Patents
Literature
60results about How to "Achieve freedom of choice" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
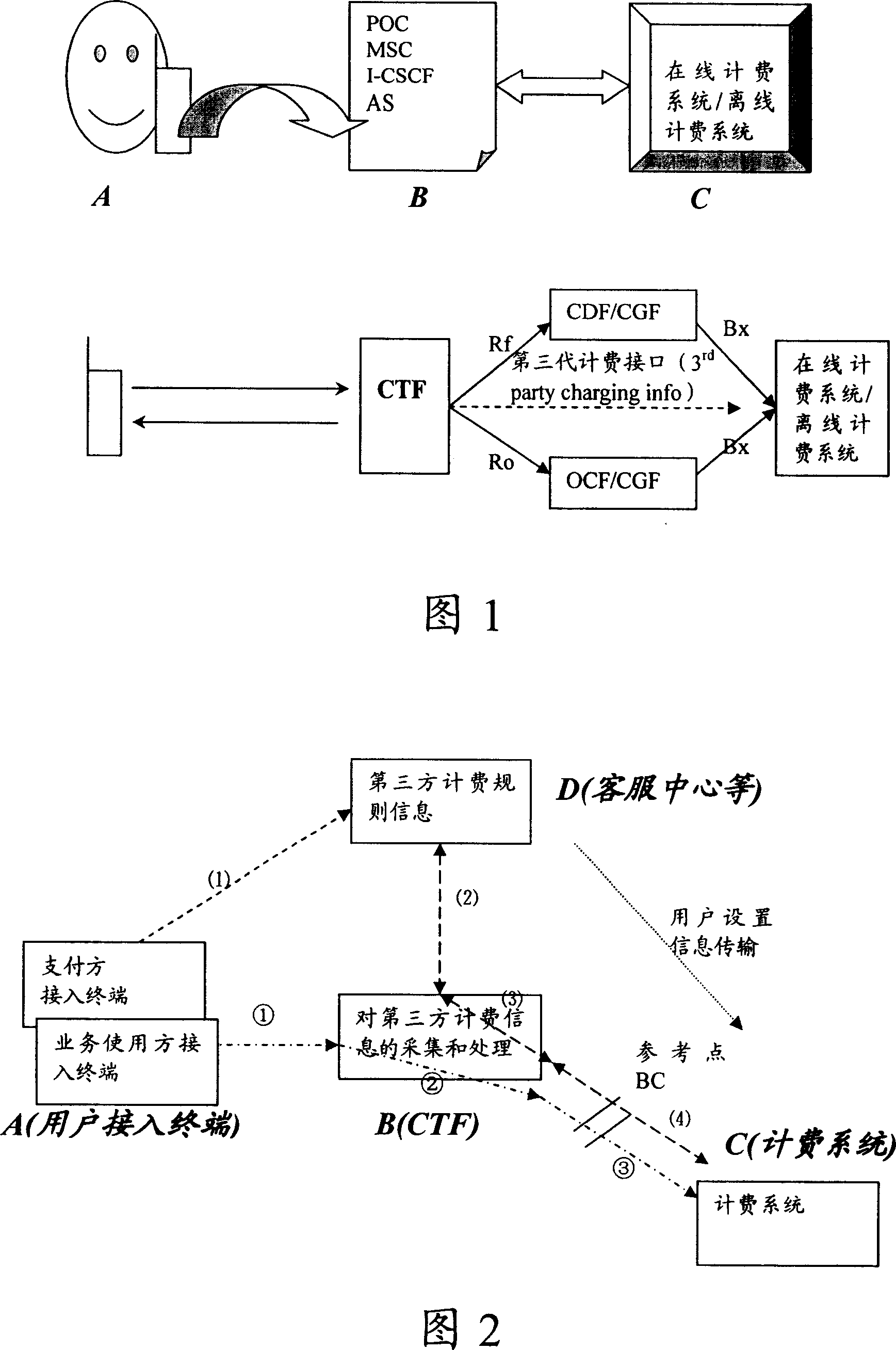
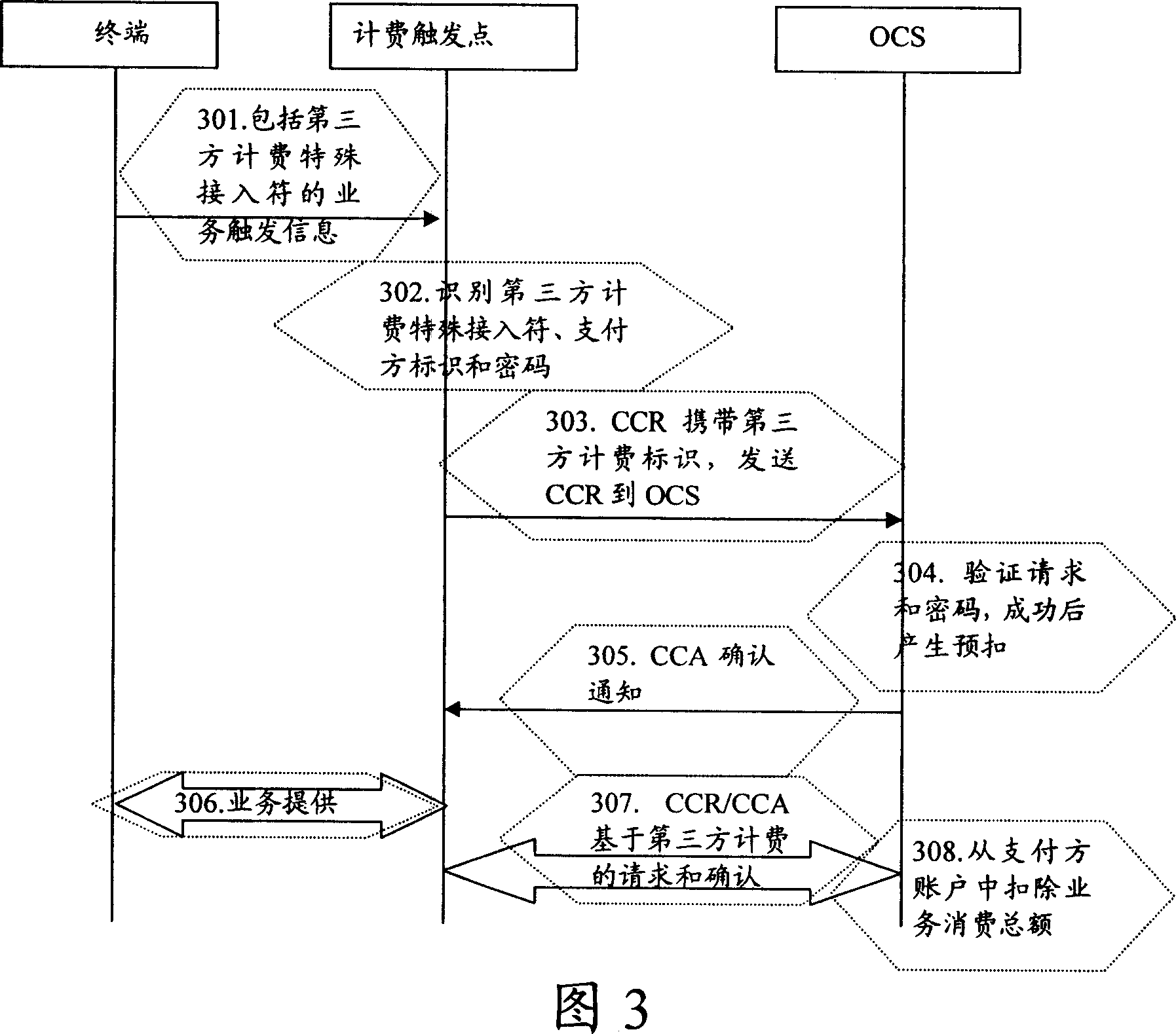
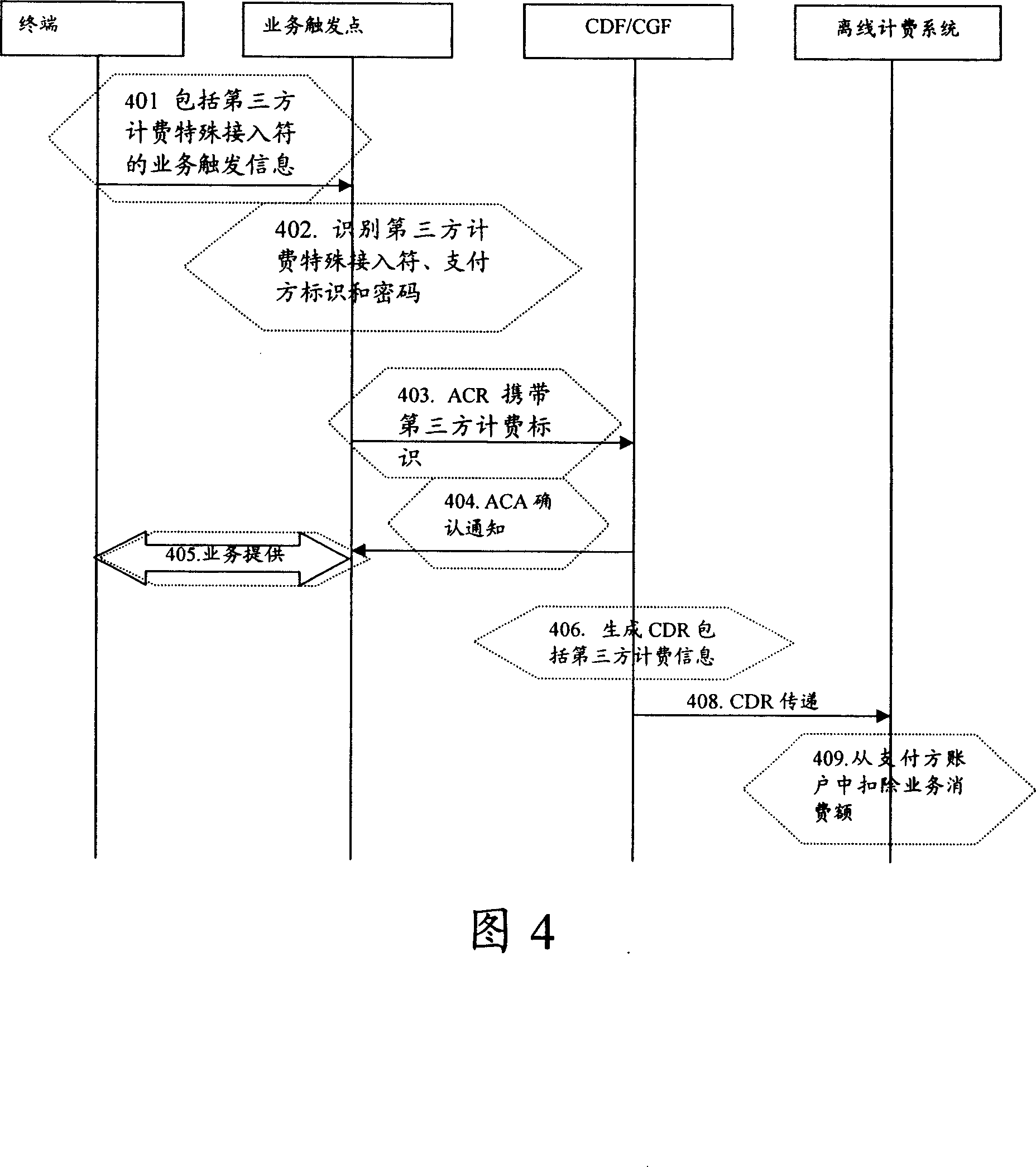
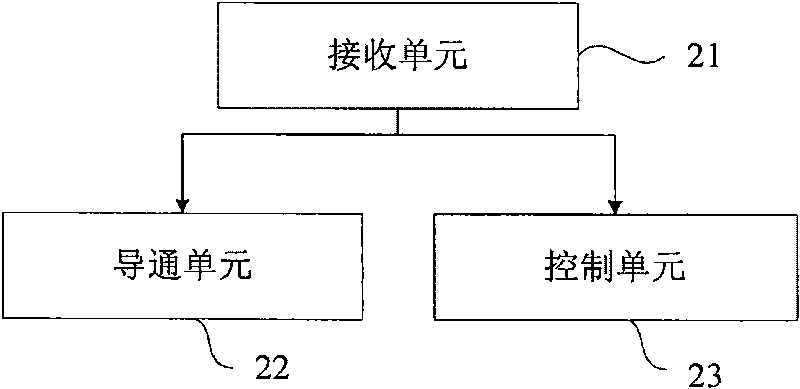
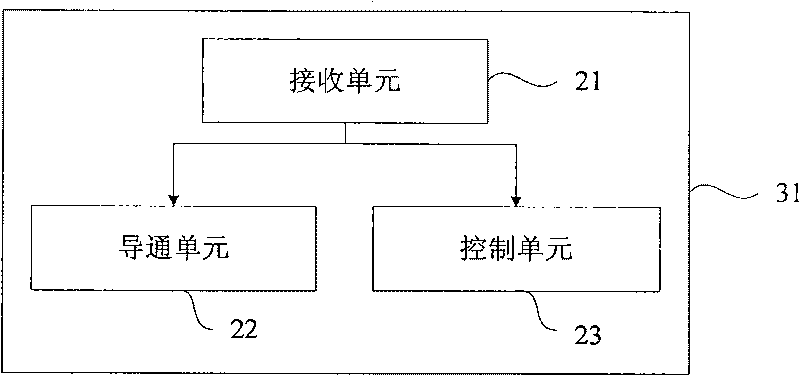
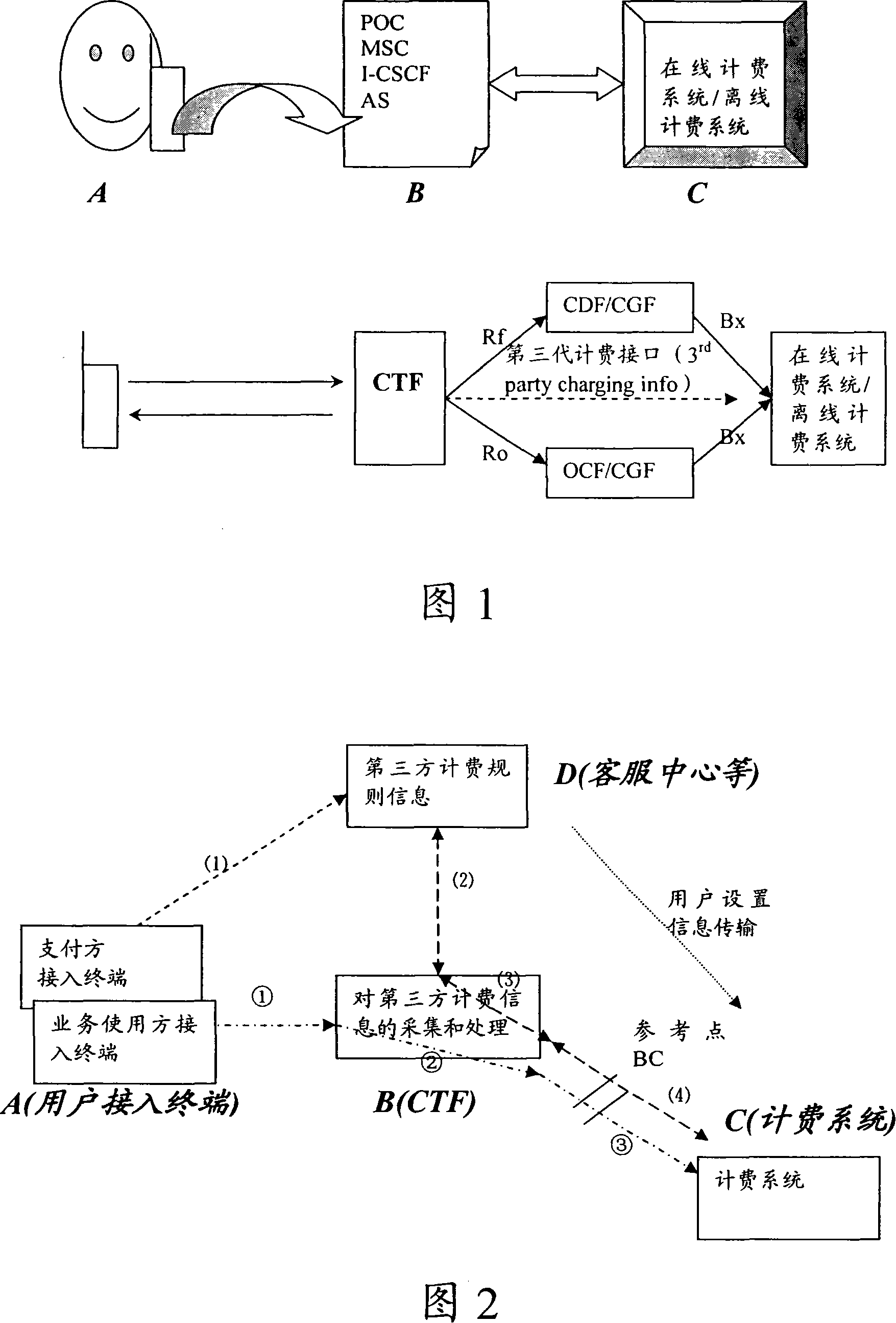
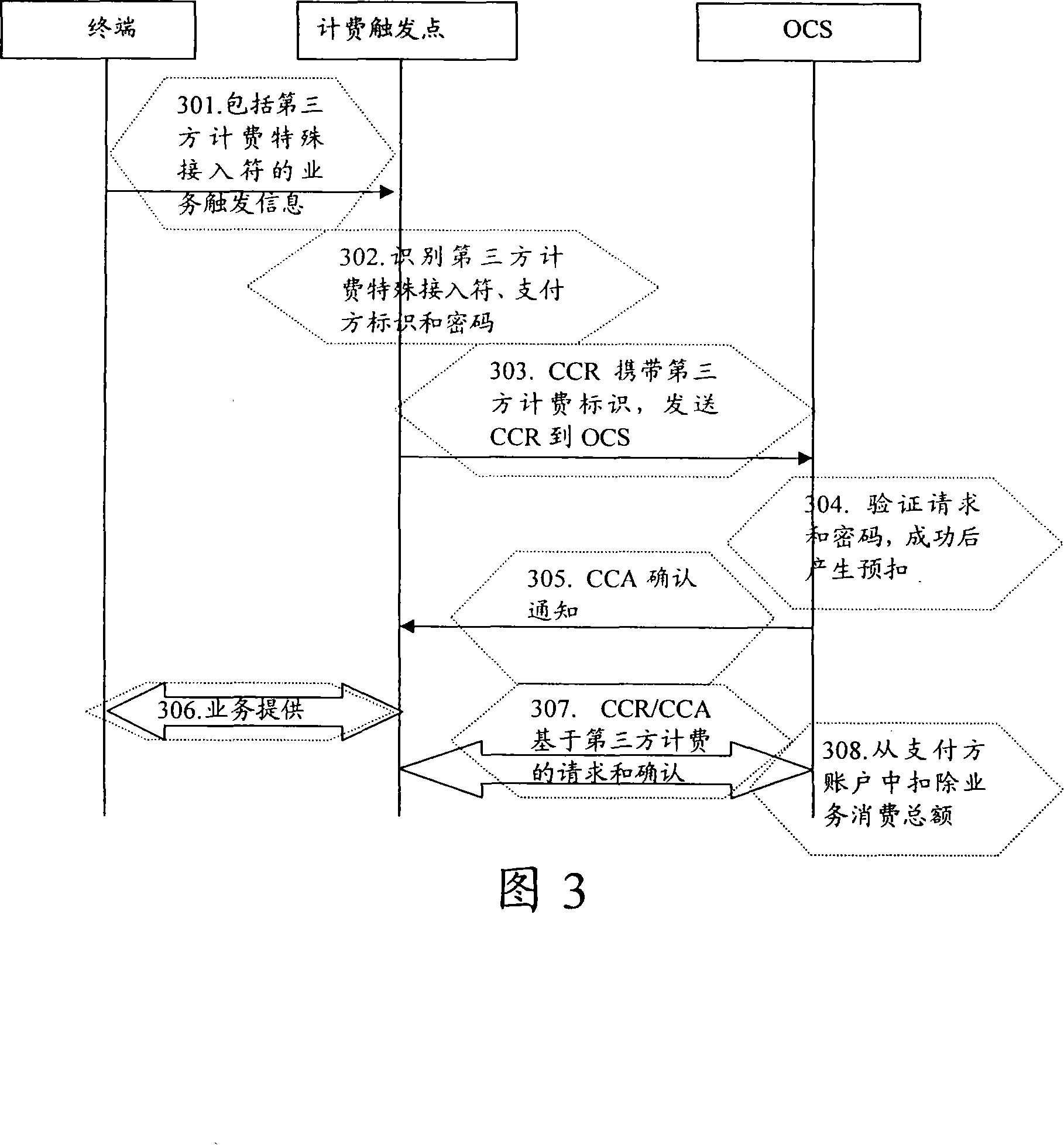
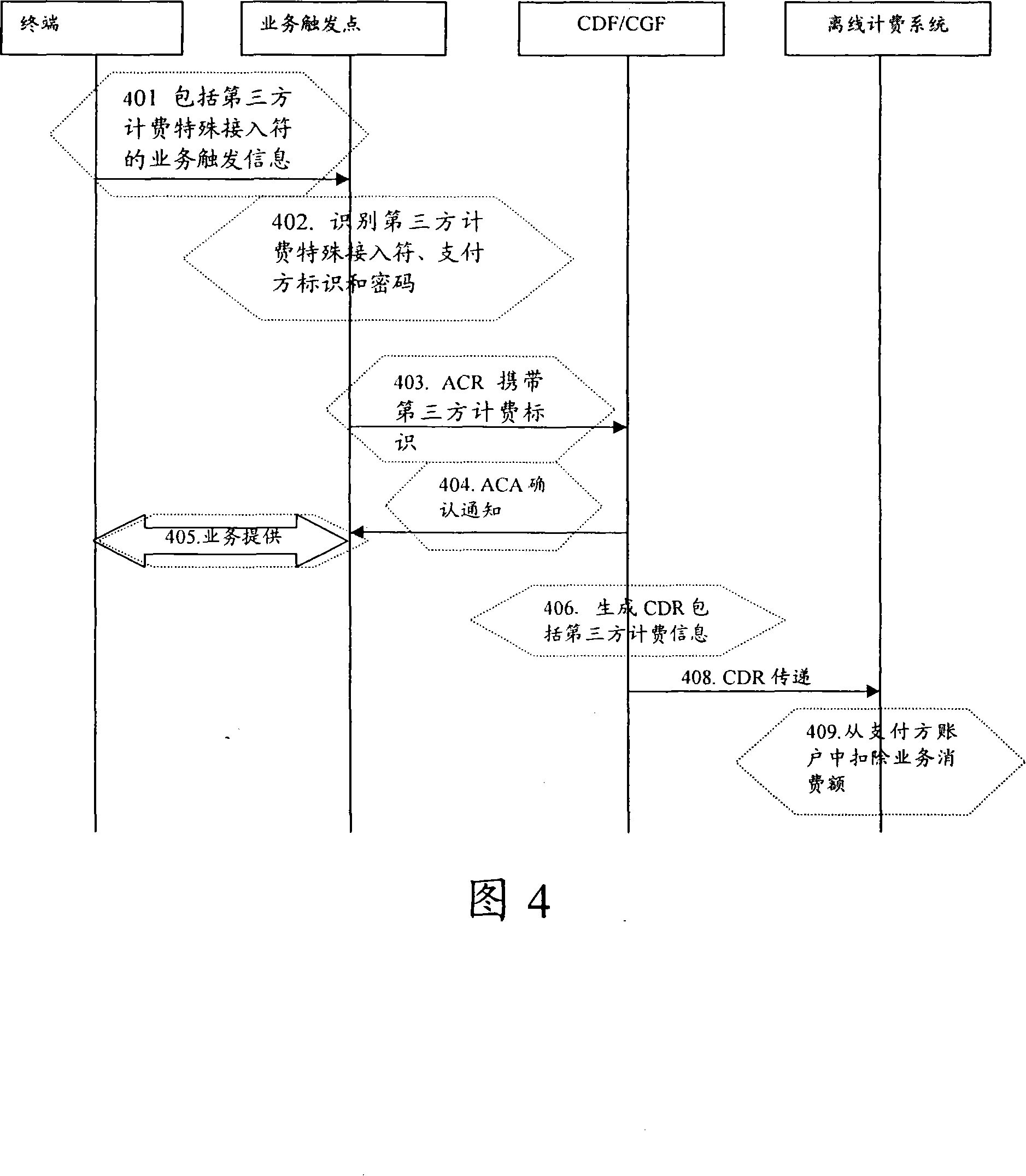
Third party charge method and system
InactiveCN101090325AFlexible third-party billingEnsure safetyMetering/charging/biilling arrangementsTelephonic communicationBilling systemInternet privacy
This invention discloses a method for charging to a third party including: a paying party sets its own payment line and cipher and shares related information to a service using party, which inputs a special access symbol, ID of the paying party or cipher for charging to the third party, a service applied server judges if the attribute of the user is the third party charging, if it is the third party charging service request needing to pay by other account, it collects its information such as MSISDN, account number information or cipher of the paying party and transmits it to a charge system via a charge access protocol, the charge system deducts the fee from the third party account according to the information. This invention also discloses another kind of method for charging to a third party and a system.
Owner:HUAWEI TECH CO LTD
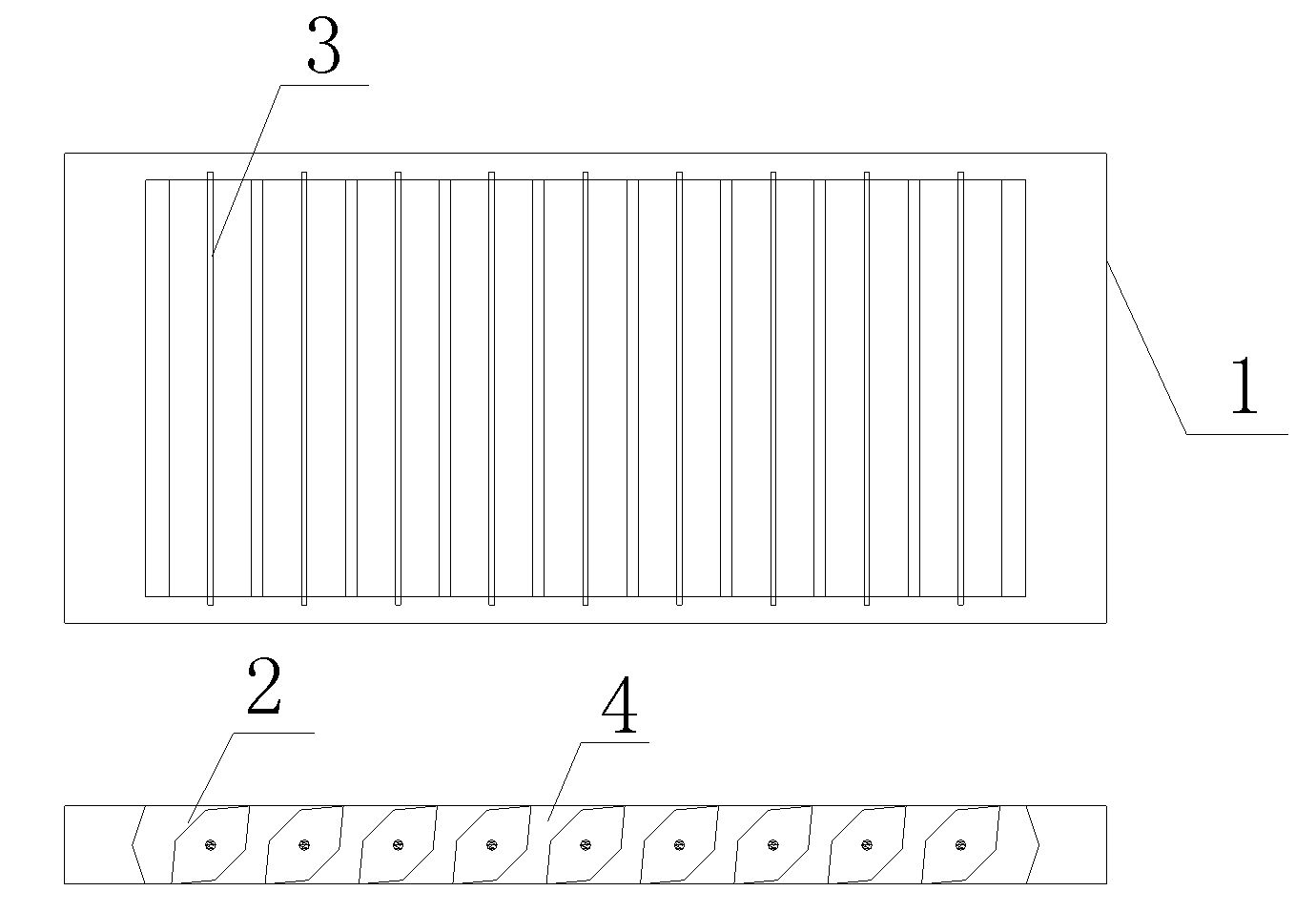
Ventilation pressure leakage sound barrier module
ActiveCN101858061AImprove noise reductionChange directivityNoise reduction constructionSound barrierFree rotation
The invention discloses a ventilation pressure leakage sound barrier module, belonging to the noise reduction field. The module comprises a ring-shaped rigidity frame, a plurality of regulation screws and noise elimination vanes which correspond to the regulation screws and can freely rotate around the regulation screws; the noise elimination vanes are arranged in the ring-shaped frame regularly to form a shutter structure which can freely rotate. The module of the invention adopts the rotatable design of the noise elimination vanes, can reach more excellent ventilation noise elimination effect by regulating the wind facing angle of the module, realize free selection of different noise elimination ranges and ventilation amount, and simultaneously also changes the direction of noise radiation; in addition, the rotatable design of the noise elimination vanes can lead the module to automatically regulate the ventilation angle of the noise elimination vanes according to different travelling directions of vehicles, thus achieving the best noise elimination and pressure leakage effects.
Owner:ZISEN ENVIRONMENTAL TECHNOLOGY CO LTD
Micro light emitting diode transfer substrate and method, display panel and preparation method
ActiveCN109449260ARealize batch transferSimplify the transfer processSolid-state devicesSemiconductor devicesProcess complexityLight-emitting diode
The invention discloses a micro light emitting diode transfer substrate and method, a display panel and a preparation method, and aims at simplifying the process complexity of micro light emitting diode preparation. The micro light emitting diode transfer substrate comprises a first substrate and an electrically induced deformation layer which is located on the first substrate, wherein the electrically induced deformation layer is used for ensuring that the thickness of the electrically induced deformation layer in a to-be-transferred area of a to-be-transferred micro light emitting diode is greater than the thickness of the electrically induced deformation layer in a not-to-be-transferred area corresponding to the micro light emitting diode.
Owner:BOE TECH GRP CO LTD
Image processing method and mobile terminal
ActiveCN106780401AAvoid duplicationAchieve freedom of choiceImage enhancementModifying/creating image using manual inputImaging processingComputer graphics (images)
The invention provides an image processing method and a mobile terminal. The method comprises the steps of obtaining a source image and a plurality of target image processing effect templates; processing the source image according to the target image processing effect templates, and obtaining a plurality of images with different target image processing effects; according to the images with different target image processing effects, generating target images with a plurality of images, and receiving an adjustment trigger operation on the first image in the images in the target images by a user; according to the adjustment trigger operation, putting the first image in a to-be-adjusted state; receiving adjusting operations on the first image by the user, and adjusting the display state of the first image according to the adjusting operations. The process makes the processed images have a repetitive beauty, avoids repetitive operations of the user, and simplifies the operation process.
Owner:VIVO MOBILE COMM CO LTD
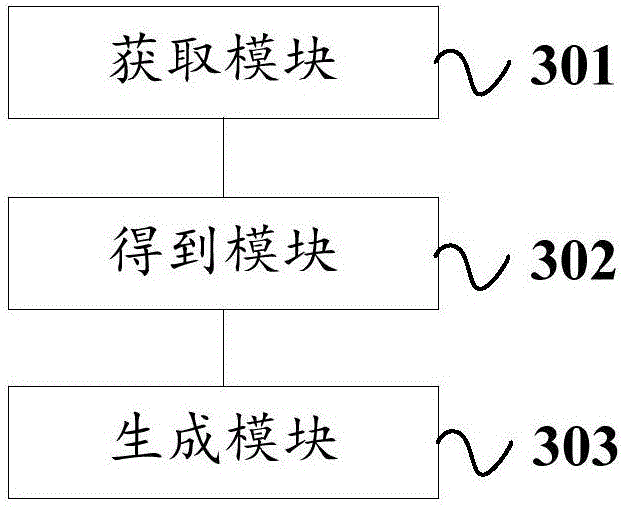
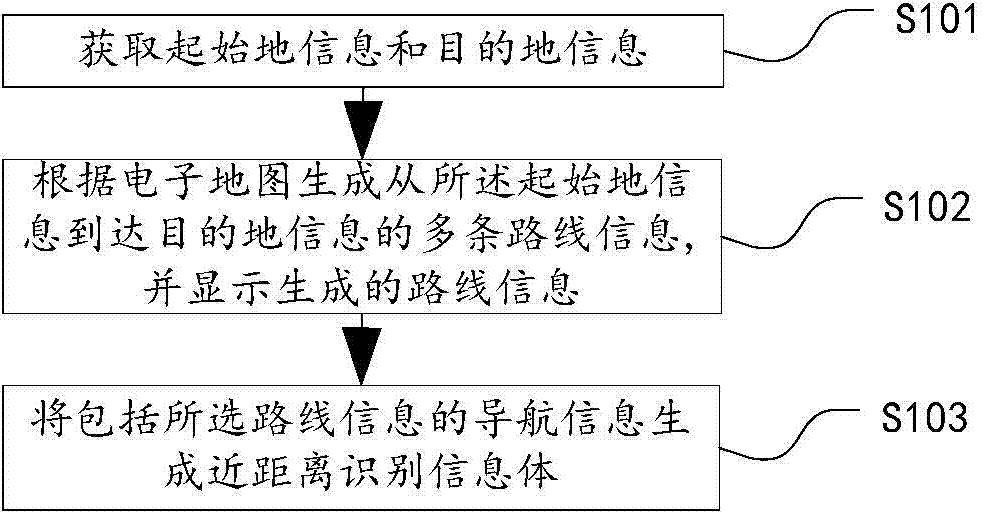
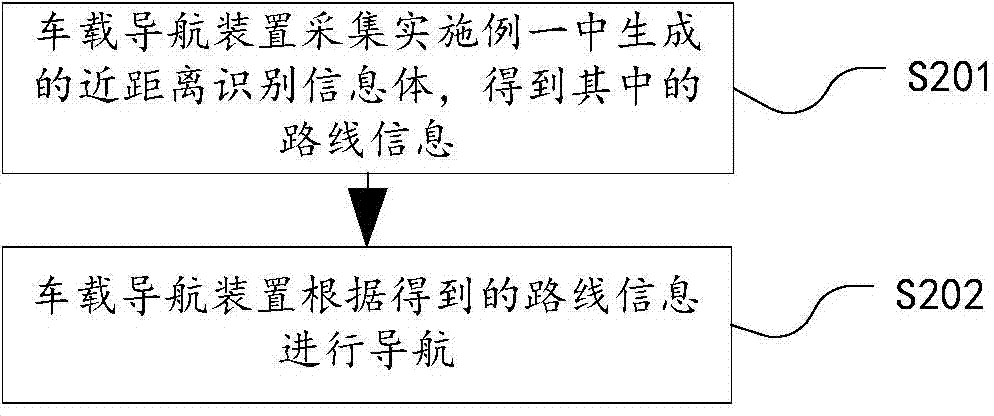
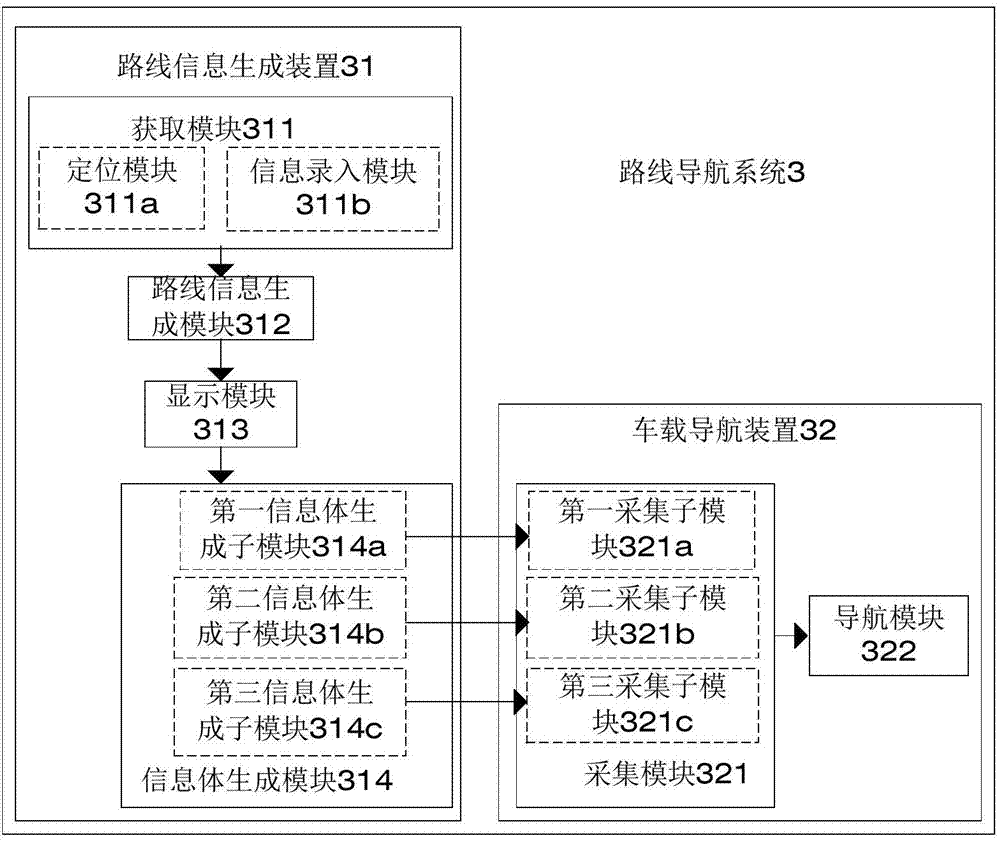
Route information generation, navigation method, device and system
InactiveCN103575286AImprove navigation accuracyImprove user experienceInstruments for road network navigationElectronic mapInformation transfer
The invention discloses a route information generation, navigation method, device and system. The route information generation method comprises the following steps: acquiring originating location information and destination location information; generating information on a plurality of routes from the originating location information to the destination location information according to an electronic map and displaying; generating short-distance recognition information body from the navigation information including the information on the selected route. The route navigation method comprises the following steps: acquiring the short-distance recognition information body by a vehicle-mounted navigation device to obtain route information therein; navigating the obtained route information. According to the technical scheme, the technical problem that the route navigation method is not complete in the prior art is solved, on one hand, the free selection of the routes by a user is realized, particularly under the condition of taking a taxi, on the other hand, the route information selected by the user can be conveniently transmitted to other equipment capable of collecting the short-distance recognition information body, such as to the vehicle-mounted navigation device, so that the vehicle-mounted navigation device can acquire the navigation route from the route information generation device.
Owner:YULONG COMPUTER TELECOMM SCI (SHENZHEN) CO LTD
Charging system and method
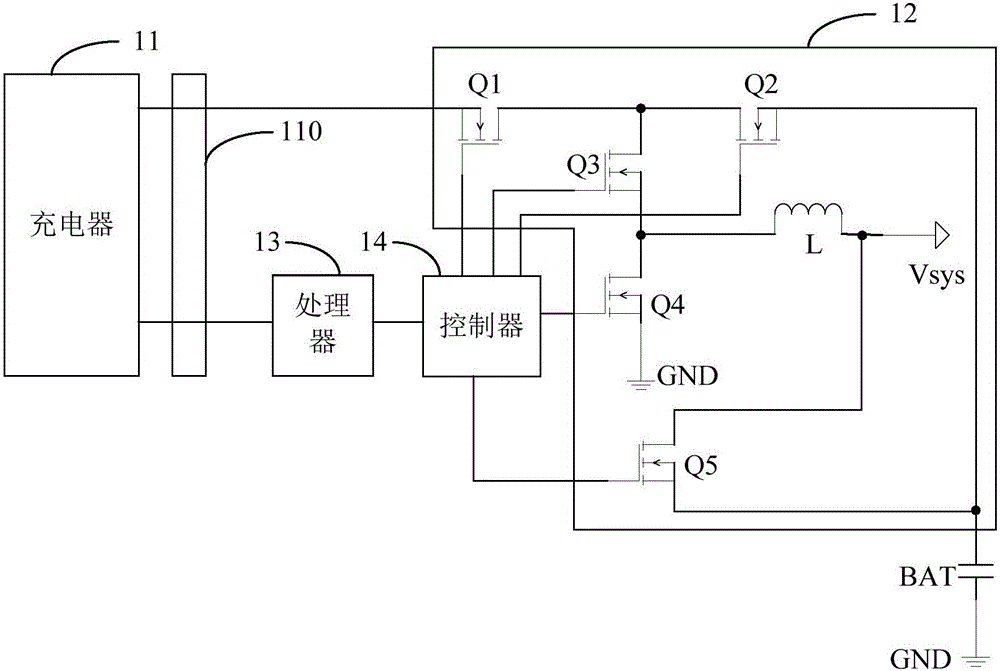
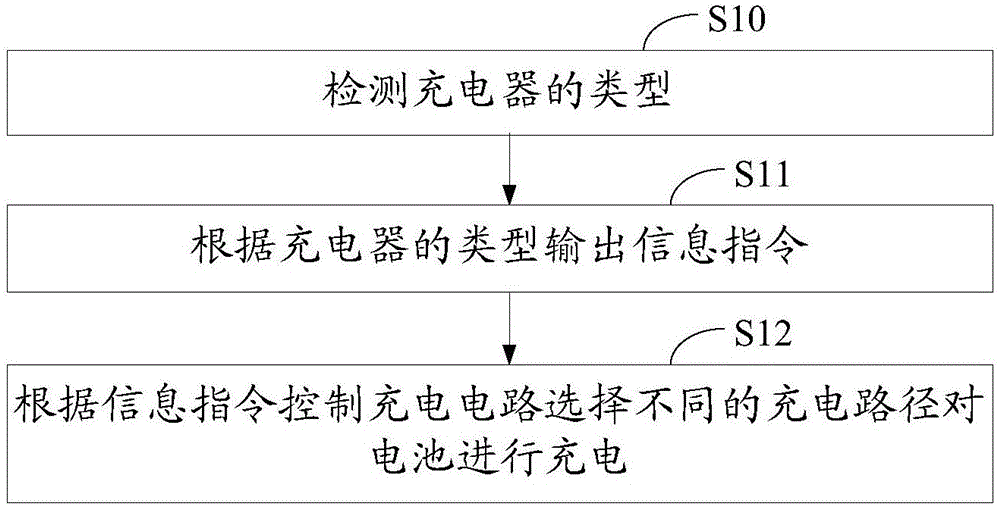
InactiveCN106786838AAchieve freedom of choiceCircuit authenticationElectric powerBattery chargerElectrical and Electronics engineering
The invention discloses a charging system and method. The charging system comprises a charger, a charging circuit, a processor and a controller, wherein the charger is used for outputting a charging signal and charging a battery; the charging circuit is connected with the charger; the charging signal is used for charging the battery through the charging circuit; the processor communicates with the charger and is used for identifying the type of the charger and outputting an information instruction according to the type of the charger; and the controller is connected with the processor and the charging circuit and is used for controlling the charging signal to charge the battery through different charging paths in the charging circuit according to the information instruction. According to the charging system and method, target of being simultaneously compatible with two charging schemes can be achieved through identifying the type of the charger and controlling charging through different charging paths, and free selection of the charger is achieved.
Owner:SHENZHEN TINNO WIRELESS TECH
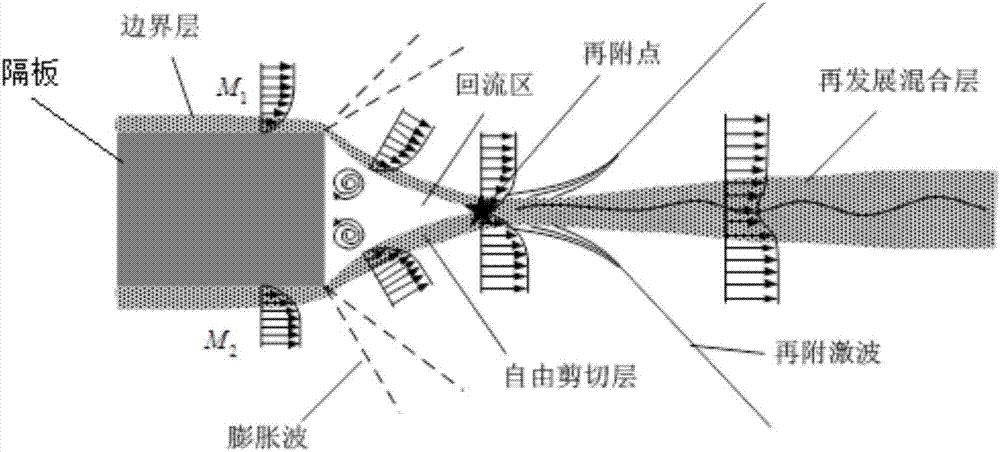
Active control device of supersonic blunt trailing edge mixed layer
The invention provides an active control device of a supersonic blunt trailing edge mixed layer. By arranging a jet flow active control structure in the trailing edge of a partition plate, active control and passive control are combined, the problem that as for active control, vibrating plates are prone to melting is avoided, and various control modes can be freely selected. The active control device achieves the effects that active control and passive control are combined and complement each other.
Owner:NAT UNIV OF DEFENSE TECH
Program on-demand system and method based on one-way set top box
ActiveCN102387406AImprove the digital TV experienceIncrease incomeSelective content distributionNetwork terminationBroadcasting
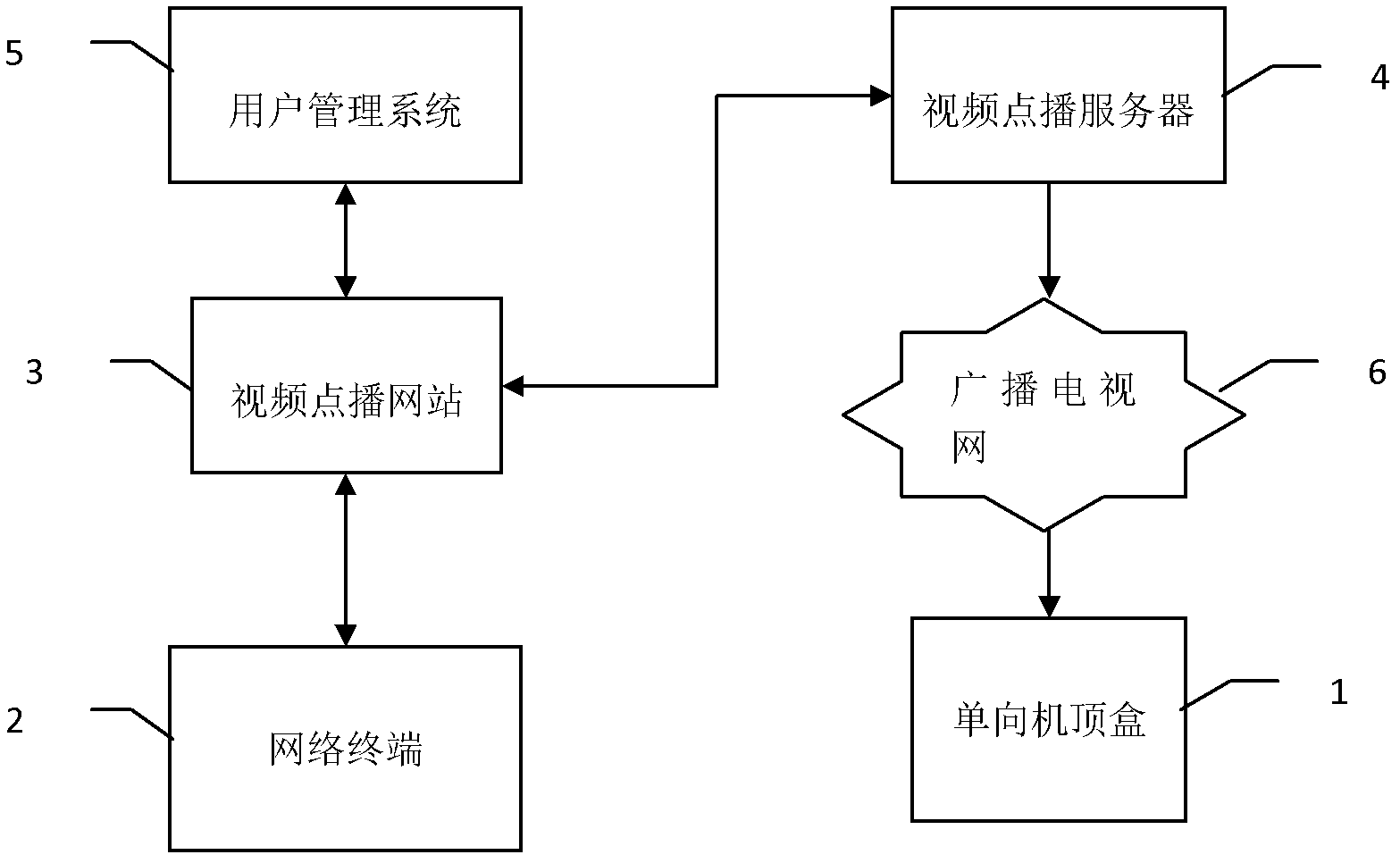
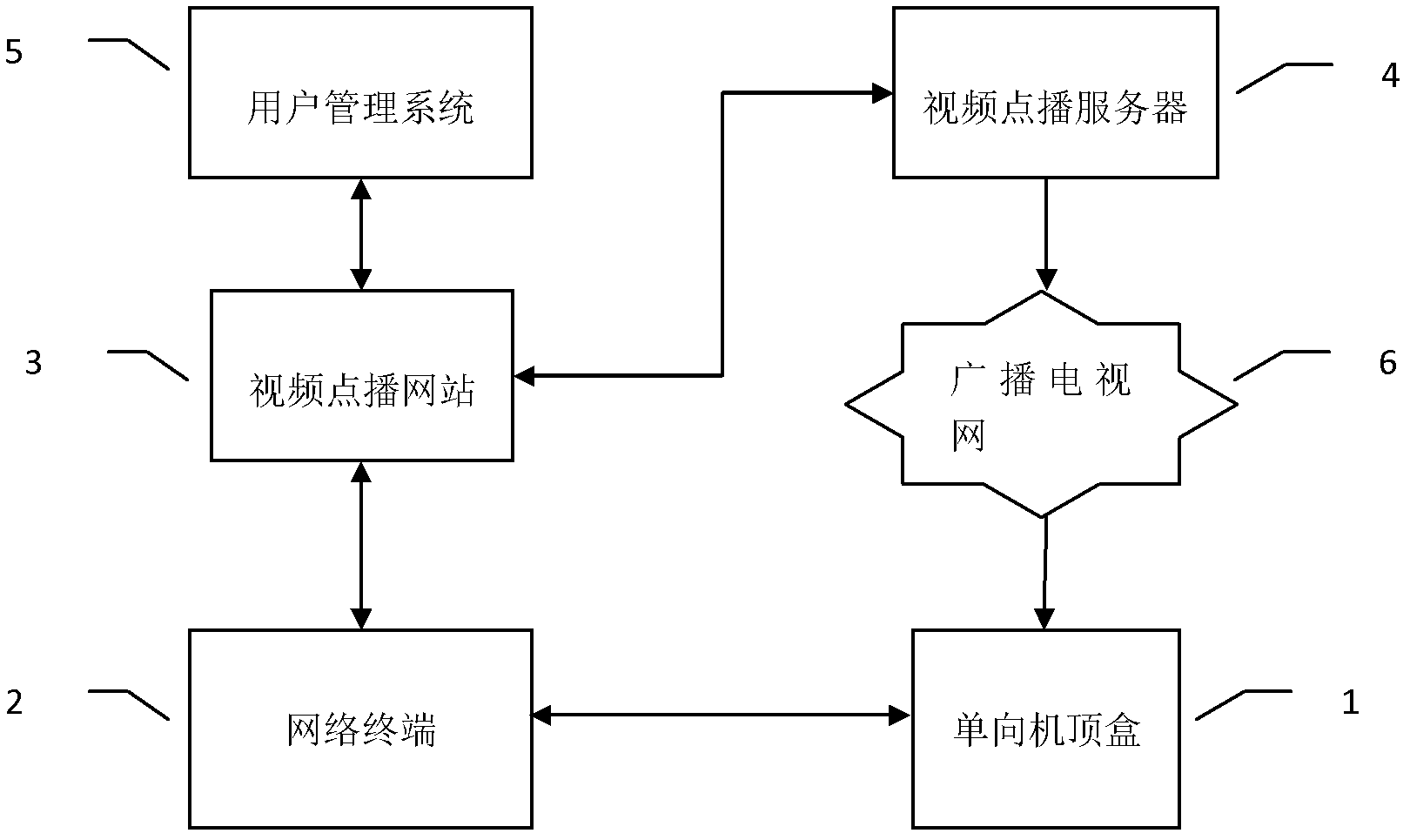
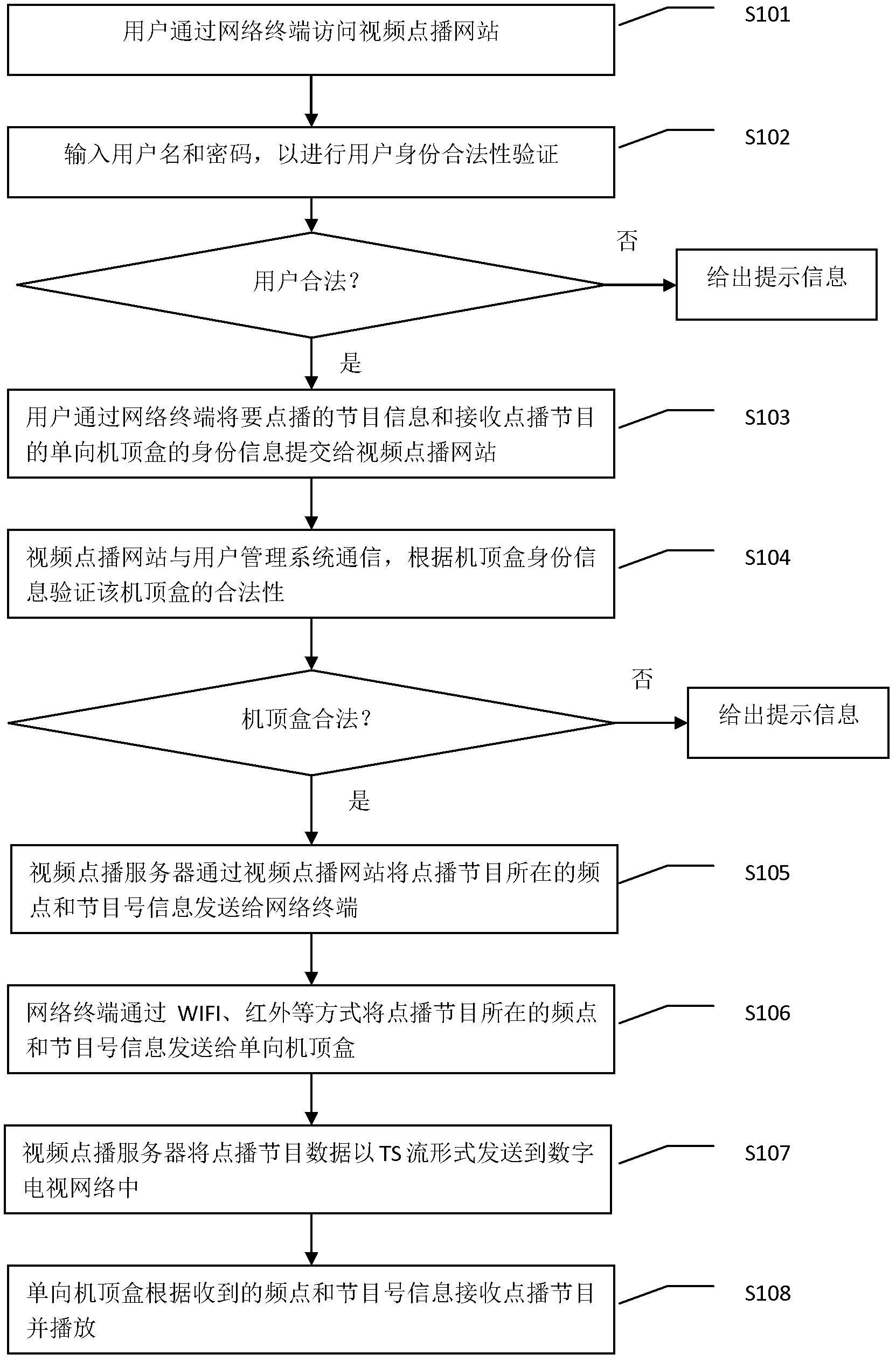
The invention discloses a program on-demand system and method based on a one-way set top box. The program on-demand system comprises the one-way set top box, a network terminal, a video on-demand website, a video on-demand server, a user management system and a broadcast television network, wherein the network terminal carries out two-way communication with the video on-demand website through an IP (Internet Protocol); the video on-demand website carries out two-way communication with the video on-demand server; meanwhile, the video on-demand website also carries out two-way communication with the user management system; and the video on-demand server carries out one-way communication with the one-way set top box through the broadcast television network. According to the program on-demand system and method disclosed by the invention, on-demand information of the one-way set top box is transmitted to the video on-demand website through the network terminal and on-demand program is transmitted to the one-way set top box through the video on-demand server, so that a program on-demand function of the one-way set top box is achieved; upgrade of set top box hardware is avoided, so that the popularization cost of video on-demand services is reduced; and a user and the set top box are legally verified so as to ensure the security of the video on-demand services.
Owner:SHANDONG TAIXIN ELECTRONICS CO LTD
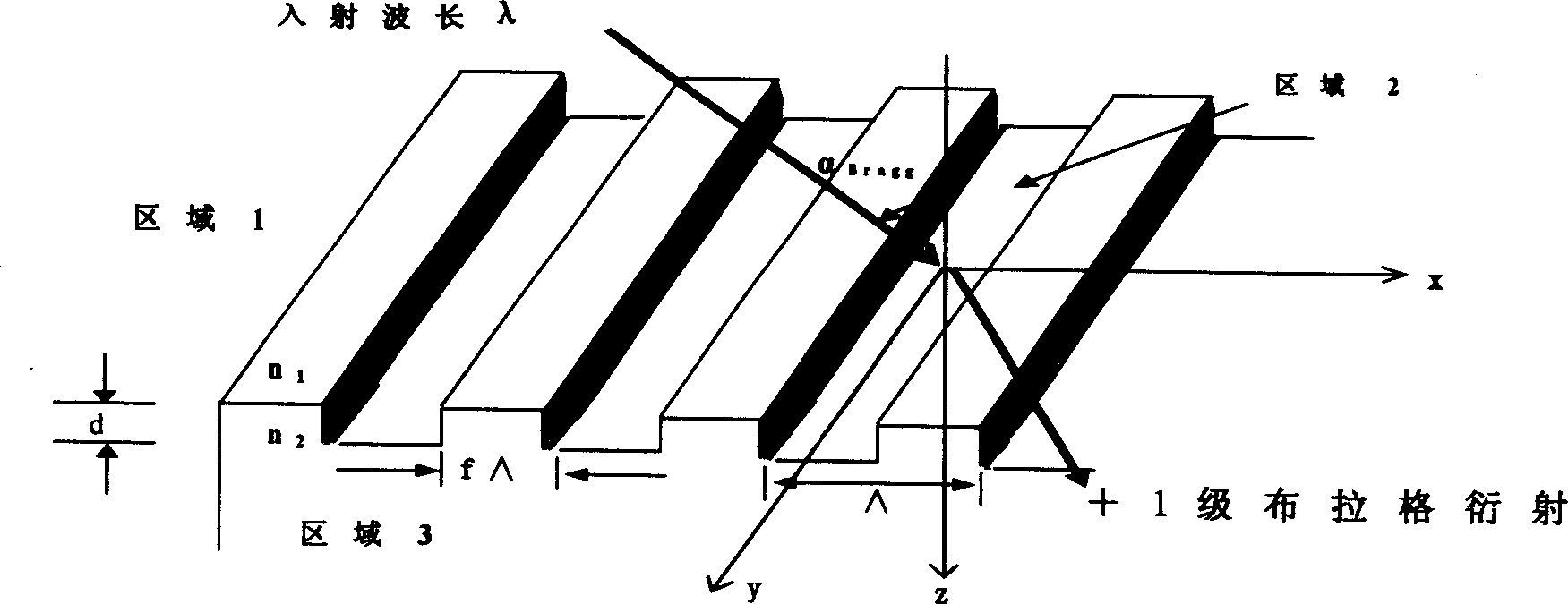
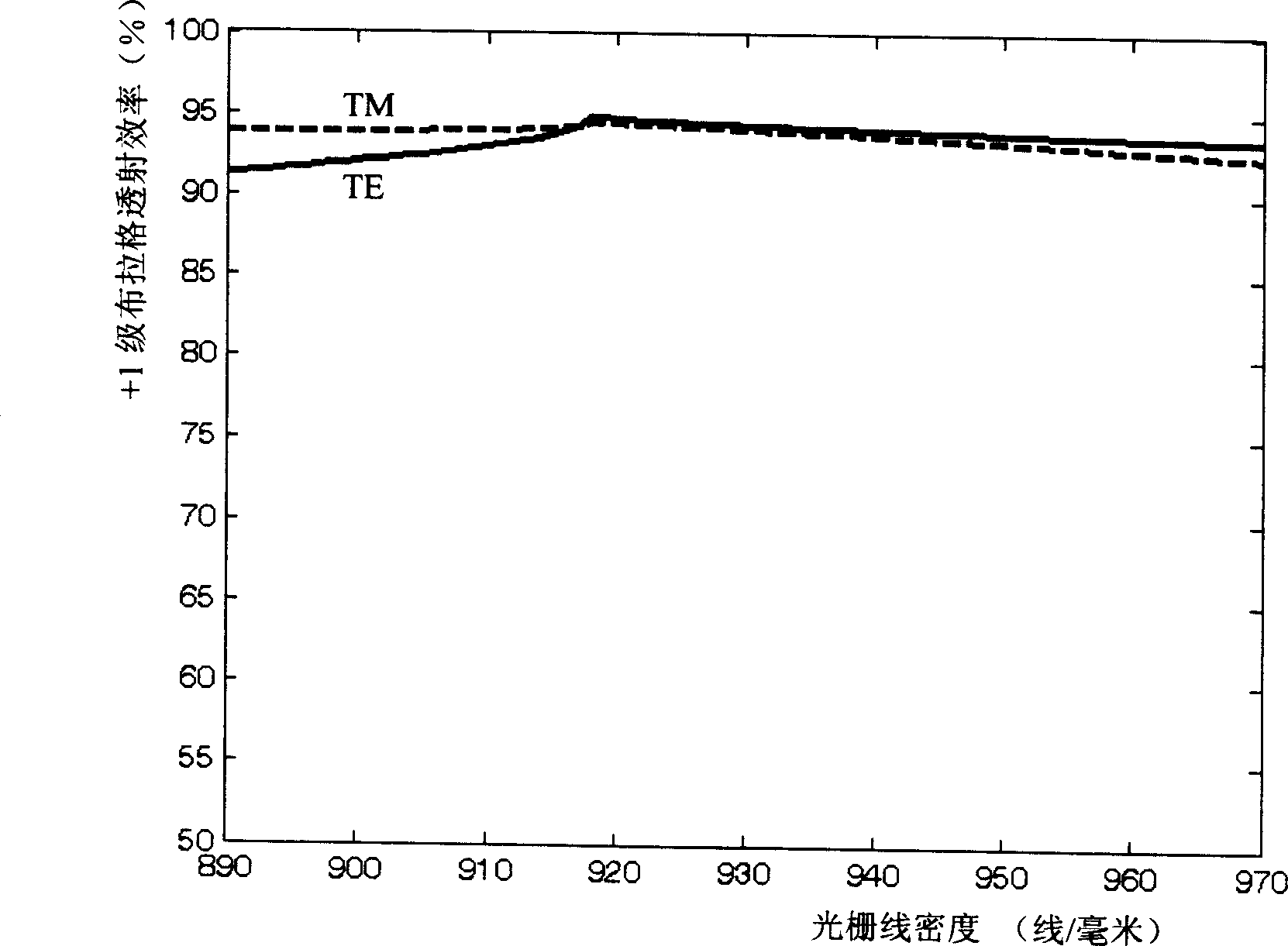
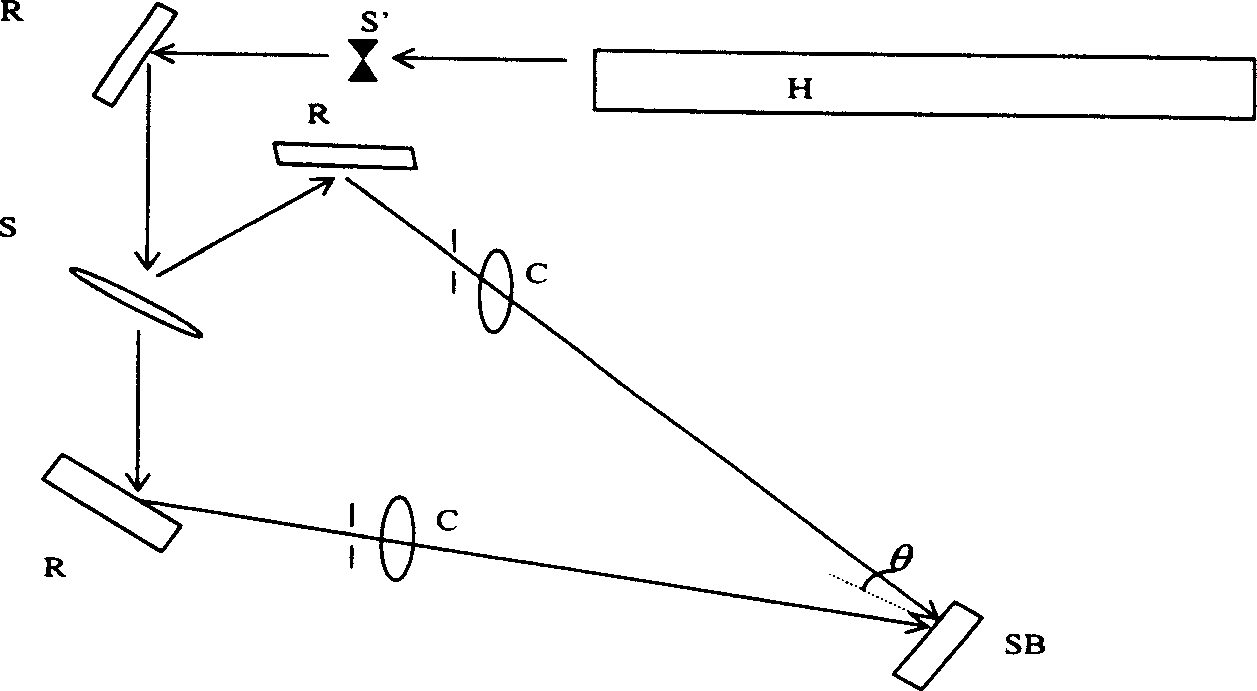
High-diffraction efficiency quartz transmission grating of 1053 nano wavelength
A quartz transmission grating of high efficiency of diffraction is a high density, rectangle quartz grating of deep eclipse. The grating's wave-length is 1053 nms and its line density 895-965 lines / millimeter. Its depth is 1.9-2.1 micron while the ratio of the filled space to empty space by the grating is 1 / 2.The invention can make the efficiency of +1 class Prague transmission and diffraction in the partial vibration directions of TE and TM be more than 85% at the same time. It realizes free choice of partial vibration model. Especially when the grating density is 920 lines / millimeter and the grating depth is 2 micron, each efficiency of TE and TM partial vibration is more than 94%. The invention is produced by the micro-electronics light chiseling craft and the deep eclipse craft. Its cost is low and it is suitable for volume-produce.
Owner:SHANGHAI INST OF OPTICS & FINE MECHANICS CHINESE ACAD OF SCI
Loading method and device of field programmable gate array
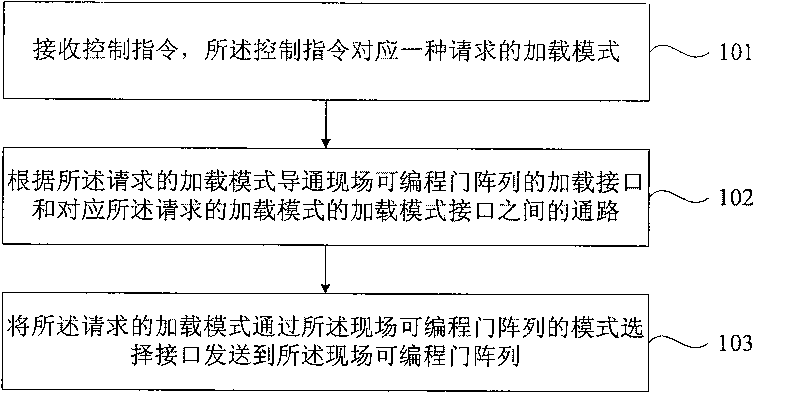
InactiveCN101763278AAchieve freedom of choiceUnable to solve the problemProgram loading/initiatingField-programmable gate arrayMode selection
The embodiment of the invention provides a loading method and a device of a field programmable gate array. The loading method comprises the following steps: receiving a control command, wherein the control command is corresponding to a requested loading mode; communicating a passage between a loading interface of the field programmable gate array and a loading mode interface of the corresponding requested loading mode; and transmitting the requested loading mode to the field programmable gate array through a mode selection interface of the field programmable gate array so as to be convenient for the field programmable gate array to load according to the loading mode. Through the method and the device, a corresponding loading mode can be selected according to the type of a loading mode and a loading mode interface of a main control device so as to freely select a loading or debugging mode to of a FPGA device and solve the problems of no matching and inconvenient matching of a development project caused by consistent loading mode of the FPGA device.
Owner:HUAWEI TECH CO LTD
Third-party billing method and system
InactiveCN101150416AFlexible BillingAchieve freedom of choiceMetering/charging/biilling arrangementsData switching by path configurationThird partyRelevant information
This invention discloses a charge method for a third party including: a paying party sets its own limit and ciphered code and shares related information to a service using party, when which accesses service, it inputs a special access symbol of charge of the third party, ID of the paying party or ciphered code, a service applied server judges if the attribute of the user is that it is charged by a third party, if so, the applied server system collects charge information of the third party and transfers the information to the charge system by a charge access protocol, which pre-deducts and deducts the fee from the account of a third party. This invention discloses another charge method and a system for third parties.
Owner:HUAWEI TECH CO LTD
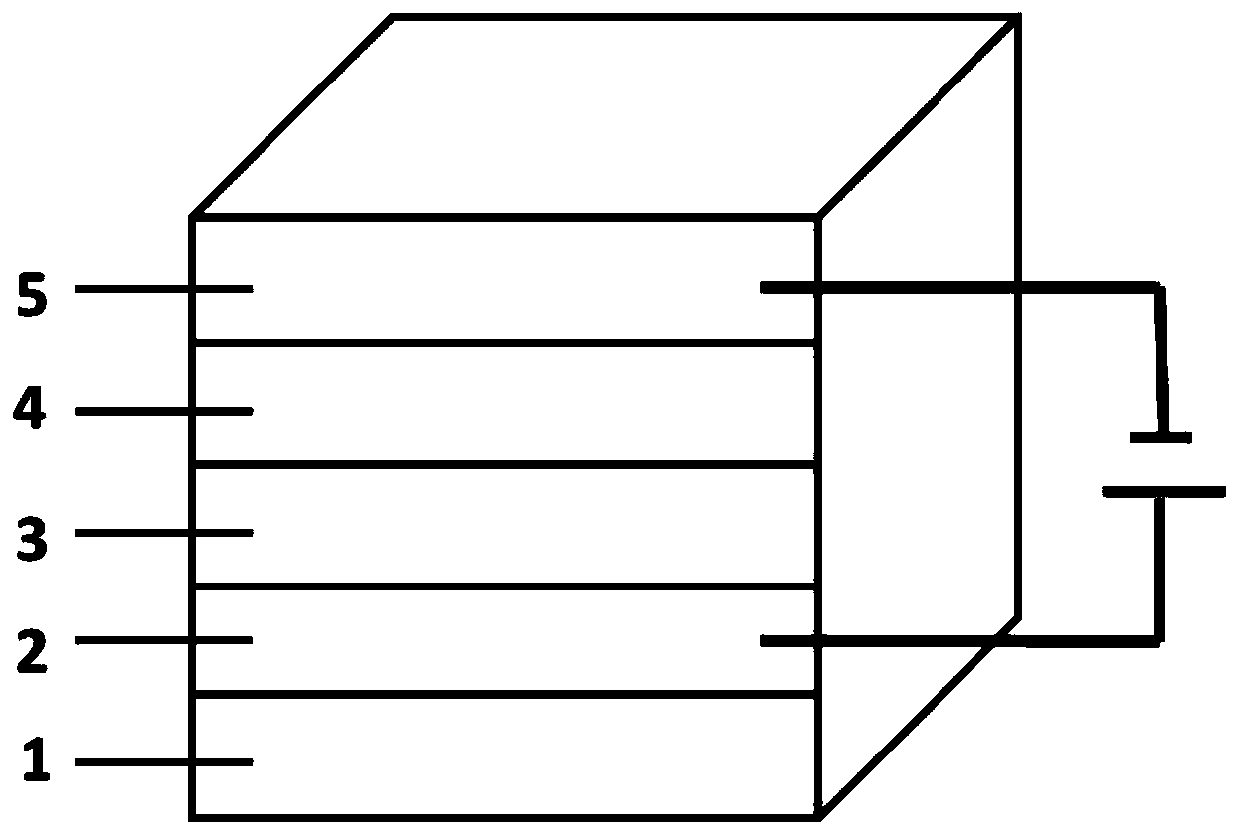
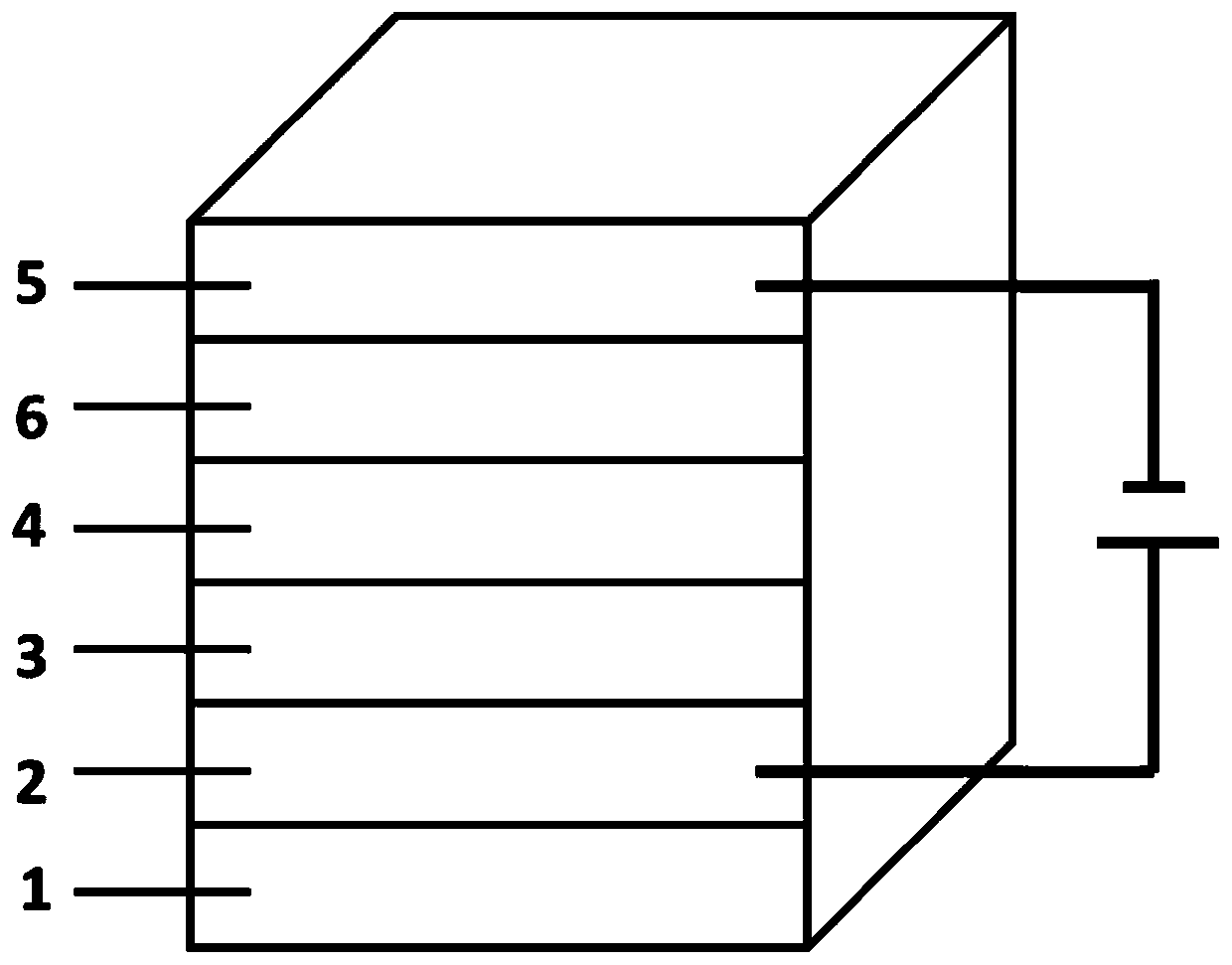
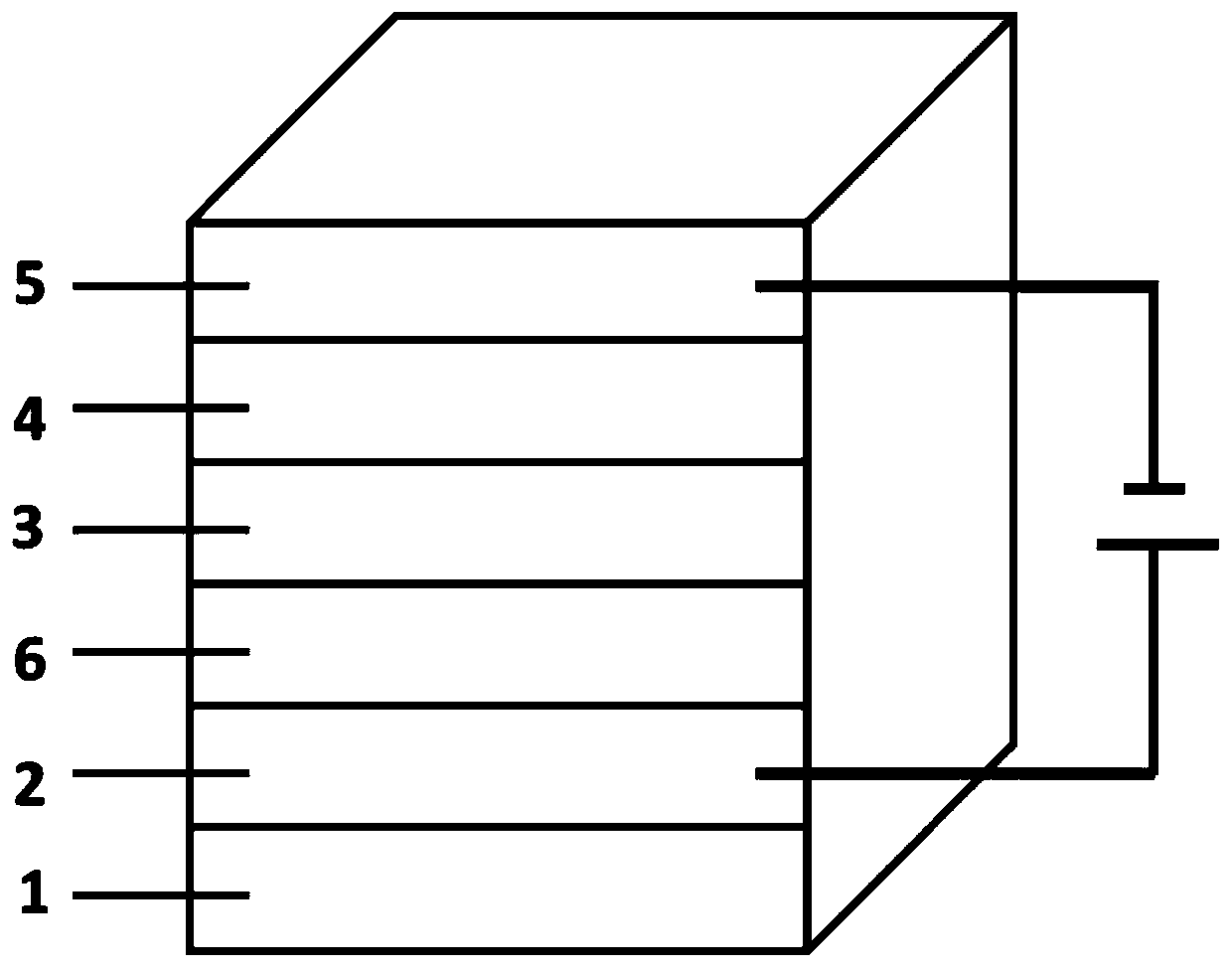
Novel self-filtering narrow spectral response organic light detector
ActiveCN110534650AImprove controllabilityAchieve freedom of choiceSolid-state devicesSemiconductor/solid-state device manufacturingSpectral responseLight filter
The invention relates to a novel self-filtering narrow spectral response organic light detector. The organic light detector sequentially comprises a substrate, a positive electrode, a P-type layer, anN-type layer and a negative electrode, the P-type layer can be further divided into a single-layer P-type layer structure and a multi-layer P-type layer structure, and in the single-layer P-type layer structure, the band gap of the P-type layer material is wider than that of the N-type layer material; in the multi-layer P-type layer structure, the band gap of at least one P-type layer material inthe P-type layer materials which are not in direct contact with the N-type layer is wider than that of the N-type layer material and / or the P-type layer material which is in direct contact with the N-type layer, and a buffer layer can be added between the positive electrode and the P-type layer and / or between the N-type layer and the negative electrode. According to the invention, a novel devicestructure and a simple preparation method are adopted, and the free selection of a detection spectrum waveband and the free adjustment of the half-peak width of a detection spectrum are achieved without a band-pass optical filter.
Owner:GUANGZHOU GUANGDA INNOVATION TECH CO LTD
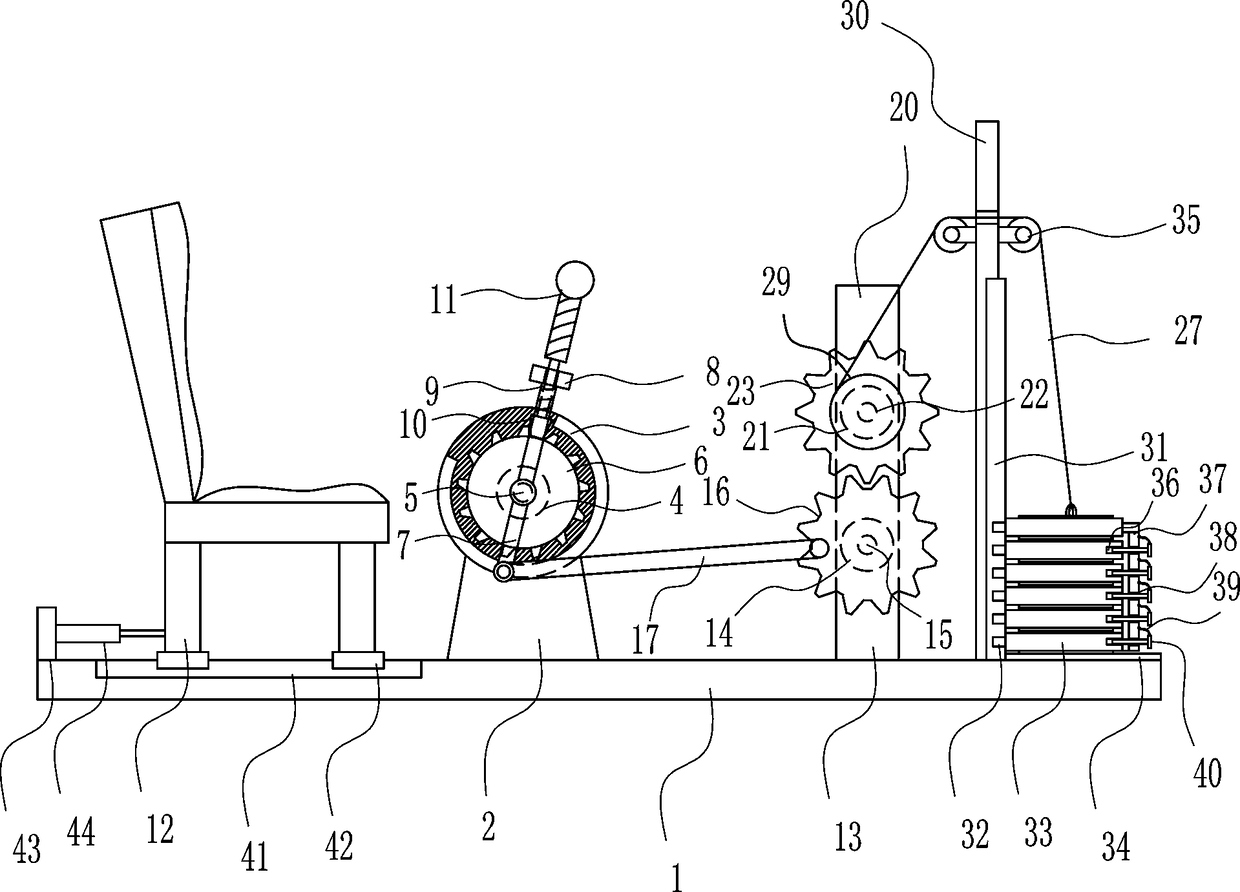
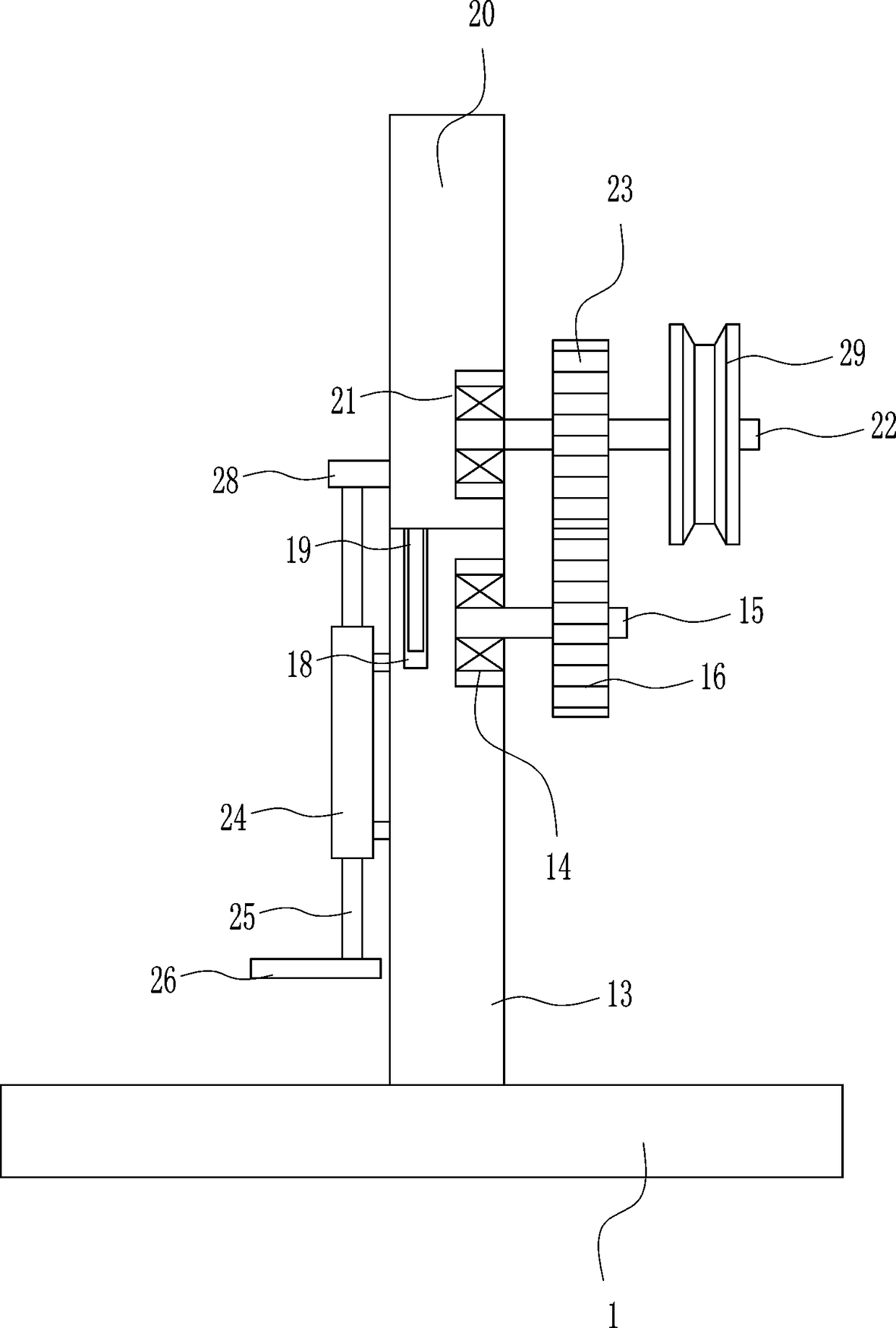
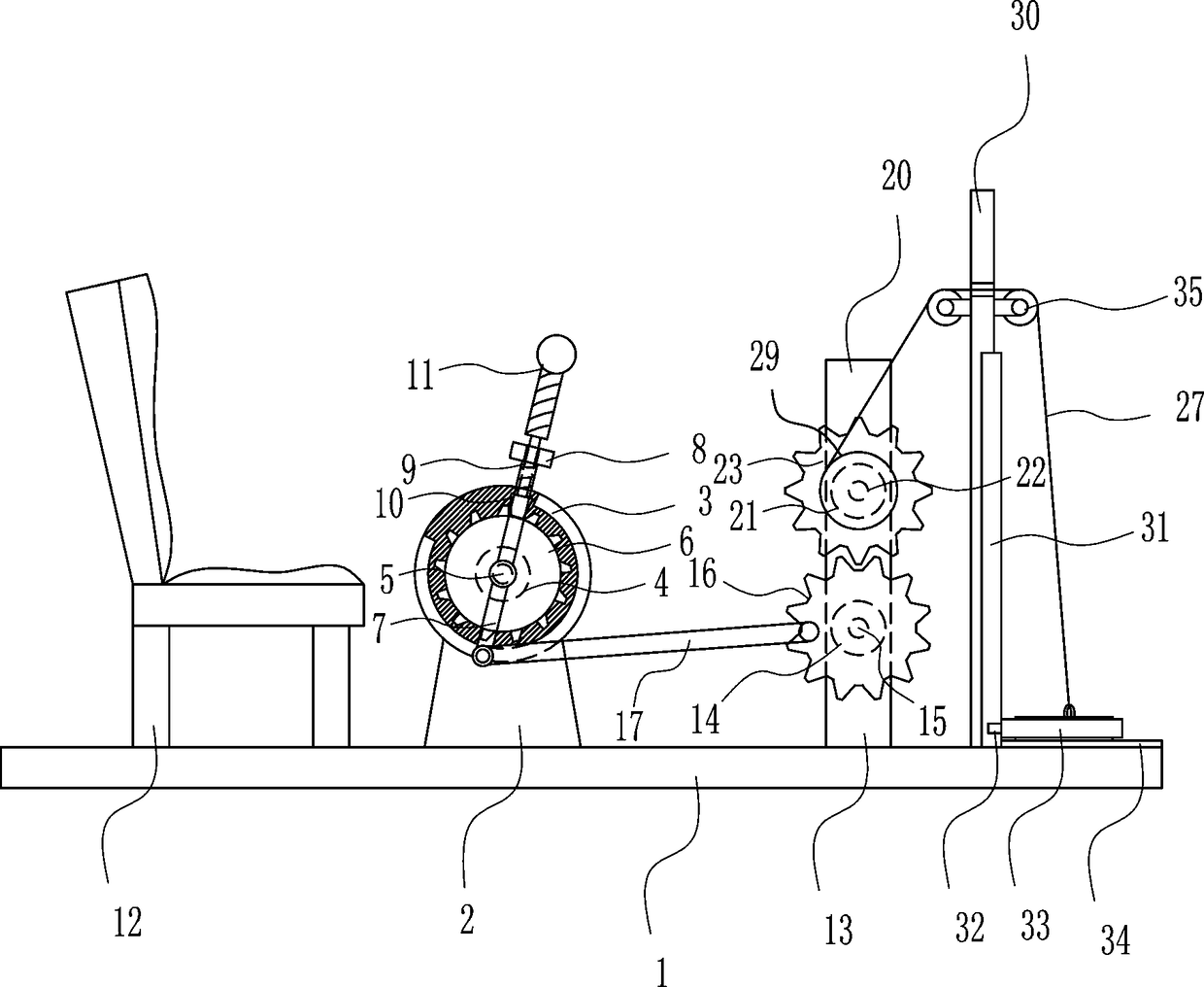
Orthopedic patient recovery exercise device
InactiveCN108187312AAchieve freedom of choicePromote recoveryMuscle exercising devicesEngineeringPhysical exercise
The invention relates to an exercise device, in particular to an orthopedic patient recovery exercise device, and aims to provide an orthopedic patient recovery exercise device which can be used indoors and is convenient to operate and good in exercise effect. The orthopedic patient recovery exercise device comprises a bottom plate, a mounting seat and the like. A seat is arranged on the left sideof the top of the bottom plate, the mounting seat is arranged in the middle of the top of the bottom plate, a fixed basin is arranged on the mounting seat, a first bearing block is arranged on the fixed basin, a first rotating shaft is arranged on the first bearing block, and a ratchet wheel is arranged on the first rotating shaft. A patient drives a weight block to rise by pulling a handle and needs large effort for pulling the handle as the weight block has a certain weight, exercise effects are achieved, and the device is convenient to operate and applicable to postoperative patients unsuitable for going out.
Owner:张冬冬 +1
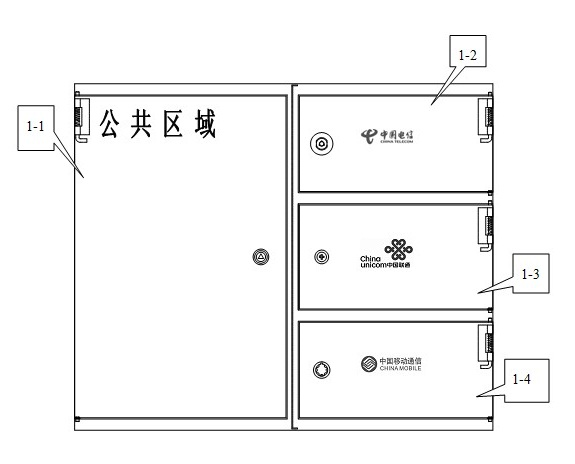
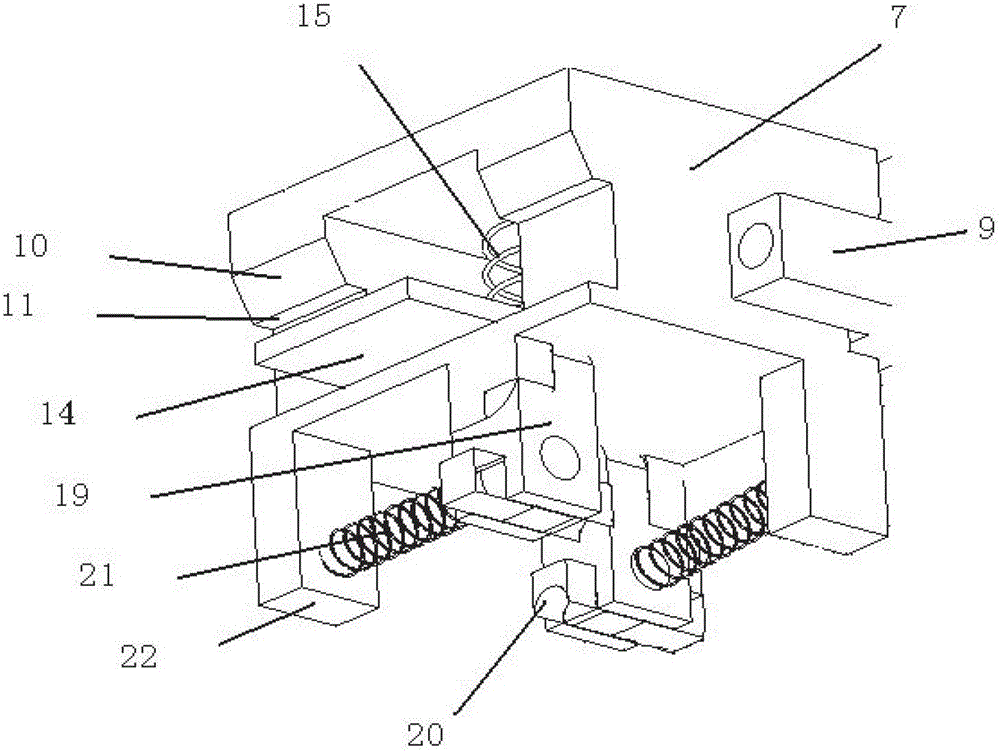
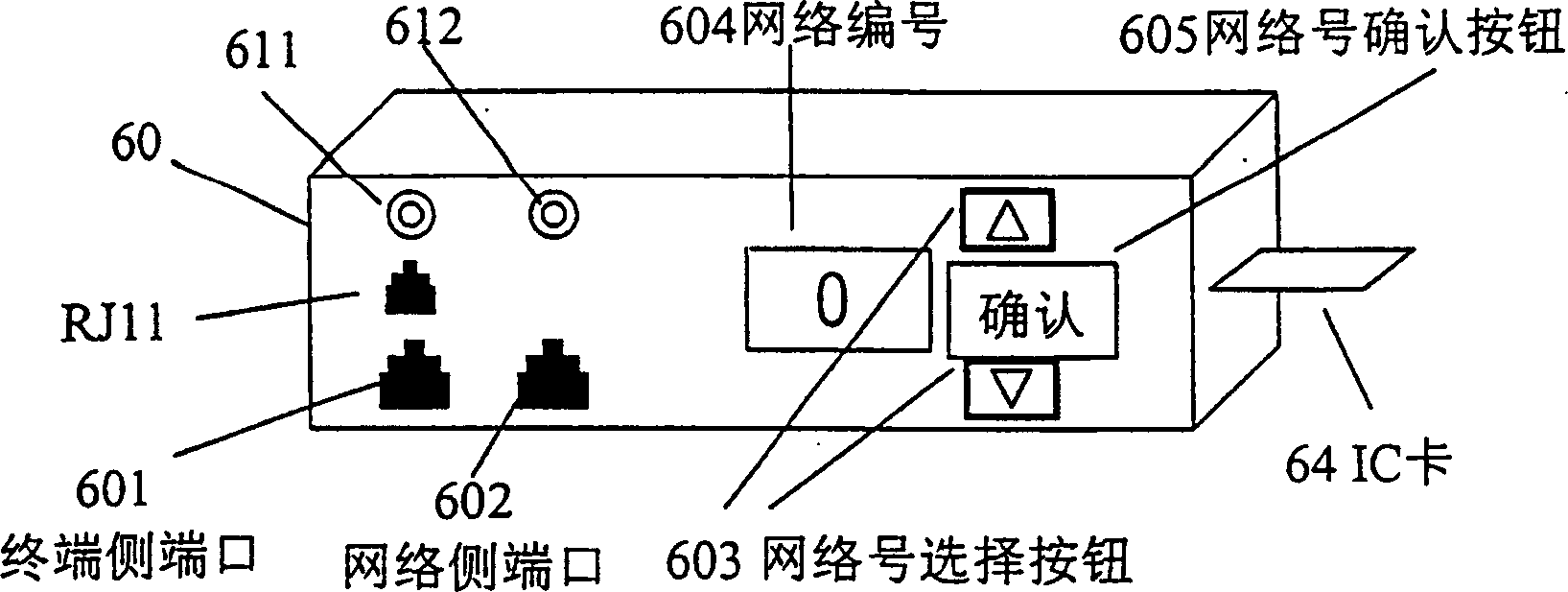
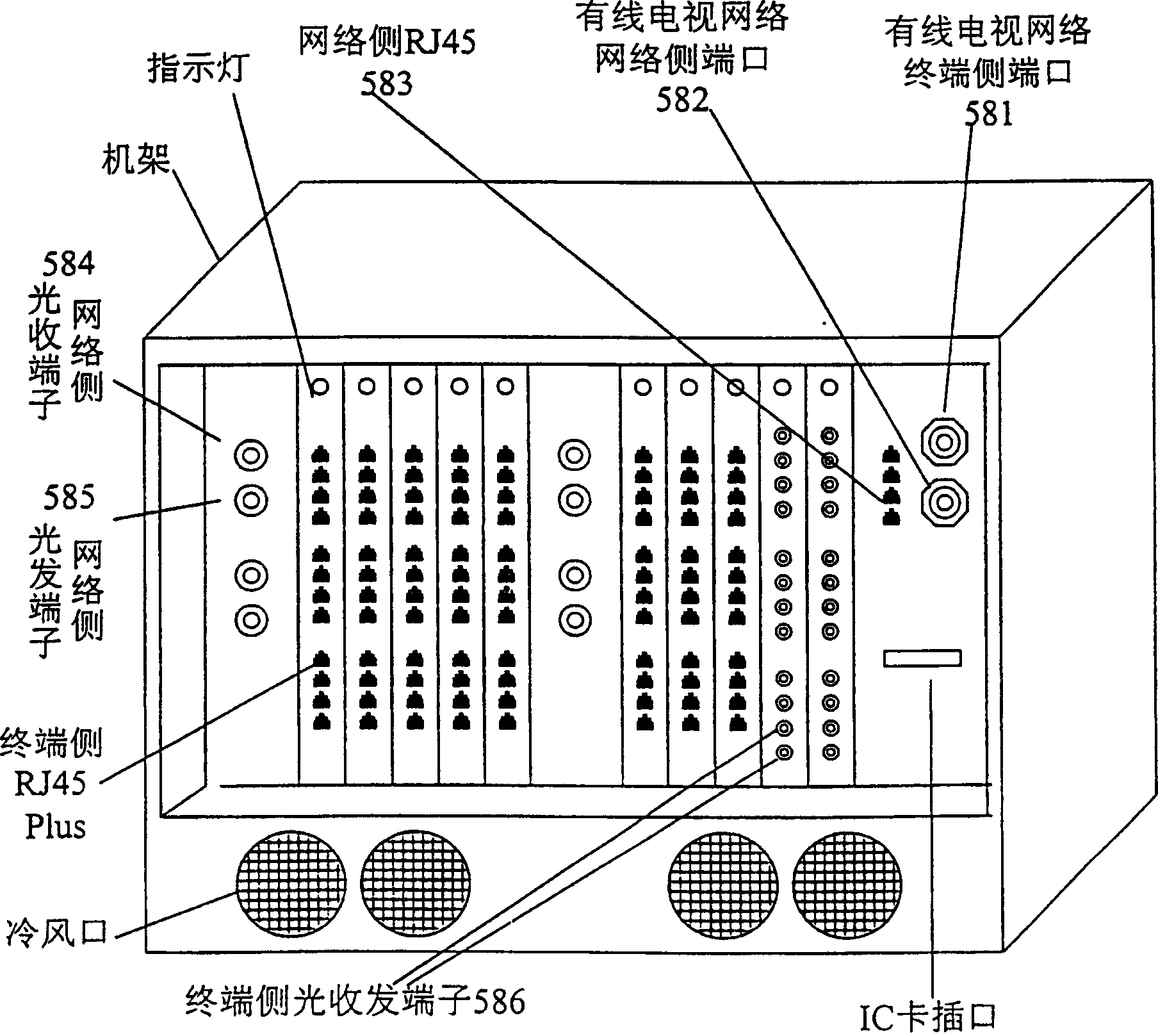
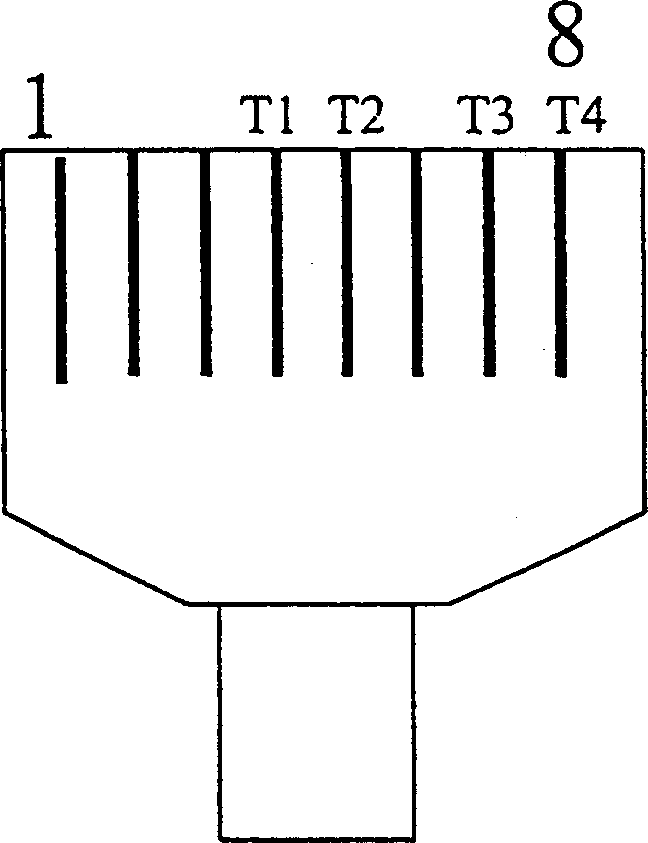
Multifunctional three-in-one network floor terminal box for FTTx (Fiber To The x)
ActiveCN102681121ASave resourcesGood for management and beautificationFibre mechanical structuresMultiple functionElectrical and Electronics engineering
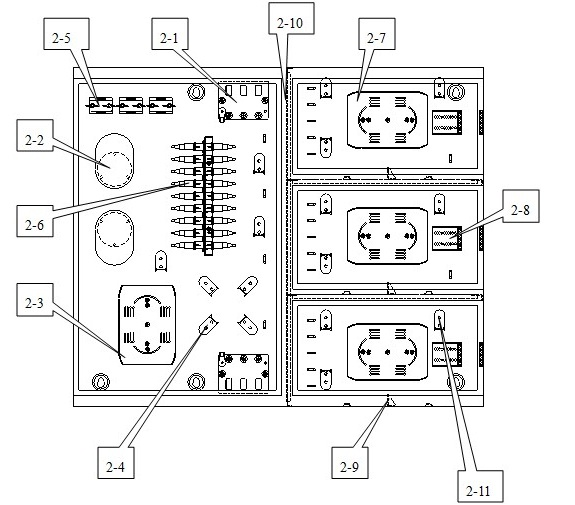
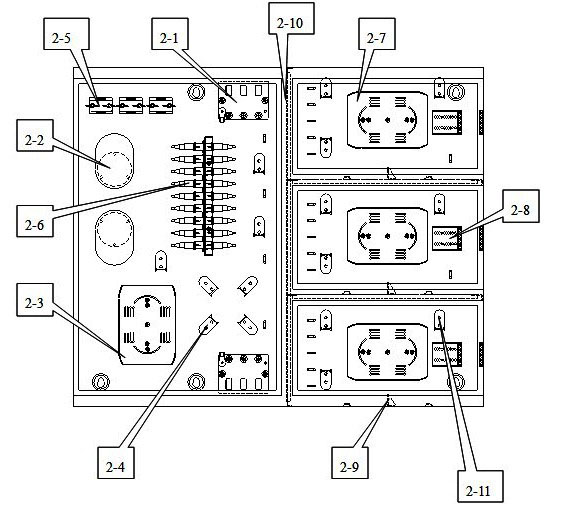
The invention discloses a multifunctional three-in-one network floor terminal box for FTTx (Fiber To The x). The floor terminal box comprises a public junction bin and special bins and is characterized in that the special bins comprise a first special bin, a second special bin and a third special bin, wherein the first special bin, the second special bin and the third special bin are same in internal structure; and each of the first special bin, the second special bin and the third special bin can be mutually connected and communicated with the public junction bin. According to the multifunctional three-in-one network floor accessing scheme for the construction of FTTx, all signals provided by three major operators are terminated to one shared three-in-one network floor terminal box which is internally provided with independent equipment bins for the three major operators, a shared butterfly-shaped optical cable is laid in a section to a user in the floor, and therefore an operator service can be replaced for the user just by operation in the floor terminal box.
Owner:NANJING HUAMAI TECH
Electronic interaction window display system
InactiveCN103164021ASimple structureNovel and reasonable designInput/output for user-computer interactionAdvertisingComputer graphics (images)Analog signal
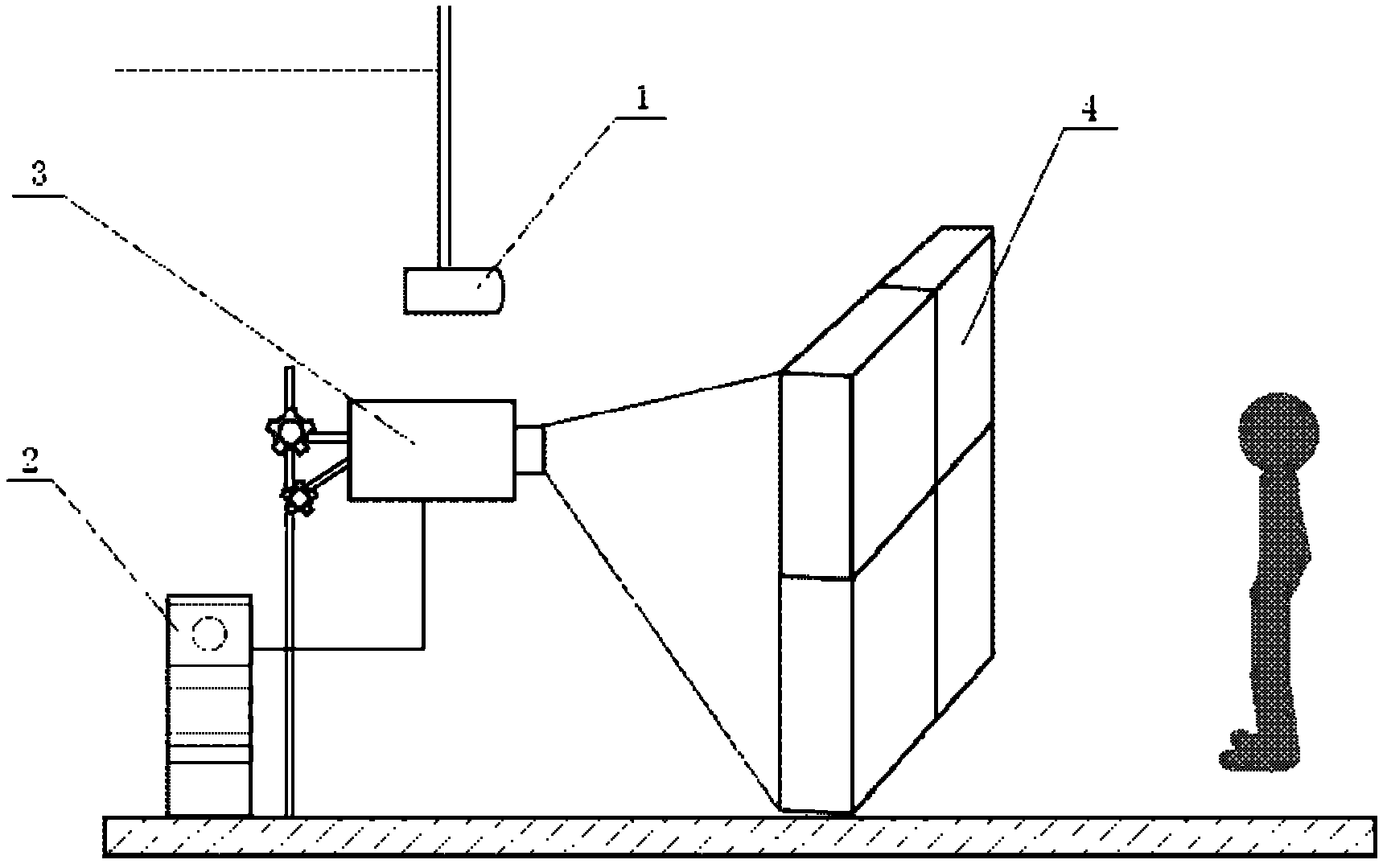
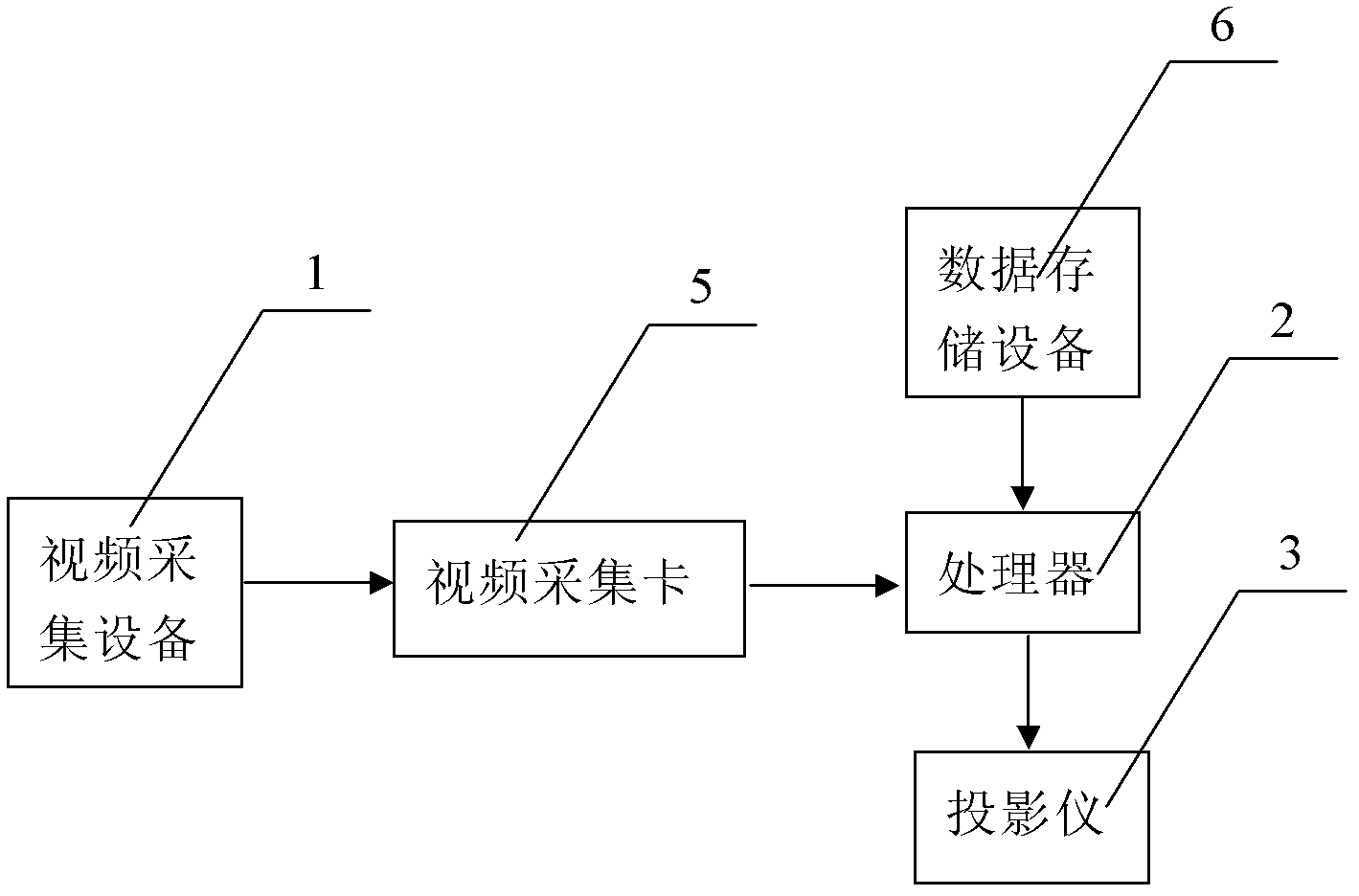
The invention discloses an electronic interaction window display system. The electronic interaction window display system comprises a video capture device, a video capture card and a processor, wherein the video capture device is used for capturing user motion change images, the video capture card is connected with the video capture device and used for converting user motion change image analog signals shot by the video capture device into digital signals, and the processor is connected with the video capture card and used for operating the digital signals output by the video capture card to obtain coordinate position parameters on a screen corresponding to change areas. The processor is connected with data storage equipment and retrieves video information stored in the data storage equipment in advance according to the coordinate position parameters calculated by the processor, the processor is connected with a projector used for projecting the video information retrieved from the data storage equipment, and a glass window model used by being matched with the projector and provided with a projection film is arranged in front of the projector. The electronic interaction window display system shocks the visual sensory organs of an audience, can show advertising publicity as a window, and is strong in interestingness and good in publicity effect.
Owner:XIAN TEKTONG DIGITAL TECH
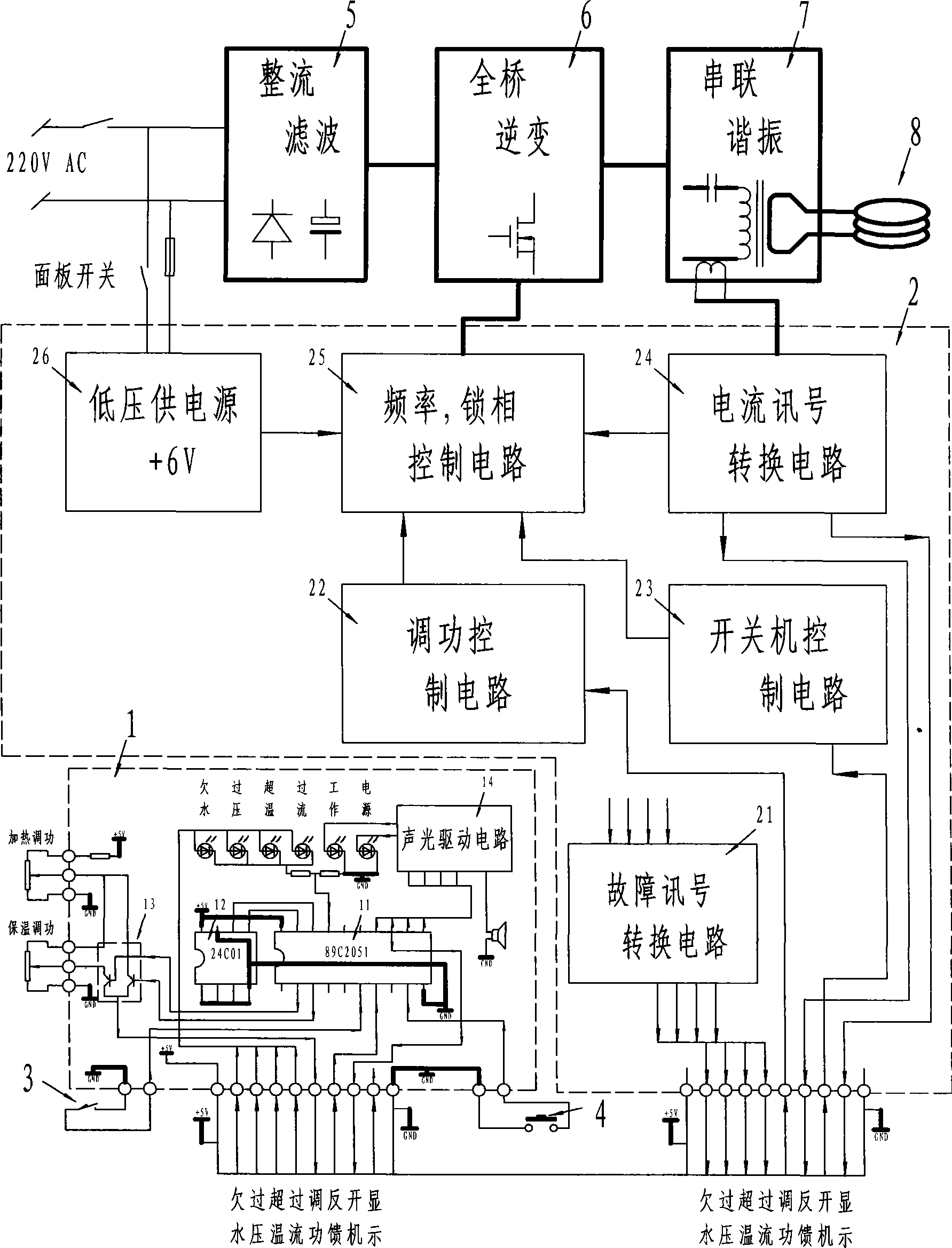
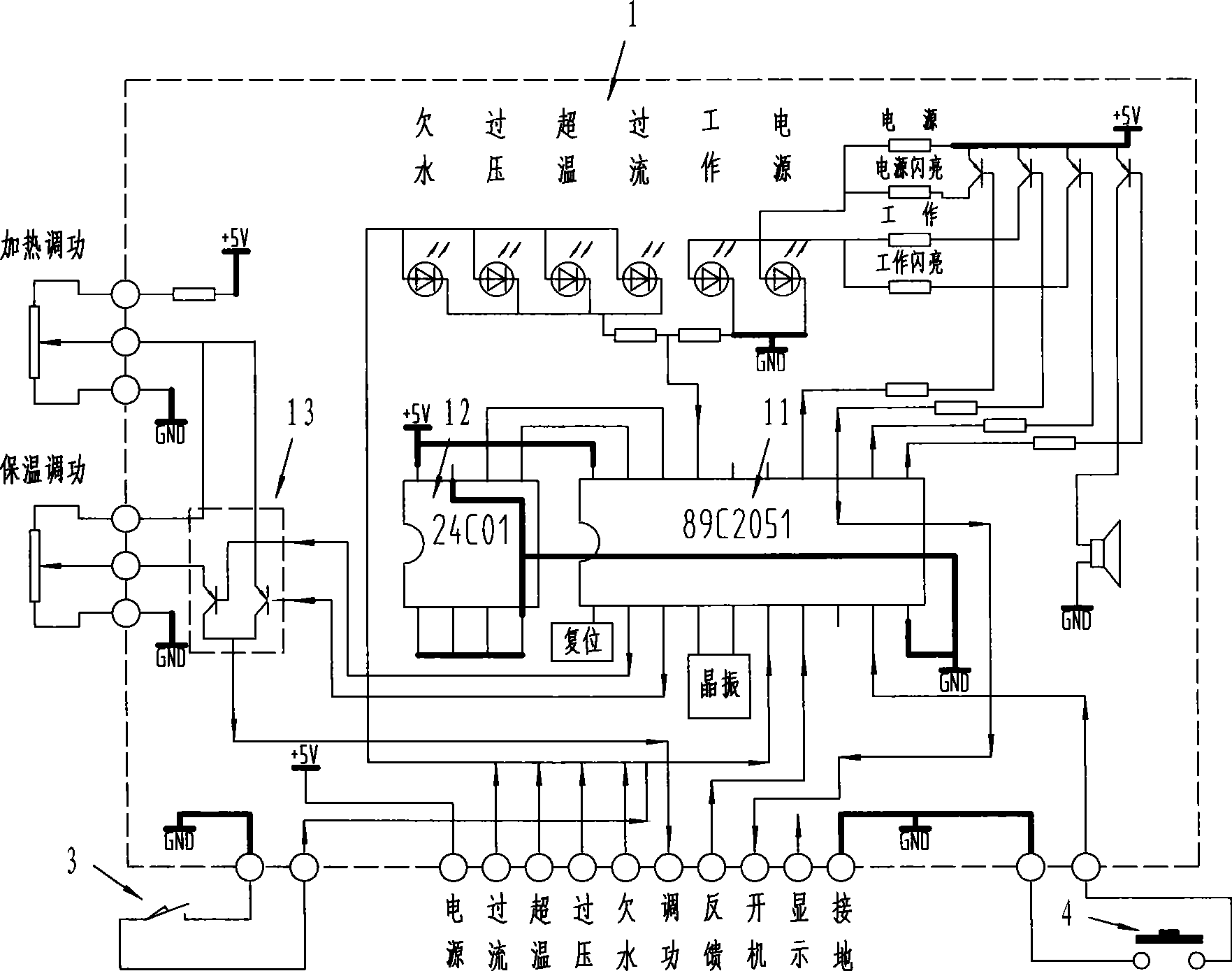
Intelligent high-frequency induction heating equipment
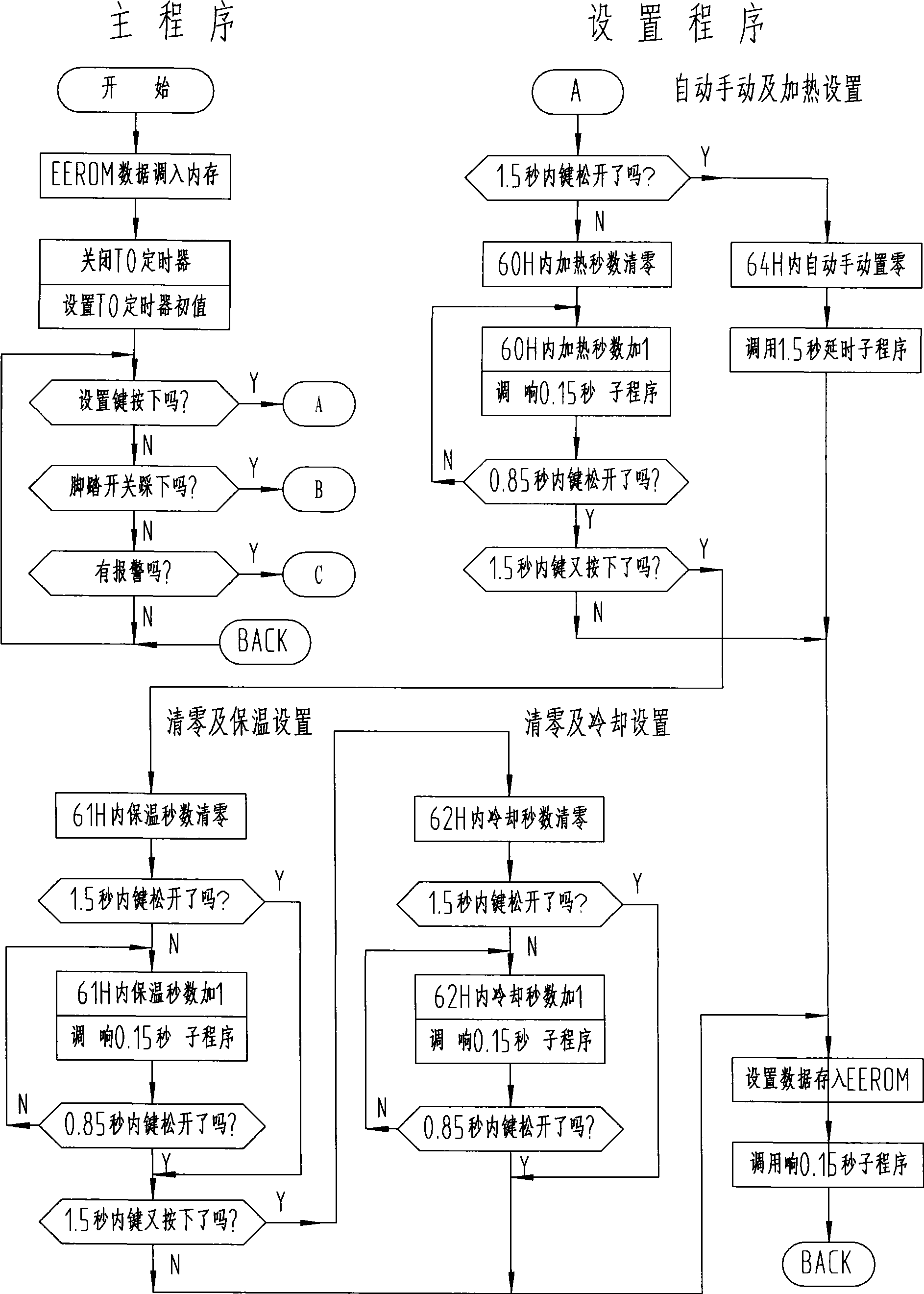
InactiveCN101370329AConstant volumeRealize fully automatic controlFluid heatersInduction heating controlMicrocontrollerAutomatic control
The invention relates to a intelligent type HF induction heating apparatus comprising set key, foot pedal switch, master control board. The master control board comprises frequency, locking phase control circuit, switching control circuit, power adjusting control circuit, and fault signal change-over circuit; the master control board is controlled by single chip machine which is provided with EEROM memorizer and heat insulation change-over circuit. Parameter is set and function is converted for the single chip machine by setting time length of pressing key and releasing key. The invention accomplishes full-automatic control, free choice for heat insulation and cooling during whole automatic control process, instant stop function and manual conversion.
Owner:钱小林
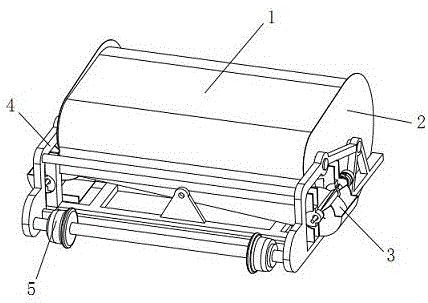
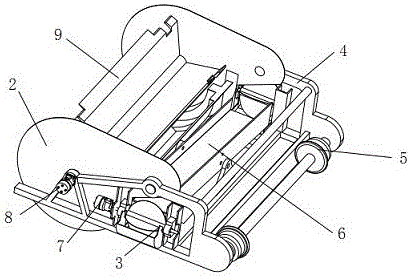
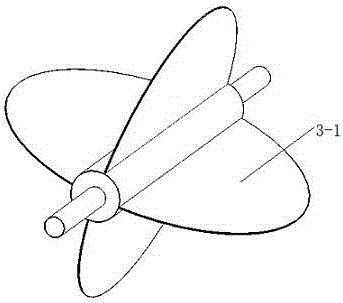
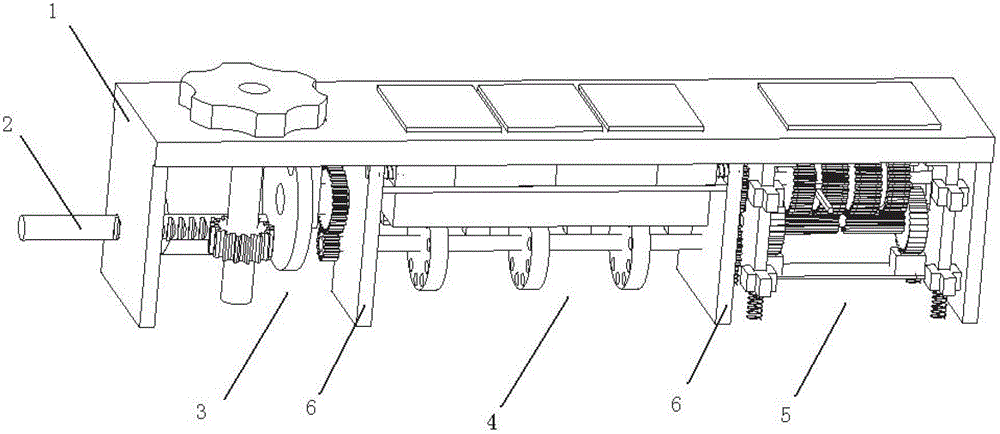
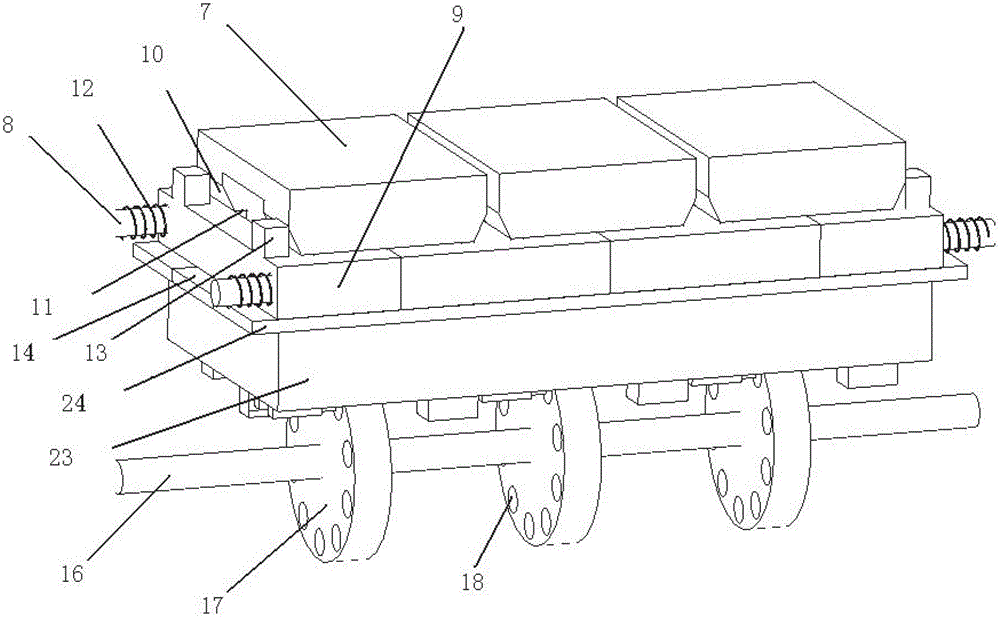
Turning plate type bidirectional sand discharging and removing device of track sand removing vehicle for both roads and railways
The invention discloses a turning plate type bidirectional sand discharging and removing device of a track sand removing vehicle for both roads and railways. The turning plate type bidirectional sand discharging and removing device of the track sand removing vehicle for both roads and railways comprises a machine frame, a sand collecting portion, a turning plate type sand discharging portion and a guide portion, wherein the machine frame is used for being connected with a driving device; the sand collecting portion is located at the front end of the machine frame; the turning plate type sand discharging portion is located at the back of the sand collecting portion and used for receiving accumulated sand conveyed by the sand collecting portion; the guide portion is located at the rear end of the machine frame; the turning plate type sand discharging portion comprises a sand guide turning plate and sand lifting portions arranged on two sides of the machine frame; the middle of the sand guide turning plate is installed with the assistance of a support installed in the middle of the machine frame, the two sides of the sand guide turning plate are installed on the inner side of the support with the assistance of a rotary shaft, and outlets in two ends of the sand guide turning plate are located over the sand lifting portions on two sides of the machine frame; the sand collecting portion conveys the accumulated sand in an area to be cleaned into the sand discharging portion; the sand discharging portion is of a turning plate type structure, and according to the turning plate type structure, the output sides can be adjusted as needed; and the different output sides correspond to the sand lifting portions on different sides, free selection of the sand discharging direction can be realized, and adaptability is higher.
Owner:SHIJIAZHUANG TIEDAO UNIV
Key type mechanical password lock and password setting method
InactiveCN102943591ABroaden your optionsSmall starting torquePuzzle locksPermutation locksTransmission systemControl system
The invention discloses a key type mechanical password lock and a password setting method. The password lock comprises a lock shell as well as a key transmission system, a resetting system and a tongue control system, wherein the key transmission system, the resetting system and the tongue control system are arranged in the lock shell; the key transmission system comprises a keyboard component and a password carrying wheel plate component; each key body of the keyboard component is corresponding to each password carrying wheel plate of the password carrying wheel plate component; each key body is divided into two key branches extending to two plate surfaces of the corresponding password carrying wheel plate, so that the two key branches are respectively matched with the two plate surfaces; a plurality of wheel plate holes are formed in each password carrying wheel plate; pin shafts are inserted in the wheel plate holes; the extending directions and the combinations of the pin shafts on the two plate surfaces bear password information; when pushing wheel blocks on the key branches can apply forces to the corresponding pin shafts during pushing the keys, the password carrying wheel plate component is pushed to rotate in a positive or negative direction; and the resetting system is driven by a linking mechanism to store energy and the tongue control system is driven by the linking mechanism to act; and the key type mechanical password lock disclosed by the invention has the advantages of modularized structure, standardized part, small pressing force, safety, reliability, low production cost and wide application range.
Owner:ANHUI UNIVERSITY OF TECHNOLOGY AND SCIENCE
System enabling user to select information networks and method thereof
InactiveCN1174579CAchieve freedom of choiceEnsure safetyData switching networksTime-division multiplexing selectionInformation networksDistributed computing
The present invention relates to a system and a method for selecting information networks by users; the system includes at least two information networks and user terminals, and also includes: a network selector connects to the user terminal to receive and transmit user parameters and requests for accessing a information network and control the user terminal; a safe exchange connectting between each network selector and information network judging if the user parameters and requests are legal, and permit accessing the required information network for the user according to the judgement results. The present invention can not only realize the selection of information network for users, but also can assure the safety of information networks.
Owner:余鲲
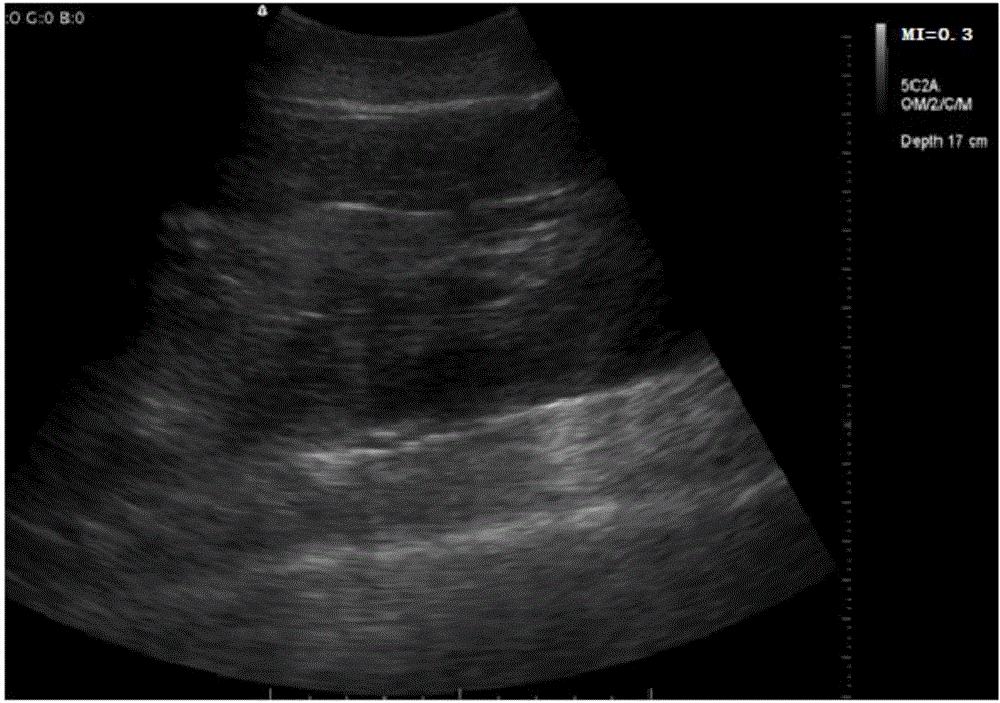
Method for achieving natural mating and spawning of acipenser dabryanus through artificial control
ActiveCN106069899AAchieve spontaneous matingProtection breachClimate change adaptationPisciculture and aquariaMatingSemen
The invention discloses a method for achieving natural mating and spawning of acipenser dabryanus through artificial control. Natural spawning, natural spermiation and male and female synchronous development and maturation are achieved through artificial control technical measures of B ultrasonic parent nondestructive identification, selection of sexually mature fishes, bed material control, water flow control and the like for the first time. The situations that in a traditional method, synthetic auxins are adopted for inducing, egg collecting and semen collecting are conducted by artificially compressing the abdomen, artificial fertilization hurts fish bodies, and artificial selection is conducted are avoided, and a great breakthrough is achieved for acipenser dabryanus protection. By means of the method for achieving natural mating and spawning of the acipenser dabryanus through artificial control, the fertility rate (60%-80%) of the acipenser dabryanus is equivalent to that of artificial propagation, due to the clustering and adhesion action of fertilized eggs, the seedling hatched rate (20%-50%) is lower than that of artificial propagation, but the survival rate (80%-90%) of later stage culture of hatched fry is higher than that of artificially propagated fry.
Owner:YANGTZE RIVER FISHERIES RES INST CHINESE ACAD OF FISHERY SCI
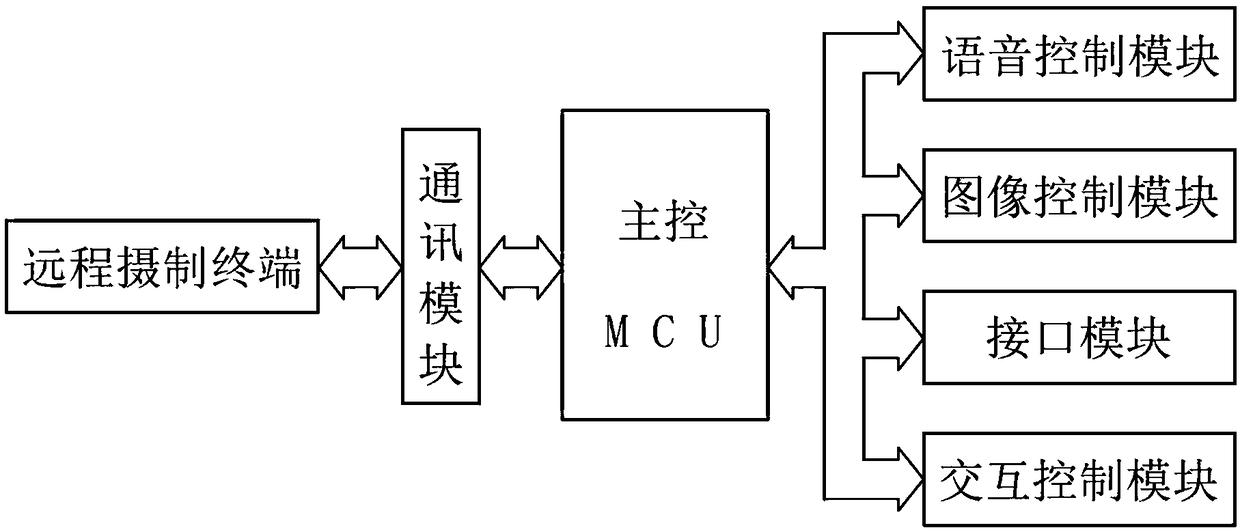
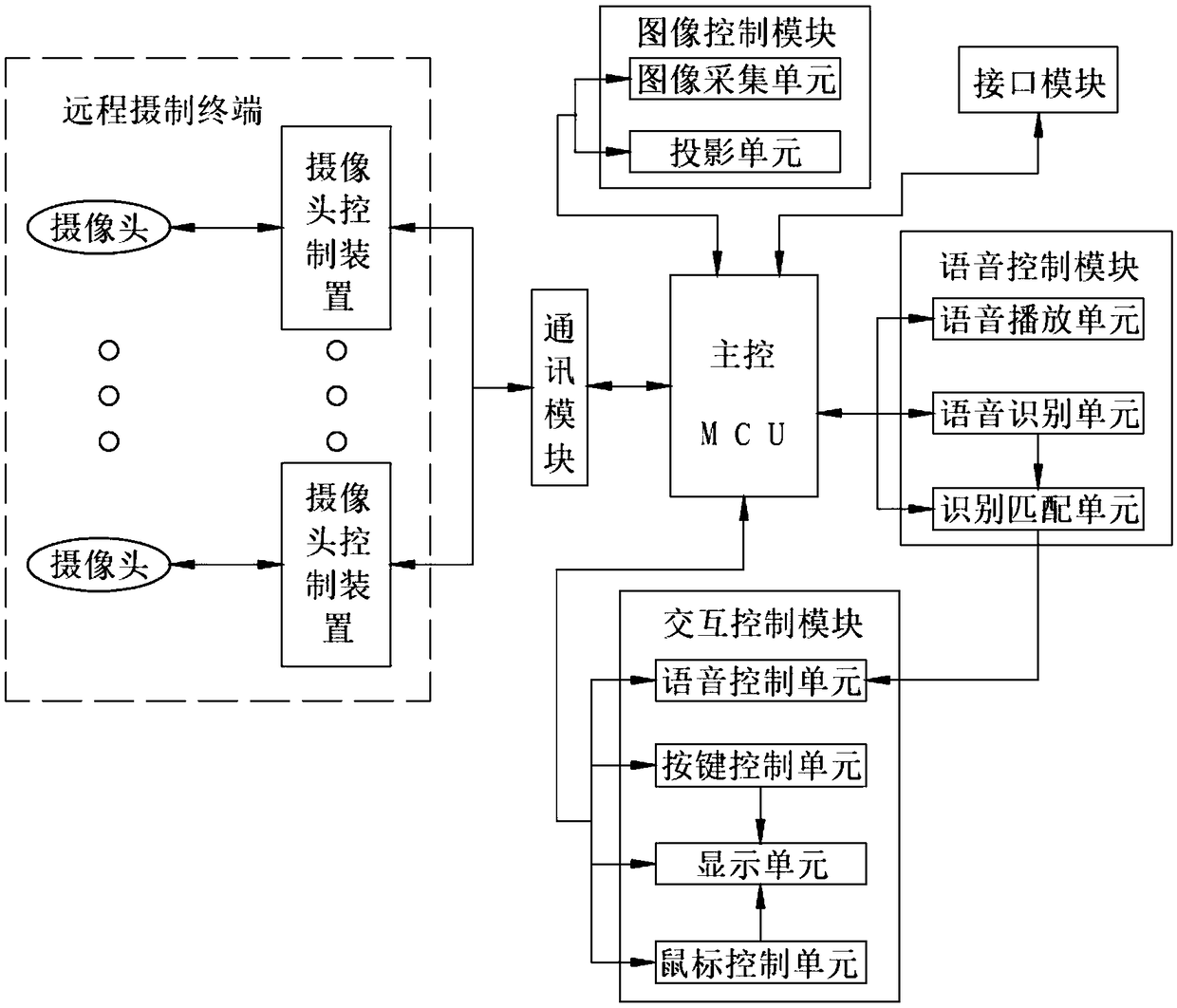
Classroom live-action reproduction interactive teaching system based on a robot projection function
InactiveCN108877347AImprove user experienceRealize intelligent interactionElectrical appliancesInteraction controlLive action
The invention discloses a classroom live-action reproduction interactive teaching system based on a robot projection function. The classroom live-action reproduction interactive teaching system is composed of a voice control module, an image control module, a remote shooting terminal, a communication module, an interface module, an interaction control module and a main control MCU. The voice control module, image control module, the communication module, the interface module and the interaction control module are connected with the main control MCU respectively and electrically; and the remoteshooting terminal and the main control MCU are in communication connection by the communication module. According to the system disclosed by the invention, the system not only serves as an image acquisition device but also serves as a remote projection device to carry out classroom live-action reproduction device; and the system also has the good interaction performance.
Owner:安徽硕威智能科技股份有限公司
Preparation method of garnet crystal film based on He ion irradiation
InactiveCN109468683AQuality improvementAchieve freedom of choicePolycrystalline material growthSingle crystal growth detailsSurface layerFilm base
The invention discloses a preparation method of a garnet crystal film based on He ion irradiation. The preparation method comprises the following steps: a) computing to obtain energy and dosage of Heion; b) performing irradiation on the garnet crystal according to the computed energy and dosage of the He ion, wherein a damaged layer divides the garnet crystal into a surface layer of an upper endand a substrate layer of a lower end; c) cleanly washing the irradiated garnet crystal and substrate material; d) fixing the substrate layer and substrate material of the garnet crystal, and mutuallyextruding together; and e) performing annealing treatment on the fixed garnet crystal and substrate material to obtain the garnet crystal film. The garnet crystal film manufactured by combining the Heion irradiation with the bonding technology is high in quality, and the free selection of the substrate material is realized.
Owner:SHANDONG JIANZHU UNIV
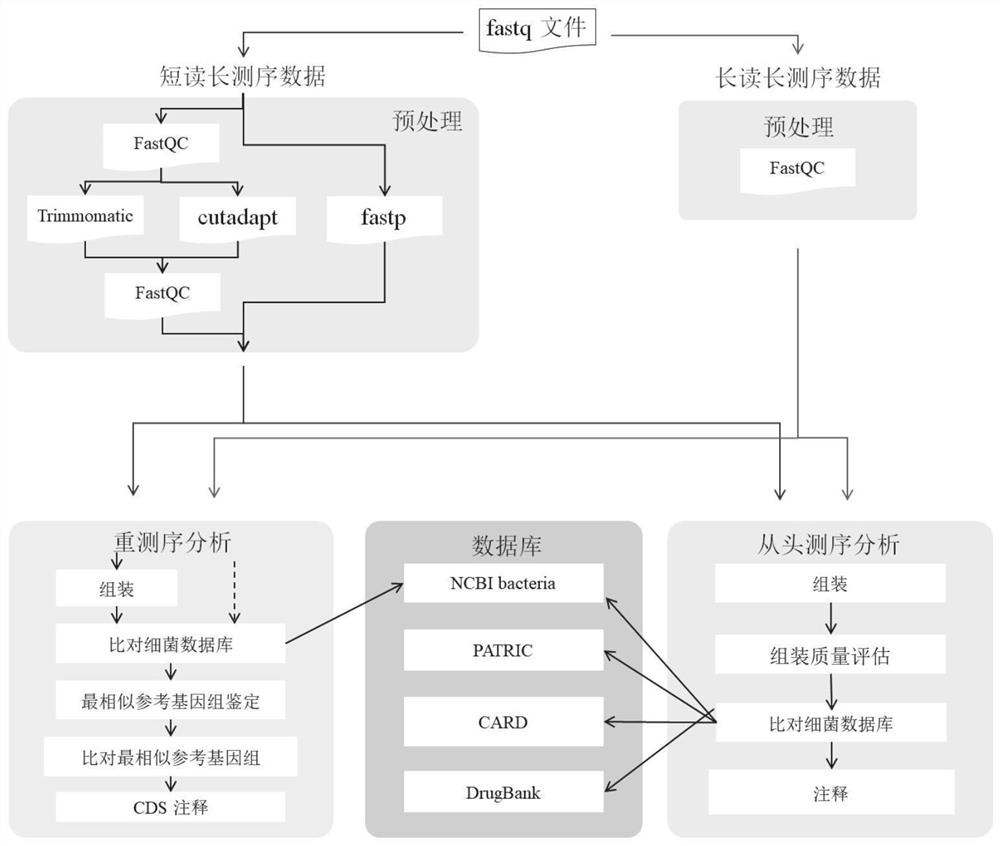
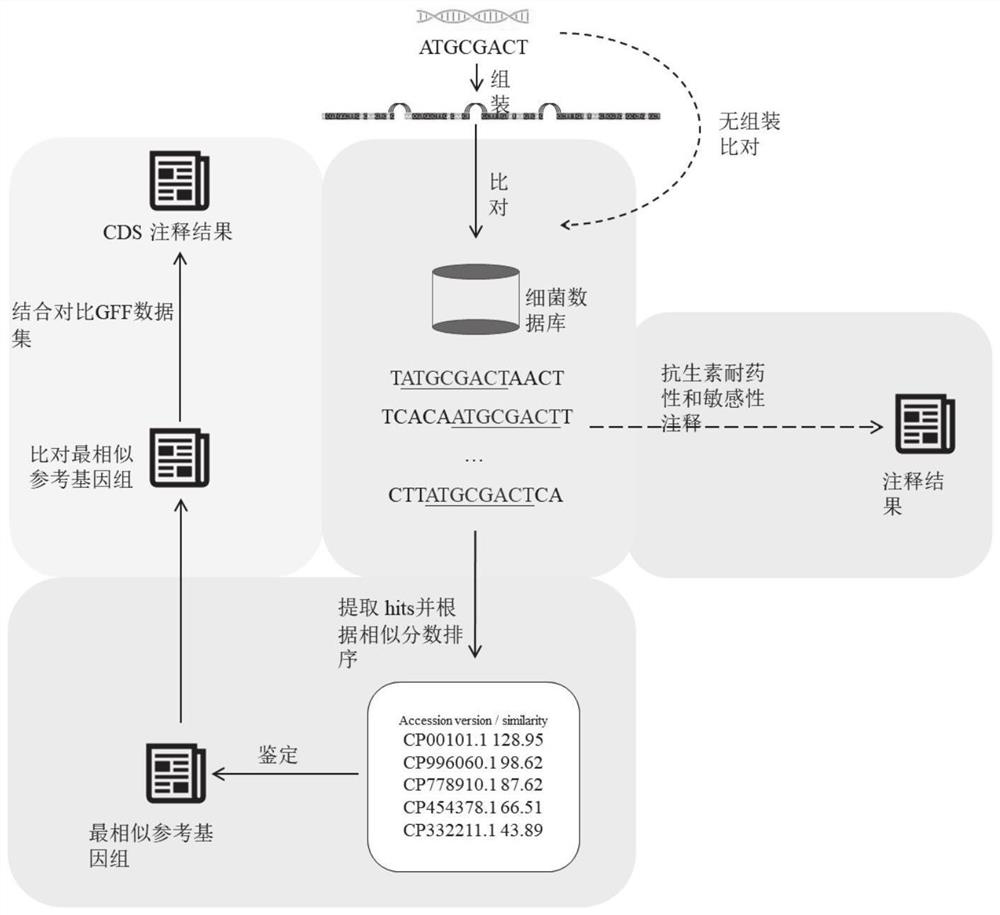
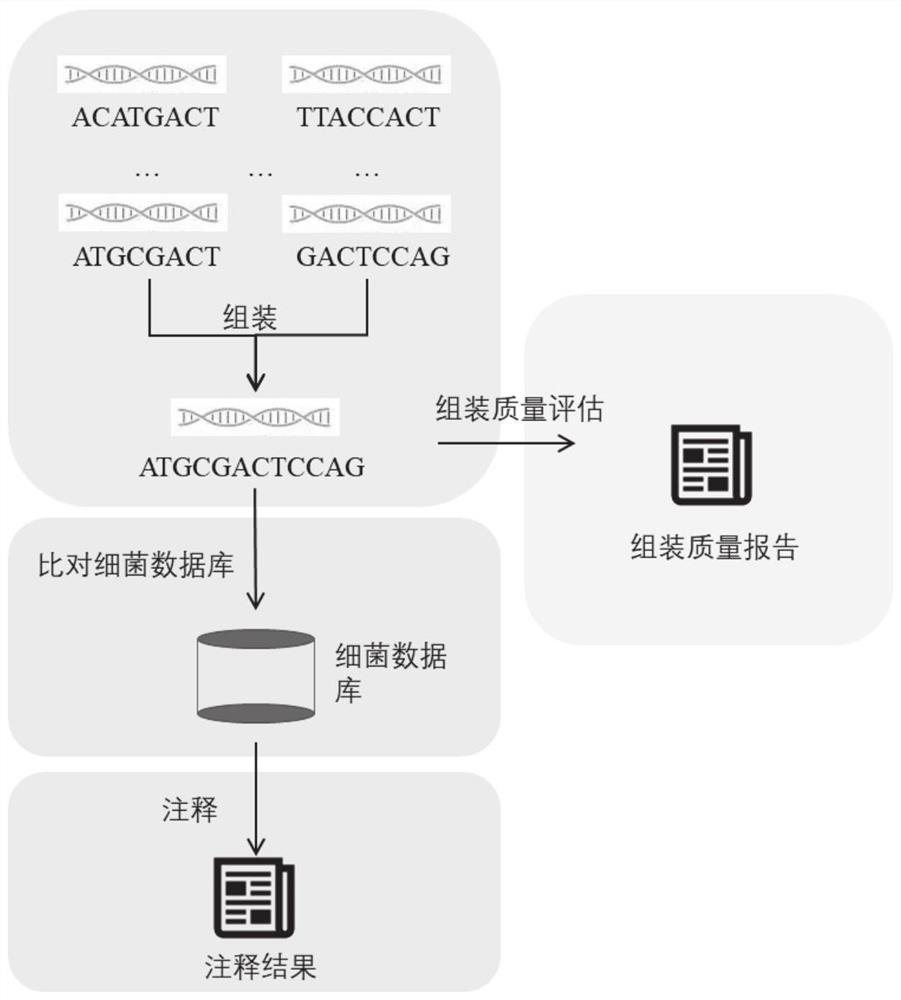
Automatic analysis method and system for sequencing data of whole genome of bacteria
PendingCN112863603AAchieve freedom of choiceImplement custom settingsSequence analysisInstrumentsSequence analysisGenomic sequencing
The invention provides an automatic analysis method for bacterial whole genome sequencing data, which comprises the following steps: acquiring bacterial genome sequencing data, and judging the type of the sequencing data; respectively carrying out corresponding preprocessing according to the type of the sequencing data; performing re-sequencing analysis and de novo sequencing analysis on the preprocessed sequencing data according to an analysis type selected by a user and preset tool software and software parameters; and realizing the identification and annotation of the whole genome of the bacteria. The scheme provides a user-friendly automated analysis method, for researchers and clinicians without professional bioinformatics knowledge, automated bioinformatics analysis steps including sequencing quality control, re-sequencing and de novo assembly, similar bacterial reference genome identification, bacterial genome annotation, and at the same time, researchers and clinicians without professional bioinformatics knowledge; meanwhile, customized bioinformatics analysis can be carried out according to short-read-length and long-read-length sequencing data generated by different platforms, and an accurate analysis result is obtained.
Owner:NANKAI UNIV
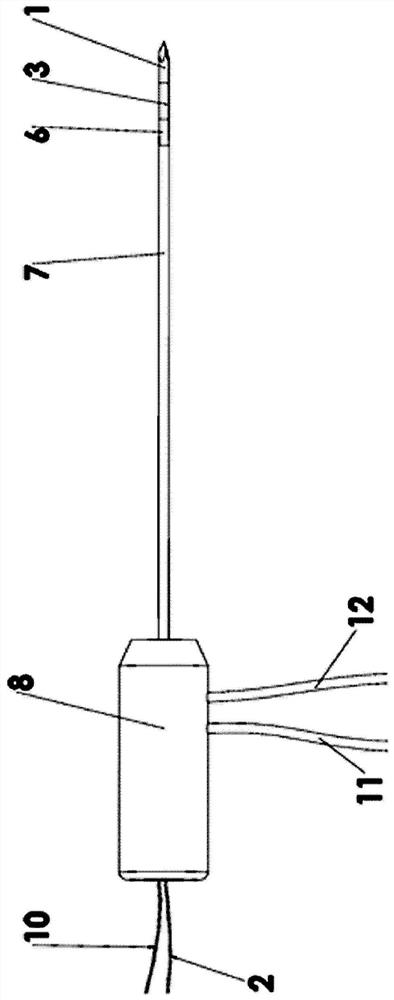
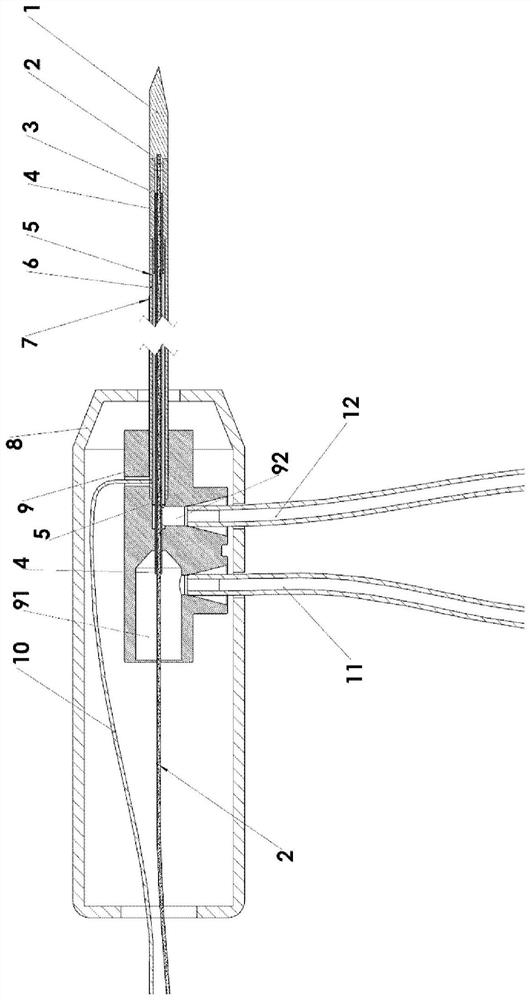
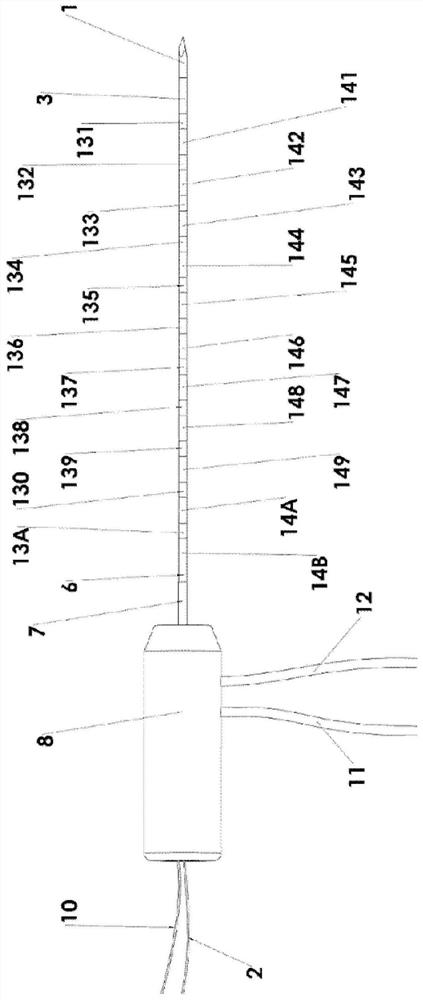
Multipole electroporation ablation needle and electroporation ablation equipment adopting multipole electroporation ablation needle
PendingCN113069201AReduce in quantityReduced collateral damageSurgical needlesSurgical instruments for heatingNuclear medicineMetal electrodes
The invention discloses a multipole electroporation ablation needle and electroporation ablation equipment adopting the multipole electroporation ablation needle. The multipole electroporation ablation needle comprises a needle rod, wherein the needle rod is provided with at least two metal electrodes; every two metal electrodes can be set into mutual electric insulation, and each metal electrode is provided with an electrode connecting conductor of which one end is connected with high-voltage output equipment and can be independently set into a positive electrode or a negative electrode at will; and in addition, the multipole electroporation ablation needle can carry out high-voltage discharging electroporation ablation between any two metal electrodes. The electroporation ablation needle can reduce insertion injuries, a focus on any position of the needle rod can be subjected to ablation under a situation that the depth of the electrode needle is not moved, surgical time is shortened, and the pain of a patient is lightened.
Owner:SINOSURGICAL HEALTHCARE TECH ZHEJIANG CO LTD
Data visualization color matching extraction method based on image clustering
PendingCN111768469AFlexible color matchingAchieve freedom of choiceImage analysisCharacter and pattern recognitionHistogramColor matching
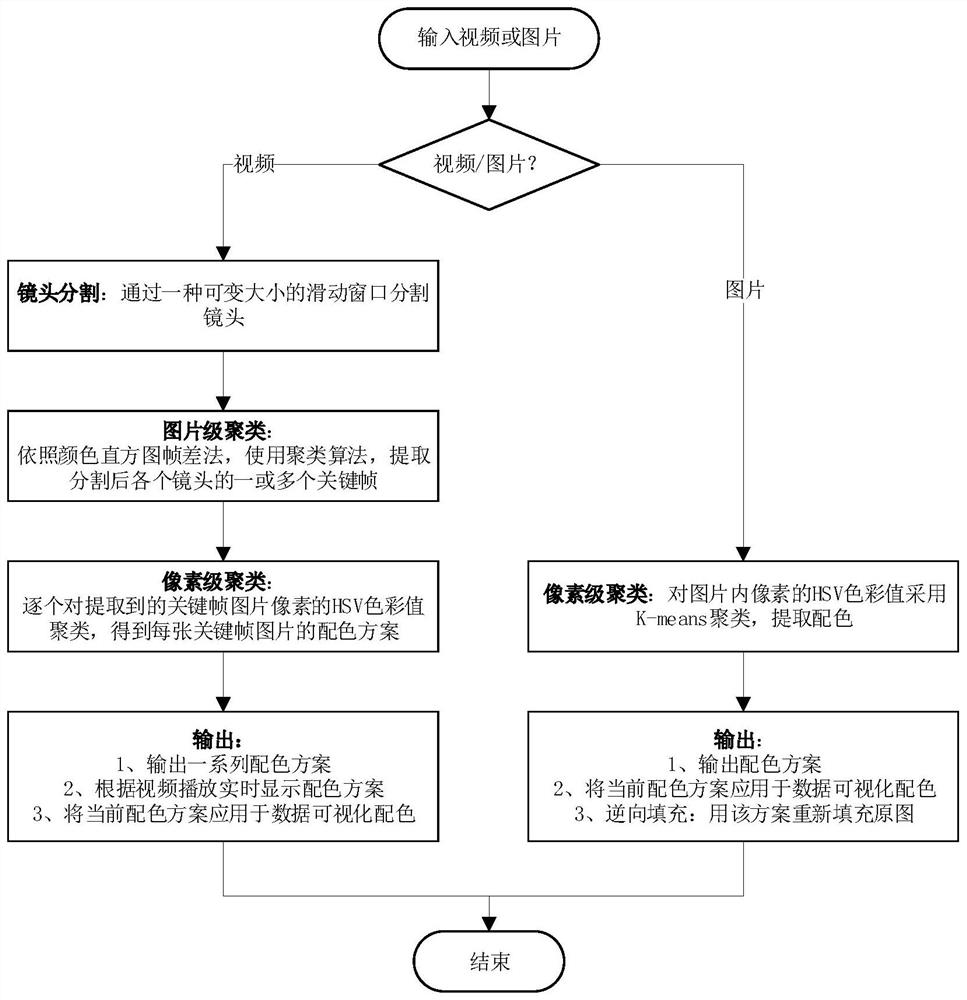
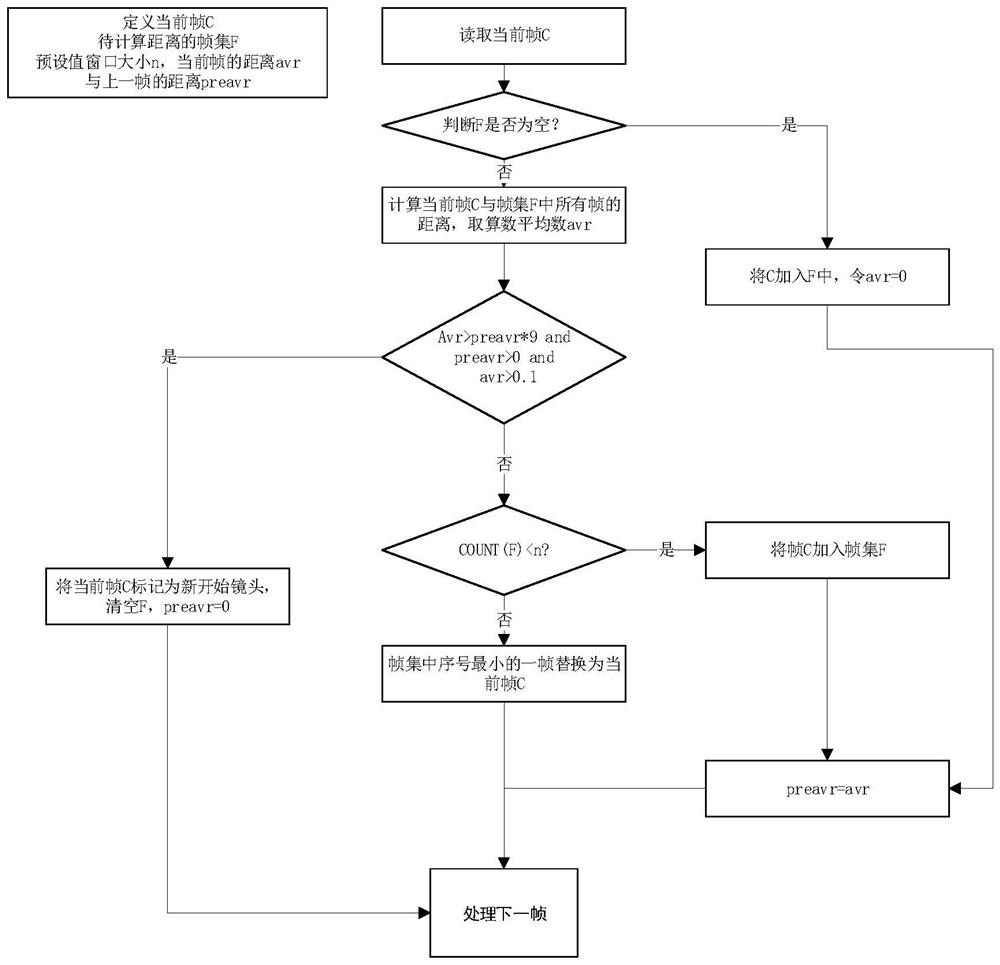
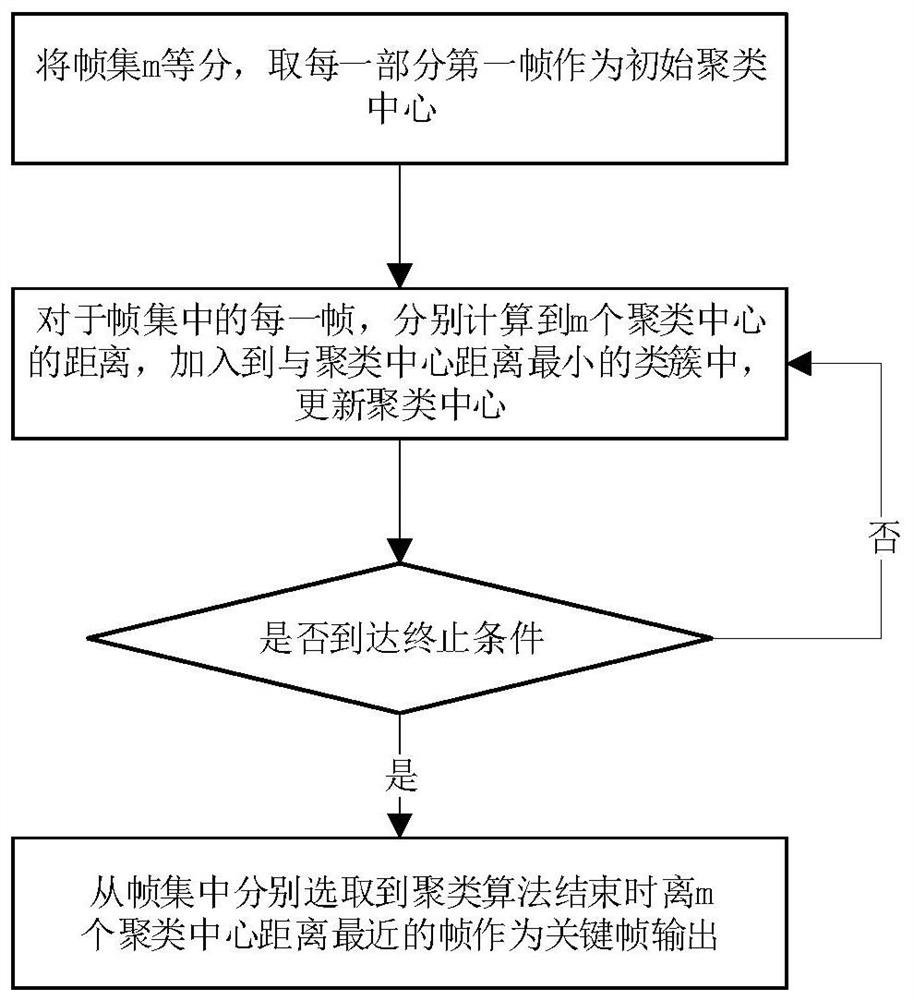
The invention discloses a data visualization color matching extraction method based on image clustering. The method comprises the steps: inputting a video or a picture; segmenting the video through asize-adjustable sliding window to obtain a plurality of shot video frames; extracting one or more key frames of each shot video frame after segmentation through a color histogram frame difference method and a clustering algorithm; clustering the HSV color values of the pixels in the extracted key frame pictures one by one to obtain a color matching scheme of each key frame picture; or clustering the HSV color values of the pixels in the input picture to obtain a color matching scheme of the picture; and outputting a color matching scheme, and applying the current color matching scheme to datavisual color matching. The method is low in time complexity and can be used for picture extraction color matching or video extraction color matching, a color matching method changing along with a video playing time sequence is actually provided based on video color matching, an original picture is reversely filled with a color matching scheme, and the subjective approximate consistency of color extraction and the original picture is verified.
Owner:COMMUNICATION UNIVERSITY OF CHINA
Vehicle positioning system and method
InactiveCN105390012AAvoid positioningRealize real-time monitoringRoad vehicles traffic controlLocation information based serviceComputer moduleFilling-in
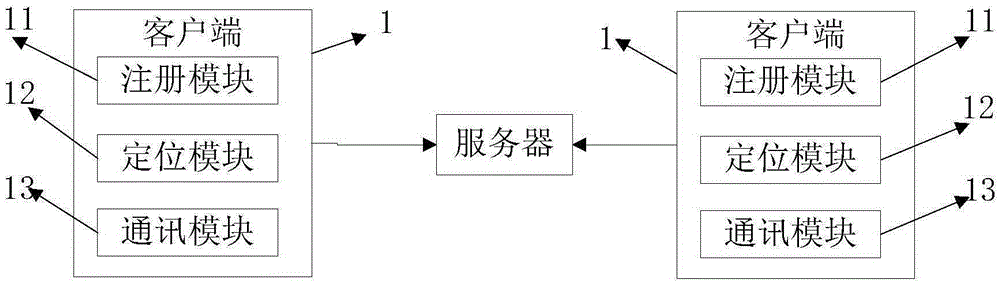
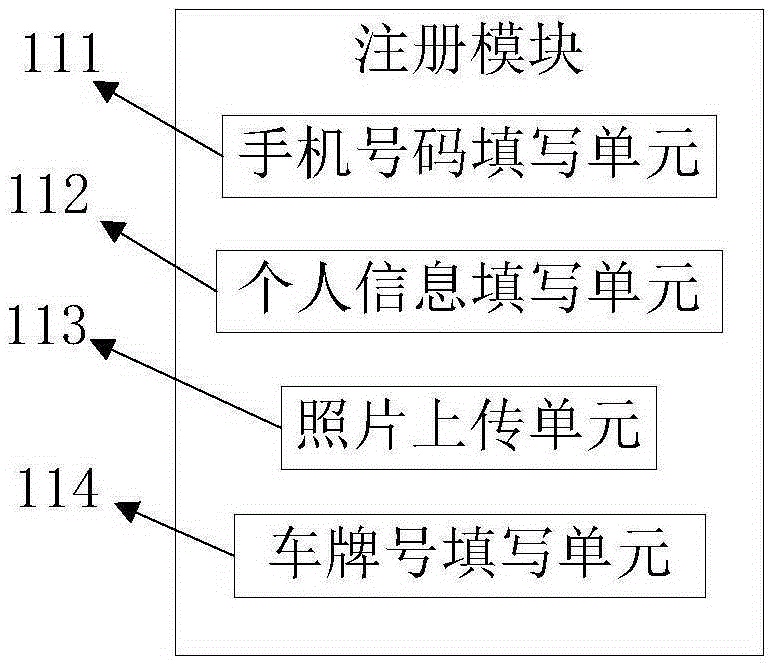
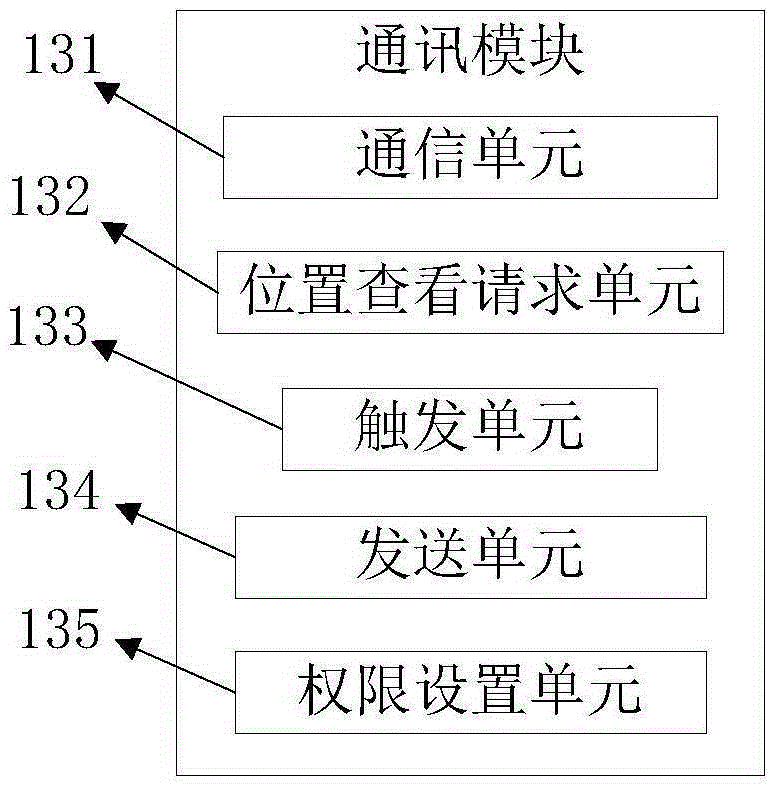
The invention relates to a vehicle positioning system and method. The system is composed of a client and a server. The client consists of a registration module, a positioning module, and a communication module; the registration module is used for filling in registration information and uploading the information to the server for storage; the positioning module is used for carrying out positioning on a position of a vehicle where a registered user is located; and the communication module is used for carrying out friend adding between the registered use and other registered users and checking the positioning information of the positions of the vehicles where the added friends are located according to positioning information checking permission set by the added friends after the friend adding requests are accepted. Therefore, real-time monitoring of a vehicle position is realized.
Owner:郭少方
Intelligent upper arm thrombus-preventing muscle massager
ActiveCN108498294AQuick fixEasy to determineBlood stagnation preventionPneumatic massageControl systemThrombus
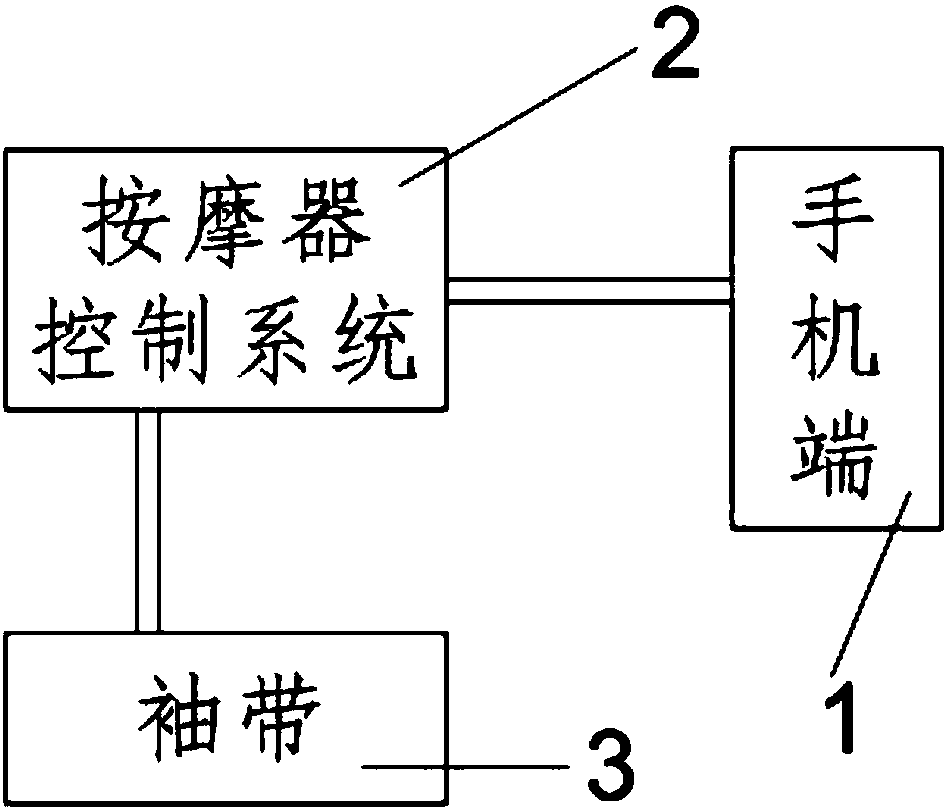
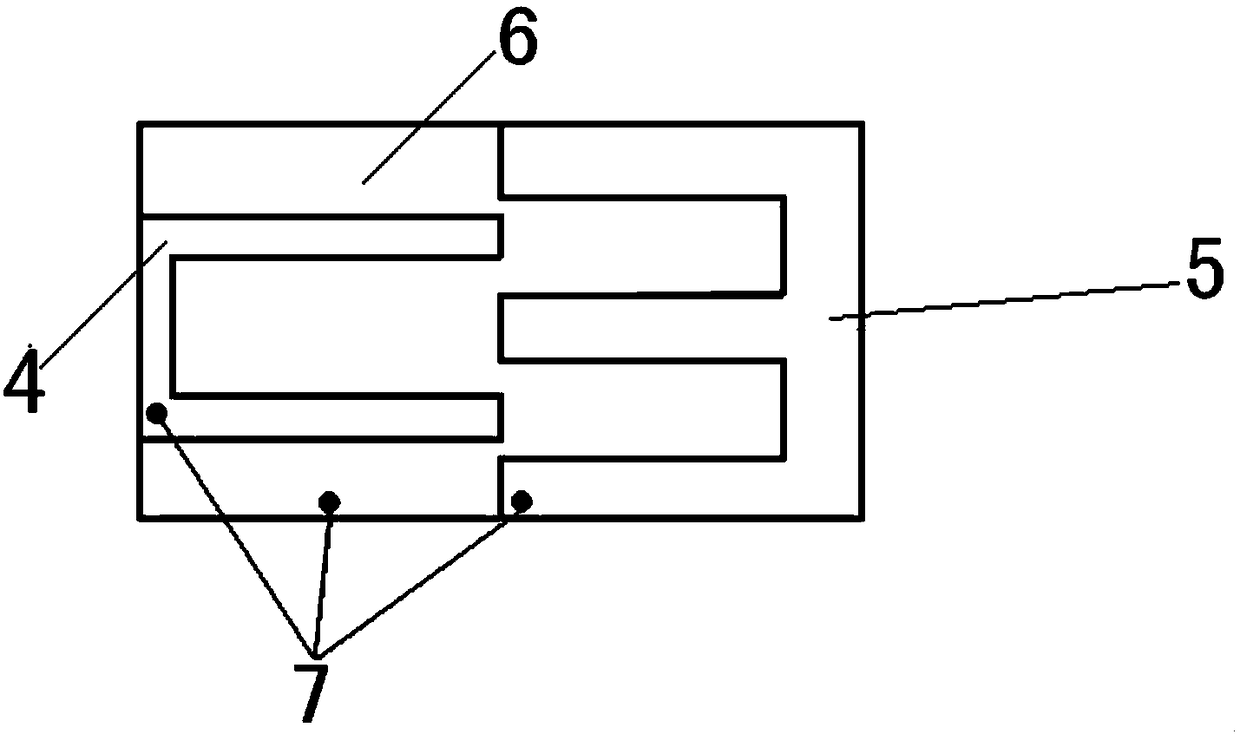
The invention provides an intelligent upper arm thrombus-preventing muscle massager. The intelligent upper arm thrombus-preventing muscle massager comprises a mobile phone side, a massager control system and a cuff, wherein the mobile phone side is provided with an APP client side, and the mobile phone side is connected with the massager control system through the APP client side; the massager control system is connected with the cuff; the massager control system is used for adjusting the massage pressure, massage time, massage speed and massage mode. The intelligent upper arm thrombus-preventing muscle massager has the advantages that the medical resources and user time are saved, the burden of medical workers and family members is alleviated, and the comfort and convenience are obtained.
Owner:THE FIRST AFFILIATED HOSPITAL OF XIAN JIAOTONG UNIV
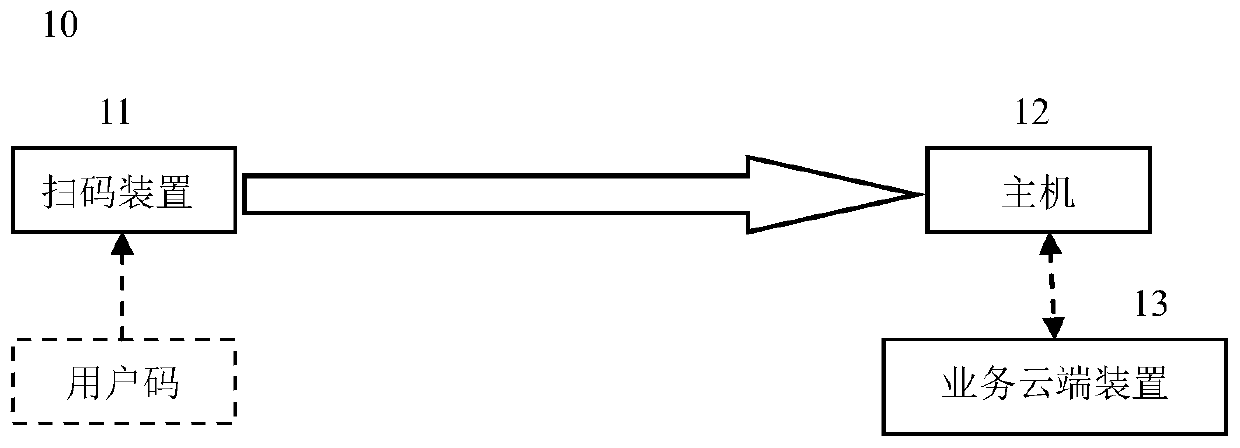
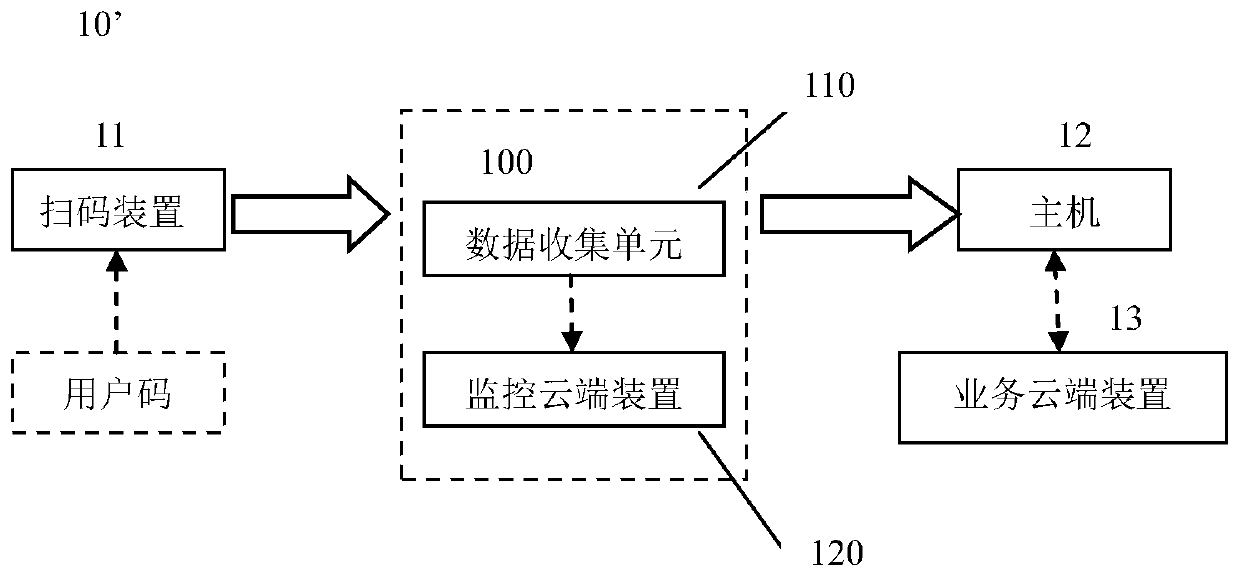
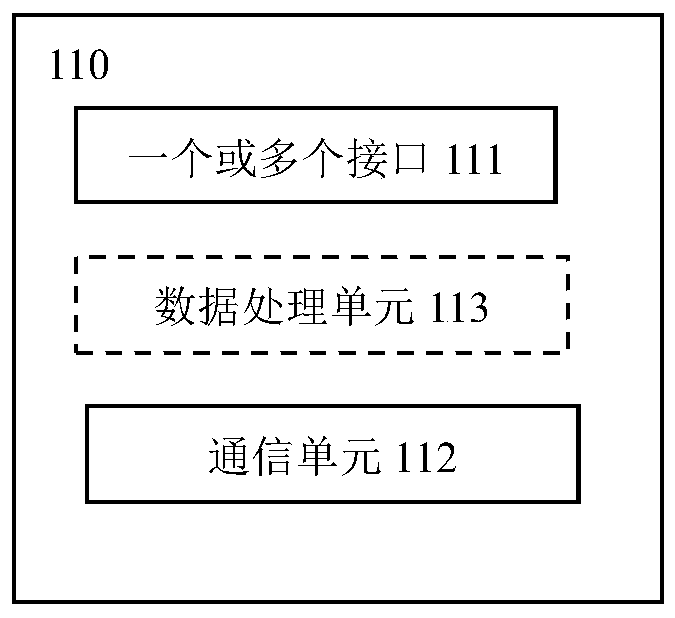
Device and method for monitoring code scanning operation performed by code scanning device
ActiveCN110009074AScan code operation monitoring/evaluationReduce in quantityCo-operative working arrangementsPayment architectureComputer hardwareCommunication unit
A device for monitoring a code scanning operation performed by a code scanning device is provided. The device includes a data collection unit configured to collect data related to the code scanning operation; and a monitoring cloud device configured to perform analysis based on the data so as to determine information related to the code scanning operation. The data collection unit further comprises a communication unit, and the communication unit is configured to send the data from the data collection unit to the monitoring cloud device. Therefore, the monitoring / evaluation of the code scanning operation executed by the merchant / service provider can be conveniently realized.
Owner:ADVANCED NEW TECH CO LTD
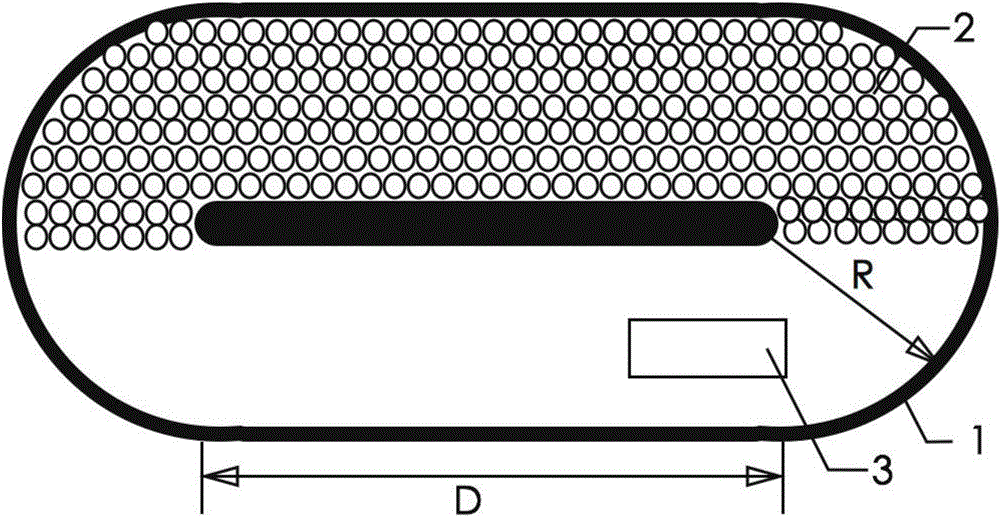
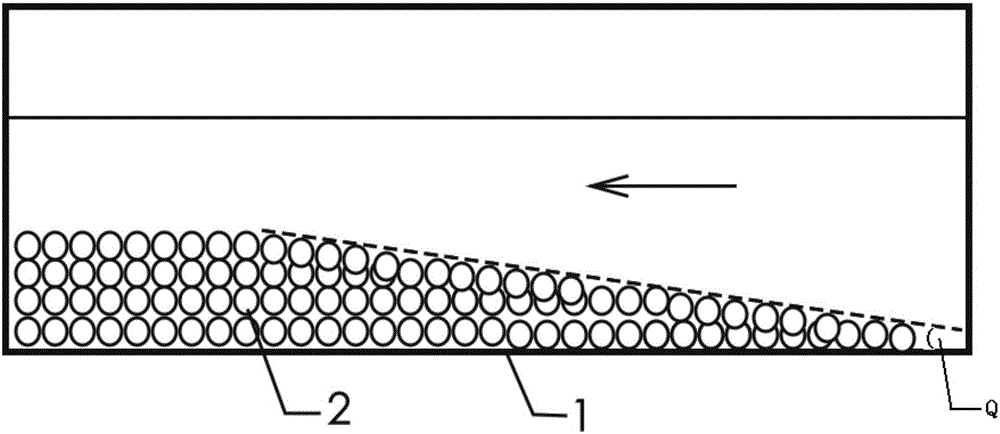
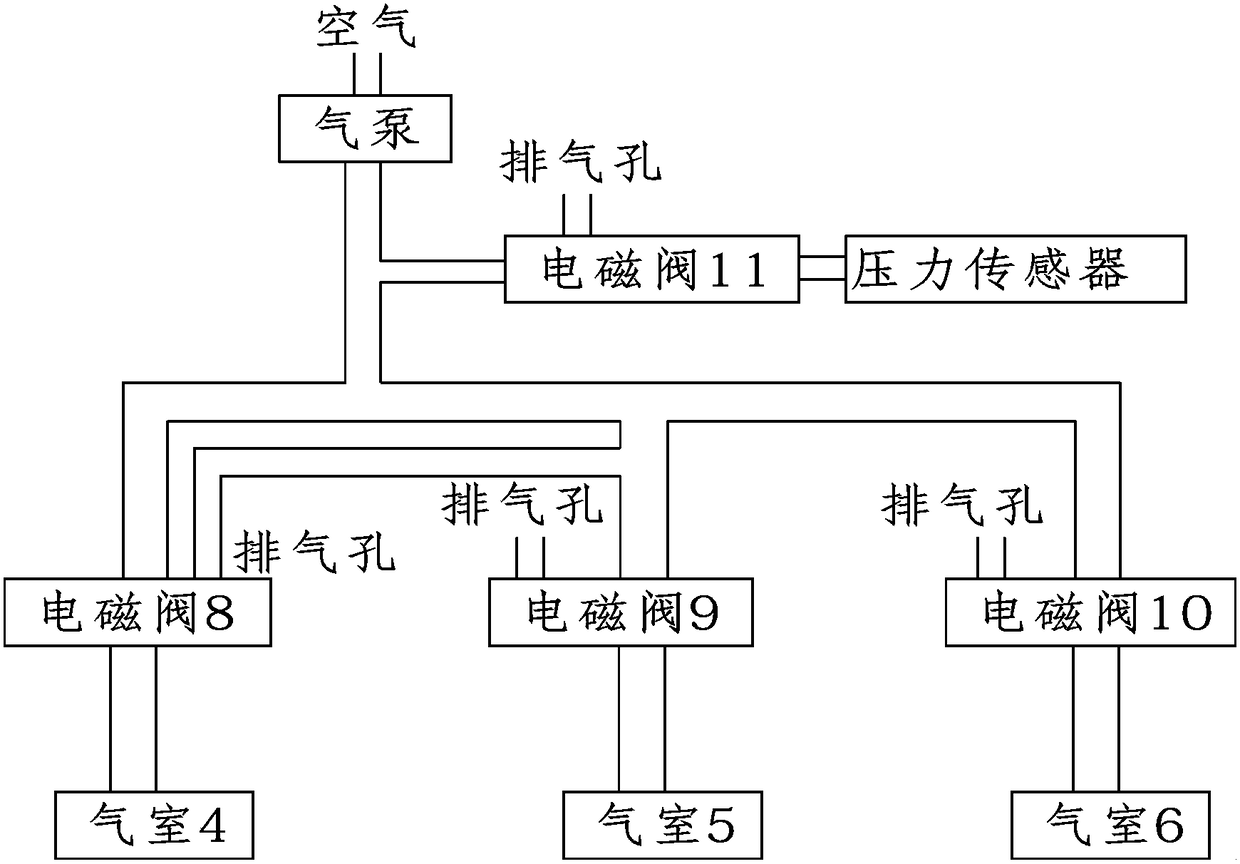
Ventilation and pressure release type sound absorption and insulation barrier for high-speed railway
ActiveCN101899817BGood ventilation and pressure reliefReduce mechanical loadNoise reduction constructionRailway tracksCarrying capacityEngineering
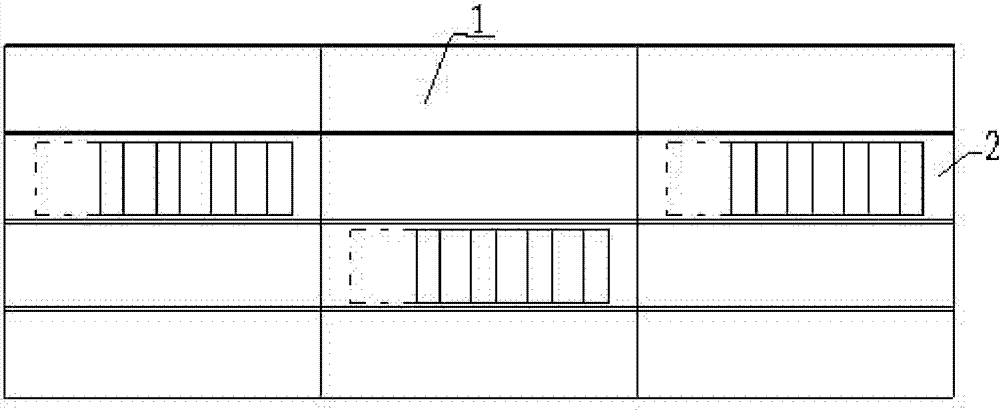
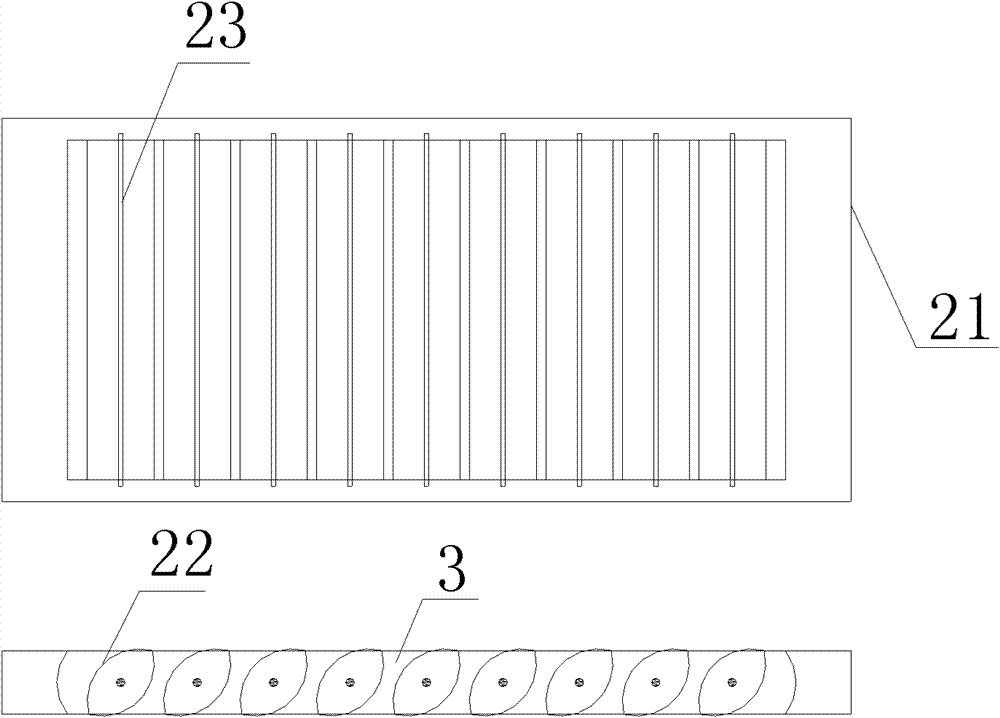
The invention discloses a ventilation and pressure release type sound absorption and insulation barrier for a high-speed railway, belonging to the field of noise reduction of the high-speed railway. The barrier of the invention comprises a sound barrier sound insulation member and at least one ventilation and pressure release type sound barrier module for ventilation and pressure release, whereinthe ventilation and pressure release type sound barrier module is embedded in the sound barrier sound insulation member and is combined with the sound barrier sound insulation member to form the ventilation and pressure release type sound absorption and insulation barrier. The invention provides a sound barrier module which takes the air pressure difference formed between the inner side and the outer side of a sound barrier when a high-speed train passes through the sound barrier as power to realize the purpose of automatically adjusting the angle of rotary muffled blades of the invention to achieve an optimal pressure release way, and the sound barrier module is combined with the sound barrier sound insulation member of the prior art to form the sound absorption and insulation barrier integrating the functions of sound insulation, noise elimination, noise reduction, ventilation and pressure release, thereby greatly improving the carrying capacity and the economical efficiency of the existing sound insulation barrier for railways.
Owner:ZISEN ENVIRONMENTAL TECHNOLOGY CO LTD
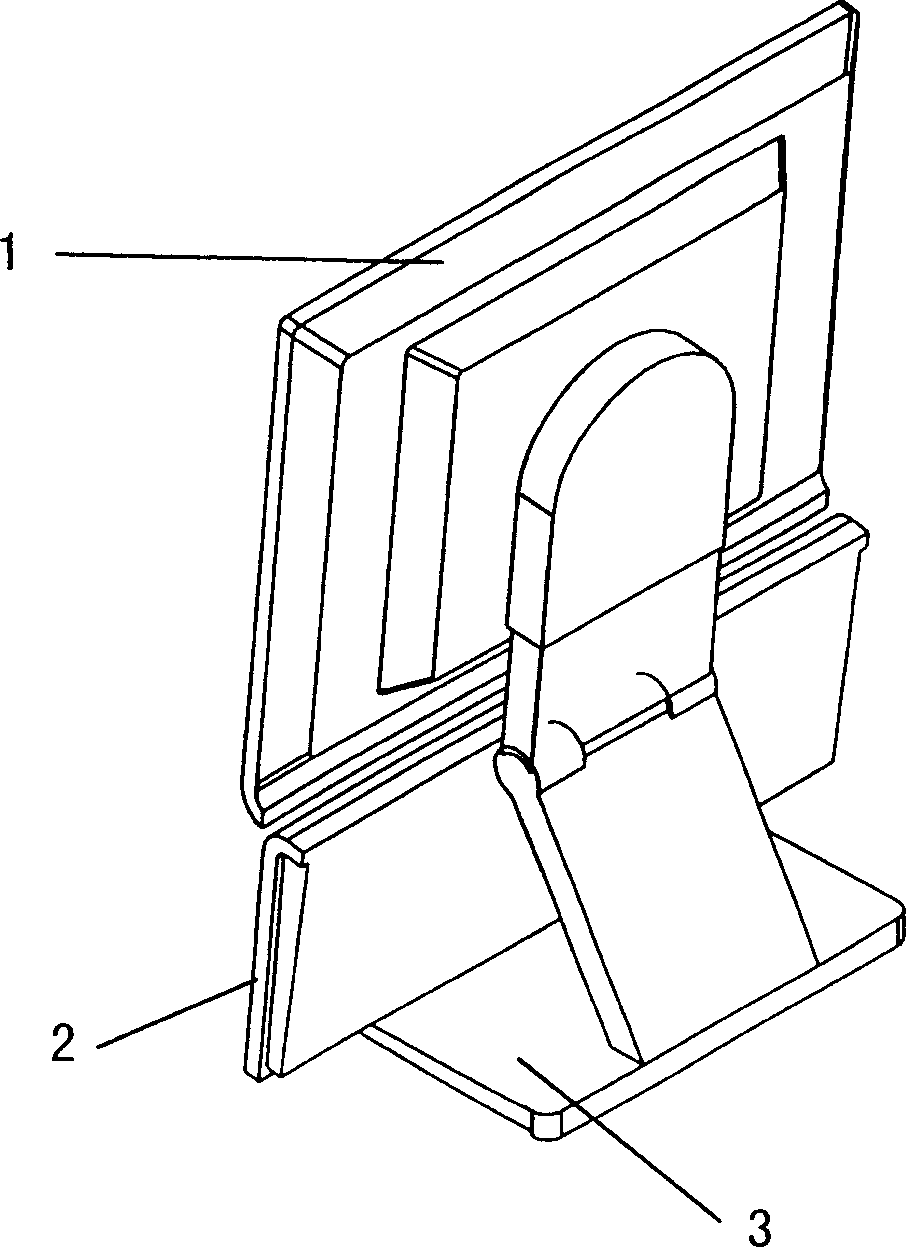
Display of computer capable of integrated assembling with keyboard
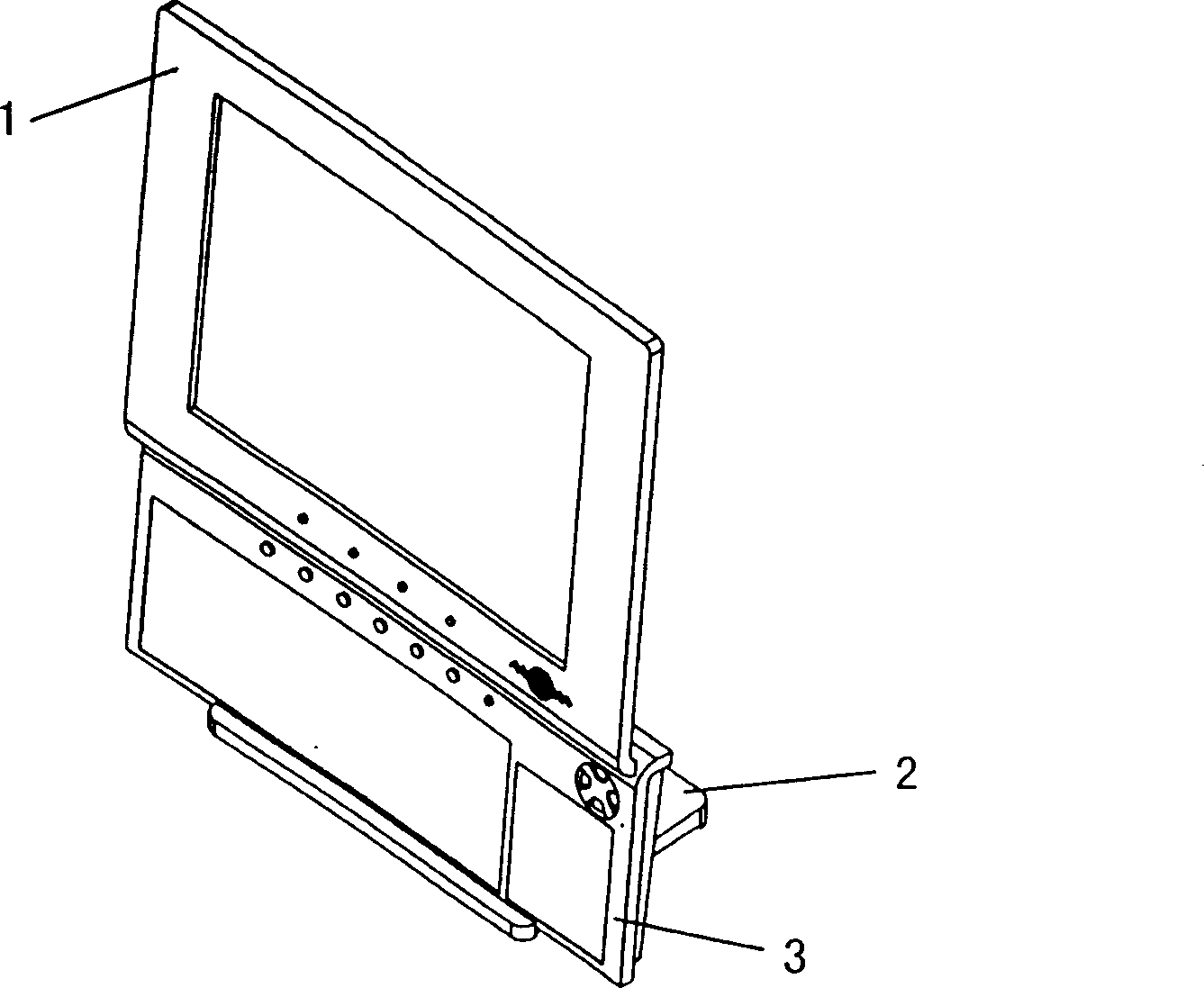
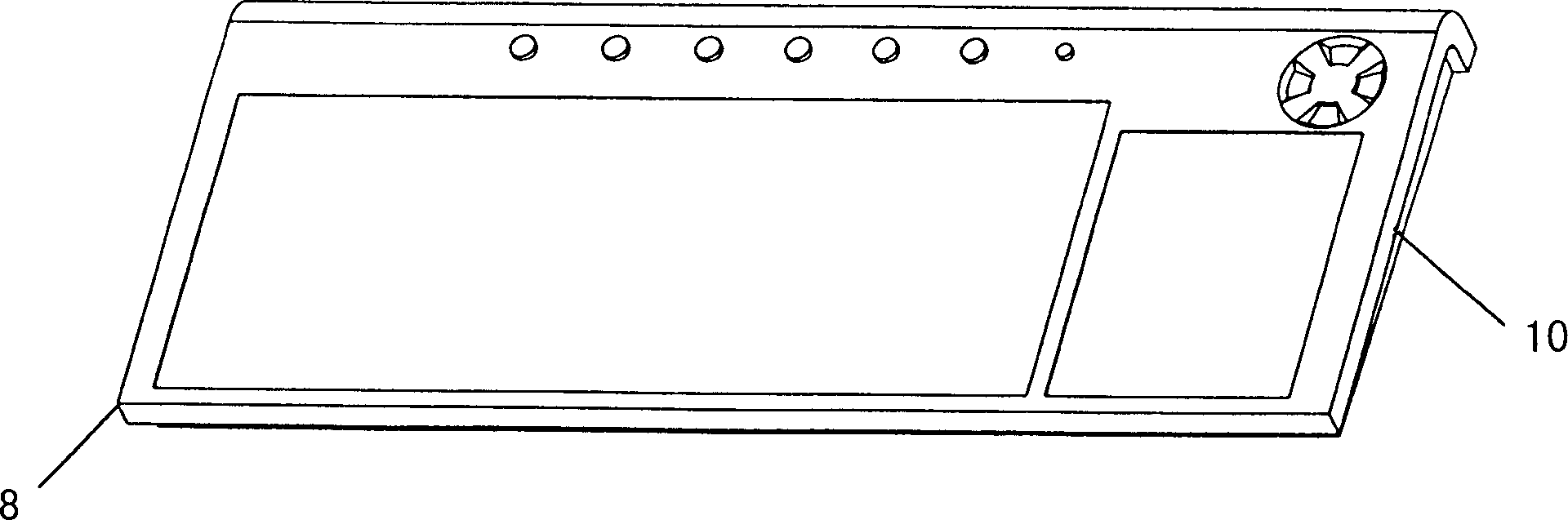
InactiveCN1434362AAchieve freedom of choiceAdapt to the development trend of home appliancesDigital data processing detailsLeading edgeDisplay device
The invention discloses a computer montor able to be integrated with the keyboard, including the monitor and the keyboard, and these is the position slot in parallel with the plane where the monitor lies set on the leading edge of the base board of the monitor pedestal, and the leading edge of the keyboard is the position one which can be inserted in the position slot and their shapes of cross section match each other in the direction perpendicular to that of the length of the position edge; there is the lift mechanism set on the top of the support stand on the back of the monitor. It realizes the free selection of the vertical using mode or the common one.
Owner:LENOVO (BEIJING) LTD
Features
- R&D
- Intellectual Property
- Life Sciences
- Materials
- Tech Scout
Why Patsnap Eureka
- Unparalleled Data Quality
- Higher Quality Content
- 60% Fewer Hallucinations
Social media
Patsnap Eureka Blog
Learn More Browse by: Latest US Patents, China's latest patents, Technical Efficacy Thesaurus, Application Domain, Technology Topic, Popular Technical Reports.
© 2025 PatSnap. All rights reserved.Legal|Privacy policy|Modern Slavery Act Transparency Statement|Sitemap|About US| Contact US: help@patsnap.com