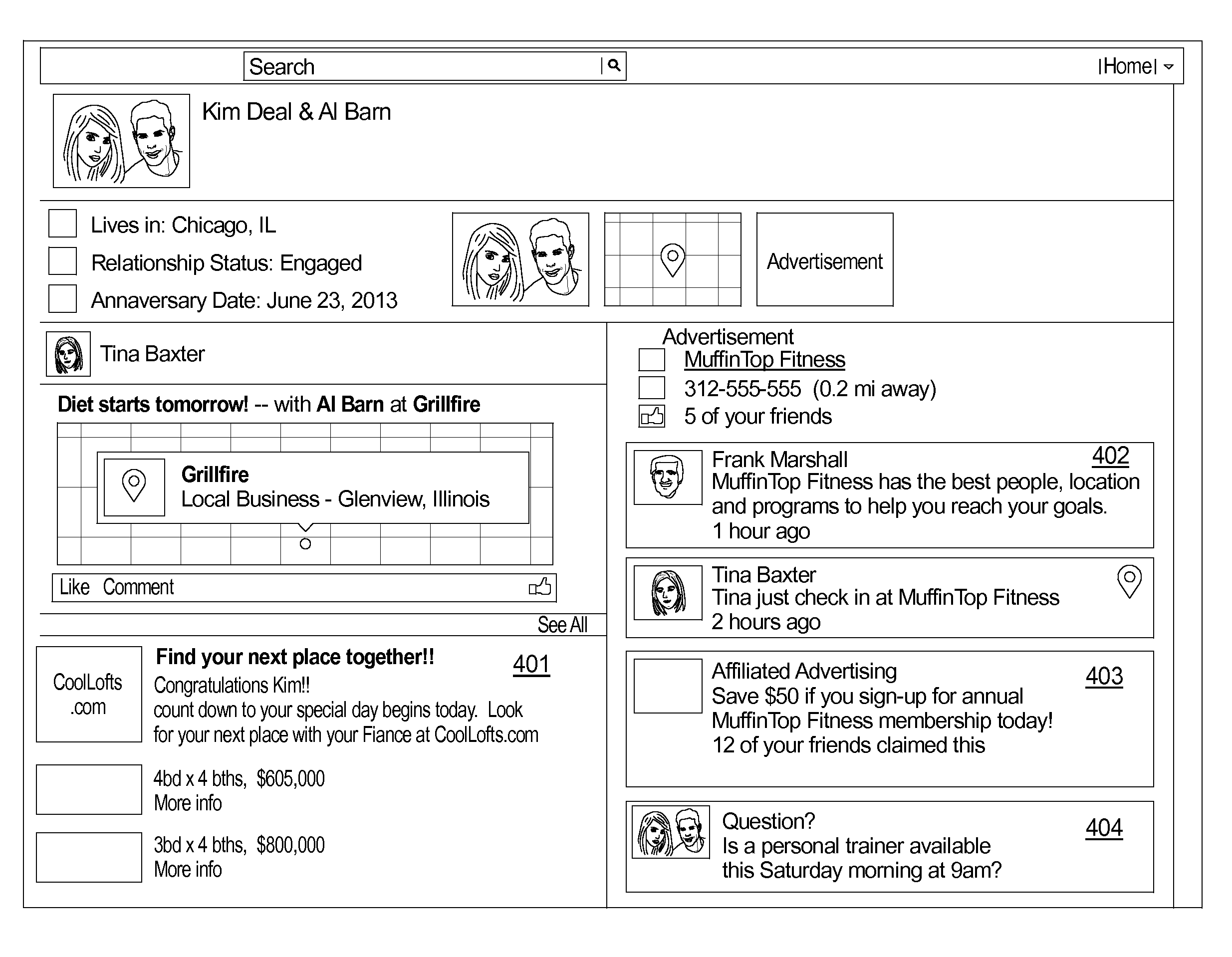
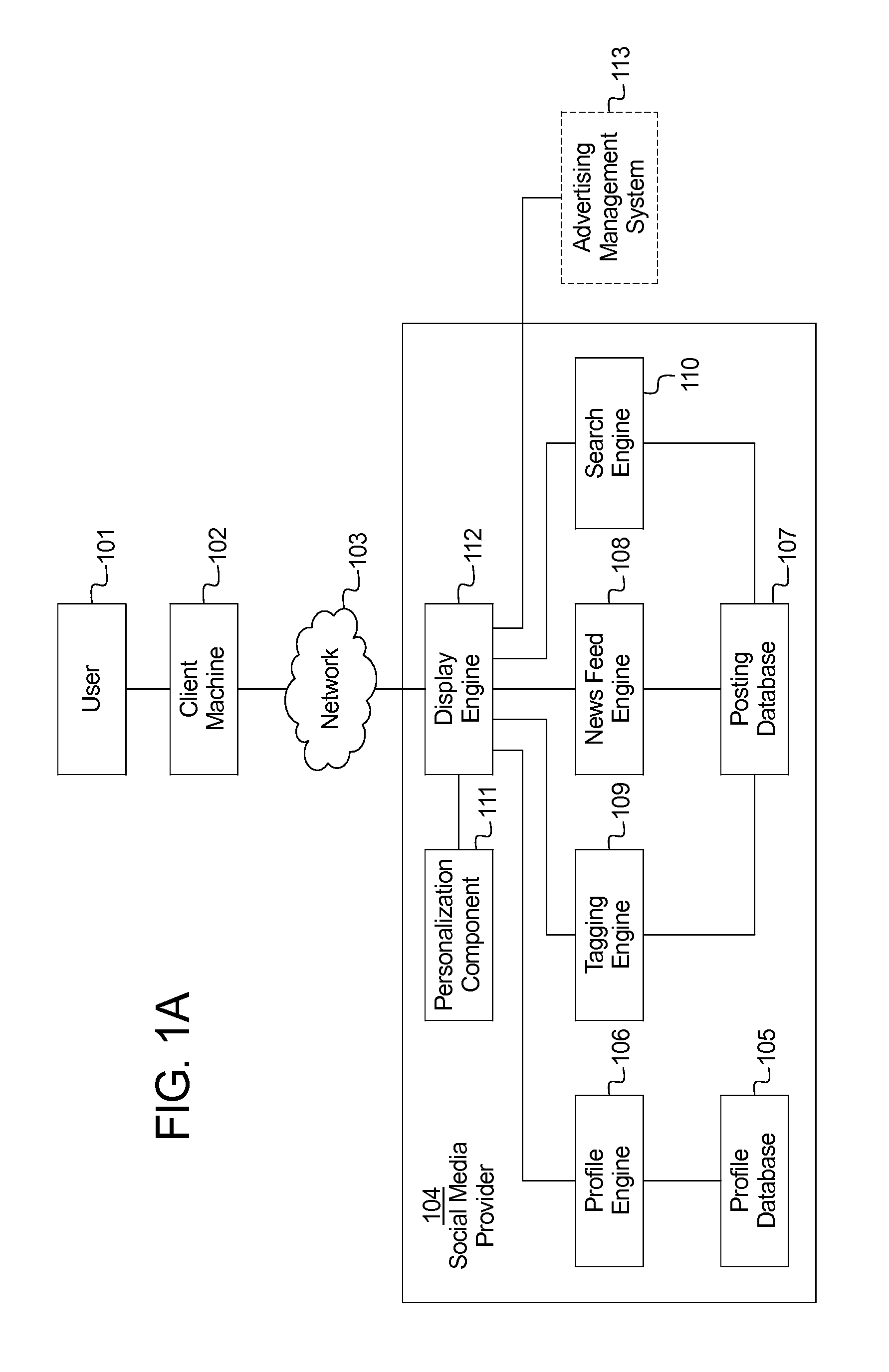
Systems and methods for organizing and displaying social media content
InactiveUS20140040029A1Process information easilyEasy to processTransmissionMarketingClient-sideMedia content
A system for dynamically presenting and organizing social media by subject matter includes: a posting database including a plurality of posts, wherein each post includes post content and one or more post tags; and a display engine posting the plurality of posts to a client machine, wherein the plurality of posts are posted in response to receiving a post tag selection from the client machine.
Owner:OMEGA INTELLIGENCE
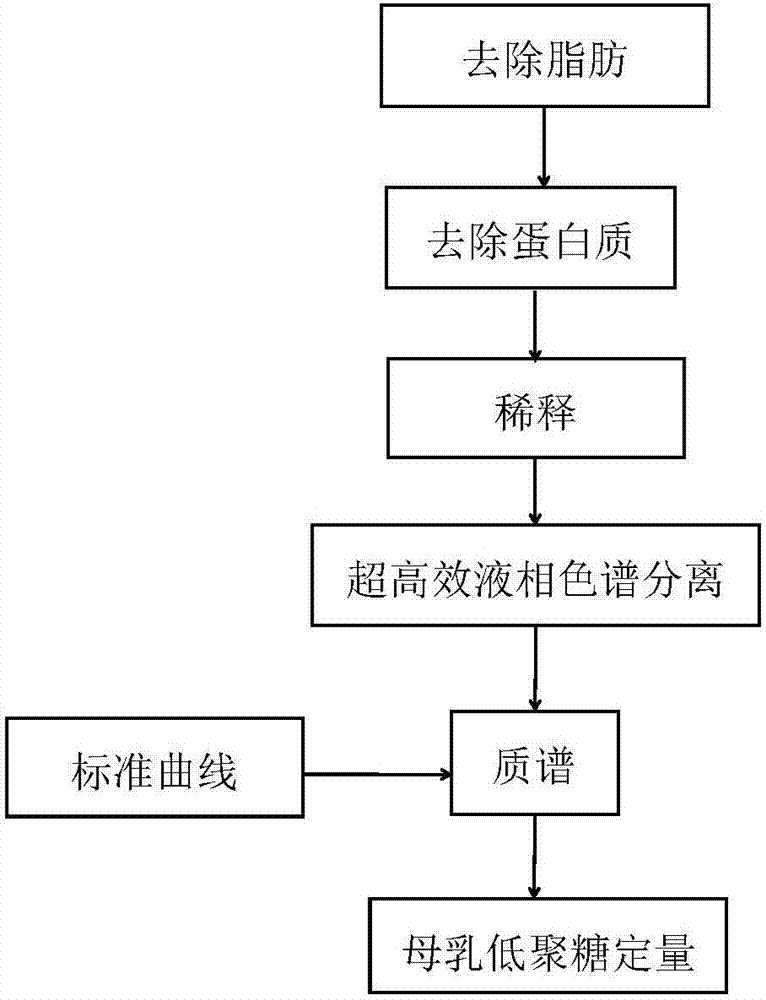
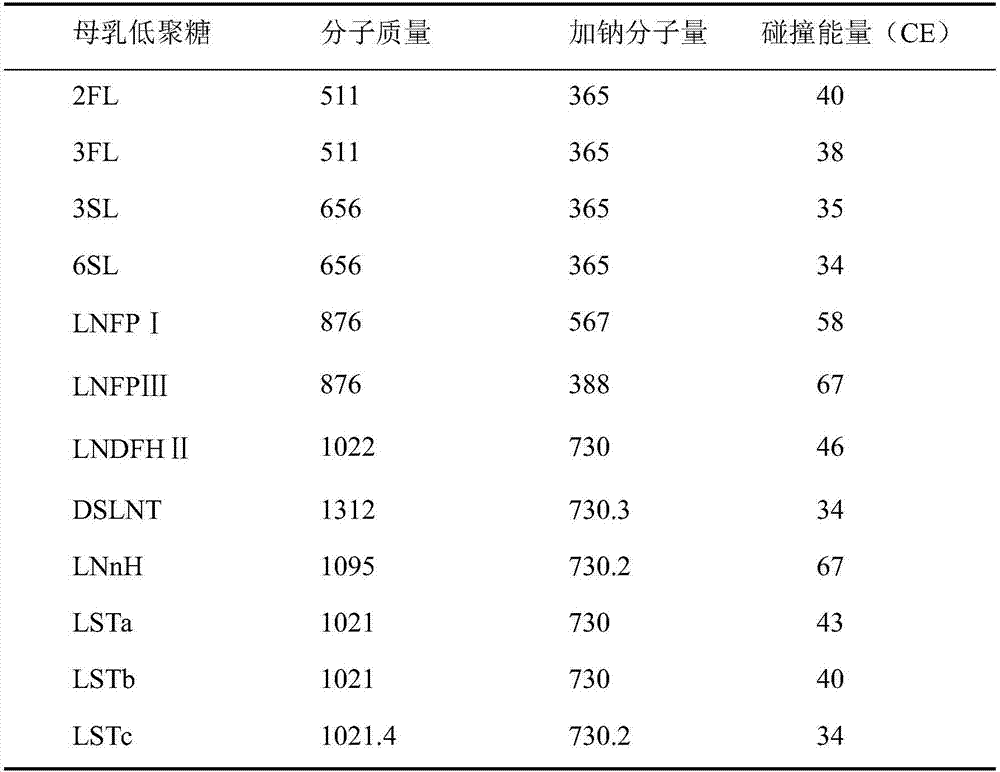
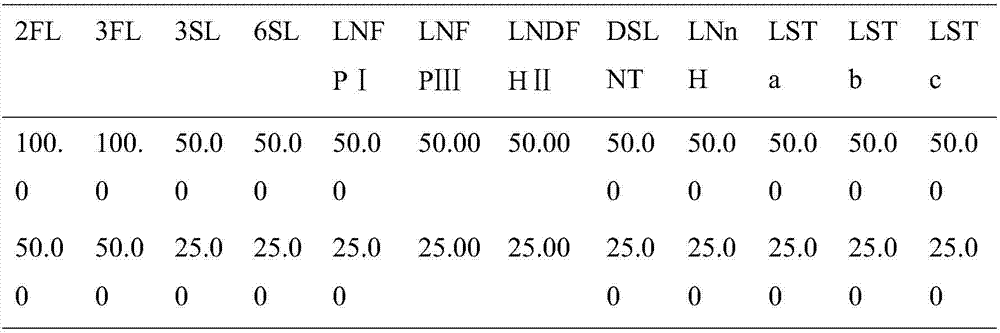
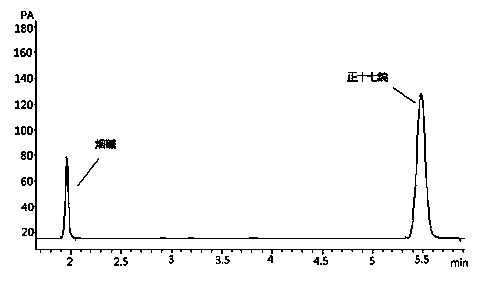
Quick qualitative and quantitative method for oligosaccharide in breast milk
ActiveCN107192771AAccurate detectionQuick checkComponent separationGradient elutionColumn temperature
The invention discloses a quick qualitative and quantitative method for oligosaccharide in breast milk. The quick qualitative and quantitative method mainly includes steps of 1, pretreating samples, to be more specific, removing fat and proteins from 150-250 micro-l of breast milk to obtain ultimate supernatant, adding ultra-pure water into the supernatant and diluting the supernatant to obtain the loaded samples; 2, establishing standard curves for standard substances by the aid of ultrahigh-performance liquid chromatography and mass spectrometry; 3, separating different components of the oligosaccharide in the breast milk in the loaded samples by the aid of ultrahigh-performance liquid mass spectrometry and carrying out quantitative analysis by the aid of mass spectrometry combined with the standard curves so as to obtain the content of the oligosaccharide in the breast milk. The ultrahigh-performance liquid chromatography is implemented by the aid of amino chromatographic columns with the sizes of 2.1*100 mm and 1.7 micrometers, 8-10 mmol / L of ammonium formate solution (A) and acetonitrile (B), and the ammonium formate solution (A) and the acetonitrile (B) are used as mobile phases; gradient elution programs are carried out by the aid of 95%-75% of B for 0-10 min or are carried out by the aid of 75% of B for 10-15 min or are carried out by the aid of 75%-65% of B for 15-20 min or are carried out by the aid of 65%-10% of B for 20-21 min or are carried out by the aid of 10% of B for 21-24 min or are carried out by the aid of 10%-95% of B for 24-25 min or are carried out by the aid of 95% of B for 25-35 min; the flow rates are 0.3 mL / min, and the column temperatures are 40-60 DEG C. The quick qualitative and quantitative method has the advantage that 12 types of oligosaccharide in the breast milk can be quickly detected by the aid of the quick qualitative and quantitative method and can be quantified by the aid of the quick qualitative and quantitative method.
Owner:INST OF AGRO FOOD SCI & TECH CHINESE ACADEMY OF AGRI SCI
Electroslag remelting process of titanium-containing alloy steel
The invention discloses an electroslag remelting process for titanium alloy steel, which comprises the following steps of: (1) preparing consumable electrodes; (2) preparing electroslag; (3) forming slag through electroslag; (4) remelting electroslag. The step (3) of slag formation comprises the following procedures of: firstly, adding one third of total slag material into a crystallizer to form slag, mixing a right amount of mixture including Al powder and FeTi powder into the rest residue material, and adding the rest residue material with the mixture into the crystallizer to form slag. Theelectroslag remelting process for titanium alloy steel does not need special slag purification process, the investment to slag purification equipment is reduced, the technological flow is shortened, and the production cost is greatly lowered. By adding the mixture of Al powder, FeTi powder and CaF2 powder into a slag pool in the process, oxygen generated by unstable oxides in the molten slag can be prevented from entering the slap pool and a molten pool to oxidize elements in steel, and the massive uncertain oxidation burning of Ti element in steel is avoided. Therefore, the content of Ti element in the titanium alloy steel obtained by the process can accurately reach the standard requirement, and the quality of the titanium alloy steel is ensured.
Owner:CHONGQING IRON & STEEL (GRP) CO LTD
Website data statistics system and method
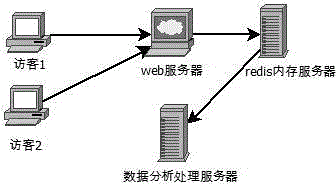
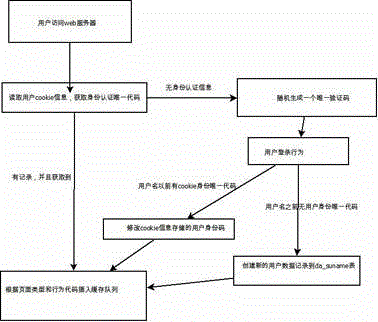
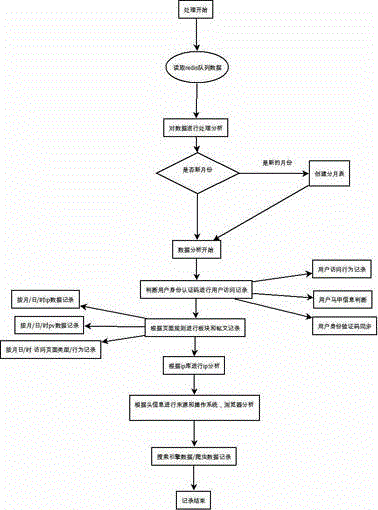
ActiveCN104869009AReduce consumptionEasy accessData switching networksHigh level techniquesEngineeringServer-side
The invention relates to a data statistics method, and particularly relates to a website data statistics system and a method, so as to carry out statistics and analysis on basic website traffic data, master a website traffic trend and fully perceive behavior habits of visitors. The method comprises steps: a data statistics code is added to a web page and a data transmission code is set for judging, building, recording and transmitting basic information of user accessing website; a background processing program is deployed for carrying out program analysis, transmitting data and carrying out packet processing on the data; according to needs, multiple table files are divided for recording the data; and the data are displayed visually, and through using a table component and a server end structure, the data content are displayed. Consumption of hardware is little, the production environment is separated for processing, and expandability is good.
Owner:QINGDAO NEWS NETWORK COMM CO LTD
Method and device for providing application information in mobile terminal device
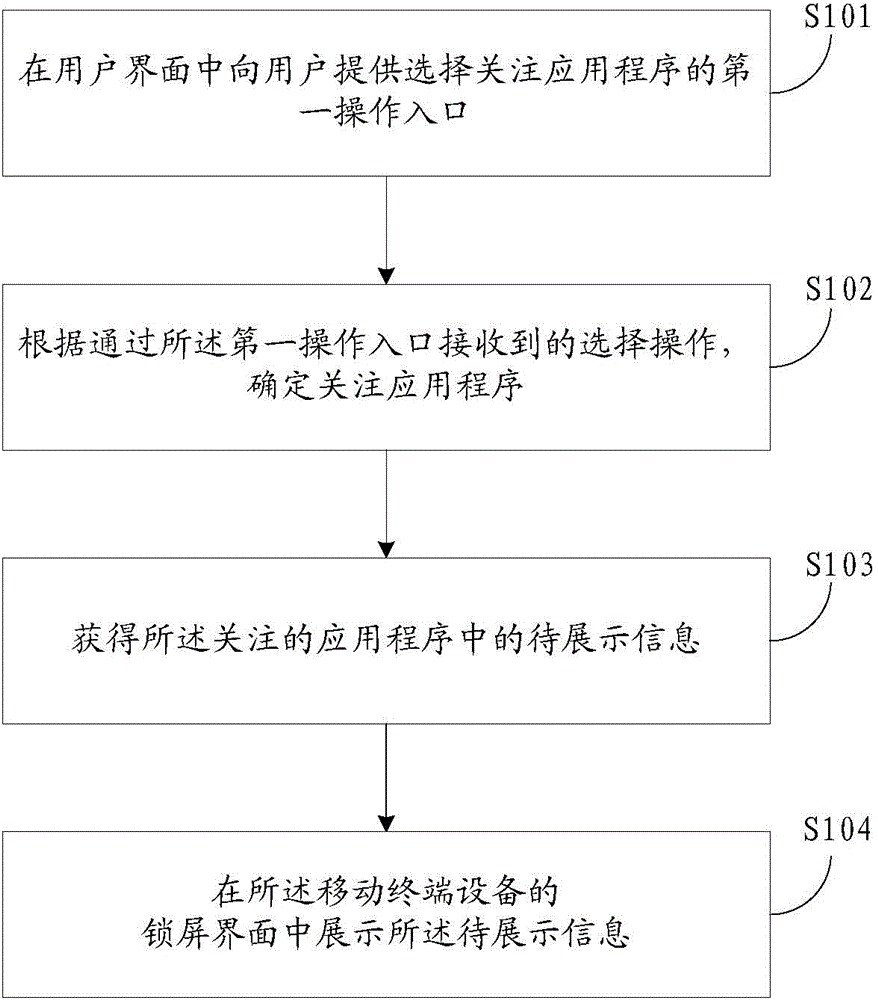
ActiveCN104639721AEasy to operateImprove experienceService provisioningSubstation equipmentTerminal equipmentLock screen
A method and an apparatus for providing application program information in a mobile terminal device are provided. The method includes: providing a first operation entrance for selecting a concerned application program to a user in a user interface; determining the concerned application program based on a selection operation received via the first operation entrance; obtaining information that is pending for display in the concerned application program; and displaying the information that is pending for display in a lock screen interface of the mobile terminal device. Using the present disclosure, the path of acquiring information for a user can be shortened.
Owner:ALIBABA GRP HLDG LTD
Collaborative media presentation service with usage rights enforcement
InactiveUS20150020153A1Increase valueAccurate contentDigital data protectionTransmissionDigital rights managementMedia server
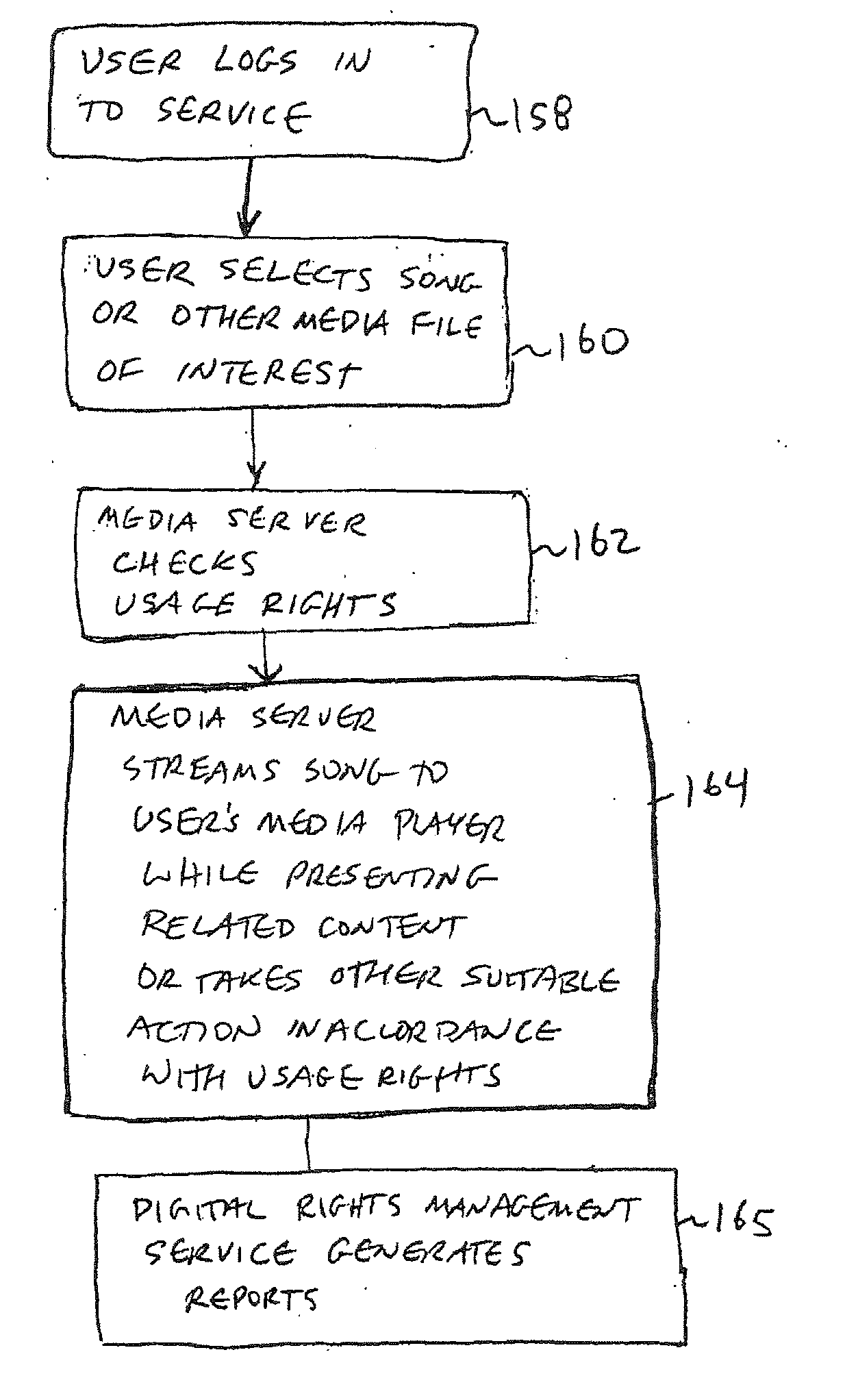
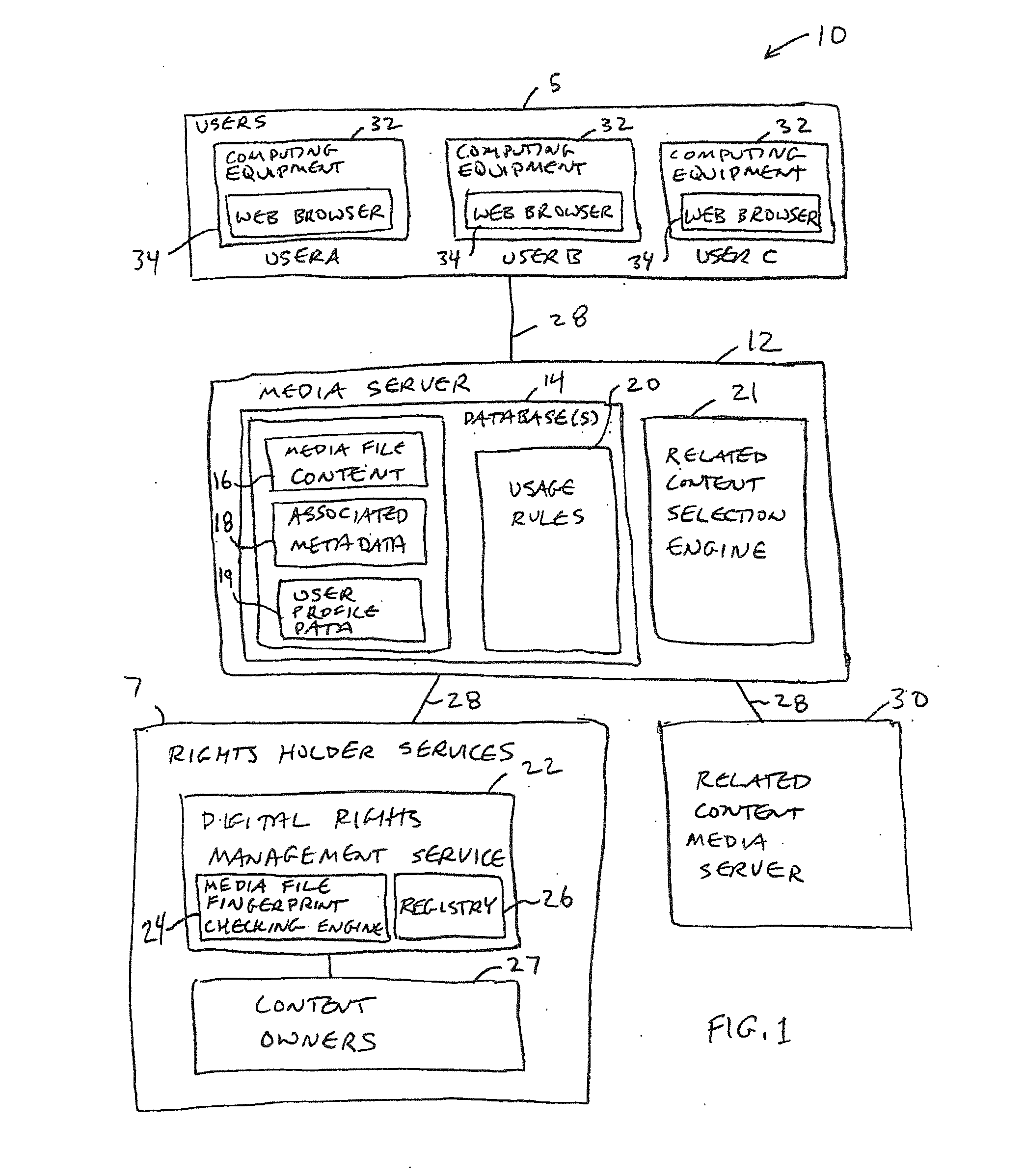
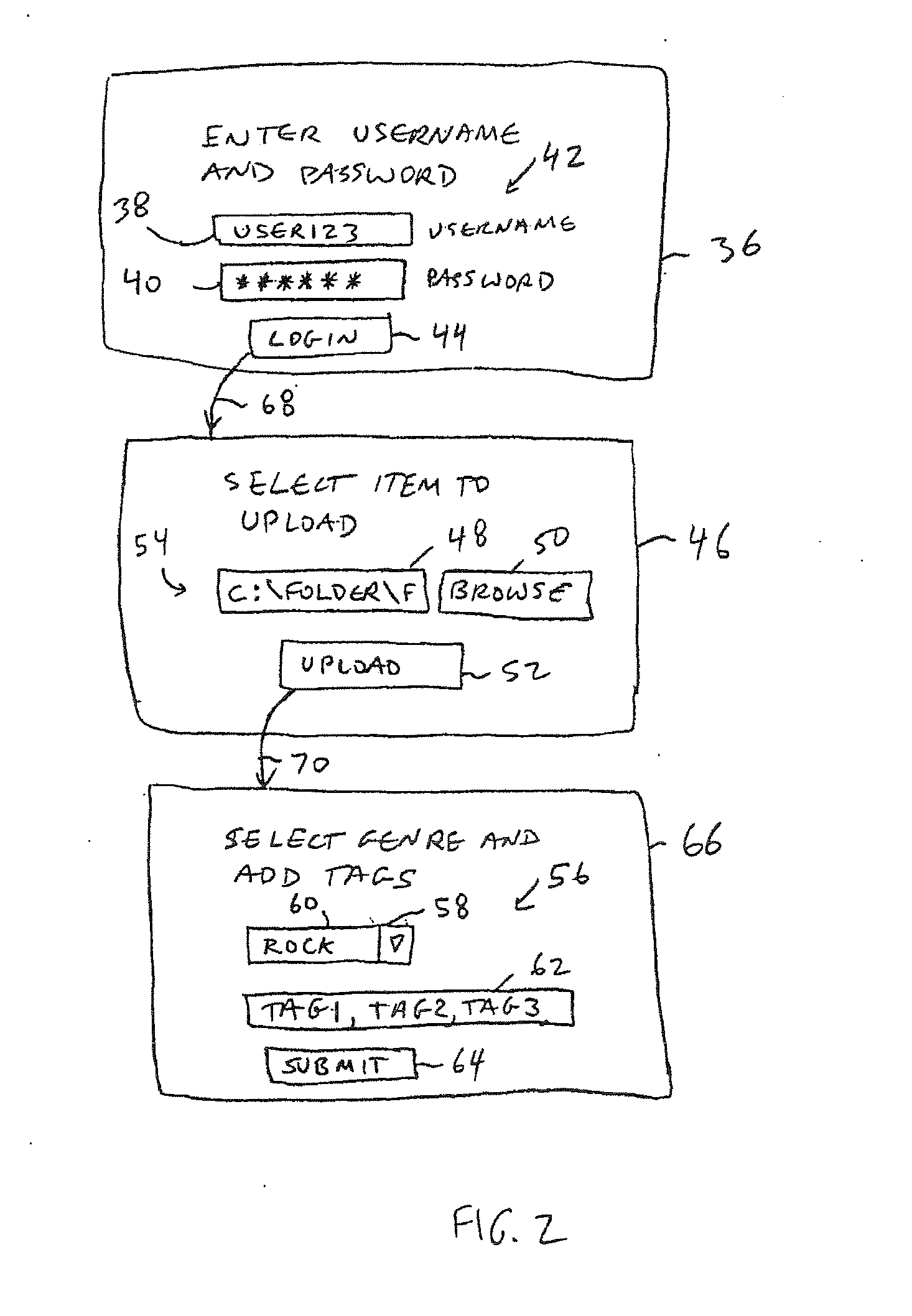
An online collaborative media presentation service is provided. Users of the online service can create highly customized profiles that contain personal profile information and information on media files and other topics of interest. Users can upload media files to a media server associated with the online service without preapproval from content owners. The rights of content owners are preserved by using a digital rights management service to identify uploaded media files. The media server submits uploaded media files to the digital rights management service to determine whether streaming of the media files is permissible. If streaming is not permitted, the media server can block an uploaded file. If streaming is permitted, the media server can make the media file available for streaming. When streaming media to users, the media server displays targeted advertisements and other related content to users. The related content adds value for the online service.
Owner:MYSPACE MUSIC
Method for detecting content of nicotine in smoke of electronic cigarette
ActiveCN104062385ANicotine content fastRapid determinationComponent separationBiochemical engineeringVapor phase chromatography
The invention discloses a method for detecting the content of nicotine in smoke of an electronic cigarette. The detection method comprises the following steps: (1) preparing an extracting agent; (2) preparing a standard series solution of the nicotine; (3) connecting a Cambridge filter and an absorption bottle with a smoking machine in series and capturing the smoke of the electronic cigarette; putting the Cambridge filter for capturing the smoke and absorption liquid into a tapered bottle and carrying out vibration extraction; (4) moving away an extracting liquid filtering membrane and analyzing filter liquid by using a gas chromatographic method; and (5) drawing a standard curve and calculating the content of the nicotine in the smoke of the electronic cigarette. The method disclosed by the invention has the advantages of high sensitivity, good selectivity, simplicity in operation and high repeatability, and can realize the accurate quantitative analysis of the nicotine in the electronic cigarette.
Owner:CHINA TOBACCO YUNNAN IND
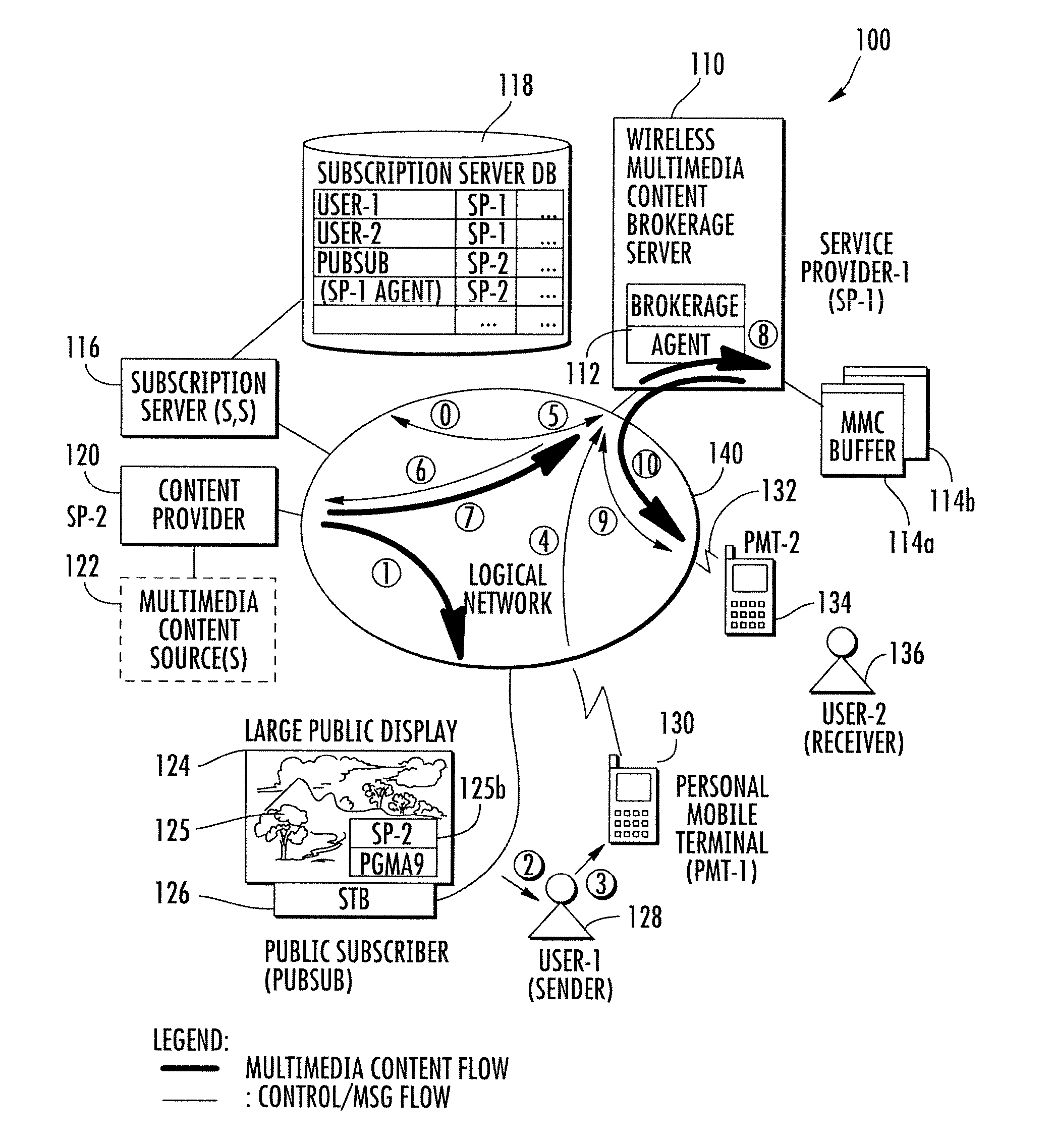
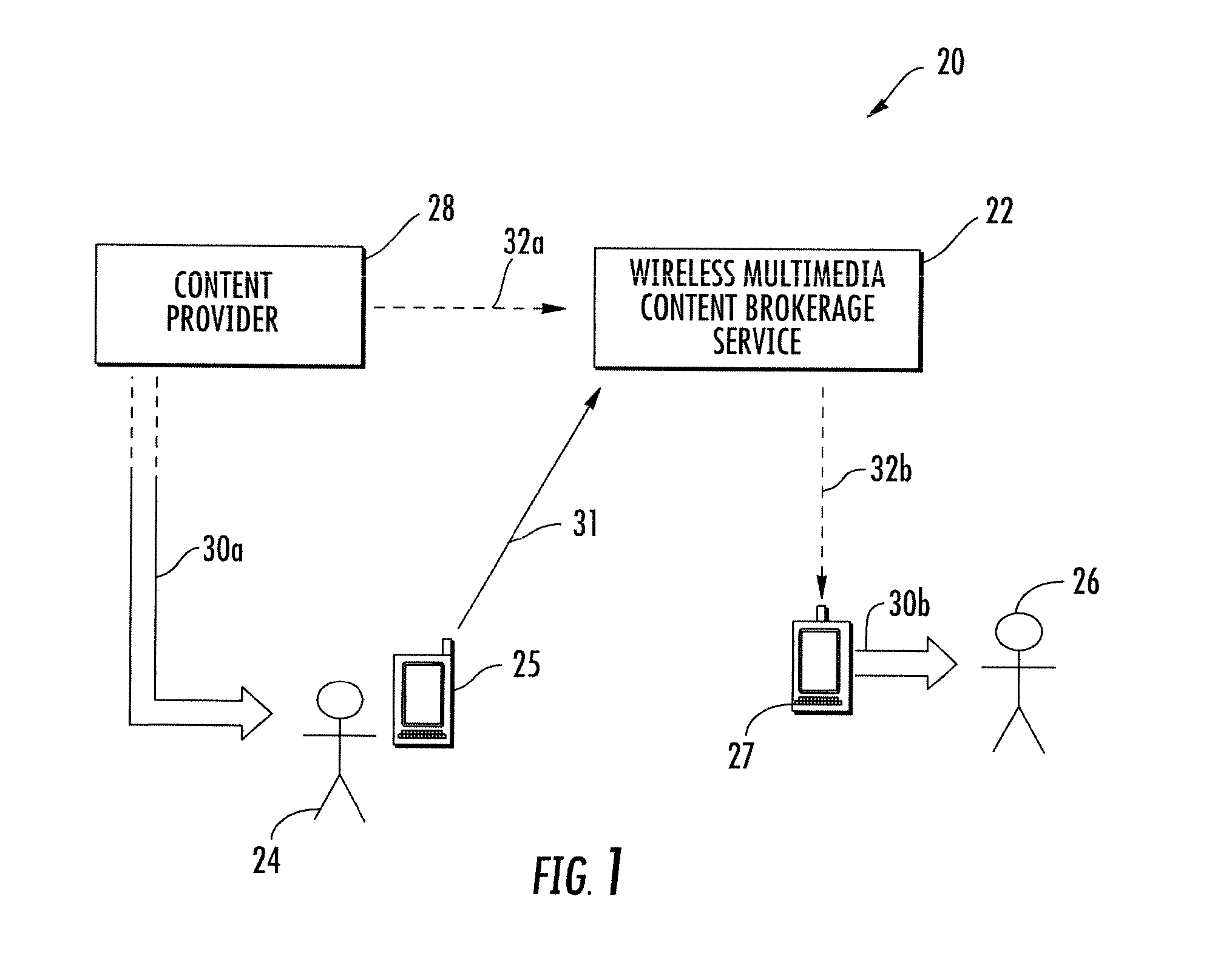
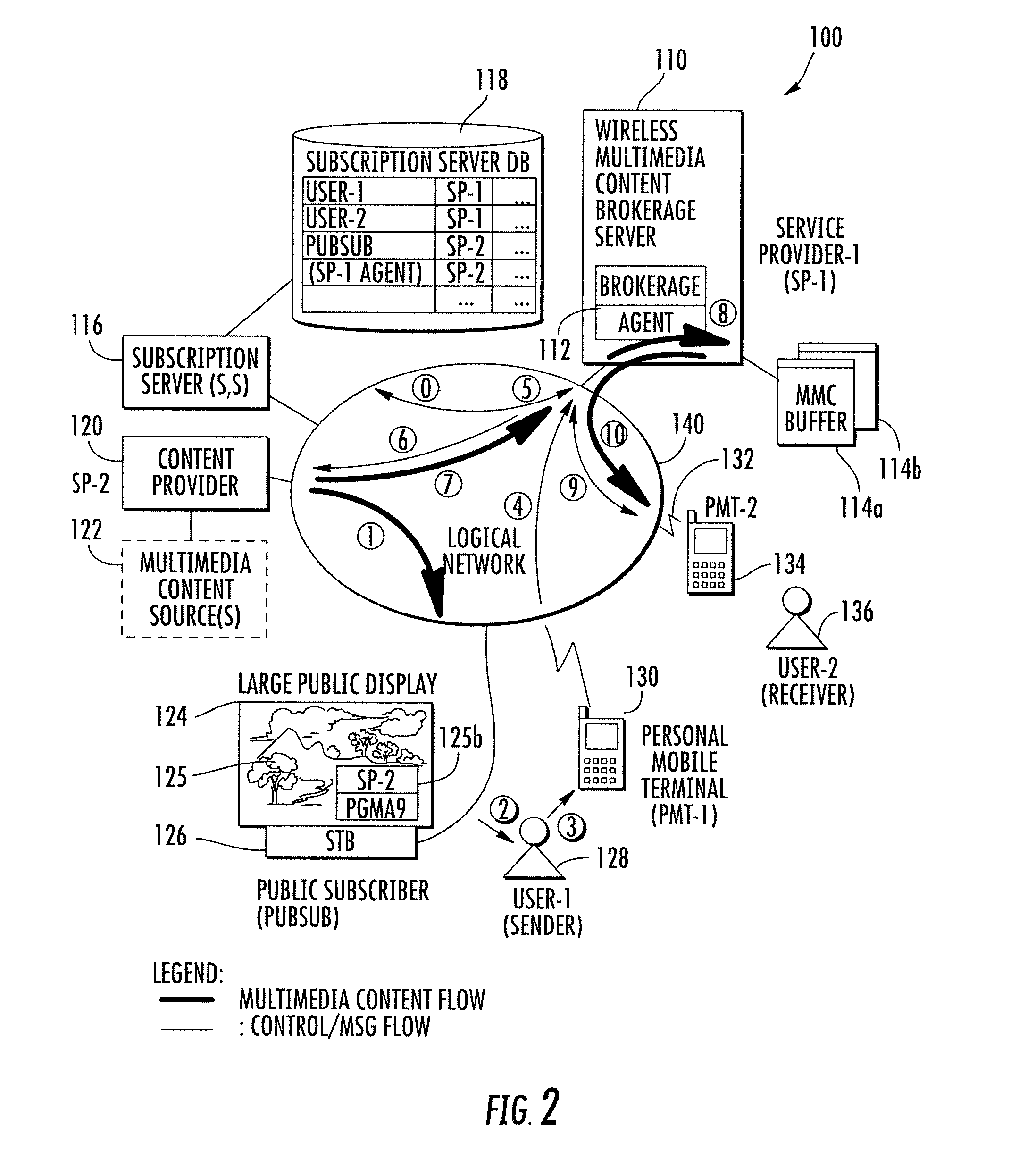
Wireless multimedia brokerage service for real time content provisioning
InactiveUS20130219076A1Accurate contentMultiple digital computer combinationsTransmissionDisplay deviceMobile device
A wireless multimedia brokerage service supports sharing of real-time multimedia content whereby a mobile device user can perceive real-time content from a device in visual proximity, such as a nearby display device, and share the content of the display device with another mobile terminal user without the need to download or otherwise directly access the real-time content. The brokerage service can handle the transactional details of obtaining rights to the real-time content and also manage establishing and terminating a real-time multimedia session with the device(s) of the recipient user(s). In some embodiments, the wireless multimedia content brokerage service can proactively obtain subscriptions to content providers based on the location of one or more users. The brokerage service can also proactively obtain and buffer real-time content after receiving a request to share the content, with the buffering allowing for content to be preserved while the recipient user or users are contacted. The content can then be pushed or otherwise provided to the recipient(s).
Owner:QURIO HLDG
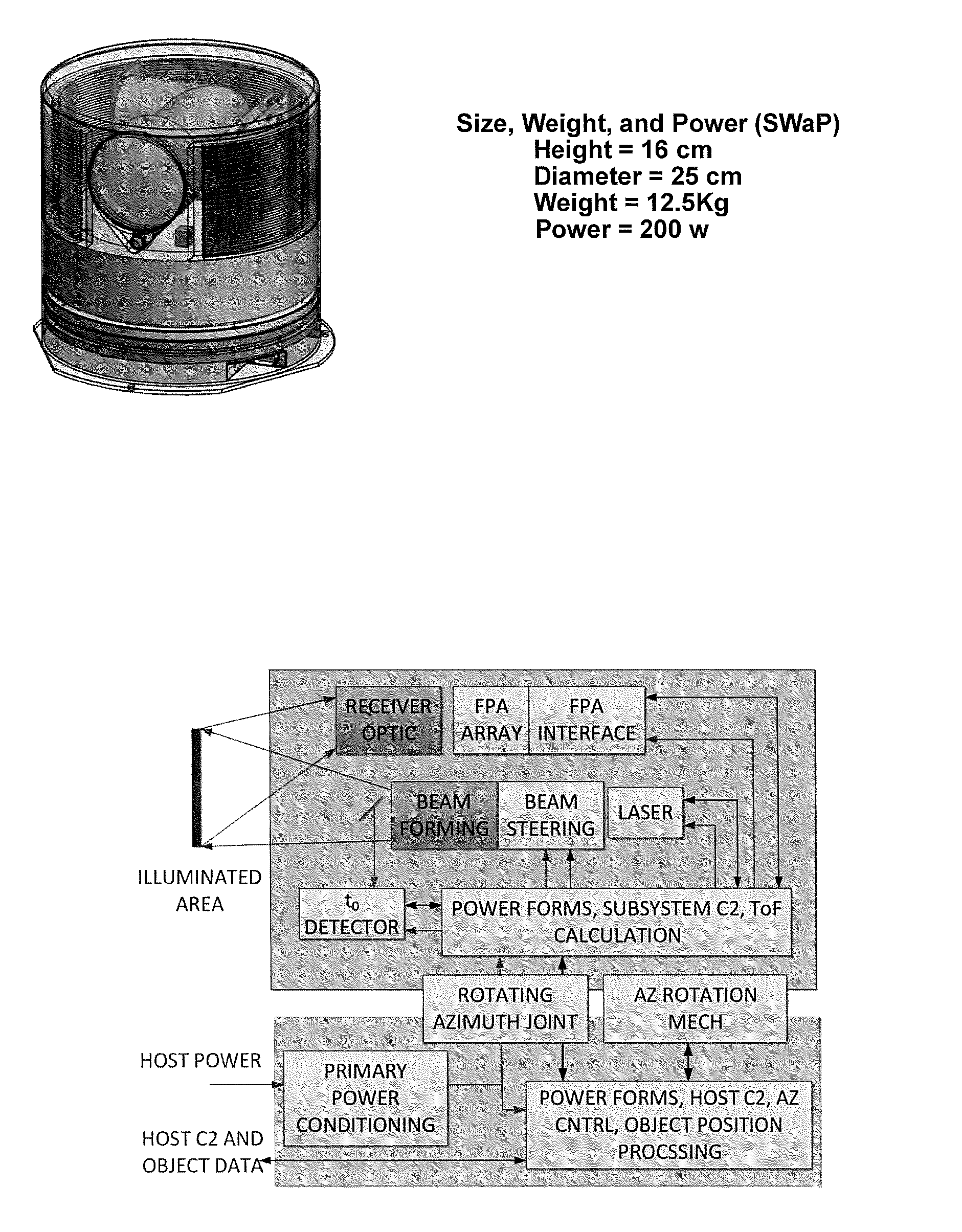
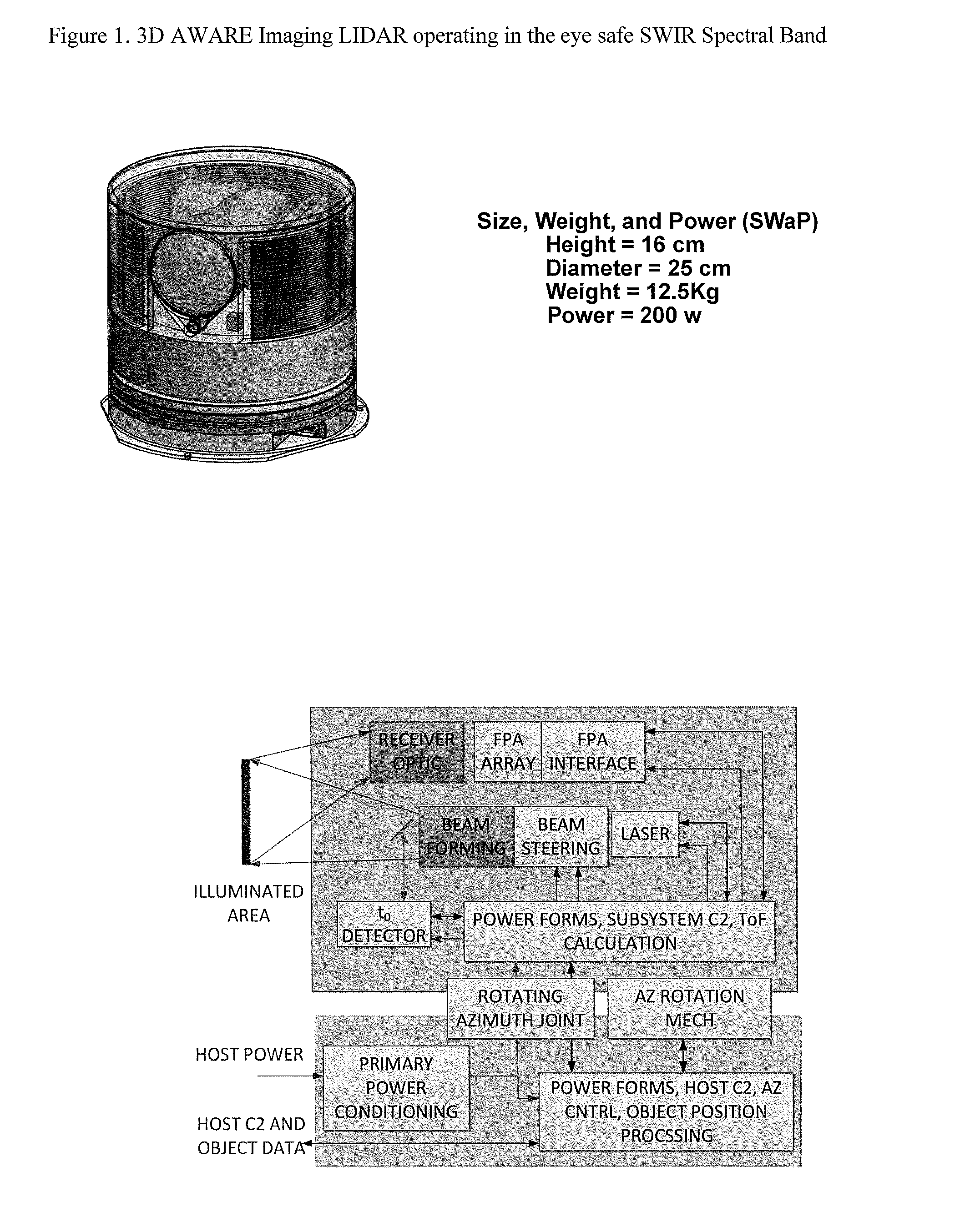
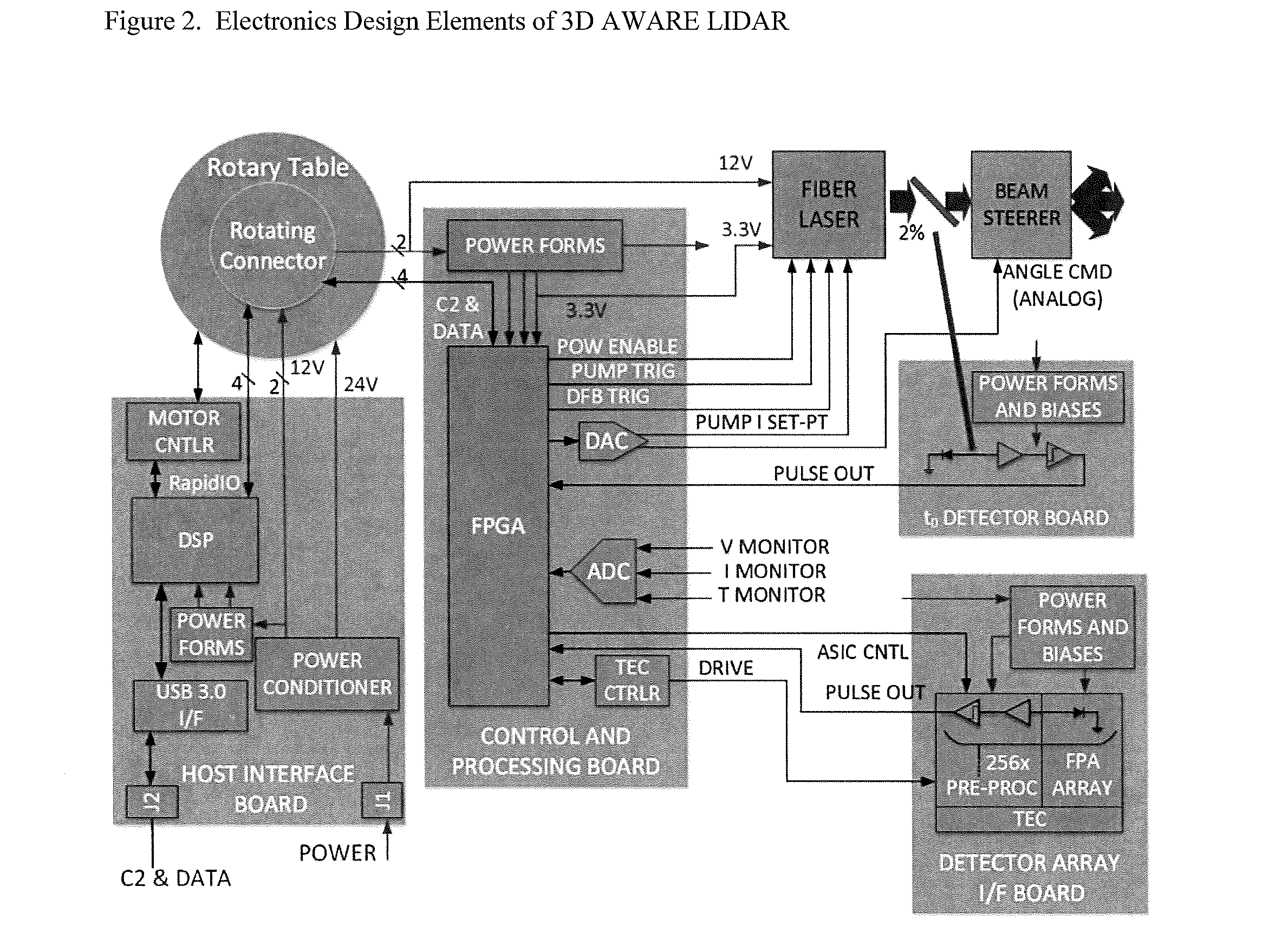
3D Active Warning and Recognition Environment (3D AWARE): A low Size, Weight, and Power (SWaP) LIDAR with Integrated Image Exploitation Processing for Diverse Applications
InactiveUS20160267669A1High resolutionImprove spatial resolutionTelevision system detailsImage analysisWide areaHigh pulse rate
An invention is disclosed for a multi-mode LIDAR sensor system that embodies a high pulse rate fiber laser operating in the SWIR wavelength at 1.5 microns, a long linear array of small SWIR sensitive detectors with very high speed readout electronics, and fully integrated methods and processing elements that perform target detection, classification, and tracking using techniques that emulate how the human visual path processes and interprets imaging data. High resolution three dimensional images are created of wide areas. Image exploitation processing methods detect objects and object activities in real time thus enabling diverse applications such as vehicle navigation, critical infrastructure protection, and public safety monitoring.
Owner:IRVINE SENSORS
Thermal spray material and process for preparing same
Owner:NIPPON YTTRIUM