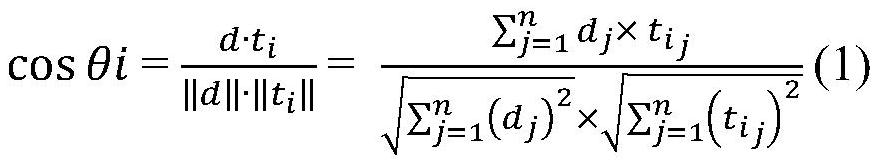Patents
Literature
32results about How to "Simple model structure" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
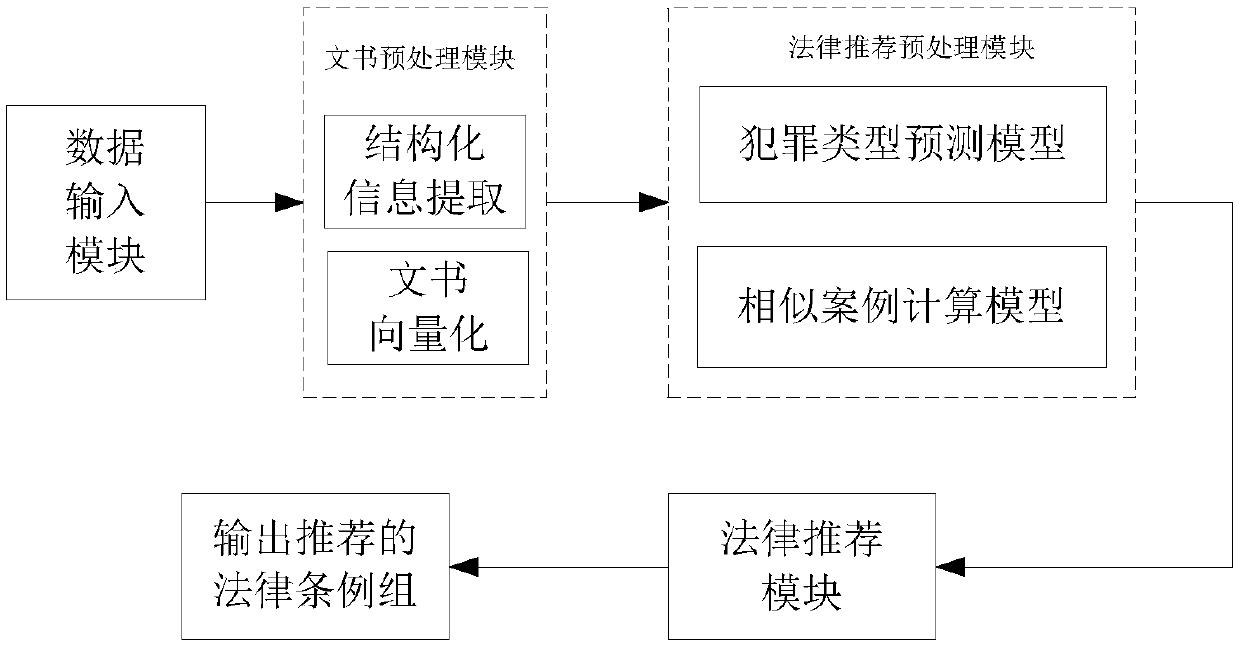
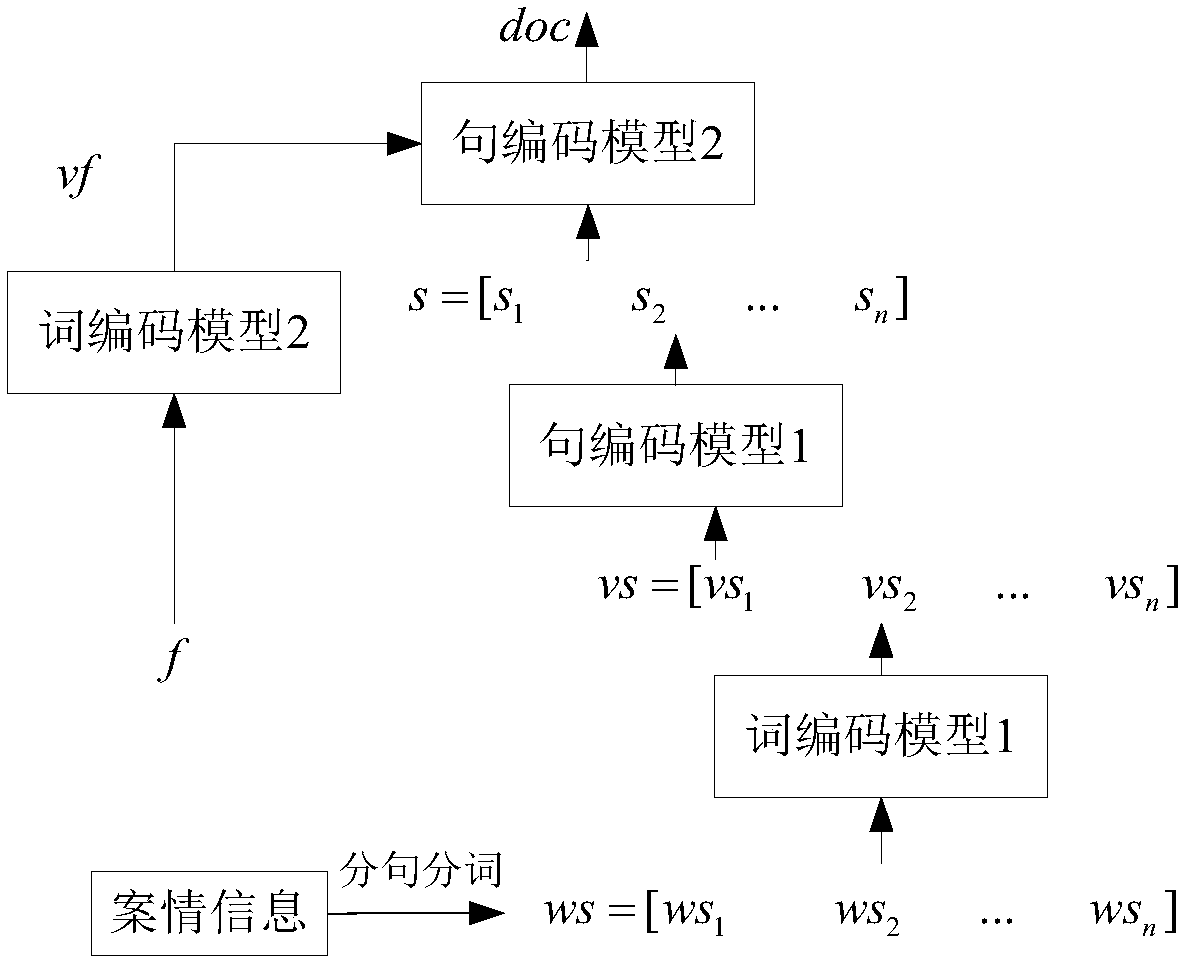
Case legal regulation recommending method and system
ActiveCN107818138ASimple model structureImprove accuracyData processing applicationsSpecial data processing applicationsPaper documentDocument preparation
The invention discloses a case legal regulation recommending method. The method comprises the steps of 1, obtaining judgment document information and basic case deciding legal regulation information,and processing sensitive information involved in a judgment document; 2, preprocessing the obtained judgment document to obtain structuralized information and non-structuralized information; 3, vectorizing the structuralized information, conducting sentence splitting and word splitting on the non-structuralized information, afterwards vectorizing the non-structuralized information, and jointly coding the vectorized structuralized information and the vectorized non-structuralized information to form judgment document vectors; 4, inputting the judgment document vectors to a criminal type prediction model to obtain a detailed type of a corresponding case and a legal regulation applied in the type of the case; 5, predicting a criminal type of the case to obtain a relevancy matrix of a legal provision, calculating legal regulation confidence coefficient applied in a similar case, and finally putting forward a legal provision recommending combination applied in the case based on the criminaltype and the similar case.
Owner:ENJOYOR COMPANY LIMITED
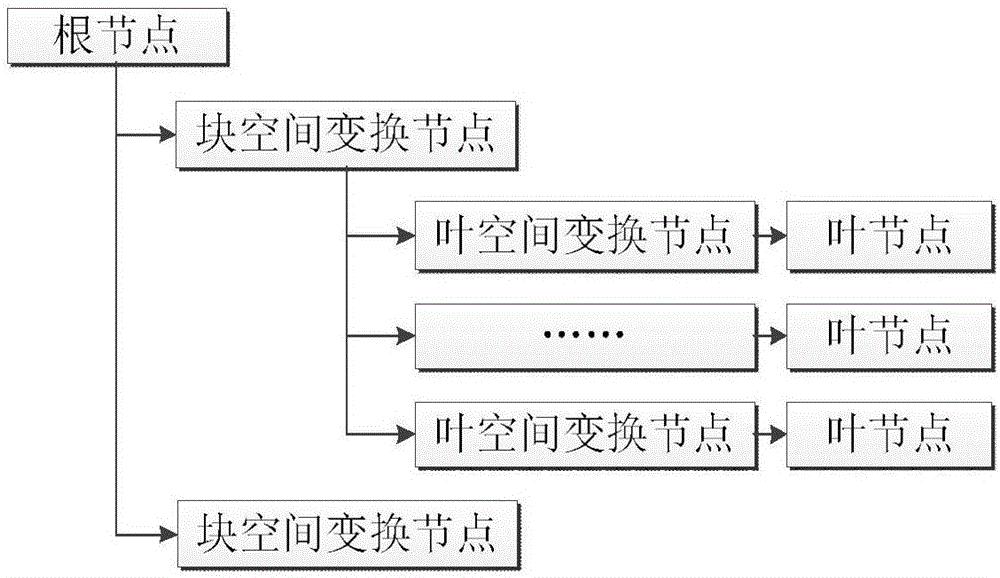
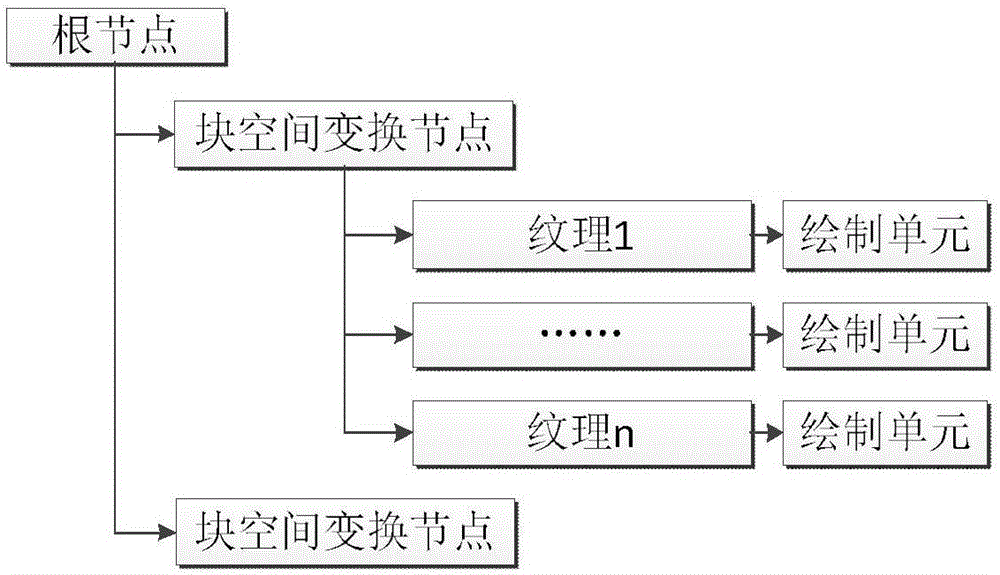
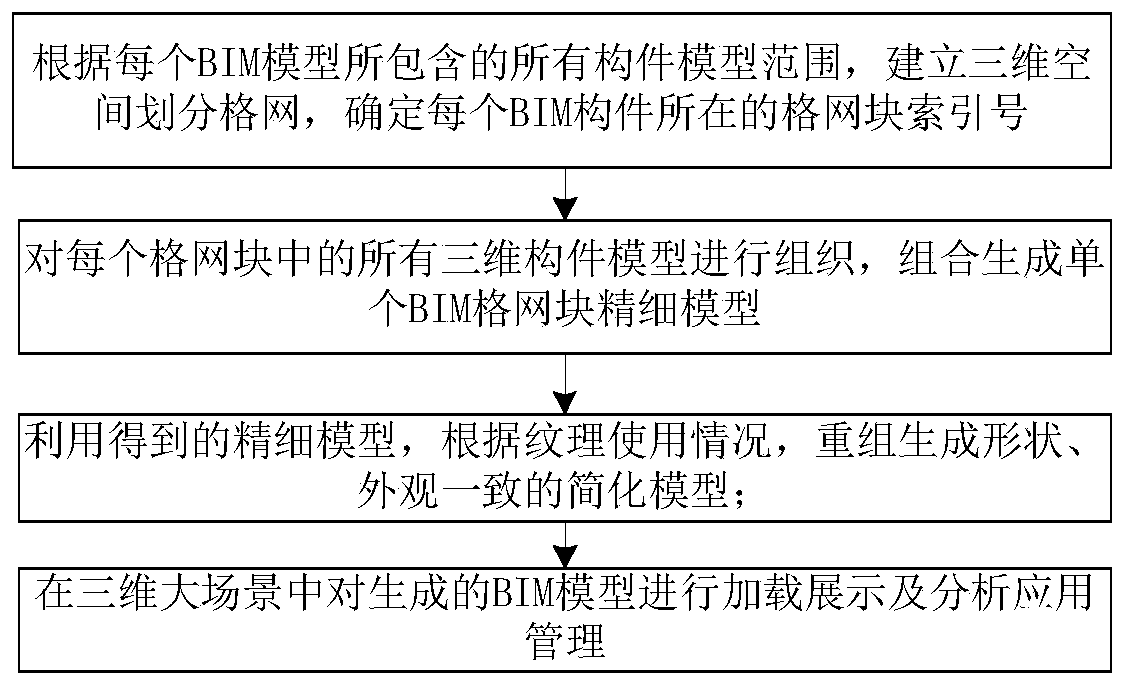
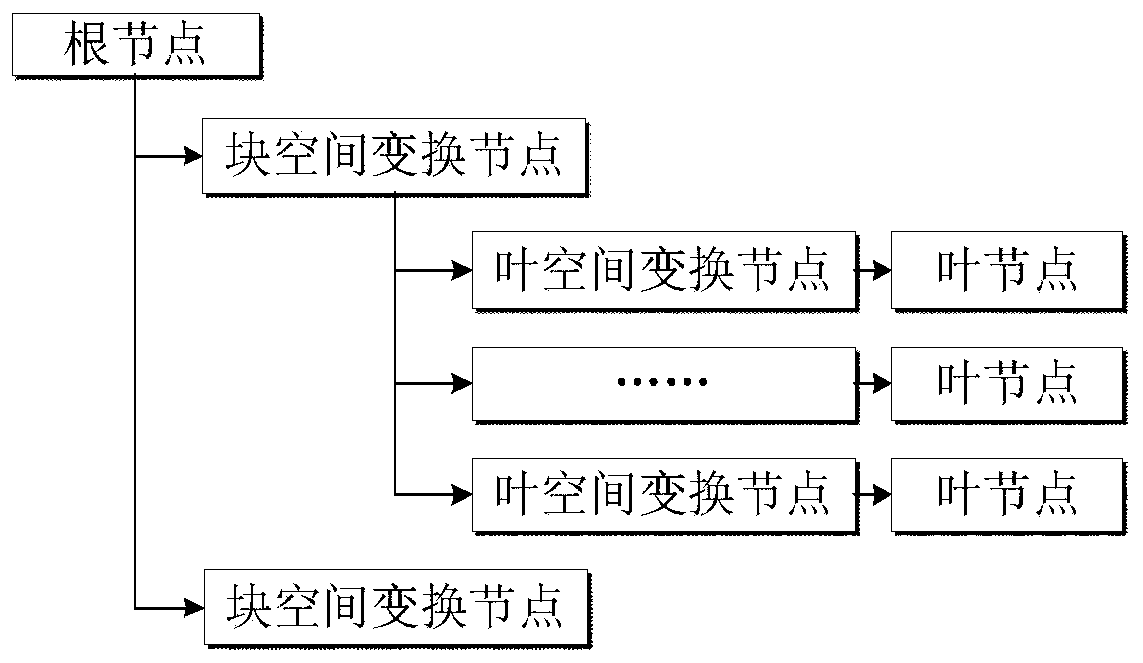
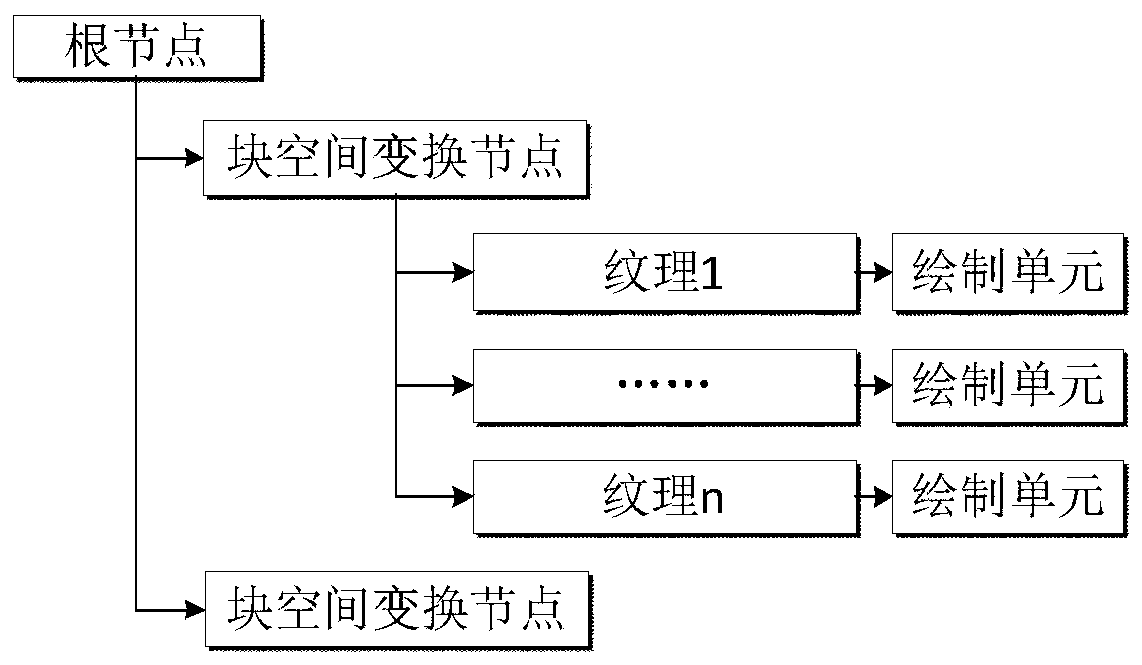
Visual implementation method of BIM model in three-dimensional large scene
ActiveCN106599493AReduce in quantityImprove display efficiencyGeometric CADSpecial data processing applicationsReduced modelThree-dimensional space
The invention discloses a visual implementation method of a BIM model in a three-dimensional large scene, and relates to the field of building information model visualization. The visual implementation method comprises the following steps: firstly, establishing a three-dimensional space division grid according to the dimension of a range of all member models contained in each BIM model; then organizing the member models in each grid block to combine and generate a precise model of a single grid block; recombining and generating a simplified model with consistent shape and appearance according to vein use conditions by using the obtained precise model; and finally, loading and displaying the precise model or the simplified model of the BIM model in the three-dimensional large scene according to a selected display mode and an index number. According to the visual implementation method disclosed by the invention, on the basis of guaranteeing the application of the BIM model, all members of each BIM model are divided according to the three-dimensional space division grid by means of the space grid idea, and the members in the grid are merged to reduce the number of the models of the index and improve the loading efficiency of the BIM model.
Owner:CHONGQING SURVEY INST
A perceptron network data classification method based on an AdaBoost algorithm
InactiveCN109726767AImprove classification accuracySimple model structureCharacter and pattern recognitionNumeric ValueActivation function
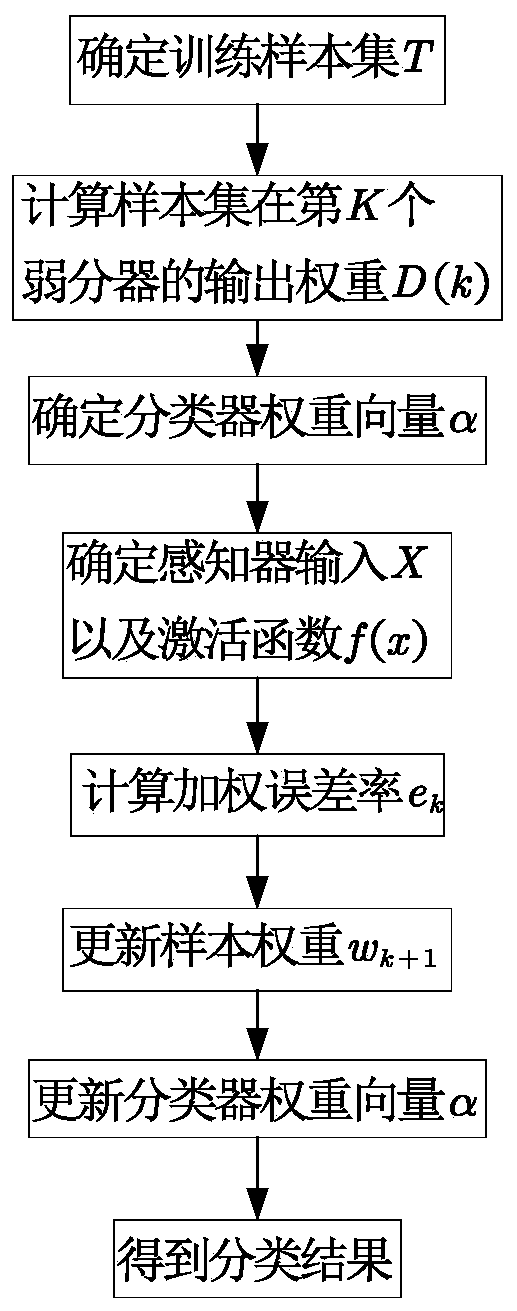
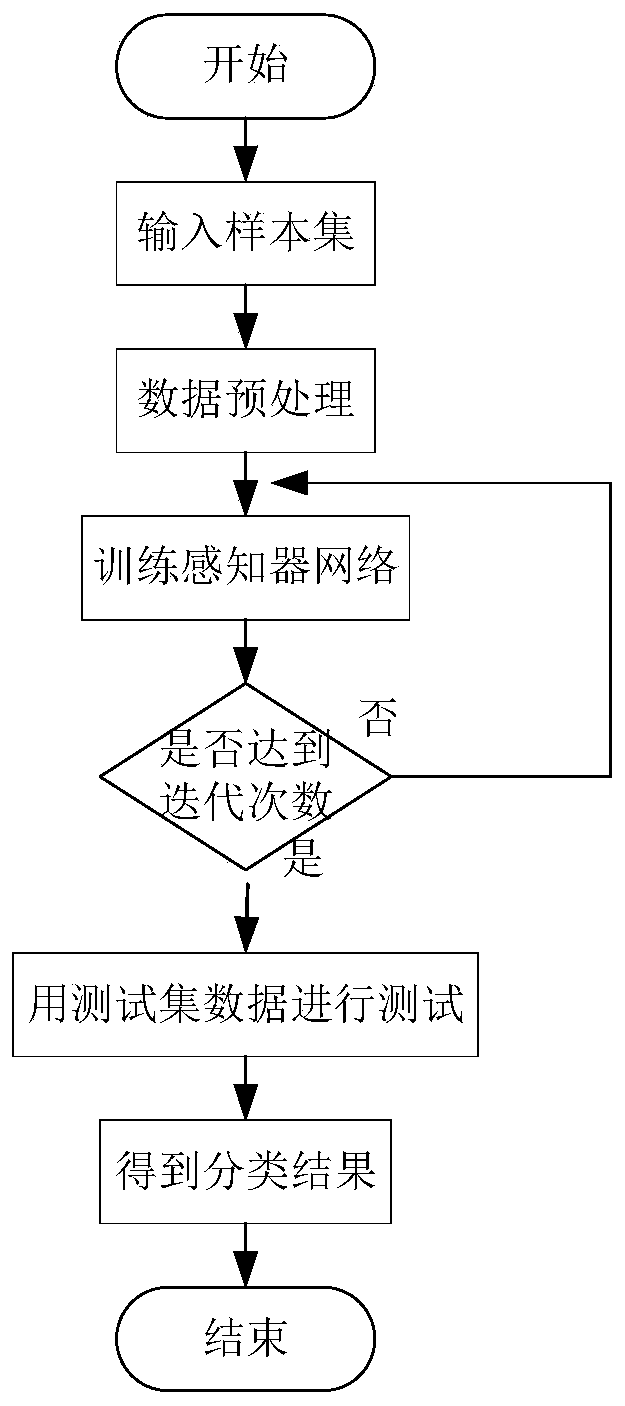
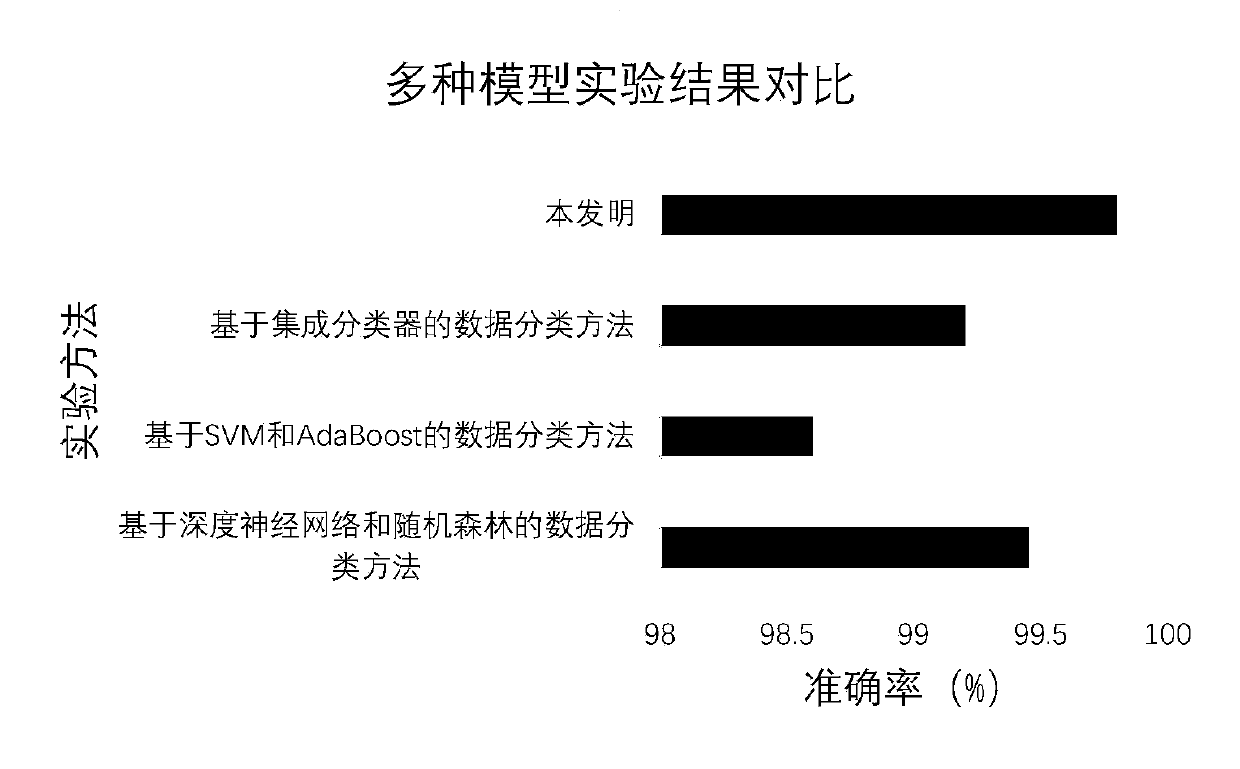
The invention relates to a perceptron network data classification method based on an AdaBoost algorithm. The perceptron network data classification method is characterized by comprising the followingsteps: (1) determining a training sample set; (2) calculating the output weight of the sample set in the weak classifier; (3) determining a classifier weight vector; (4) determining input of a sensorand an activation function; (5) calculating a weighted error rate; (6) updating the sample weight; (7) updating a classifier weight vector; And (8) taking a test set sample as an input, and sending the input to a sensor for classification to obtain a classification result. According to the method, the network connection weight of the sensor is updated and iterated by using the idea of integrated learning in the AdaBoost algorithm, and a plurality of weak classifiers are integrated into a strong classifier, so that the classification accuracy of the model is improved. According to multiple groups of data experiment results, the classification method for improving the timeliness of the model on the basis of ensuring the classification accuracy is provided for numerical classification.
Owner:胡燕祝
Tobacco tar bottle
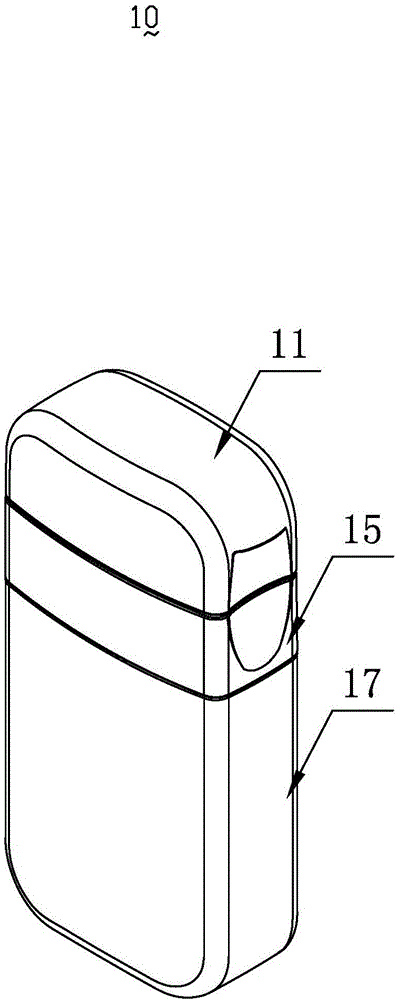
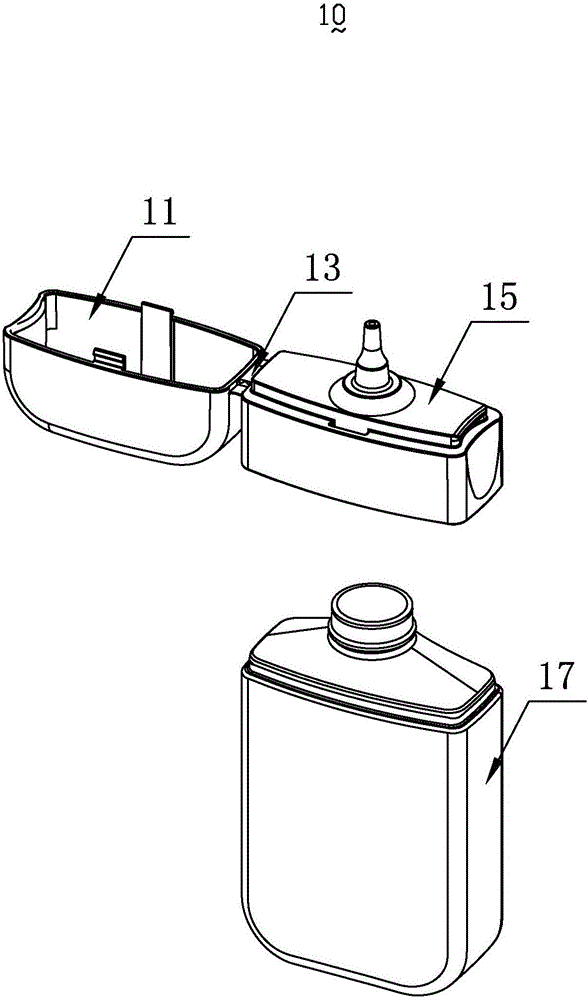
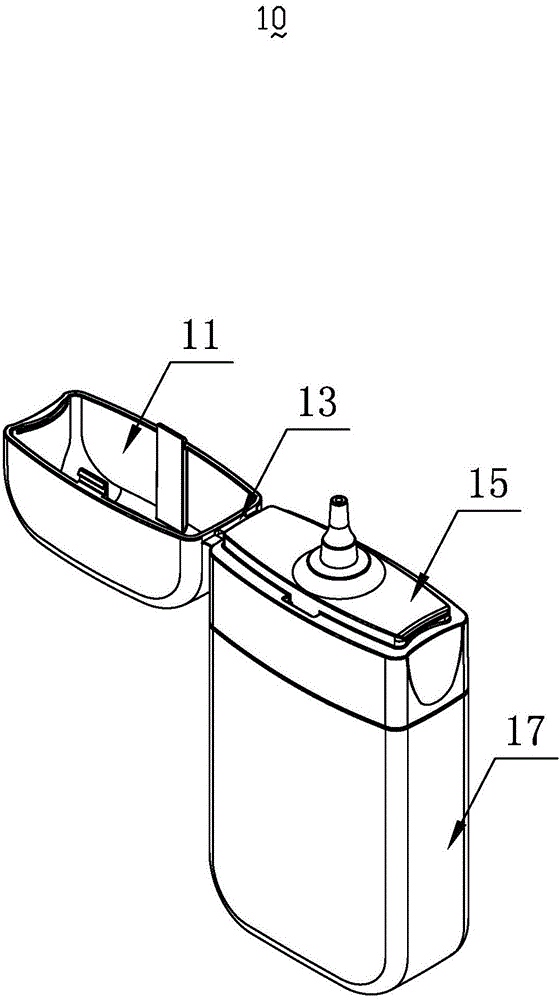
InactiveCN103818629ASimple model structureSimple production processBottlesContainer/bottle contructionTarBottle
The invention provides a tobacco tar bottle used for accommodating tobacco tar. The tobacco tar bottle comprises a bottle body storing the tobacco tar. The bottle body is provided with a bottle body mouth, a lower cover and an upper cover. The bottle body mouth is used for injecting the tobacco tar. The lower cover is provided with a bottle mouth, is connected to the bottle body mouth and decreases the flow diameter of the tobacco tar in the bottle body through the bottle mouth. The upper cover covers the bottle mouth. A second seal structure is provided between the lower cover and the upper cover and corresponds to the bottle mouth of the lower cover. A first seal structure is provided between the lower cover and the bottle body and corresponds to the bottle body mouth. A first clamping-connection structure is provided between the lower cover and the bottle body which are in fixed clamping connection through the first clamping-connection structure. The tobacco tar bottle is good in tightness, adaptive to various structures, simple in production process, high in production efficiency, free of safety risks, and simple in structure and has broad market prospect and high practicality.
Owner:RITCHY GRP
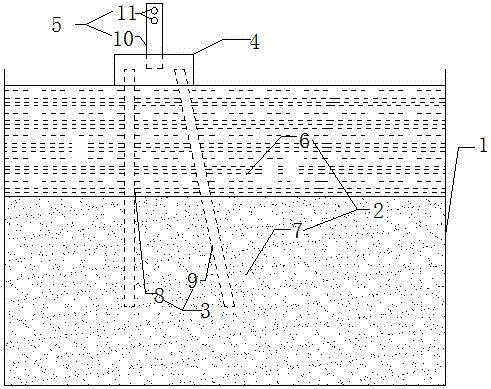
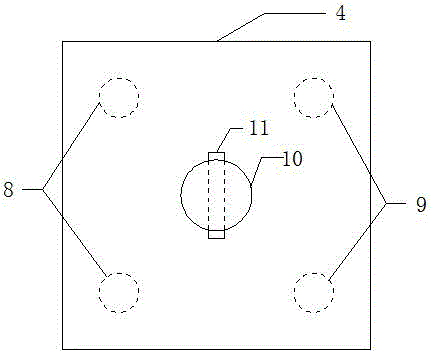
Inclined-straight alternating group piles-soil-structure interaction experiment model
InactiveCN104878786ASimple model structureEasy to makeFoundation testingSteel platesComplex problems
The invention discloses an inclined-straight alternating group piles-soil-structure interaction experiment model, belongs to the technical field of geotechnical similar simulation experiments with an aim to solve problems that the inclined-straight alternating group piles-soil-structure interaction experiment model is complex in manufacturing. The inclined-straight alternating group piles-soil-structure interaction experiment model is composed of a soil box, foundation soil, an inclined-straight alternating group pile foundation, a bearing platform and an upper portion structure; the soil box is of a hollow cuboid body provided with an upper opening and is formed by welding of steel plates; the foundation is divided into two layers, the upper layer is clay, and the lower layer is sandy soil; the inclined-straight alternating group pile foundation is formed by two vertical piles and two inclined piles, and pile bottoms are identical in elevation; the bearing platform is a cuboid body cast with concrete, and the vertical piles and the inclined piles are integrally connected through the bearing platform; the upper portion structure is composed of a single-column pier and a counter weight, the single-column pier is of a cylindrical body, two vertically-arrayed threaded rods are embedded in the upper end of the single-column pier, and the counter weight is formed by combination of weights different in weight. The inclined-straight alternating group piles-soil-structure interaction experiment model is simple in structure, convenient to machine and manufacture and high in engineering practicability.
Owner:SHANDONG UNIV OF SCI & TECH
Apple tree canopy scale nitrogen content diagnosis method
PendingCN112129709AAvoid errorsGood dimensionality reduction effectColor/spectral properties measurementsNeural architecturesLearning machineSoil science
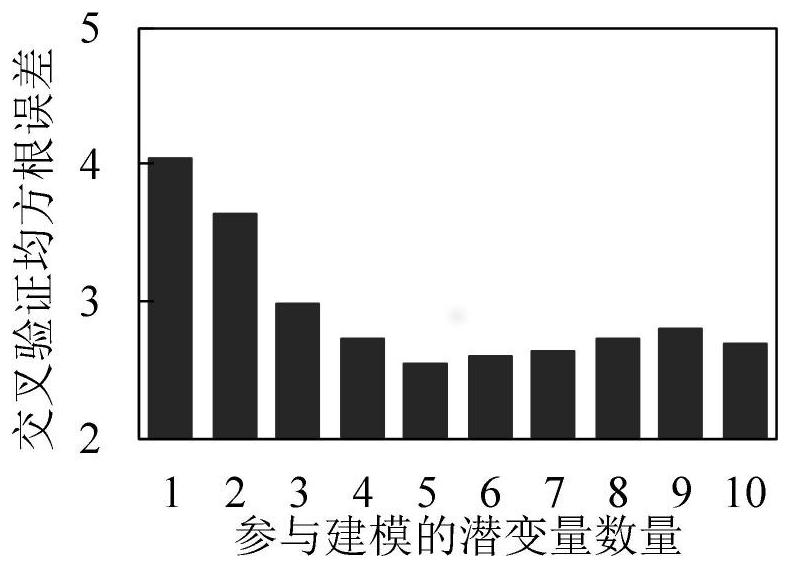
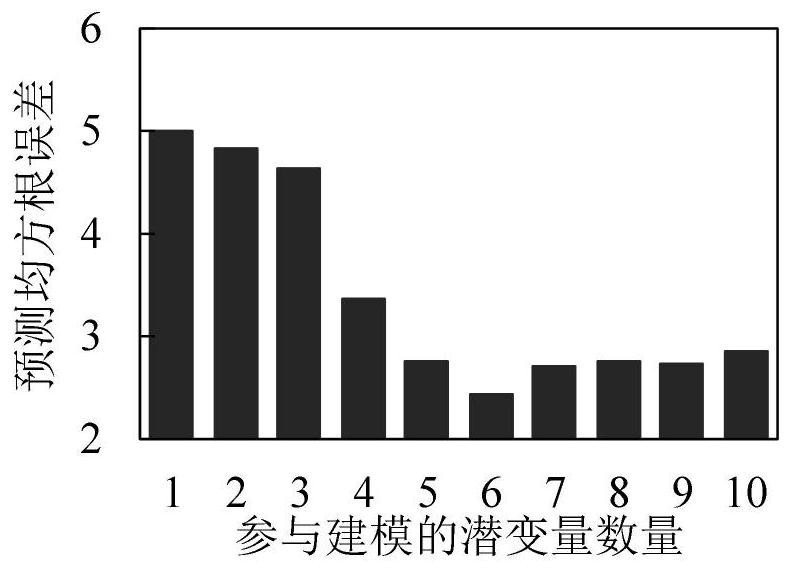
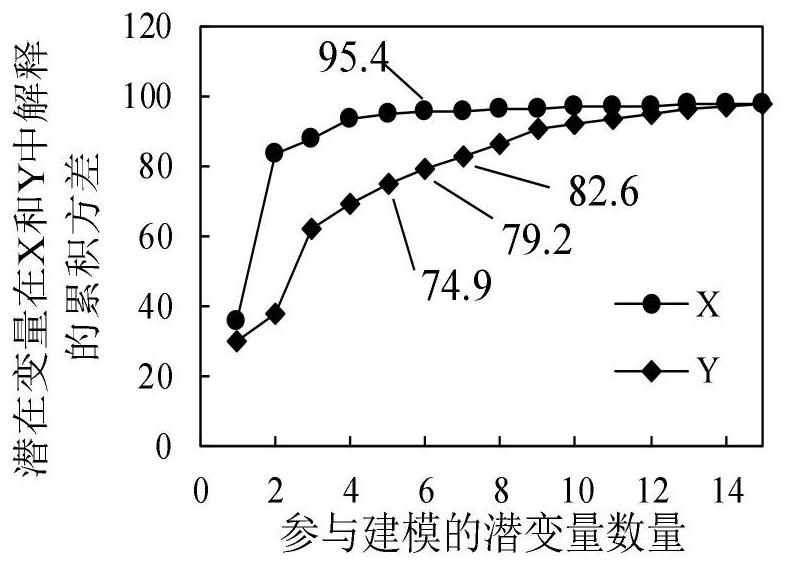
The invention discloses an apple tree canopy scale nitrogen content diagnosis method. The method comprises the following steps: determining the reflectivity of an apple tree canopy scale by adopting ahyper-spectrometer; selecting an apple tree canopy leaf sample; determining the total nitrogen content of the apple tree canopy leaf sample; identifying abnormal samples in a reflectivity data set and a total nitrogen content data set by utilizing a Monte Carlo secondary detection method; preprocessing the reflectivity data set; aiming at the obtained reflectivity data sets of 1326 wave bands, extracting variables based on a potential variable method in a partial least squares regression analysis process; establishing a model for the extracted variables by adopting an extreme learning machine; and predicting the nitrogen content of the apple tree canopy leaves by utilizing the established model. According to the method, canopy scale spectral information is collected for nitrogen conditiondiagnosis, errors caused by sample randomness due to existing leaf scale diagnosis are effectively avoided, and the diagnosis result can measure the overall canopy nitrogen condition of the apple tree.
Owner:NORTHWEST A & F UNIV
110kV transformer entity modeling method for vibration characteristic analysis
PendingCN112949126ASimple model structureMeet the accuracy requirementsDesign optimisation/simulationSpecial data processing applicationsElement modelTransformer
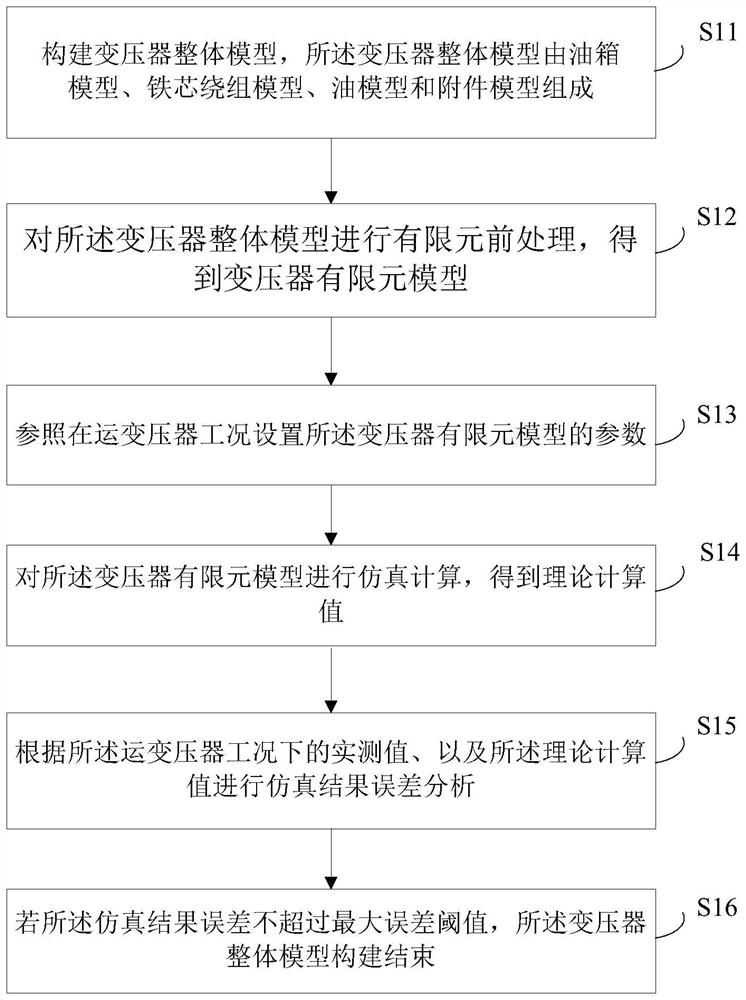
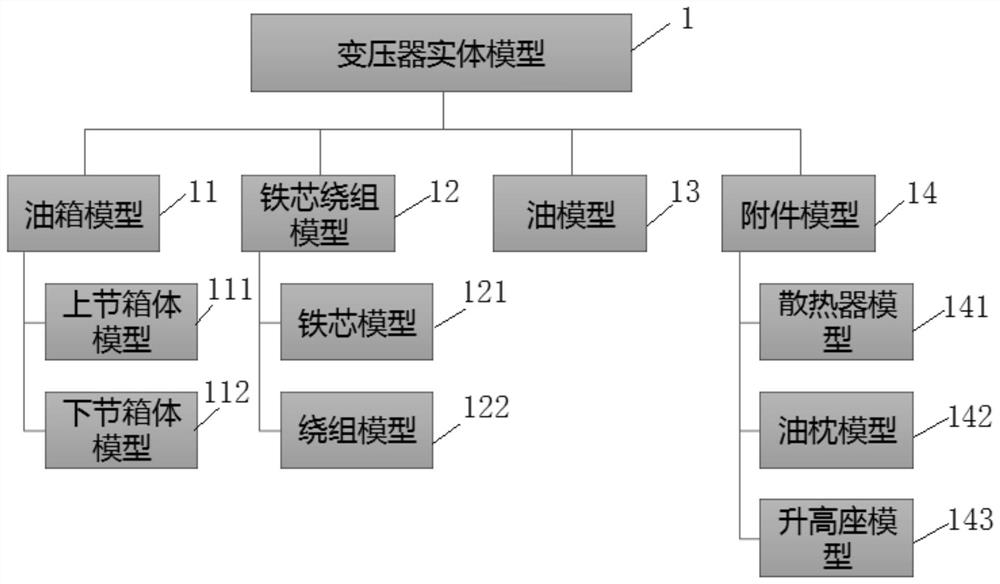
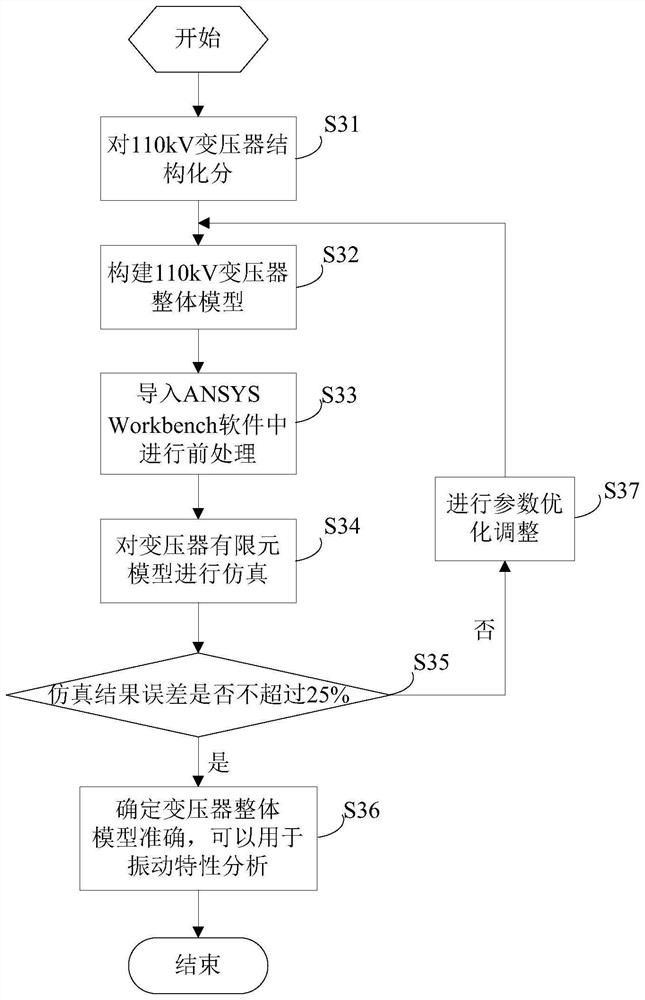
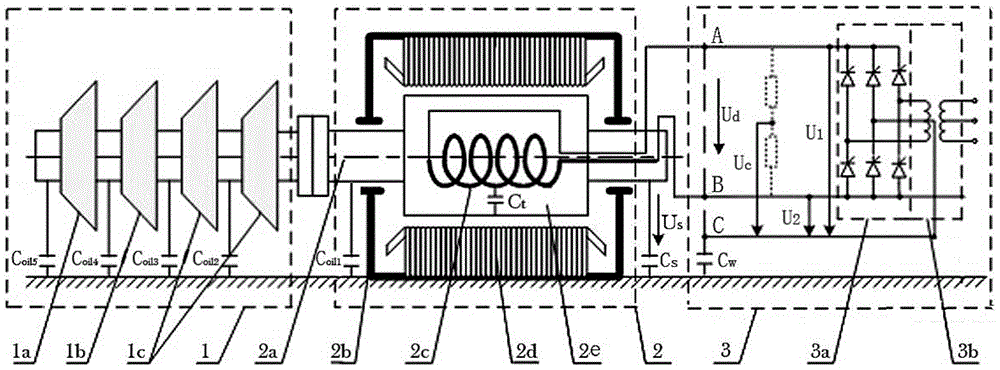
The invention provides a 110kV transformer entity modeling method for vibration characteristic analysis, and belongs to the field of transformer vibration characteristic analysis. The method comprises the steps: constructing a transformer overall model, wherein the transformer overall model is composed of an oil tank model, an iron core winding model, an oil model and an accessory model; carrying out finite element pretreatment on the overall model of the transformer to obtain a finite element model of the transformer; setting parameters of the finite element model of the transformer according to working conditions of the in-operation transformer; performing simulation calculation on the finite element model of the transformer to obtain a theoretical calculation value; performing simulation result error analysis according to the actual measurement value under the operation transformer working condition and the theoretical calculation value; and if the simulation result error does not exceed the maximum error threshold value, finishing the construction of the overall model of the transformer.
Owner:YINCHUAN POWER SUPPLY COMPANY OF STATE GRID NINGXIA ELECTRIC POWER +1
Model for modeling large steam turbine generator with function of static excitation
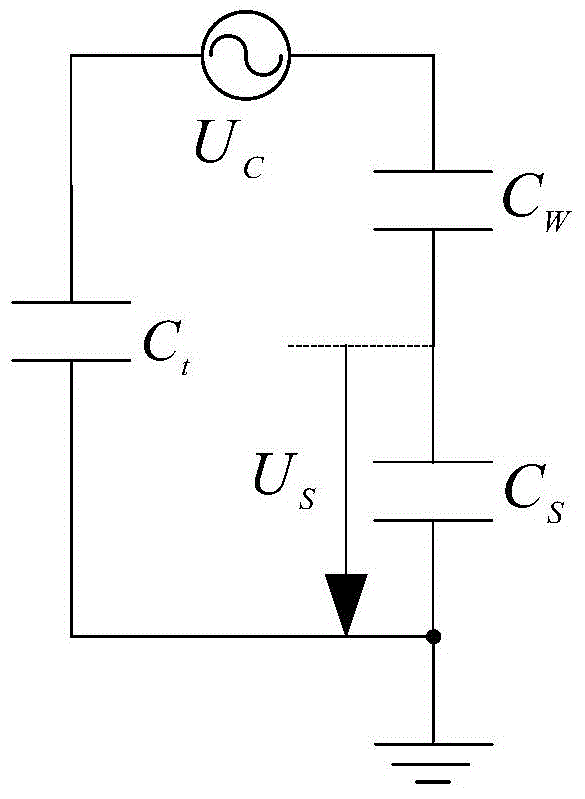
InactiveCN104156513AConvenient simulation analysisConvenient researchSpecial data processing applicationsCapacitanceEngineering
The invention discloses a model for modeling a large steam turbine generator with a function of static excitation. The model comprises a static excitation subsystem model, an excitation winding subsystem model and a rotor shafting subsystem model; the static excitation subsystem model implemented by a modeling method considers ground capacitance of a neutral point; the excitation winding subsystem model considers the influence of input-output upon an end and excitation winding loss related to frequency; the rotor shaft system subsystem model considers ground capacitance of a rotor shaft and shafting impedance of a steam turbine. The output end of the static excitation subsystem model is connected to the input end of the excitation winding subsystem model. The output end of the static excitation subsystem model is connected to the input end of the excitation winding subsystem model. The output end of the excitation winding subsystem model is connected to the input end of the rotor shafting subsystem model. The static excitation subsystem model of the generator is established, the excitation winding subsystem model and the rotor shafting subsystem model are established, shaft voltage waveform produced by a static excitation system can be acquired for the generator in a simulation manner, and simulation analysis and research for control of shaft voltage is facilitated. The model has the advantages of simple structure, easiness in implementation and high precision.
Owner:STATE GRID CORP OF CHINA +3
Pulse tube refrigerator working condition prediction method and system based on machine learning
ActiveCN112966399AAccurate predictionReduce cost of measurementDesign optimisation/simulationNeural learning methodsMeasurement costAlgorithm
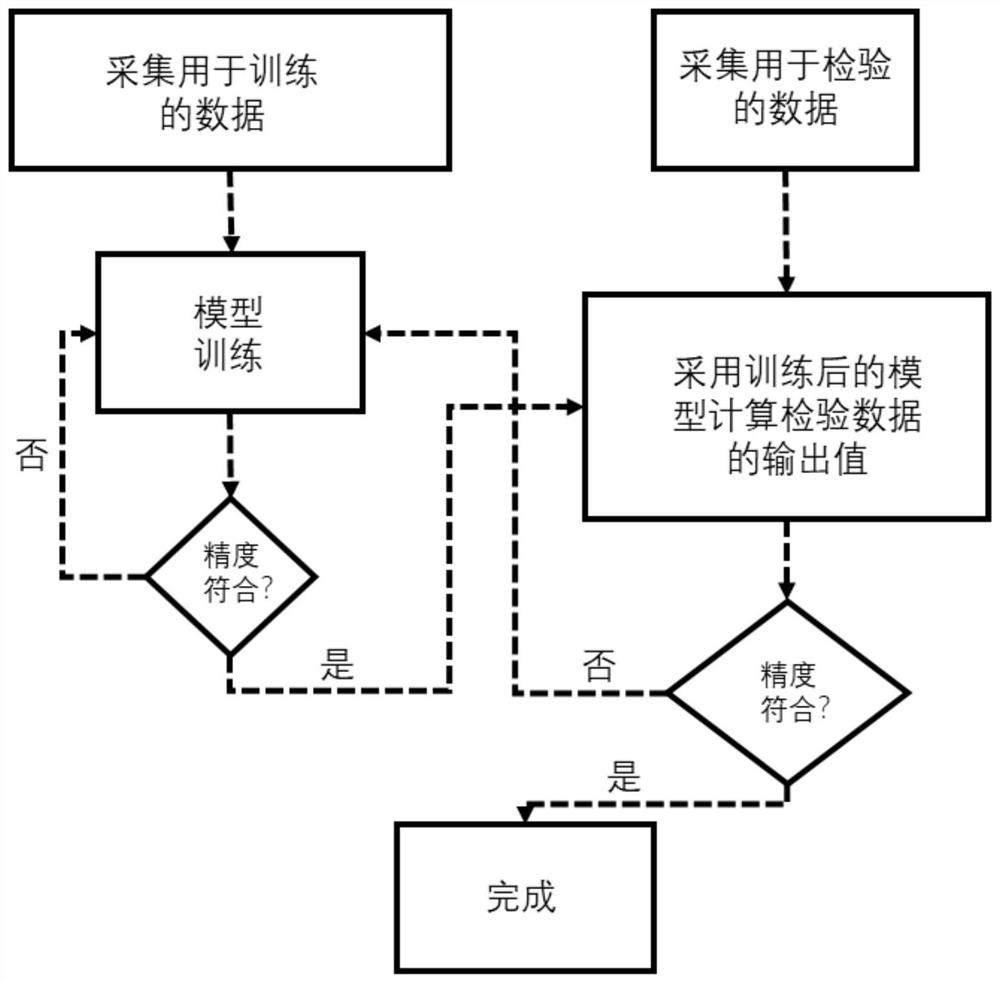
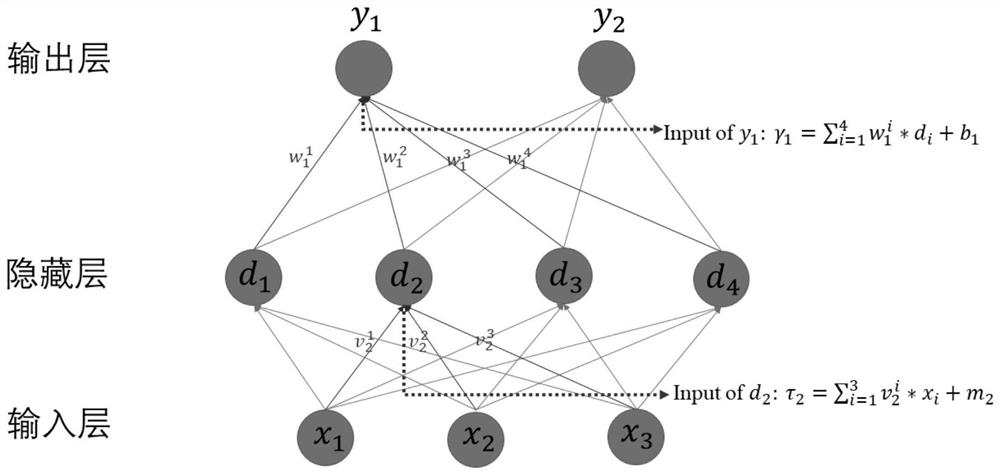
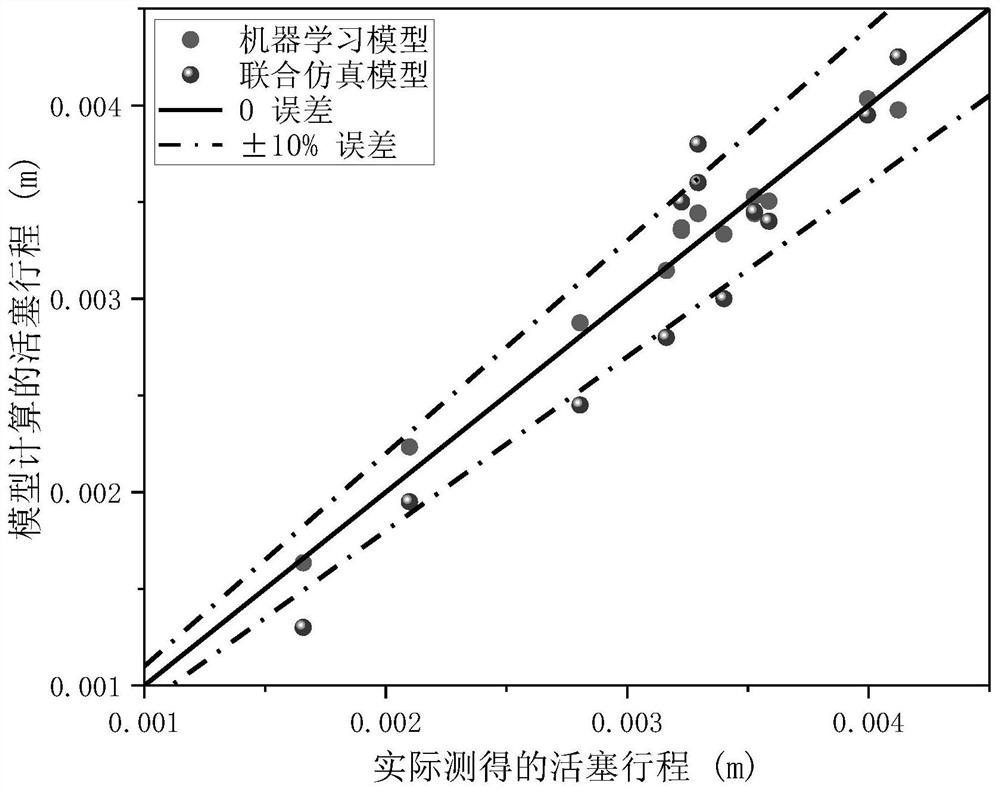
The invention relates to a pulse tube refrigerator working condition prediction method and system based on machine learning, and the method comprises the following steps: collecting working condition parameter data during the operation of a pulse tube refrigerator, and dividing the working condition parameter data into two parts: training data and inspection data; building a working condition prediction model by adopting an LM optimized back propagation algorithm based on the training data, and continuously improving the precision of the working condition prediction model through adoption of an iteration method; and inputting the test data into the working condition prediction model, comparing a predicted value calculated by the working condition prediction model with actually measured data, and verifying the precision of the working condition prediction model. The system comprises a data acquisition module, a learning training module and an inspection module. According to the invention, the training learning model is set up by collecting the working condition parameter data during operation of the pulse tube refrigerator, the piston stroke and the pressure amplitude, which are difficult to measure, of the given PTC are accurately predicted, the measurement cost of the method is lower than that of an additionally-installed sensor, and the deviation between a predicted value and an actual value is small.
Owner:ZHANGJIAGANG INST OF IND TECH SOOCHOW UNIV +1
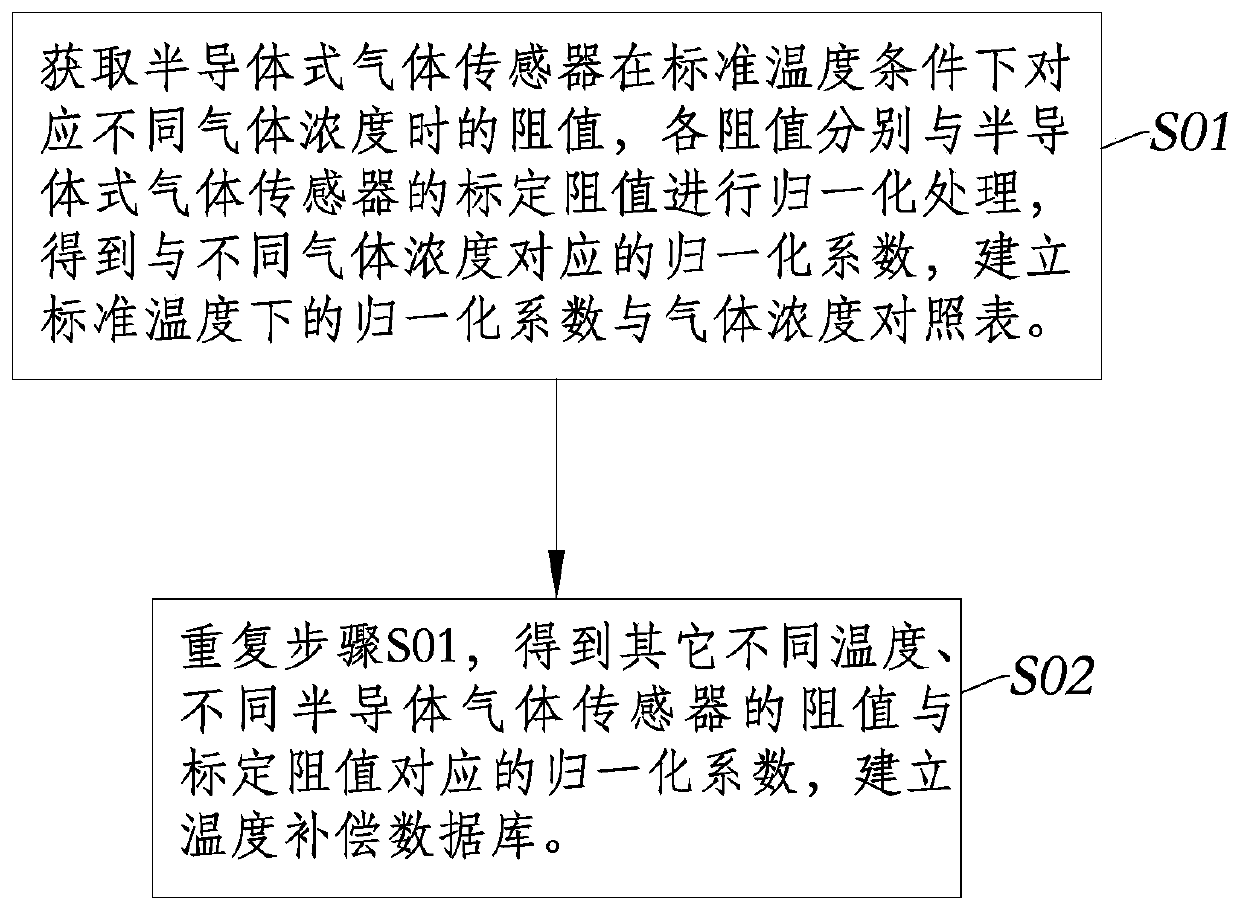
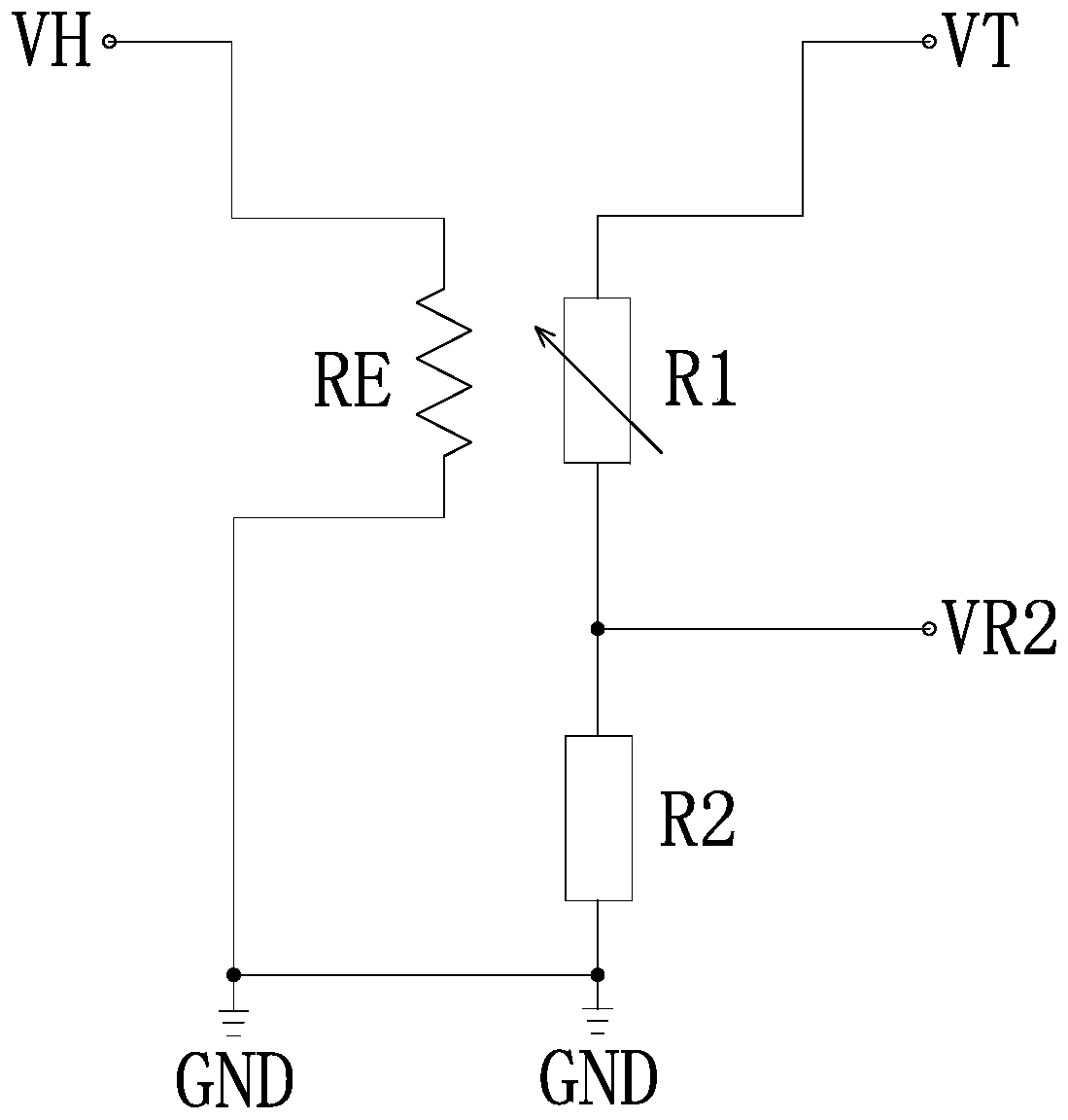
Full-range temperature compensation method of semiconductor gas sensor
ActiveCN110220945ASimple model structureReduce difficultyMaterial resistanceEngineeringGas concentration
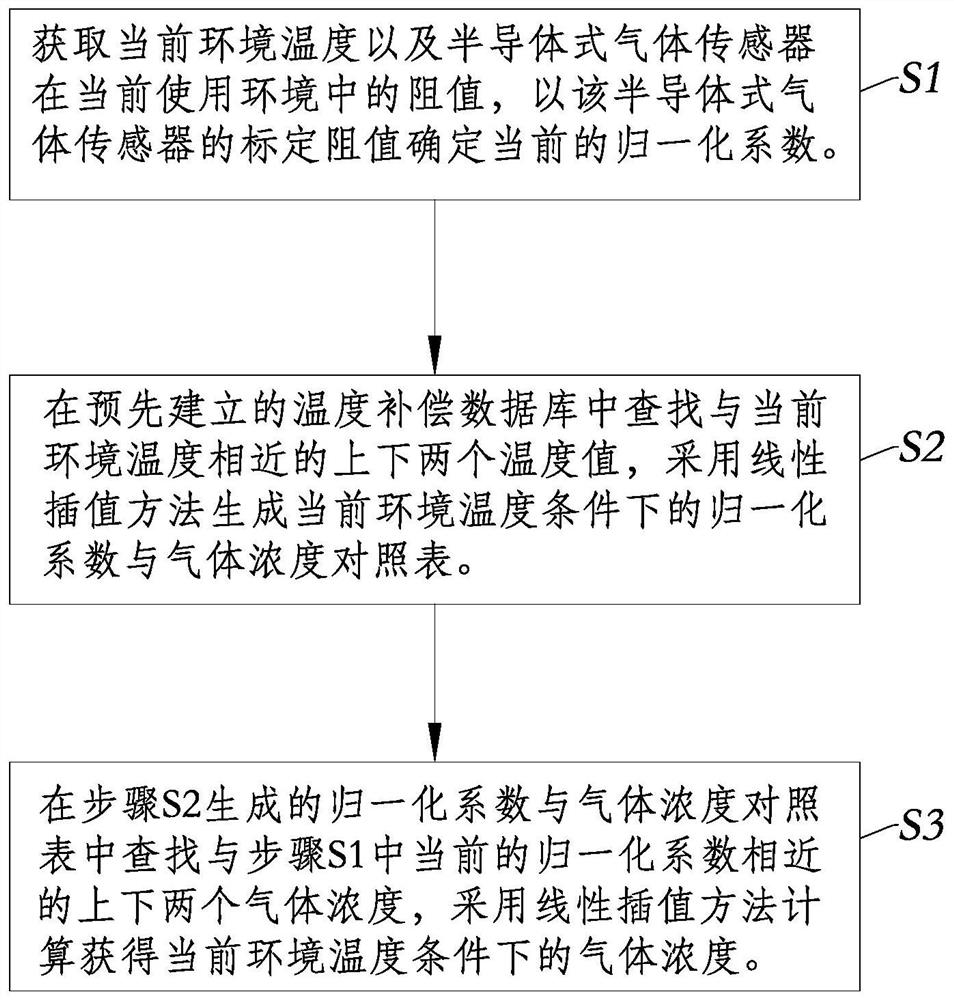
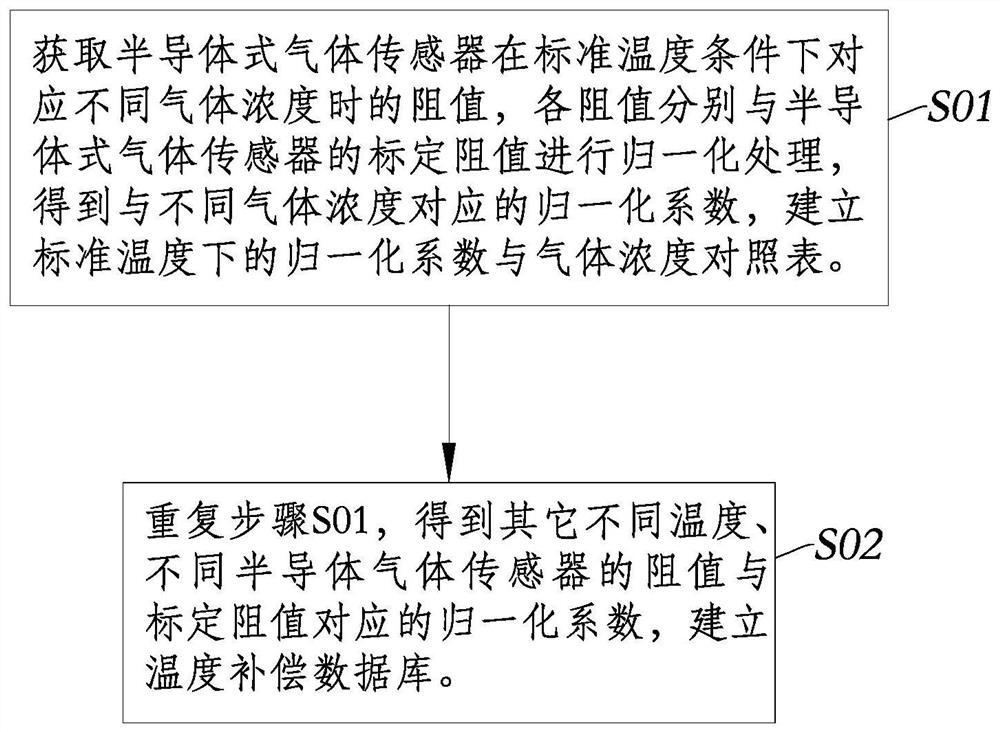
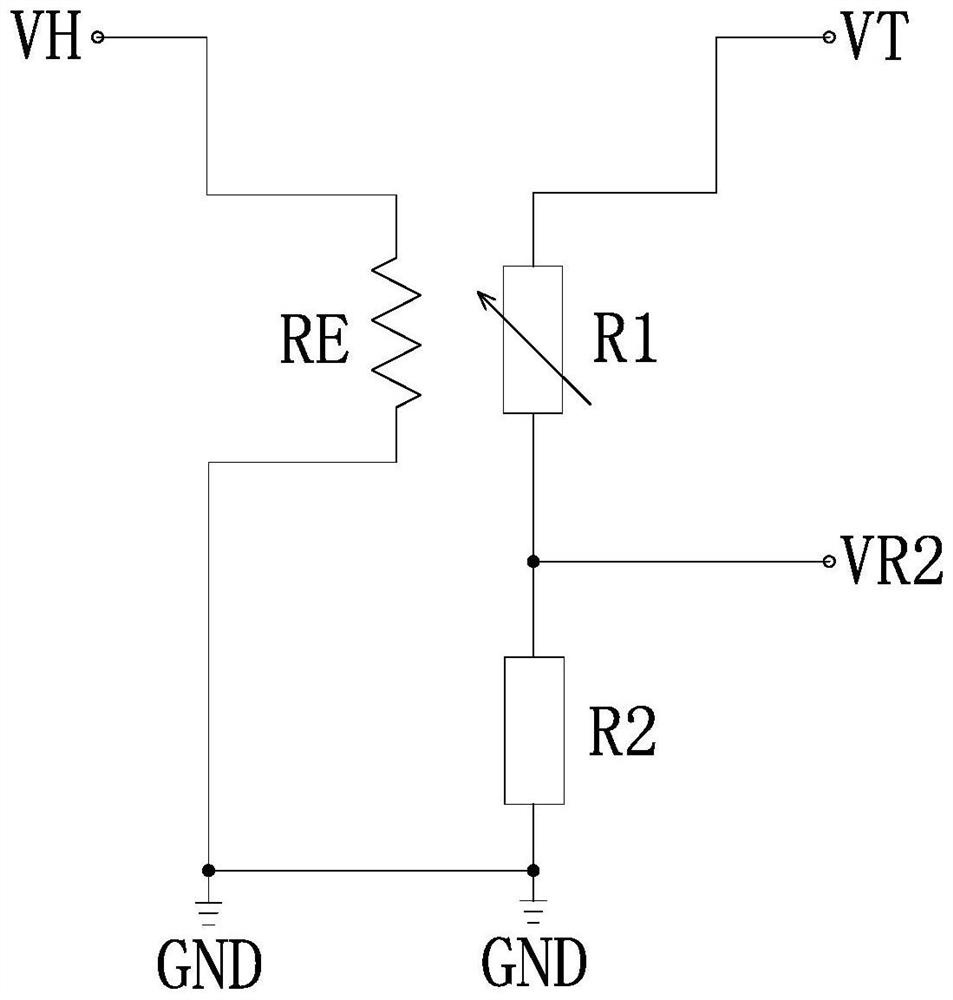
The invention discloses a full-range temperature compensation method of a semiconductor gas sensor, belongs to the technical field of gas sensor detection, and solves the defect that a conventional semiconductor gas sensor cannot be used for gas concentration output. The full-range temperature compensation method comprises the following steps of: S1, acquiring a current environment temperature andthe resistance value of the semiconductor gas sensor in the current use environment, and determining the current normalization coefficient according to the calibration resistance value of the semiconductor gas sensor; S2, searching an upper temperature value and a lower temperature value which are close to the current environment temperature in a pre-established temperature compensation database,and generating a normalization coefficient and gas concentration comparison table under the current environment temperature condition by adopting a linear interpolation method; and S3, searching theupper gas concentration and the lower gas concentration which are close to the current normalization coefficient in the step S1 from the normalization coefficient and gas concentration comparison table generated in the step S2, and performing calculating to obtain the gas concentration under the current environment temperature condition by adopting the linear interpolation method.
Owner:GOLDCARD HIGH TECH
A visualization method of BIM model in 3D large scene
ActiveCN106599493BImprove loading efficiencyImprove display efficiencyGeometric CADDesign optimisation/simulationReduced modelThree-dimensional space
The invention discloses a visual implementation method of a BIM model in a three-dimensional large scene, and relates to the field of building information model visualization. The visual implementation method comprises the following steps: firstly, establishing a three-dimensional space division grid according to the dimension of a range of all member models contained in each BIM model; then organizing the member models in each grid block to combine and generate a precise model of a single grid block; recombining and generating a simplified model with consistent shape and appearance according to vein use conditions by using the obtained precise model; and finally, loading and displaying the precise model or the simplified model of the BIM model in the three-dimensional large scene according to a selected display mode and an index number. According to the visual implementation method disclosed by the invention, on the basis of guaranteeing the application of the BIM model, all members of each BIM model are divided according to the three-dimensional space division grid by means of the space grid idea, and the members in the grid are merged to reduce the number of the models of the index and improve the loading efficiency of the BIM model.
Owner:CHONGQING SURVEY INST
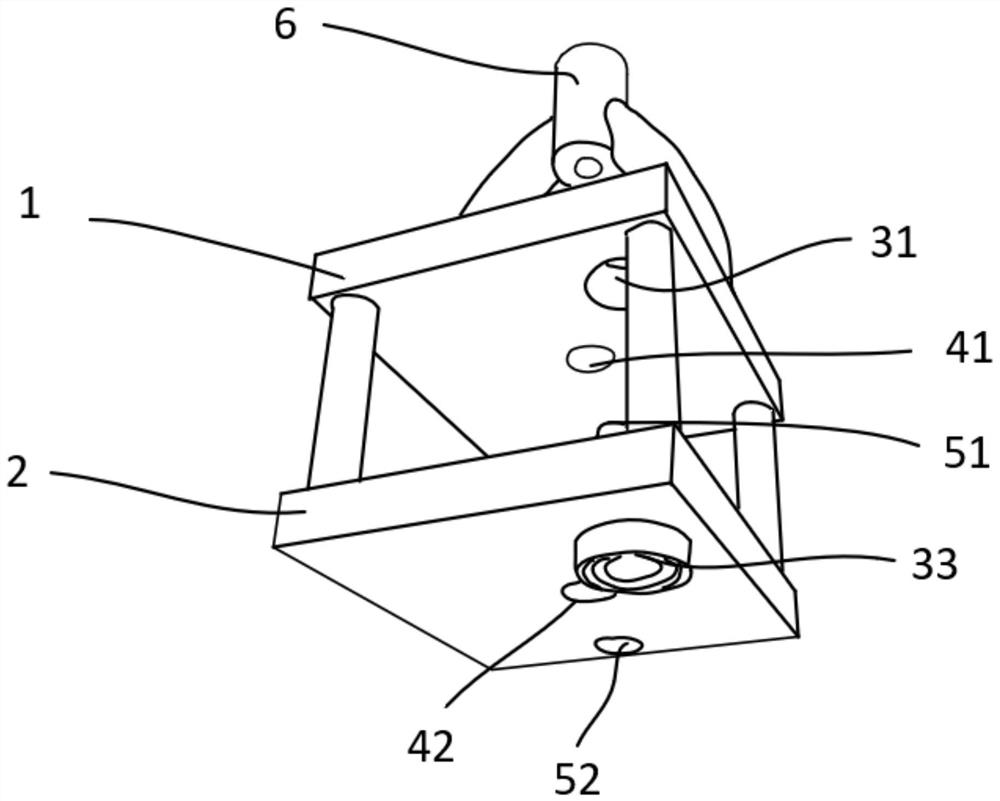
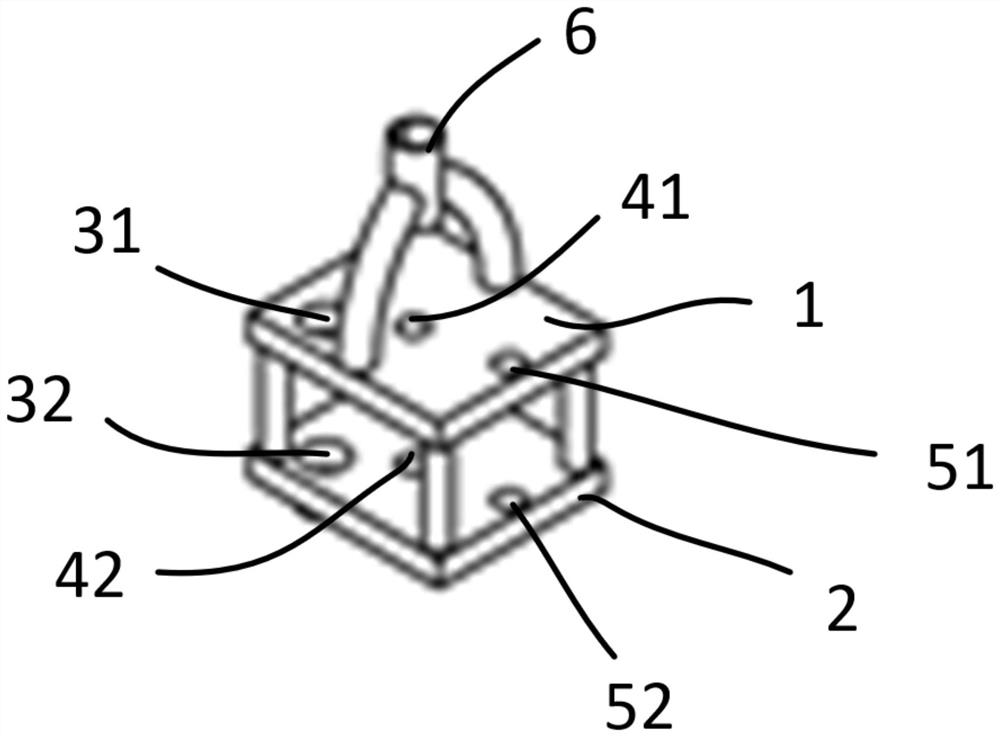
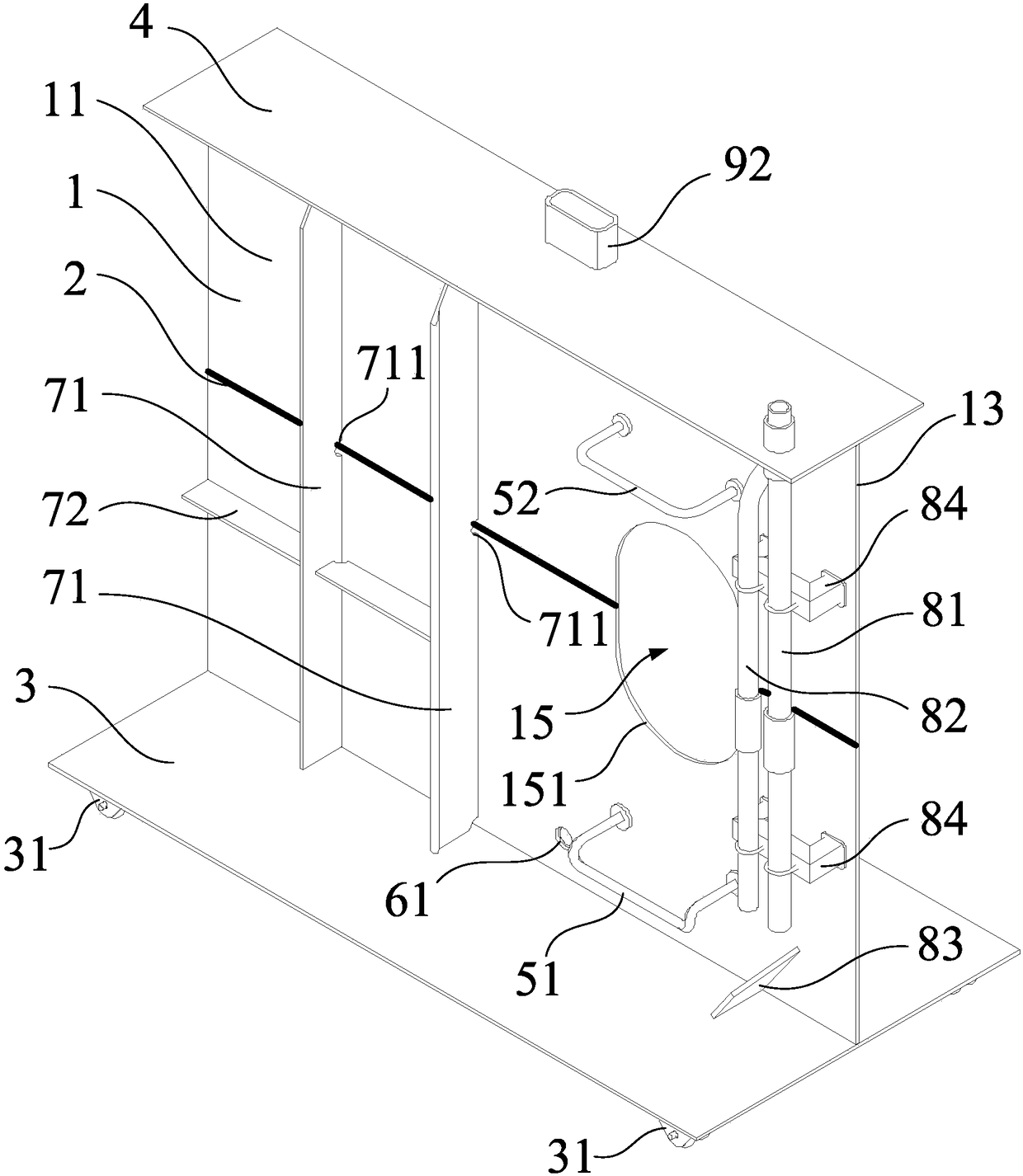
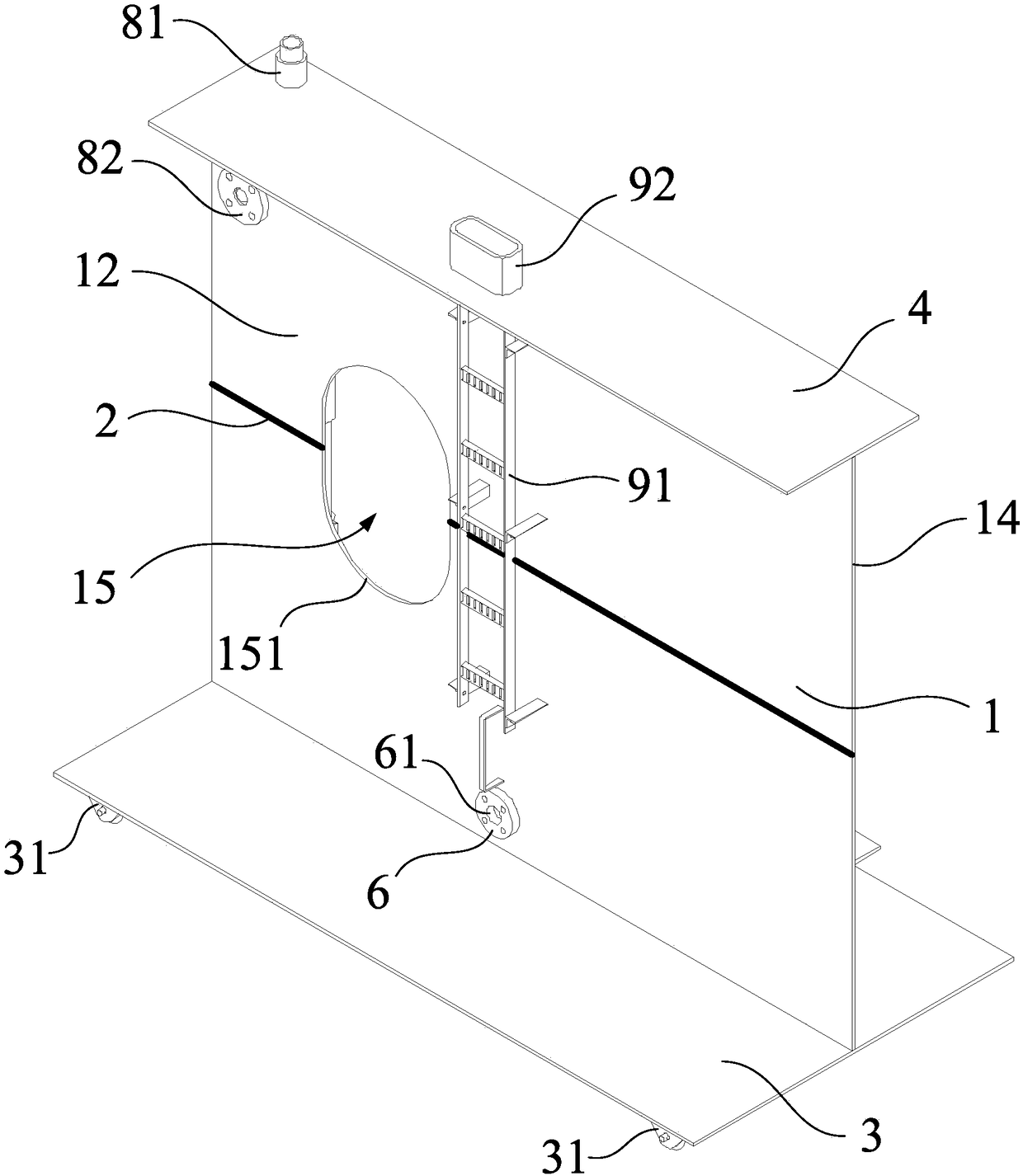
Integrated stepping type electrode support
PendingCN114305427AStable structureReduce in quantityDiagnostic recording/measuringSensorsEngineeringElectrode array
The invention provides an integrated stepping type electrode support which comprises a first supporting plate, a second supporting plate and a stepping assembly, the first supporting plate is parallel to the second supporting plate, a first through hole is formed in the first supporting plate, a second through hole is formed in the second supporting plate, and the stepping assembly penetrates through the first through hole and the second through hole. The stepping assembly is used for conducting stepping driving on the electrode support. The electrode array is fixedly connected with the stepping assembly, so that when the stepping assembly is adjusted to conduct stepping driving on the electrode support, the electrode array and the electrode support can advance at the same time. Furthermore, the electrode bracket is also provided with a placement part for accommodating the optical fiber head, so that the electrode bracket is more stable in structure and not easy to break in the experiment process.
Owner:SHENZHEN INST OF ADVANCED TECH CHINESE ACAD OF SCI
Instant heating type steam boiler
InactiveCN101101110AShorten the lengthSimple structureSteam generation heating methodsFlash steam boilersEngineeringSteam generation
The invention relates to an instantaneous heating type steam boiler, comprising: heater with connecting end, water pipe with water inlet and steam outlet, main body with heater and water pipe, wherein the heater and water pipe are built in the main body, and the connecting end, water inlet and steam outlet are all bare outside the main body. The heater and water pipe are built in the main body to avoid the problem of the existing technique that it is essential that complicated flow pipeline is set on the main body in order to increase the steam production. The mold structure of heating type steam boiler and manufacturing process is simple.
Owner:韩京姬
Command word control method and equipment
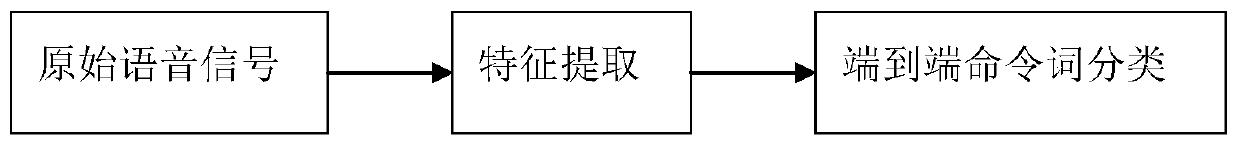
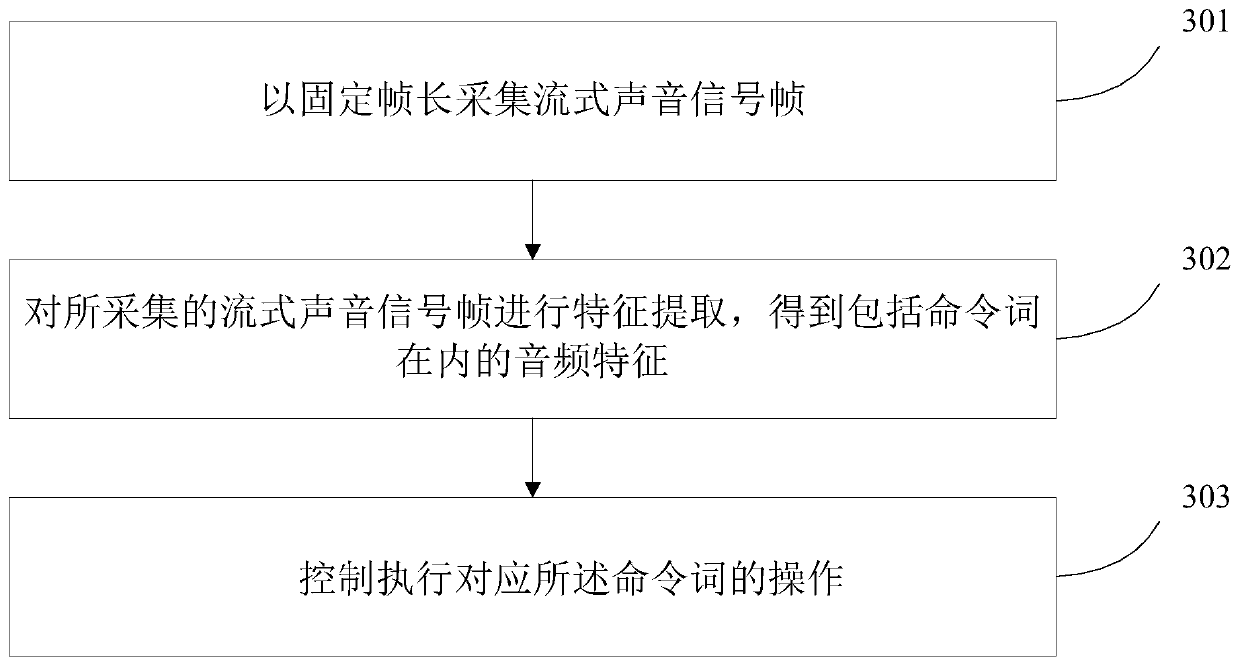
ActiveCN110556099AReduce the amount of data training and the number of parametersSimple model structureSpeech recognitionFixed frameSpeech recognition
The invention discloses a command word control method and equipment. According to the method, firstly, streaming sound signal frames are collected at fixed frame length; next, feature extraction is performed on the collected streaming sound signal frames to obtain audio features including command words; and operation corresponding to the command words is controlled to be executed.
Owner:MOBVOI INFORMATION TECH CO LTD
Contact network operation and maintenance method and device based on space-time distribution technology
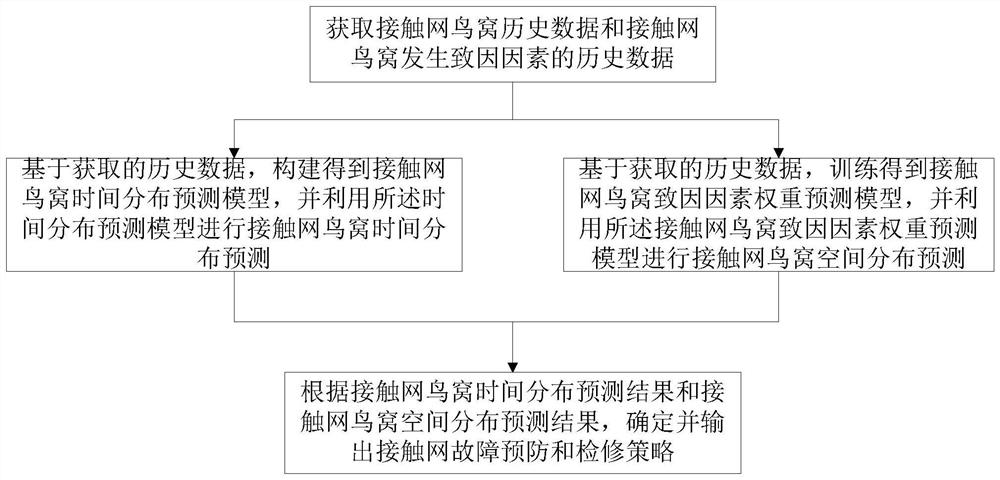
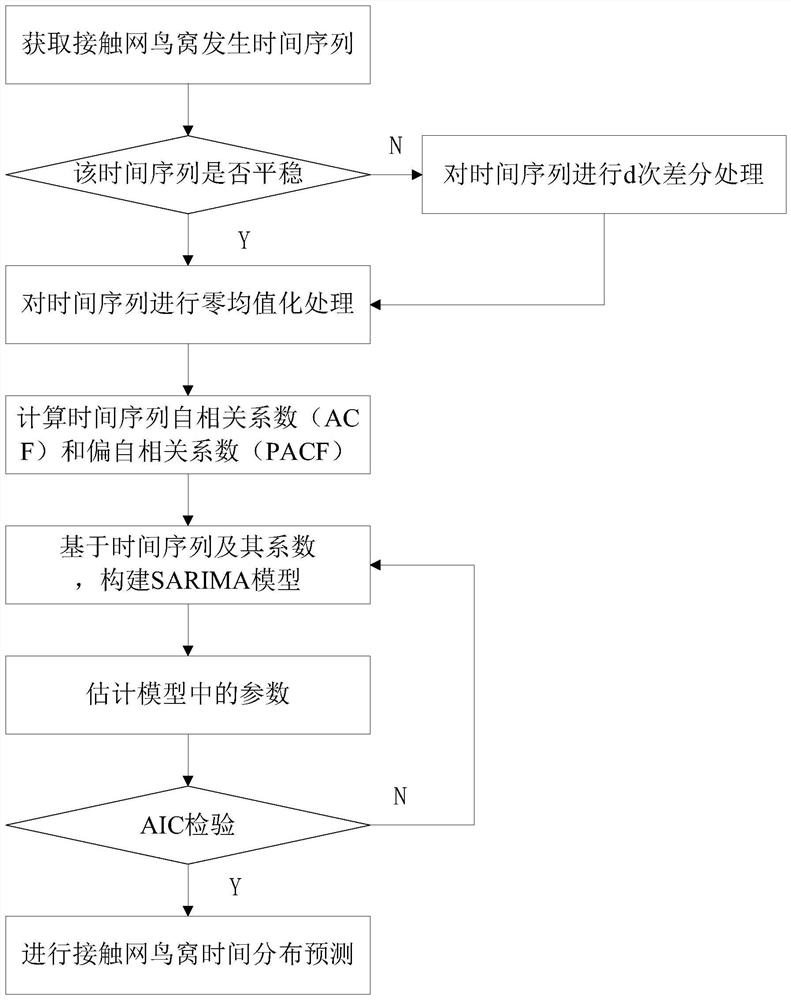
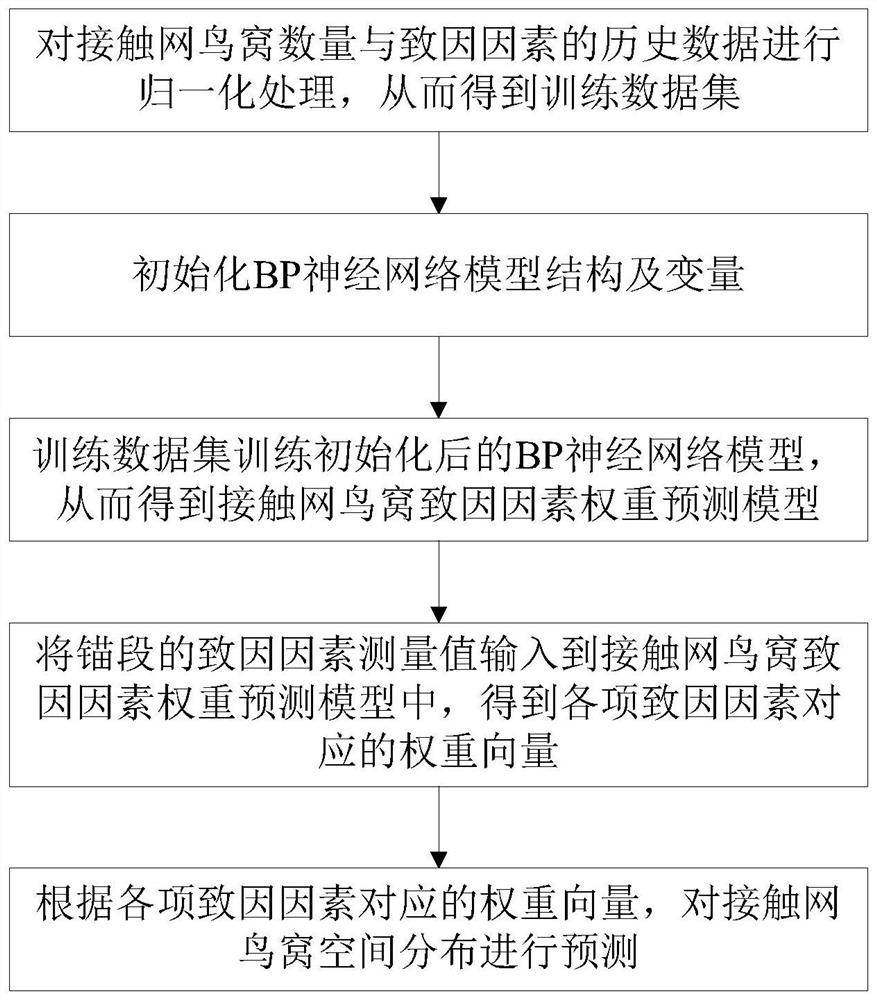
PendingCN114444798ASimple model structureFunction is easy to implementForecastingNeural architecturesTime distributionMaintenance strategy
The invention discloses an overhead line system operation and maintenance method and device based on a space-time distribution technology. The method comprises the following steps: acquiring overhead line system bird nest historical data and overhead line system bird nest generation cause factor historical data; on the basis of the acquired historical data, constructing a catenary nest time distribution prediction model, and performing catenary nest time distribution prediction by using the time distribution prediction model; based on the acquired historical data, training to obtain a catenary bird nest cause factor weight prediction model, and performing catenary bird nest spatial distribution prediction by using the catenary bird nest cause factor weight prediction model; and determining and outputting an overhead line system fault prevention and maintenance strategy according to the overhead line system bird nest time distribution prediction result and the overhead line system bird nest space distribution prediction result. According to the method, the bird nest distribution of the contact network is accurately predicted from the two dimensions of time and space, so that more comprehensive and reliable data support is provided for fault prediction and maintenance of the contact network.
Owner:CHENGDU TANGYUAN ELECTRICAL APPLIANCE
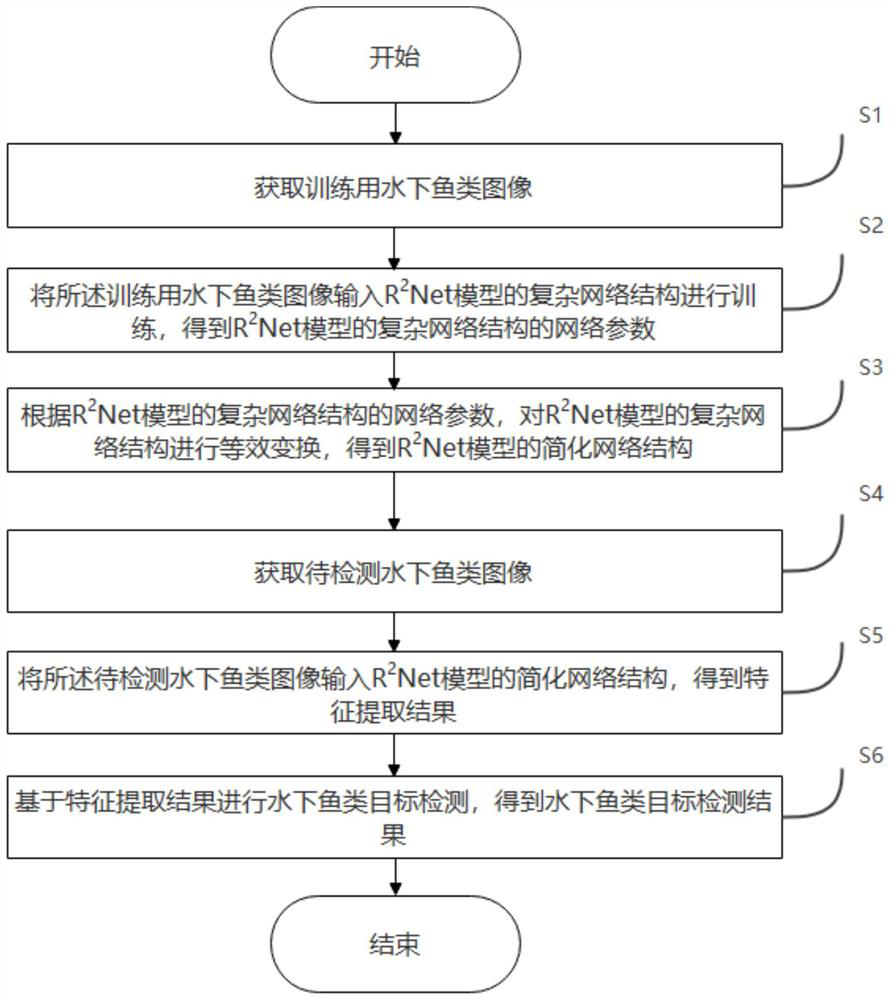
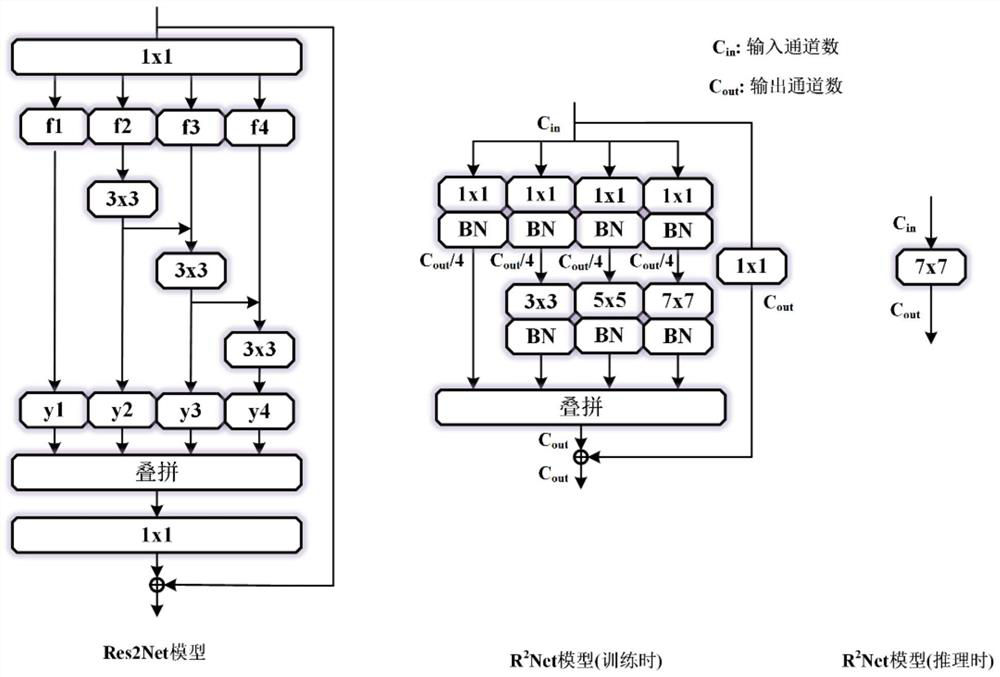
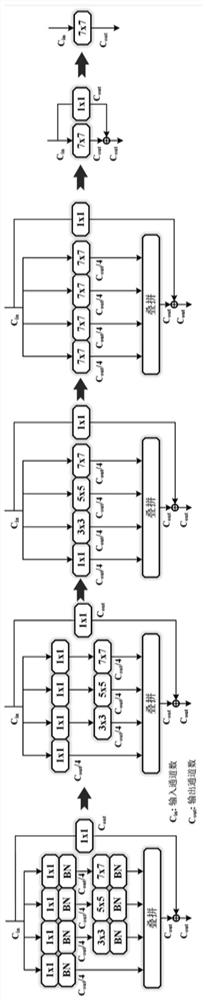
Underwater fish target detection method and device based on R2Net, and storage medium
ActiveCN113554092ASimple model structureReduce model sizeClimate change adaptationCharacter and pattern recognitionComplex networkBiology
The invention provides an underwater fish target detection method and a device based on R2Net, and a storage medium, and relates to the technical field of target detection. When underwater fish target detection is carried out, the method comprises steps: firstly, acquiring an underwater fish image for training; inputting the underwater fish image for training into a complex network structure of an R2Net model for training to obtain network parameters of the complex network structure of the R2Net model; performing equivalent transformation on the complex network structure of the R2Net model according to the network parameters of the complex network structure of the R2Net model to obtain a simplified network structure of the R2Net model; wherein the simplified network structure of the R2Net model is a Kmax * Kmax equivalent convolution kernel; obtaining a to-be-detected underwater fish image; inputting the to-be-detected underwater fish image into the simplified network structure of the R2Net model to obtain a feature extraction result; and performing underwater fish target detection based on the feature extraction result to obtain an underwater fish target detection result. According to the invention, the R2Net model is used for underwater fish target detection, and high-precision rapid underwater fish target detection is realized.
Owner:DALIAN OCEAN UNIV
Voice synthesis method and device, computer device and computer readable storage medium
ActiveCN111357015ASimple model structureSimple structureNatural language translationMachine learningSynthesis methodsEngineering
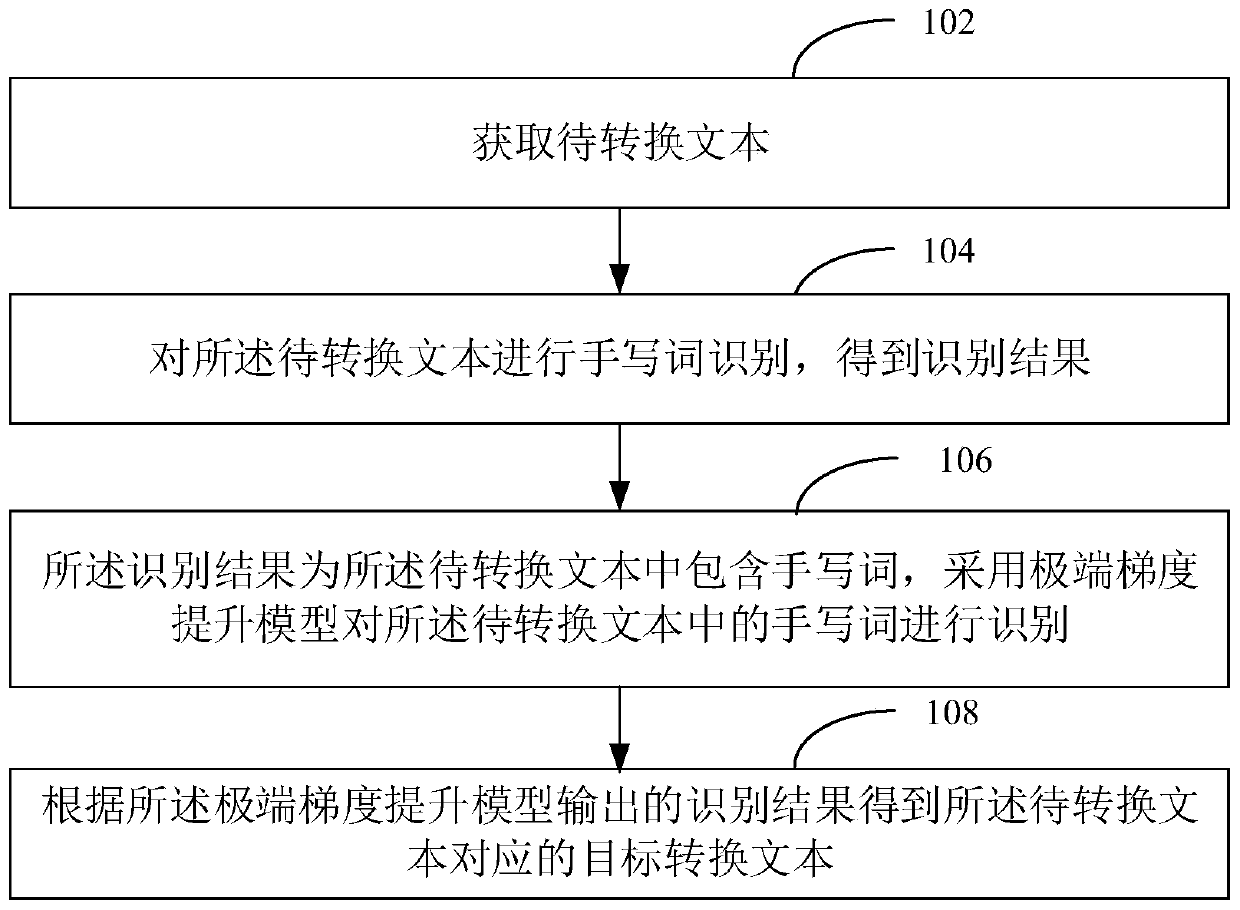
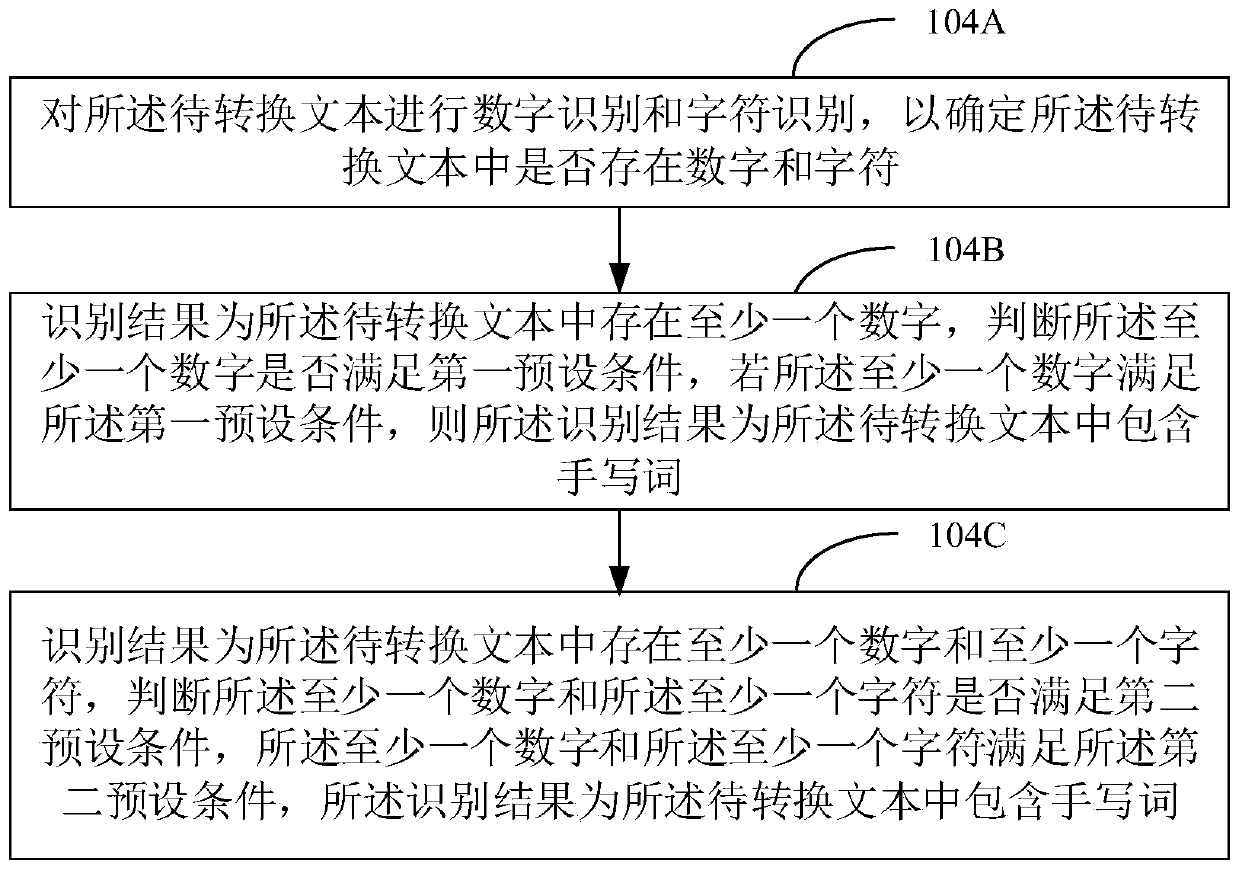
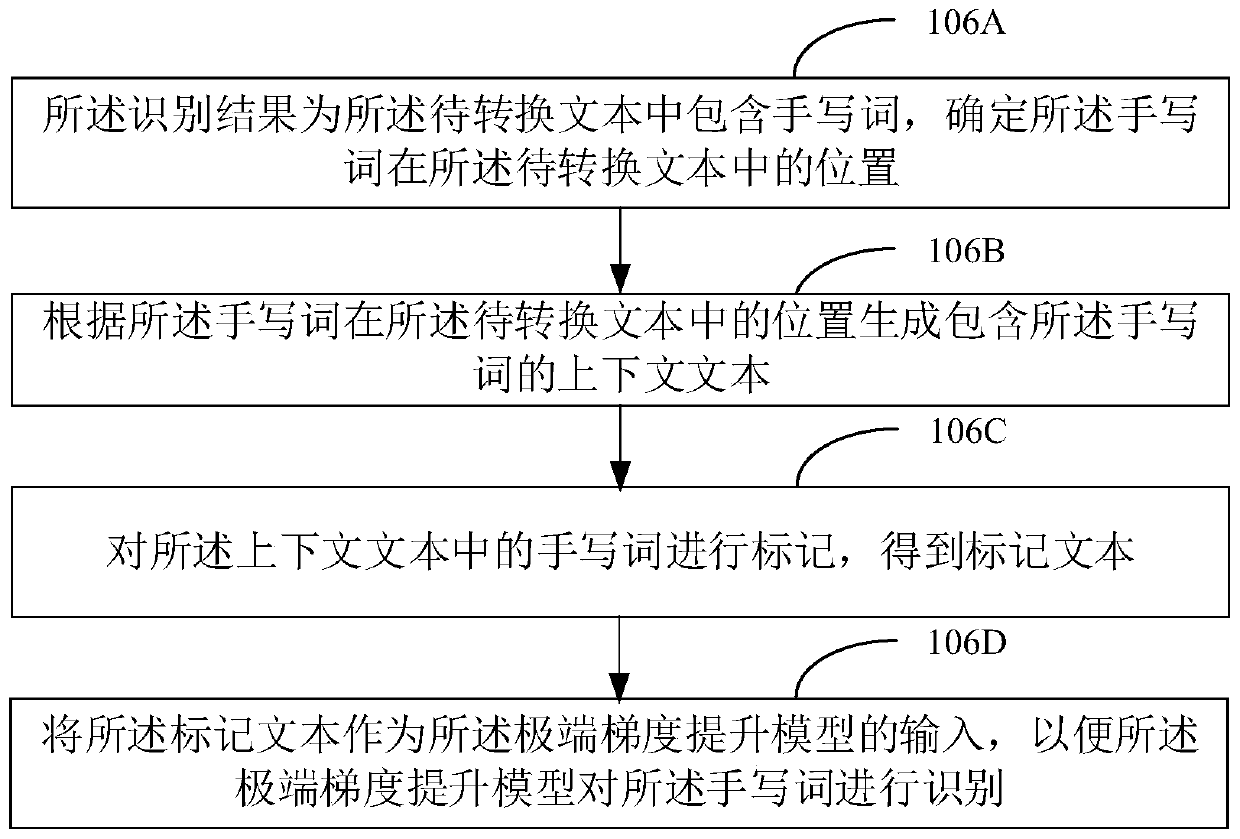
The invention discloses a text conversion method and device, a computer device and a computer readable storage medium. The method comprises the steps of obtaining a to-be-converted text; performing handwritten word recognition on the text to be converted to obtain a recognition result, wherein the recognition result is that the to-be-converted text contains handwritten words, and recognizing the handwritten words in the to-be-converted text by adopting an extreme gradient lifting model; and obtaining a target conversion text corresponding to the to-be-converted text according to a recognitionresult output by the extreme gradient lifting model. Compared with deep learning, the method is high in recognition speed and high in recognition accuracy.
Owner:北京优必选智能机器人有限公司
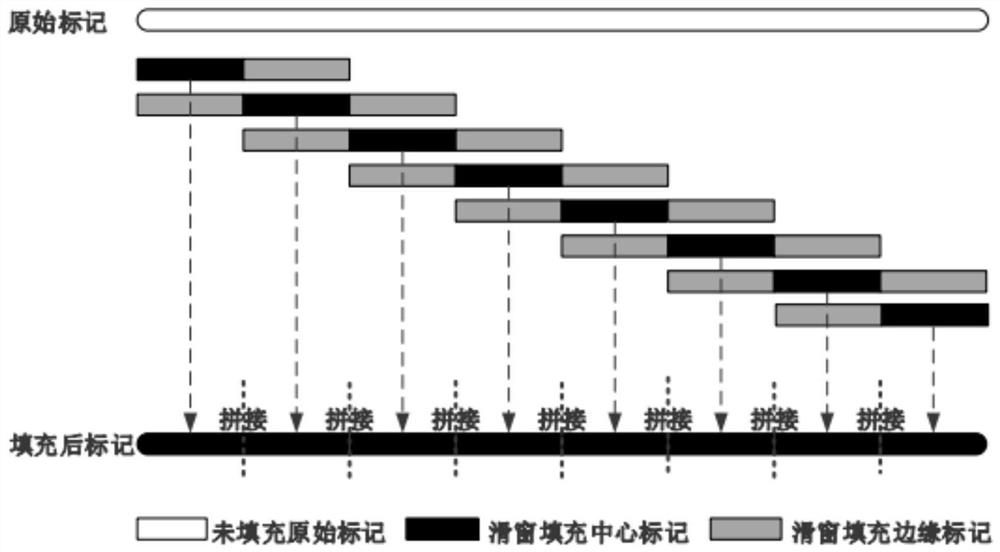
Missing mark filling method based on sliding window sparse convolution denoising auto-encoder
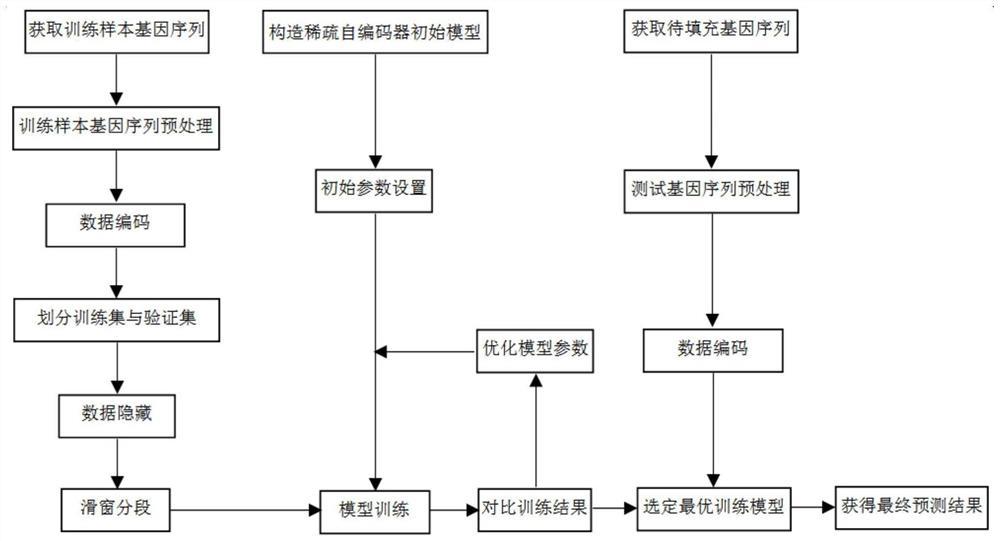
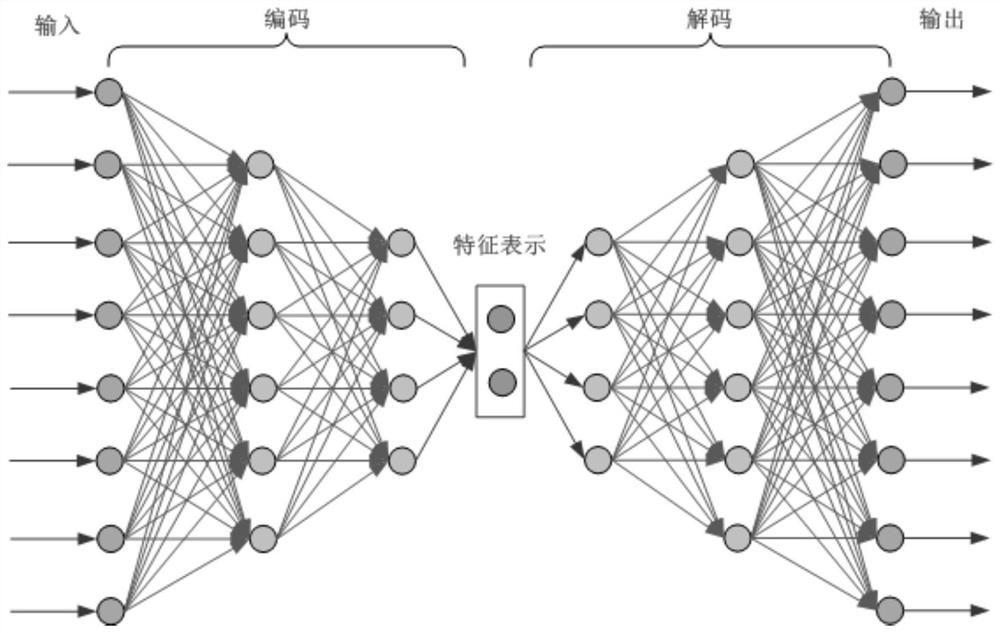
PendingCN114864004AImprove fill accuracySimple model structureBiostatisticsSequence analysisNetwork modelAutoencoder
The invention provides a missing marker filling method based on a sliding window sparse convolution denoising auto-encoder, which comprises the following steps: firstly, carrying out numerical conversion and one-hot coding on known gene data, dividing a training set and a verification set, then building a sparse convolution neural network model, carrying out missing marker filling on a gene sequence by adopting a segmented sliding window mode, and finally, obtaining a missing marker filling result; through overlapping of windows, central area prediction results with sufficient data features are obtained and spliced, and filling results of edge areas are abandoned. And then the filling precision of the missing mark is calculated, and the hyper-parameter of the neural network is adjusted according to the feedback result of the early-stage training. In practical application, gene filling results of multiple species such as corn and rice show that the filling precision of the method is remarkably higher than that of traditional algorithms such as KNN and SVD. The method is high in filling precision, has the advantages of simple model structure and high training efficiency, and has a wide application prospect in the field of gene sequence analysis.
Owner:YANGZHOU UNIV
Ship coating training model
InactiveCN109410731ASimple model structureFacilitate teaching demonstration and group trainingEducational modelsMarine engineeringWelding joint
The invention discloses a ship coating training model. The ship coating training model comprises a wall panel, the wall panel is used for simulating an uncoated surface of a ship compartment, and thewall panel has oppositely disposed front and rear surfaces and a first side, a second side, a third side and a fourth side sequentially connected end to end. The first side, the second side, the thirdside and the fourth side are all adjacent to the front surface and the rear surface, and the length direction of the wall panel is parallel to the first side and the third side. The front surface andthe rear surface each are provided with at least one weld joint, and the length direction of the at least one weld joint is parallel to that of the wall panel. The ship coating training model has a simple structure and simulates a coating environment inside a ship hull, and is very convenient to carry out teaching demonstration and group training.
Owner:上海外高桥造船海洋工程项目管理有限公司
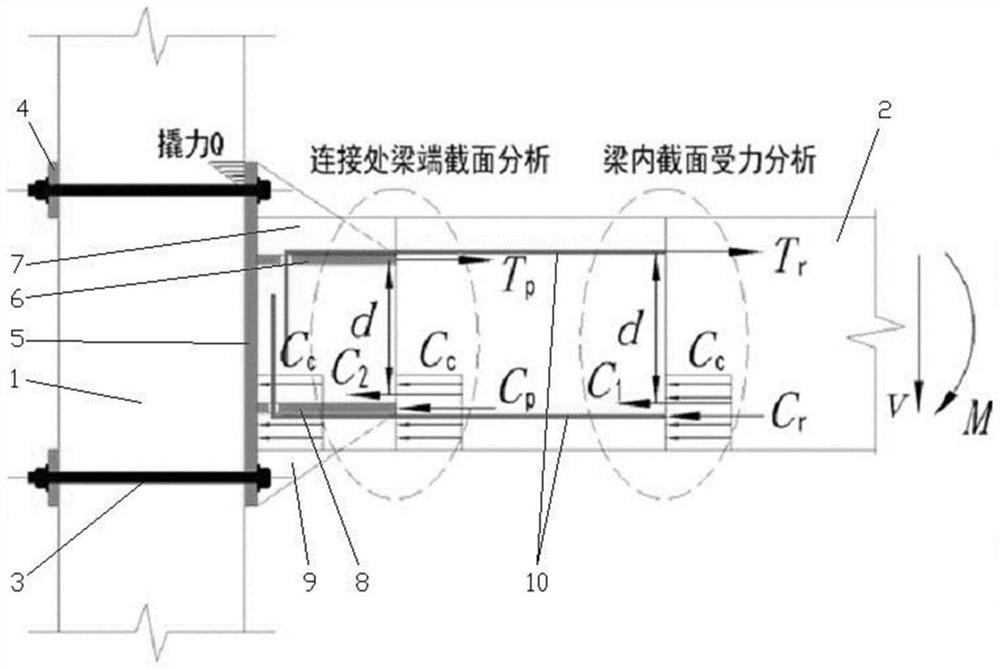
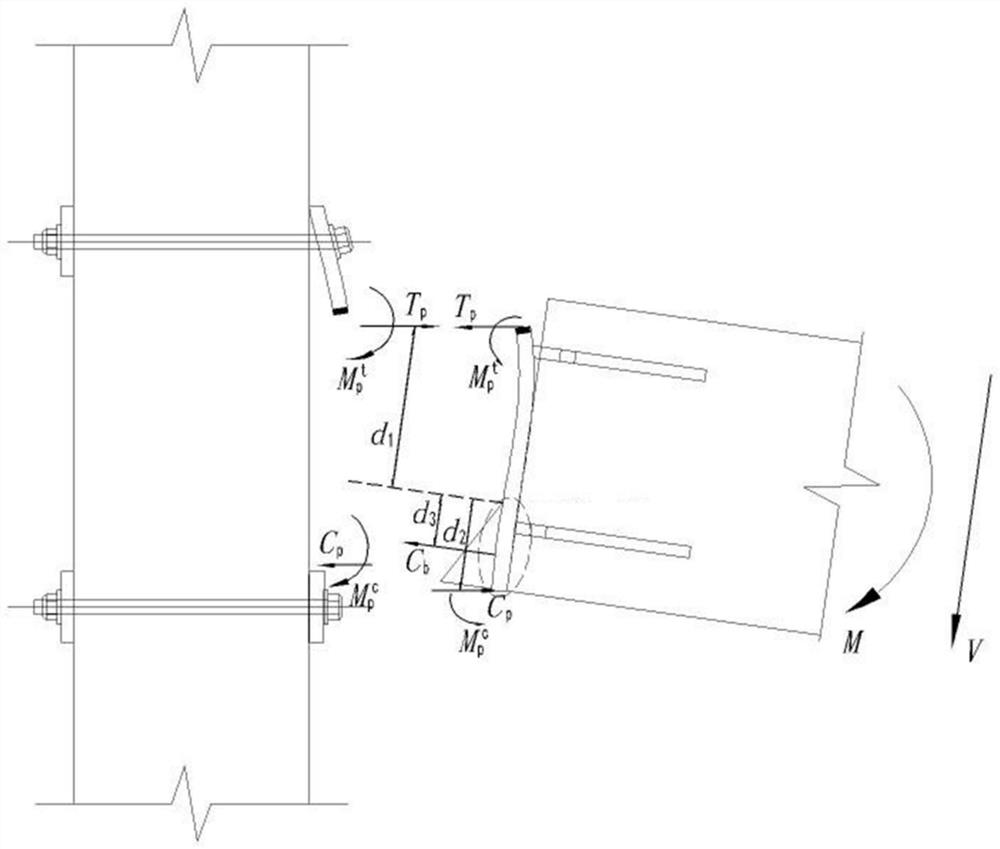
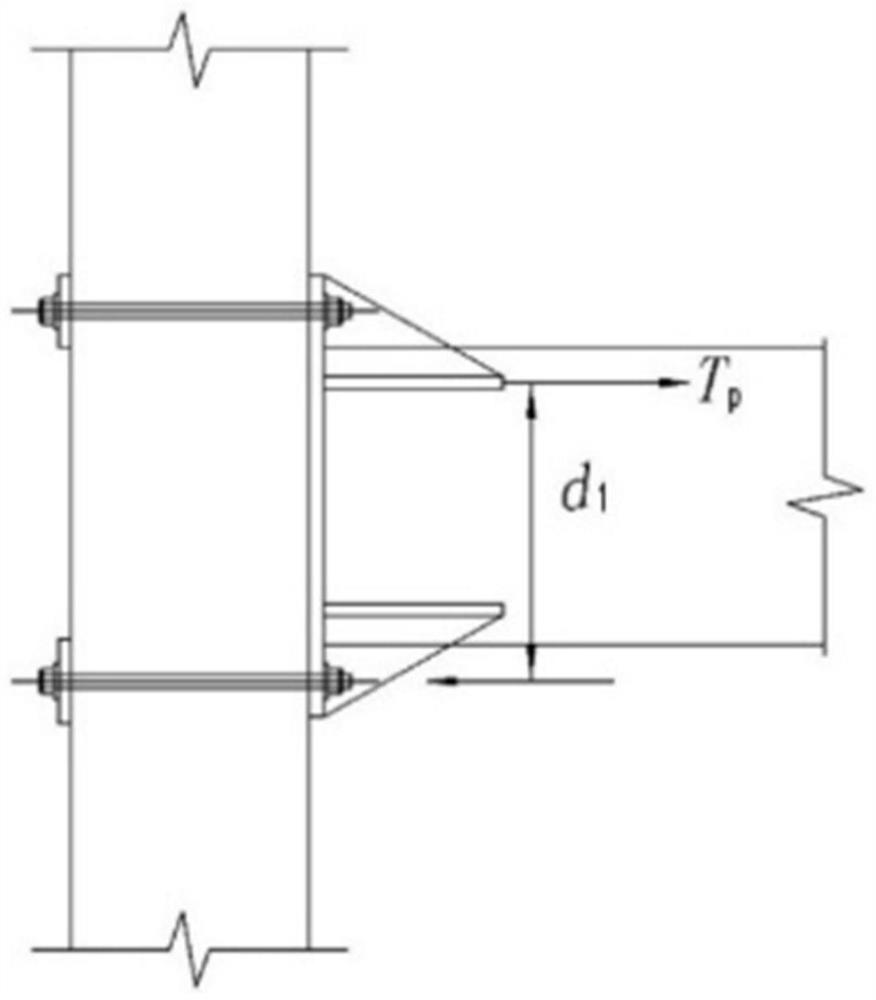
Design method of semi-rigid connection joint of low multi-layer fabricated concrete beam column
PendingCN112417552ASimple model structureEasy and fast installationGeometric CADDesign optimisation/simulationGroutEnergy consumption
The invention discloses a design method of a semi-rigid connection joint of a low multi-layer fabricated concrete beam column. The design method comprises the following steps: 1) establishing a beam column semi-rigid joint model on the basis of semi-rigid connection structure analysis; 2) analyzing the failure form of the beam-column semi-rigid joint model, and determining the flexural capacity ofjoint connection; 3) determining the flexural capacity of the beam end concrete section; and 4) determining the flexural capacity of the beam column semi-rigid node. The beam column semi-rigid jointbuilt through the design method is simple in structure, secondary concrete pouring or grouting is not needed, and rapid installation is facilitated. According to the invention, through reasonable design, the damage mode is controllable, and the safety of the connection node is ensured. Different from a traditional cast-in-place frame joint which generates plastic hinge energy consumption through abeam end, the semi-rigid joint adopted by the invention generates energy consumption through a relative corner generated by a beam column, the section of the beam end or a joint connecting piece is mainly damaged, the integrity of the column is kept, and the beam component can be replaced.
Owner:CHINA CONSTR SEVENTH ENG DIVISION CORP LTD +1
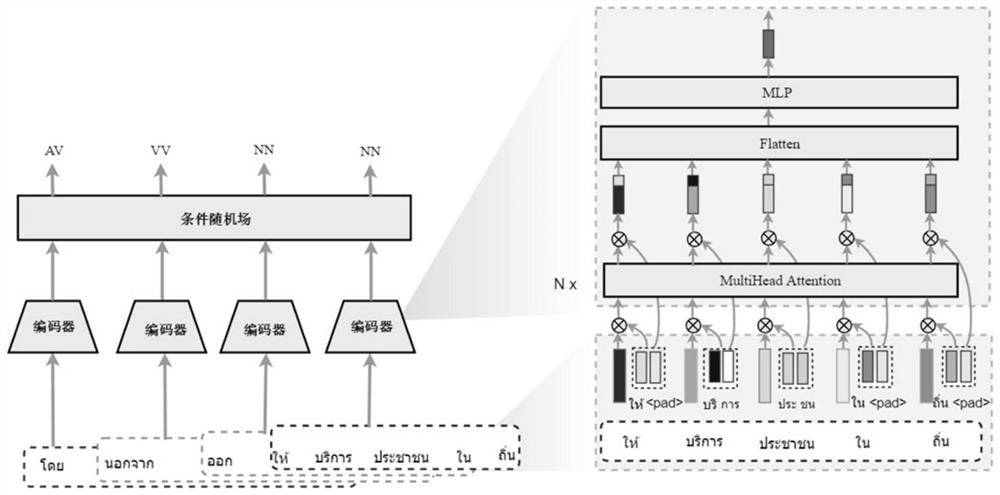
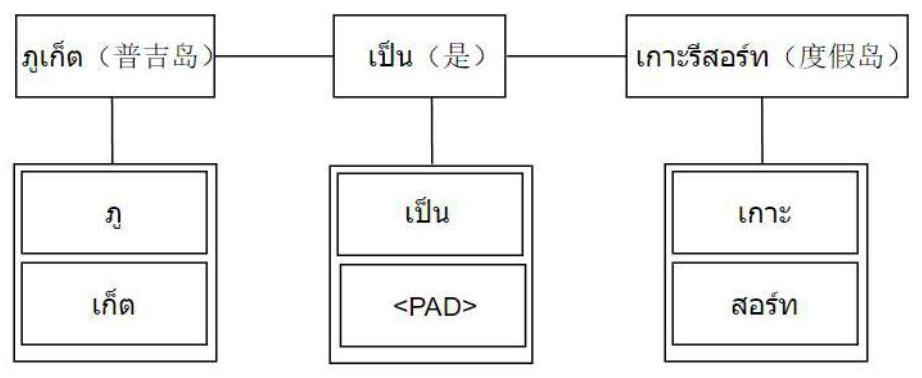
Thai and Burmese part-of-speech tagging method for fusing word-syllable pairs by using a local multi-head attention mechanism
ActiveCN113901210ASimple model structureImprove part-of-speech predictionNatural language data processingNeural architecturesConditional random fieldPart of speech
The invention relates to a Thai and Burmese part-of-speech tagging method for fusing word-syllable pairs by using a local multi-head attention mechanism, and belongs to the field of natural language processing. The method comprises the following steps: preprocessing a Thai or Burmese text data set; selecting word-syllable pair features as model input in a windowing mode; learning context features from the word-syllable pair sequence by using a local multi-head attention mechanism; and finally, modeling a part-of-speech dependency relationship through a conditional random field, and predicting a part-of-speech tag. Experimental results for part-of-speech tagging data sets of Thai and Burmese show that compared with a current optimal model, the method integrates syllables as morphological features of words, facilitates learning of context features of unknown words, and relieves the influence of wrong tagging of the unknown words on the performance of the model. Moreover, a local multi-head self-attention mechanism is adopted, so that the model can obtain richer local dependence features, and a better tagging result is obtained in a part-of-speech tagging task.
Owner:KUNMING UNIV OF SCI & TECH
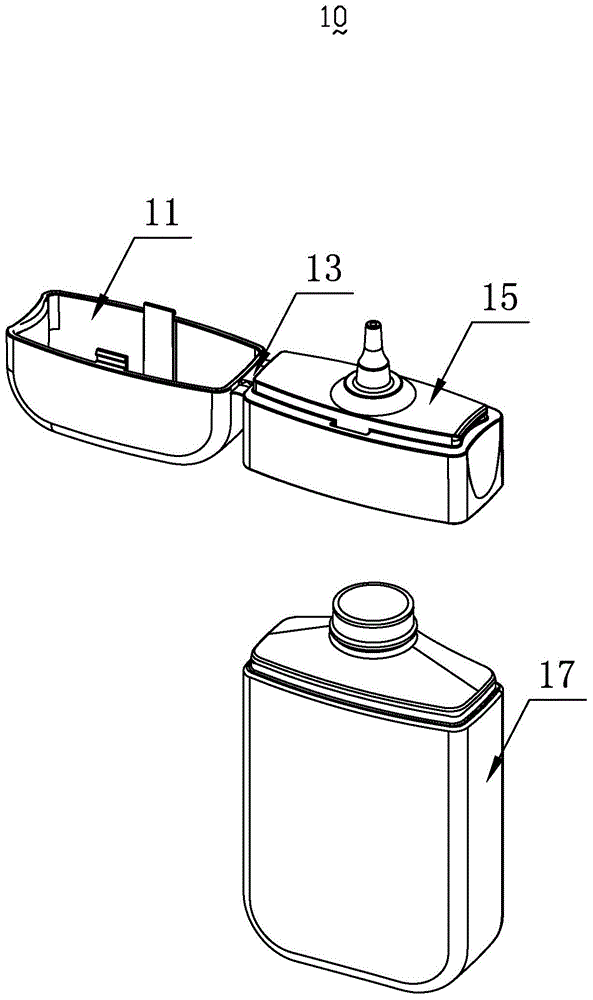
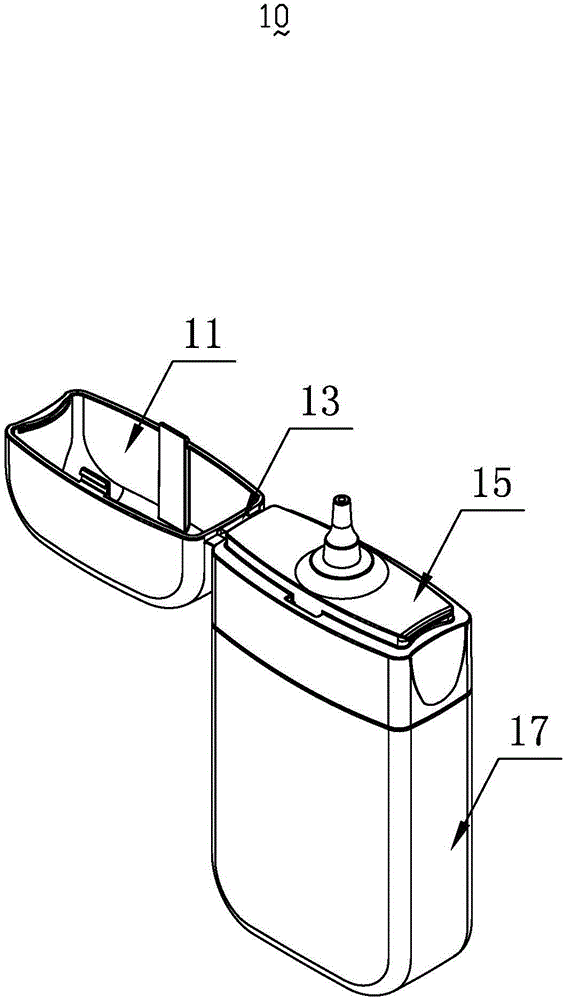
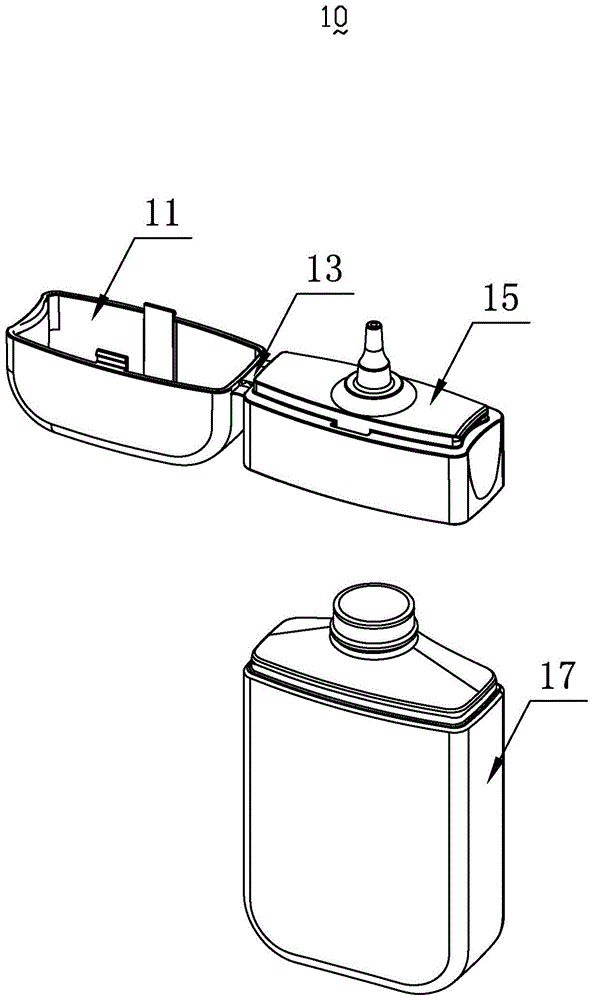
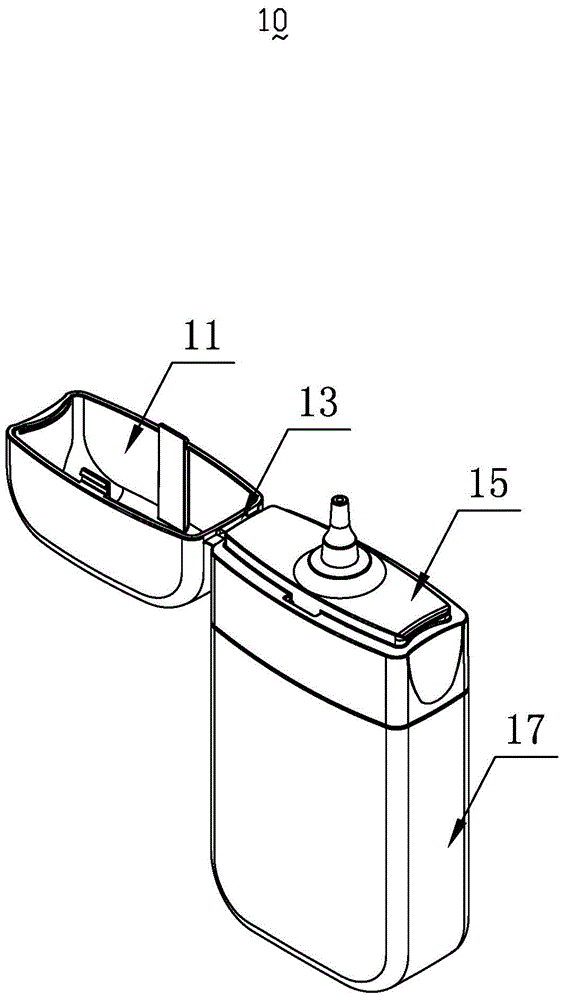
a smoke oil bottle
InactiveCN103818629BSimple model structureSimple production processBottlesContainer/bottle contructionTarBottle
The invention provides a tobacco tar bottle used for accommodating tobacco tar. The tobacco tar bottle comprises a bottle body storing the tobacco tar. The bottle body is provided with a bottle body mouth, a lower cover and an upper cover. The bottle body mouth is used for injecting the tobacco tar. The lower cover is provided with a bottle mouth, is connected to the bottle body mouth and decreases the flow diameter of the tobacco tar in the bottle body through the bottle mouth. The upper cover covers the bottle mouth. A second seal structure is provided between the lower cover and the upper cover and corresponds to the bottle mouth of the lower cover. A first seal structure is provided between the lower cover and the bottle body and corresponds to the bottle body mouth. A first clamping-connection structure is provided between the lower cover and the bottle body which are in fixed clamping connection through the first clamping-connection structure. The tobacco tar bottle is good in tightness, adaptive to various structures, simple in production process, high in production efficiency, free of safety risks, and simple in structure and has broad market prospect and high practicality.
Owner:RITCHY GRP
A model training method and related device
ActiveCN113887245BIncrease training speedEasy to adjustSemantic analysisNeural architecturesPattern recognitionMedicine
Owner:TENCENT TECH (SHENZHEN) CO LTD
Water pollution early warning method, system and equipment based on deep learning and storage medium
ActiveCN114742183ASimple model structureReduce training complexityGeneral water supply conservationCharacter and pattern recognitionMonitoring siteFeature fusion
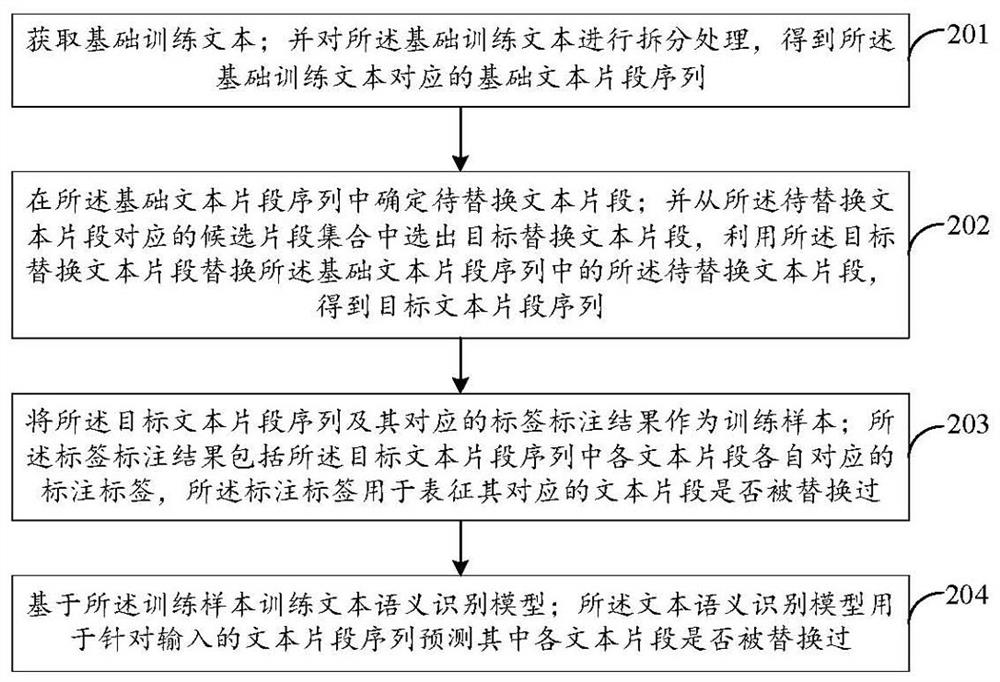
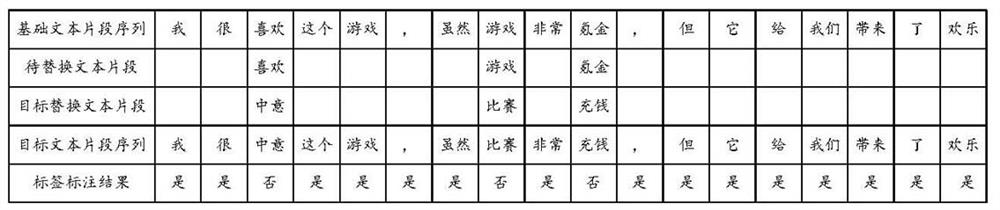
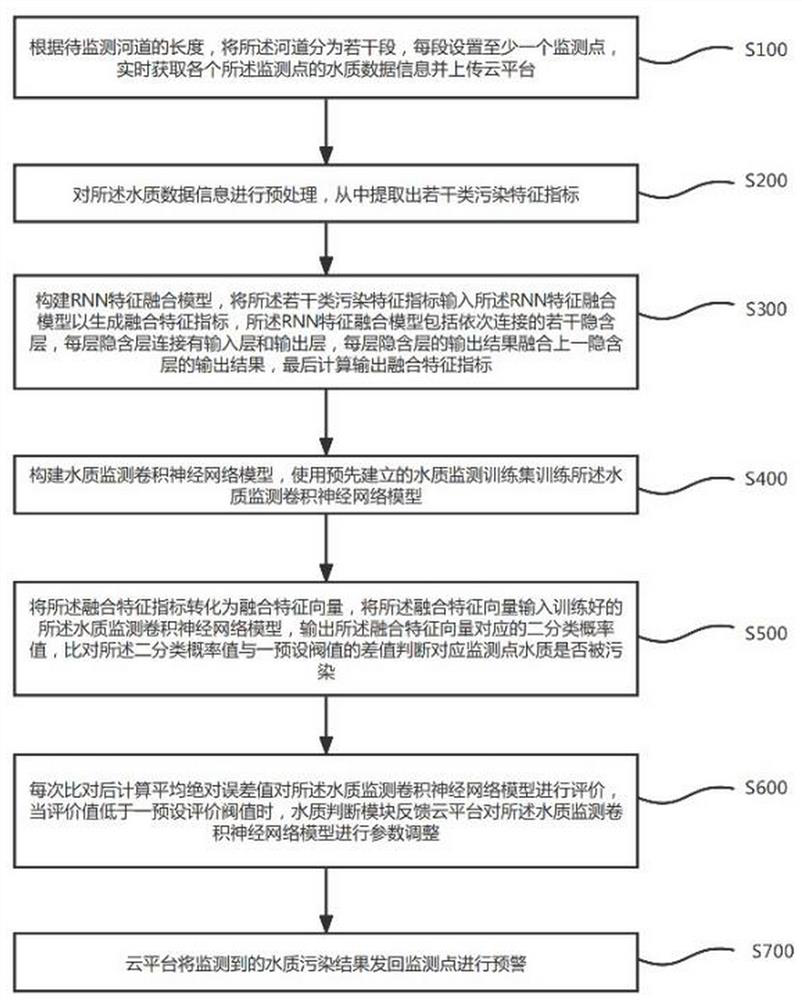
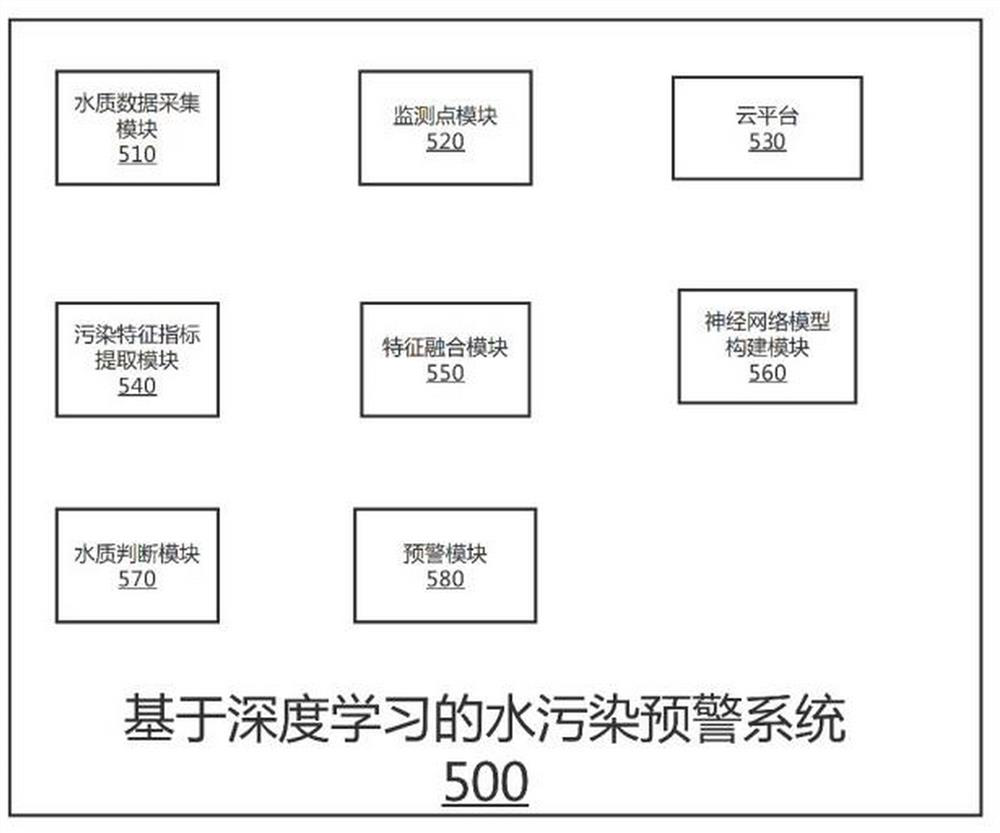
The invention discloses a water pollution early warning method, system and device based on deep learning and a storage medium, and the method comprises the steps: setting a plurality of monitoring points in a river channel, and obtaining the water quality data information of each monitoring point; extracting a plurality of pollution characteristic indexes from the water quality data information; constructing an RNN feature fusion model, and inputting the pollution feature indexes into the RNN feature fusion model to generate fusion feature indexes; constructing a water quality monitoring convolutional neural network model, and training the convolutional neural network model; converting the fusion feature index into a fusion feature vector, inputting the fusion feature vector into the trained water quality monitoring convolutional neural network model, and outputting a probability value corresponding to the fusion feature vector; comparing a difference value between the binary probability value and a preset threshold value to judge whether the water quality of the corresponding monitoring point is polluted or not; and the monitoring point module which monitors the water quality pollution carries out early warning. The water pollution monitoring efficiency can be effectively improved, and early warning can be performed in time.
Owner:广东中浦科技有限公司
a liquid bottle
InactiveCN103786942BSimple model structureSimplify the process of makingClosuresBottlesEngineeringBottle
The invention provides a liquid bottle which is used for containing liquid substances. The liquid bottle comprises an upper cap, a lower cap and a bottle body. The lower cap is located between the upper cap and the bottle body. The bottle body and the lower cap are sealed and clamped. The liquid bottle is suitable for liquid devices of different shapes and different structures, easy to process and produce, high in assembly speed, good in sealing effect, and convenient to use and carry.
Owner:RITCHY GRP
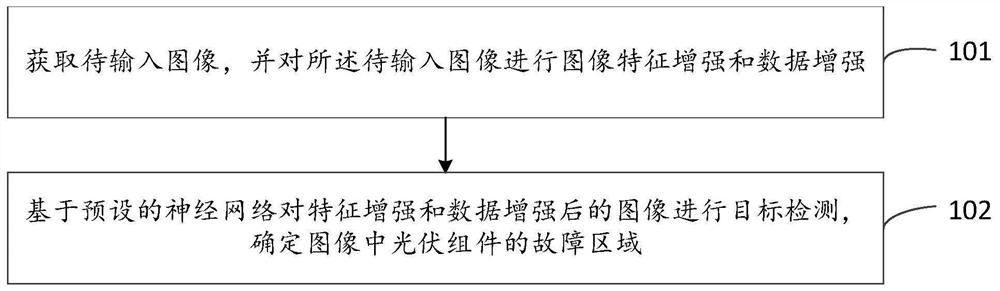
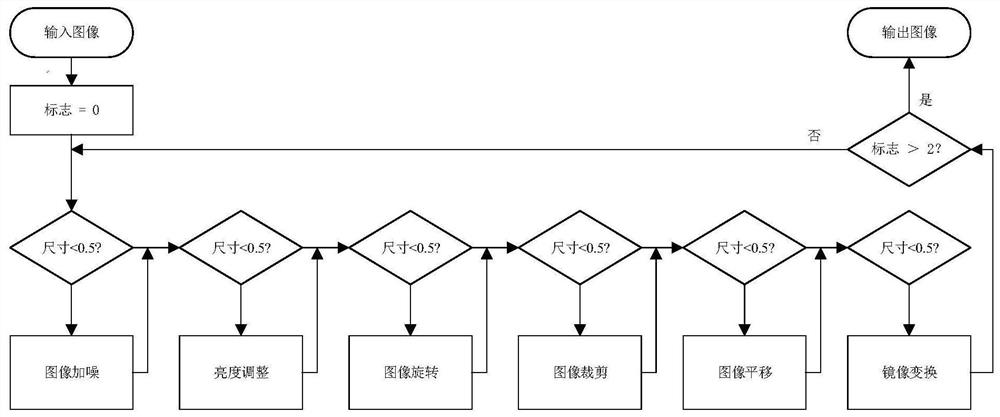
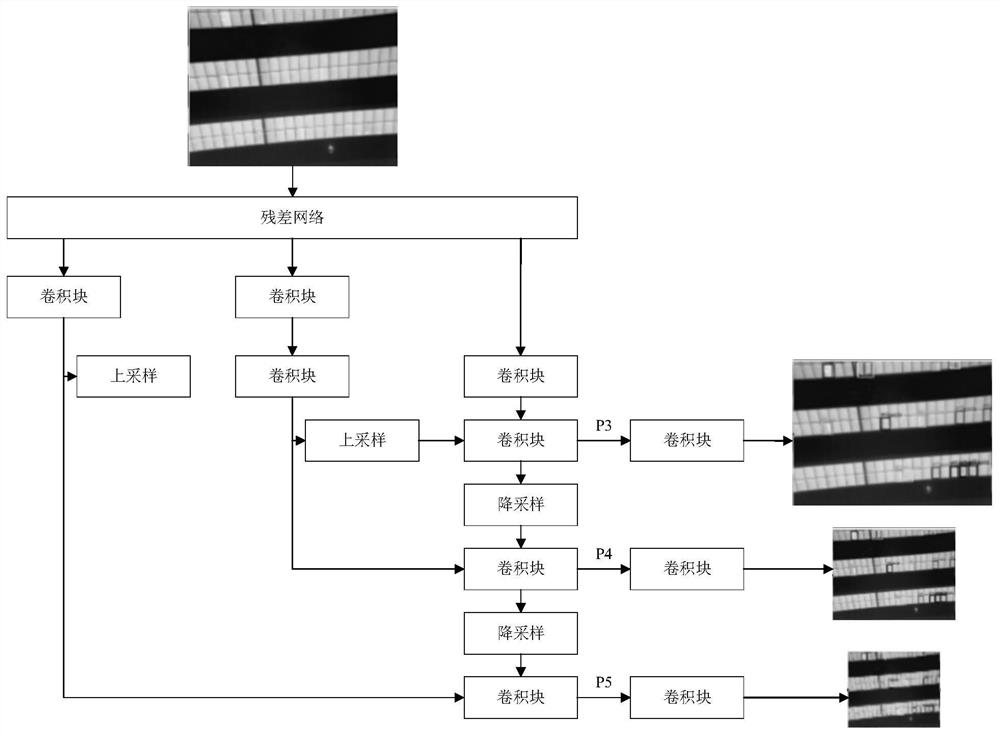
Photovoltaic module fault area image detection method and system
PendingCN113469955ASimple model structureImprove detection accuracyImage enhancementImage analysisElectrical and Electronics engineeringImage detection
The invention provides a photovoltaic module fault area image detection method and system, and the method comprises the steps: obtaining a to-be-input image, and carrying out image feature enhancement and data enhancement of the to-be-input image; and performing target detection on the image after feature enhancement and data enhancement based on a preset neural network, and determining a fault area of a photovoltaic module in the image. According to the photovoltaic module fault area image detection method and system provided by the embodiment of the invention, a novel neural network is provided to detect the photovoltaic module image, rich context information in a feature map is fused, and a distraction mechanism is combined, so that the model structure is simplified, the image detection accuracy is improved, and the detection speed is improved.
Owner:车嘉祺
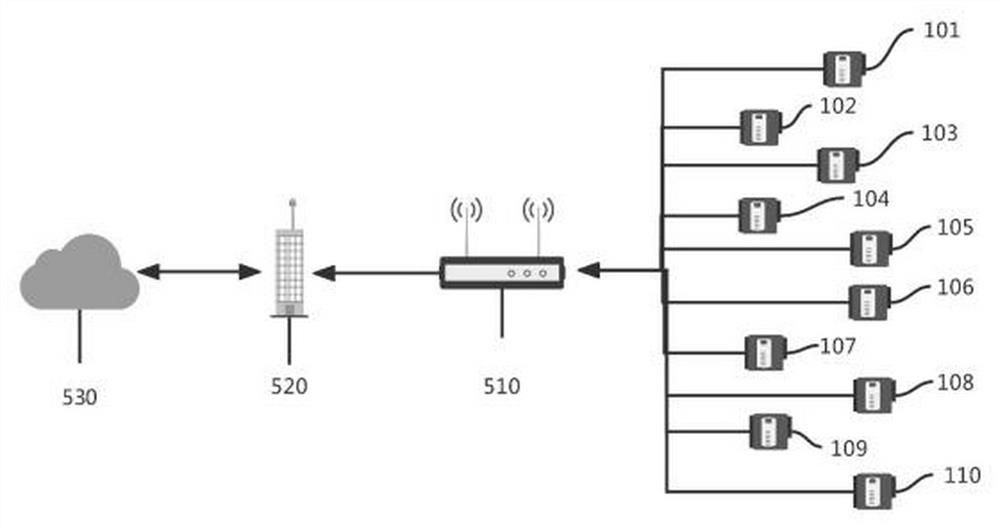
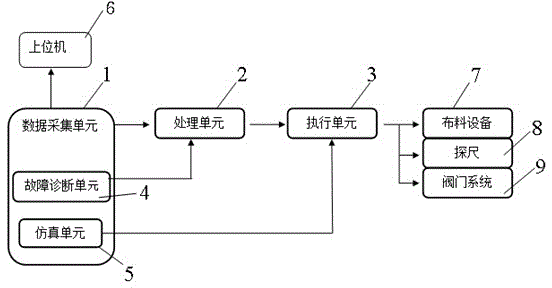
Furnace top information collecting system of blast furnace based on field bus and information collecting method thereof
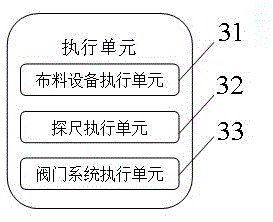
ActiveCN102591305BSimple model structureAvoid signal jamTotal factory controlProgramme total factory controlAutomatic controlProcessing
The invention discloses a furnace top information collecting system of a blast furnace based on a field bus and an information collecting method of the furnace top information collecting system of the blast furnace, which belong to the field of automatic control in iron-making. The information collecting system provided by the invention comprises an upper computer, a data collecting unit, a processing unit and an executing unit. The upper computer, the data collecting unit, the processing unit and the executing unit are connected orderly. The data collecting unit connects the information collecting points at the furnace top of the blast furnace through the field bus and transmits the collected information to the processing unit finally. The information collecting points at the furnace top of the blast furnace comprise a material distributing device, a stock rod and a valve system. The system and method of the invention simplify the structural mode of current information collection. With a field bus connecting a plurality of devices, connecting cables and costs for installation and maintenance are saved. Meanwhile, the system avoids signal blockage caused by gathering of hundreds of detection signals of detection points and control points at the inlet of the control system, thereby increasing the reliability of the system.
Owner:ANHUI MASTEEL ENG & TECH GRP
Full range temperature compensation method for semiconductor gas sensor
ActiveCN110220945BSimple model structureReduce difficultyMaterial resistancePhysical chemistryGas concentration
The invention discloses a full-scale temperature compensation method for a semiconductor gas sensor, belongs to the technical field of gas sensor detection, and solves the defect that the existing semiconductor gas sensor cannot be used for gas concentration output. The full-range temperature compensation method of the present invention includes the following steps: S1. Obtain the current ambient temperature and the resistance value of the semiconductor gas sensor in the current use environment, and determine the current normalization coefficient based on the calibrated resistance value of the semiconductor gas sensor; S2, search the upper and lower temperature values similar to the current ambient temperature in the pre-established temperature compensation database, and use the linear interpolation method to generate the normalization coefficient and gas concentration comparison table under the current ambient temperature condition; S3, generate in step S2 Look up the upper and lower gas concentrations that are similar to the current normalization coefficient in step S1 in the normalization coefficient and gas concentration comparison table, and use the linear interpolation method to calculate and obtain the gas concentration under the current ambient temperature condition.
Owner:GOLDCARD HIGH TECH
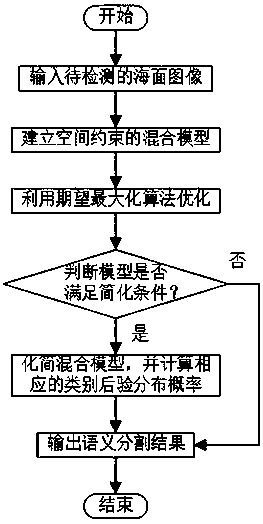
Sea surface image semantic segmentation method based on space constraint hybrid model capable of being simplified
PendingCN110363777AImprove accuracySimple model structureImage enhancementImage analysisColor imageExpectation–maximization algorithm
The invention discloses a sea surface image semantic segmentation method based on a space constraint hybrid model capable of being simplified. The sea surface image semantic segmentation method comprises the following steps: (1) inputting a sea surface color image to be detected; (2) supposing that the sea surface image has three main semantic regions including sky, coast / haze and seawater and a potential obstacle region, and establishing a spatial constraint hybrid model according to the three main semantic regions; (3) optimizing the spatial constraint hybrid model by using an expectation maximization algorithm (EM); (4) calculating a KL distance (Kullback-Leibler divergence) of Gaussian distribution of sky, coast / haze categories, and if the KL distance is smaller than a set threshold value, simplifying the spatial constraint hybrid model; and (5) outputting a sea surface image semantic segmentation result. According to the method, semantic segmentation can be effectively carried outon the sea surface image, and the method has the advantages of being high in speed and good in robustness.
Owner:SHANGHAI UNIV
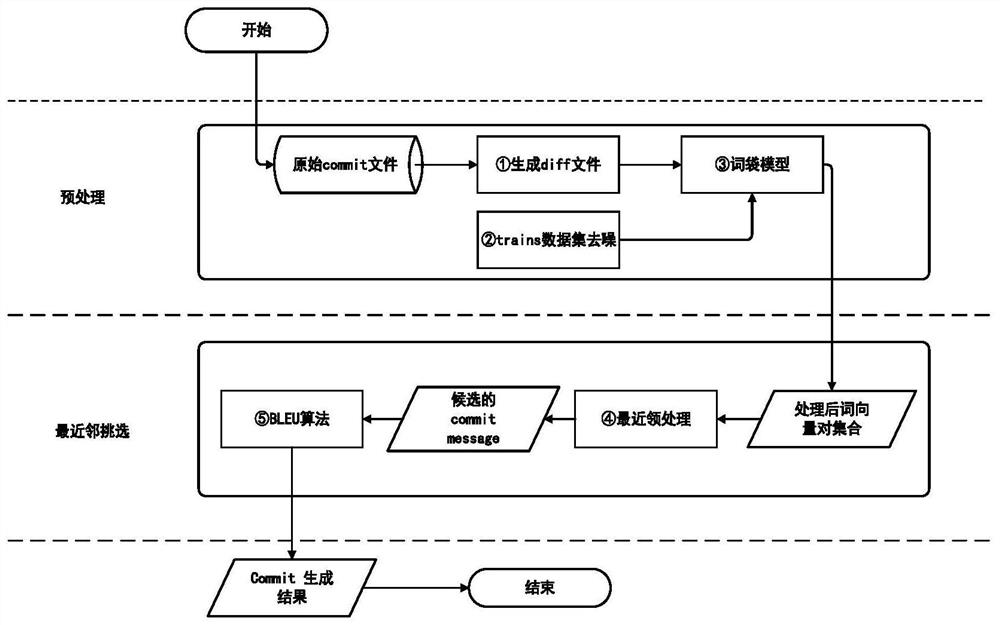
A Method for Automatically Generating Code Change Log Based on Nearest Neighbor Algorithm
InactiveCN111090460BAdd preprocessing stageSimple model structureProgram documentationAlgorithmNear neighbor
The invention discloses a code change log automatic generation method based on the nearest neighbor algorithm, which belongs to the field of code change log automatic generation. The method includes: preprocessing of input data, preprocessing of training set data, obtaining a set of word frequency vector pairs through bag-of-words model, calculating candidate intermediate results through KNN algorithm, calculating BLEU‑4 values, and finally obtaining output results. This method has the characteristics of simple model structure, strong interpretability, no training of the model, greatly reduced actual running time compared with NMT, insensitivity to noise, and strong robustness.
Owner:ZHEJIANG UNIV