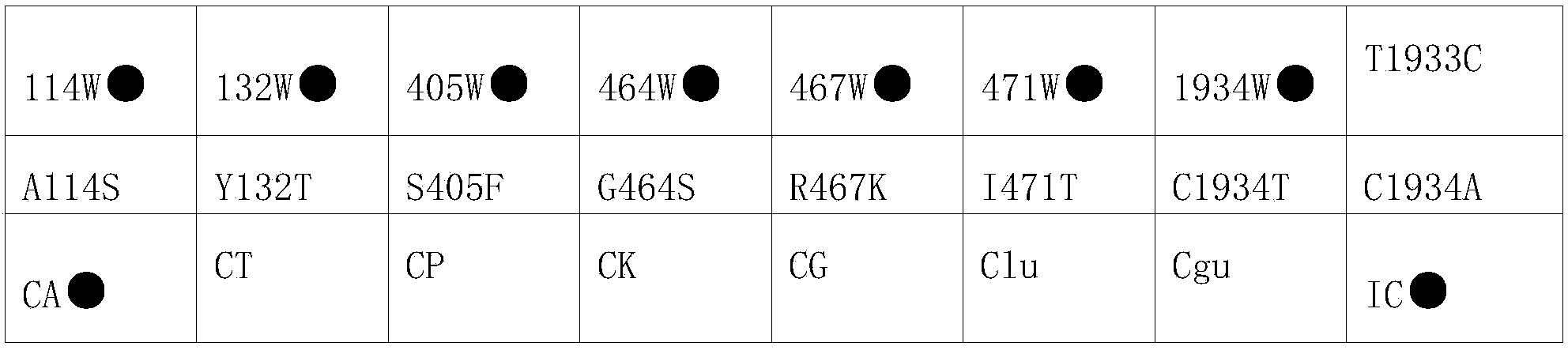Patents
Literature
254 results about "Candida dubliniensis" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
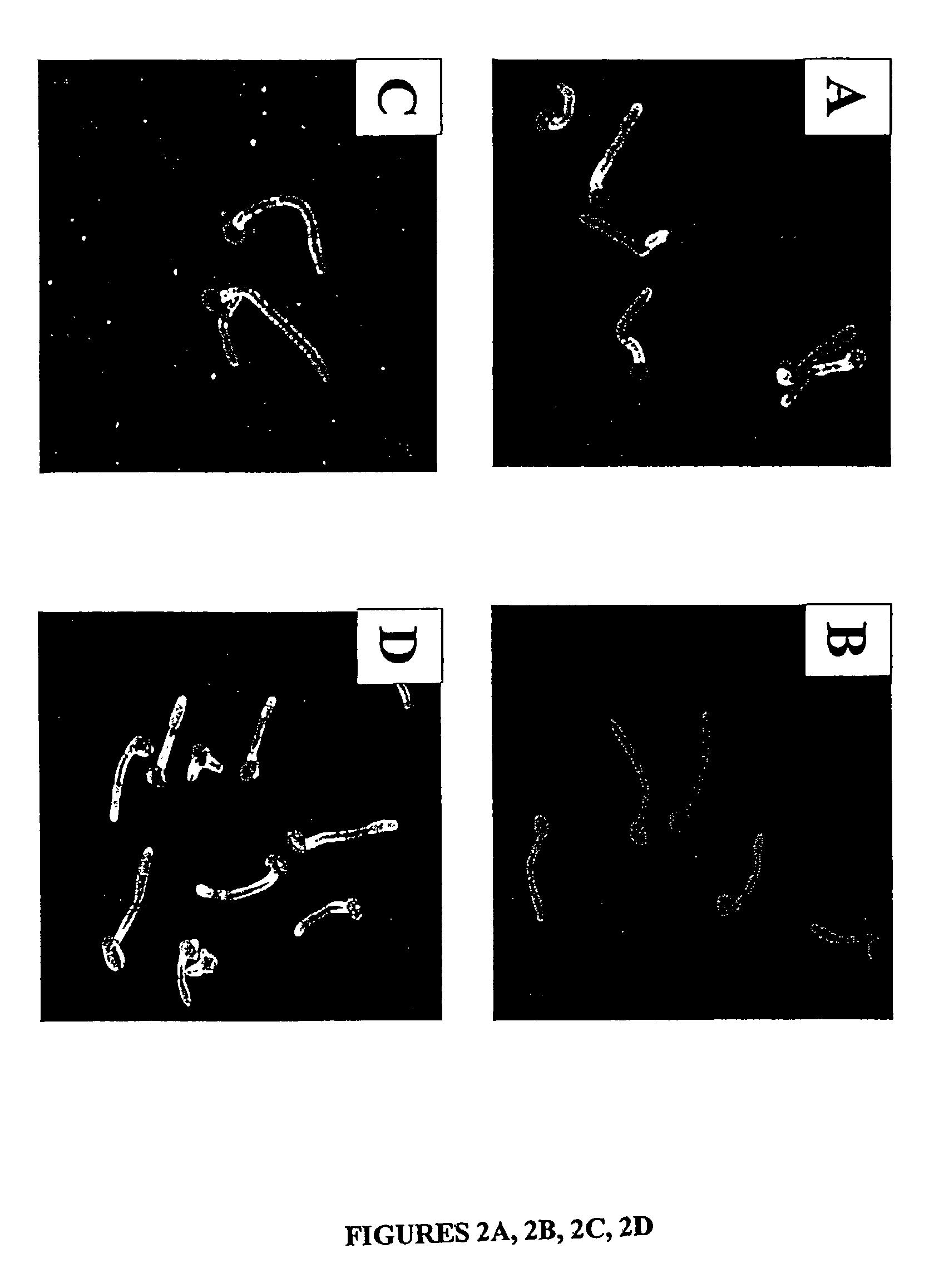
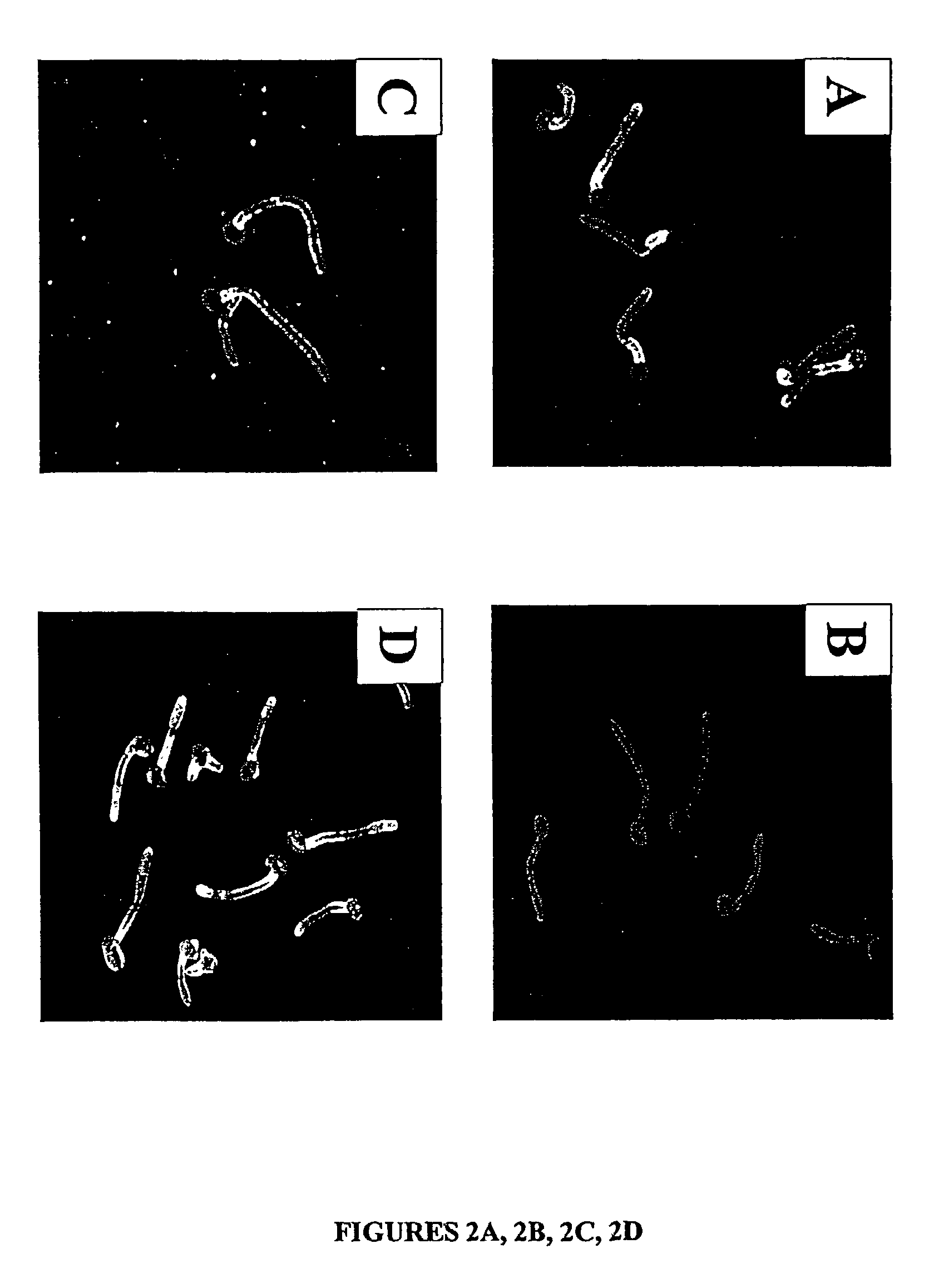
Candida dubliniensis is a fungal opportunistic pathogen originally isolated from AIDS patients. It is also occasionally isolated from immunocompetent individuals. It is a dimorphic yeast of the genus Candida, very closely related to Candida albicans but forming a distinct phylogenetic cluster in DNA fingerprinting. It is most commonly isolated from oral cavities, and is also occasionally found in other anatomical sites.
Nucleic acid probes and methods for detecting clinically important fungal pathogens
InactiveUS6858387B1Function increaseAttached with easeSugar derivativesMicrobiological testing/measurementNucleic Acid ProbesAspergillus flavus
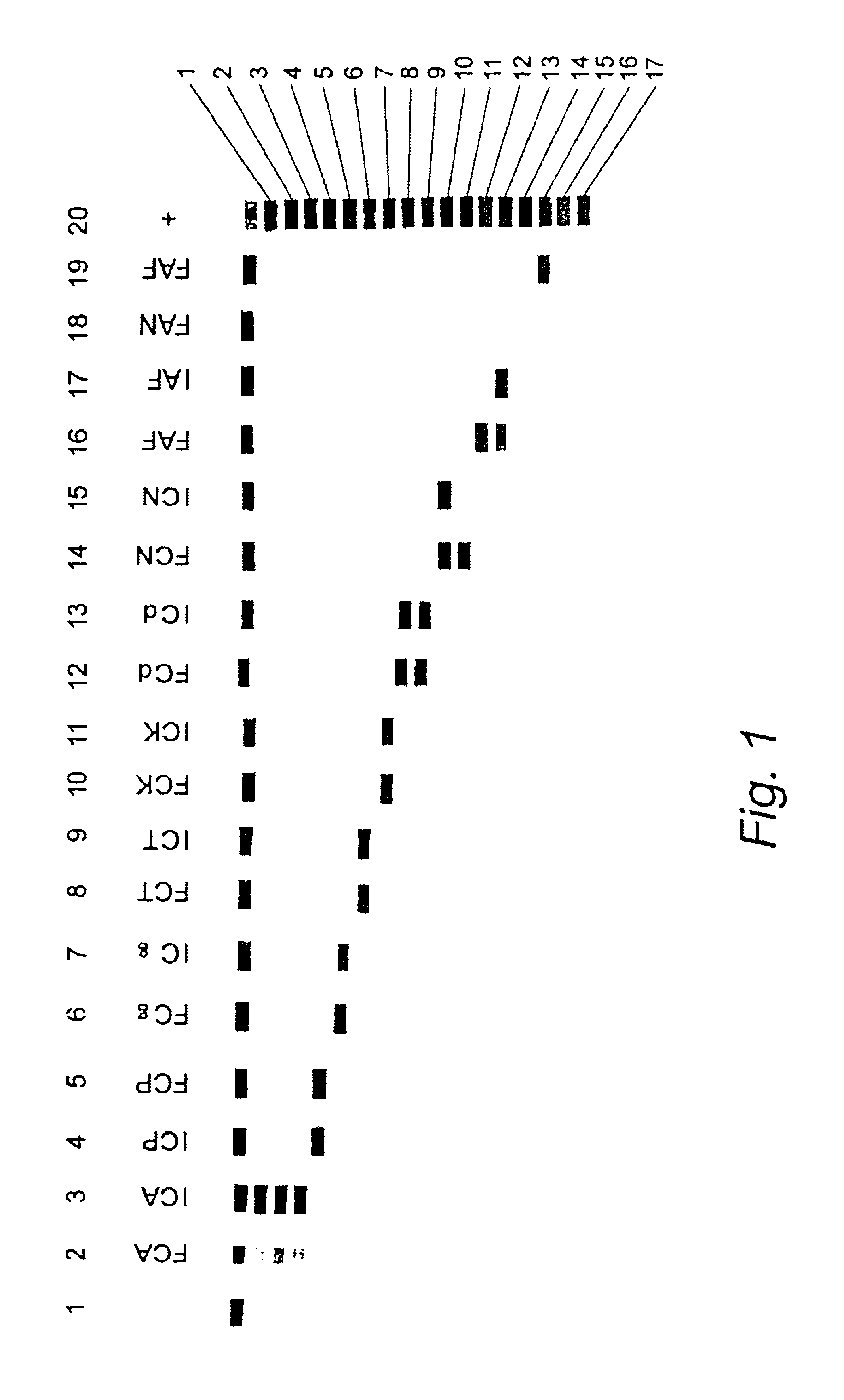
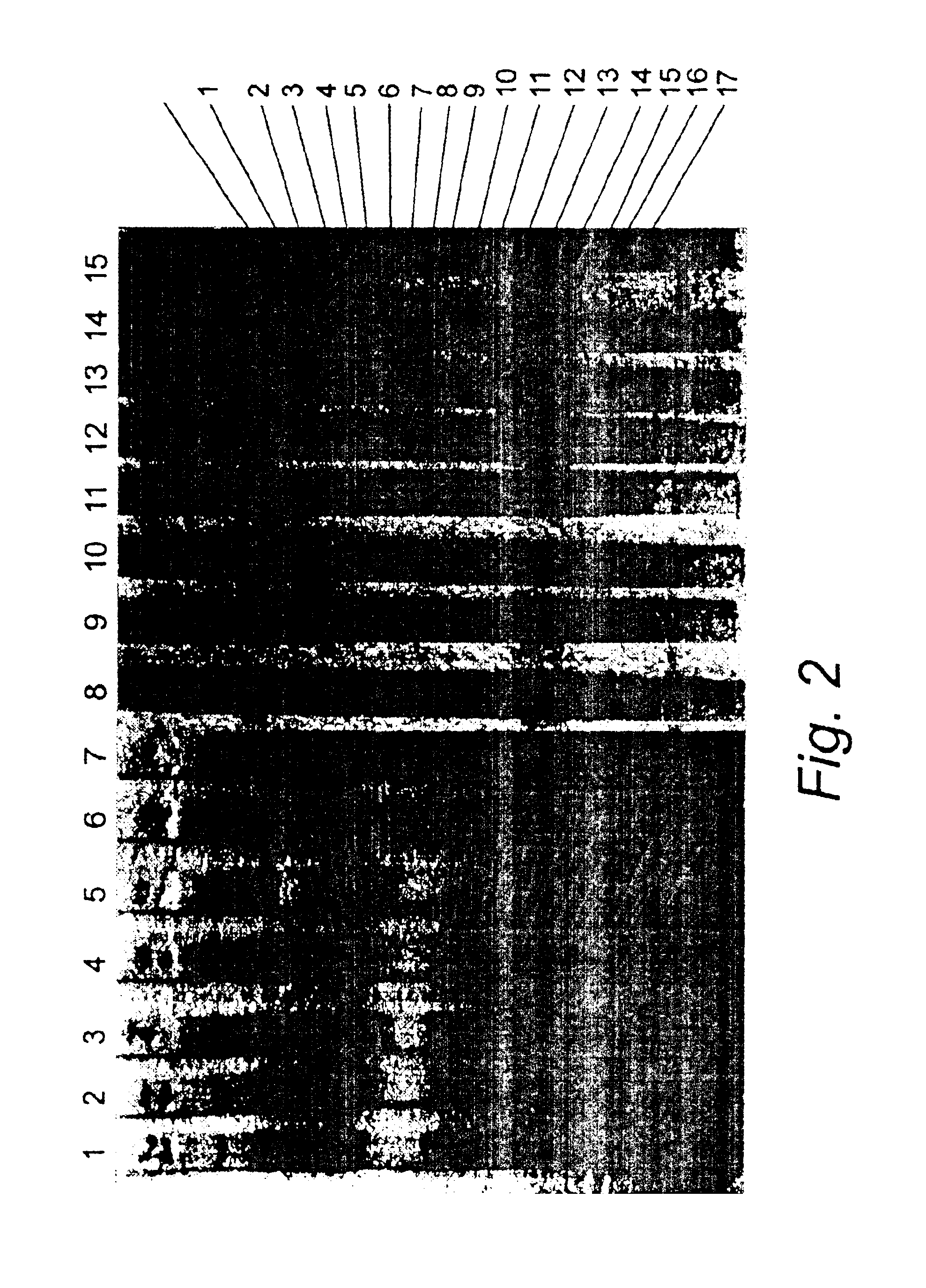
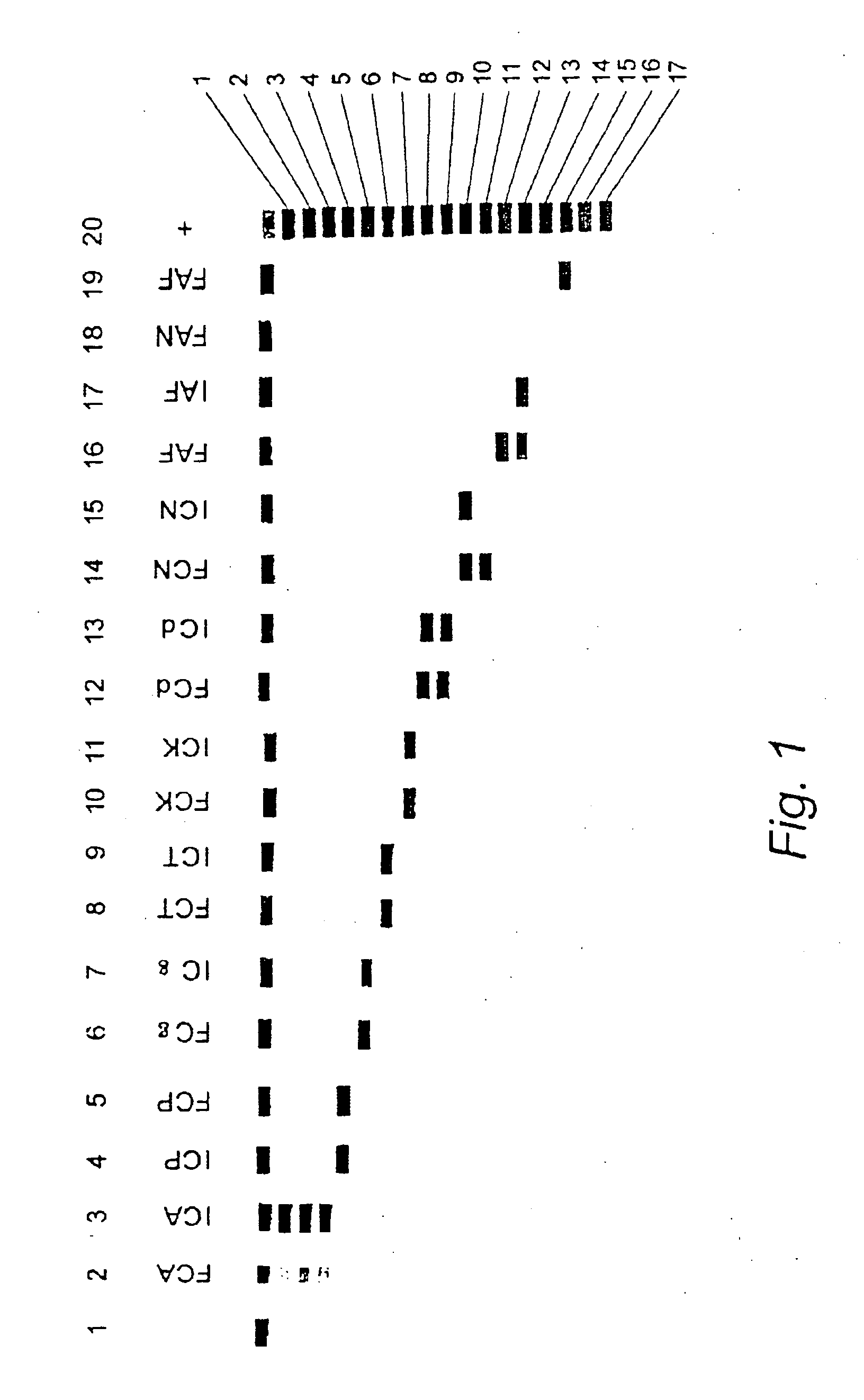
The current invention relates to the field of detection and identification of clinically important fungi. More particularely, the present invention relates to species specific probes originating from the Internal Transcribed Spacer (ITS) region of rDNA for the detection of fungal species such as Candida albicans, Candida parapsilosis, Candida tropicalis, Candida kefyr, Candida krusei, Candida glabrata, Candida dubliniensis, Aspergillus flavus, Aspergillus versicolor, Aspergillus nidulans, Aspergillus fumigatus, Cyptococcus neoformans and Pneumocystis carinii in clinical samples, and methods using said probes.
Owner:ENTERPRISE IRELAND +2
Pharmaceutical compositions and methods to vaccinate against candidiasis
InactiveUS20030124134A1Treat prevent alleviateToxic reductionPeptide/protein ingredientsSnake antigen ingredientsHospitalized patientsVirulent characteristics
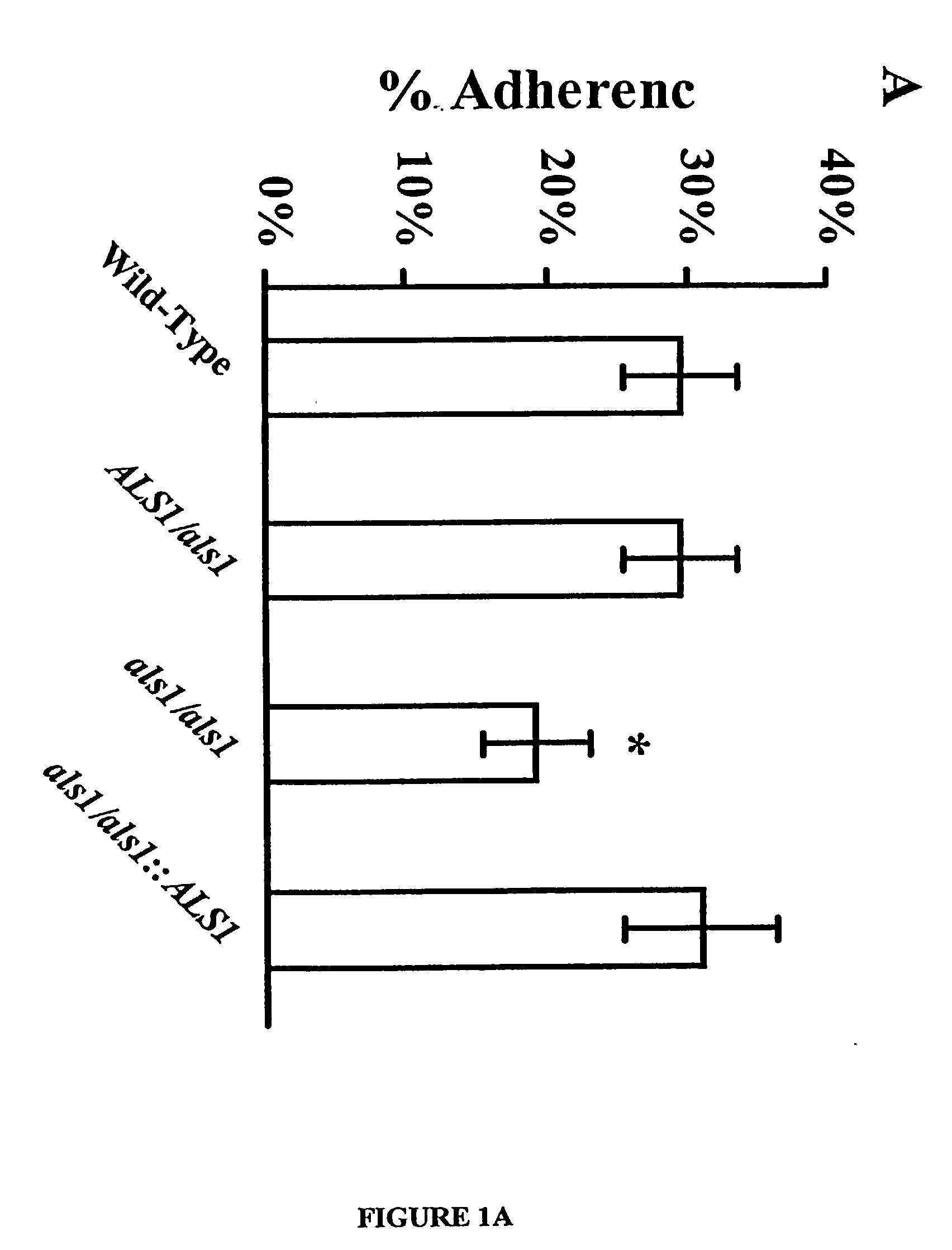
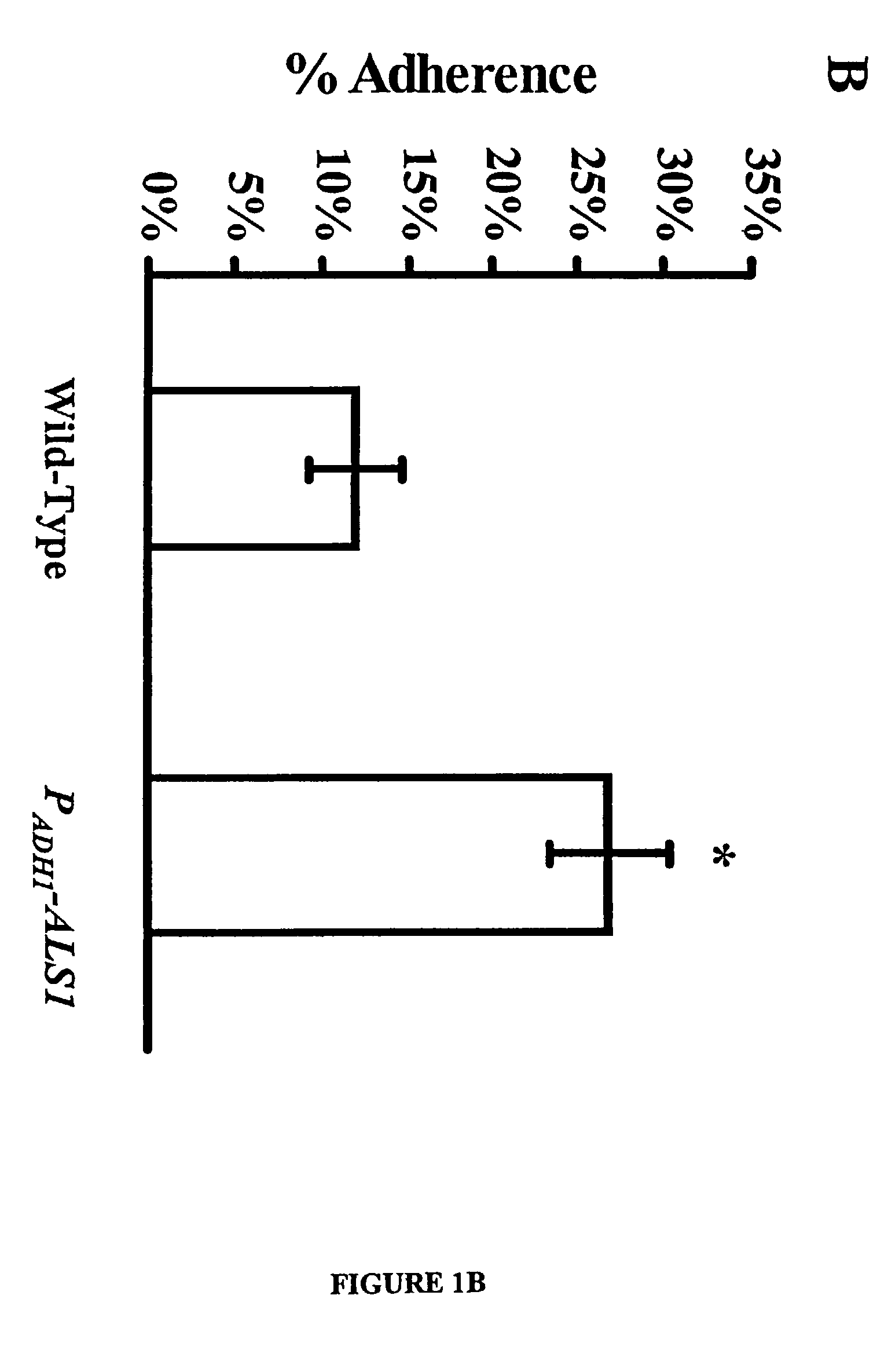
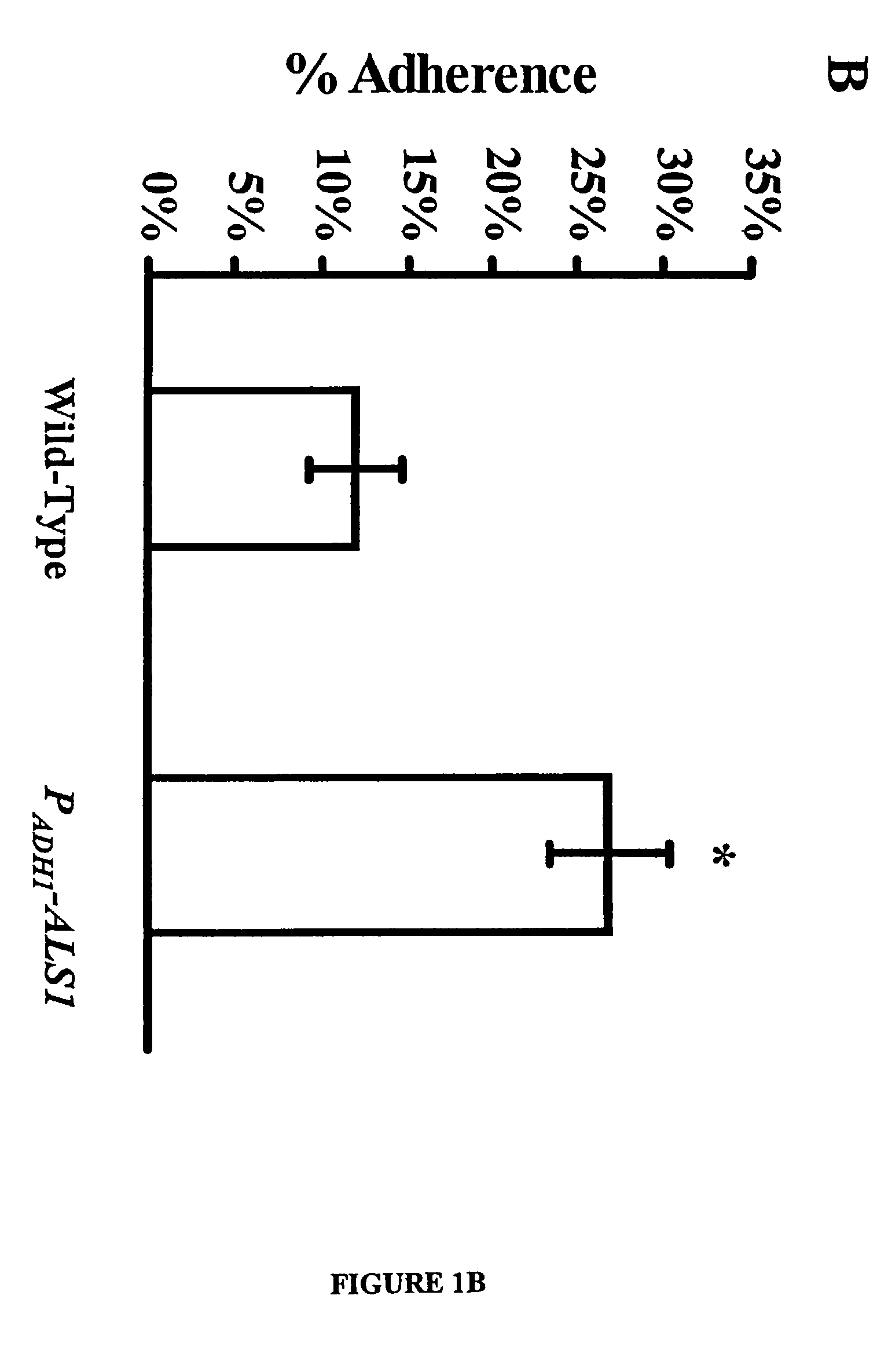
A Candida albicans bloodstream infections cause significant morbidity and mortality in hospitalized patients. Filament formation and adherence to host cells are critical virulence factors of C. albicans. Multiple filamentation regulatory pathways have been discovered, however the downstream effectors of these regulatory pathways remain unknown. The cell surface proteins in the ALS group are downstream effectors of the filamentation regulatory pathway. Particularly, Als1p mediates adherence to endothelial cells in vitro and is required for virulence. The blocking of adherence by the organism is described resulting from the use of a composition and method disclosed herein. Specifically, a pharmaceutical composition comprised of a gene, gene product, or specific antibody to the ALS gene family is administered as a vaccine to generate an immune response capable of blocking adherence of the organism.
Owner:LOS ANGELES BIOMEDICAL RES INST AT UCLA HARBOR MEDICAL CENT +1
Species-specific, genus-specific and universal DNA probes and amplification primers to rapidly detect and identify common bacterial and fungal pathogens and associated antibiotic resistance genes from clinical specimens for diagnosis in microbiology laboratories
InactiveUS20040185478A1Reduce usageDetermine rapidly the bacterial resistance to antibioticsMicrobiological testing/measurementFermentationBacteroidesNeisseria meningitidis
DNA-based methods employing amplification primers or probes for detecting, identifying, and quantifying in a test sample DNA from (i) any bacterium, (ii) the species Streptococcus agalactiae, Staphylococcus saprophyticus, Enterococcus faecium, Neisseria meningitidis, Listeria monocytogenes and Candida albicans, and (iii) any species of the genera Streptococcus, Staphylococcus, Enterococcus, Neisseria and Candida are disclosed. DNA-based methods employing amplification primers or probes for detecting, identifying, and quantifying in a test sample antibiotic resistance genes selected from the group consisting of blatem, blarob, blashv, blaoxa, blaZ, aadB, aacC1, aacC2, aacC3, aacA4, aac6'-lla, ermA, ermB, ermC, mecA, vanA, vanB, vanC, satA, aac(6')-aph(2''), aad(6'), vat, vga, msrA, sul and int are also disclosed. The above microbial species, genera and resistance genes are all clinically relevant and commonly encountered in a variety of clinical specimens. These DNA-based assays are rapid, accurate and can be used in clinical microbiology laboratories for routine diagnosis. These novel diagnostic tools should be useful to improve the speed and accuracy of diagnosis of microbial infections, thereby allowing more effective treatments. Diagnostic kits for (i) the universal detection and quantification of bacteria, and / or (ii) the detection, identification and quantification of the above-mentioned bacterial and fungal species and / or genera, and / or (iii) the detection, identification and quantification of the above-mentioned antibiotic resistance genes are also claimed.
Owner:GENEOHM SCI CANADA
Composition comprising a Lactobacillus pentosus strain and uses thereof
A novel probiotic strain termed Lactobacillus pentosus NCIMB 41114 is capable of preventing growth of pathogenic microorganisms in the GI tract. In particular, this bacterium inhibits Candida overgrowth, and can be employed to prevent and treat associated diseases, including IBS and thrush.
Owner:READING SCHOOL OF FOOD BIOSCI FOOD MICROBIAL SCI UNIT UNIV OF
Species-specific, genus-specific and universal DNA probes and amplification primers to rapidly detect and identify common bacterial and fungal pathogens and associated antibiotic resistance genes from clinical specimens for diagnosis in microbiology laboratories
InactiveUS20030049636A1Low costRapid positioningMicrobiological testing/measurementFermentationBacteroidesNeisseria meningitidis
DNA-based methods employing amplification primers or probes for detecting, identifying, and quantifying in a test sample DNA from (i) any bacterium, (ii) the species Streptococcus agalactiae, Staphylococcus saprophyticus, Enterococcus faecium, Neisseria meningitidis, Listeria monocytogenes and Candida albicans, and (iii) any species of the genera Streptococcus, Staphylococcus, Enterococcus, Neisseria and Candida are disclosed. DNA-based methods employing amplification primers or probes for detecting, identifying, and quantifying in a test sample antibiotic resistance genes selected from the group consisting of blatem, blarob, blashv, blaoxa, blaZ, aadB, aacC1, aacC2, aacC3, aacA4, aac6'-lla, ermA, ermB, ermC, mecA, vanA, vanB, vanC, satA, aac(6')-aph(2''), aad(6'), vat, vga, msrA, sul and int are also disclosed. The above microbial species, genera and resistance genes are all clinically relevant and commonly encountered in a variety of clinical specimens. These DNA-based assays are rapid, accurate and can be used in clinical microbiology laboratories for routine diagnosis. These novel diagnostic tools should be useful to improve the speed and accuracy of diagnosis of microbial infections, thereby allowing more effective treatments. Diagnostic kits for (i) the universal detection and quantification of bacteria, and / or (ii) the detection, identification and quantification of the above-mentioned bacterial and fungal species and / or genera, and / or (iii) the detection, identification and quantification of the above-mentioned antibiotic resistance genes are also claimed.
Owner:BERGERON MICHEL G +3
Methods and compositions for detecting and identifying species of Candida
ActiveUS20080102449A1Avoid disadvantagesMicrobiological testing/measurementFermentationForward primerNucleotide
Methods and compositions useful in the detection and identification of species of Candida are disclosed. These species include Candida albicans, Candida glabrata, Candida parapsilosis, and Candida tropicalis, each of which is a causative agent for vaginal candidiasis. The compositions of the invention are combinations of oligonucleotides. These oligonucleotides include pairs of forward and reverse primers for polymerase chain reactions, wherein each primer pair is capable of priming the synthesis of an amplicon specific to one of Candida albicans, Candida glabrata, Candida parapsilosis, and Candida tropicalis, but preferably is not capable of priming the synthesis of an amplicon specific to any of the other three species. In preferred embodiments, the forward primers of the primer pairs have identical sequences, while each reverse primer of the primer pairs has a unique sequence relative to all of the other reverse primers; or the reverse primers of the primer pairs have identical sequences, while each forward primer of the primer pairs has a unique sequence relative to all of the other forward primers. These unique primer sequences account for the species specificity of the resultant amplicons. The oligonucleotides also include probes capable of detecting these amplicons, and sequencing primers for determining, in primer extension reactions, the nucleotide sequences contained within the amplicons. In preferred methods of the invention, a biological sample is tested for the presence of at least one isolate of Candida albicans, Candida glabrata, Candida parapsilosis, and Candida tropicalis by isolating nucleic acid from the sample, attempting a polymerase chain reaction in a mixture containing this nucleic acid and a plurality of these primer pairs, ascertaining whether any amplicon is produced in the mixture using an oligonucleotide probe, and determining the sequence of any resultant amplicon using the sequencing primers. The detection of an amplicon indicates that the sample contains at least one isolate of Candida albicans, Candida glabrata, Candida parapsilosis, or Candida tropicalis, and the nucleotide sequence data is used to determine which of these four Candida species is present.
Owner:MEDICAL DIAGNOSTIC LAB
Plant lactobacillus strain and its application
InactiveCN1888051AImprove cleanlinessImprove adhesionBacteriaBacteria material medical ingredientsEscherichia coliStaphylococcus cohnii
The present invention discloses one plant lactobacillus strain and its application. The plant lactobacillus, Lactobacillus plantarum L323 CGMCC No. 1329, is separated from Chinese pregnant woman's vaginal secretion, and has relatively high bacteriotasis on common vaginal pathogens, such as staphylococcus aureus, candida albicans, colibacillus and vaginal Gardnar bacillus, and acid producing and H2O2 producing capacity higher than other lactobacillus. The plant lactobacillus, Lactobacillus plantarum L323 CGMCC No. 1329, may be used in preparing vaginal microbial preparation for preventing and / or treating vaginal infectious diseases.
Owner:TIANYOUDA BIO ENG SCI & TECH BEIJING +1
Modified coconut oils with broad antimicrobial spectrum
InactiveUS20100016430A1Improve antibacterial propertiesAntibacterial agentsBiocideEscherichia coliCandida famata
Owner:MALAYSIAN AGRI RES & DEV INST MARDI
Fluorescence quantitative PCR primer, probe and kit for detecting ordinary pathogenic fungi
InactiveCN104450936AOvercome limitationsSolve the problem of fungal infectionMicrobiological testing/measurementDNA/RNA fragmentationFluorescenceCandida tropicalis
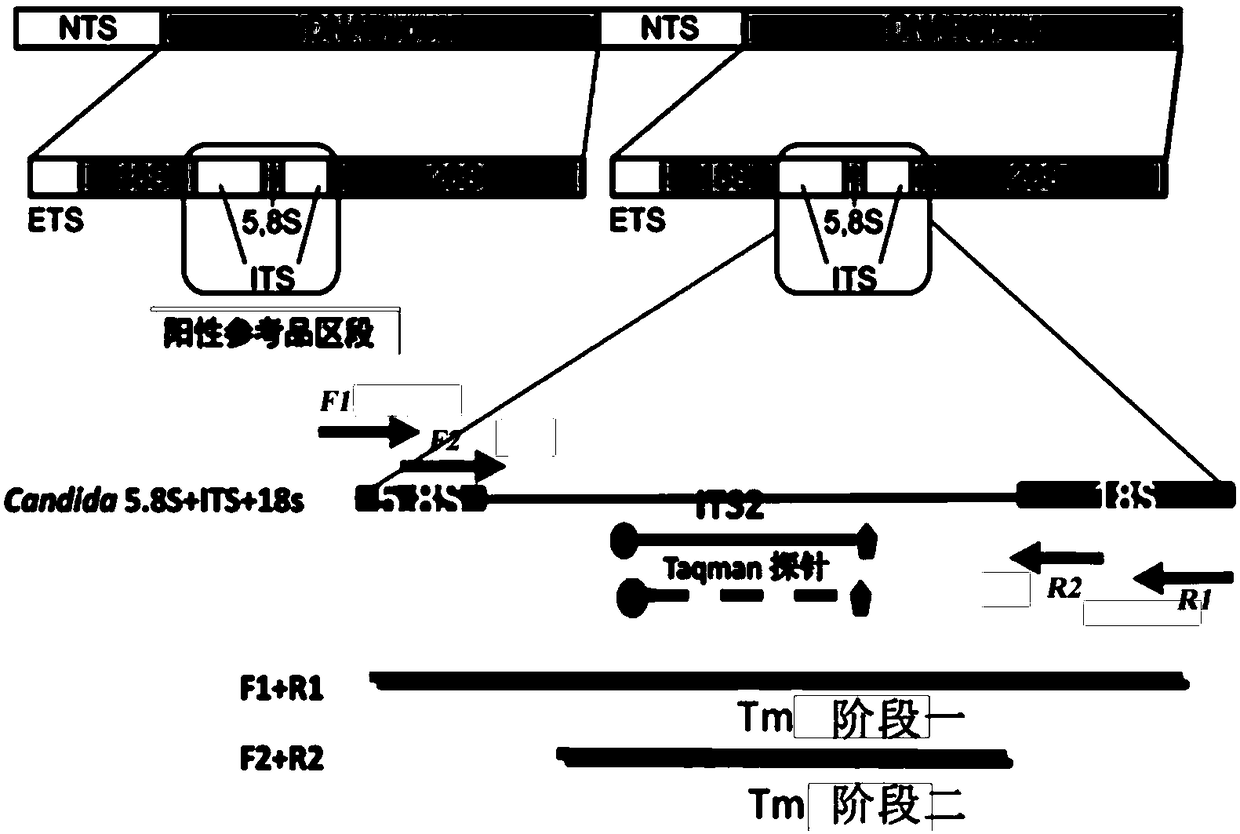
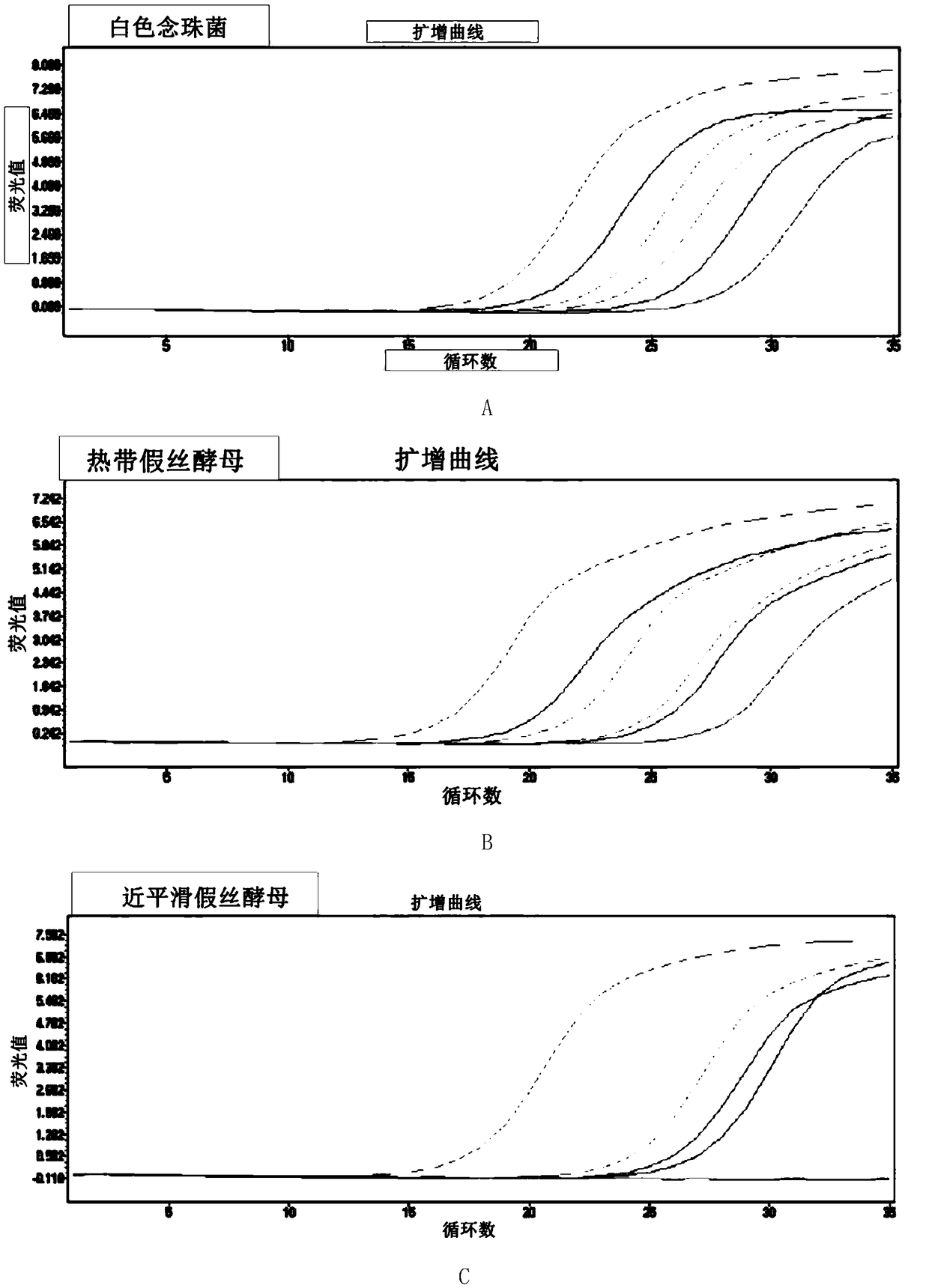
The invention discloses a fluorescence quantitative PCR primer, a probe and a kit for detecting ordinary pathogenic fungi. A specific primer and a TaqMan probe are independently designed, and a fluorescence PCR detection method which can be used for simultaneously detecting 15 clinically ordinary pathogenic fungi, including 8 candida mycoderma bacteria (candida albicans, candida glabrata, candida parapsilosis, candida kefyr, candida sake, candida kruse, candida guilliermind and candida tropicalis), four aspergilli (aspergillus niger, aspergillus flavus, aspergillus terreus and aspergillus fumigates), cryptococcus, rhizopus oryzae and mucor circinelloides, is established. The method is relatively high in sensitivity and specificity, and has significant meanings for early diagnosis and treatment on invasive infections with fungi.
Owner:天津宝瑞生物技术有限公司
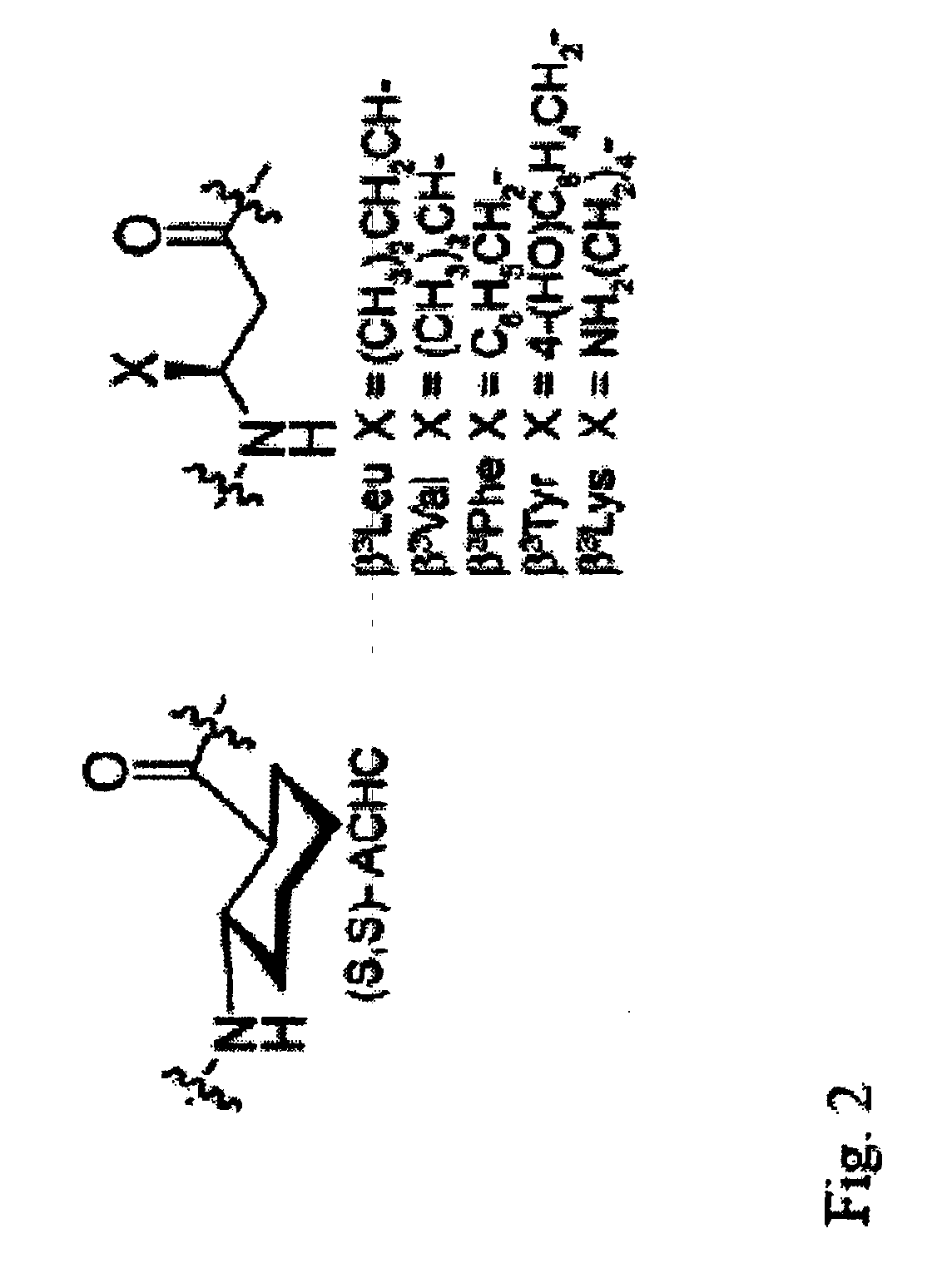
Beta-peptides with antifungal activity
The present invention is directed to the design, synthesis and use of various β-peptides exhibiting antifungal activity. The β-peptides are relatively short in length, adopt globally amphiphilic conformations, and cause little lysis of human red blood cells at concentrations that kill Candida albicans, a common human fungal pathogen.
Owner:WISCONSIN ALUMNI RES FOUND
Detection method of invasive fungus infection, detection kit and application
ActiveCN109055502ASimplify detectionSimplify operabilityMicrobiological testing/measurementDNA/RNA fragmentationCell freeFluorescence
The invention provides a fast multiplex PCR identification diagnosis detection method of invasive fungus infection based on cfDNA (cell free DNA). The method can be used for identifying the invasive infection caused by clinic common and high-incidence Candida albicans, Candida tropicalis, Candida parapsilosis, Candida krusei, Candida glabrata and Aspergillus fumigtus. An amplification primer is designed according to characteristic genome segments of each fungus category and species; a detection fluorescence probe of a strain can be distinguished according to the amplification segment design; the real time PCR can be performed on a sample to be tested; high sensitivity of nested PCR and high specificity and multi-target performance advantages of multiple fluorescent hybrid probe PCR are integrated for identifying the fungus strain. The invention also provides a PCR diagnosis kit for the invasive fungus infection and application thereof.
Owner:DIASYS DIAGNOSTIC SYST SHANGHAI
Weissella cibaria XHR1 and applications thereof and weissella cibaria XHR1 containing pickled vegetable
ActiveCN105255787AImprove micro-ecological levelExpansion of industrial productionBacteriaMicroorganism based processesWeissella cibariaCandida famata
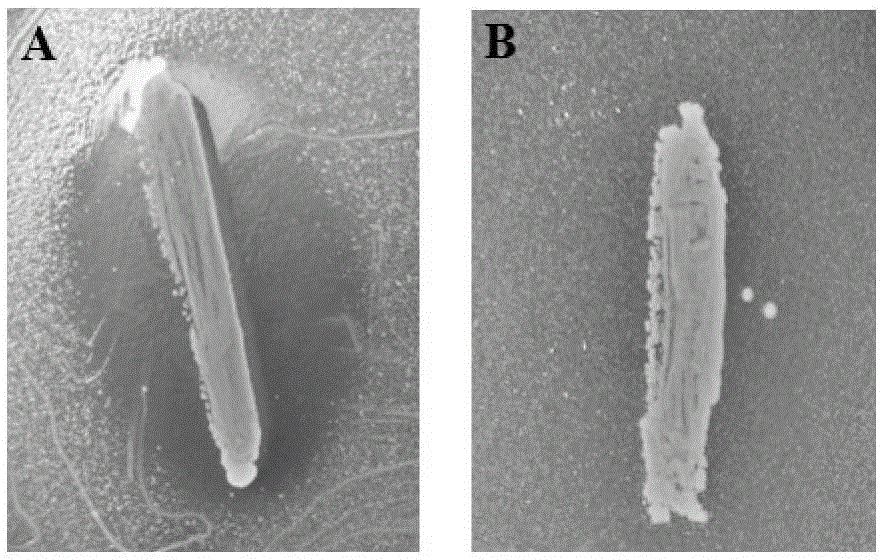
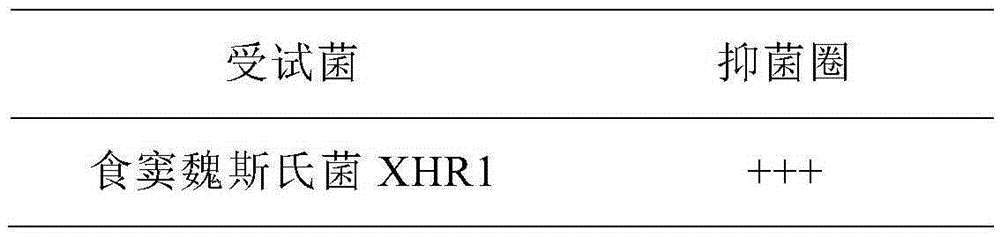
The invention provides a weissella cibaria XHR1 and applications thereof and a weissella cibaria XHR1 containing pickled vegetable. The preparation of the pickled vegetable is implemented through the steps of adding a culture solution of weissella cibaria XHR1 and bacterial powder of lactobacillus plantarum CICC 20764 into a pickled vegetable fermentation starting solution for pickling vegetables, so that the final concentration of weissella cibaria XHR1 is 10<6>-10<7> cfu / mL eventually, and then the final concentration of lactobacillus plantarum CICC 2076 is 10<5>-10<6> CFU / mL; and then, adding vegetables into the pickled vegetable fermentation starting solution, and carrying out vegetable pickling according to a traditional vegetable fermentation process, so that the pickled vegetable disclosed by the invention is obtained finally. According to the invention, prebiotic lactobacillus for inhibiting the growth of Candida albicans is applied to the fermentation of Sichuan pickled vegetables, so that the consumer group is wider, and the micro ecological level of alimentary canals of wide consumers can be improved, thereby inhibiting Candida albicans diseases from the perspective of public health.
Owner:XIHUA UNIV
Nucleic acid probes and methods for detecting clinically important fungal pathogens
InactiveUS20050164243A1Faster and simpler to performSuitable for automationSugar derivativesMicrobiological testing/measurementNucleic Acid ProbesAspergillus flavus
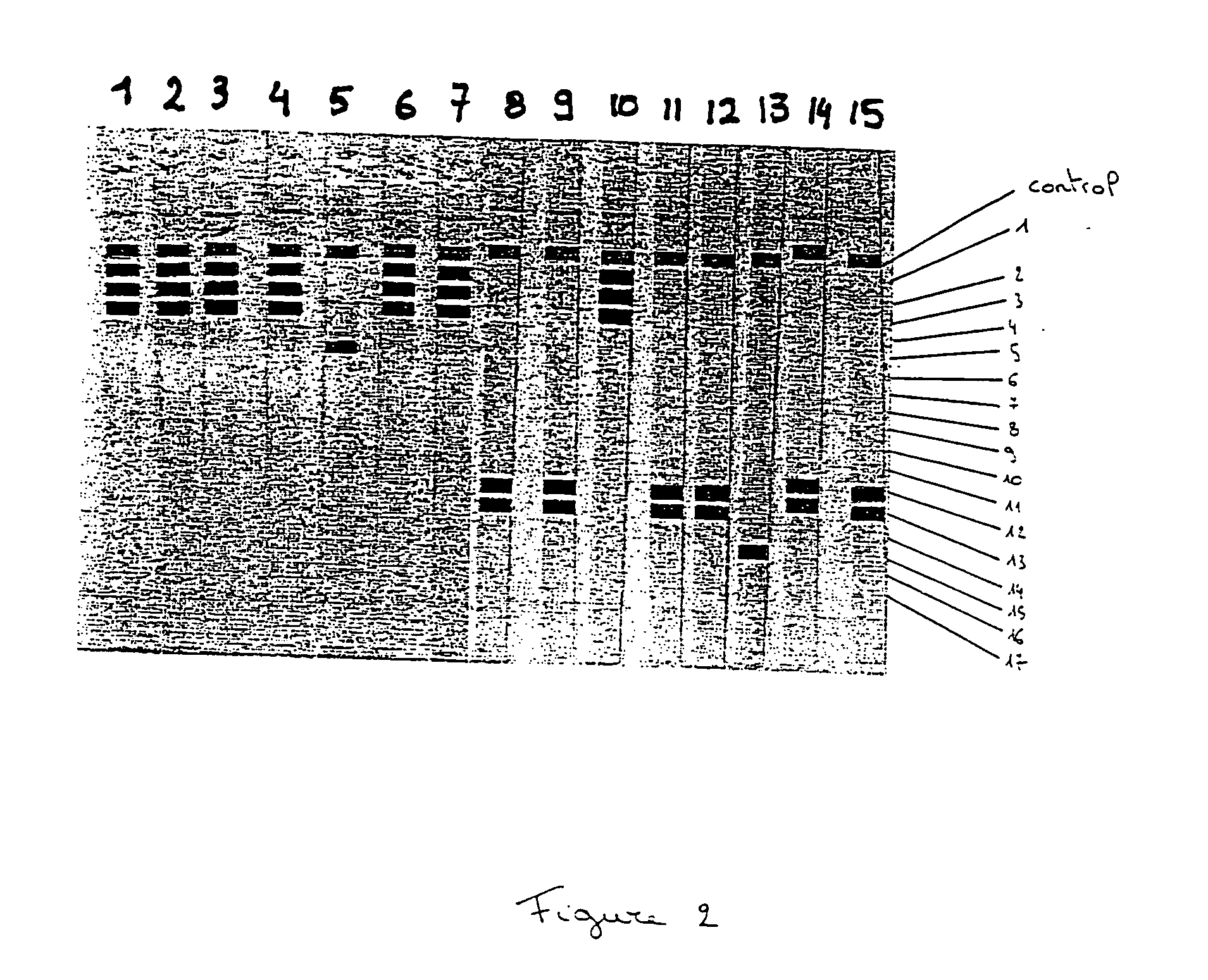
The current invention relates to the field of detection and identification of clinically important fungi. More particularely, the present invention relates to species specific probes originating from the Internal Transcribed Spacer (ITS) region of rDNA for the detection of fungal species such as Candida albicans, Candida parapsilosis, Candida tropicalis, Candida kefyr, Candida krusei, Candida glabrata, Candida dubliniensis, Aspergillus flavus, Aspergillus versicolor, Aspergillus nidulans, Aspergillus fumigatus, Cryptococcus neoformans and Pneumocystis carinii in clinical samples, and methods using said probes.
Owner:INNOGENETICS NV +2
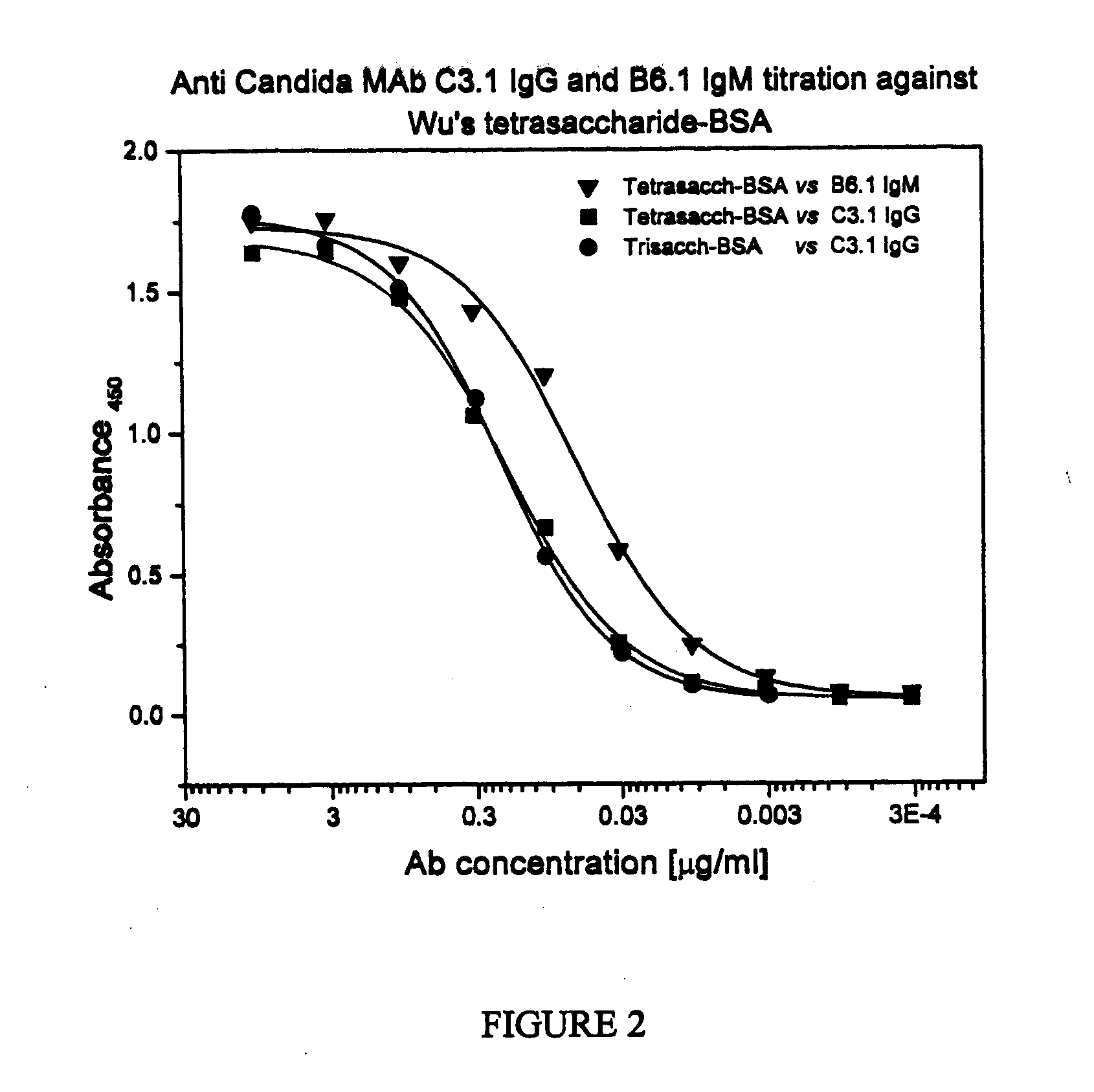
Synthetic Anti-Candida Albicans Oligosaccharide Based Vaccines
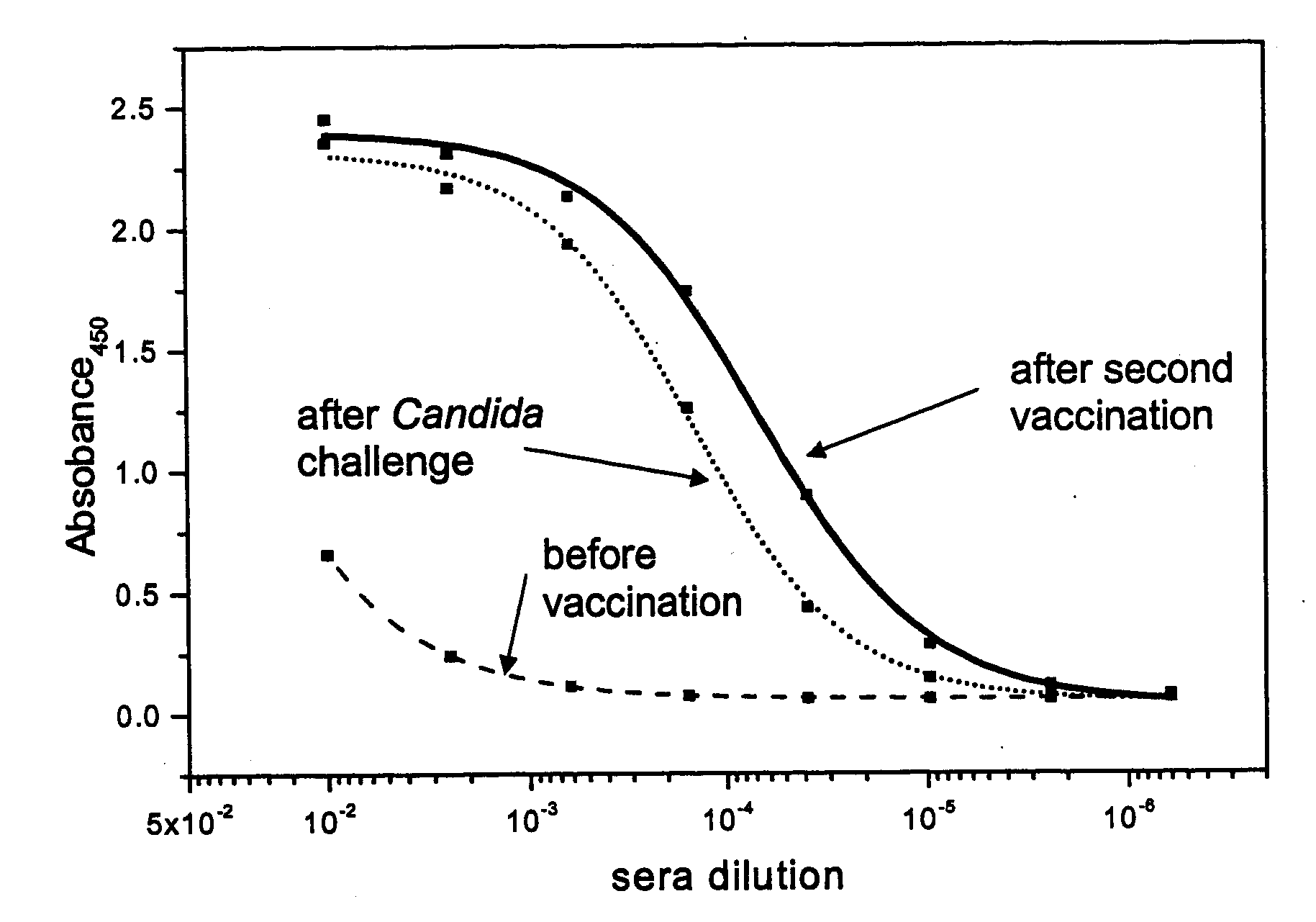
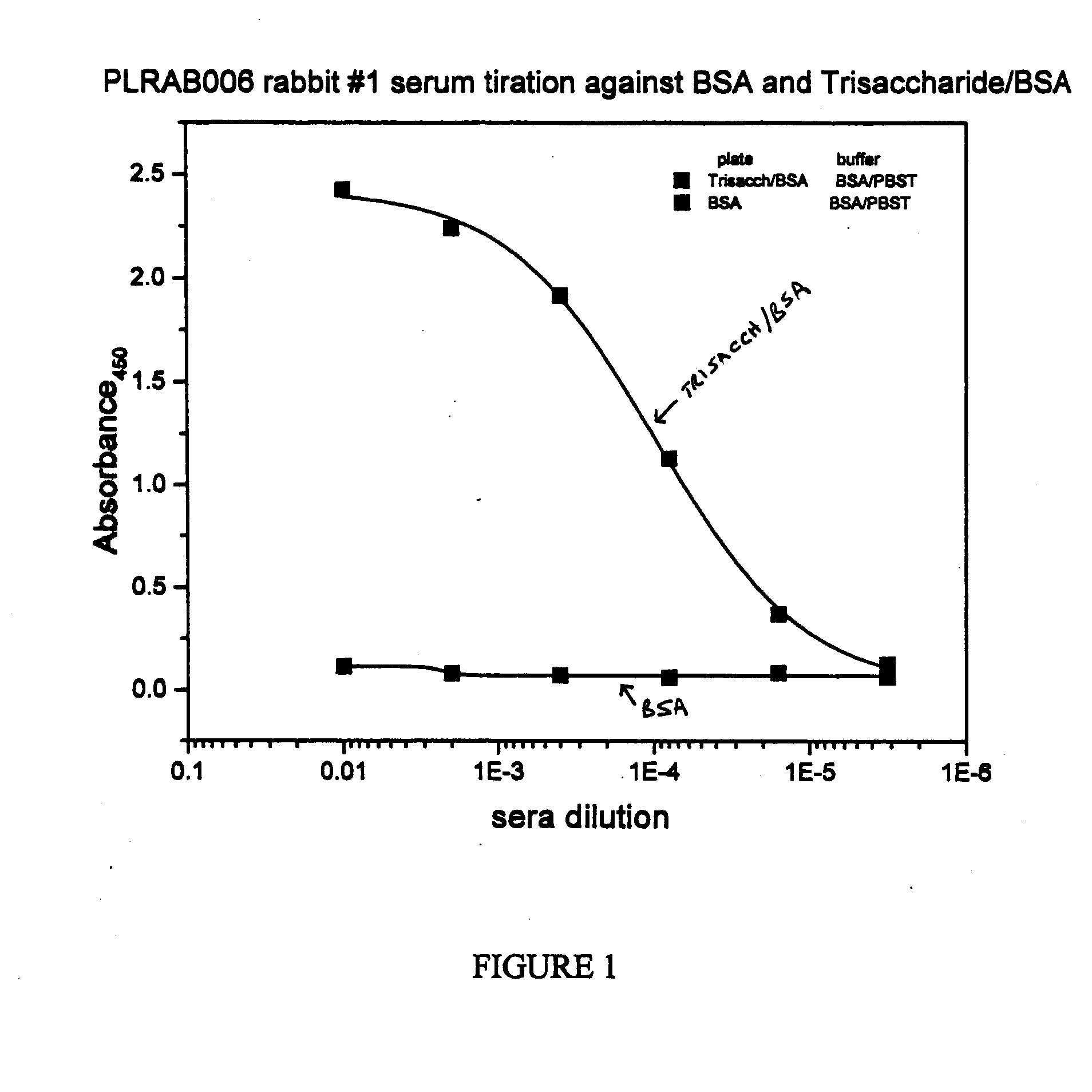
The present invention provides immunogenic oligosaccharide compositions and methods of making and using them. In particular, the compositions comprise native O-linked and S-linked oligosaccharides coupled to a protein carrier via a linker, wherein the resultant conjugate elicits a protectively immunogenic response, particularly in vaccines against pathogenic Candida species and more particularly against Candida albicans. Preferably the pathogenic Candida species are those that possess cell wall oligosaccharide compositions similar to the β-mannan component of Candida albicans cell walls.
Owner:THE GOVERNORS OF THE UNIV OF ALBERTA
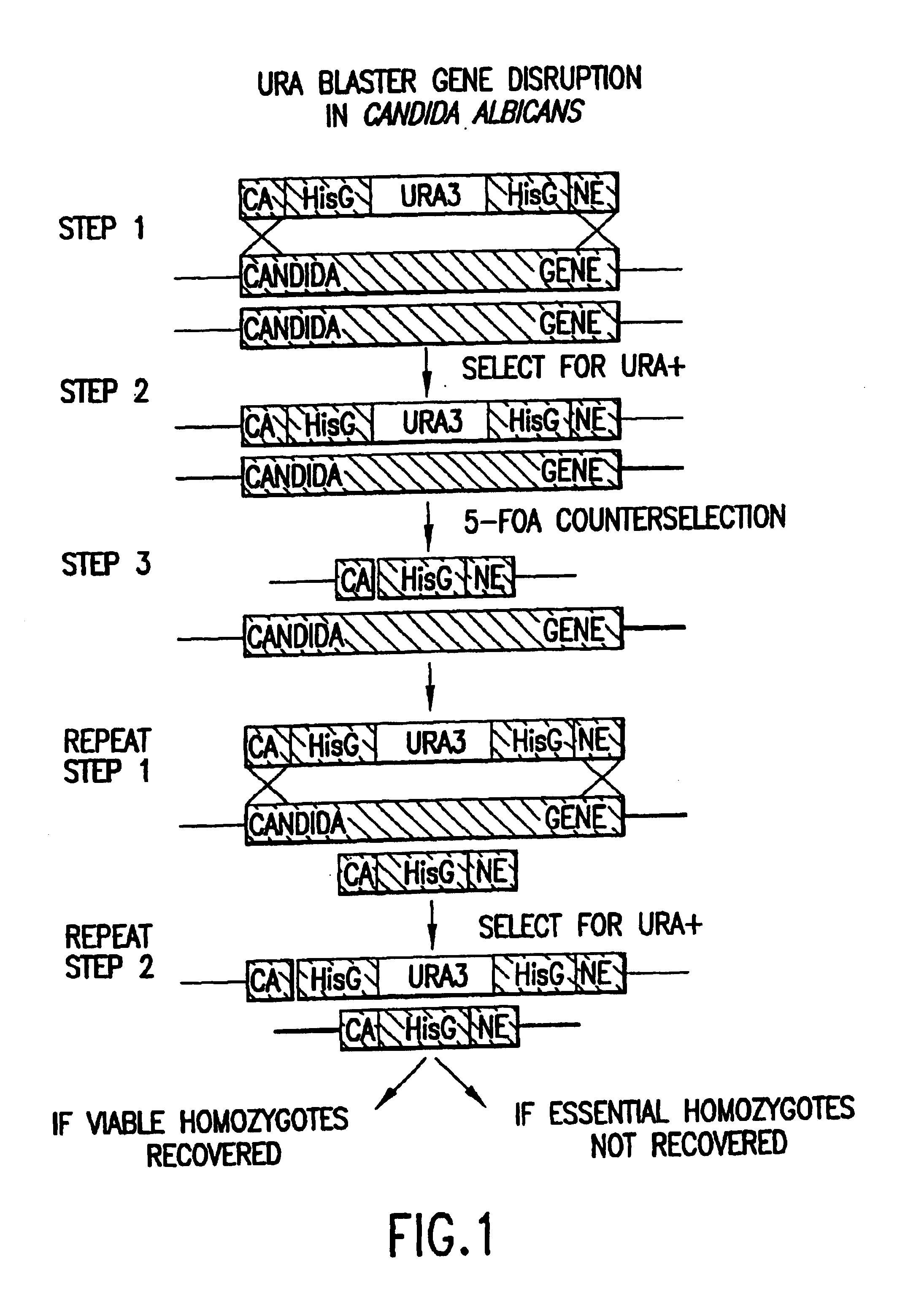
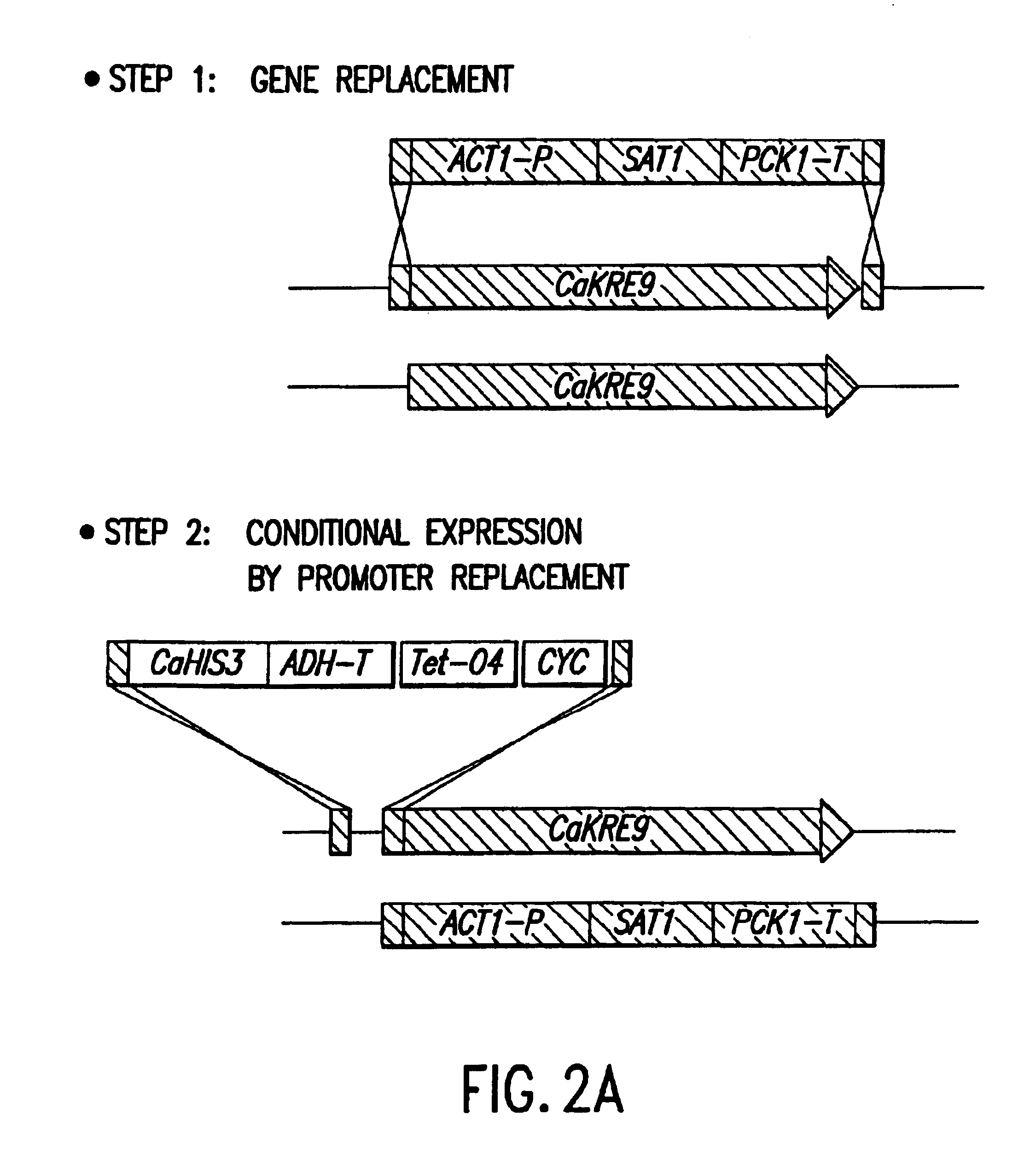
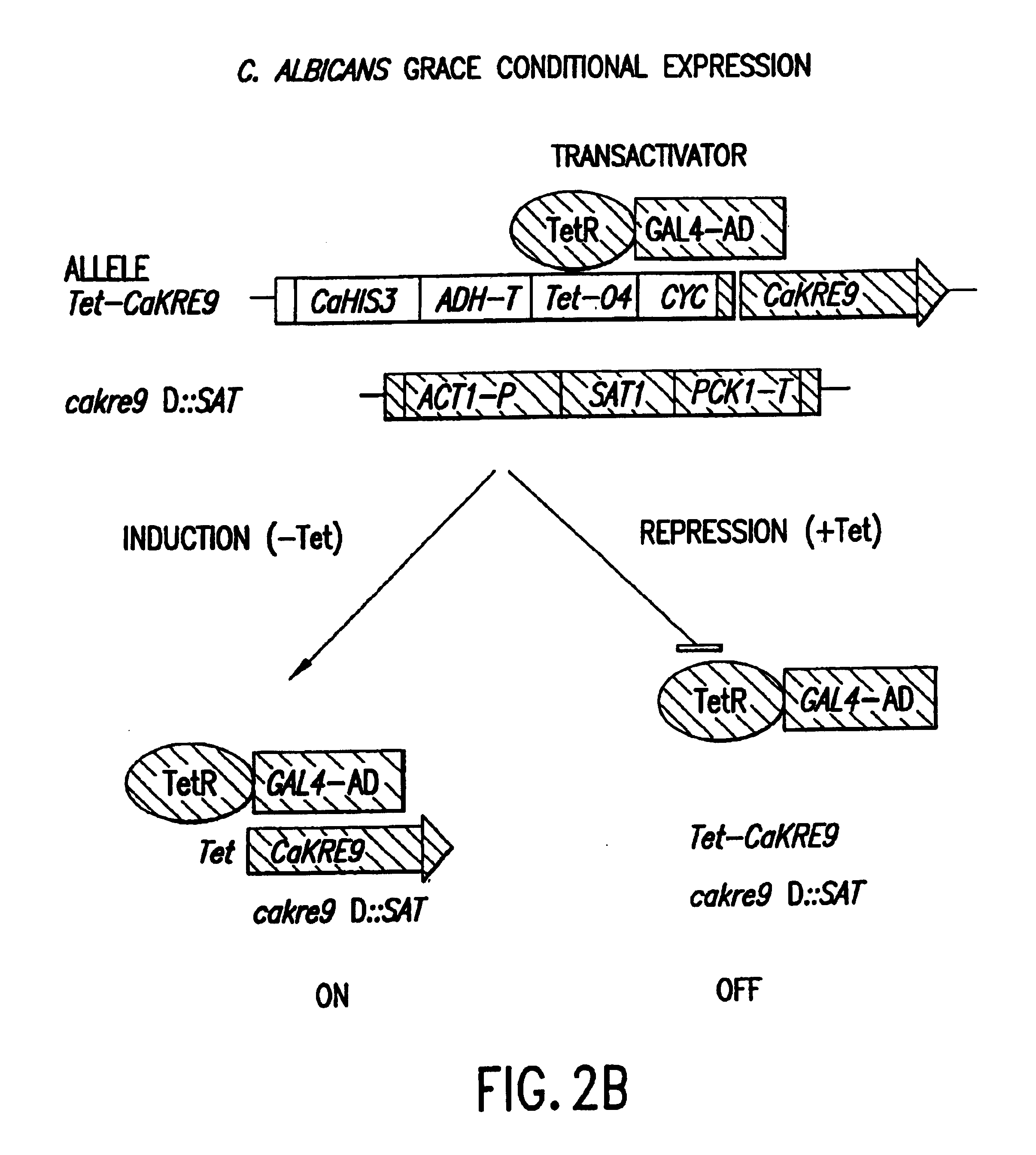
Gene disruption methodologies for drug target discovery
InactiveUS6783985B1Efficient and effectiveAntibody mimetics/scaffoldsMicroorganismsVirulent characteristicsScreening method
The present invention provides methods and compositions that enable the experimental determination as to whether any gene in the genome of a diploid pathogenic organism is essential, and whether it is required for virulence or pathogenicity. The methods involve the construction of genetic mutants in which one allele of a specific gene is inactivated while the other allele of the gene is placed under conditional expression. The identification of essential genes and those genes critical to the development of virulent infections, provides a basis for the development of screens for new drugs against such pathogenic organisms. The present invention further provides Candida albicans genes that are demonstrated to be essential and are potential targets for drug screening. The nucleotide sequence of the target genes can be used for various drug discovery purposes, such as expression of the recombinant protein, hybridization assay and construction of nucleic acid arrays. The uses of proteins encoded by the essential genes, and genetically engineered cells comprising modified alleles of essential genes in various screening methods are also encompassed by the invention.
Owner:MERCK & CO INC
Antibody for detecting aspergillus
ActiveCN101149377AEasy to operateStrong specificityFused cellsBiological testingPenicillium marneffeiCandida famata
This invention provides the capture antibody and the detection antibody to detect the aspergillus, the two antibody is the single clone antibody of the cross oncocyte CCTCC NO.C200731 and CCTCC NO.C200730, they can unites the glucoprotein whose molecular weight is 25-100kDa in the cell wall of the aspergillus, the reaction with the Penicillium marneffei, white oidiomycosis, smoothness oidiomycosis, tropic oidiomycosis, close smoothness oidiomycosis, Candida krusei are all negative. The antibody in this invention can make into the immunity reagent to detect the aspergillus in common use.
Owner:ZHUJIANG HOSPITAL SOUTHERN MEDICAL UNIV
Combined detection kit for vaginitis
InactiveCN102253206AIncreased sensitivityImprove featuresMaterial analysis by observing effect on chemical indicatorCandida vaginitisBacterial vaginosis
The invention aims to provide a combined detection kit for vaginitis, capable of evaluating vaginal micro-ecological environment, vaginal cleanliness, and pathogens of bacterial vaginosis, Candida vaginitis, trichomonas vaginitis of the tested person simultaneously. Ihe following technical scheme adopted by the invention, a combined detection kit for vaginitis is provided and comprises a reaction device, diluent and chromogenic liquid, the reaction device is a hole type box-shaped reaction device with six reacting blind holes, in which a hydrogen peroxide concentration detection reagent pad, a leukocyte esterase detection reagent pad, a neuraminidase detection reagent pad, a proline aminopeptidase detection reagent pad, an acetyl glucosaminidase detection reagent pad and pH detection test paper are arranged respectively. Through the combined detection, the vaginitis caused by most of the BV (Bacterial Vaginosis) pathogens, the Candida albicans and the trichomonas can be detected, BV missing detection caused by the simple detection for one kind of enzyme activity is avoided largely, and the detection reliability and accuracy are increased.
Owner:泰普生物科学(中国)有限公司
Compositions and methods for detection of Candida species
The invention features compositions and methods for the detection of Candida species in biological samples, such as human tissue samples. The invention features species specific nucleic acid probes that can hybridize to target nucleic acid molecules in Candida species and can be used in a variety of detection assays, such as fluorescence based assays or NMR based assays. The compositions and methods of the invention can be used to rapidly and accurately detect one or more Candida species in patients and can be used to assist in point-of care-decisions for treating Candida infections.
Owner:T2 BIOSYST
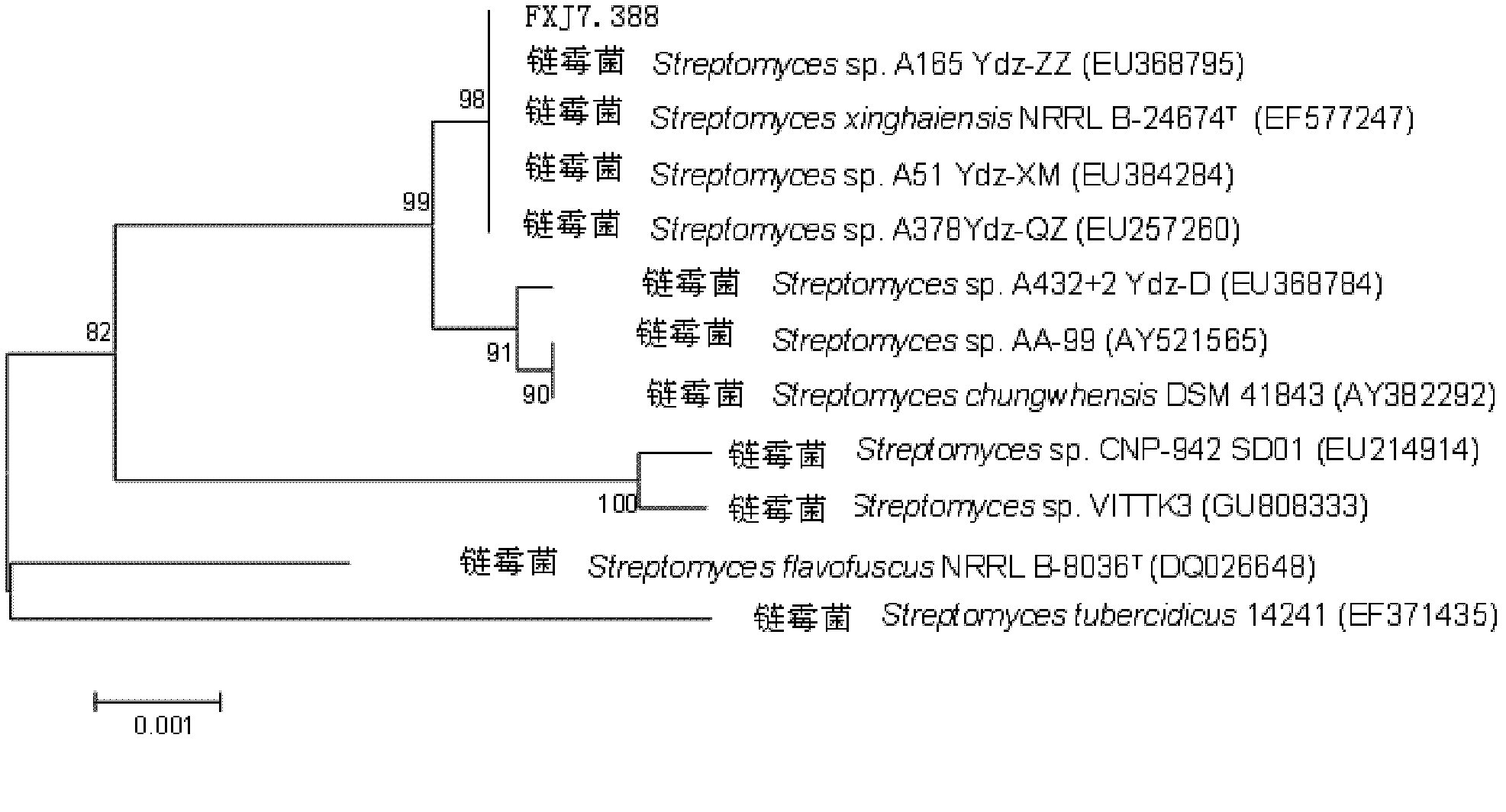
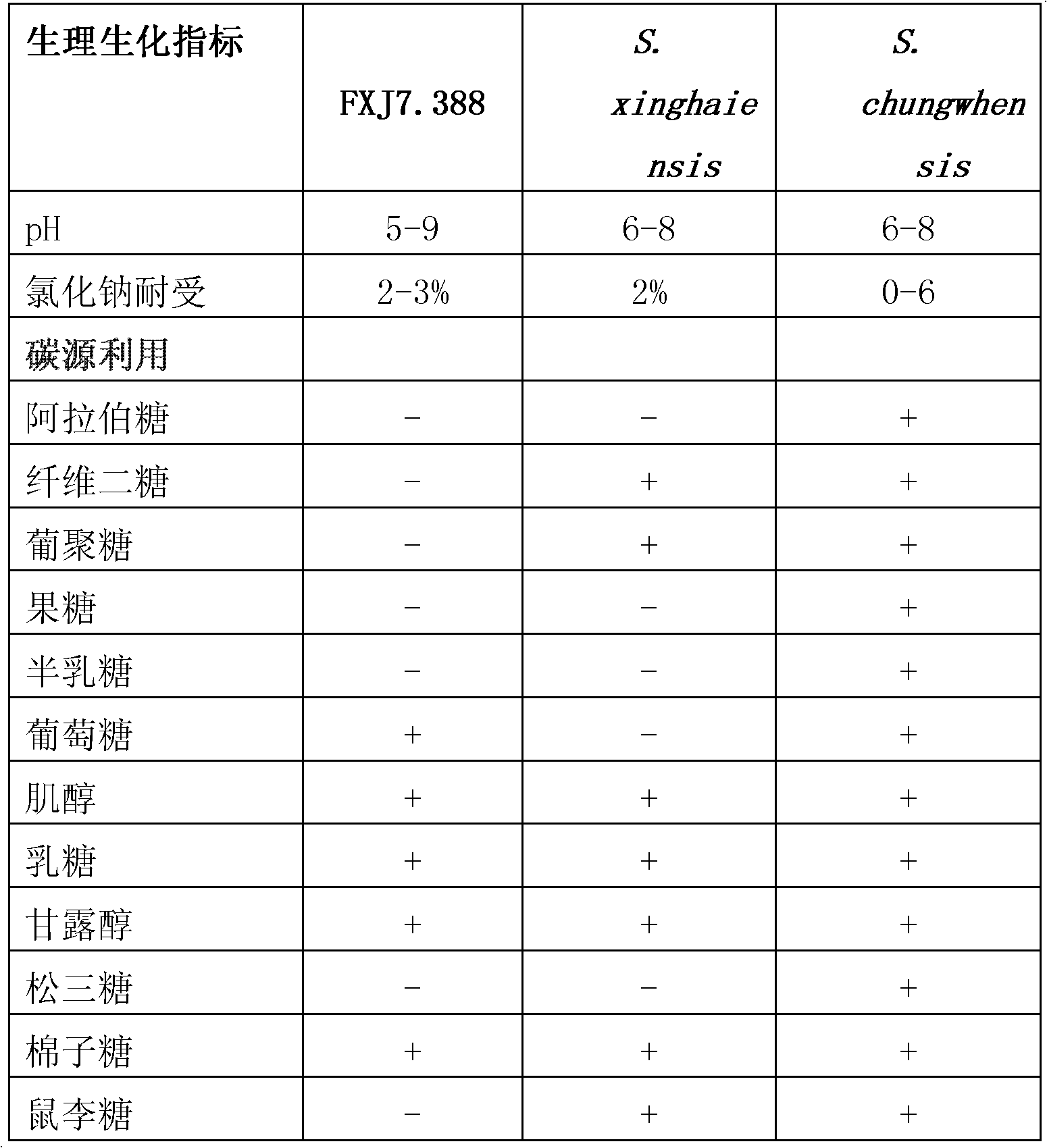
Marine Streptomyces strain and use thereof
InactiveCN102220271AHigh antibacterial activityAntibacterial agentsAntimycoticsCandida famataHigh activity
The invention provides a marine Streptomyces strain and use thereof. The strain provided by the invention is Streptomyces sp. FXJ7.388, and the collection number of the strain is CGMCC No.4474. The invention also provides the use of the strain in the preparation of an antibacterial product. The antibacterial effect is to inhibit gram-positive bacteria, including Multi-resistant Staphylococcus aureus (MRSA 1-1), staphylococcus aureus CGMCC1.2386, bacillus subtilisGCMCC1.2428 and / or Candida albicans 2.538. Experiments prove that the Streptomyces sp. FXJ7.388 separated by the invention has the advantage of resisting gram-positive bacteria and Candida albicans, namely having high activity for resisting MRSA, staphylococcus aureus AND bacillus subtilis and certain inhibiting effect on Candida albicans.
Owner:INST OF MICROBIOLOGY - CHINESE ACAD OF SCI
Primer probe combination for simultaneously detecting four types of candida and kit thereof
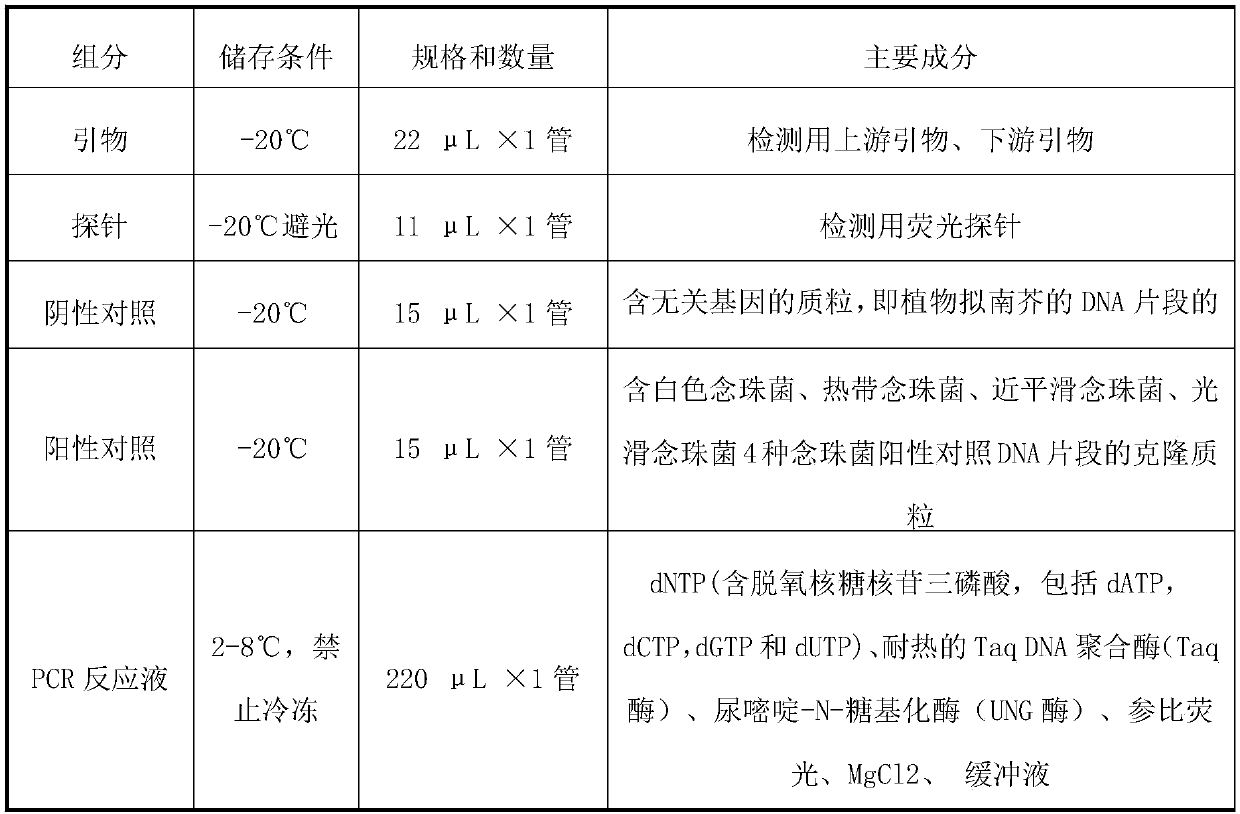
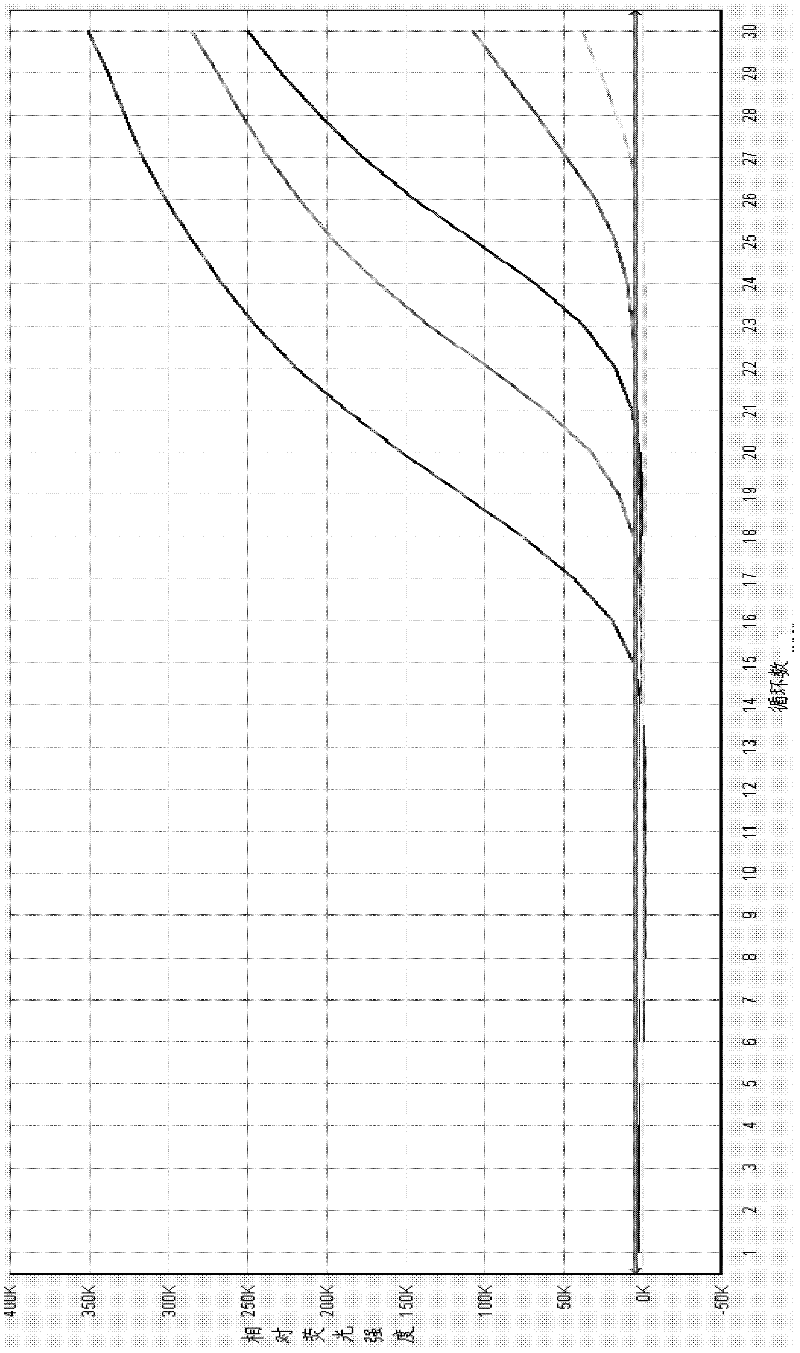
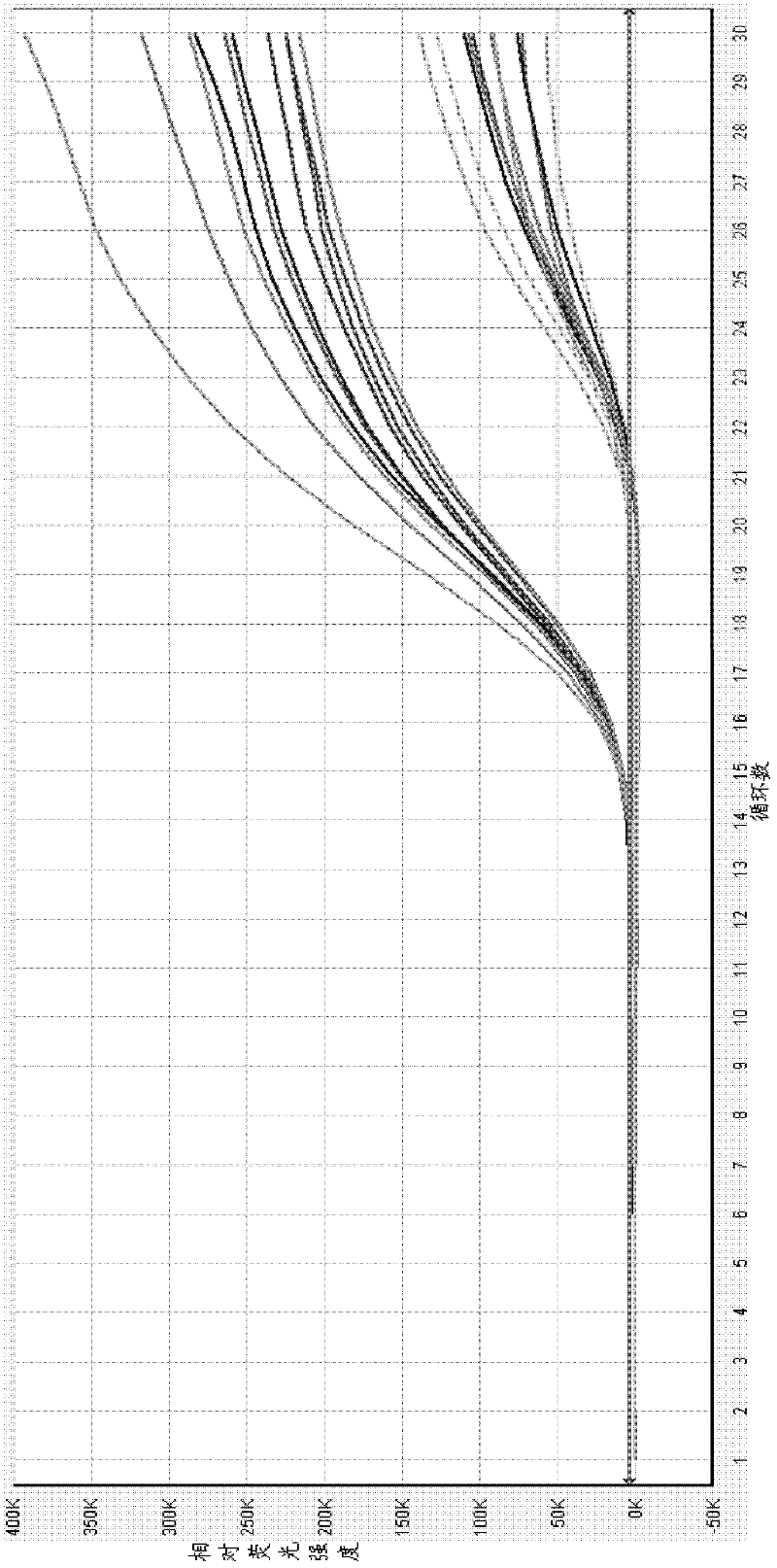
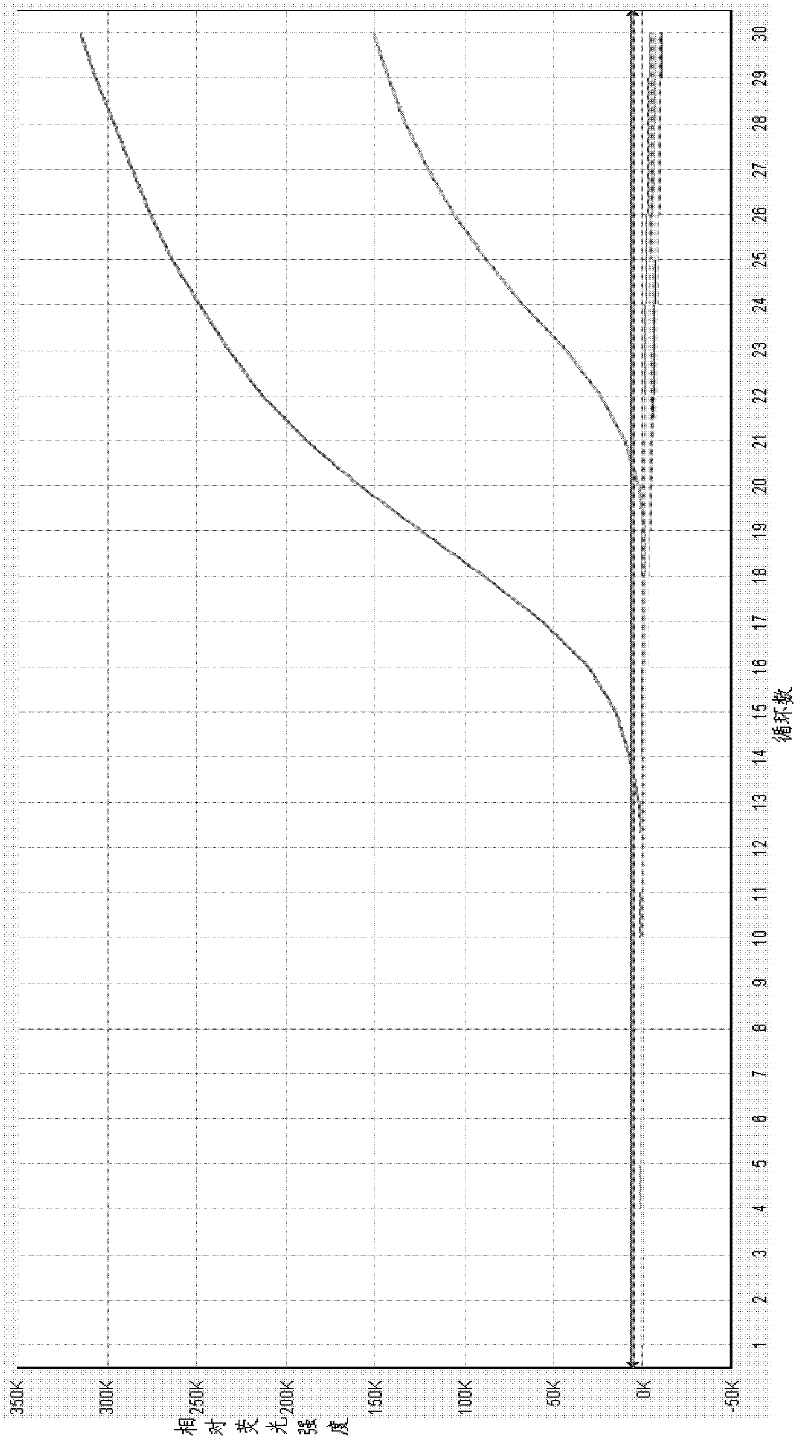
ActiveCN108034745AMicrobiological testing/measurementMicroorganism based processesMicroorganismFluorescence
The invention provides a primer probe combination for simultaneously detecting four types of candida, which belongs to the technical field of microbe detection. The primer probe combination comprisesprimers and a probe, the primer is an upstream primer and a downstream primer, a sequence of the upstream primer is shown as SEQ ID No.1; the sequence of the downstream primer is shown as SEQ ID No.2,and the probe is the sequence labeled with a report fluorescence group and a quenching fluorescence group shown as SEQ ID No.3. The invention provides a pair of primers and a probe which specificallyand simultaneously detect four types of candida such as Candida albicans, Candida tropicalis, candida glabrata and candida parapsilosis.
Owner:杭州缔蓝生物技术有限公司
Insect antimicrobial peptide Thanatin derivant, producing method and uses of the same
InactiveCN101173005AIncrease varietyHigh antibacterial activityAntibacterial agentsAntimycoticsK pneumoniaeHemolysis
The invention relates to a group of insect Thanatin ramifications with antibacterial action and the salt which is applicable for medicinal use, as well as a preparation method and the use thereof. Approved by the results of a sterilization experiment, a hemolysis experiment and an irritable partial stimulation experiment, the insect Thanatin ramification has bigger antibacterial activity than Thanatin to colon bacillus, Candida albicans, Klebsiella pneumoniae, and staphylococcus aureus. The hemolysis reaction is not incurred when the concentration is 2.5mg / ml without acute toxicity stimulation reaction.
Owner:沈子龙
Anticmicrobial denture cleaning compositions
A denture cleansing composition includes a monoperoxysulfate compound, an effective amount of a sequestering agent, such as a citrate compound, for removal of calculus and to provide a pH to the composition in solution (water) of about 3 to 5, and an effective amount of an antimicrobial agent, such as a benzoate compound, to provide antimicrobial activity to the composition to effectively kill bacteria, or other microorganisms found on the dentures. Tests conducted show that the composition is particularly effective in killing microbial strains of Streptococcus mutans, Streptococcus pyogenes, Candida albicans and Actinomyces viscosus within 20 minutes of contact.
Owner:PROTECH PROFESSIONAL PROD +1
Polymerase chain reaction (PCR) fluorescence detection kit and detection method for candida albicans
ActiveCN102417931AEasy extractionEasy to operateMicrobiological testing/measurementMicroorganism based processesFluorescenceNucleotide
The invention relates to the field of microbe detection, in particular to a polymerase chain reaction (PCR) fluorescence detection kit for candida albicans. According to the technical scheme, the fluorescence detection kit for the candida albicans comprises a PCR reaction solution; the PCR reaction solution comprises primers A and a fluorescence probe A; the primers are divided into an upstream primer A and a downstream primer A; and the nucleotide sequence of the upstream primer A is shown as SEQ ID NO.1, the nucleotide sequence of the downstream primer A is shown as SEQ ID NO.2, and the nucleotide sequence of the fluorescence probe A is shown as SEQ ID NO.3. The kit has the advantages of high sensitivity and specificity, stability, timeliness, convenience for operation and the like.
Owner:泰普生物科学(中国)有限公司
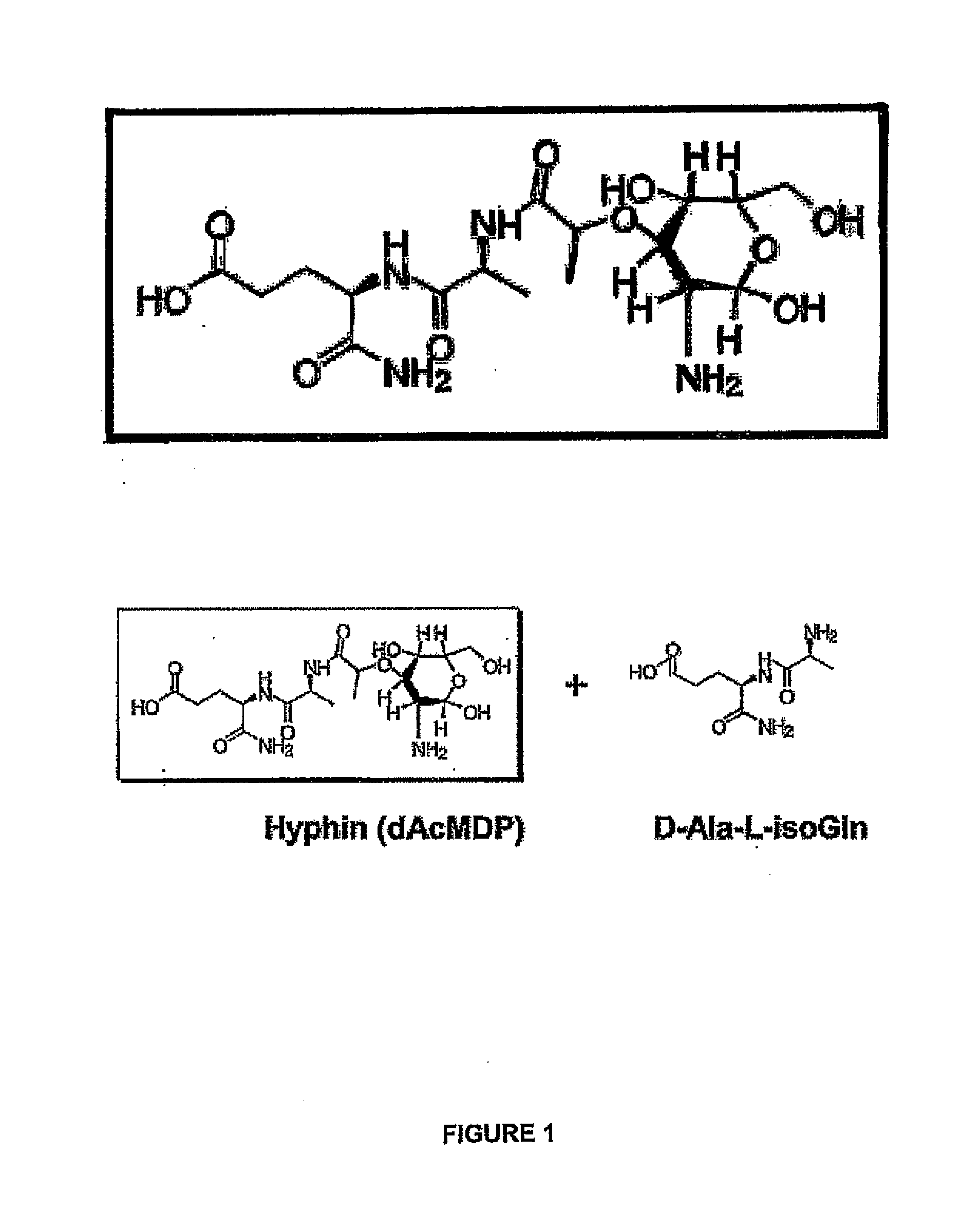
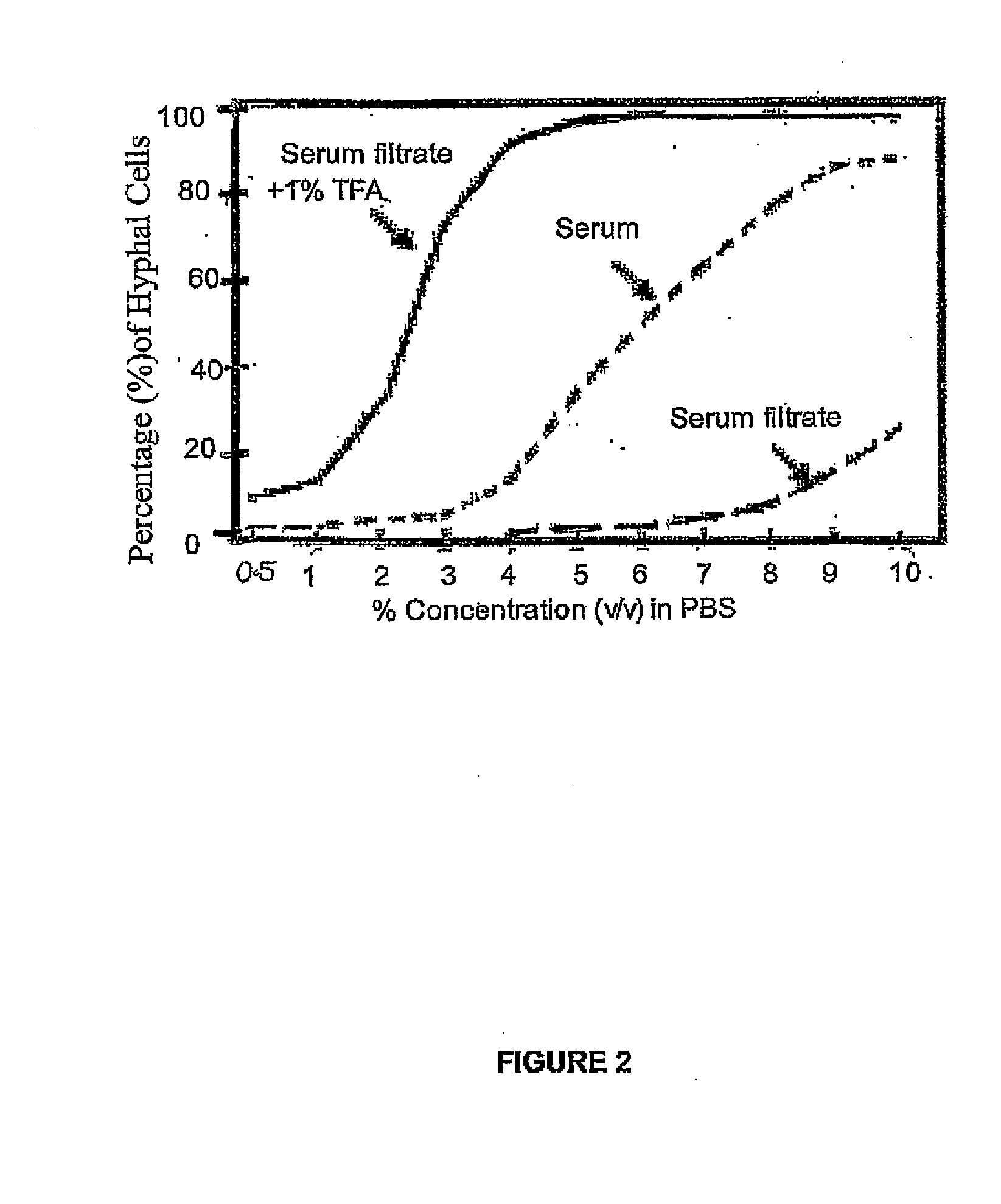
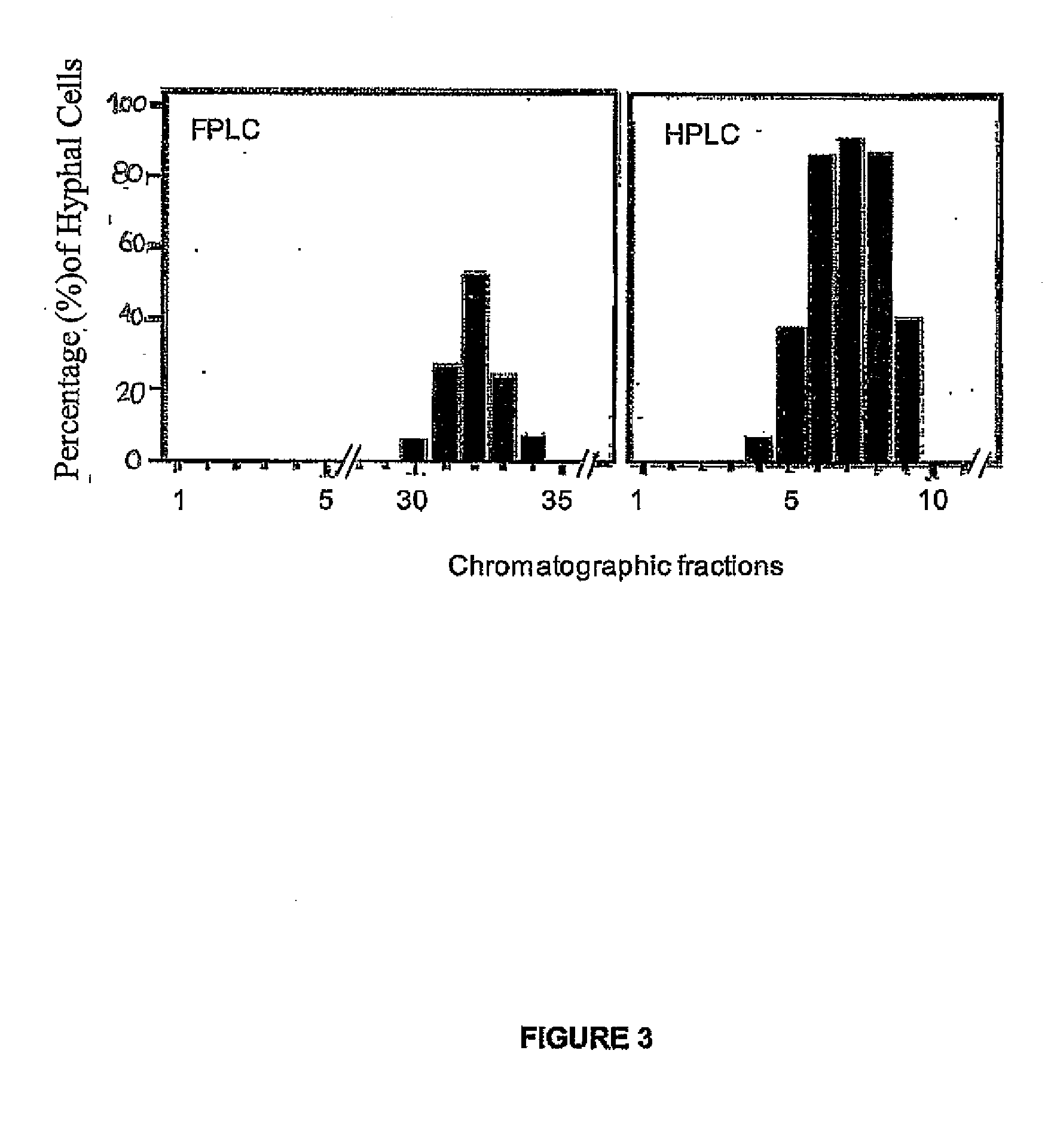
Modulators of Candida Hyphal Morphogenesis and Uses Thereof
The invention relates to modulation of fungal morphology between yeast-to-hyphal growth transition by controlling muramyl-L-alanine concentration and uses thereof.
Owner:AGENCY FOR SCI TECH & RES
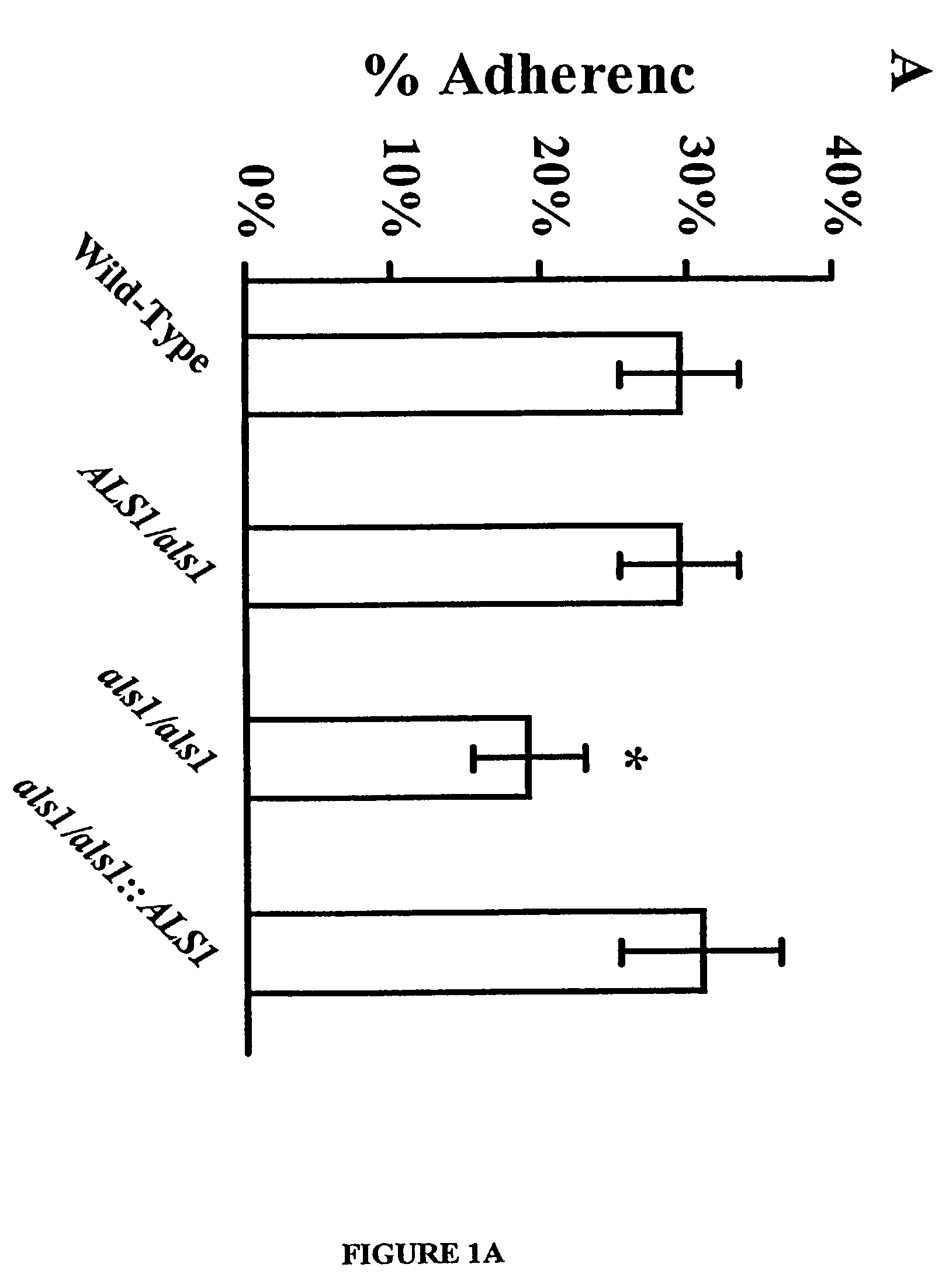
Pharmaceutical compositions and methods to vaccinate against candidiasis
InactiveUS8541008B2Toxic reductionReduce adhesionPeptide/protein ingredientsSnake antigen ingredientsHospitalized patientsVirulent characteristics
A Candida albicans bloodstream infections cause significant morbidity and mortality in hospitalized patients. Filament formation and adherence to host cells are critical virulence factors of C. albicans. Multiple filamentation regulatory pathways have been discovered, however the downstream effectors of these regulatory pathways remain unknown. The cell surface proteins in the ALS group are downstream effectors of the filamentation regulatory pathway. Particularly, Als1p mediates adherence to endothelial cells in vitro and is required for virulence. The blocking of adherence by the organism is described resulting from the use of a composition and method disclosed herein. Specifically, a pharmaceutical composition comprised of a gene, gene product, or specific antibody to the ALS gene family is administered as a vaccine to generate an immune response capable of blocking adherence of the organism.
Owner:LOS ANGELES BIOMEDICAL RES INST AT UCLA HARBOR MEDICAL CENT +1
Triple nucleic acid test kit
ActiveCN106319055AHigh similarityPrimer design is difficultMicrobiological testing/measurementMicroorganism based processesNucleic acid testCandida famata
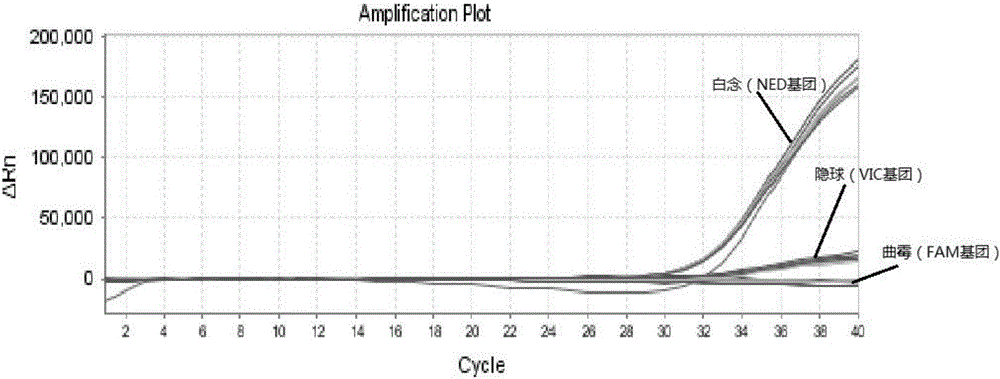
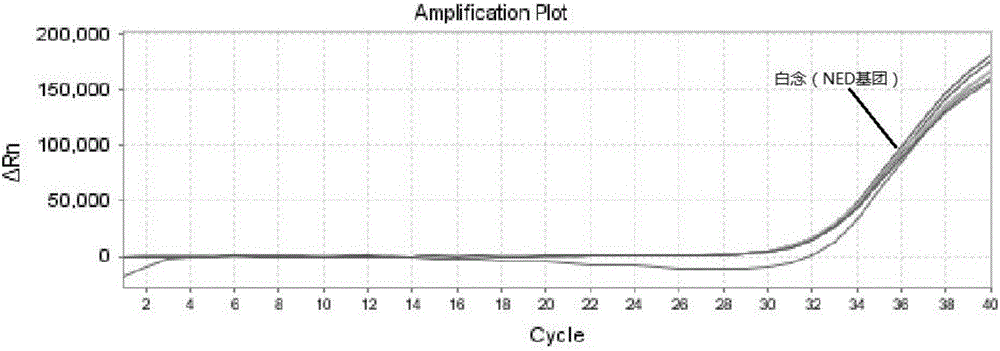
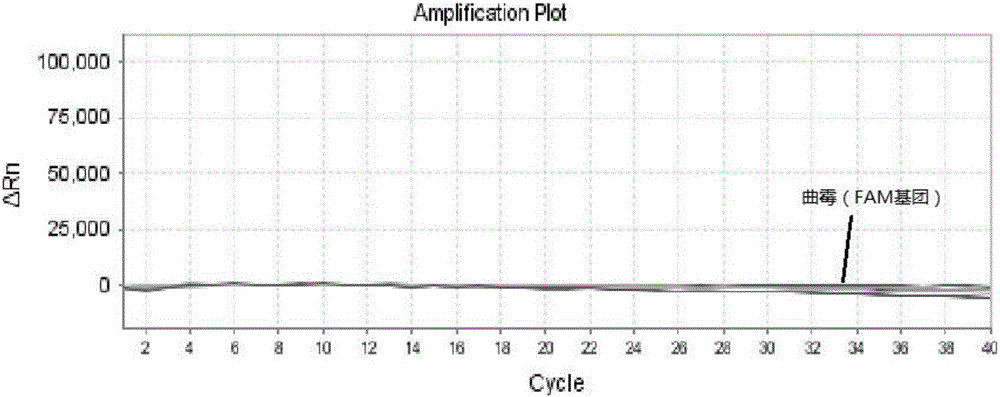
The invention relates to a triple nucleic acid test kit. The kit comprises a thallus lysate, a positive quantitative reaction liquid, a negative control reaction liquid and a thallus test reaction liquid, wherein the thallus test reaction liquid contains specific primers, specific probes and doping fluorescent dye for aspergillus, Cryptococcus neoformans and candida albicans, the three specific oprimers are designed according to areas of ITS sequences of aspergillus, Cryptococcus neoformans and candida albicans, FAM, VIC and CY-5 are selected for mark report groups of the three specific probes, MGB serves as a quenching group, the excitation wavelength of the FAM group is 492 nm, and the emission wavelength is 517 nm; the excitation wavelength of the VIC group is 557 nm and the emission wavelength is 574 nm; the excitation wavelength of the CY-5 group is 646 nm and the emission wavelength is 664 nm. The triple nucleic acid test kit for aspergillus, Cryptococcus neoformans and candida albicans can assist in diagnosing infection caused by aspergillus, Cryptococcus neoformans and candida albicans.
Owner:BEIHAI SINLON BIOTECH CO LTD
Recombinant Bacillus subtilis for efficiently synthesizing acetylglucosamine
ActiveCN106479945AEasy to buildEasy to useBacteriaMicroorganism based processesCandida famataGenetic engineering
The invention discloses recombinant Bacillus subtilis for efficiently synthesizing acetylglucosamine and belongs to the field of genetic engineering. Recombinant Bacillus subtilis BSGN6-PxylA-glmS is used as a starting strain, glucosamine synthetic route is enhanced by overexpressing glucosamine acetylase coding gene (CaGNA1) from Candida albicans SC5314 or knocking out unnecessary area skin at the same time, and accumulated acetylglucosamine Bacillus subtilis genetically-engineered bacteria are obtained; the yield is 21% or 94% higher than that of CaGNA1 genetically-engineered bacteria from overexpressed Saccharomyces cerevisiae, and the basis is laid for the further metabolic engineering modification of Bacillus subtilis.
Owner:SHANDONG RUNDE BIOTECH CO LTD
Cytochrome P450 monooxygenase CYP52A2B from Candida tropicalis
InactiveUS7049112B2Increase productionHigh copy numberBacteriaSugar derivativesCandida tropicalisCandida famata
Novel genes have been isolated which encode cytochrome P450 and NADPH reductase enzymes of the ω-hydroxylase complex of C. tropicalis 20336. Vectors including these genes, transfected host cells and transformed host cells are provided. Methods of producing of cytochrome P450 and NADPH reductase enzymes are also provided which involve transforming a host cell with a gene encoding these enzymes and culturing the cells. Methods of increasing the production of a dicarboxylic acid and methods of increasing production of the aforementioned enzymes are also provided which involve increasing in the host cell the number of genes encoding these enzymes. A method for discriminating members of a gene family by quantifying the expression of genes is also provided.
Owner:COGNIS IP MANAGEMENT GMBH
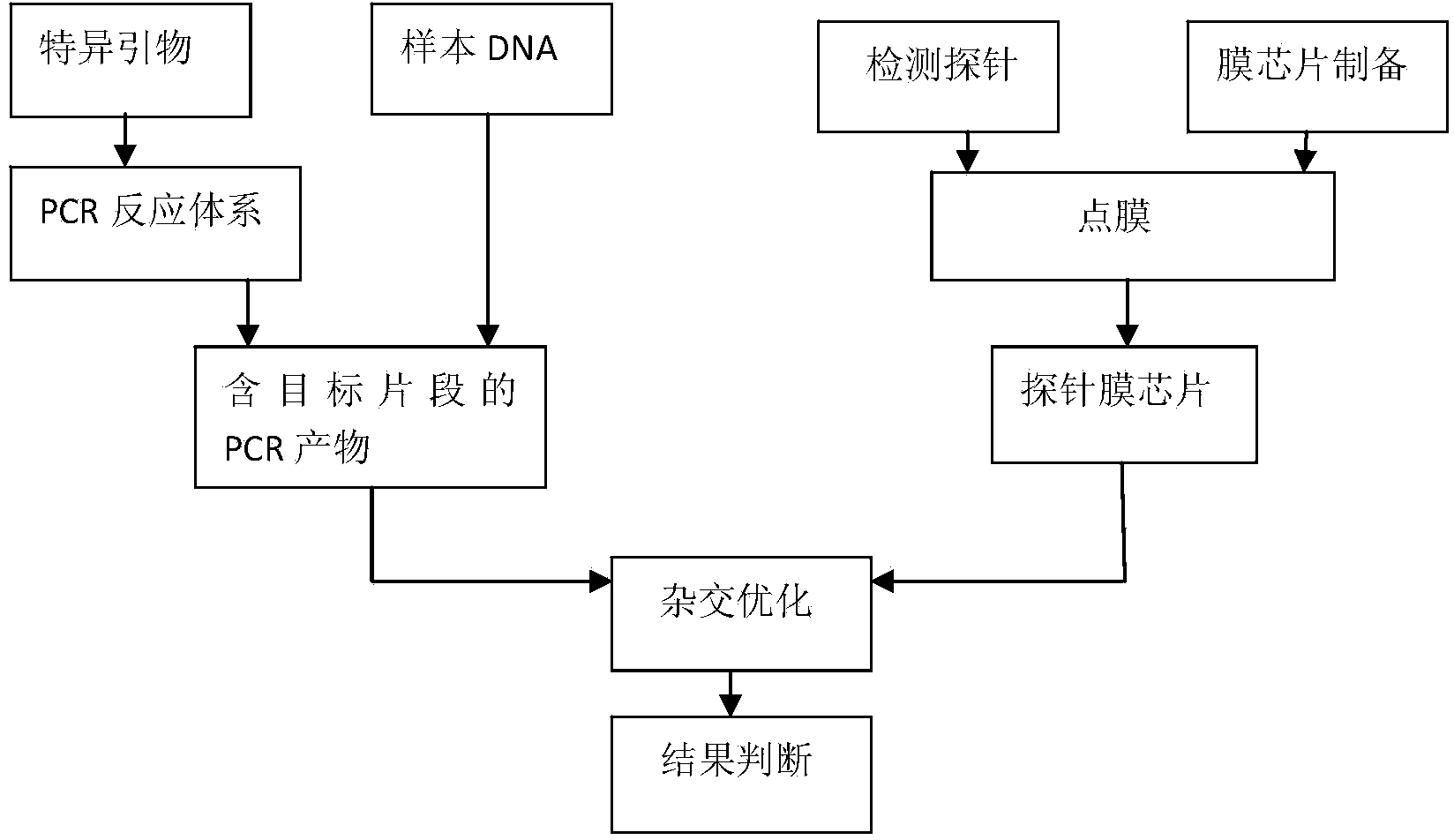
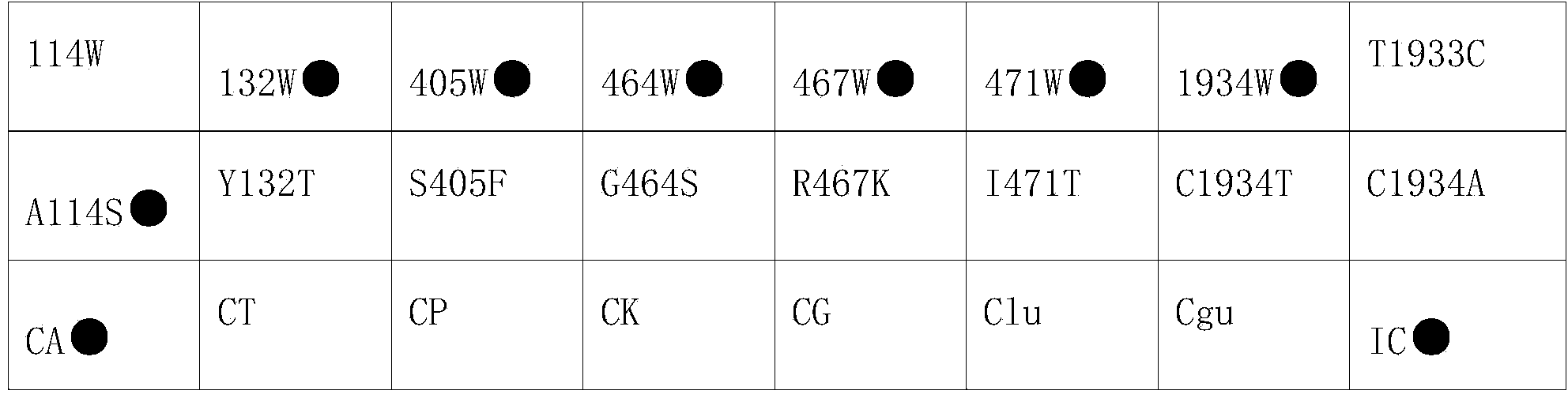
Nucleic acid film strip and kit for identification of candida species and detection of gene mutation
ActiveCN104263833AReliable resultsHigh detection sensitivityMicrobiological testing/measurementMicroorganism based processesNucleic Acid ProbesCandida famata
The invention relates to a diagnostic reagent of candida and particularly relates to a nucleic acid film strip and a kit capable of simultaneously performing identification of candida species and detection of gene mutation. The nucleic acid film strip for identification of candida species and detection of gene mutation comprises a substrate and nucleic acid probes immobilized on the substrate, wherein the nucleic acid probes comprise a nucleic acid probe used for identification of candida species and having base sequences shown in SEQ ID NO: 1-7, a nucleic acid probe used for detection of gene mutation of candida albicans and having base sequences shown in SEQ ID NO: 8-23 and an internal reference nucleic acid probe having a base sequence shown in SEQ ID NO: 24. The nucleic acid film strip and the kit are used for identification of the species and detection of drug-resistant mutant genes according to the gene sequences of candida, judging from molecular structure, results are accurate and reliable and the detection sensitivity is high; the detection time is short and the detection can be completed only just in eight hours.
Owner:杭州美联医学股份有限公司 +2
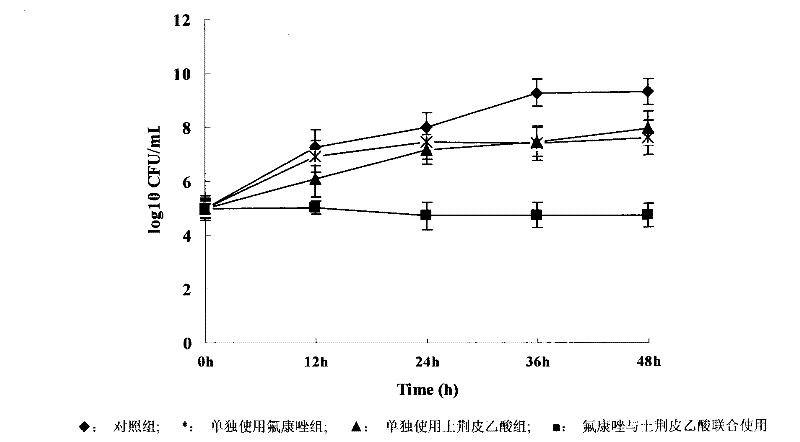
A synergist with synergistic antifungal effect with fluconazole
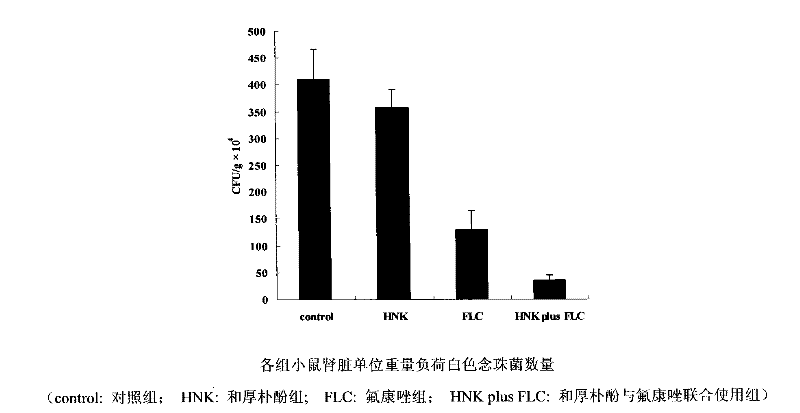
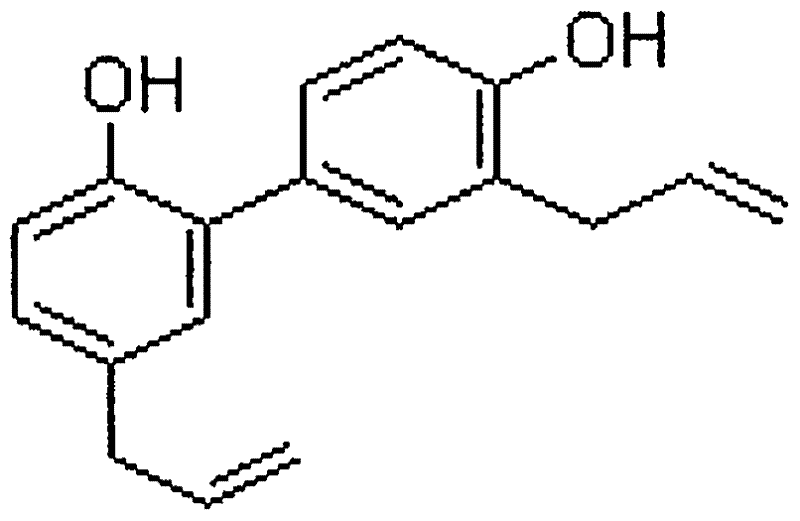
The invention discloses a novel application of honokiol and fluconazole in antifungal cooperation. According to the invention, the fungi are candida albicans. According to the invention, a standard macrodilution drug sensitivity experiment recommended by CLSI is adopted, in vitro sensitivity of honokiol towards clinically separated drug resistant candida albicans is evaluated with systematic methods such as a checkerboard macrodilution method and a time-kill curve method. Also, an in vitro synergistic effect of honokiol and fluconazole in clinic drug resistant fungi resistance is evaluated. Further, with a mice infection and treatment experiment, an in vivo synergistic effect of honokiol and fluconazole in clinic drug resistant fungi resistance is evaluated.
Owner:JILIN UNIV