Patents
Literature
105 results about "Genetic systems" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
Biological control
InactiveUS20030213005A1Not affectAvoidance of sterilisationClimate change adaptationOther foreign material introduction processesBiotechnologyBiological body
The invention relates to a non-human multicellular organism carrying a dominant lethal genetic system, the lethal effect of which is conditional, wherein the lethal effect of the lethal system occurs in the natural environment of the organism.
Owner:ISIS INNOVATION LTD
Biological Control
InactiveUS20080115233A1Avoidance of sterilisationSolution to short lifeClimate change adaptationVector-based foreign material introductionMulticellular organismOrganism
The invention relates to a non-human multicellular organism carrying a dominant lethal genetic system, the lethal effect of which is conditional, wherein the lethal effect of the lethal system occurs in the natural environment of the organism.
Owner:OXFORD UNIV INNOVATION LTD
Method for regeneration plant of tung oil tree leaf
ActiveCN103385168AIncrease resistanceIncrease productionPlant tissue cultureHorticulture methodsTerra firmaVernicia fordii
The invention discloses a method for regeneration plant of tung oil tree leaf, and belongs to the technology field of rapid propagation of tung oil tree. The method comprises the steps of adventitious bud induction directly by tung oil tree leaf, subculture, multiplication culture, strong seedlings culture, rooting culture, acclimatization and transplantation, which not only provides a path for rapid breeding of excellent tung oil tree plants, and lays a solid foundation for improving resistance of the tung oil tree, tung oil tree output and properties of oil quality later on, and for building a tung oil tree genetic system.
Owner:CENTRAL SOUTH UNIVERSITY OF FORESTRY AND TECHNOLOGY
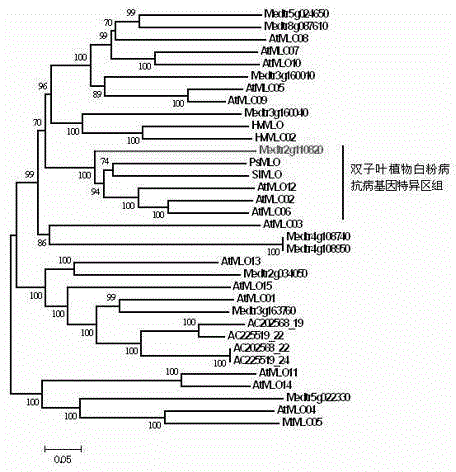
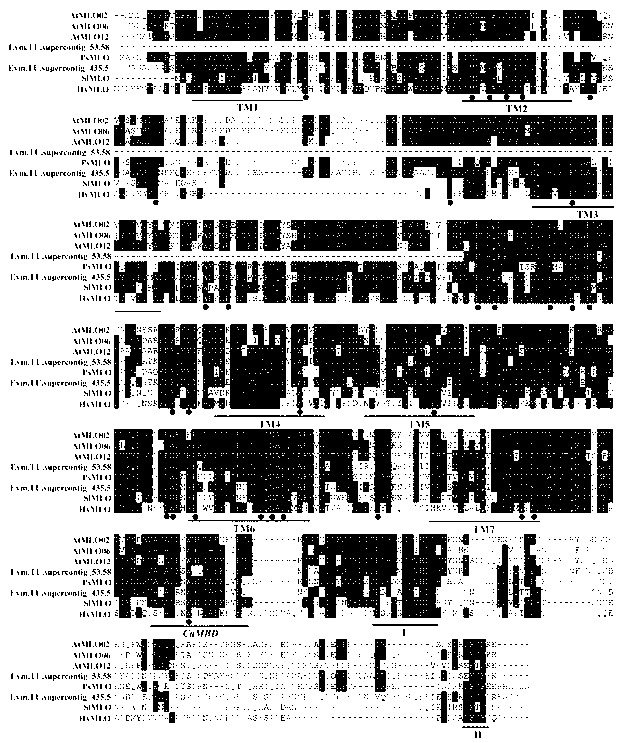
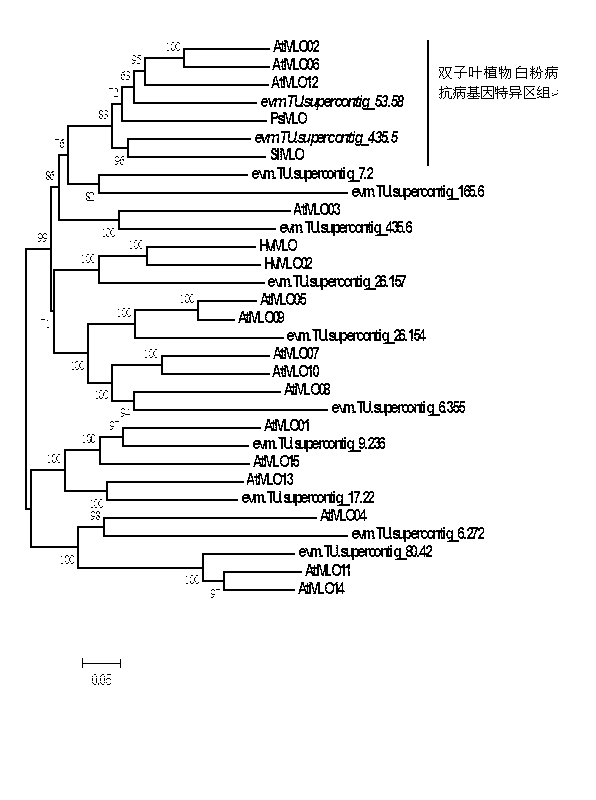
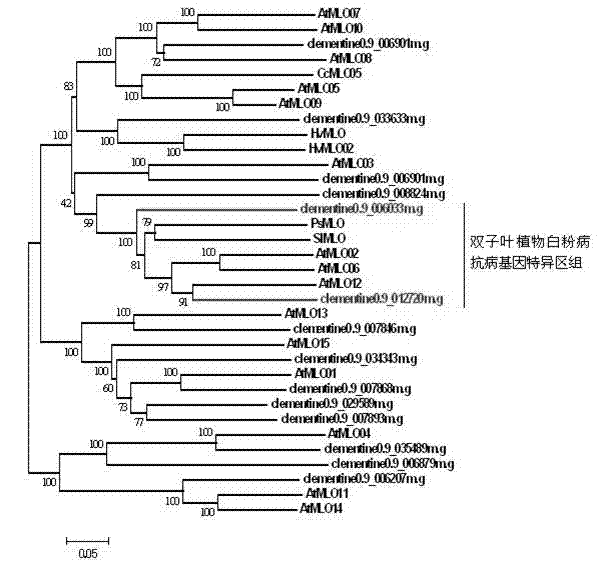
Rapid identification of powdery mildew resistance Lotus corniculatus genes utilizing comparative genomics
InactiveCN102703463AShorten digging cycleRapid identificationMicrobiological testing/measurementPlant peptidesBiotechnologyGenetics genomics
The invention relates to rapid identification of powdery mildew resistance Lotus corniculatus genes utilizing comparative genomics, relates to knowledge of subjects such as plant comparative genomics, genetics and bioinformatics, and belongs to the technical field of plant biotechnology. The rapid identification of powdery mildew resistance Lotus corniculatus genes utilizing the comparative genomics mainly includes the steps of firstly, downloading full genomic sequences of Lotus corniculatus and collecting MLO (mildew resistance locus o) genes; secondly, identifying the MLO genes; thirdly, identifying phylogenetic relationship of the MLO genes; and fourthly comparing the MLO powdery mildew resistance genes. By the rapid identification, mining cycle of the powdery mildew resistance genes is shortened, and the powdery mildew resistance genes can be identified quickly. Corresponding co-separation functional marks (SNP (single nucleotide polymorphism), Scar and the like) can be developed through candidate powdery mildew resistance genes identified, the rapid identification is also available for molecular marker-assisted selection of the powdery mildew resistance genes, and accuracy is high. The rapid identification can also be used with other molecular markers for resistance genes to create multiresistance breeding materials, and accordingly breeding period is shortened, breeding efficiency is improved, and basis for elaborating powdery mildew resistance Lotus corniculatus molecular mechanisms is laid.
Owner:常熟市支塘镇新盛技术咨询服务有限公司
Application of bioinformatics in fast identifying powdery mildew resistance gene of carica papaya l
InactiveCN102719448AShorten digging cycleRapid identificationMicrobiological testing/measurementPlant peptidesBiotechnologyPapaya family
The invention relates to fast identification of a powdery mildew resistance gene of carica papaya l, relating to subject knowledge including plant comparative genomics, genetics, bioinformatics and the like, belonging to the field of plant biotechnology sciences. Application of the invention mainly comprises the steps: 1) downloading complete genome sequence of the carica papaya l and acquiring MLO-type genes; 2) identifying the MLO-type genes; 3) researching phylogenetic relationships of the MLO-type genes; and 4) comparing MLO-type powdery mildew resistance genes. The invention has the advantages that discovery period of the powdery mildew resistance gene of the carica papaya l is effectively shortened, which assists in fast identification of the powdery mildew resistance gene; that identified candidate powdery mildew resistance genes are developed with corresponding co-segregation functional markers (SNP, SCAR, etc.), which can be used in fast auxiliary selection of molecular markers of the powdery mildew resistance gene with high accuracy; that other molecular markers of resistance genes can be combined to formulate multi-resistance breeding materials, which shortens breeding years and improves breeding yields; and that a foundation is laid to illustrate the powdery mildew resistance gene of the carica papaya l.
Owner:常熟市支塘镇新盛技术咨询服务有限公司
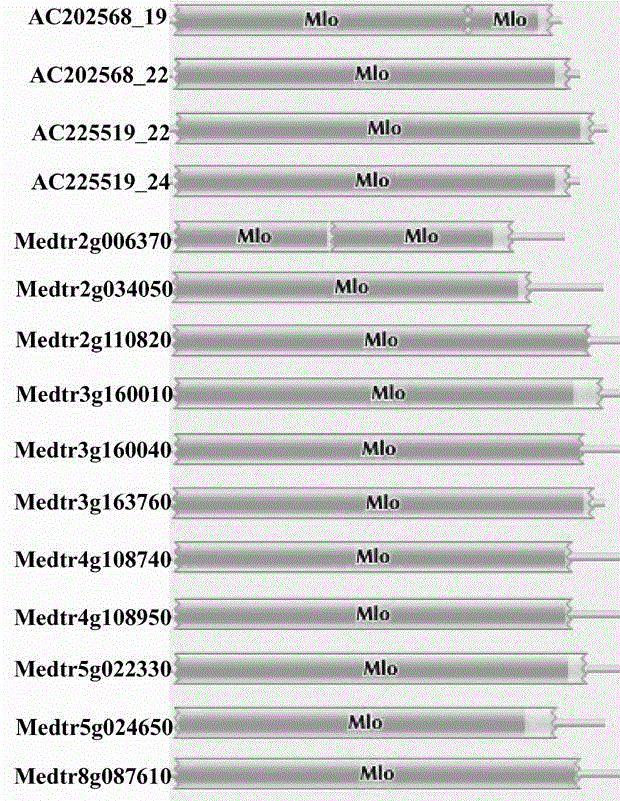
Rapid identification of powdery mildew gene of Medicago truncatula by utilizing comparative genomics
InactiveCN102719445AQuick assist selectionImprove accuracyMicrobiological testing/measurementPlant peptidesBiotechnologyRapid identification
The present invention is rapid identification of a powdery mildew gene of Medicago truncatula, relates to discipline knowledge of plant comparative genomics, genetics and bioinformatics and belongs to the technical field of plant biotechnology. The invention mainly comprises the following steps: 1) download of Medicago truncatula genome sequence and collection of an MLO gene; 2) identification of MLO type gene; 3) MLO gene phylogenetic relationship; and 4) comparison of MLO type powdery mildew gene. The invention effectively shortens a mining cycle of the powdery mildew gene of Medicago truncatula and facilitates the rapid identification of the powdery mildew gene. The identified candidate powdery mildew genes can be used to develop corresponding coseparation functional markers (SNP and SCAR, etc.), and can also be used quickly for molecular marker assisted selection of powdery mildew resistant gene, and has high accuracy. Combined with other disease resistant gene molecular markers, the identified candidate powdery mildew genes can be used in development of multiresistance breeding material, so as to shorten the breeding period and improve breeding efficiency. The invention lays foundation to elaborate molecular mechanism of powdery mildew resistance of Medicago truncatula.
Owner:常熟市支塘镇新盛技术咨询服务有限公司
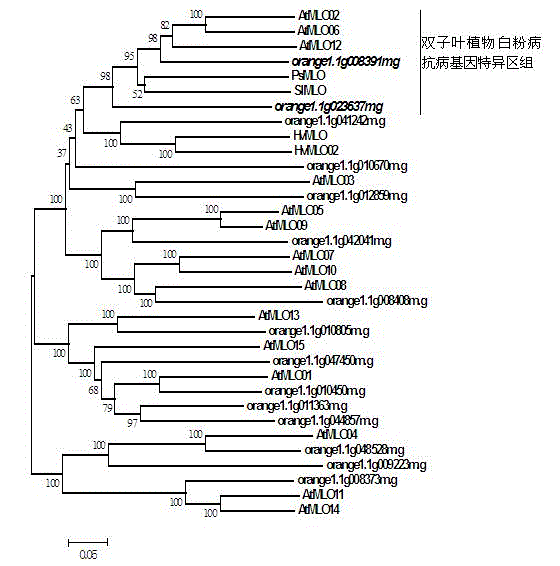
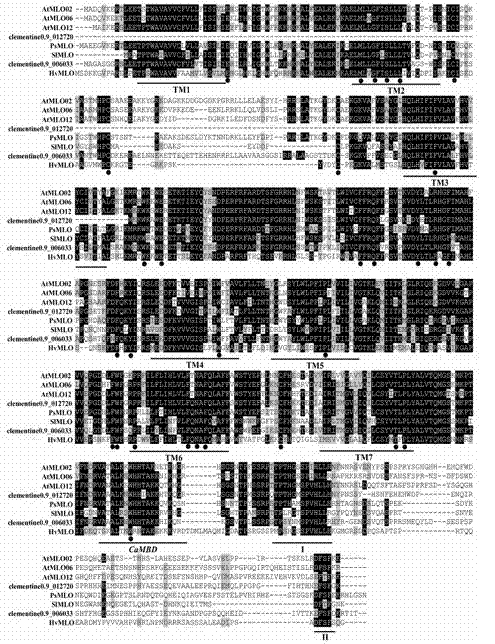
Method for rapidly identifying orange MLO powdery mildew resistance gene
InactiveCN102719446AShorten digging cycleRapid identificationMicrobiological testing/measurementPlant peptidesBiotechnologyRapid identification
The present invention is a method for rapidly identifying an orange MLO powdery mildew resistance gene, relating to plant comparative genomics, genetics, bioinformatics and other disciplines of knowledge, and belonging to the field of plant biotechnology science. The method in the invention mainly comprises the following steps of: 1) download of an orange full genome sequence and collection of an MLO gene; 2) identification of the MLO gene; 3) identification of a candidate MLO powdery mildew gene through phylogenetic relationship of the MLO gene; and 4) comparison of the MLO powdery mildew gene. The method in the invention can effectively shorten a mining cycle of the orange powdery mildew gene, facilitating rapid identification of the powdery mildew gene; the method can develop corresponding cosegregation functional markers (SNP, SCAR and the like) through the identified candidate powdery mildew gene and can also be used in fast molecular marker-assisted selection of the powdery mildew resistance gene with high accuracy; the method can also create multiple breeding materials with resistance property through combination of other resistance gene molecular markers, shortening a breeding period and improving breeding efficiency; the method can further lays a foundation for describing an orange powdery mildew resistance molecular mechanism.
Owner:常熟市支塘镇新盛技术咨询服务有限公司
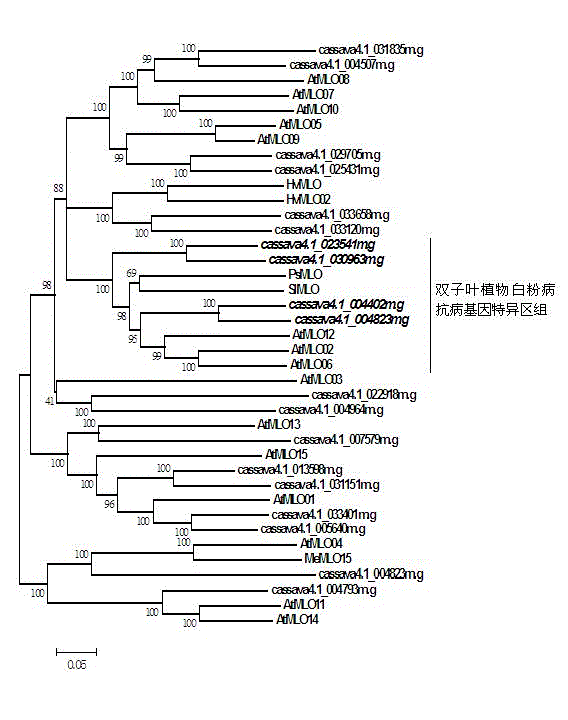
Method for quickly identifying manihot esculenta mildew-resistance locus (MLO) gene by applying comparative genomics
InactiveCN102796745AShorten digging cycleRapid identificationMicrobiological testing/measurementPlant peptidesDiseaseGenetics genomics
The invention discloses a method for quickly identifying a manihot esculenta mildew-resistance locus (MLO) gene, relates to knowledge of plant comparative genomics, genetics, bioinformatics and the like and belongs to the field of plant biotechnology science. The method mainly comprises the following steps of: 1) downloading a manihot esculenta whole genome sequence, and collecting the MLO gene; 2) identifying the MLO gene; 3) identifying the MLO gene according to the MLO gene phylogenetic relationship; and 4) comparing the MLO genes. By the method, the discovery cycle of the manihot esculenta MLO gene is shortened effectively, and the MLO gene can be quickly identified; corresponding coseparation functional markers (single nucleotide polymorphism (SNP) and specific combining ability (SCA)) can be developed by the identified candidate MLO gene, and the method can be quickly used for molecular marker auxiliary selection of the MLO gene, and is high in accuracy; by combining other disease-resistant gene molecular markers, multiresistance breeding materials can be prepared, the breeding cycle is shortened, and the breeding efficiency is improved; and the foundation is laid for elaborating a manihot esculenta MLO gene molecular mechanism.
Owner:常熟市支塘镇新盛技术咨询服务有限公司
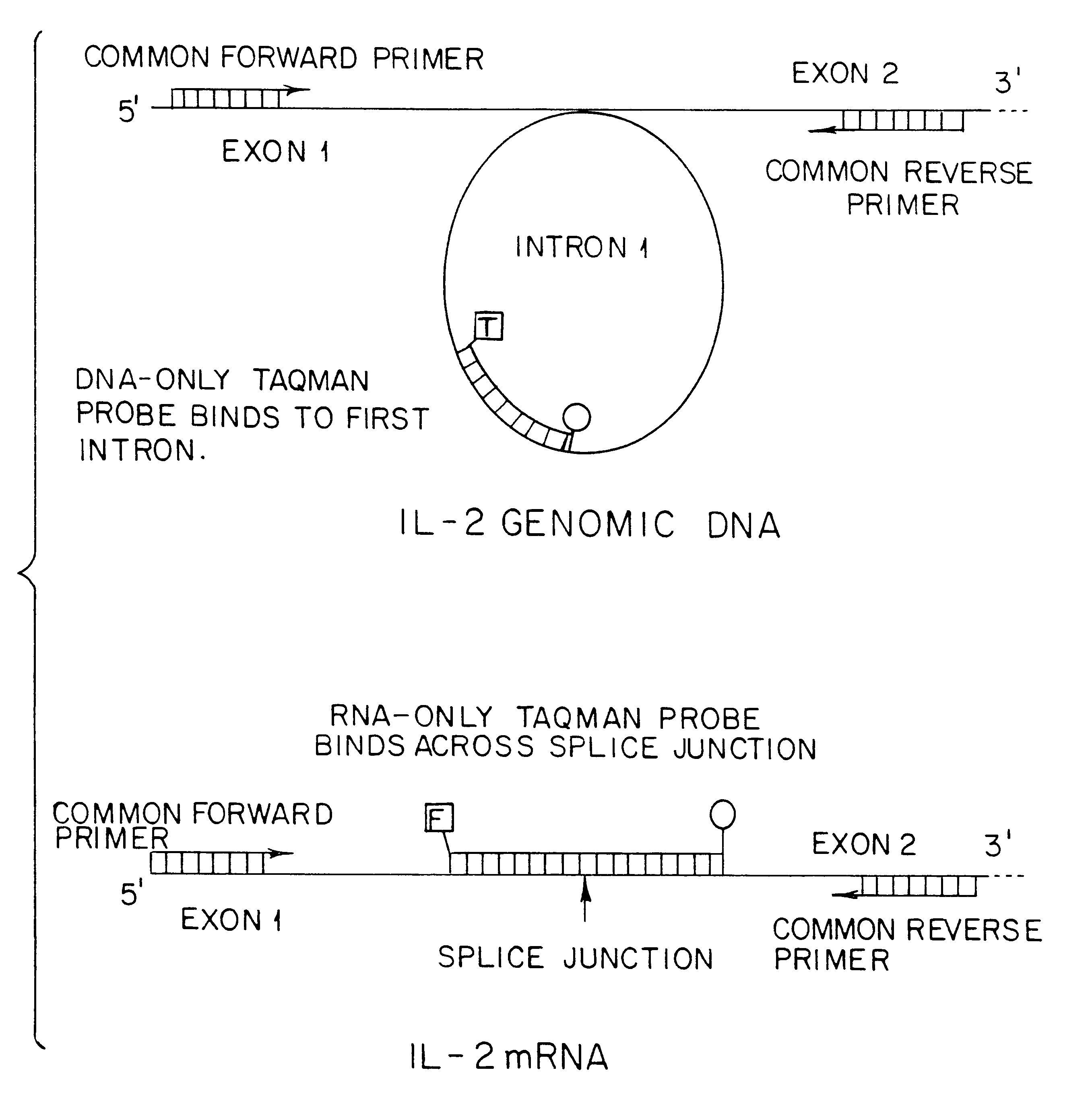
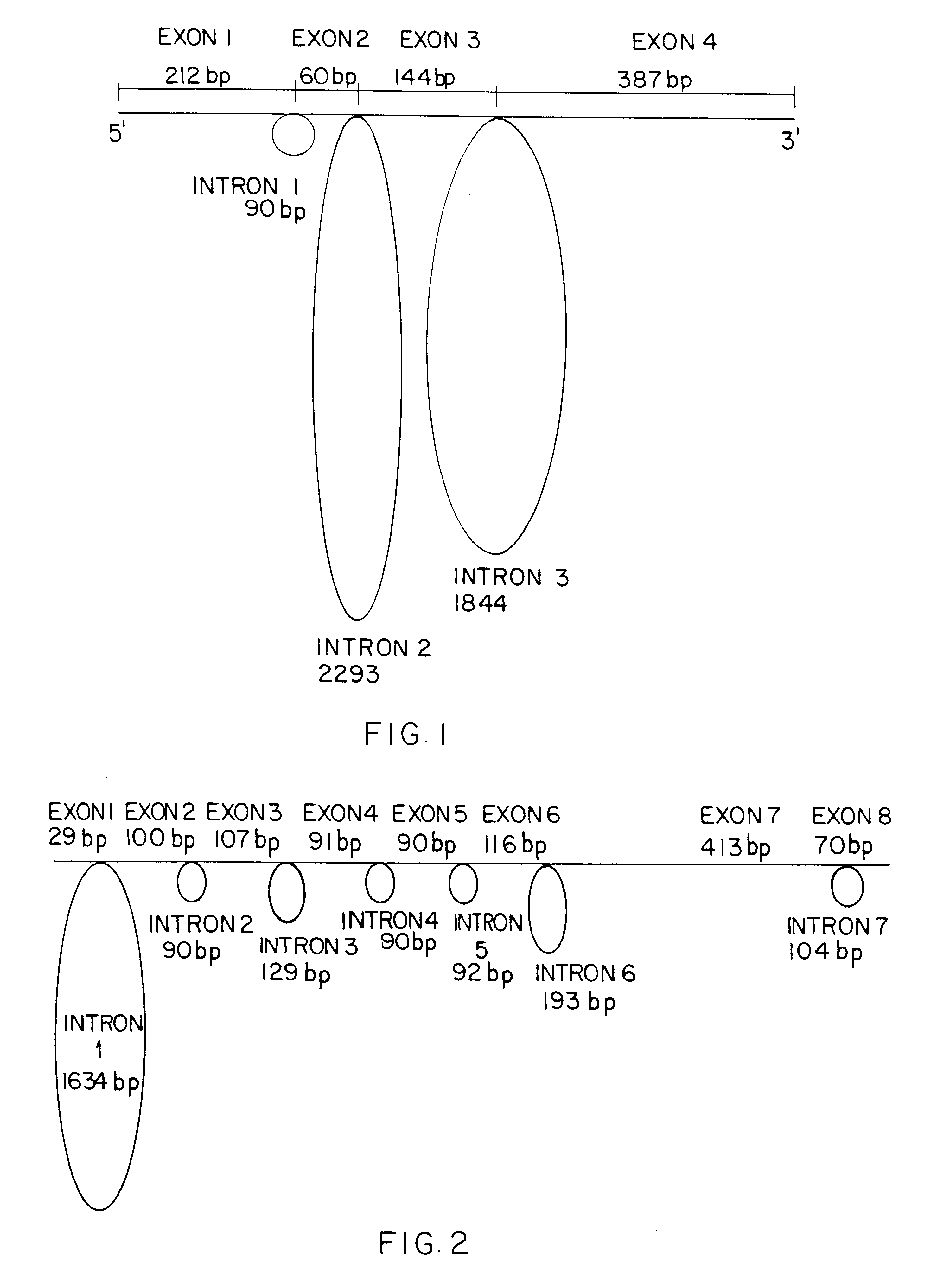
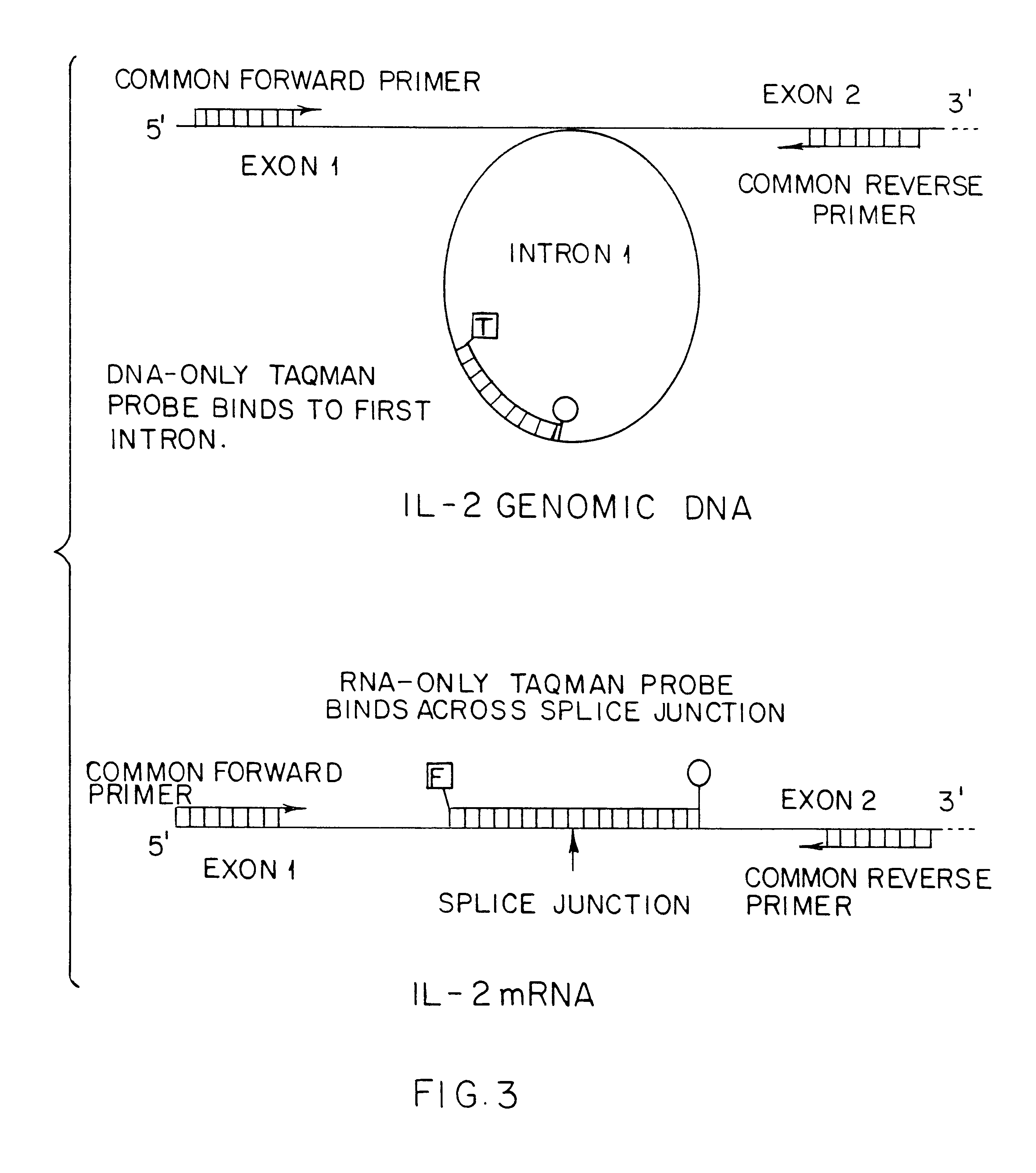
Quantitation of RNA transcripts using genomic DNA as the internal amplification competitor
InactiveUS6258543B1Improve accuracyHigh sensitivitySugar derivativesMicrobiological testing/measurementNucleic acid amplification techniqueIntein
This invention is directed to methods for the quantitative measurement of specific gene expression levels in biological samples. In one embodiment, methods for the quantitative monitoring of gene expression without either co-amplification of an added template or use of an endogenous constitutive transcript are provided. The former involves a duplex amplification reaction in which a single set of primers is used to amplify both genomic DNA and expressed mRNA from the same gene sequence. These primers are targeted for sequences flanking the splice junction and intron sequences for the mRNA and DNA respectively. By their use, any suitable nucleic acid amplification technology yields mRNA and DNA amplimers which are distinguishable by length and sequence heterogeneity. These amplimers are also present in the final amplification reaction in ratios which are dependent upon the ratios of the expressed mRNA to the DNA in the sample, allowing the quantitation of mRNA in a sample which is normalized to the number of copies of genomic DNA since the genomic DNA acts as the internal quantitation standard, and in effect yields the amount of mRNA per cell. Any detection methodology which can detect amplimers of different lengths or sequences can be used for post amplification quantitation. This strategy may be employed for any gene system in which the mRNA sequence differs from the original genomic DNA sequence. In another embodiment, methods for the quantitative monitoring of gene expression involving determining the ratio of genomic material to expressed mRNA without nucleic acid amplification and / or primer binding are described. This method includes the independent and direct detection of the genomic material and mRNA by complementary probe binding without prior amplification. Additional methods for quantitative measurement of gene expression rates without RNA transcription region introns are described. The invention is useful in gene expression determination in research and commercial applications.
Owner:APPL BIOSYSTEMS INC
Rapid identification of MLO (mildew resistance locus o) powdery mildew resistance Poncirus trifoliata genes
InactiveCN102703464AShorten digging cycleRapid identificationMicrobiological testing/measurementPlant peptidesBiotechnologyRapid identification
The invention relates to rapid identification of powdery mildew resistance Poncirus trifoliata genes, relates to knowledge of subjects such as plant comparative genomics, genetics and bioinformatics, and belongs to the technical field of plant biotechnology. The rapid identification mainly includes the steps of firstly, downloading full genomic sequences of Poncirus trifoliate and collecting MLO genes; secondly, identifying the MLO genes; thirdly, identifying phylogenetic relationship of the MLO genes; and fourthly, comparing the MLO powdery mildew resistance genes. By the rapid identification, mining cycle of the powdery mildew resistance Poncirus trifoliata genes is shortened effectively, and the powdery mildew resistance genes can be identified quickly. Corresponding co-separation functional marks (SNP (single nucleotide polymorphism), Scar and the like) can be developed through candidate powdery mildew resistance genes identified, the rapid identification is also available for molecular marker-assisted selection of the powdery mildew resistance genes, and accuracy is high. The rapid identification can also be used with other resistance gene molecular markers to create multiresistance breeding materials, breeding period is shortened, breeding efficiency is improved, and basis for elaborating powdery mildew resistance Poncirus trifoliata molecular mechanisms is laid.
Owner:常熟市支塘镇新盛技术咨询服务有限公司
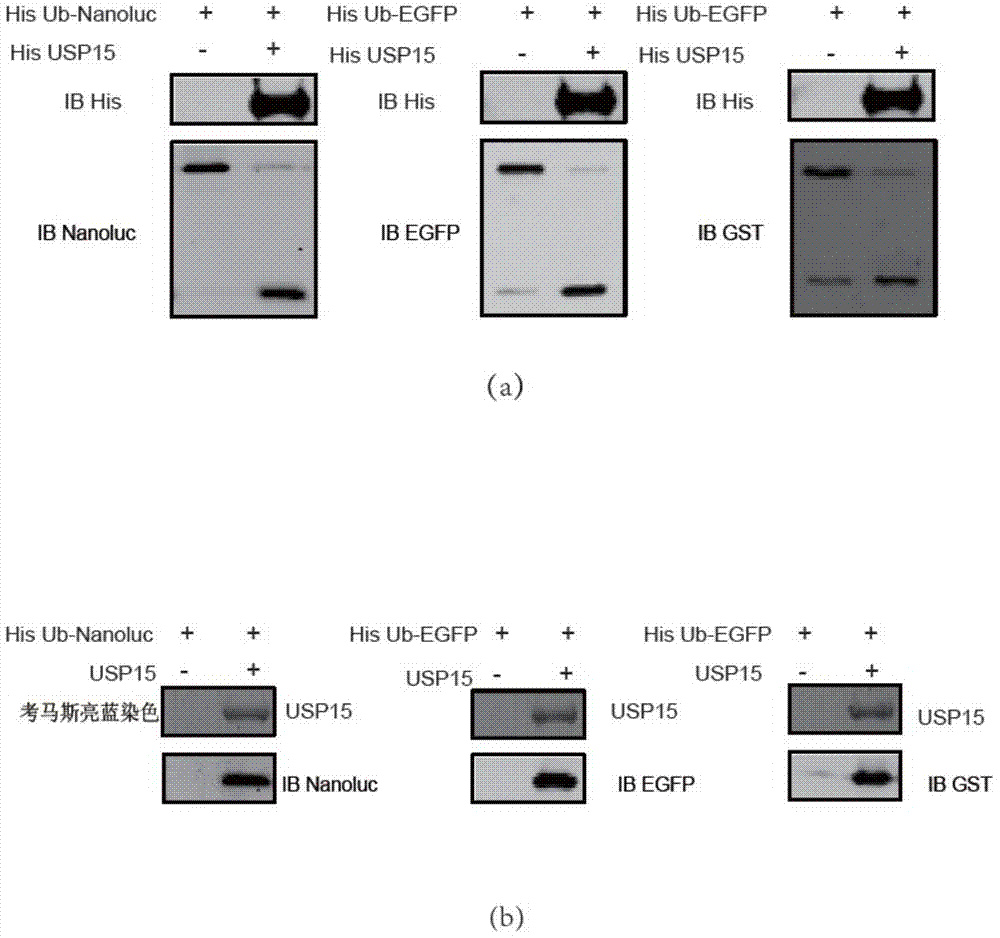
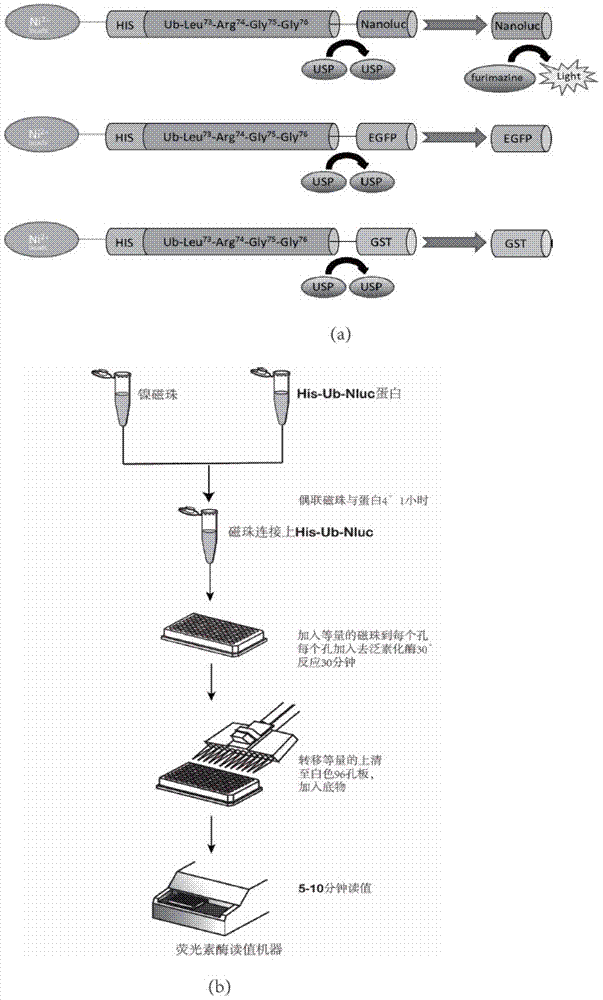
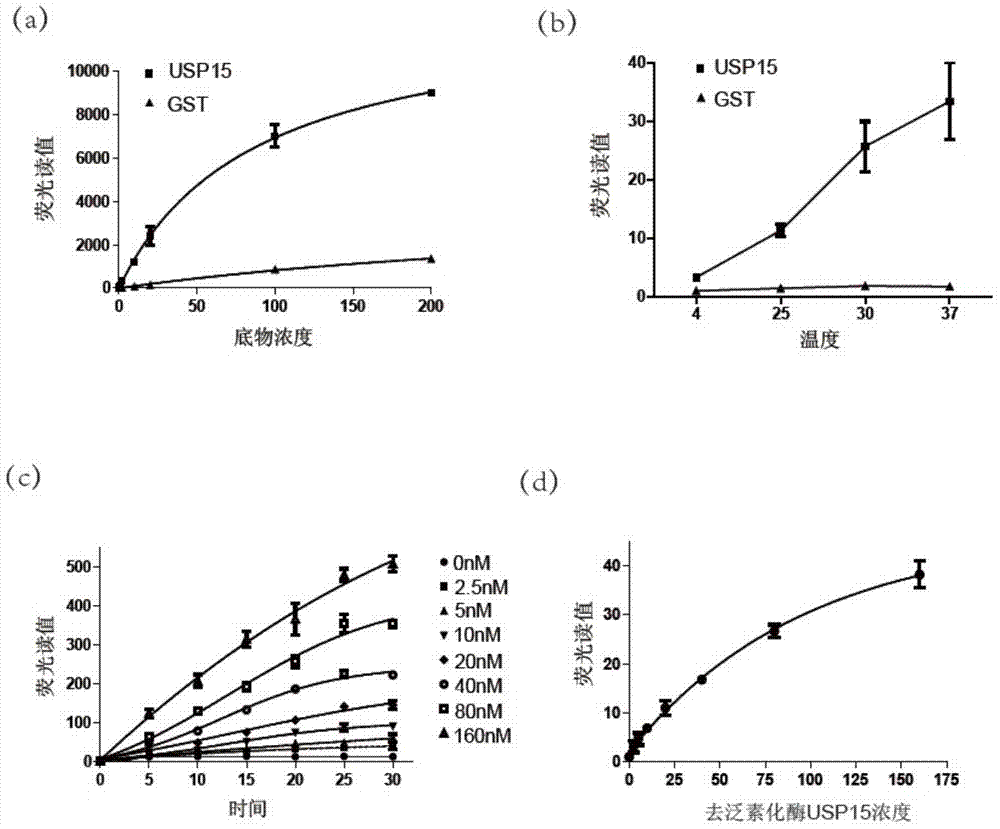
Ub-Nanoluc reporter gene system and Ub-Ub-GS-Nanoluc reporter gene system, constructions and applications thereof
InactiveCN104844704AMicrobiological testing/measurementOxidoreductasesLuciferase GeneEnzyme inhibitor
The present invention discloses an Ub-Nanoluc reporter gene system and an Ub-Ub-GS-Nanoluc reporter gene system, and applications of the Ub-Nanoluc reporter gene system and the Ub-Ub-GS-Nanoluc reporter gene system in deubiquitination enzyme activity detection and deubiquitination enzyme inhibitor throughput-screening, wherein the Ub-Nanoluc reporter gene DNA sequence is represented by SEQ ID NO.1, and the Ub-Ub-GS-Nanoluc reporter gene DNA sequence is represented by SEQ ID NO.4. According to the present invention, the high sensitivity characteristic of luciferase Nanoluc is utilized to provide the method for deubiquitination enzyme activity detection and deubiquitination enzyme inhibitor throughput-screening by using the chemical luminescent reporter gene so as to provide the new action target point and the new ideal for screening and research of new drugs including anticancer drugs, and provide broad application prospects, great enormous benefits, and great social benefits.
Owner:EAST CHINA NORMAL UNIV
Compositions, methods, and kits useful for the diagnosis and treatment of spinal muscular atrophy
InactiveUS20050100923A1Easy splicingImprove the level ofNanotechBacteriaSurvival of motor neuronSmn gene
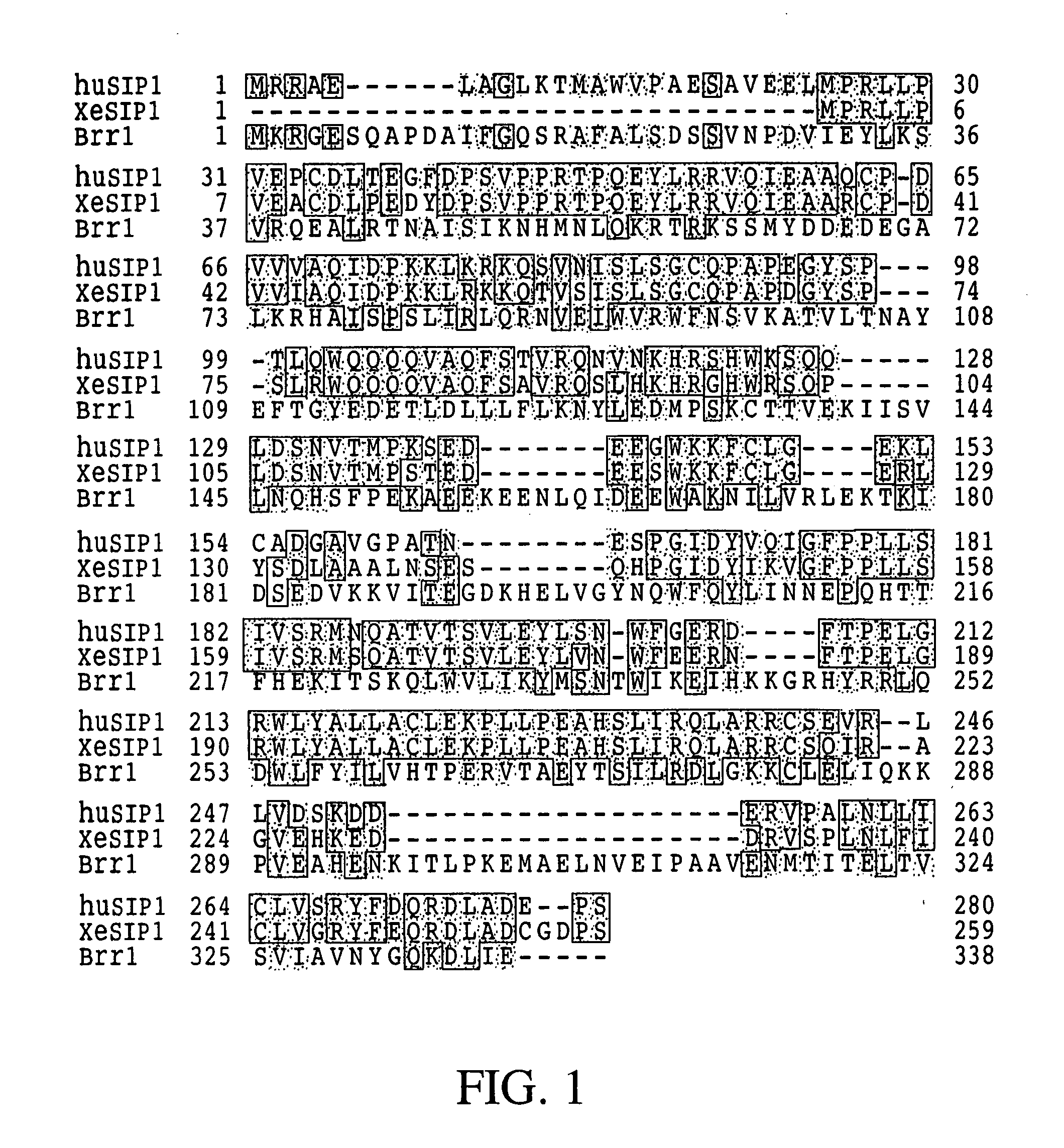
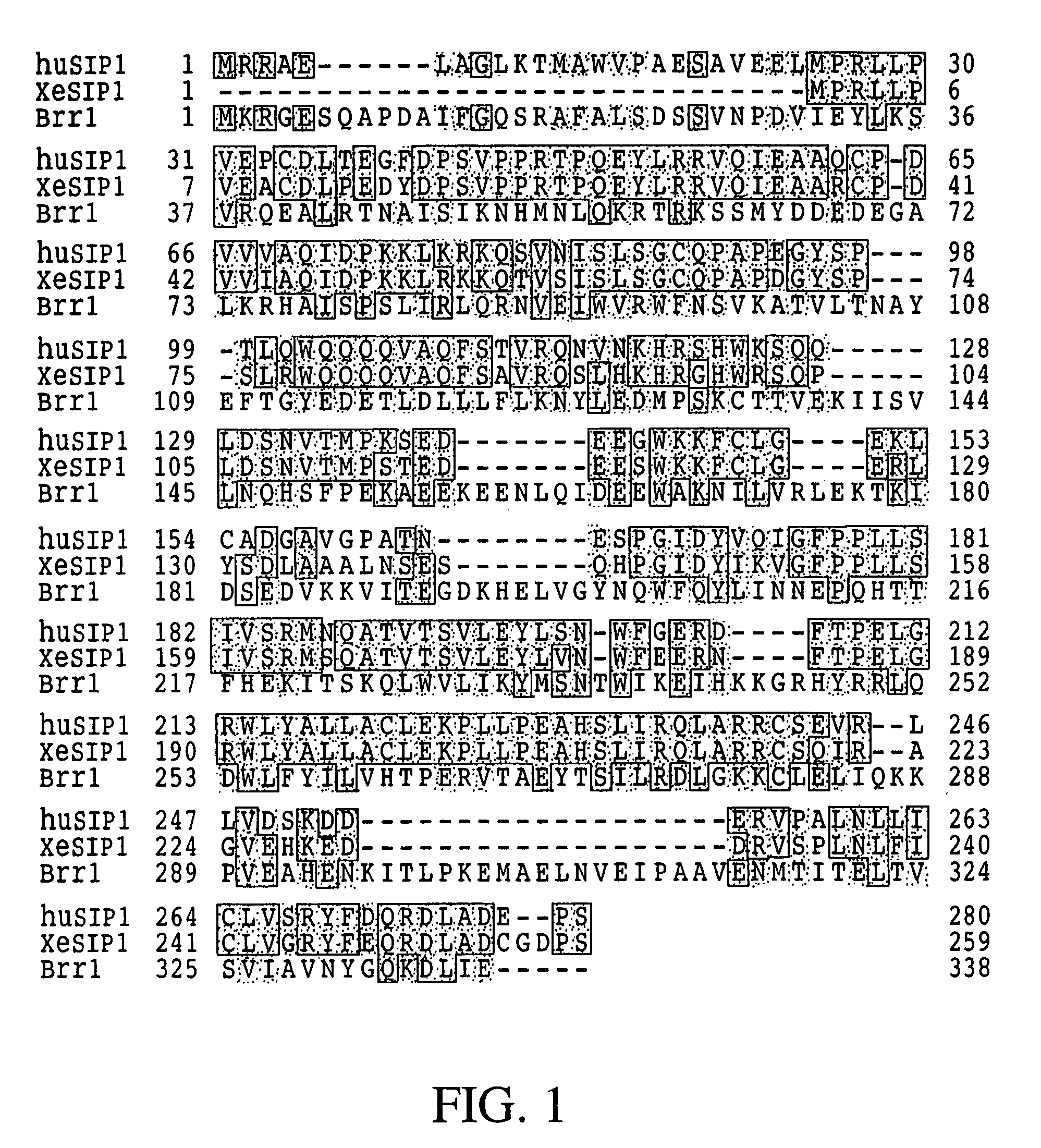
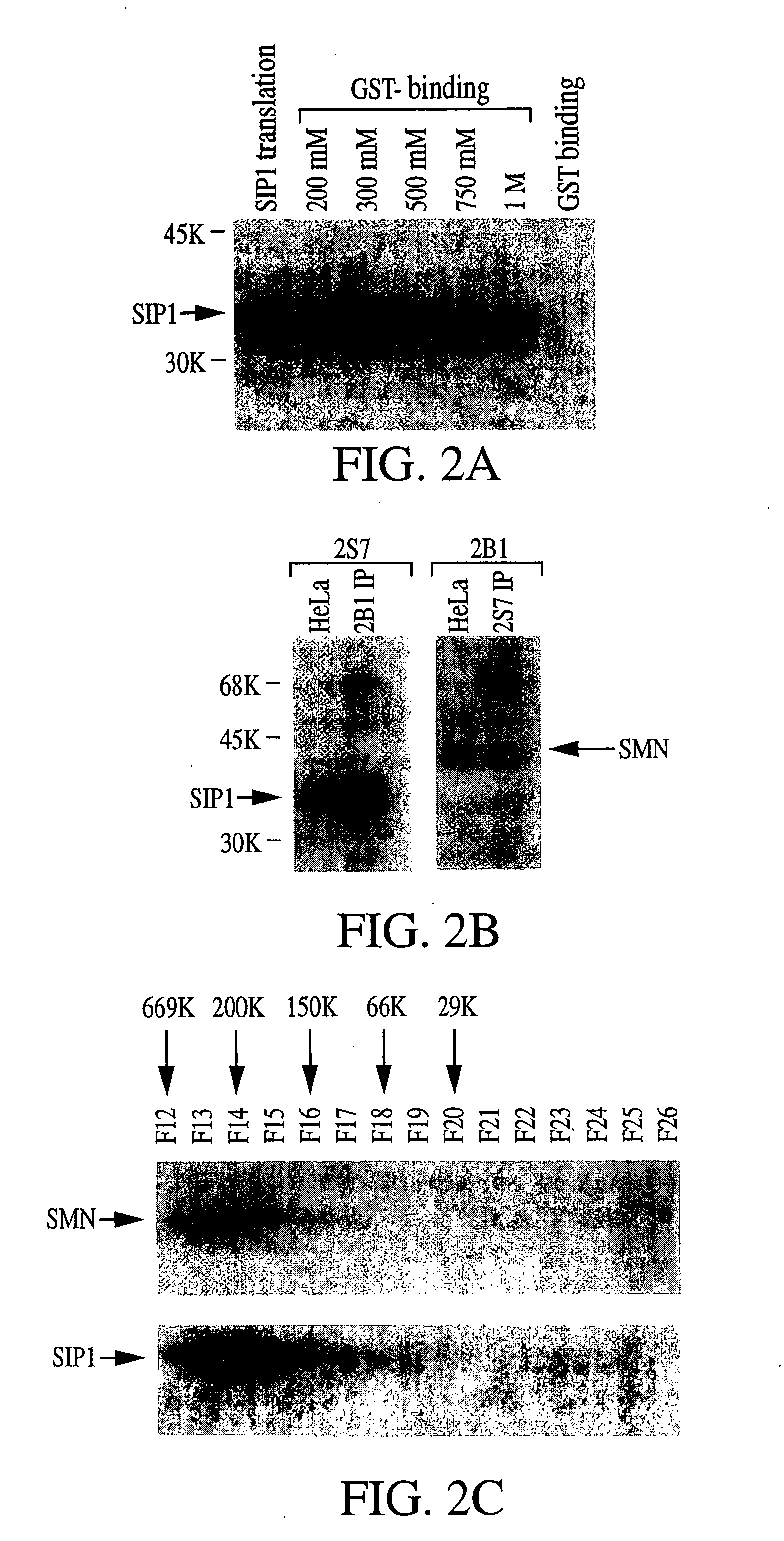
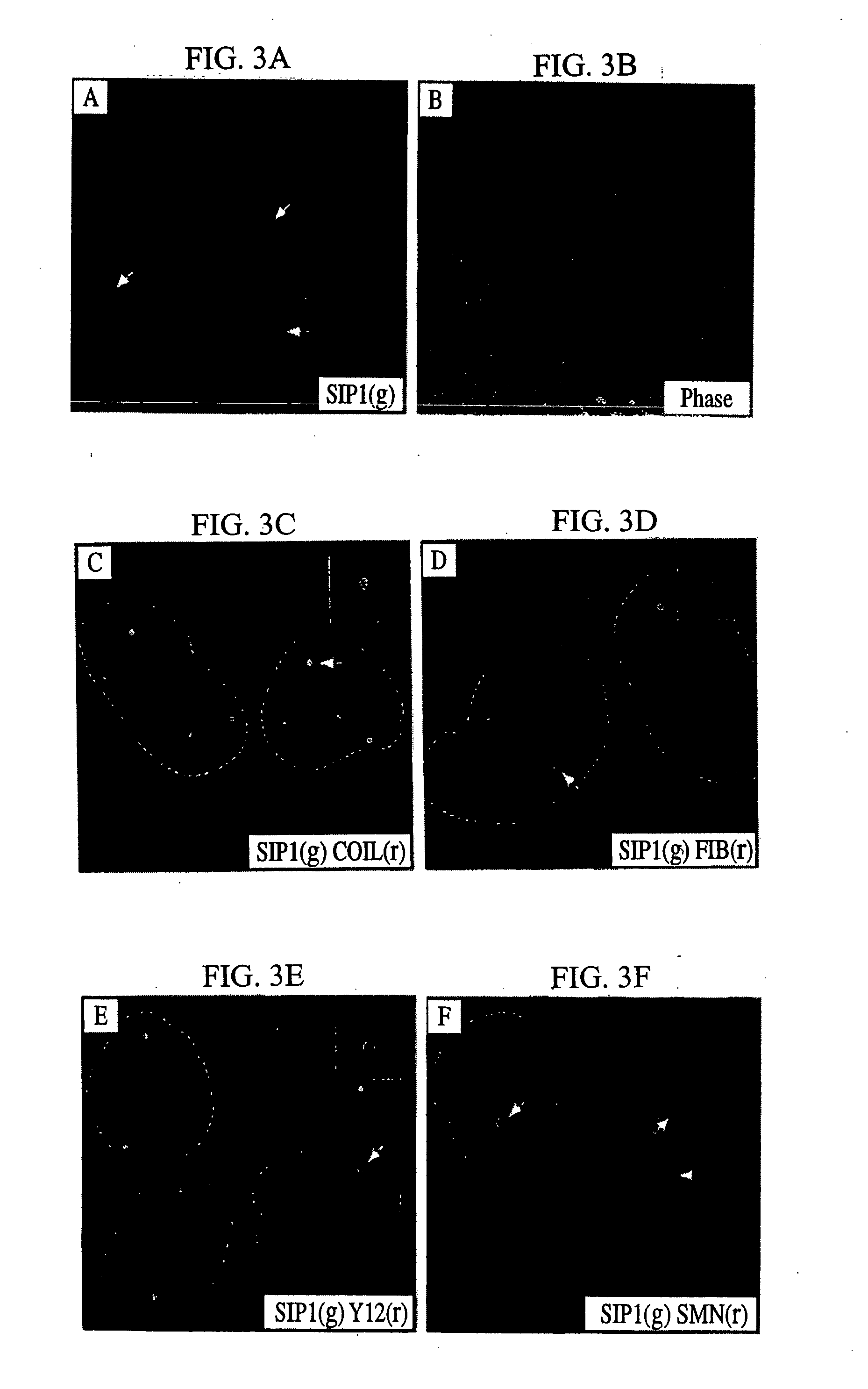
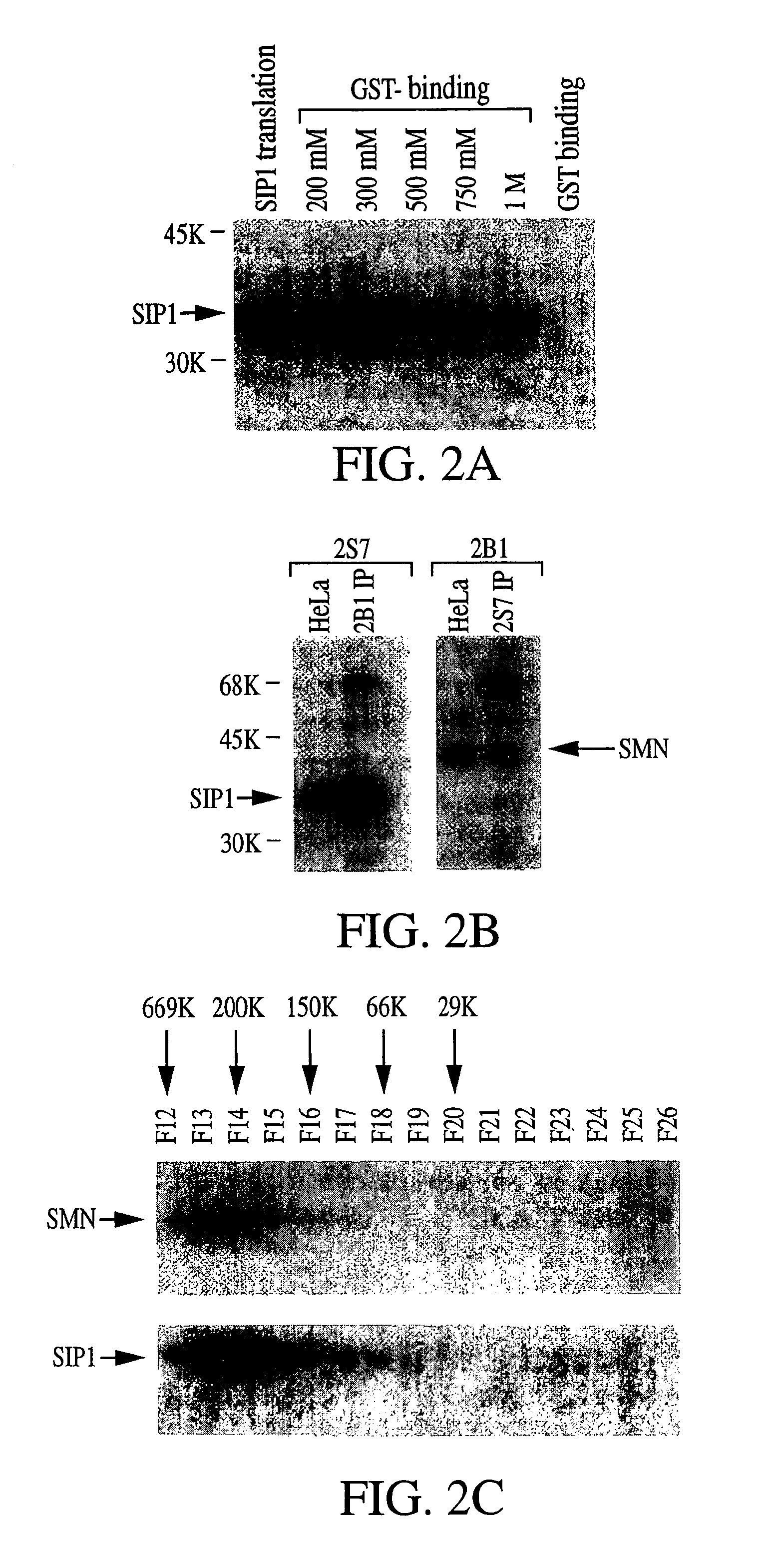
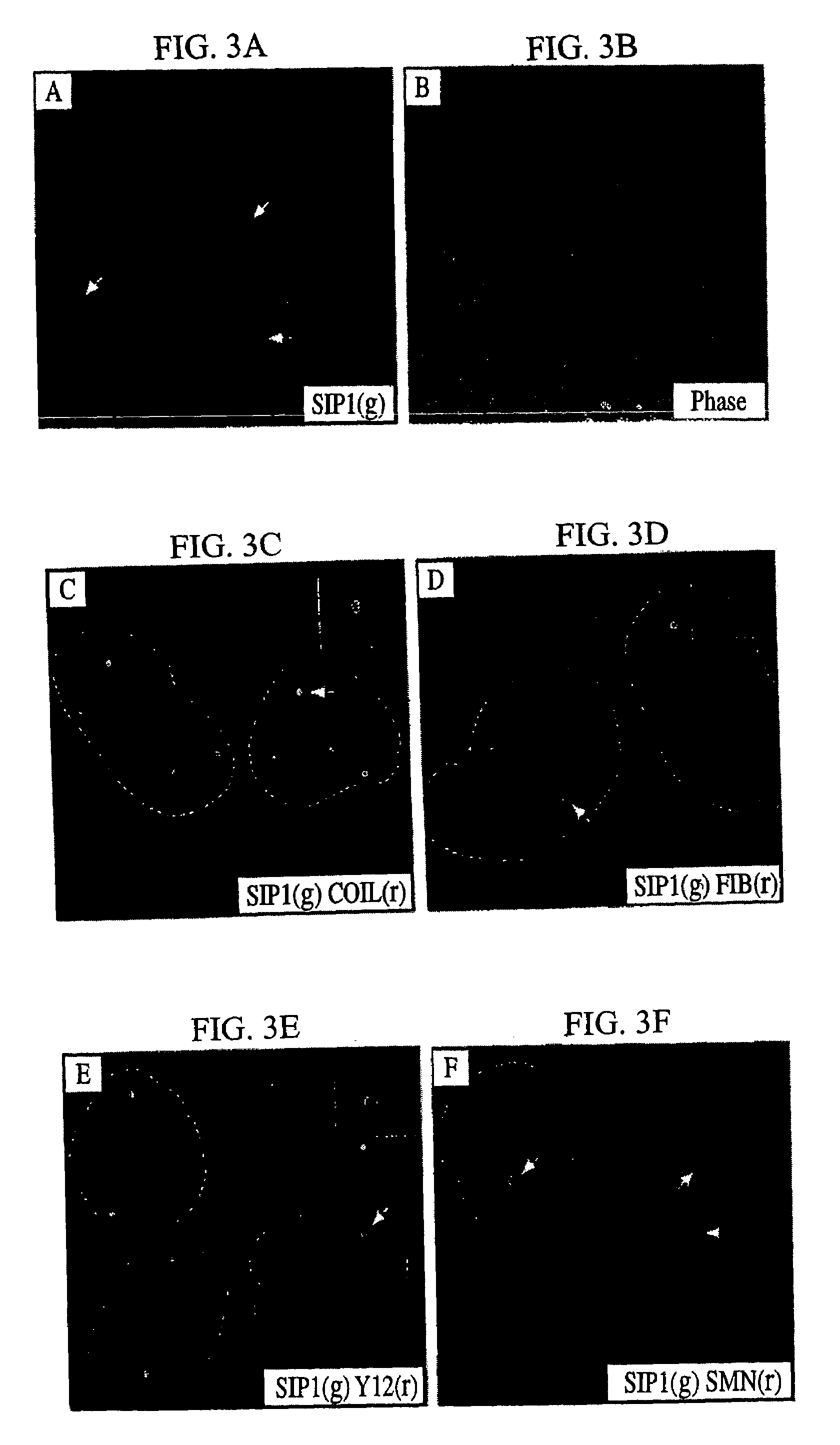
The invention relates to an isolated nucleic acid encoding a eukaryotic Survival of Motor Neuron-Interacting Protein 1 (SIP1), compositions comprising SIP1 and SIP1 and the spinal muscular atrophy (SMA) disease gene product Survival of Motor Neuron protein (SMN), and diagnostic and therapeutic assays directed to SMA. The invention also relates to another protein that specifically interacts with SMN and is a component of gems, designated Gemin3, and the nucleic acid encoding the protein. Additionally, the invention relates to a novel cell line wherein the endogenous SMN genes have been deleted and where an exogenous nucleic acid encoding SMN has been inserted into the cell such that expression of SMN in the cell is under the control of an inducible promoter. This novel cell line provides a stable genetic system for the study of SMA and for the development of SMA therapeutics.
Owner:THE TRUSTEES OF THE UNIV OF PENNSYLVANIA
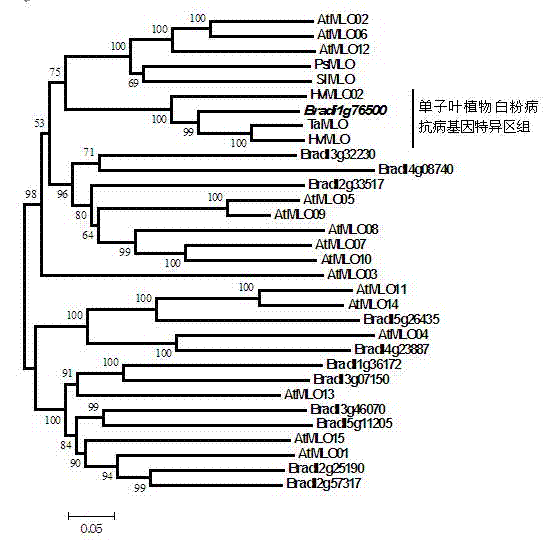
Rapid identification of anti-powdery-mildew gene of Brachypodium distachyon
InactiveCN102703462AShorten digging cycleRapid identificationMicrobiological testing/measurementPlant peptidesBiotechnologyBrachypodium sylvaticum
The invention relates to rapid identification of anti-powdery-mildew gene of Brachypodium distachyon, relates to the knowledge of plant comparative genomics, genetics, biological information and other subjects, and belongs to the scientific field of plant biotechnology. The rapid identification of anti-powdery-mildew gene of Brachypodium distachyon comprises the main steps of: (1) downloading full genome sequences of Brachypodium distachyon and collecting MLO (mildew resistance locus o) type gene; (2) identifying the MLO type gene; (3) analyzing the MLO type gene historical development relation; and (4) comparing the MLO type powdery mildew gene. According to the rapid identification of anti-powdery-mildew gene of Brachypodium distachyon provided by the invention, the excavation period of the anti-powdery-mildew gene of Brachypodium distachyon is effectively shortened, and the rapid identification of the powdery mildew gene is facilitated; the identified powdery mildew gene can be used for developing corresponding coseparation functional markers (SNP (single nucleotide polymorphism), SCAR (sequence-characterized amplified region), and the like), and can also be rapidly used for assisted selection of molecule markers of anti-powdery-mildew gene, and the accuracy is high; development of multi-resistance breeding materials can be carried out by combining with other anti-disease gene molecule markers, the breeding time is shortened and the breeding efficiency is improved; and foundation is established for expounding the molecular mechanism of the anti-powdery-mildew gene of Brachypodium distachyon.
Owner:常熟市支塘镇新盛技术咨询服务有限公司
Transgenic vector system capable of promoting cell transplantation and gene expression and applications of transgenic vector system
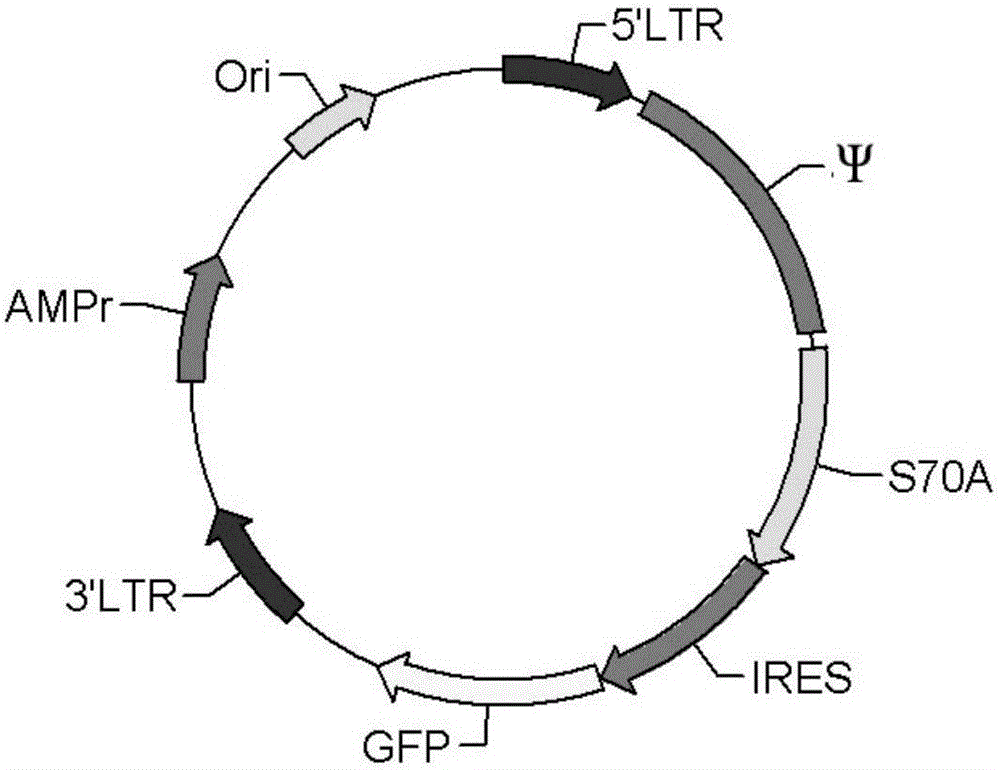
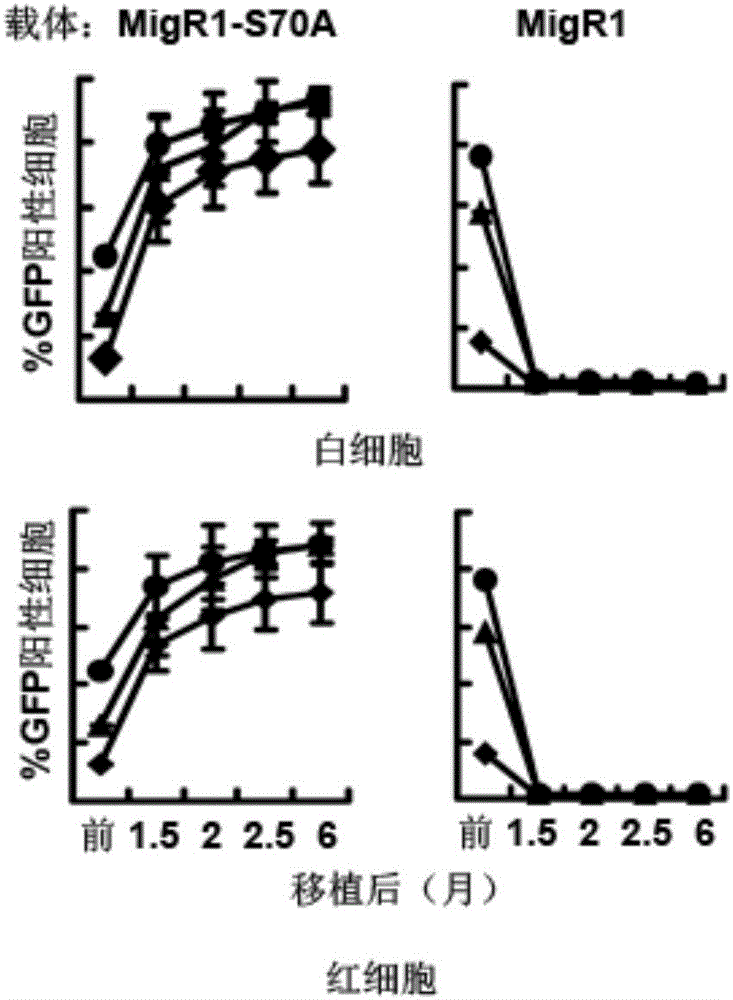
ActiveCN106480090AEasy to make successAccelerated deathNucleic acid vectorVector-based foreign material introductionCell survivalGene Modification
The invention discloses a transgenic vector system capable of promoting cell transplantation and gene expression. The transgenic vector system comprises a screening gene system, a target gene expression system, and a vector used for bearing the two systems, wherein the screening gene system is an expression system of a gene capable of promoting cell division or cell survival or a mutant of the gene, or a silencing system of a gene capable of inhibiting division or promoting cell death. Compared with the prior art, the system has the advantages that firstly, the limitation that whether the target gene has the growth vigor or not is eliminated, secondly, the screening occurs on both the stem cell level and the dedifferentiating cell level, and therefore, the gene modification cells can be stably maintained all the life or the number of the gene modification cells can be increased; thirdly, medicine screening is not needed, and therefore, the influences of the toxicity of the medicines are avoided; moreover, the mutant only promoting the cell survival but not promoting the cell mutation is adopted, and therefore, the risk of causing cancer is avoided. Therefore, the system has great application values in gene and cell therapy and the manufacturing of animal bioreactors or animal models based on adult (stem) cells.
Owner:上海百英生物科技股份有限公司
Induction of tung tree hypocotyls callus and method for efficiently regenerating plants
ActiveCN104094848AStrong embryonicHigh activityPlant tissue cultureHorticulture methodsHypocotylTerra firma
The invention discloses induction of tung tree hypocotyls callus and a method for efficiently regenerating plants, and belongs to the field of plant cell engineering. According to the invention, through the processes of inducing the callus by the hypocotyl of the tung oil tree, inducing an adventitious bud by the callus, performing transgenerational multiplication culture, performing strong seedling culture, performing rooting culture, performing acclimatization and transplantation and the like, a way for quickly breeding good tung oil tree plants is provided, and a firm foundation for later improving the resistance of the tung oil tree, accelerating the molecular breeding process of the tung oil tree, improving the oil quality and other properties, and establishing the genetic system of the tung oil tree in a genetic engineering method is built.
Owner:CENTRAL SOUTH UNIVERSITY OF FORESTRY AND TECHNOLOGY
Purine nucleosidase
InactiveUS6066484AReduce the amount requiredIncrease volumeSugar derivativesBacteriaMicroorganismHaemophilus
Novel purine nucleosidase derived from Ochrobactrum anthropi microorganisms, a gene system coding therefor, uses therefor and particularly its use in the production of beer.
Owner:SUNTORY HLDG LTD
Isolated polypeptide deletion mutants of survival of motor neuron-interacting protein 1
InactiveUS7459309B2Improve the level ofEasy splicingNanotechBacteriaSurvival of motor neuronNeuron survival
The invention relates to an isolated nucleic acid encoding a eukaryotic Survival of Motor Neuron-Interacting Protein 1 (SIP1), compositions comprising SIP1 and SIP1 and the spinal muscular atrophy (SMA) disease gene product Survival of Motor Neuron protein (SMN), and diagnostic and therapeutic assays directed to SMA. The invention also relates to another protein that specifically interacts with SMN and is a component of gems, designated Gemin3, and the nucleic acid encoding the protein. Additionally, the invention relates to a novel cell line wherein the endogenous SMN genes have been deleted and where an exogenous nucleic acid encoding SMN has been inserted into the cell such that expression of SMN in the cell is under the control of an inducible promoter. This novel cell line provides a stable genetic system for the study of SMA and for the development of SMA therapeutics.
Owner:THE TRUSTEES OF THE UNIV OF PENNSYLVANIA
Method for screening pig disease resistant breeds, and application thereof
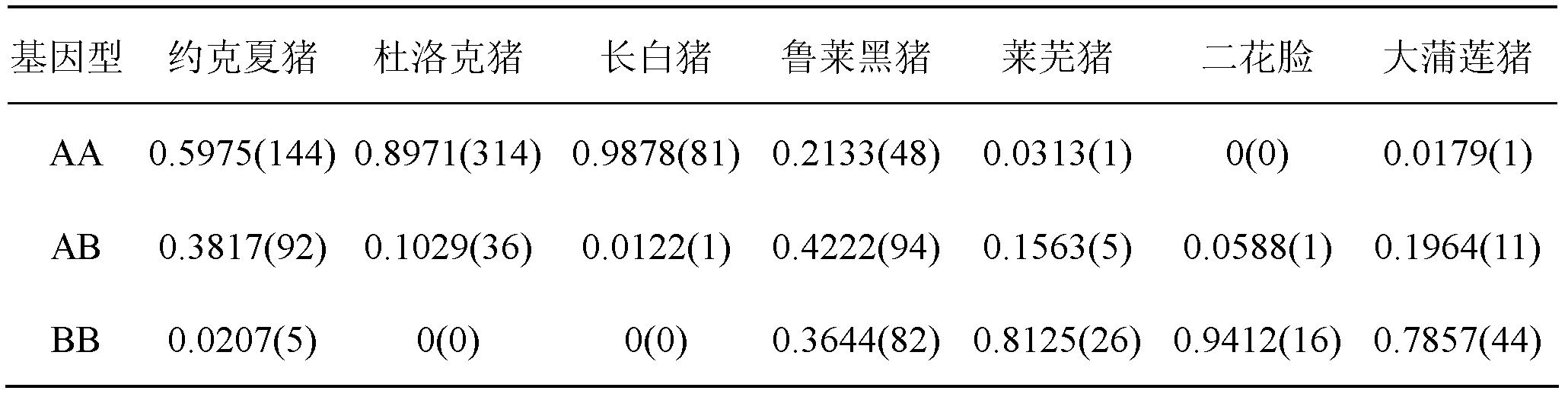
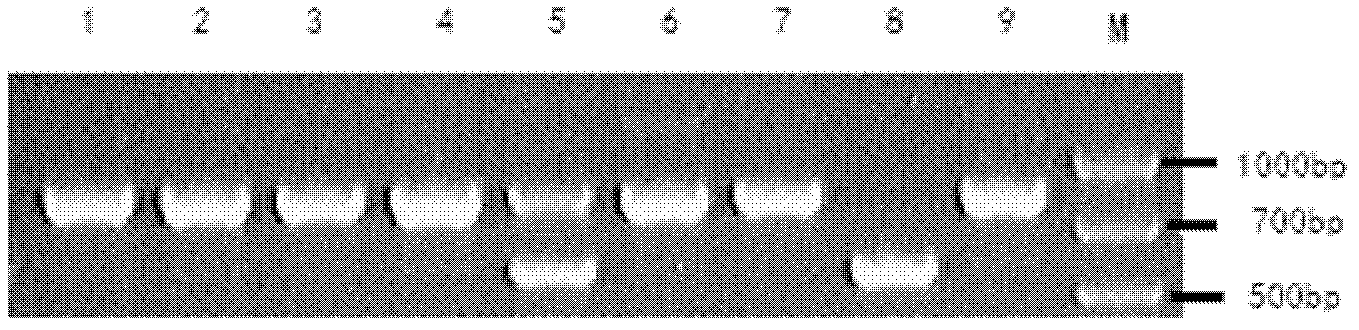
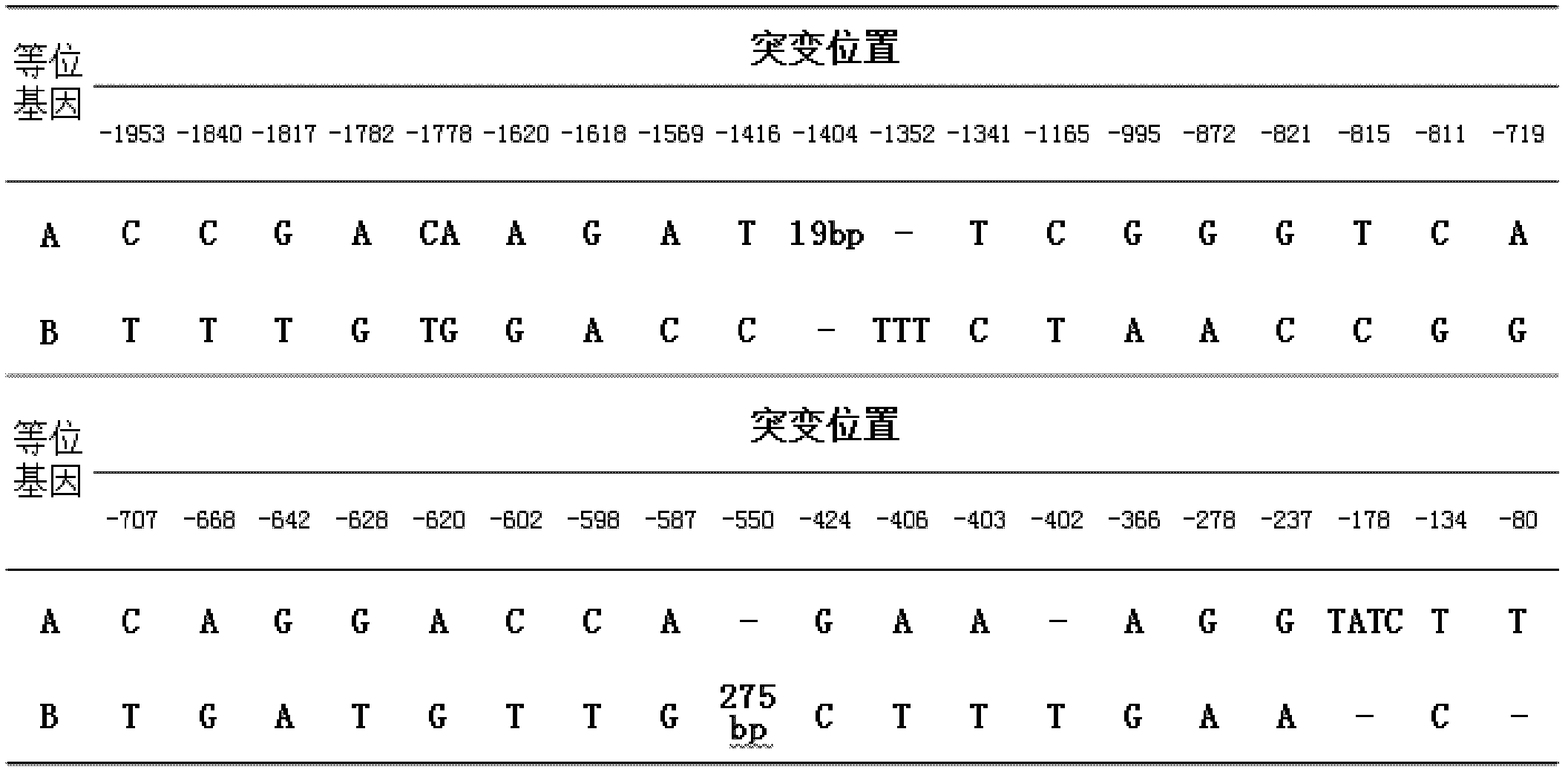
The invention relates to the field of molecular genetics, and specifically relates to a molecular marking method for a pig MX1 gene 5' control region mutable site, and an application thereof in pig disease resistance breeding. According to the invention, a plurality of differences are found in MX1 5' control regions of different breeds of pigs. Among Chinese local pig breeds, a 275bp segment inserted gene is a dominant genotype. Among western pig breeds, the frequency of the genotype is low. With a luciferase reporter assay system, it is found that, the activity of a promoter with an inserted275bp segment is substantially higher than that of another. The existence of the 275bp segment in a pig genome MX1 control region can serve as a molecular mark related to pig disease resistant characters. The method is simple and fast, and is not influenced by the environment. With the method, early-stage breed selection can be realized.
Owner:SHANDONG AGRICULTURAL UNIVERSITY
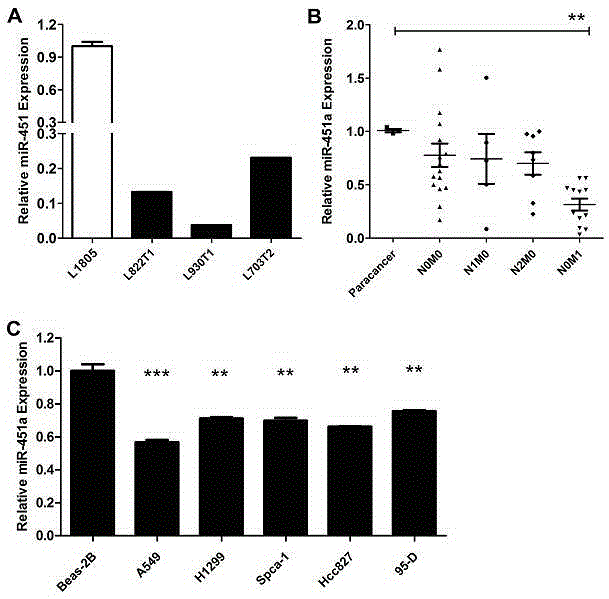
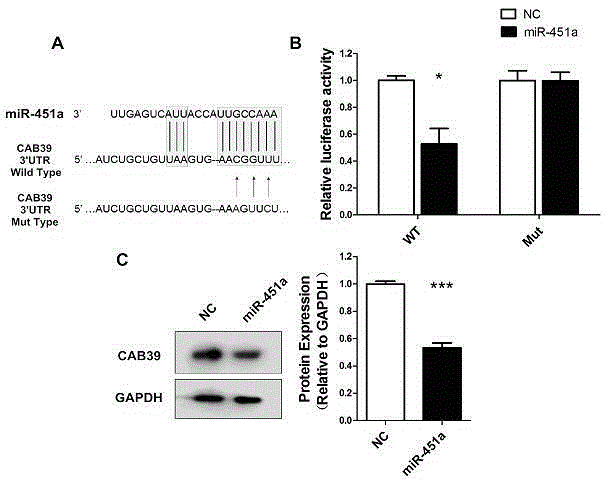
Effect of miR-451a cells in non-small cell lung cancer
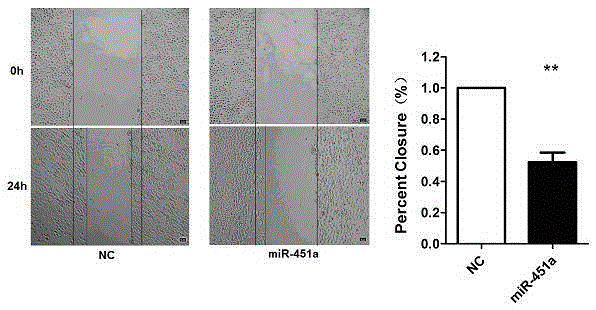
The invention relates to the effect of miR-451a cells in the non-small cell lung cancer. Migration of A549 cells can be significantly inhibited through overexpression of miR-451a in the A549 cells. By means of the bioinformatics analysis means, it is predicted that translation of the target gene mRNA is inhibited or the target gene mRNA is directly degraded by miR-451a through regional complementation with 3'-UTR of the target gene CAB39 mRNA; it is determined that the CAB39 gene is the target gene of miR-451a through double-luciferase reporter gene system analysis and Western Blot experiment verification. Finally, it is verified through the Transwell Assay and Flow Cytometry Assay experimental means that miR-451a performs the function of inhibiting migration of cells by reducing the CAB39 gene in the A549 cells, and a novel accurate treating researching and treating scheme is provided. By means of the effect, important research guidance and application value is provided for using miRNA for diagnosing and treating the non-small cell lung cancer and searching for drug targets in the fields such as tumor immunotherapy clinically.
Owner:SHANGHAI UNIV
Quick propagation method for akebia trifoliata stem
ActiveCN104855294AHigh rooting ratePromote growthPlant tissue cultureHorticulture methodsAKEBIA TRIFOLIATA STEMSeedling
The invention provides a quick propagation method for akebia trifoliata stem. The quick propagation method comprises such steps as explant disinfection, primary culture, multiplication culture, rooting culture, acclimatization and transplant. The quick propagation method has the advantages that the explant pollution rate is low, the propagation speed is high, and the culture cycle is short; not only is a way provided for quick propagation of elite akebia trifoliata stem plants, but also industrialized seedling culture of the akebia trifoliata stem and follow-up genetic system transformation become possible.
Owner:CENTRAL SOUTH UNIVERSITY OF FORESTRY AND TECHNOLOGY
Coincidence reporter gene system
Disclosed is a nucleic acid comprising a nucleotide sequence encoding (i) two or more reporters comprising a first reporter and a second reporter that is different from the first reporter; and (ii) one or more ribosomal skip sequences, wherein a ribosomal skip sequence is positioned between the first and second reporters, wherein the first and second reporters are stoichiometrically co-expressed from the nucleotide sequence and the nucleic acid does not comprise a cytomegalovirus-immediate early (CMV-IE) promoter. Also disclosed are methods of screening test compounds for ability to modulate a biological activity of interest using the nucleic acid, as well as related recombinant expression vectors, host cells, and populations of cells.
Owner:UNITED STATES OF AMERICA
Pif promoter capable of being activated by pinctada martensii MSX (muscle segment homeobox) gene and application thereof
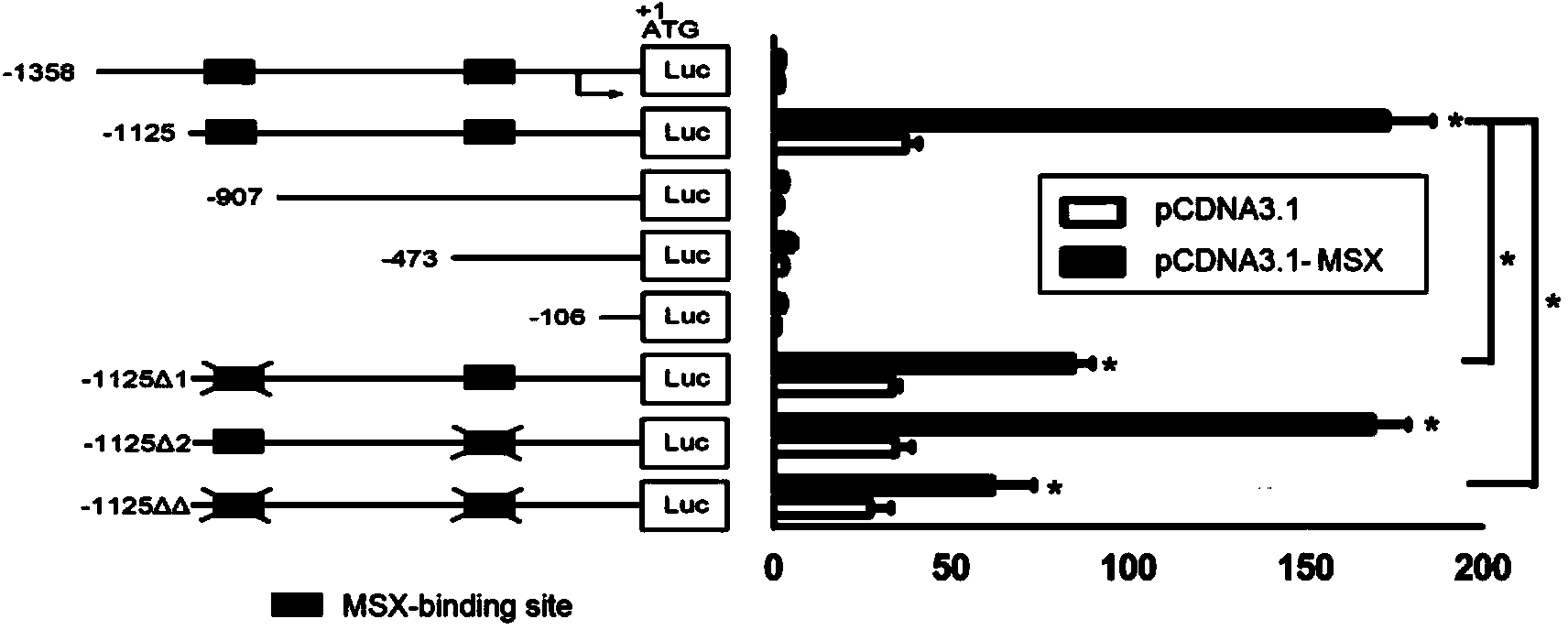
ActiveCN103820453AVector-based foreign material introductionDNA/RNA fragmentationNucleotideBinding site
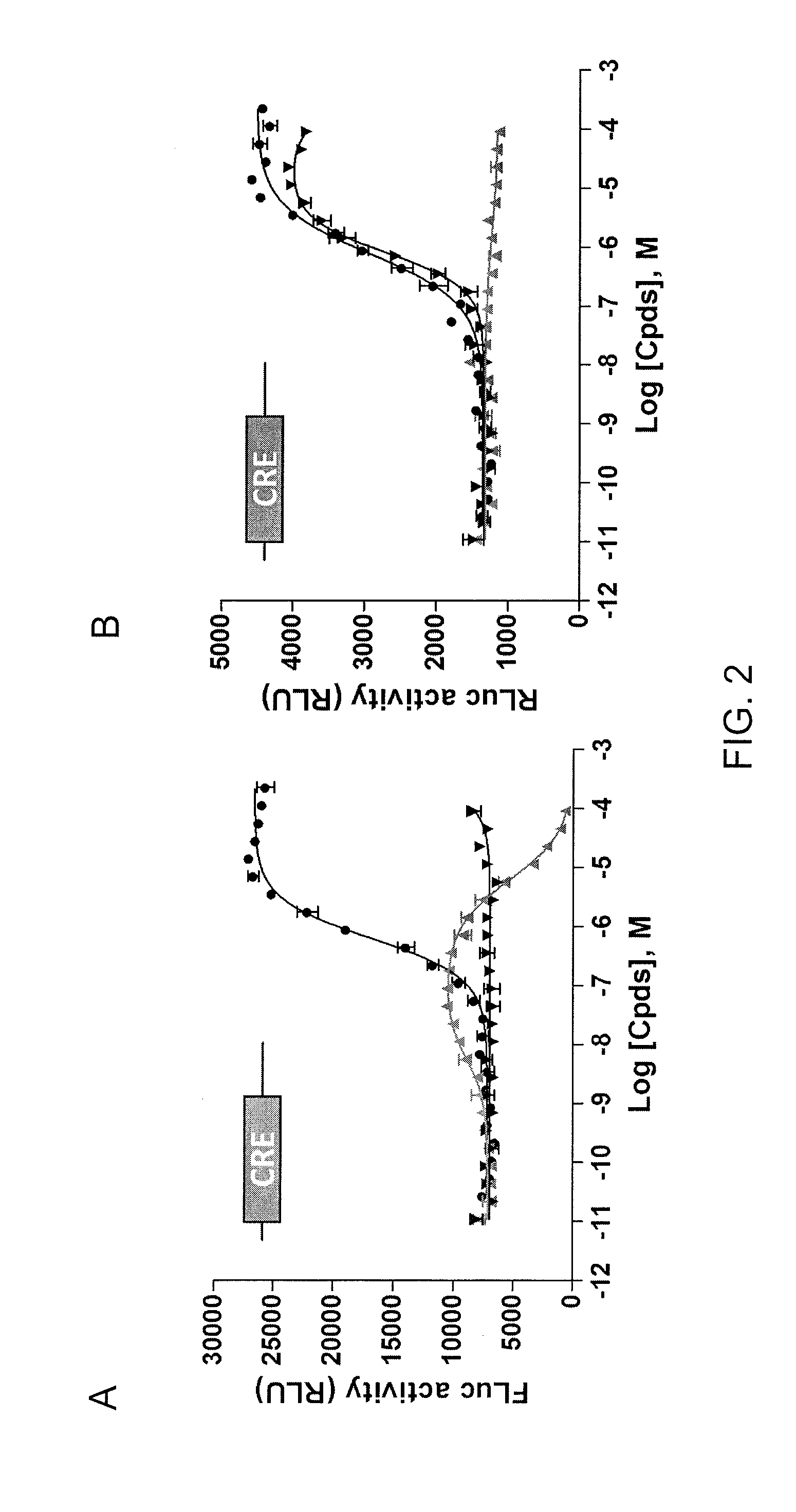
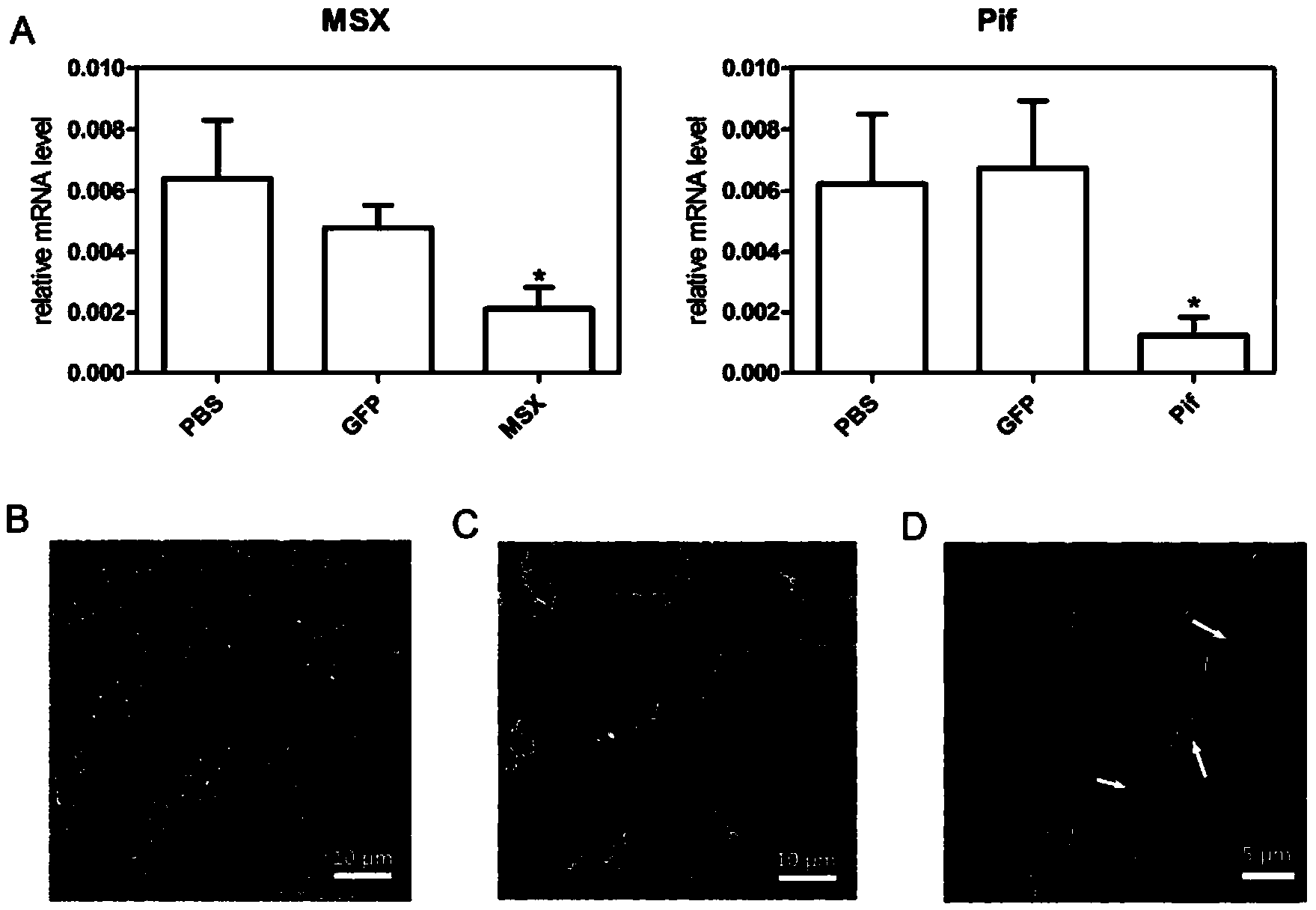
The invention discloses a Pif promoter capable of being activated by pinctada martensii MSX (muscle segment homeobox) genes and application of the Pif promoter. The nucleotide sequence of the Pif promoter is shown as SEQ ID NO.1. A Pif promoter sequence is obtained through amplification by a genome walking method. The Pif promoter has a binding site 1 and a binding site 2, which can be bound to the MSX genes. Through the establishment of wild type carriers and mutant type carriers of Pif promoters of different fragment lengths, a dual-luciferase reporter gene system is applied to identify the fact that the MSX genes act on the Pif promoter through the binding site 1. The Pif promoter can start the downstream target gene expression, thereby promoting the growth of pearls or pearl oysters.
Owner:SOUTH CHINA SEA INST OF OCEANOLOGY - CHINESE ACAD OF SCI
Method for obtaining regeneration plants by use of camellia oleifera hypocotyl
ActiveCN104938335AStrong seedling effect is goodHigh rooting ratePlant tissue cultureHorticulture methodsHypocotylCamellia oleifera
The invention discloses a method for obtaining regeneration plants by use of camellia oleifera hypocotyl and belongs to the field of plant cell engineering. The method mainly comprises the step that adventive buds are induced by camellia oleifera hypocotyl. Through multiplicative subculture, strong seedling culture, rooting culture, acclimatization and transplant and the like of the adventive buds, not only is a new way provided for quick propagation of favorable camellia oleifera plants, but also a solid foundation is laid for improving the resistivity of camellia oleifera, accelerating the breeding process of camellia oleifera molecules and improving the oil quality and other properties by the gene engineering method, as well as building a camellia oleifera genetic system.
Owner:CENTRAL SOUTH UNIVERSITY OF FORESTRY AND TECHNOLOGY
Rapid Identification of pear powdery mildew resistance gene
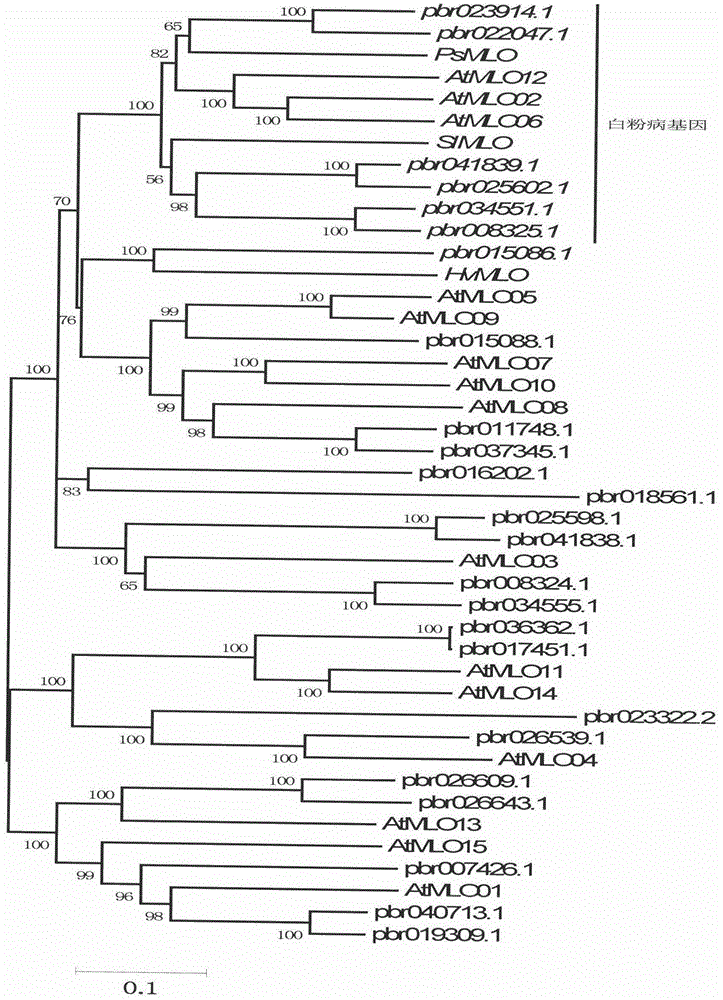
InactiveCN104561027AShorten digging cycleImprove identification efficiencyMicrobiological testing/measurementGenetic engineeringAgricultural scienceRapid identification
The invention relates to rapid identification of a pear powdery mildew resistance gene, and relates to the subject knowledge of plant comparative genomics, genetics and bioinformatics, belonging to the field of science of biotechnology. The rapid identification of the pear powdery mildew resistance gene comprises the following main steps of: 1) downloading of a complete genome sequence of a pear and collection of an MLO gene; 2) identification of the MLO gene; 3) MLO gene phylogenetic relationship; and 4) comparison of an MLO powdery mildew gene. The rapid identification of the pear powdery mildew resistance gene, provided by the invention, has the advantages of effectively shortening the pear powdery mildew gene mining period, and being favorable for the rapid identification of the powdery mildew gene; corresponding co-segregation functional markers (SNP, SCAR and the like) can be developed through the identification of candidate powdery mildew genes, and the co-segregation functional markers can be rapidly used for the molecular marker-assisted selection of the powdery mildew resistance gene, so that the accuracy is high; by combining other powdery mildew resistance gene molecular markers, a multi-resistance breeding material can be created, the breeding period can be shortened, and the breeding efficiency can be improved; a foundation is laid for explaining a pear powdery mildew resistance molecular mechanism.
Owner:CHANGSHU CITY DONGBANG DONGDUN VEGETABLE PROFESSION COOP
Reagents and Methods for Treating Cancer
ActiveUS20110293698A1High potencyImprove efficacyOrganic active ingredientsFermentationLung cancerOvarian cancer cells
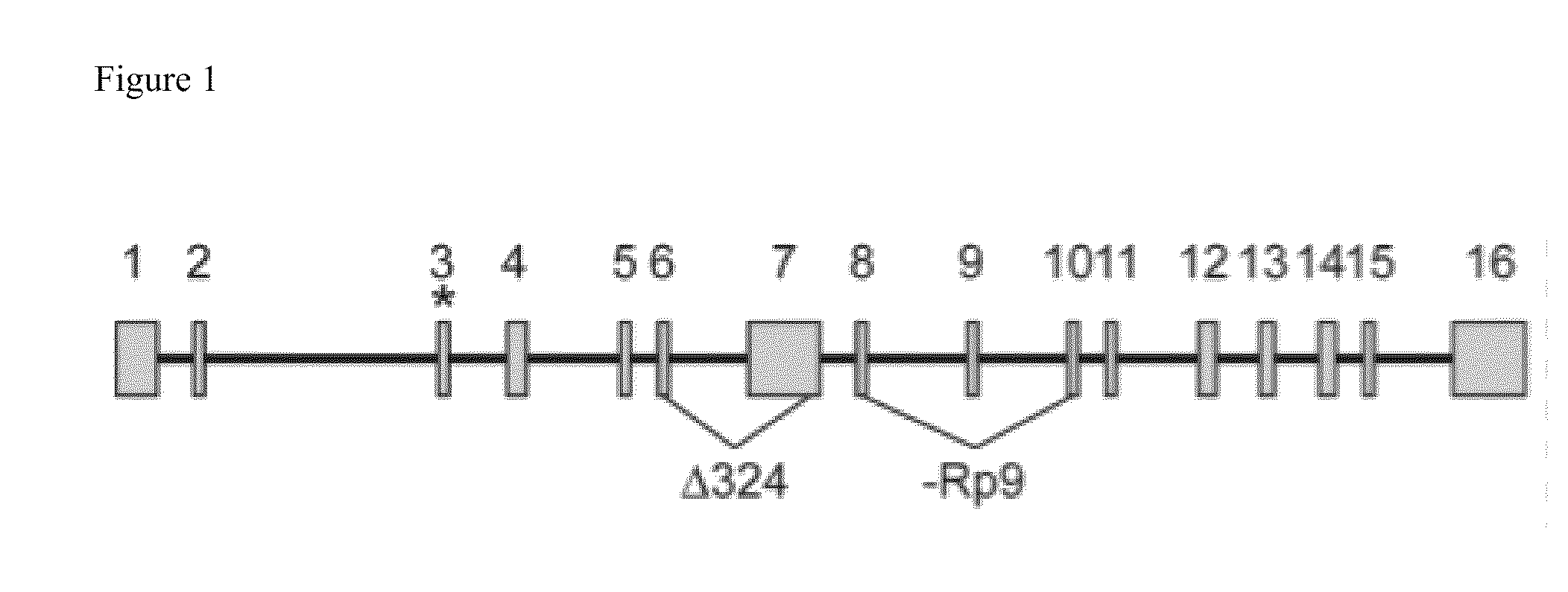
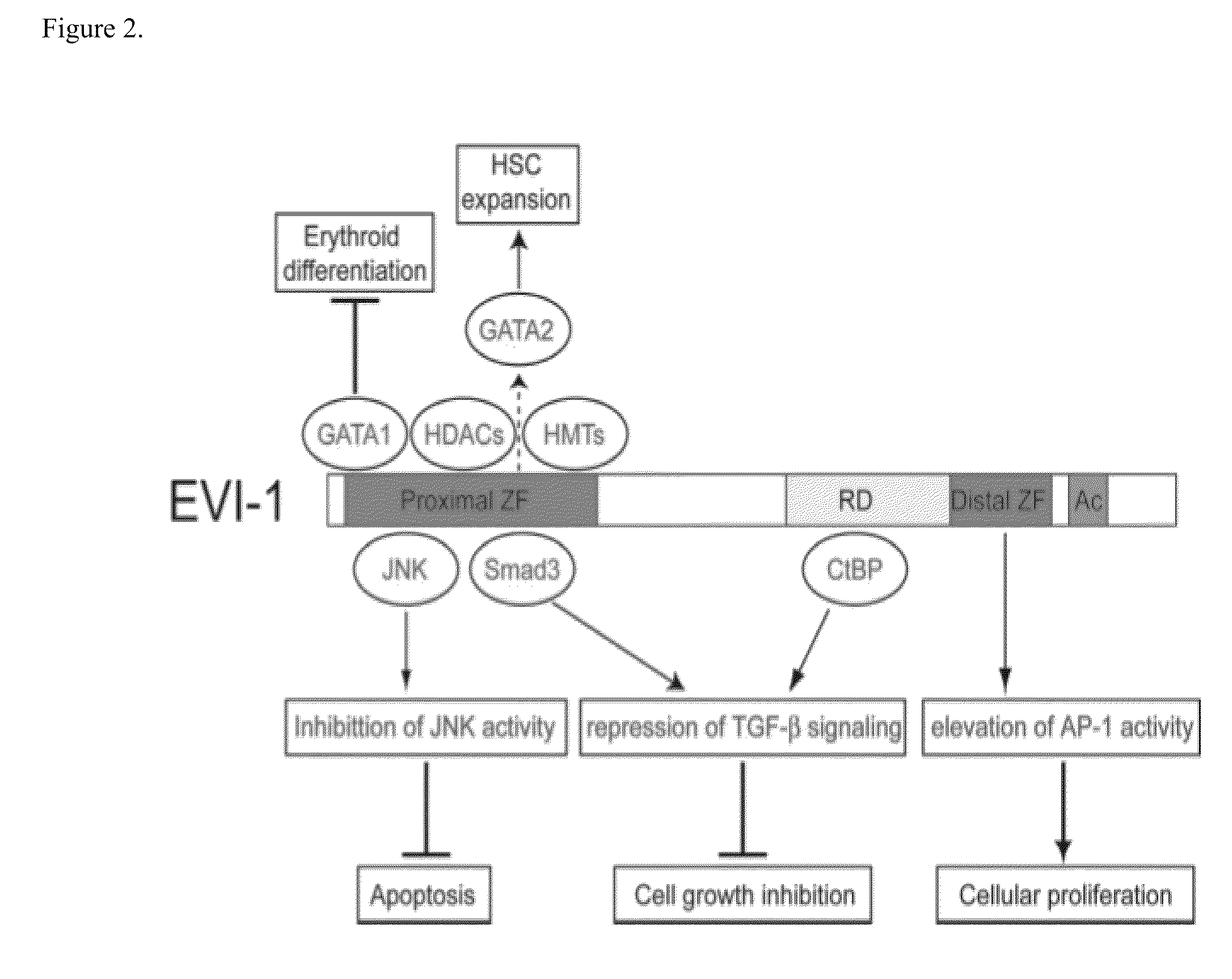
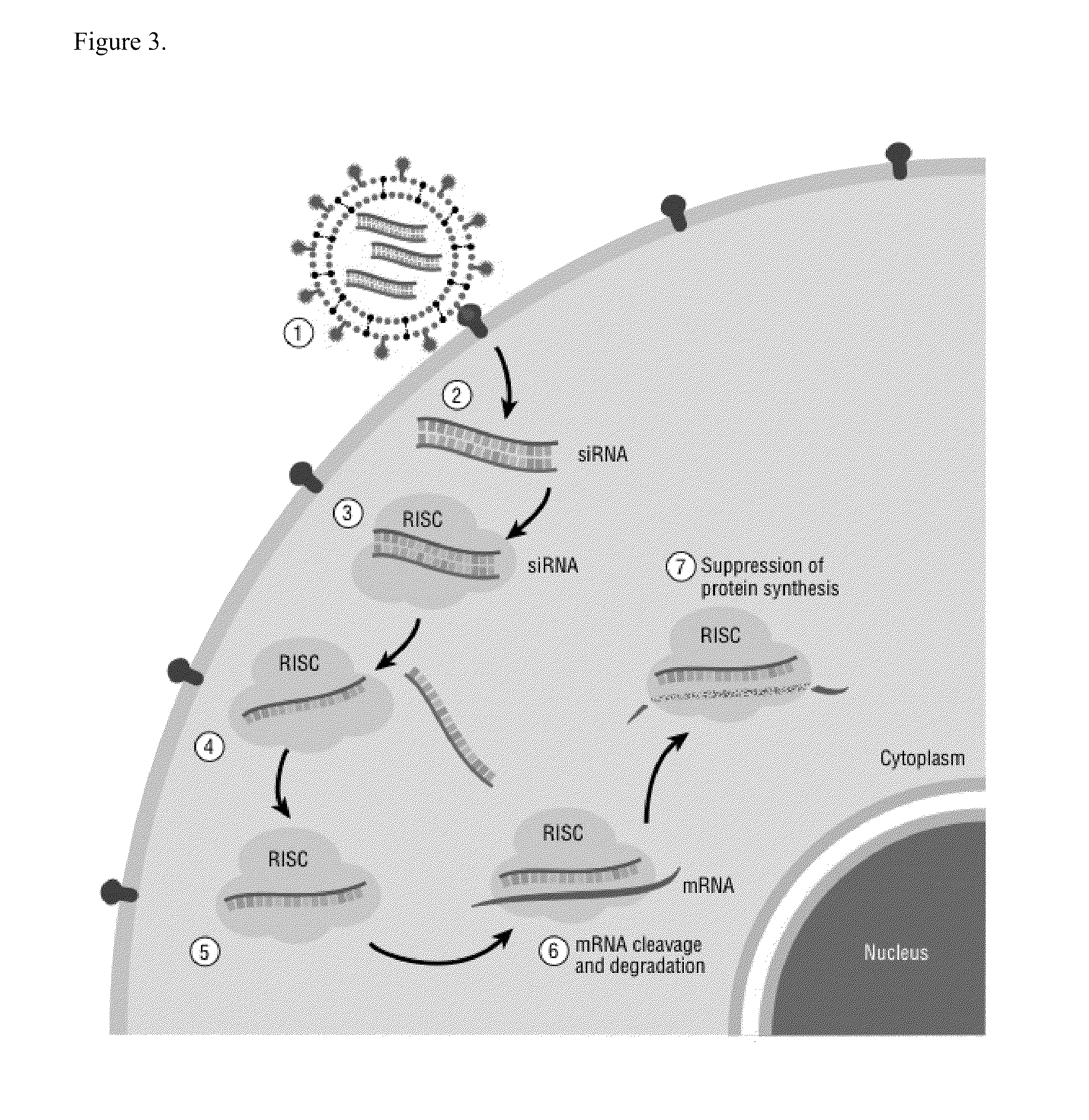
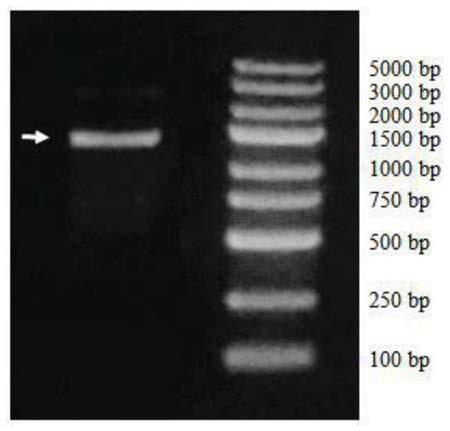
This invention describes a genetic system for targeting the EVI1 gene in mammalian cells. The EVI1 gene is an oncogenic transcription factor that, when expressed, accelerates cell division and inhibits death of cells. Nucleotide sequences that block the expression of EVI1 and drug delivery systems for them are described. These nucleotide sequences cause a block in cell growth and division and trigger death of mammalian cells, including lung and ovarian cancer cells.
Owner:NANOONCOLOGY ASSIGNMENT FOR THE BENEFIT OF CREDITORS INC +1
Method for efficiently and directly reproducing plants by virtue of vernicia fordii petioles
ActiveCN104054579ADirect regeneration of adventitious buds is rareStrong resistancePlant tissue cultureHorticulture methodsVernicia fordiiPlant cell
The invention discloses a method for efficiently and directly reproducing plants by virtue of vernicia fordii petioles, and belongs to the field of plant cell engineering. The method mainly comprises the processes of direct induction of adventitious buds by virtue of the vernicia fordii petioles, secondary reproduction culturing, strong seedling culturing, rooting culturing, acclimatization and transplantation and the like. According to the method, means is provided for the rapid reproduction of high-quality vernicia fordii plants, a solid foundation is also laid for improving the resistance, the yield of vernicia fordii and characters such as oil quality and the like by virtue of a genetic engineering method and establishing a vernicia fordii genetic system, and many means and technical support are provided for the culturing of transgenic vernicia fordii.
Owner:CENTRAL SOUTH UNIVERSITY OF FORESTRY AND TECHNOLOGY
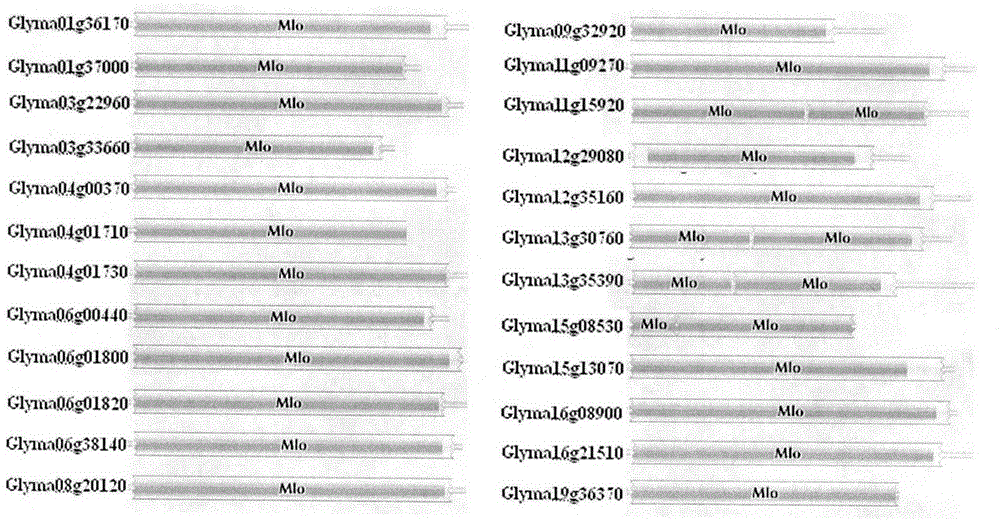
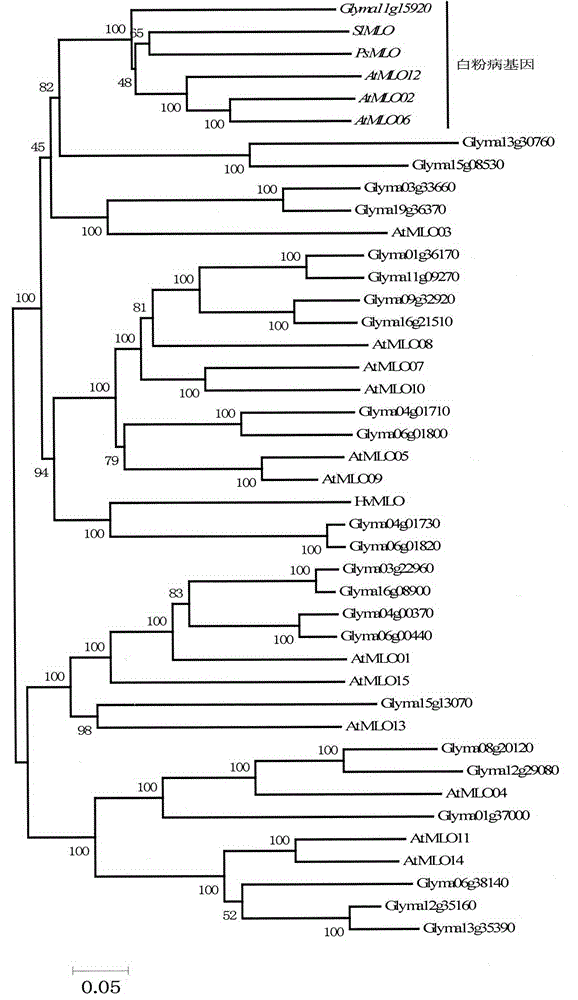
Rapid identification of soybean anti-powdery mildew gene by using candidate gene strategy
InactiveCN104593481AShorten digging cycleImprove identification efficiencyMicrobiological testing/measurementBiostatisticsCandidate Gene Association StudyRapid identification
The invention provides rapid identification of soybean anti-powdery mildew gene, relates to knowledge in the disciplines of plant comparative genomics, genetics and bioinformatics, and belongs to the scientific field of plant biotechnology. The invention includes the steps of: 1) downloading the soybean whole genome sequence and acquiring MLO type gene; 2) identifying the MLO genotype; 3) identifying MLO gene phylogenetic relationship; and 4) comparing MLO-type powdery mildew gene. The invention effectively shortens the gene mining cycle for soybean powdery mildew, and is conducive to the rapid identification of anti-powdery mildew gene; corresponding coseparation functional markers (SNP, scar, etc.) are developed through the identified candidate powdery mildew genes, and the identified candidate powdery mildew genes can also be used for fast molecular marker assisted selection of anti-powdery mildew genes and have high accuracy; and combined with other resistant gene molecular markers, multi-resistance breeding material can be created, so as to shorten the breeding period and improve breeding efficiency. Therefore, the invention lays foundation for the explanation of powdery mildew resistant molecular mechanism of soybean.
Owner:JIANGSU CHANGSHU MODERN AGRI IND PARK DEV
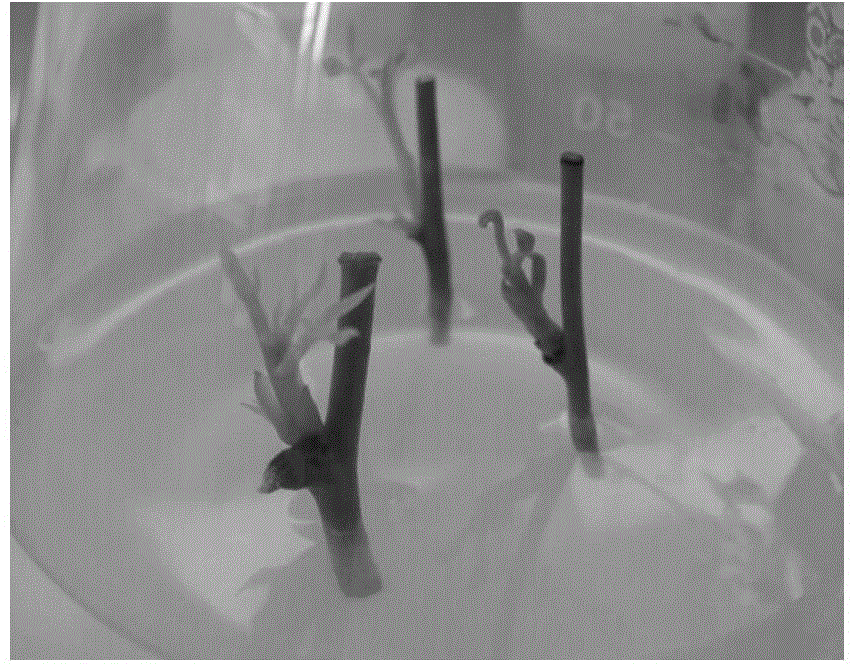
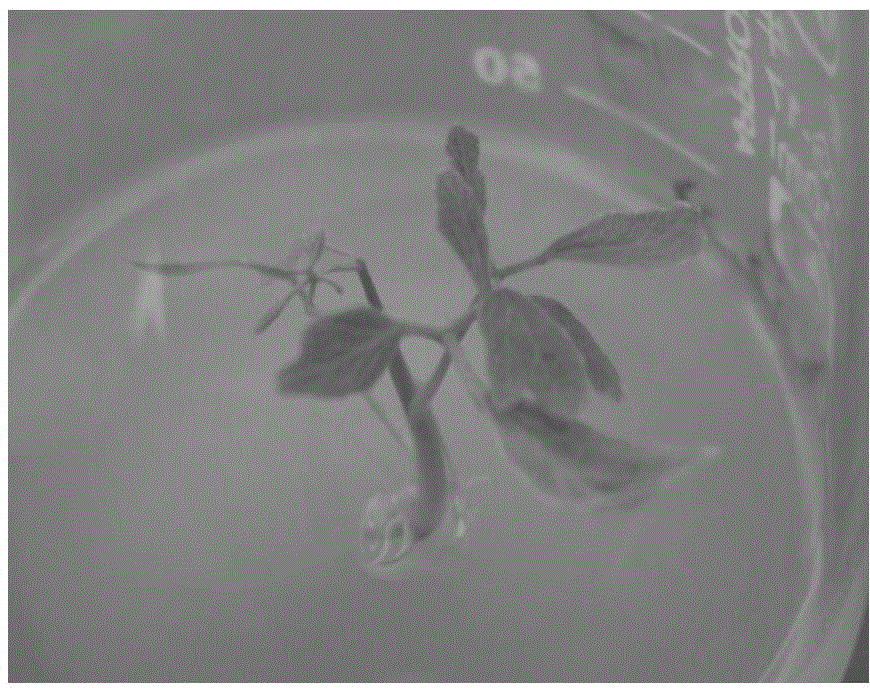
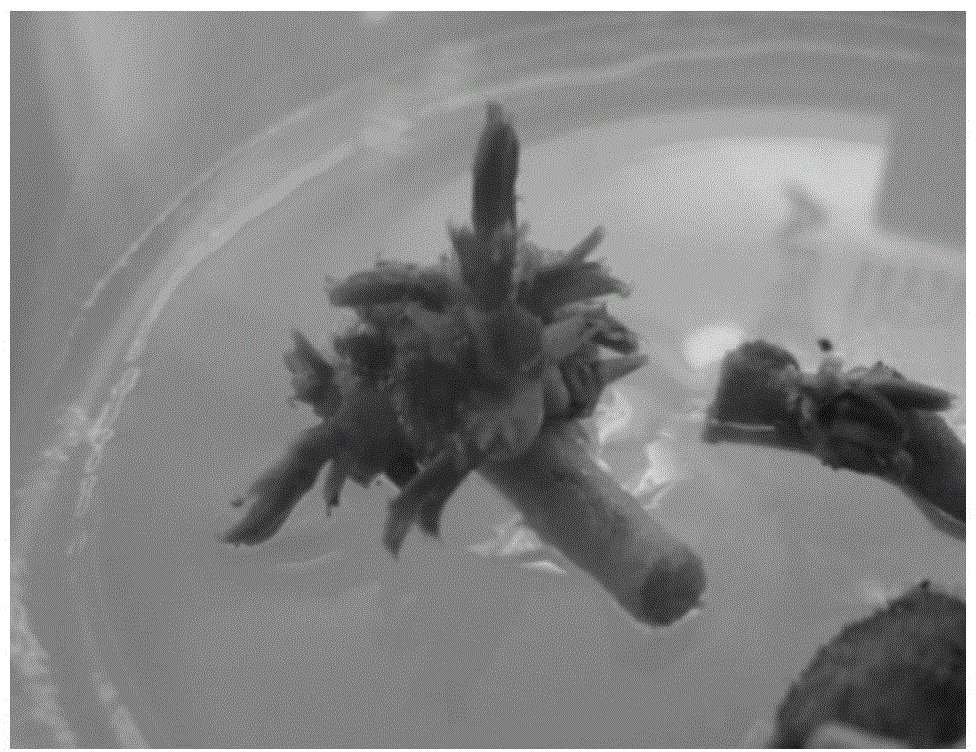
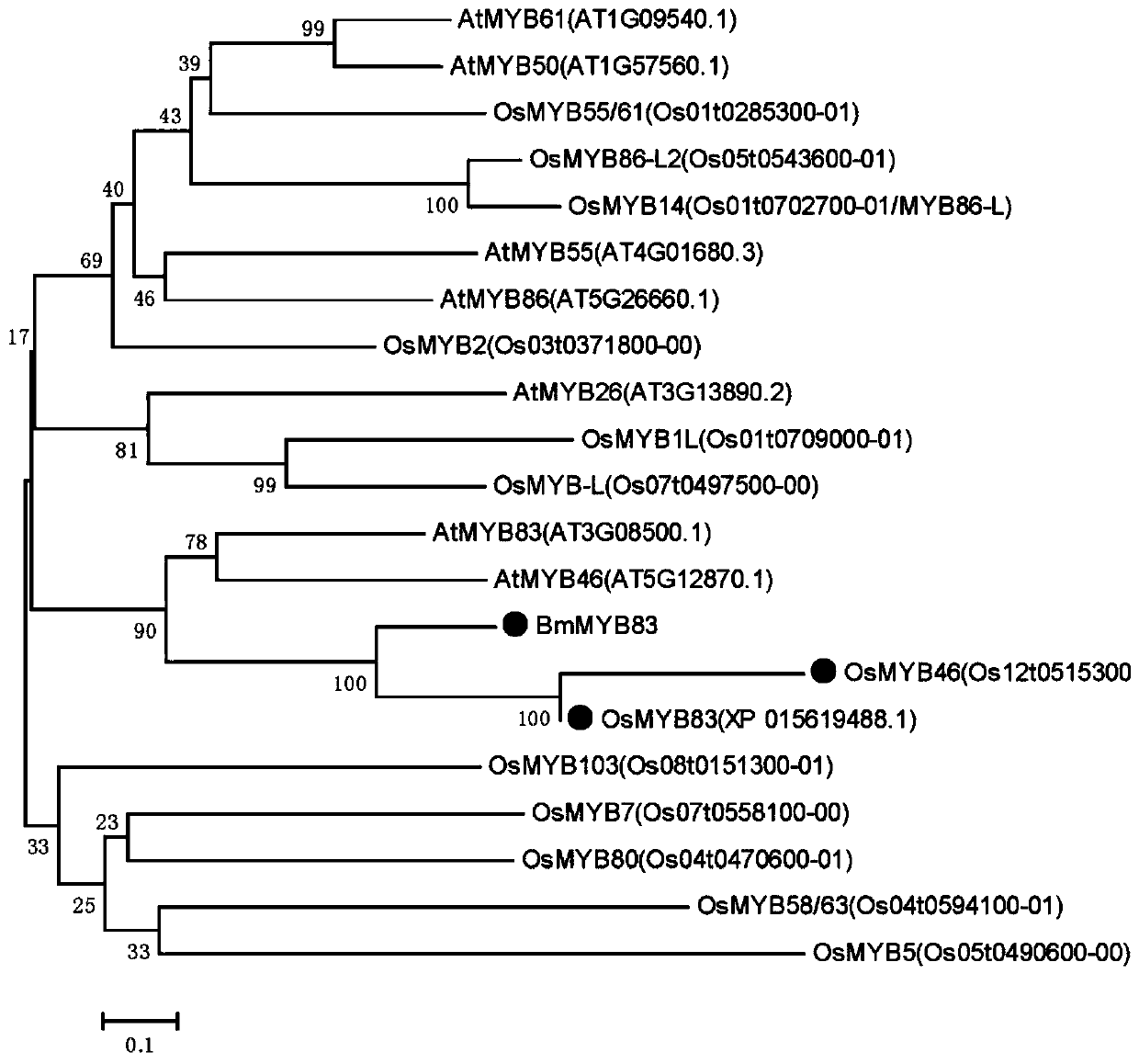
Bambusa multiplex transcription factor BmMYB83 gene and application thereof
The invention discloses a bambusa multiplex transcription factor BmMYB83 gene and application thereof and belongs to the technical field of plant gene engineering. The transcription factor BmMYB83 disclosed by the invention belongs to an MYB transcription factor family, is derived from bambusa multiplex and has a nucleotide sequence shown in SEQ ID NO.1, and the amino acid sequence of a protein encoded by the gene is shown in SEQ ID NO.2. Through gene cloning expression and comparison analysis, the evolutionary status of an encoding gene system of the bambusa multiplex transcription factor BmMYB83 is revealed, a method for using the gene is also provided, plant type variation, such as planting dwarfing, leaf curling and number increase of scapes, of arabidopsis plants caused through over expression of the gene, and root lengths of rice can be also reduced. The gene disclosed by the invention plays a significant role in plant property improvement.
Owner:NANJING FORESTRY UNIV
Protein family phylogenetic analysis method based on amino acid sequence alignment
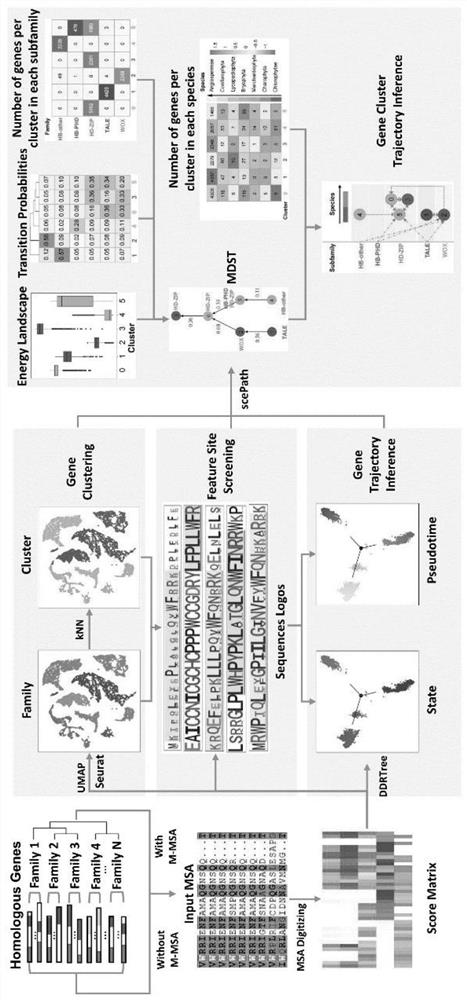
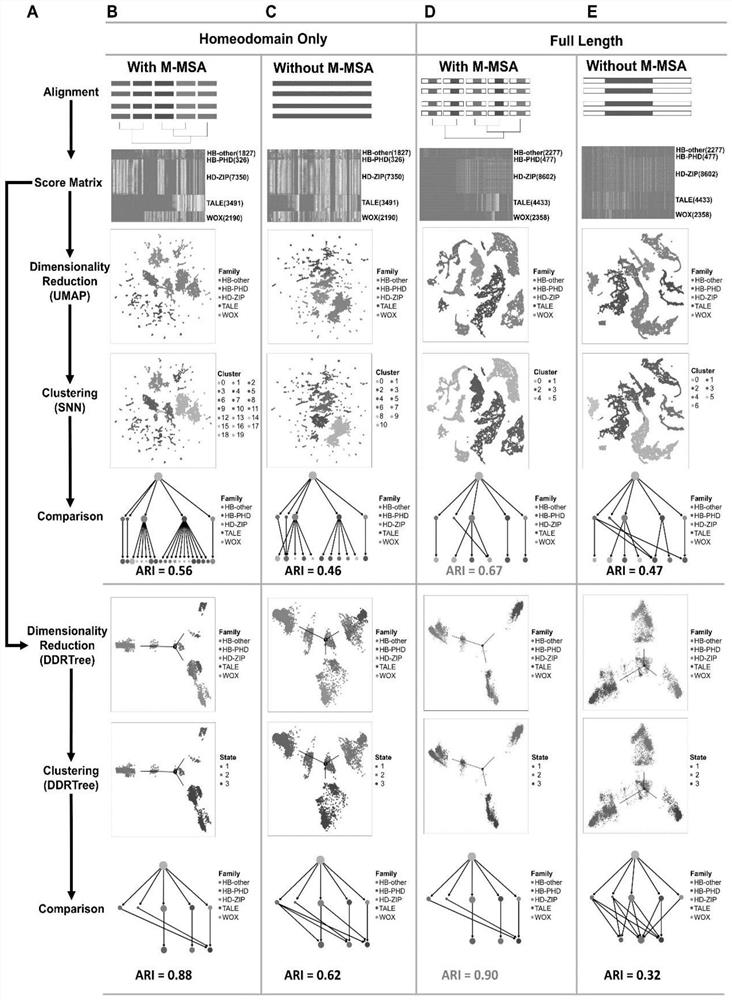
PendingCN114882949AEnsure cluster stabilityClustering is fastCharacter and pattern recognitionProteomicsAmino acid sequence alignmentGene cluster
The invention discloses a protein family phylogenetic analysis method based on amino acid sequence alignment, which comprises the following steps: obtaining a combined multi-sequence alignment result based on an amino acid sequence alignment fusion method; digitizing a combined multi-sequence comparison result, and constructing a fractional matrix; performing dimension reduction and clustering processing on the fractional matrix to obtain an input sequence; identifying a specific site and a conserved site of the input sequence; performing quasi-time analysis on the input sequence to obtain a track sequence of the input sequence; and obtaining a development trajectory of the input sequence based on the trajectory sorting. According to the method, fractional matrix construction and dimension reduction analysis are carried out through the sequence site features, so that the clustering and evolutionary relationship between gene families is deduced, the sequence gene clustering speed is effectively increased under the condition that the sequence clustering stability is guaranteed, and a new tool and method are provided for gene phylogenetic analysis and development trajectory analysis.
Owner:HUAZHONG AGRI UNIV
High-infection quasi-influenza virus preparation as well as special coding sequence and application thereof
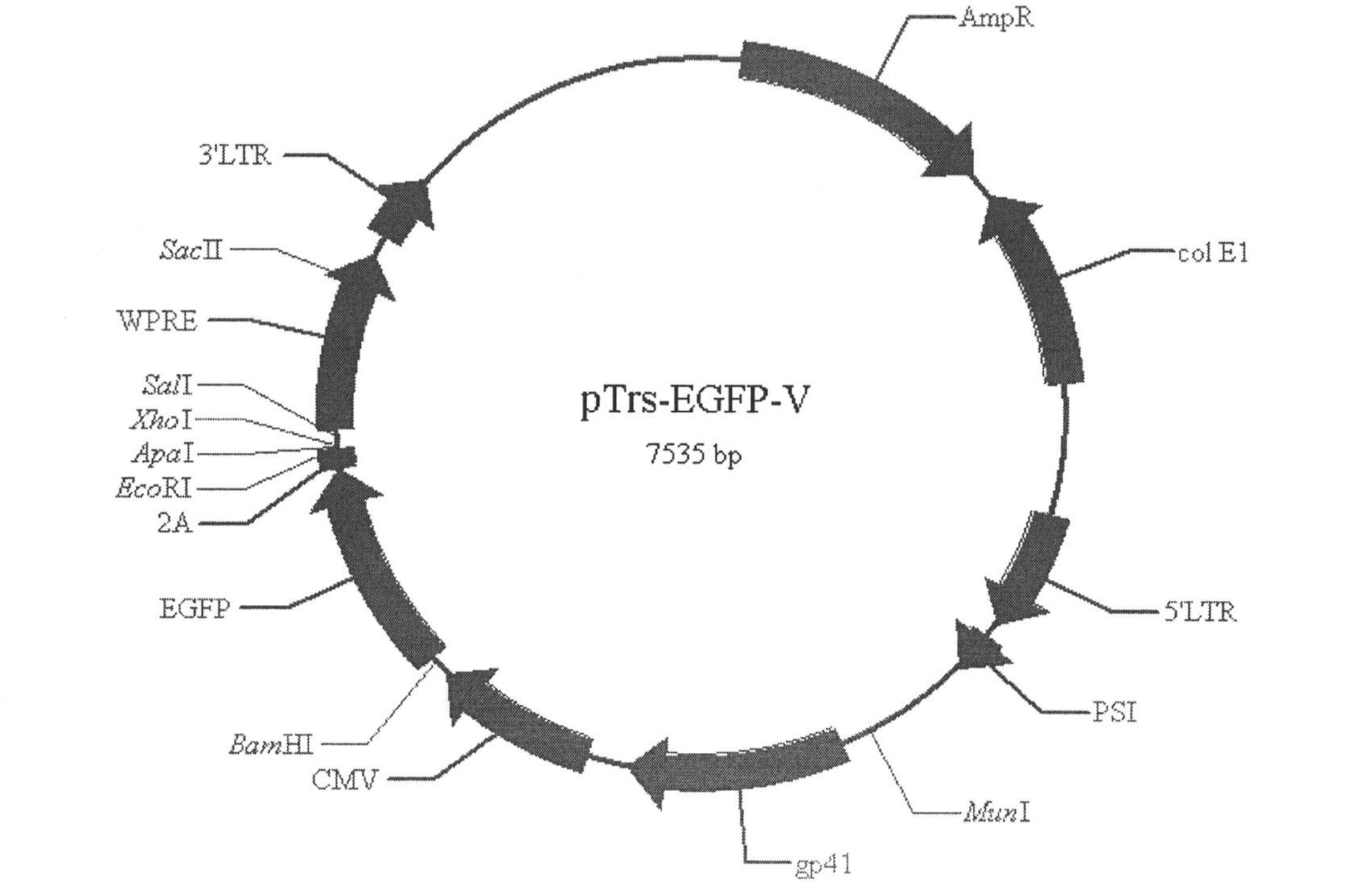
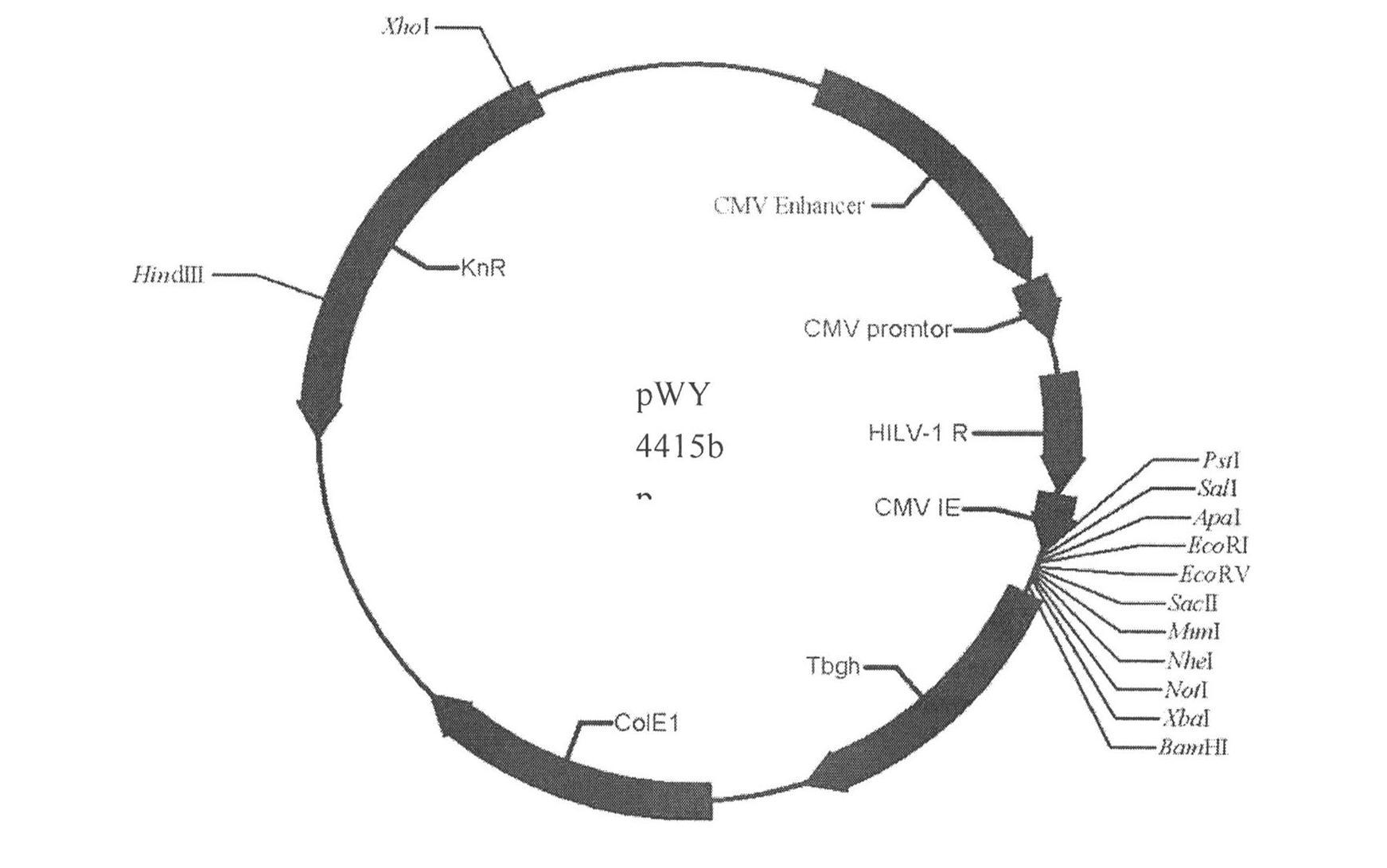
InactiveCN102234635AHigh physiological propertiesConsistent antigenicityMicrobiological testing/measurementVirus peptidesHighly pathogenicViral Membrane Proteins
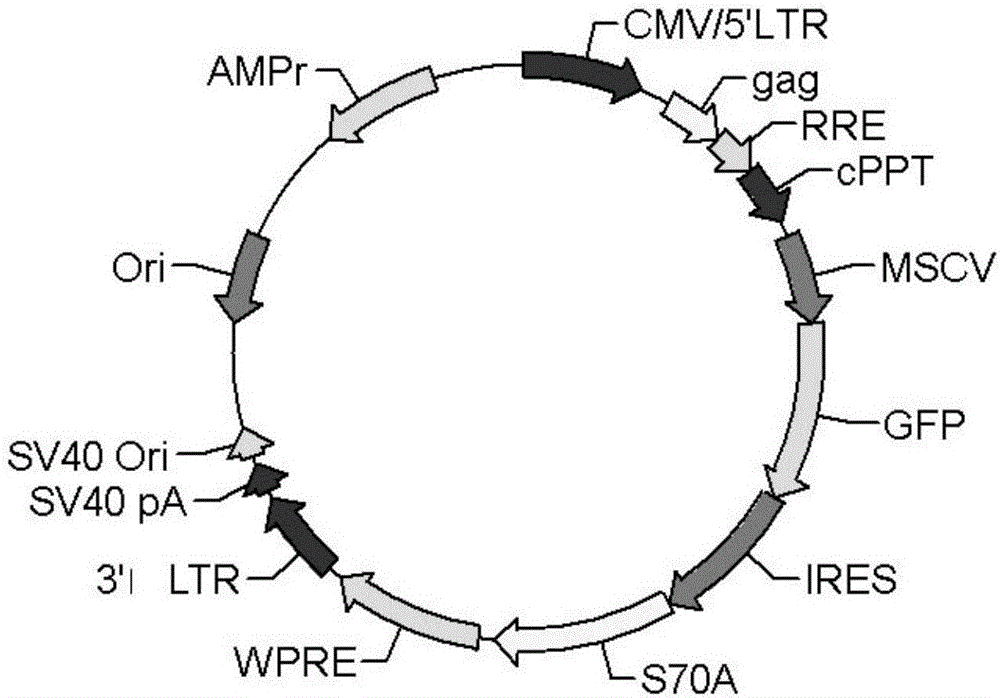
The invention particularly relates to a high-infection quasi-influenza virus system which is mainly characterized in that carriers of LTR(Long Terminal Repeat) containing HIV(Human Immunodeficiency Virus)-1, promoters, gag-pol and other elements and high-infection influenza virus membrane protein expression plasmids are subjected to co-transfection and packaging to form quasi-influenza viruses which have infection characteristics and antigenicity of high-infection influenza virus membrane protein. The system is used as a convenient and safe platform for researches on the physiological characteristics of highly pathogenic influenza virus membrane protein and researches on neutralizing antibodies. The system is used as an efficient gene introduction carrier, a target gene can be inserted into a gene-containing system reconstructed through packaging signals of the HIV-1, and the target gene can be introduced into various cells through quasi-virus infection according to retrovirus integration rules. After the cells are infected by quasi-viruses, reporter genes are integrated and expressed through gag-pol protein of the HIV-1, and the inhibition to the gag-pol function by medicaments can be reflected through the expression of the reporter genes, thus the system can be used as an evaluation platform for gag-pol medicaments.
Owner:中国疾病预防控制中心病毒病预防控制所