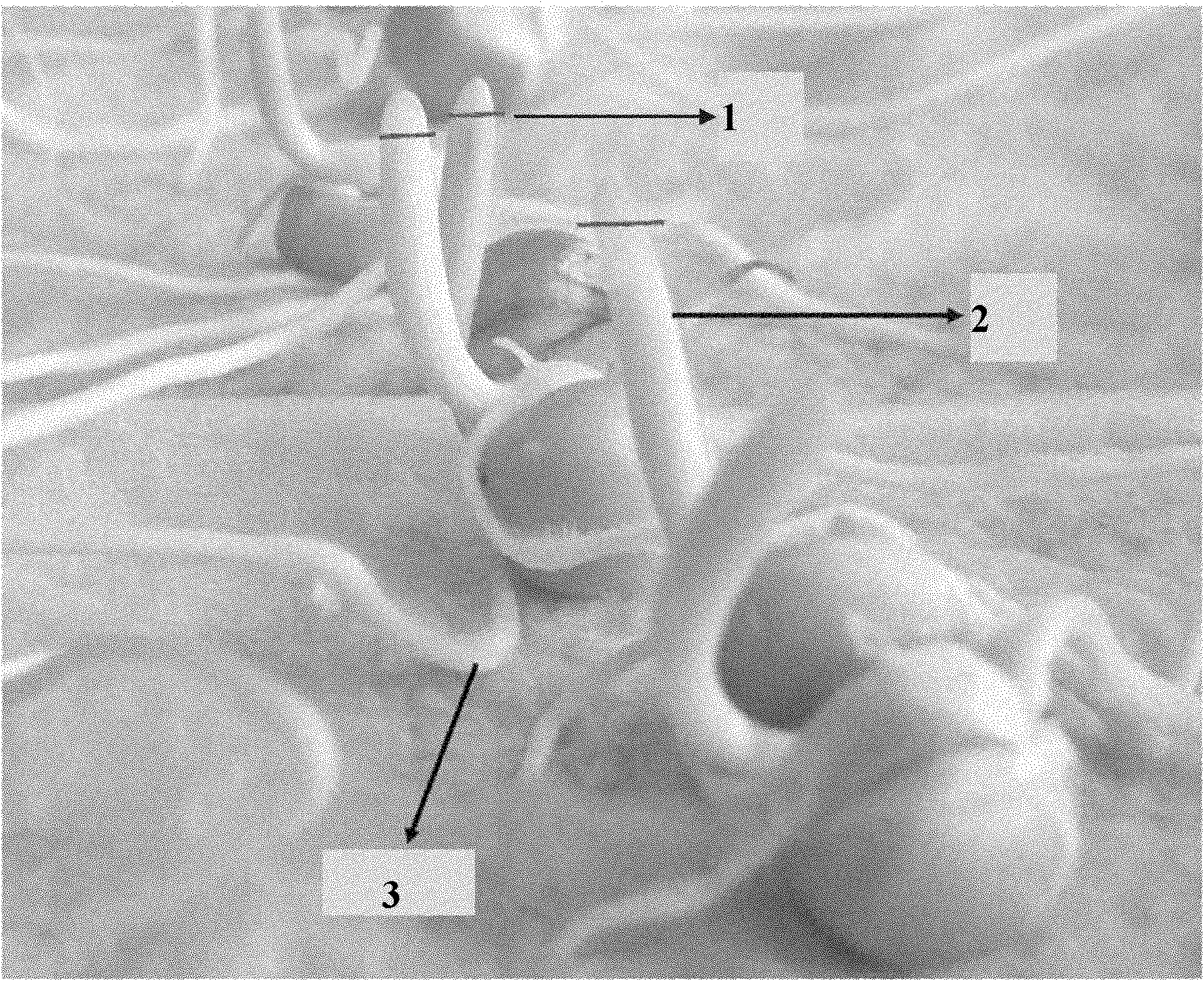Patents
Literature
1794results about How to "Improve breeding efficiency" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
Rice blast resistance gene Pi9 gene specificity molecular marker Pi9SNP as well as preparation and application thereof
ActiveCN103305510AStrong specificityImprove throughputMicrobiological testing/measurementDNA preparationAgricultural scienceNucleotide
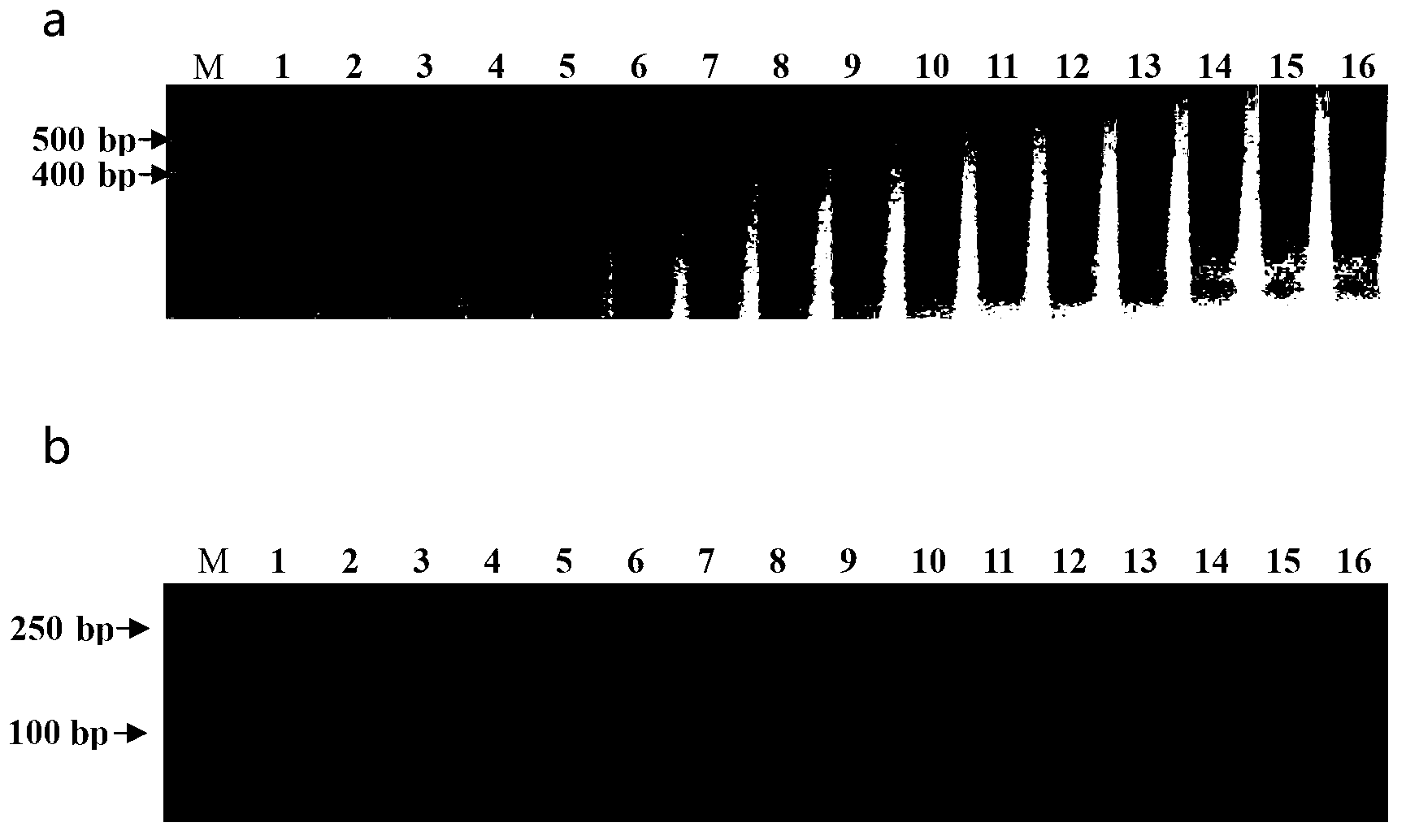
The invention discloses a rice blast resistance gene Pi9 gene specificity molecular marker Pi9SNP as well as preparation and application thereof. The molecular marker Pi9SNP is prepared by amplifying from total DNA (Deoxyribonucleic Acid) of a rice blast resistant variety with rice blast resistance gene Pi9 by using a primer pair F1 and R1 so as to obtain a nucleotide segment I, and carrying out digestion on the nucleotide segment I by using restriction enzyme Hind III, so as to obtain a nucleotide segment II which is in specificity banding pattern with the obtained rice blast resistance gene Pi9. The molecular marker is the first one which is developed for marking Pi9 gene specificity SNP (Signal Nucleotide Polymorphism) for an internal sequence of the Pig gene, and has the following advantages that the specificity is high, the cost is low and the flux is high in practical application, and the molecular marker is widely applied to colonies of different genetic backgrounds. By utilizing the molecular marker, the efficiency in auxiliary molecular marker selective breeding, gene pyramiding breeding and transgenosis breeding is improved.
Owner:PLANT PROTECTION RES INST OF GUANGDONG ACADEMY OF AGRI SCI
Functional molecular marker for rice anti-blast gene Pi9 and application thereof
ActiveCN103146695AImprove breeding efficiencyImprove accuracyMicrobiological testing/measurementPlant genotype modificationBiotechnologyNucleotide
The invention provides a functional molecular marker for rice anti-blast gene Pi9 and application thereof, belonging to the field of crop molecular genetic breeding science. The invention finds out that the rice anti-blast gene Pi9 has a 10bp insertion / deletion site positioned between the gene promoter -516 and -517, wherein the nucleotide sequence is disclosed as SEQ ID NO.1; and on such basis, the invention develops a method of the functional molecular marker for gene Pi9. By detecting the functional molecular marker, the invention can accurately detect whether genomes of different rice species contain the Pi9 functional gene and detect the homozygotic state, and can be used for screening rice hybrid transformed descendant plants to enhance the breeding efficiency of the rice anti-blast material, thereby effectively controlling the size of the breeding population, obviously saving the breeding and screening cost and obtaining the anti-blast rice species containing the Pi9 functional gene.
Owner:BIOLOGICAL TECH INST OF FUJIAN ACADEMY OF AGRI SCI
Gene-specific molecular marker Pi2SNP of rice blast-resistant gene Pi2 as well as preparation method and application thereof
ActiveCN103320437AStrong specificityHigh sequence identityMicrobiological testing/measurementDNA preparationAgricultural scienceEnzyme digestion
The invention discloses a gene-specific molecular marker Pi2SNP of a rice blast-resistant gene Pi2 as well as a preparation method and application thereof. The molecular marker Pi2SNP is a nucleotide fragment II in a specific banding pattern with the rice blast-resistant gene Pi2 obtained by amplifying F1 and R1 from the total DNA (Deoxyribose Nucleic Acid) of the rice blast-resistant variety carrying rice blast-resistant gene Pi2 by use of a primer to obtain a nucleotide fragment I and then performing enzyme digestion of the nucleotide fragment I by use of restriction endonuclease Pst I or Hinf I. The molecular marker is the first Pi2 gene-specific SNP marker developed for the sequence in the Pi2 gene, and has the advantages of high specificity and low cost and high flux in practical application, and can be widely applied to the colonies with different inheritance backgrounds. By adopting the molecular marker, the utilization efficiency of the gene in molecular marker-assisted selective breeding, gene pyramiding breeding and transgenic breeding can be improved.
Owner:PLANT PROTECTION RES INST OF GUANGDONG ACADEMY OF AGRI SCI
Upland cotton No. 4 chromosome and SNP molecular markers associated with fiber strength
ActiveCN106929574AShorten the breeding cycleImprove breeding efficiencyMicrobiological testing/measurementMolecular breedingSnp markers
The invention belongs to the technical field of cotton molecular breeding, and discloses SNP molecular markers associated with fiber strength of upland cotton as well as detection of the SNP molecular marker and application of the SNP molecular marker. The SNP molecular marker takes a cotton stable RIL group as a material, and is obtained through a genome re-sequencing method. The SNP markers are utilized to perform molecular marker assisted breeding, so that the breeding period can be greatly shortened, cotton fiber strength is enhanced, and the breeding efficiency is improved.
Owner:INST OF COTTON RES CHINESE ACAD OF AGRI SCI
Method for shortening juvenile period of apple hybrid seedlings
The invention discloses a method for shortening the juvenile period of apple hybrid seedlings, comprising the following steps of: culturing rootstock seedlings, culturing dwarf interstock seedlings, culturing apple hybrid seedlings, grafting seedlings at a high position, performing final-period management on trees and the like. According to the method, can be realized, and the semi-finished seedlings grafted at a high position after planting can blossom and fruit after growing to the 4th year, and have the fruiting plant rate up to 20% in the 5th year and up to 40% in the 6th year. Traditional seedlings need 8-10 years of juvenile period, so the juvenile period of the seedlings is shortened by 4 years; and compared with a traditional method, the plant rate after the juvenile period is greatly increased as well. According to the invention, measures of curing dwarf interstock seedlings early, sowing seedlings, taking the high-position buds of the seedlings and grafting the high-position buds to the dwarf interstock seedlings, and then planting, and other steps are adopted, and interstock grafting is performed on the apple seedlings, thus greatly shortening the juvenile period of the apple seedlings, shortening breeding process and fruiting time, increasing breeding efficiency, and saving lots of field management cost and resource investment simultaneously.
Owner:SHIJIAZHUANG POMOLOGY INST OF HEBEI ACADEMY OF AGRI & FORESTRY SCI
Application of C-type natriuretic peptide to promotion on in vitro maturation of bovine oocyte
ActiveCN102899286AImprove in vitro development abilityAccelerate the breeding and propagation technology system of fine varietiesNew breed animal cellsCell culture active agentsMeiosis InhibitionBrain natriuretic peptide
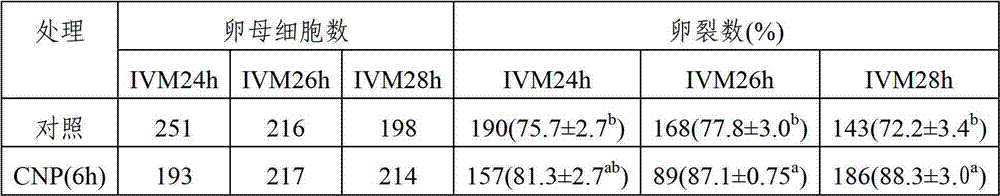
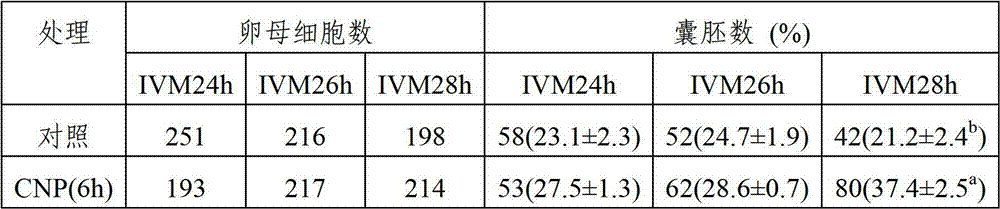
The invention relates to application of a C-type natriuretic peptide (CNP) to promotion on in vitro maturation of bovine oocyte. The invention provides a culture liquid for in vitro maturation of bovine oocyte; and a conventional culture liquid is used as a matrix containing C-type natriuretic peptide. According to the invention, CNP is used for in vitro inhibition on bovine oocyte meiosis and premature treatment to promote synchronization of oocyte nuclear maturation and cytoplast maturation, and improve ability of ectogenesis. The CNP can be promoted and applied as a novel meiosis inhibitor to animal husbandry, so as to accelerate fine breed breeding and an expanding propagation technique system. As a biological activity peptide substance, the CNP provided by the invention has toxic effect on oocyte less than that of a chemically synthesized meiosis inhibitor.
Owner:CHINA AGRI UNIV
Method for producing pre-basic seeds of potatoes
InactiveCN102550411AEfficient heat transferGood for hardening seedlingsHorticulture methodsPlant tissue cultureEngineeringSolanum tuberosum
The invention discloses a method for producing pre-basic seeds of potatoes. The method comprises the steps as follows: laboratory production of potato tube seedlings, sugar-free culture and fast propagation in a tube seedling greenhouse, and matrix and liquid culture for producing pre-basic seeds of potatoes. According to the invention, a polypropylene film bag takes the place of a traditional glass container, so that the light transmission and the air permeability are good, the cost is low, the weight is light, and the carrying is simpler; the mode of matrix and liquid culture achieves the purpose of propagating pre-basic seeds in a layered manner, so that not only can vain growth of the overground parts be controlled effectively, and growth and development of creeping stems of plants are ensured, but also forming and expanding of tubers at the top ends can be promoted effectively, and the operation is simple and easy; the pre-basic seeds can be grown in a plurality of seasons in a year, the quantity of potatoes borne in a unit area and the yield are improved greatly, the cost is reduced considerably, the propagation efficiency is improved, and industrial and mass production of virus-free miniature seed potato of potatoes can be realized.
Owner:SICHUAN AAS HORTICULTURE RES INST
Screening method for breeding strong cold-resisting new rice variety
InactiveCN1411701AReduce blindnessImprove predictabilityPlant genotype modificationAgricultural scienceScreening method
The present invention discloses a screening method for breeding strong cold-resisting new rice cavity, it uses wile rice as female parent and uses cultured rice as male parent, and adopts the following steps: making sexual hybridization to obtain hybrid, planting to obtain seed, culturing said seed to grow two-leaves and one-heart, and making treatment for 48-72 hr. in refrigerator or biological culture box of -2- +1 deg.C, restoring for 5-10 days at normal temp. in the room, transplanting living seedling into field, after it is grown up and ripened, hibernating and screening at low temp. -5-+1 deg.C, using hibernated plant with regeneration capacity to make multiple hybridization or backcross, making its progeny undergo the indoor and outdoor low-temp. treatment and screen at -5 - +1 deg.C, making family selectino and obtaining said new variety.
Owner:江西省农业科学院水稻研究所
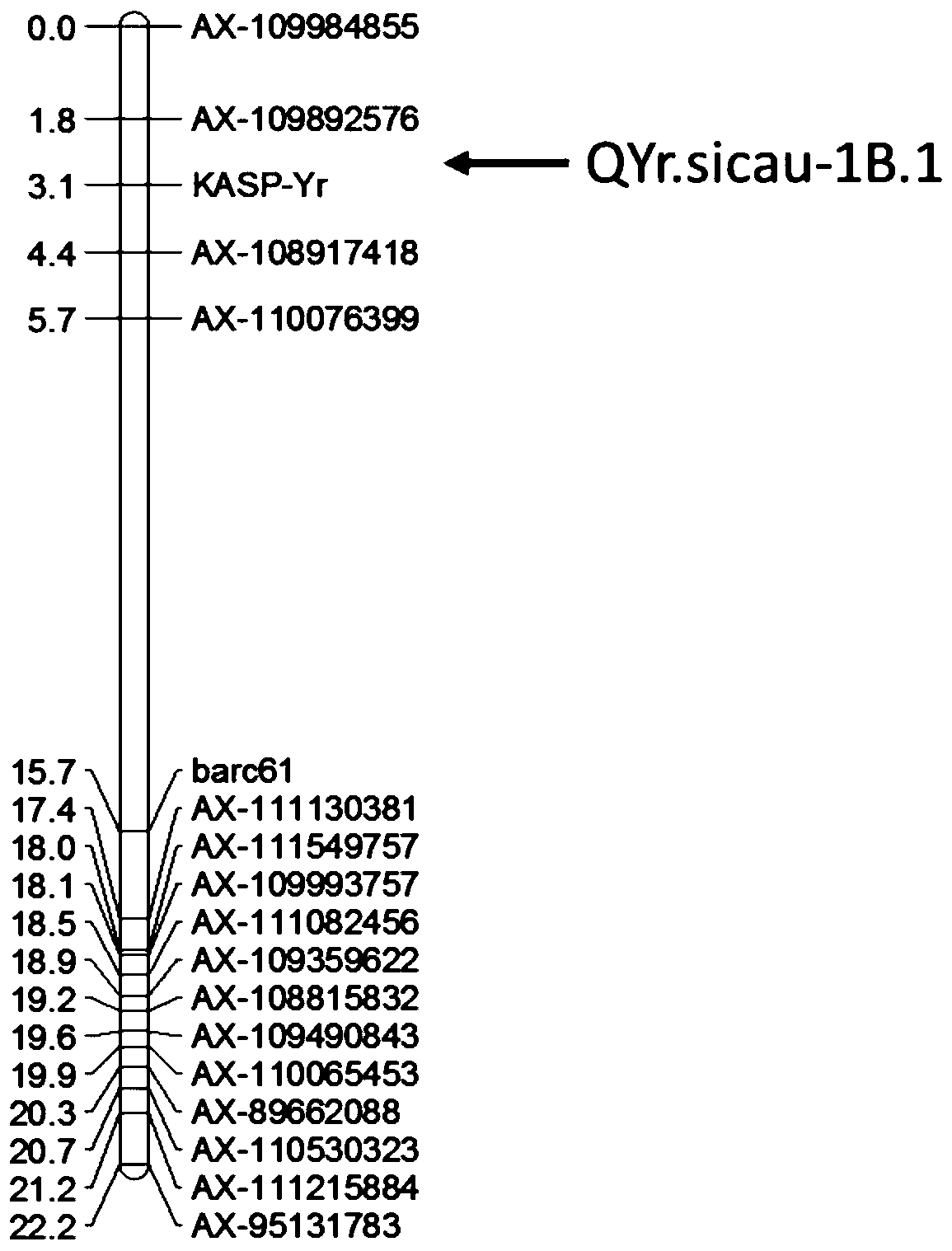
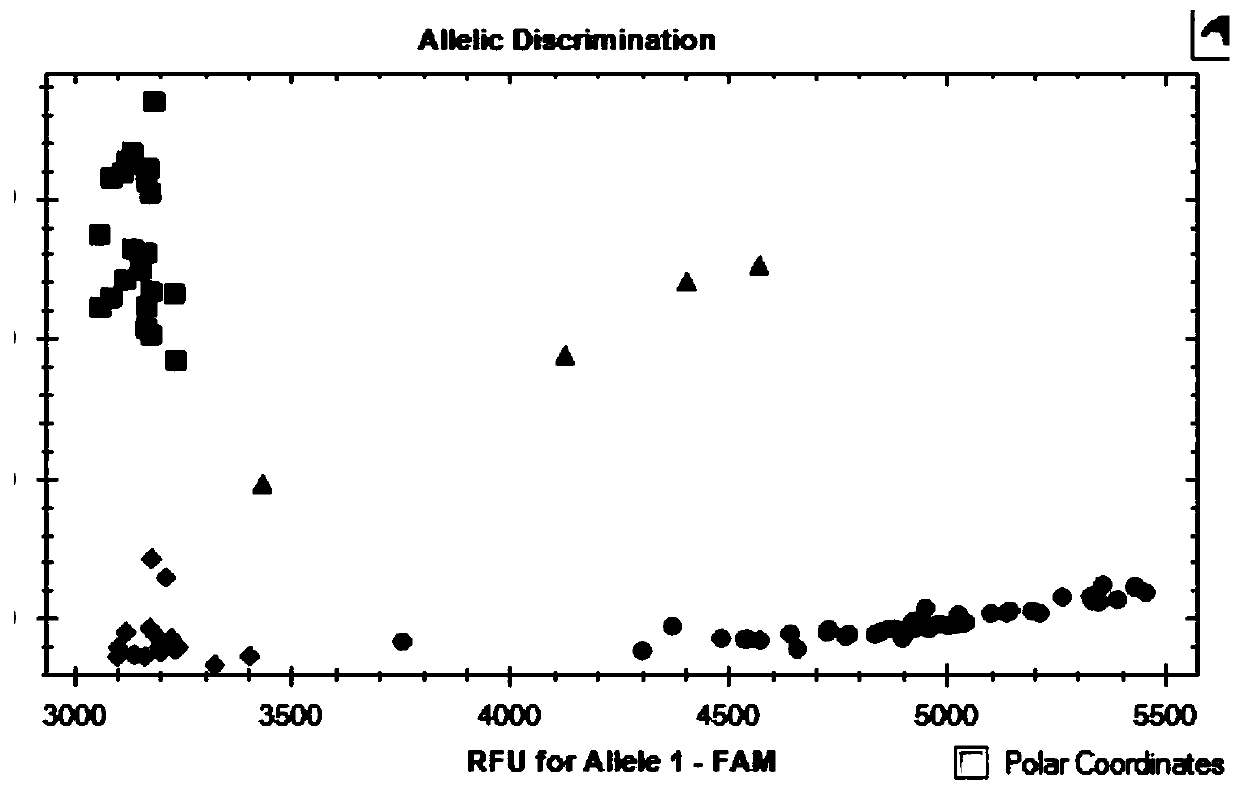
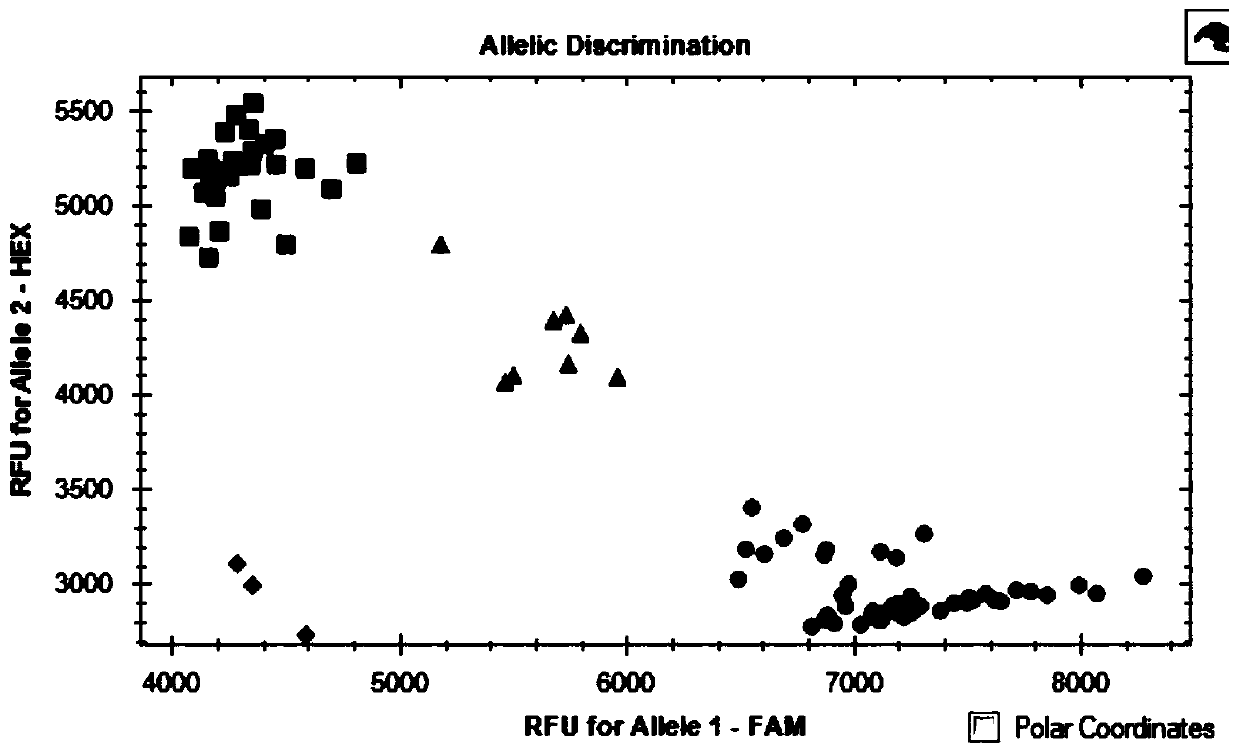
SNP molecular marker linked with wheat stripe rust resistance gene QYr.sicau-1B-1 and application thereof
ActiveCN109706263AQuick filterConvenient Assisted BreedingMicrobiological testing/measurementDNA/RNA fragmentationAgricultural scienceMolecular breeding
The invention belongs to the technical field of molecular breeding, and provides an SNP molecular marker linked with a wheat stripe rust resistance gene QYr.sicau-1B.1 and application thereof. The SNPmolecular marker is KASP-Yr, which can be obtained by amplifying a primer having a nucleotide sequence as shown in SEQ ID NO.1-3. Detection analysis indicates that the molecular marker can be used for accurately tracking wheat stripe rust resistant QTL QYr.sicau-1B.1 and predicting the stripe rust resistance characteristics of wheat, so that molecular designed breeding can be performance conveniently. The invention further discloses a method for identifying the molecular marker linked with wheat stripe rust resistance QTL QYr.sicau-1B.1. By utilizing the method, the stripe rust resistance prediction accuracy can be reinforced to quickly screen wheat variety or strain with stripe rust resistance for breeding, the breeding process for high-yield variety of wheat can be greatly accelerated.
Owner:SICHUAN AGRI UNIV
SNP molecular marker associated with chromosome 6 and fiber strength of upland cotton
ActiveCN106868131AShorten the breeding cycleImprove breeding efficiencyMicrobiological testing/measurementDNA/RNA fragmentationAgricultural scienceMolecular breeding
The invention belongs to the technical field of molecular breeding of cotton and discloses an SNP molecular marker associated with fiber strength of upland cotton as well as detection and application thereof. The SNP molecular marker is obtained by taking stable RIL group of cotton as the material through a genome resequencing method. When the SNP markers are used for carrying out molecular marker-assisted breeding selection, the breeding period can be greatly shortened; the breeding efficiency of the cotton can be improved; the cotton fiber strength can be improved.
Owner:INST OF COTTON RES CHINESE ACAD OF AGRI SCI
Plant growing device
InactiveCN104053355AConcentration homogenizationTemperature homogenizationMechanical apparatusLighting and heating apparatusVolumetric Mass DensityEngineering
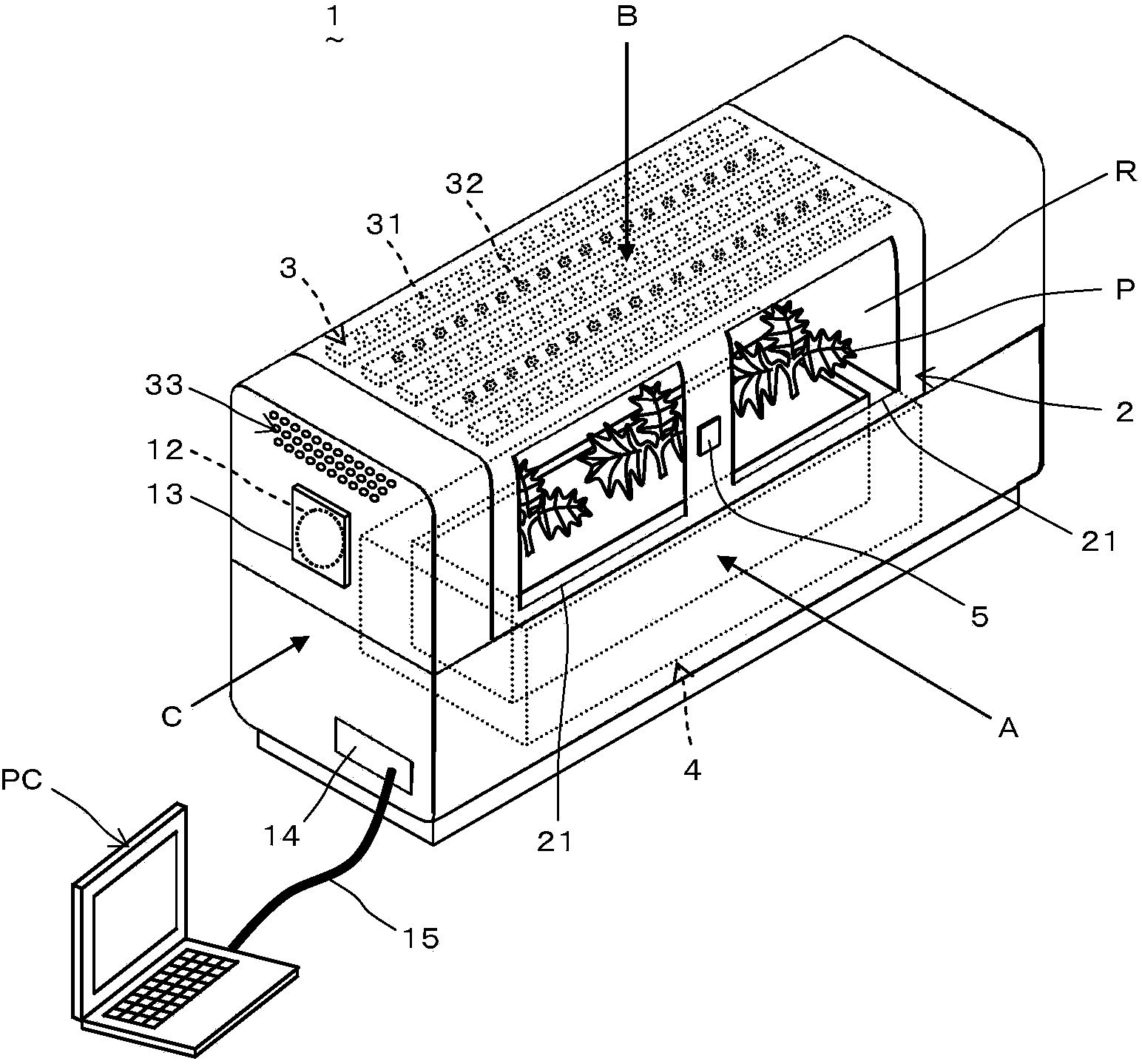
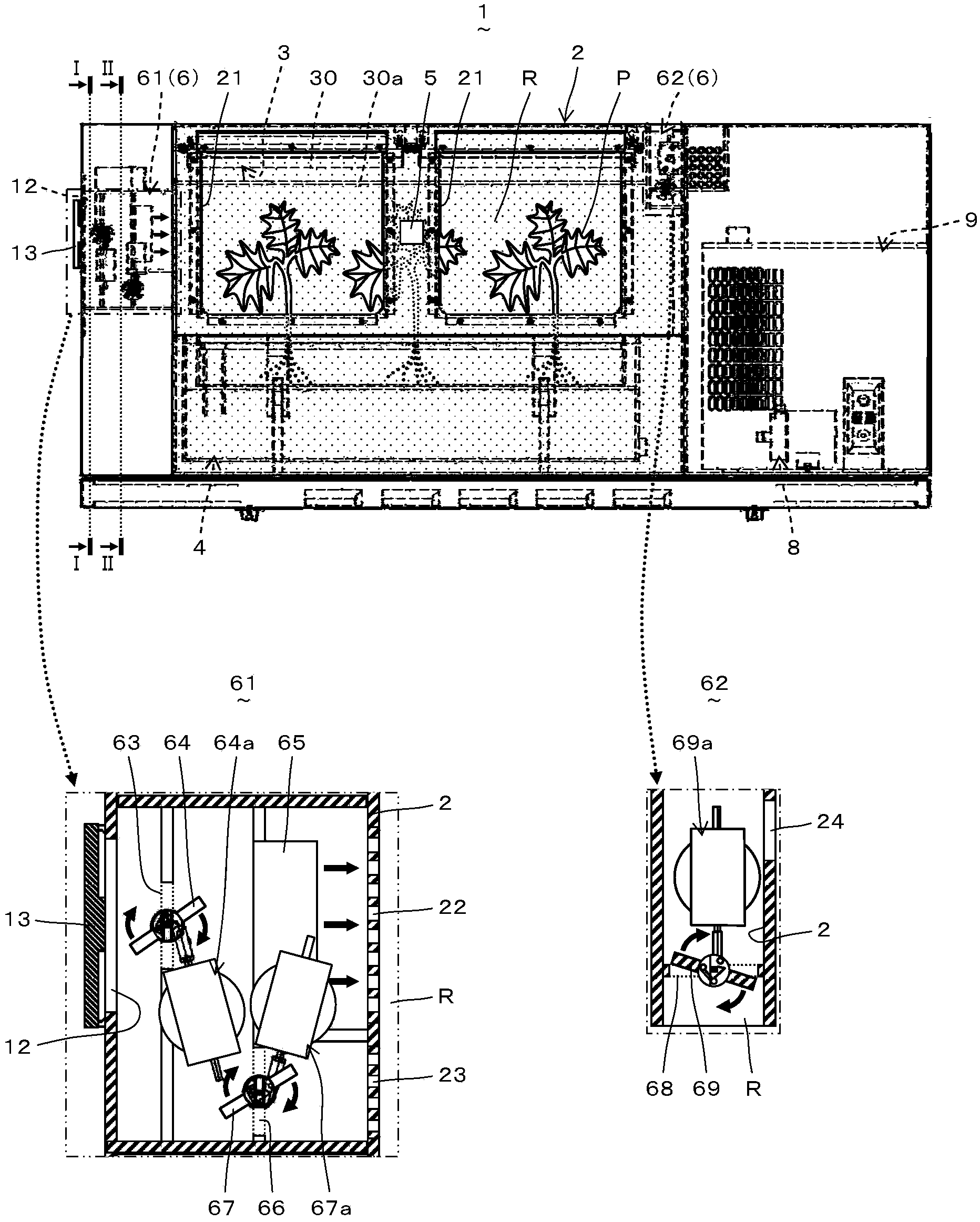
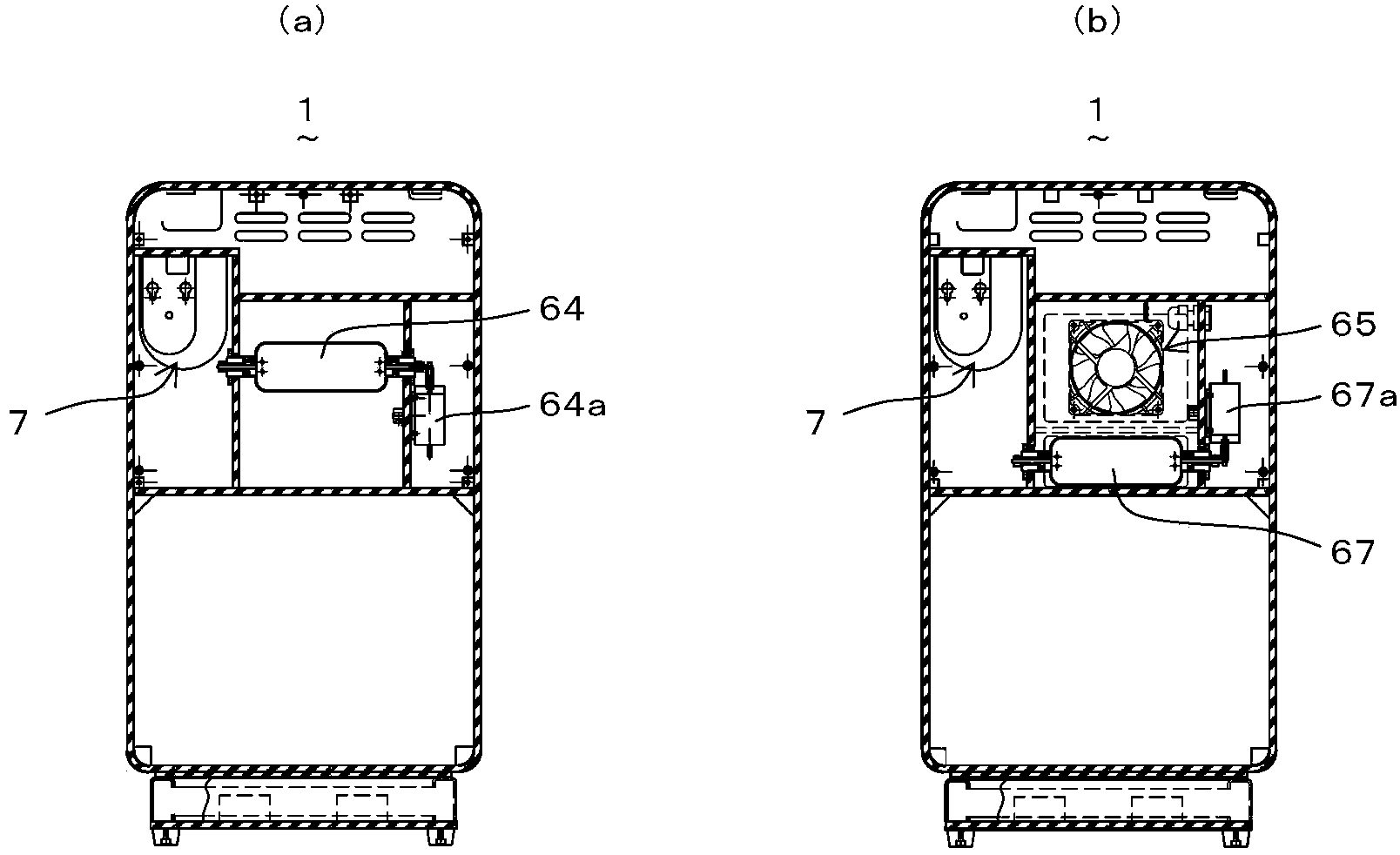
A plant growing device (1) is provided with a ventilation unit (6) comprising an intake unit (61) for taking air into a growing chamber (R) and an exhaust unit (62) for exhausting air from the growing chamber (R). The intake unit (61) has an intake port (63), an intake damper (64) for opening and closing the intake port (63), a blower (65) for sending air in the direction of the growing chamber (R), a passage (66) for guiding air to the upstream side of the blower (65), and a circulation damper (67) for opening and closing the passage (66). The exhaust unit (62) has an exhaust port (68) and an exhaust damper (69) for opening and closing the exhaust port (68). The ventilation unit (6) operates under either an intake and exhaust mode for taking air into and exhausting air from the growing chamber (R) or a circulation mode in which air is caused to circulate in the growing chamber (R). By switching between these modes, CO2 is taken into the growing chamber (R) and gas density, temperature, and humidity within the growing chamber (R) can be made uniform, and as a result, the growing efficiency of a plant (P) can be improved.
Owner:PANASONIC CORP
Method for doubling management of corn haploid
InactiveCN102177846AHigh doubling efficiencyImprove breeding efficiencyPlant genotype modificationCell buddingBud
The invention discloses a method for doubling management of corn haploid. The method for doubling management of corn haploid comprises the following steps: (1) cutting the tip of coleoptile of the corn haploid bud, but not damaging the main leaf and the growing point to obtain the corn haploid bud with coleoptile tip cut; and (2) soaking the corn haploid bud with coleoptile tip cut with a mixed solution of colchicine, dimethyl sulfoxide and water to obtain a chromosome doubled corn bud. The method for doubling management of the corn haploid can remarkably improve the doubling efficiency of the corn haploid, and the tassel pollination rate is over 30 percent which is much higher than the contrast tassel pollination rate (lower than 5 percent). The method solves the vital technical problem of low doubling efficiency in the haploid breeding process, so that the haploid breeding efficiency is greatly improved, and large-scale application of the haploid breeding technology becomes possible.
Owner:CHINA AGRI UNIV
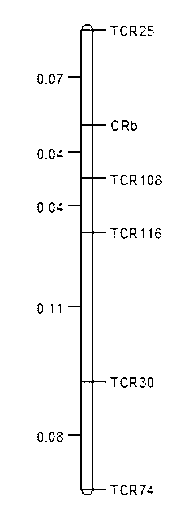
Clubroot-resistant Chinese cabbage gene CRb closely-linked molecular markers, primers and selection method of clubroot-resistant plant
InactiveCN103275972AImprove breeding efficiencyShorten the breeding processMicrobiological testing/measurementDNA/RNA fragmentationBiotechnologyResistant genes
The invention provides 5 clubroot-resistant Chinese cabbage gene CRb closely-linked molecular markers, primers and a selection method of a clubroot CRb-resistant plant. The 5 clubroot-resistant Chinese cabbage gene CRb closely-linked molecular markers comprise TCR25, TCR108, TCR116, TCR30 and TCR74. Distances of the gene CRb respectively to the TCR25, the TCR108, the TCR116, the TCR30 and the TCR74 are 0.07cM, 0.04cM, 0.08cM, 0.19cM and 0.27cM. The 5 clubroot-resistant Chinese cabbage gene CRb closely-linked molecular markers can be used for molecular marker-assistant selection of clubroot-resistant genes of brassica such as Chinese cabbage and Shanghai Bak choi. In assistant selection, the 5 clubroot-resistant Chinese cabbage gene CRb closely-linked molecular markers are used simultaneously so that theoretical selection accuracy is 100%. The 5 clubroot-resistant Chinese cabbage gene CRb closely-linked molecular markers have good repeatability, high reliability and a low detection cost and saves time and labor.
Owner:SHENYANG AGRI UNIV
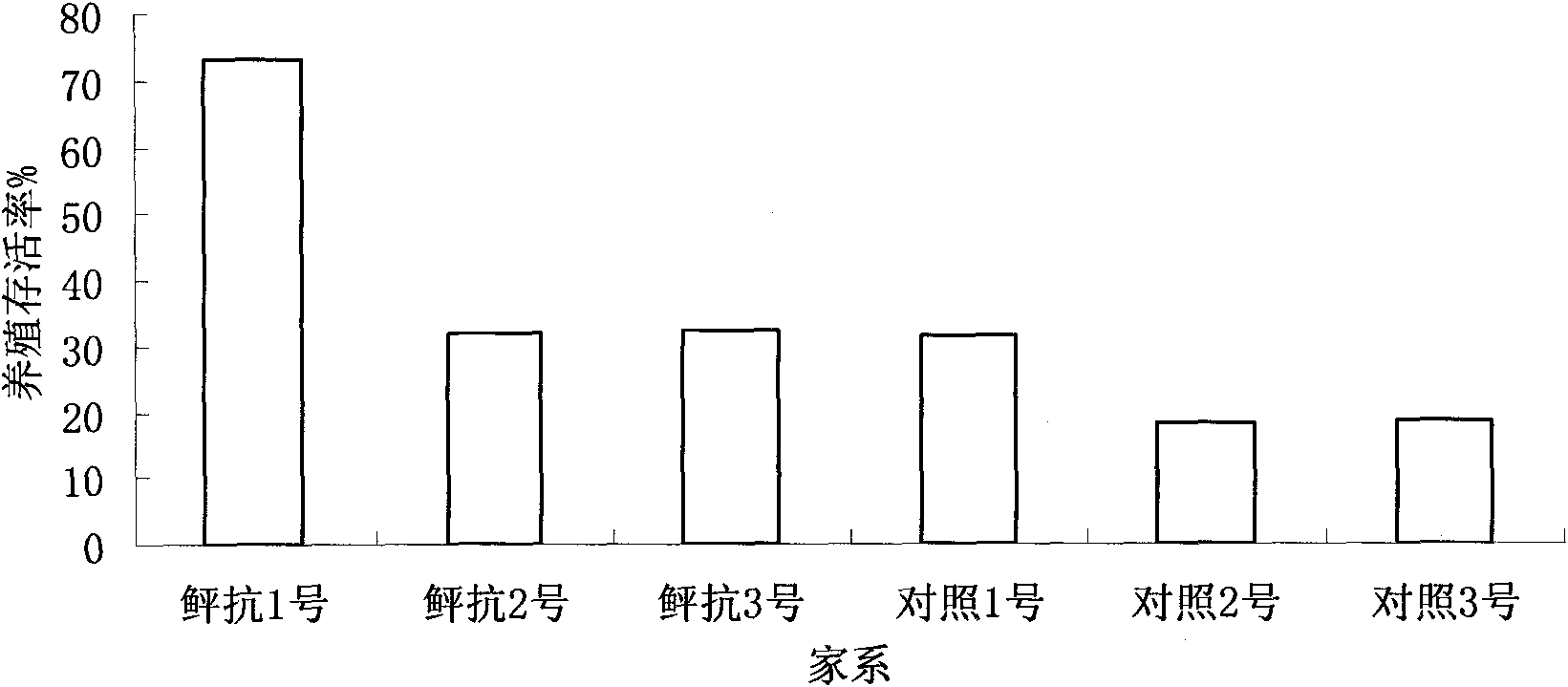
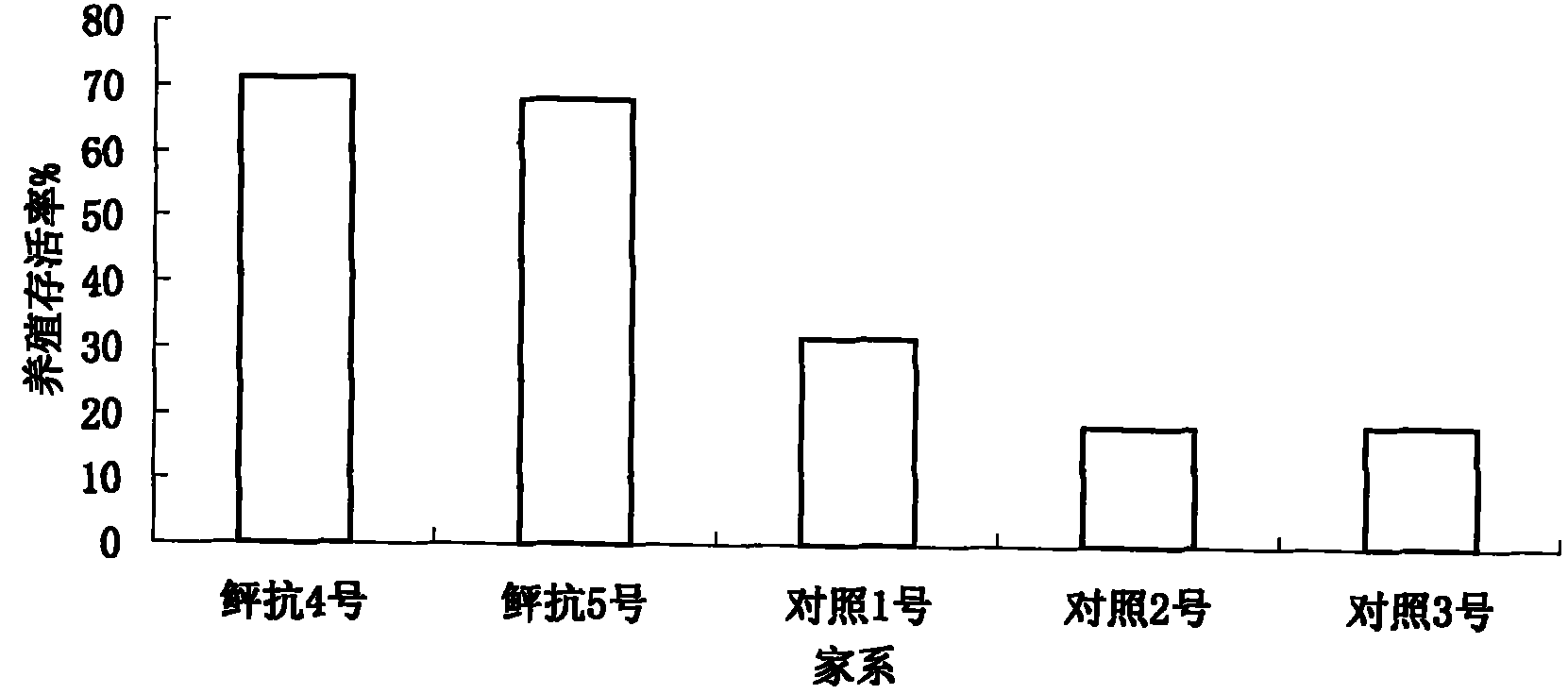
Selective breeding method for disease-resistant superior strains of paralichthys olivaceus
ActiveCN101911918AGood effectIncreased disease powerClimate change adaptationPisciculture and aquariaSocial benefitsMarine fish
The invention relates to a selective breeding method for disease-resistant superior strains of paralichthys olivaceus, which comprises group selective breeding and family selective breeding of the paralichthys olivaceus and is characterized by also comprising molecular marker-assisted selective breeding, manual gynogenesis selective breeding or inbreeding selective breeding of a disease-resistant paralichthys olivaceus family. Multiple kinds of biotechnology such as group selective breeding, intraspecific hybridization, family selective breeding, molecular marker-assisted selective breeding, manual gynogenesis selective breeding, inbreeding and the like is comprehensively adopted; a selective breeding method for the disease-resistant superior strains of marine fishes in China is established for the first time by taking the paralichthys olivaceus as a material; the superior strains of the paralichthys olivaceus of which the vibrio anguillarum resistance is obviously improved are bred; and the breeding survival rate is improved by 20 to 47 percent and the growth rate is improved by over 20 percent. The method opens up a new technical approach for the selective breeding of the disease-resistant strains of the fishes, is suitable for popularization and application to all cultured fishes, has great significance and application value for breeding the disease-resistant superior strains of the cultured fishes, and can generate great economic and social benefits.
Owner:YELLOW SEA FISHERIES RES INST CHINESE ACAD OF FISHERIES SCI
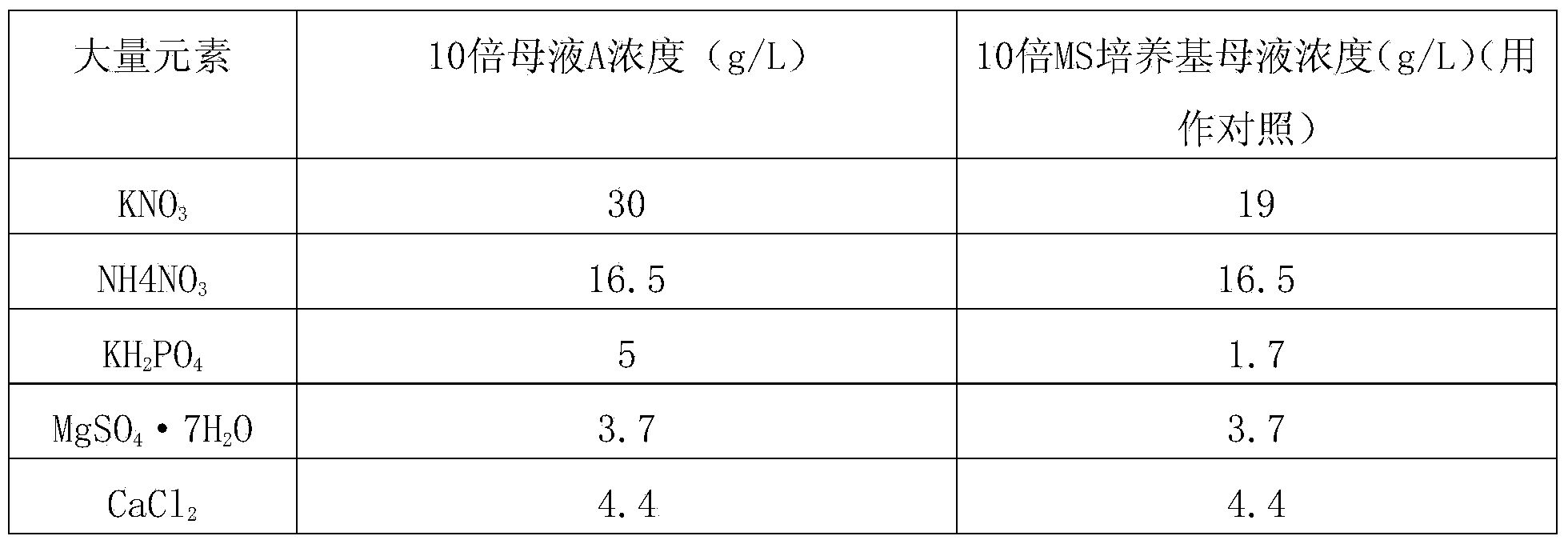
Method for improving culturing efficiency of japonica rice anther
ActiveCN103461141AImprove seedling rateImprove breeding efficiencyPlant tissue cultureHorticulture methodsAgricultural scienceJaponica rice
The invention discloses a method for improving the culturing efficiency of japonica rice anther. The method comprises the steps that firstly, culture medium mother solutions of the japonica rice anther are prepared according to formulas of the culture medium mother solutions of the japonica rice anther, then the various mother solutions are utilized to prepare an induction culture medium, a differentiation culture medium and a rooting culture medium all of which are used in different periods, and the culture media in the different periods are correspondingly applied to induction, green plantlet differentiation and rooting of calluses of the japonica rice anther respectively. By the adoption of the method, the percentage of seeding emergency of culturing of the japonica rice anther can be improved, the rice breeding progress is quickened, and the seed selection efficiency of good variety of rice is improved.
Owner:赫聪
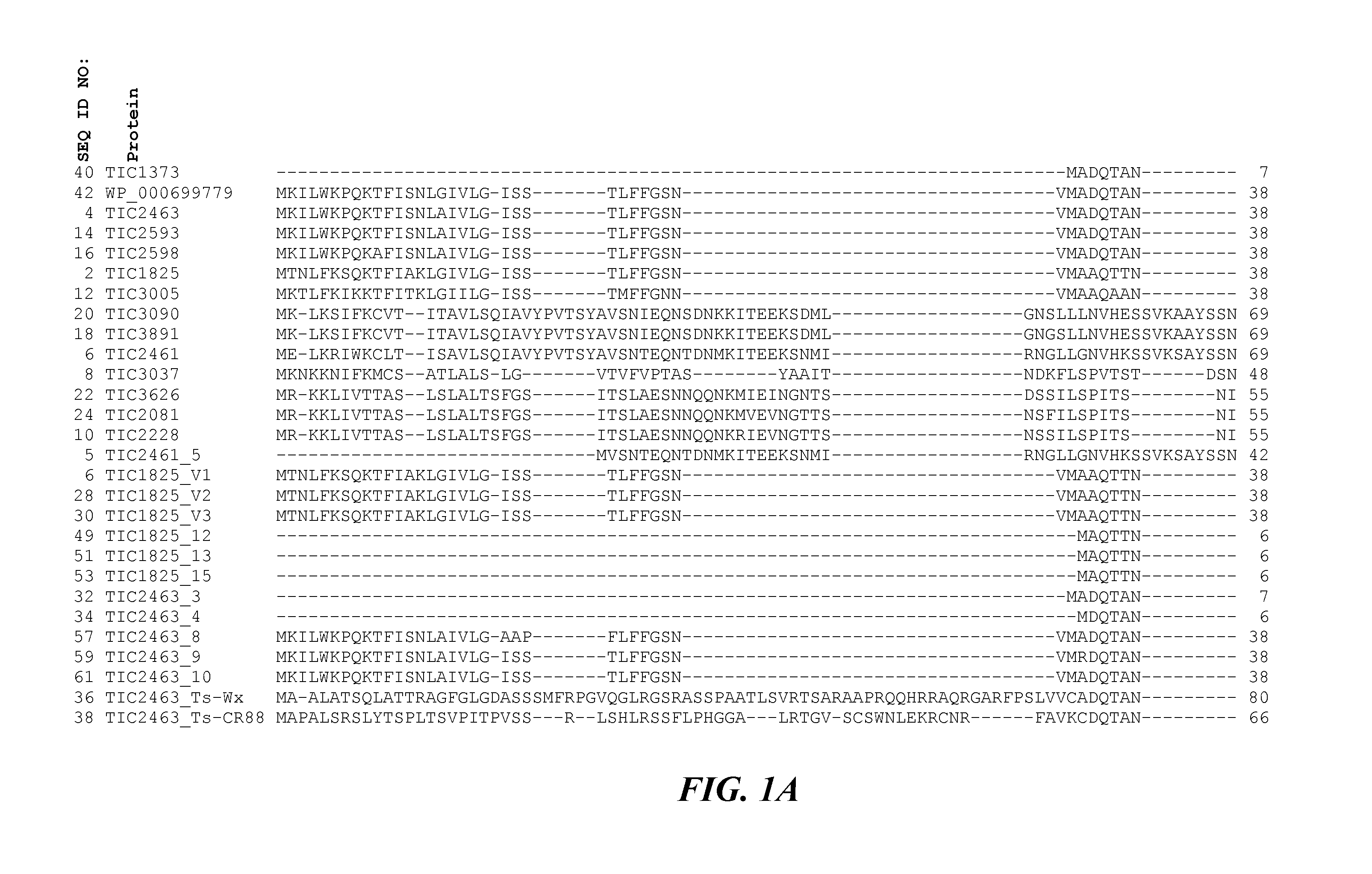
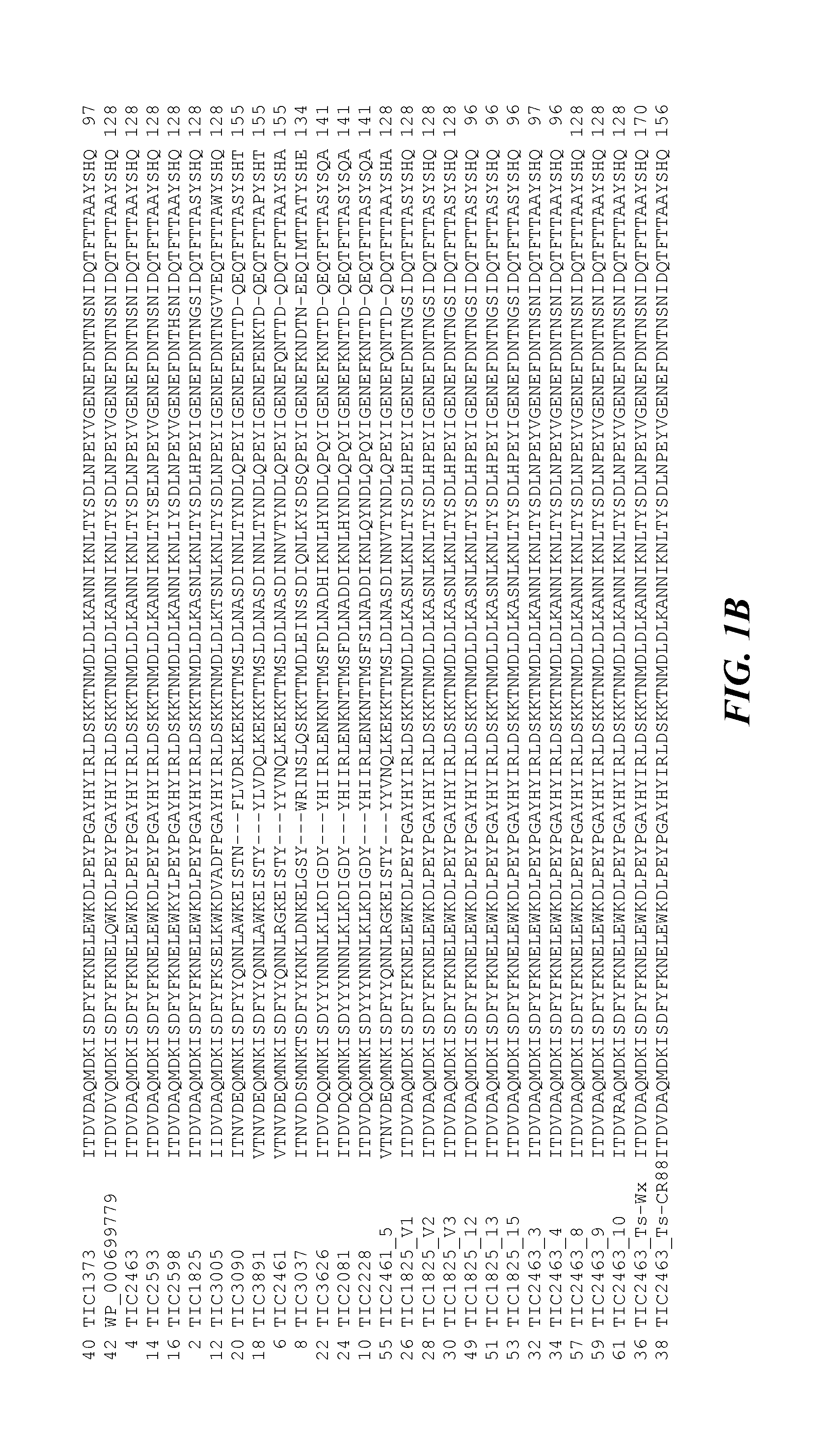
Pesticidal Toxin Proteins Active Against Coleopteran Insects
ActiveUS20150274786A1Low costImprove breeding efficiencyBiocidePeptide/protein ingredientsNucleotideInsect pest
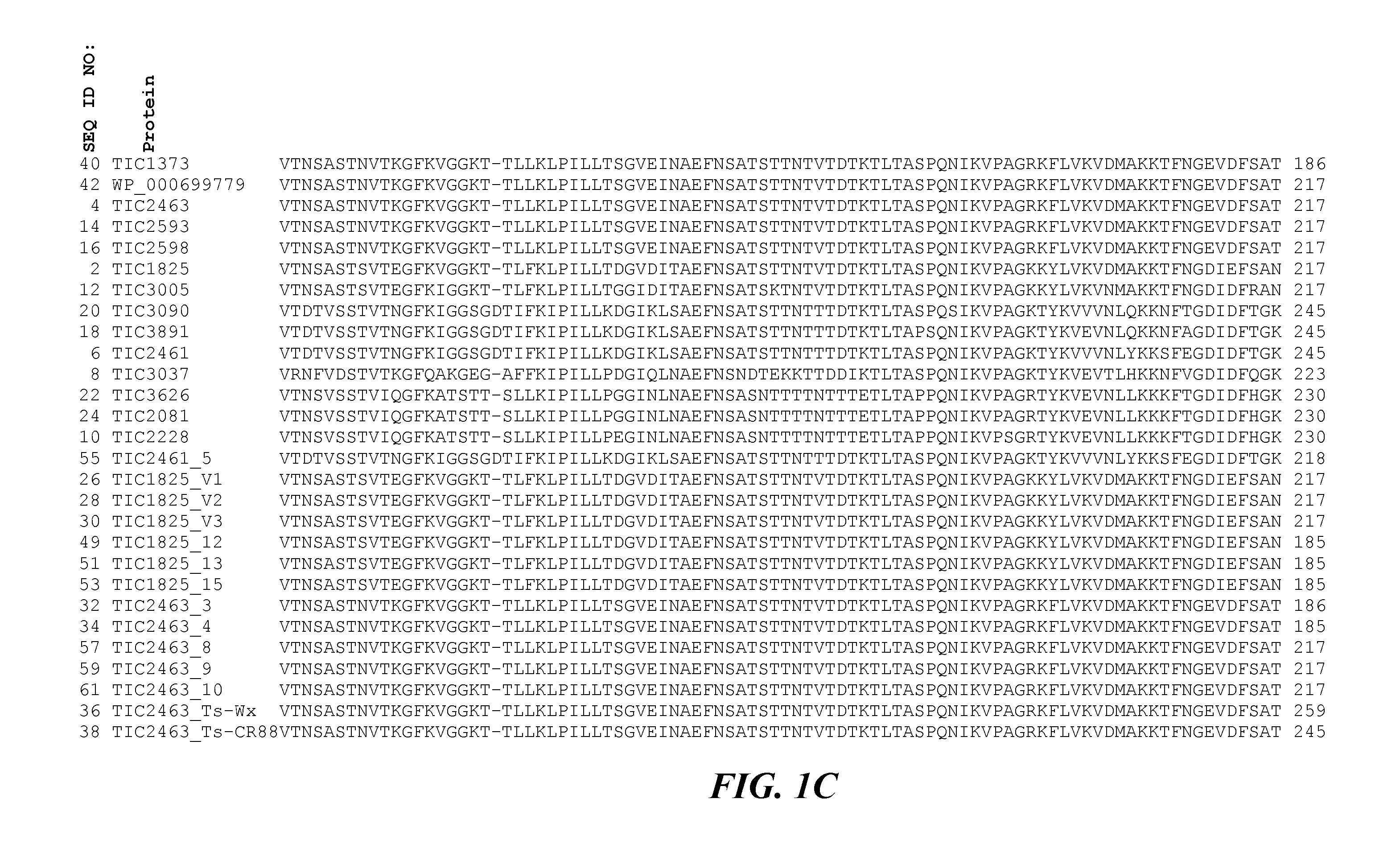
The invention generally relates to the field of insect inhibitory toxin proteins. A novel class of proteins exhibiting insect inhibitory activity against agriculturally relevant pests of crop plants and seeds are disclosed. Insecticidal activity is particularly effective against the Coleopteran order of insect pests. Plants, plant parts, and seed are provided containing a polynucleotide construct encoding one or more of the toxin proteins disclosed herein. The proteins are referred to herein variously as the TIC2463-related toxin protein class or family, the TIC2463-related toxin proteins, the TIC2463-related protein genus, toxin proteins related to the TIC2463 toxin protein, proteins related to TIC2463, TIC2463-related toxin polypeptides, TIC2463-related pesticidal proteins, and the like.
Owner:MONSANTO TECH LLC
Rice dwarf-related protein and coding gene and application thereof
ActiveCN101607989AShorten the breeding periodImprove breeding efficiencyFungiBacteriaWAS PROTEINAgricultural science
The invention discloses a rice dwarf-related protein and a coding gene and application thereof. The protein is protein (a) or (b): protein (a) is formed by an amino acid sequence shown as a sequence 3 of a sequence list; and protein (b) is formed by substituting and / or lacking and / or adding one or more amino acids in the amino acid sequence shown as the sequence 3 of the sequence list, related to rice dwarf and derived from the protein (a). The invention also provides the gene for coding the protein and a recombinant expression vector containing the gene, a transgenic cell system or recombinant bacteria. The gene is utilized to breed rice with reduced strain height. The rice dwarf-related protein and the coding gene have extremely important values of shortening the breeding period, improving the breeding efficiency and enriching rice dwarf sources.
Owner:INST OF GENETICS & DEVELOPMENTAL BIOLOGY CHINESE ACAD OF SCI
A hybrid rice breeding method
InactiveCN1951172ANo sterile cytoplasmic monotonyNo side effectsPlant genotype modificationRice cultivationHeterosisGene
The invention relates to a method for cultivating cross rice. Wherein, it comprises that (1), selecting the light-temperature sensitive sterile system with low-temperature sterile or short-light low-temperature sterile; (2), selecting the sterile masculinity as gene system; (3), crossing sterile system S with sterile masculinity system S', to obtain stable sterile masculinity S / S' F1; (4), screening and recovering system R; (5), using S / S' F1 as mother type to be crossed with recover system R, to find the needed cross combination, to produce cross rice. The invention can improve the match property of sterile system and reduce produce cost.
Owner:NAT ENG & TECHNOLOGICAL RES CENT OF INTERBREED PADDY +1
Breed conservation, screening and breed production method in tilapia complete set lines as well as between two lines of complete sets
ActiveCN102630617ALower conditionsEasy to operateClimate change adaptationPisciculture and aquariaTilapiaBase population
The invention relates to a breed conservation, screening and breed production method in tilapia complete set lines, comprising the following steps of: construction of complete set line base populations rich in hereditary variation, establishment of operation procedures of family production and management in the complete set lines, marking of individuals in the complete set lines, genetic evaluation analysis of important production traits and establishment of individual mating selection programs in the complete set lines. The invention also provides a hybrid breed production method between twolines of complete sets, which is established by utilizing the method. The method disclosed by the invention has the characteristics of high pertinence, high selection breeding efficiency, capability of effectively preventing parent inbreeding depression of the complete set lines and the like; the harvest weight of the hybrid fine breed of tilapia is averagely increased by more than 30% than that of control groups; and the culture survival rate is also obviously higher than that of the control groups. According to the invention, a new technical approach is developed for cultivation of fish fine breeds; particularly a new selection breeding method is applied in the breed conservation, the screening and the breed production of the complete set lines in fish hybrid breeding; and the method isalso suitable for being popularized and applied in all cultured fishes and has broad popularization and application prospects.
Owner:YELLOW SEA FISHERIES RES INST CHINESE ACAD OF FISHERIES SCI
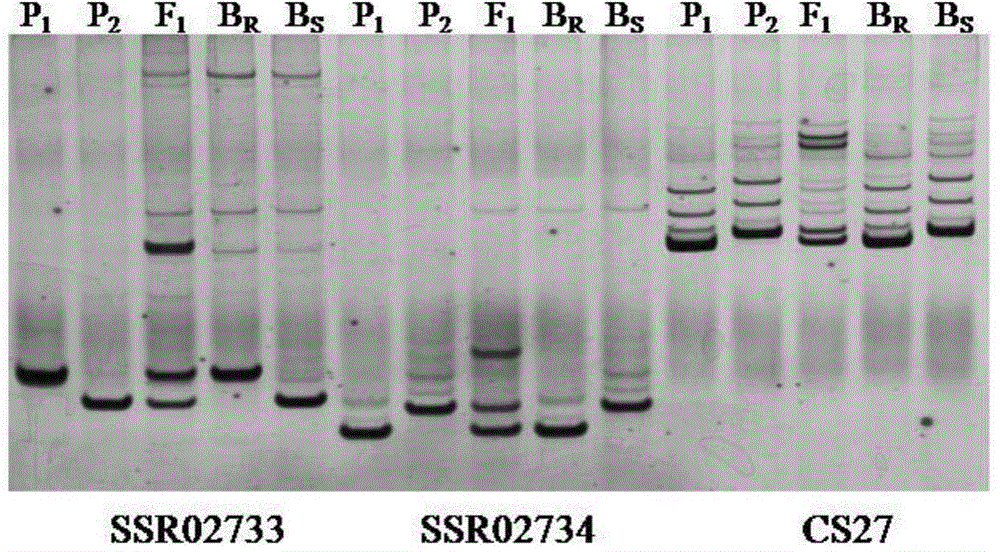
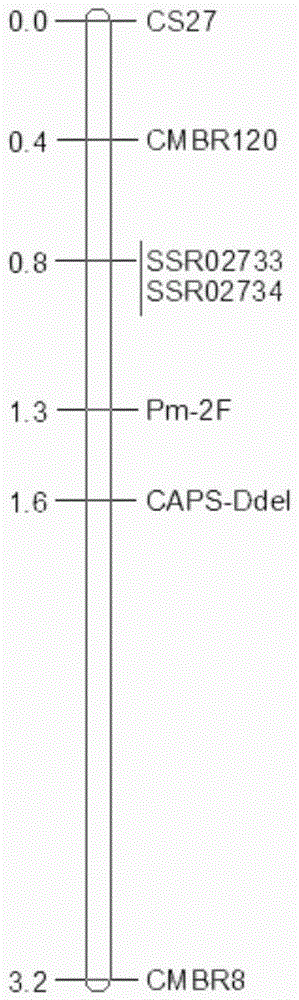
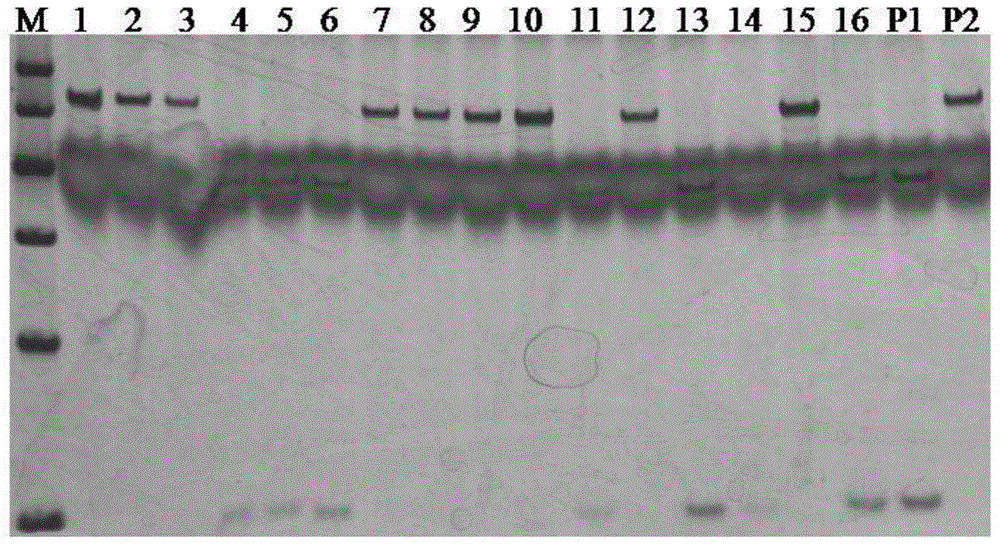
Series molecular markers closely linked with muskmelon powdery mildew resistant gene Pm-2Fand acquisition method of series molecular markers
InactiveCN104630215AShorten the breeding cycleImprove breeding efficiencyDNA preparationDNA/RNA fragmentationGenetic engineeringCancer research
The invention belongs to the field of genetic engineering and provides series molecular markers closely linked with a muskmelon powdery mildew resistant gene Pm-2F and an acquisition method of the series molecular markers. Each molecular marker contains at least one nucleotide base sequences shown in SEQ No:1-8 in a sequence table. The acquisition method of the series molecular markers comprises the following steps: selecting a test material, primarily positioning the muskmelon powdery mildew resistant gene Pm-2F, acquiring a site SNP / Indel linked with target characters, and acquiring molecular markers closely linked with the muskmelon powdery mildew resistant gene Pm-2F, so that the molecular markers closely linked with the muskmelon powdery mildew resistant gene Pm-2F are obtained. When the molecular markers are used for early screening and identification of a powdery mildew resistant variety, the breeding period can be greatly shortened, and the breeding efficiency is improved, so that the series molecular markers closely linked with the muskmelon powdery mildew resistant gene Pm-2F have important theory and practice significance.
Owner:BEIJING ACADEMY OF AGRICULTURE & FORESTRY SCIENCES +1
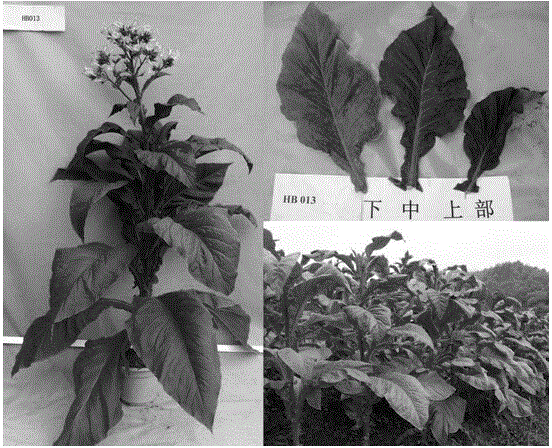
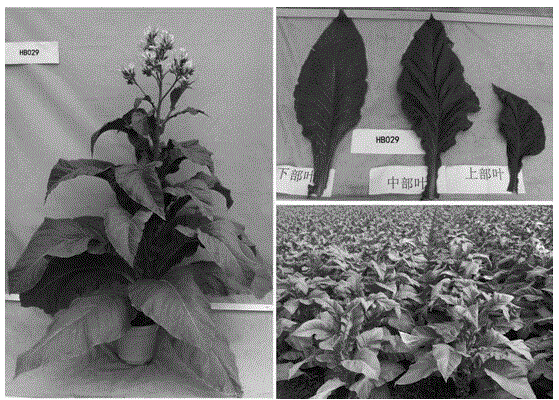
Molecular label from high-quality variety Xinluzao No.24 and relevant with cotton fiber strength
ActiveCN102181442AImprove breeding efficiencyLarge additive effectMicrobiological testing/measurementDNA/RNA fragmentationPopulationCotton fibre
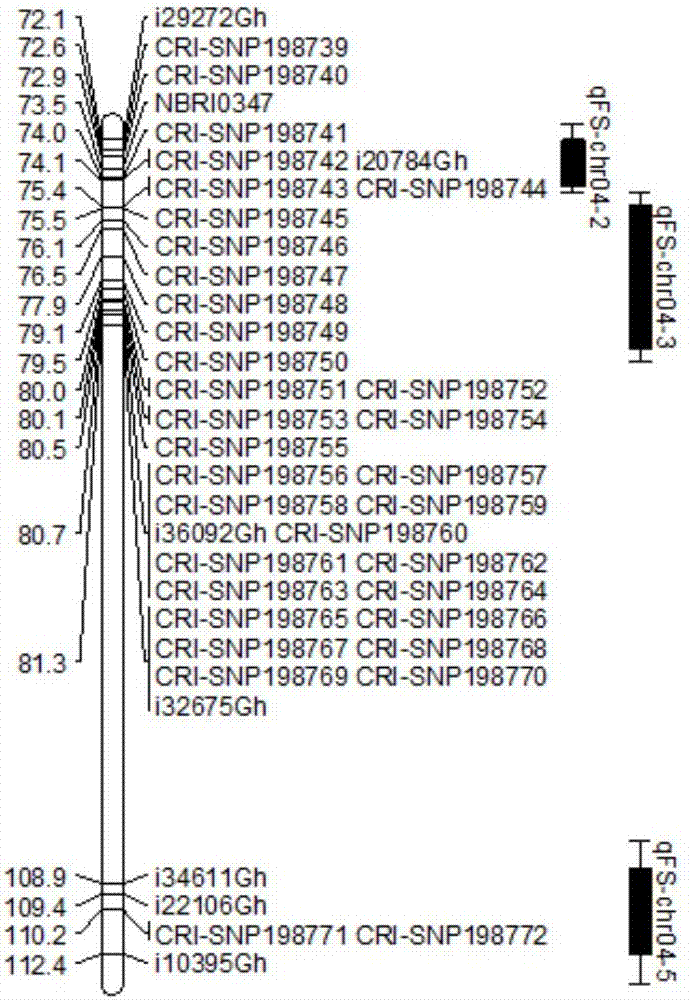
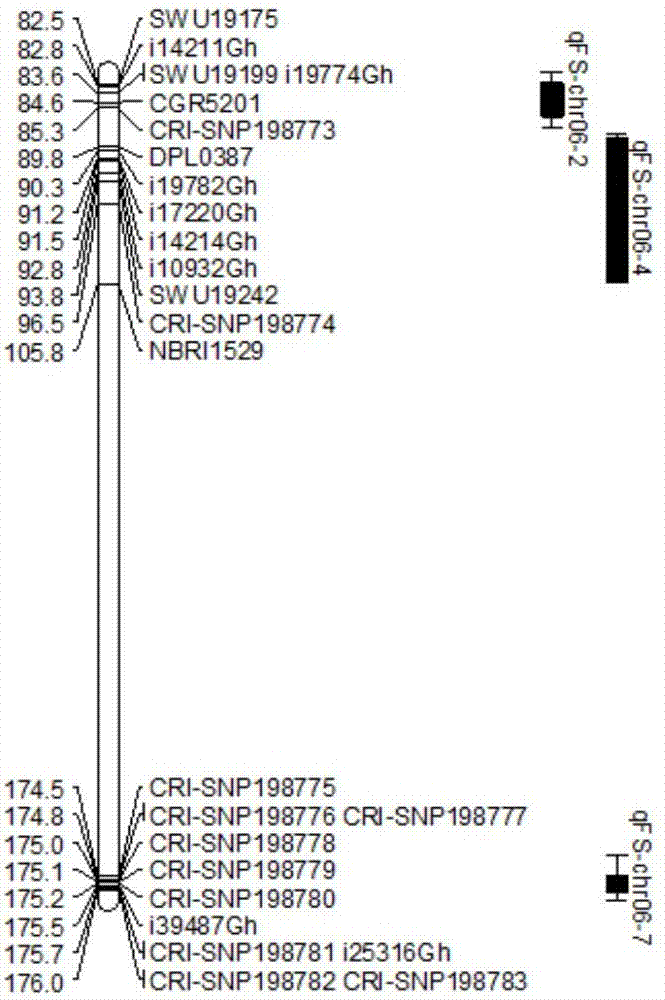
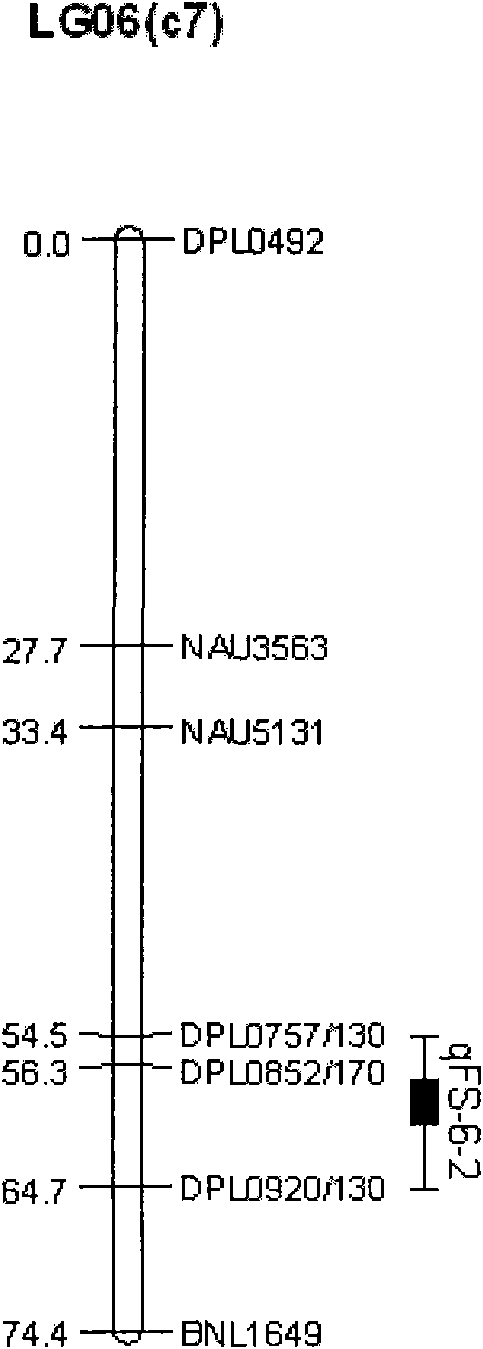
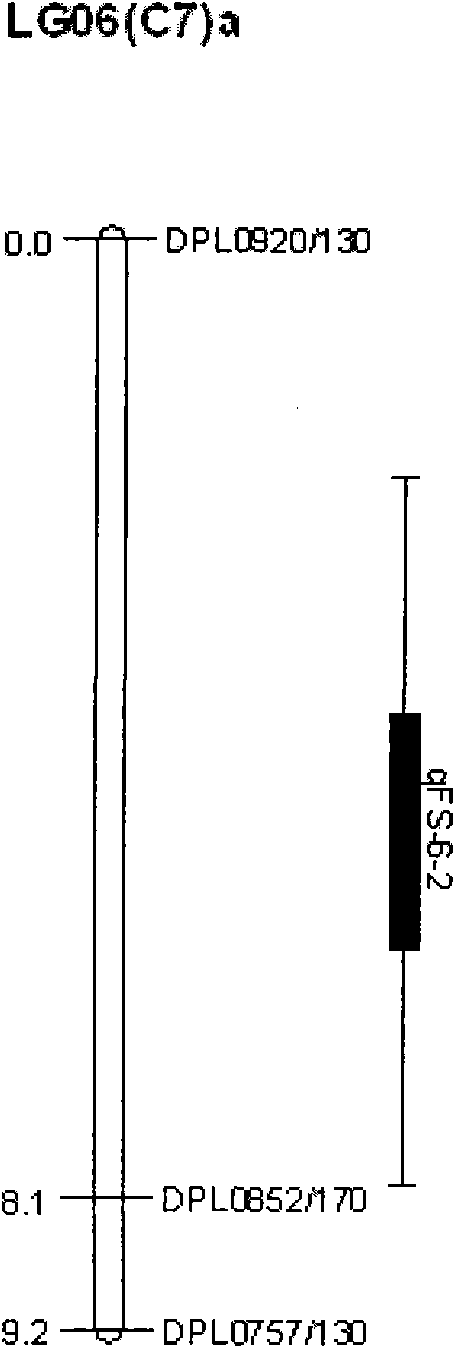
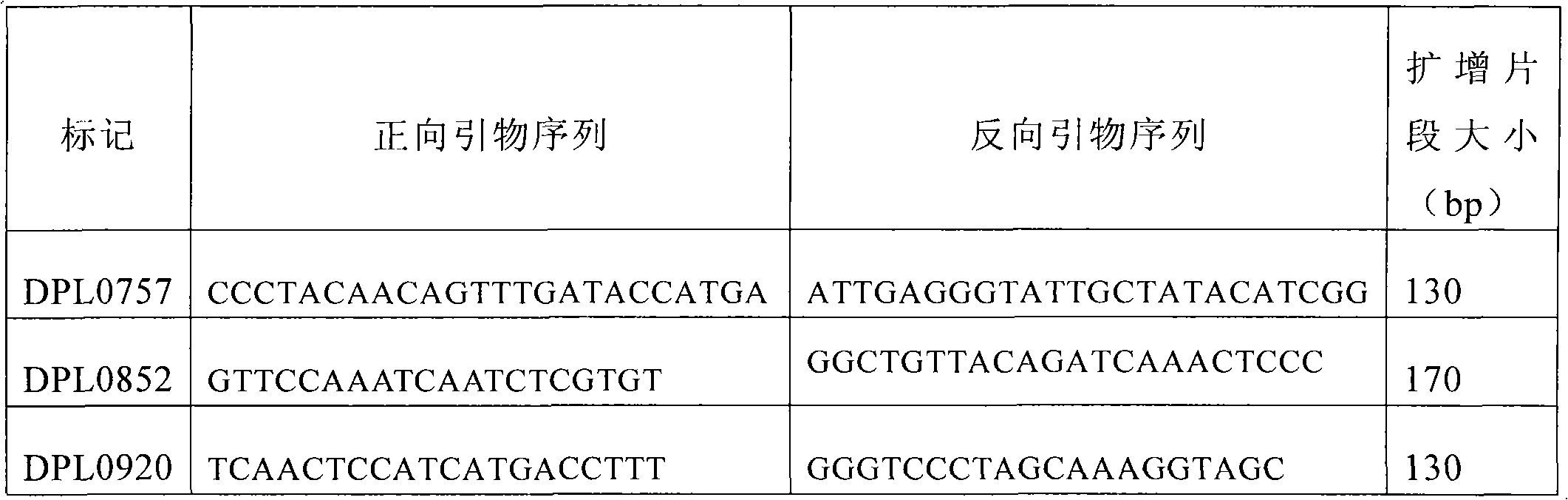
The invention discloses a molecular label linked with a major gene of a high-strength cotton fiber, which is obtained by the following method: taking the Xinluzao No.24 as a male parent, taking Lunianyan No.28 and Ji cotton 516 as female parents; respectively constructing two segregation populations of F2 and F2:3; carrying out polymorphism screening between parents (Lunianyan 28 and Xinluzao 24)by utilizing an SSR (simple sequence repeat) primer; constructing a genetic linkage map by utilizing F2; carrying out two generation quantitative trait locus (QTL); selecting an SSR label linked withthe high-strength cotton fiber to analyze the DNA polymorphism of the parent Xinluzao 24 and the Ji cotton 516; and constructing the genetic linkage map and positioning QTL; the fiber strength major QTL (qFS-62) can be stably detected in two populations and generations, and is positioned between the label interval of DPL0757-DPL0920 of chromosome c7; a synergtic gene is from the parent Xinluzao No.24; and the linking label of qFS-6-2 is DPL0757130, DPL0852170 and DPL0920130. By utilizing the label provided by the invention, molecule label assisted selection is carried out, thus improving the selection efficiency of the high-strength cotton fibre.
Owner:INST OF COTTON RES CHINESE ACAD OF AGRI SCI
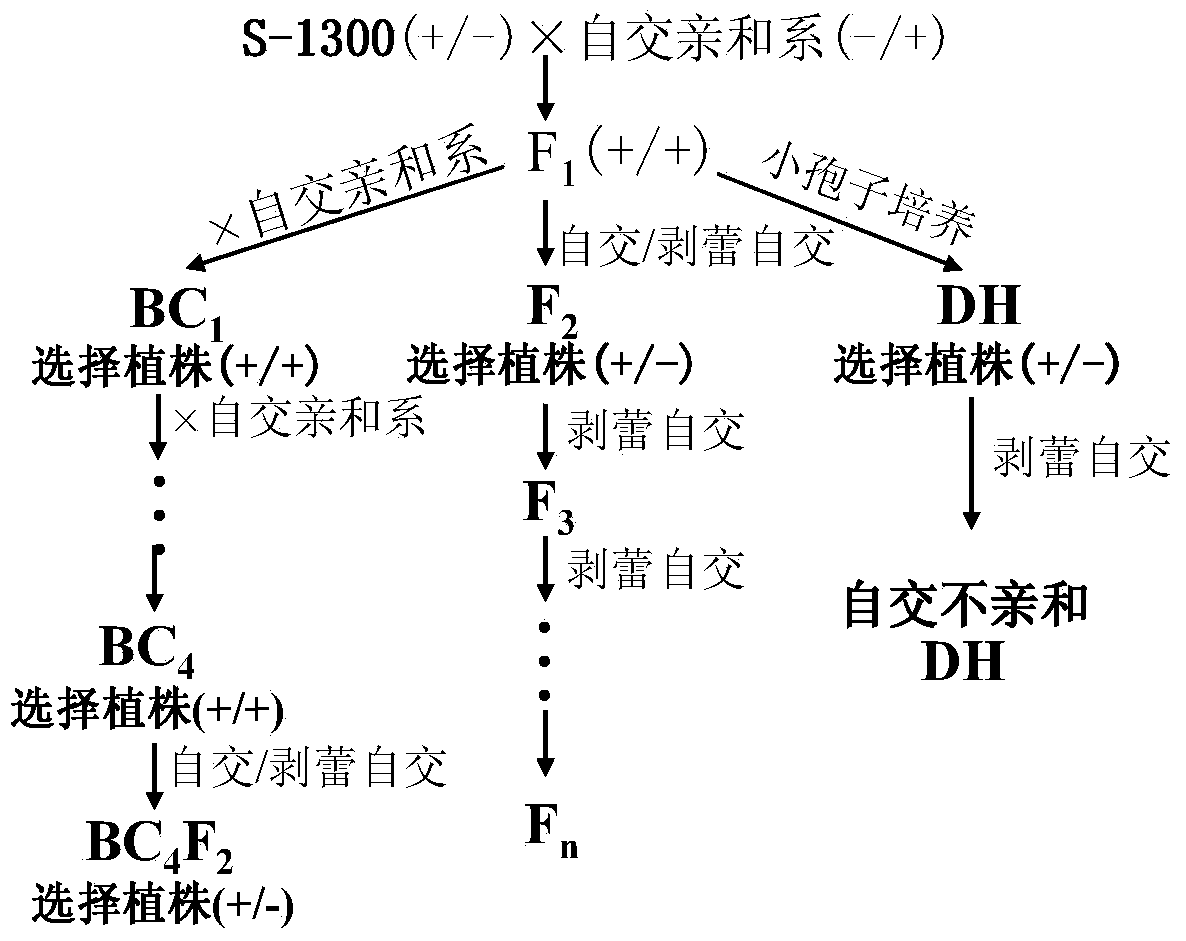
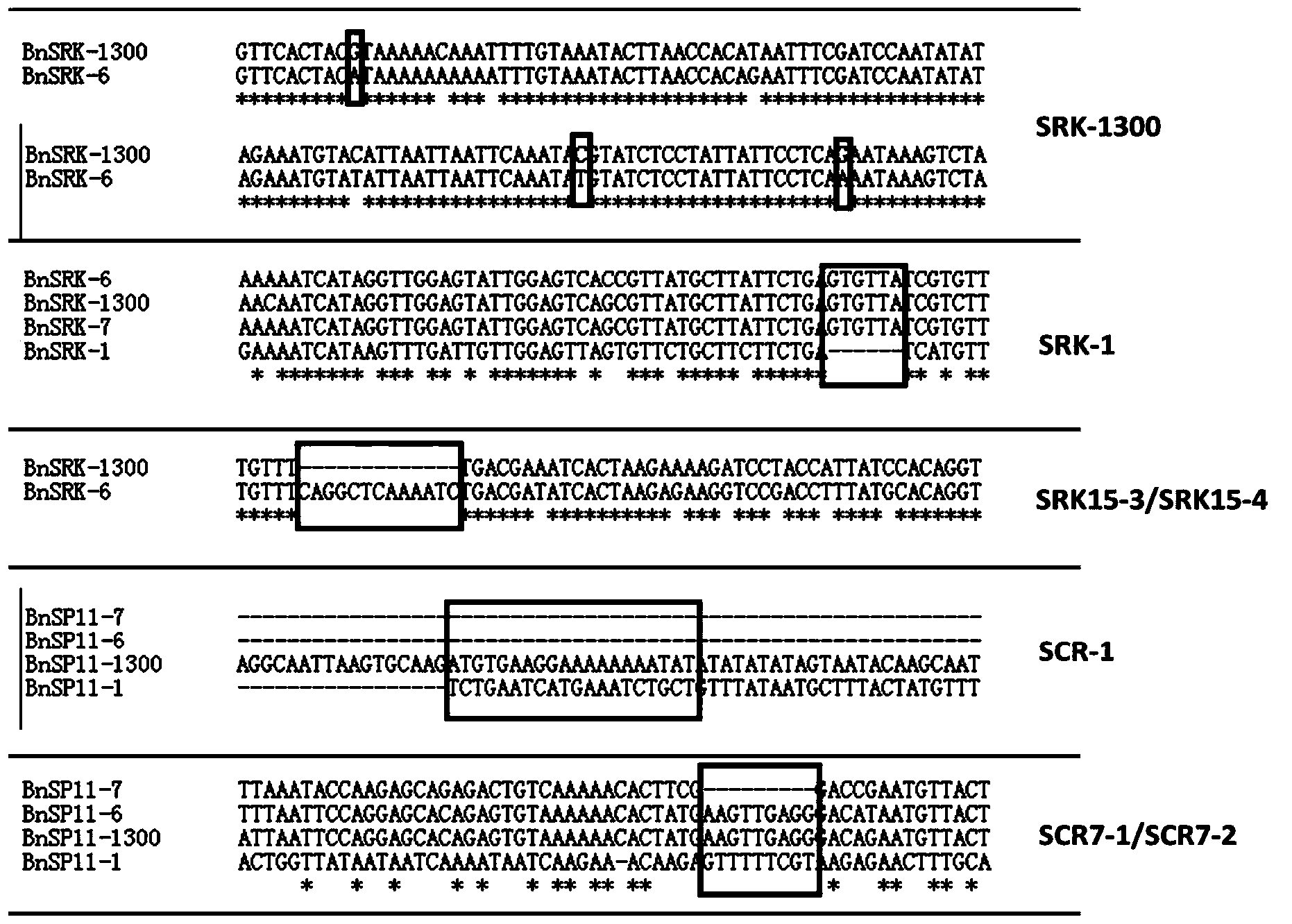
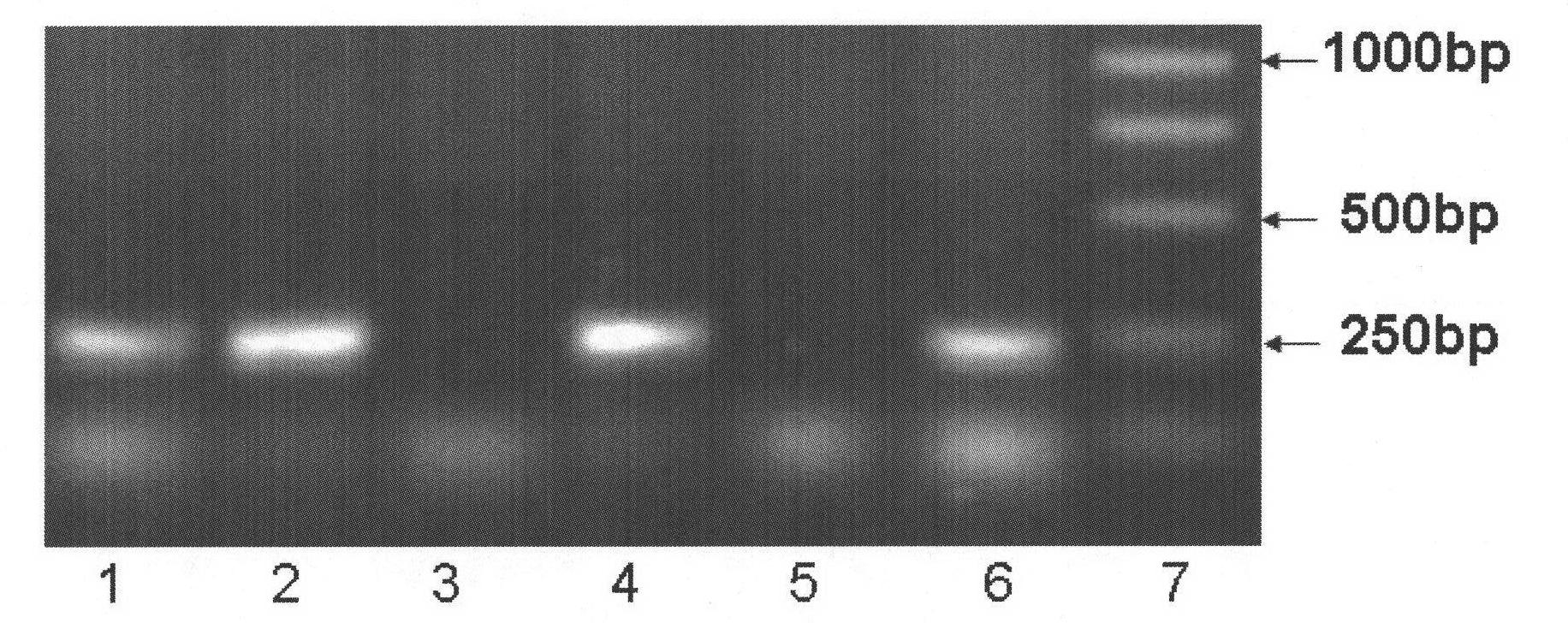
Molecular detection method of Brassica napus self-incompatible S-locus haplotype
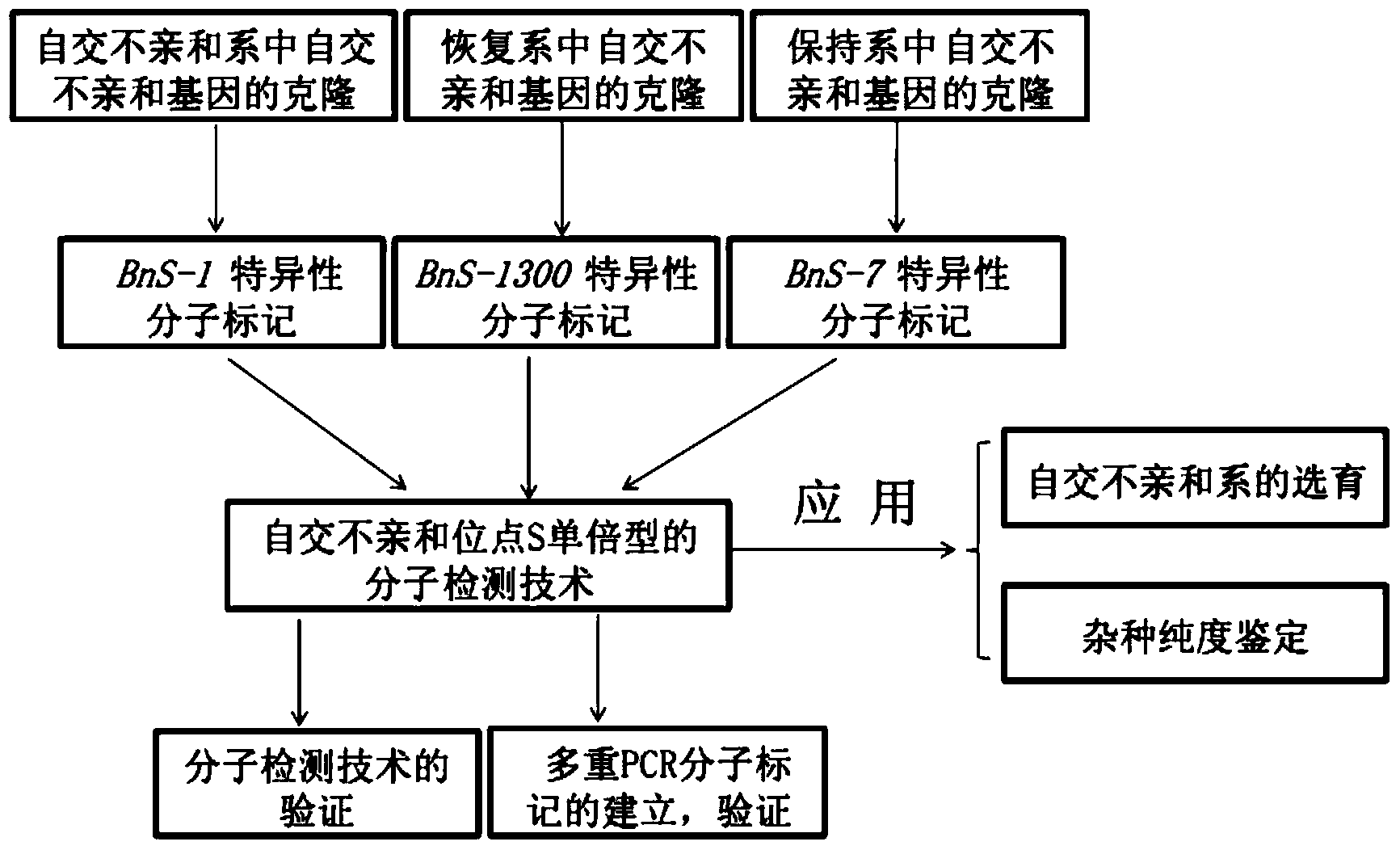
InactiveCN103451283AEasy to useSolve the problem of low coordinationMicrobiological testing/measurementDNA/RNA fragmentationBrassicaDuplex pcr
The invention belongs to the field of molecular breeding of rapeseeds, and relates to a molecular detection method of Brassica napus self-incompatible S-locus haplotype, a molecular marker and application. An SCAR (sequence characterized amplified regions) molecular marker system used for identifying the Brassica napus self-incompatible S-locus haplotype is prepared and includes four SCAR molecular markers, namely, SRK1-1, SCR1-1, SRK-1300 and SCR7-2 which can be combined to obtain two pairs of double PCR (polymerase chain reaction) markers as SRK1-1 and SRK-1300 as well as SCR7-2 and SRK-1300; the PCR marker combination can be used in a combination manner or independent manner; the DNA (deoxyribonucleic acid) sequences of the PCR marker combination are shown as SEQ ID NO: 6, SEQ ID NO:7, SEQ ID NO: 8 and SEQ ID NO: 9. According to the invention, a practical marker and an utilization method are provided for the molecular breeding of the rapeseeds.
Owner:HUAZHONG AGRI UNIV
Method for high performance collective breeding non-nuclear grape with disease resistant capability
InactiveCN1415189AImprove breeding efficiencyRapid cultivationPlant phenotype modificationPlant tissue cultureDiseaseHigh resistance
An efficient aggregate method for breeding seedless grap is disclosed, which features that the conventional hybridizing technique, the embryo retriever technique and molecular marker breeding technique are integrated. Its advantages are short breeding period shortened from 7-9 years to 3-4 months, high resistance to disease, high seedless percentage, and high quality of grape.
Owner:NORTHWEST A & F UNIV
Breeding method for multi-resistance flue-cured tobacco hybrid
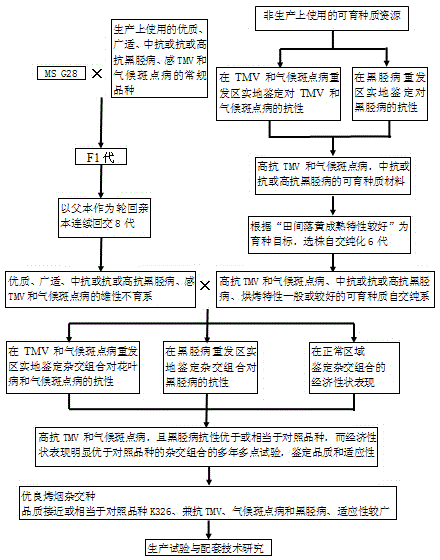
InactiveCN104885930ASolve quality problemsAddressing Adaptive IssuesPlant genotype modificationDiseasePlant disease
The invention provides a breeding method for a multi-resistance flue-cured tobacco hybrid. The breeding method for the multi-resistance flue-cured tobacco hybrid comprises the steps of firstly using MSG28 as a female parent, using high-quality, widely applicable and blackleg-resistant varieties which are sensitive to TMV and weather fleck and are used for production as recurrent parents, performing eight generations of backcrossing to breed male sterile lines, using fertile materials which are resistant to blackleg and highly resistant to TMV and weather fleck as original resources, performing six generations of selfing purification to breed selfing pure lines with common or better curing characteristics, then using the bred male sterile lines as female parents and the selfing pure lines as male parents to breed hybrid combinations, respectively evaluating resistance to different diseases and economic performance by using land parcels or areas seriously attacked by different diseases and normal land parcels in producing areas, and finally performing 2-3 years of multi-point tests for quality and adaptability evaluation to the hybrid combinations which are simultaneously resistant to different diseases and have good economic performance. By adopting the method, the aggregation of multiple characters such as high quality, simultaneous resistance to foliage diseases and root and stalk diseases and good adaptability can be realized, the breeding efficiency is improved.
Owner:HUBEI TOBACCO SCI RES INST
SSCP marker closely linked with major wheat scab resistance QTL and application thereof
InactiveCN101892307AReduce the waste of manpower and material resourcesImprove breeding efficiencyMicrobiological testing/measurementMicroorganism based processesSingle-strand conformation polymorphismSequence analysis
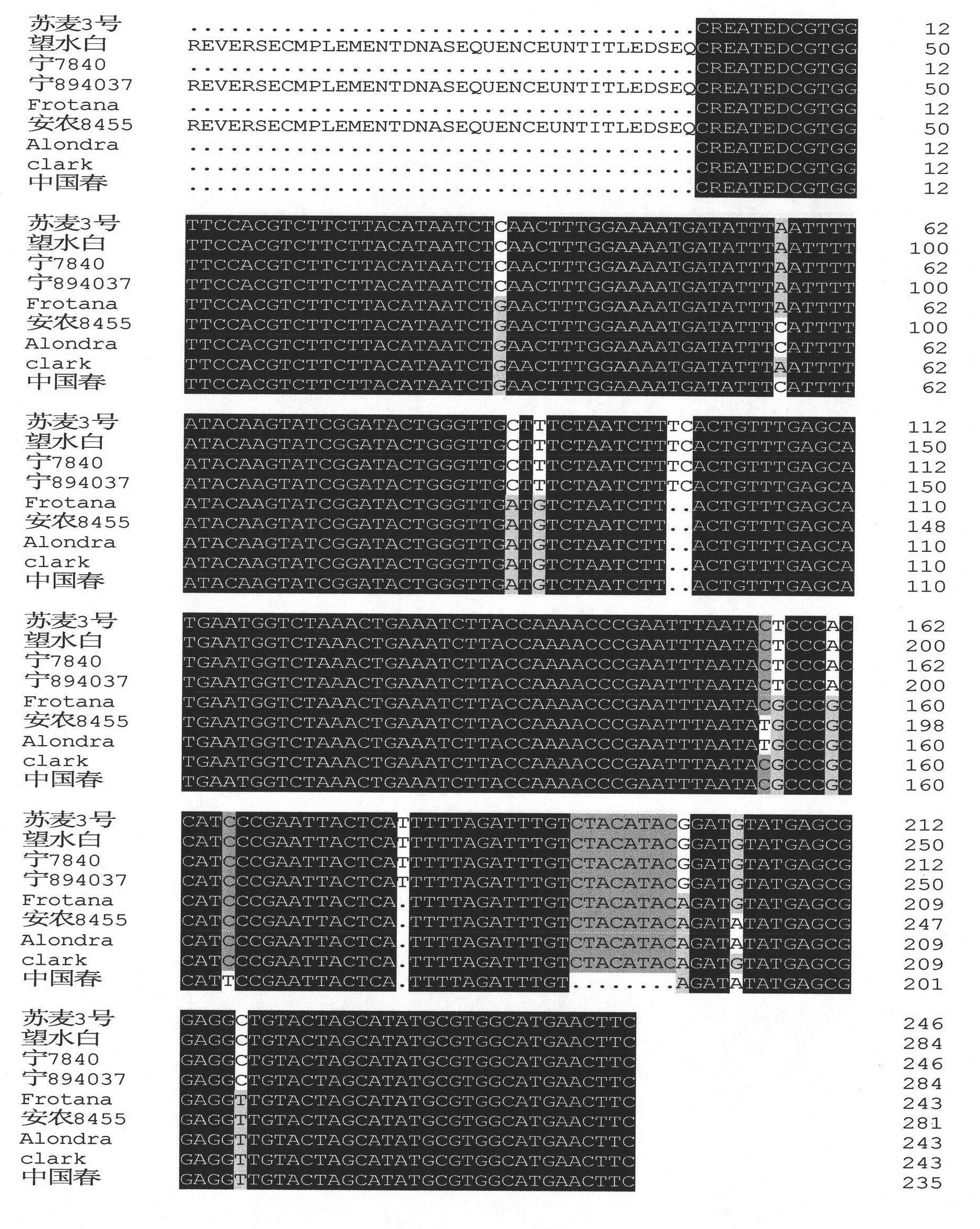
The invention relates to a single strand conformation polymorphism (SSCP) marker closely linked with a major wheat scab resistance quantitative trait locus (QTL). The SSCP marker is characterized in that: scab resistance candidate genes of a scab resistance QTL in a 3BS area of a wheat variety or strain resisting scab is PCR amplified by using a primer; after denaturalization, a PCR-amplified product has different single strand conformation; and the SSCP marker closely linked with the major wheat scab resistance QTL is established for detecting a genotype of the wheat variety or strain during breeding. The SSCP marker has the advantages of: overcoming the disadvantages that the wheat scab resistance screening can only be authenticated in a flowering period and is easily influenced by the environment in conventional breeding, predicting and screening wheat plants with the scab resistance by detecting a molecular marker at a seedling stage, eliminating disease plants, reducing waste of labor and materials and improving the breeding efficiency. Compared with an ABI DNA sequence analysis meter-based marker, the SSCP marker closely linked with the major wheat scab resistance QTL has the advantages of simple and convenient operation, low cost and same sensitivity.
Owner:JIANGSU ACADEMY OF AGRICULTURAL SCIENCES
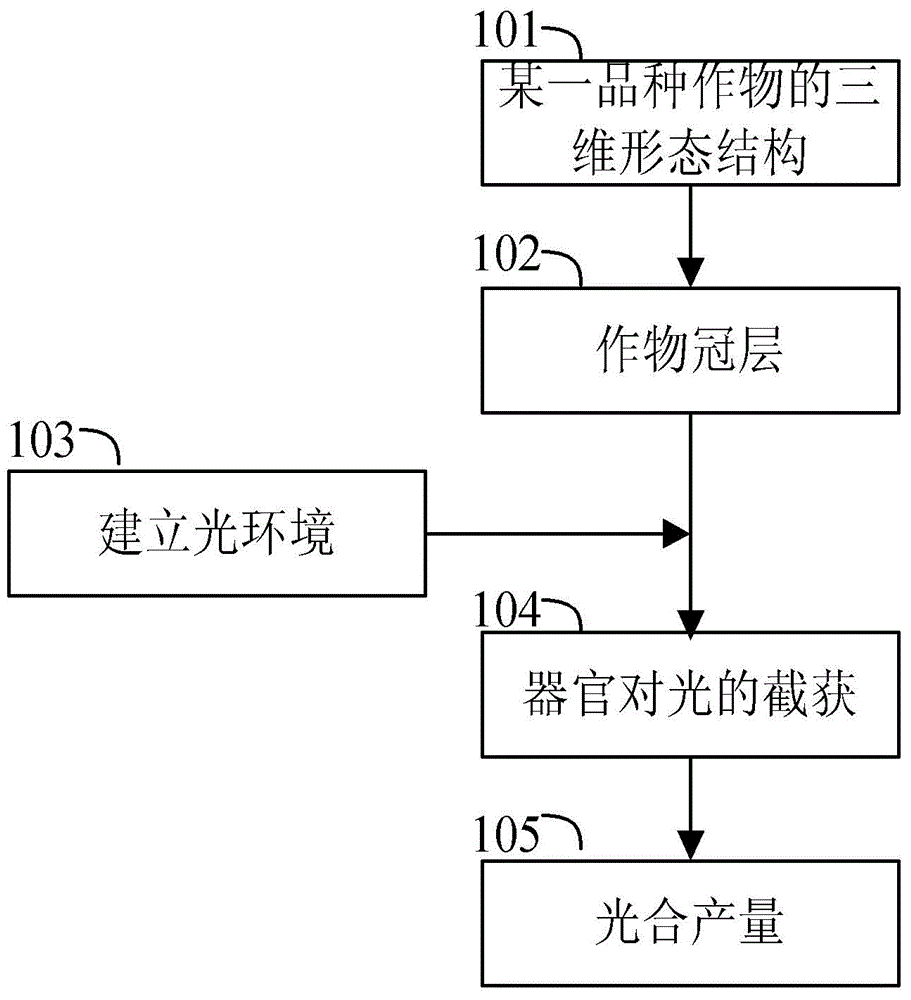
Variety analyzing method based on canopy light distributed computing
InactiveCN104615867AReduce labor costsReduce experiment costSpecial data processing applicationsKey factorsBiology
The invention relates to a variety analyzing method based on canopy light distributed computing. According to the method, by simulating interception and capturing for light by the crop light environment and a crop canopy, analyzing the difference of light distribution in the canopy, aiming at obtaining a high-yield variety structure, quantizing and analyzing the relation between character parameters related to the canopy structure and the high crop yield, and determining the key factors forming the yield difference, a guidance is provided for variety breeding. Compared with a traditional method, the variety analyzing method based on canopy light distributed computing has the advantages of saving time and labor and being easy to operate, and a modeling result has important practical value in the fields such as variety analyzing, computer virtual simulation and experiments.
Owner:QINGDAO ACADEMY OF INTELLIGENT IND
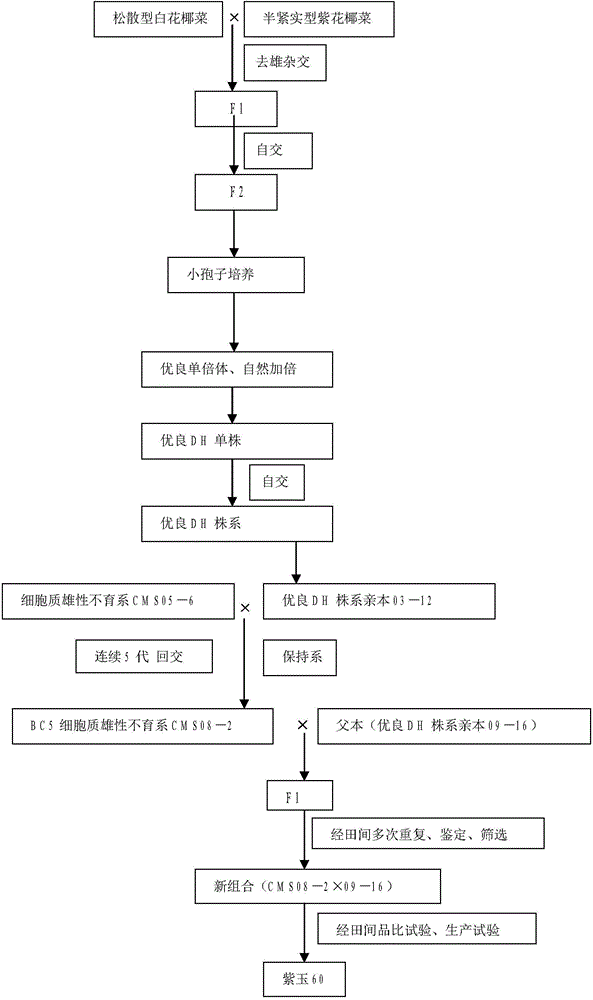
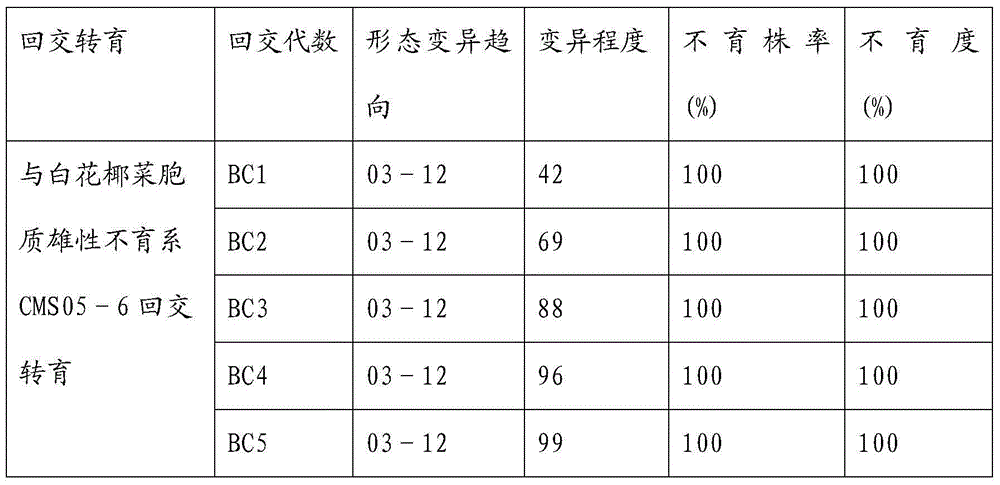
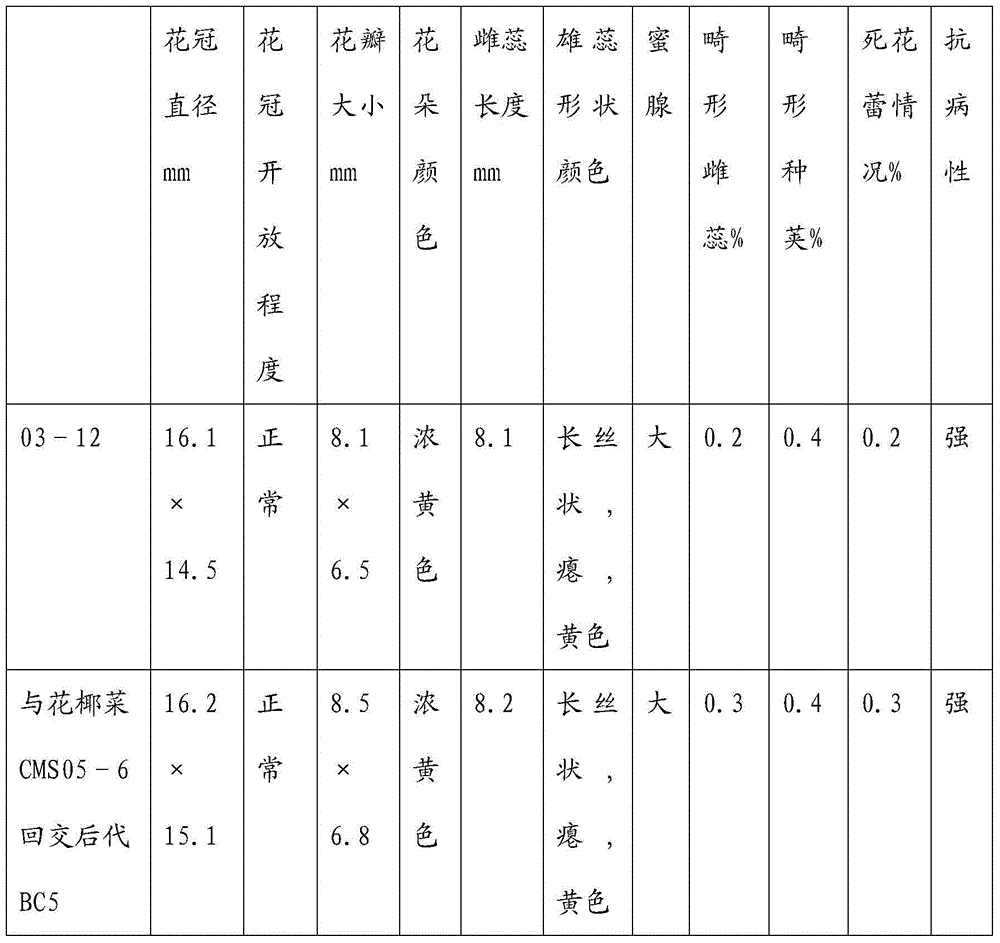
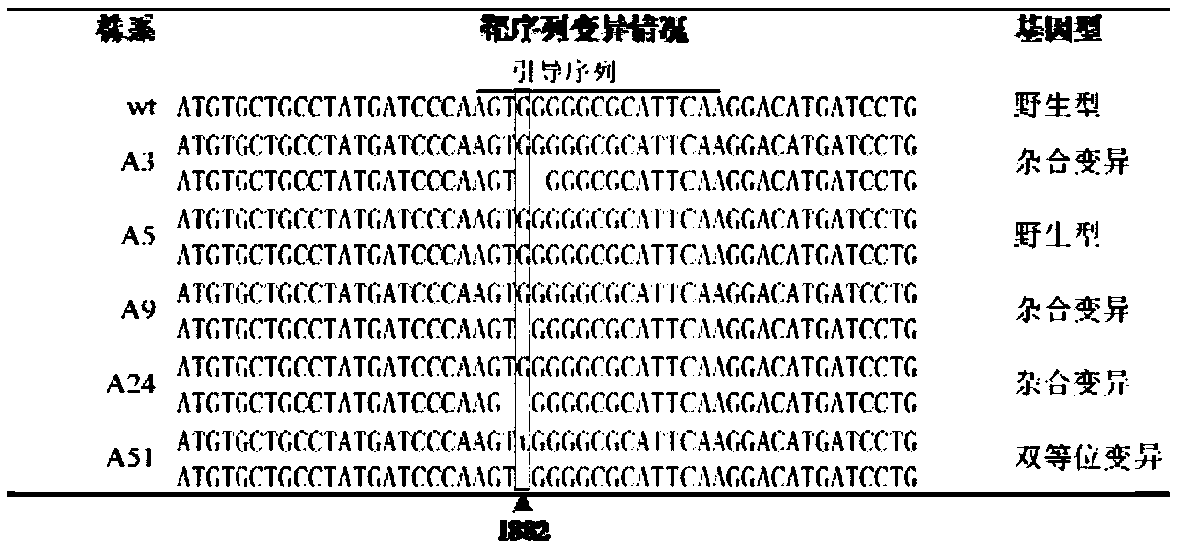
Breeding and cultivating method for early-matured, disease-resistant and loose purple cauliflowers
ActiveCN103975849AOvercoming developmental abnormalitiesImprove developmentPlant genotype modificationSporeSource material
The invention relates to a breeding and cultivating method for a plant, in particular relates to a breeding and cultivating method for loose purple cauliflowers, and belongs to the technical field of plant cultivation. The breeding and cultivating method comprises the steps of hybridizing a loose purple cauliflower breeding material with a semi-compact purple cauliflower breeding material to obtain seeds F1, and selfing single strains with high comprehensive agronomic characters after the seeds F1 are sowed to obtain seeds F2; performing microspore culture on single loose purple cauliflowers through flower buds suitable for the microspore culture from the mononuclear edging stage to the double-core early stage to grow a large number of normal regenerated plants, selfing excellent DH plants of the loose purple cauliflowers to obtain a loose purple cauliflower excellent DH strain 03-12 serving as a recurrent parent which is hybridized with a cauliflower cytoplasmic male sterility source material CMS05-6, performing five generations of backcross transformation and selection to form a cytoplasmic male sterile line CMS08-2, and hybridizing the DH strain serving as a parent with the excellent cytoplasmic male sterile line CMS08-2.
Owner:杨广林
Applications of ALS (acetolactate synthase) mutant type protein based on gene editing technology and ALS mutant type protein gene in plant breeding
ActiveCN109097346ANo significant change in basic agronomic traitsImprove breeding efficiencyMicrobiological testing/measurementTransferasesAgricultural scienceAcetolactate synthase
The invention discloses a rice ALS (acetolactate synthase) mutant type protein, a mutant type gene and applications of the rice ALS mutant type protein and the mutant type gene. The amino acid sequence of the ALS mutant type protein has following mutation: the 628-poisiton amino acid corresponding to the amino acid sequence of rice ALS has the mutation. The invention further discloses a breeding method of creating herbicide-resistant rice by gene editing. The CRISPR / Cas9 gene editing technology is used for editing the ALS gene for the first time, a T-DNA-knockout new material with the stably inherited herbicide resistant characteristic can be obtained in the T2 generation through offspring screening, and basic agronomic traits of the new material have no obvious change. Compared with breeding based on chemical mutagenesis, hybrid transform breeding and the like, the orderly improved molecular breeding technology based on gene editing has the advantages of being rapid, accurate, efficient and the like, and by means of combination with gene function marked genotype selection, breeding efficiency can be greatly increased and breeding progress is substantially accelerated.
Owner:JIANGSU ACAD OF AGRI SCI
Method for rapid in-vitro propagation of Crassulaceae plant
InactiveCN104273028AImprove germination rateImprove breeding efficiencyPlant tissue cultureHorticulture methodsNormal growthBud
The present invention discloses a method for rapid in-vitro propagation of a Crassulaceae plant. The method comprises the steps of carrying out aseptic germination of seeds, inducing adventitious bud, and carrying out in-vitro rooting of the adventitious buds. The method allows in-vitro germinated seedlings to normally grow and to be induced within a short period of time to continuously differentiate from original single-seed single seedlings into single-seed multiple seedlings, so the propagation cycle is greatly shortened, and the growth cycle of the differentiated seedlings is greatly shortened. The method provides a stable technical platform for improvement of the breeding efficiency of high-quality varieties of the Crassulaceae plant.
Owner:SHANGHAI INST OF BIOLOGICAL SCI CHINESE ACAD OF SCI
Method for reducing juvenile period of hybrid tangerine
InactiveCN101884294AShortened childhoodReduce R&D costsSeed and root treatmentCultivating equipmentsDiseaseTangerine Tree
The invention relates to a method for reducing juvenile period of hybrid tangerine, which comprises the following steps: culturing hybrid seedlings till the hybrid seedlings grow two true leaves and two stages of roots; performing high grafting of the hybrid seedlings onto tangerine branches which have closer relationships with grafts, is free from diseases and insects and 100 centimeters above the ground and have diameters more than 3 centimeters; and finally, performing final-period management such as tree shape control and girdling for promoting promoting flower-formation. When the method is used, the juvenile period of hybrid tangerine trees is reduced to about 2 years, and the juvenile period of some hybrid tangerine trees is reduced to 1 year. Compared with the conventional cultivation method of hybrid tangerine breeding seedlings, the method has the advantages of greatly reducing breeding period and accelerating the breeding of tangerine hybridization breeding.
Owner:CITRUS RES INST SOUTHWEST UNIV