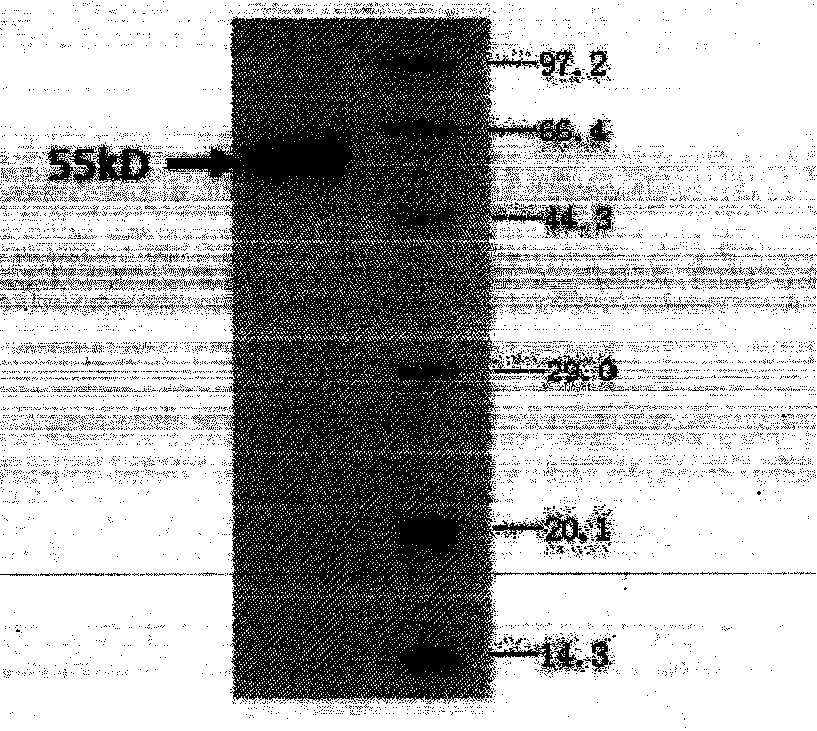Patents
Literature
147 results about "Specific identification" patented technology
Efficacy Topic
Property
Owner
Technical Advancement
Application Domain
Technology Topic
Technology Field Word
Patent Country/Region
Patent Type
Patent Status
Application Year
Inventor
Specific identification is a method of finding out ending inventory cost. It requires a detailed physical count, so that the company knows exactly how many of each goods brought on specific dates remained at year end inventory.
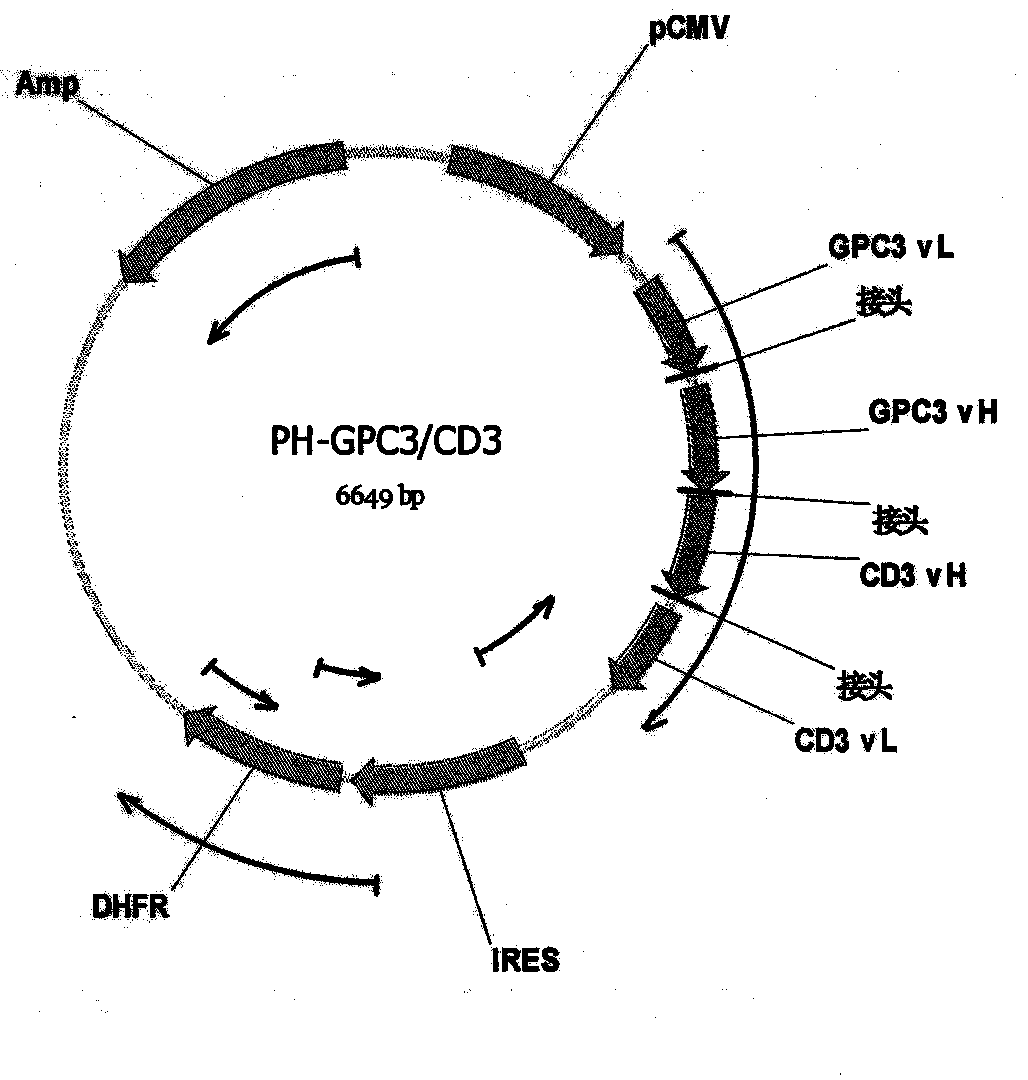
Bispecific antibody aiming at phosphatidylinositols protein polysaccharide-3 and T cell antigen
InactiveCN103833852ABacteriaImmunoglobulins against cell receptors/antigens/surface-determinantsAntigenNucleotide
The first aspect of the invention relates to a bispecific antibody, which comprises a first functional domain for specific identification of phosphatidylinositol protein polysaccharide-3, a second domain for specific identification of human T cell antigen CD3, and a connection for connecting the functional domains. The second aspect of the invention relates to a nucleotide sequence encoding the above antibody. The third aspect of the invention relates to a carrier containing the above nucleotide sequence, and includes an expressive vector. The fourth aspect of the invention relates to a eukaryotic or prokaryotic expression system containing the above carrier. The fifth aspect of the invention relates to application of the above antibody to preparation of medicament for treating or preventing tumor.
Owner:SHANGHAI INST OF ONCOLOGY
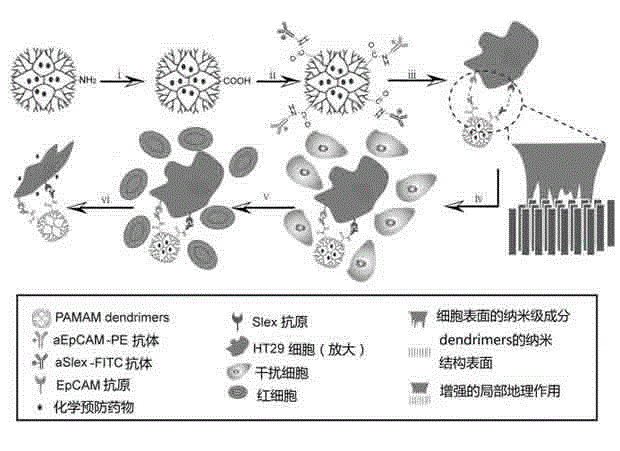
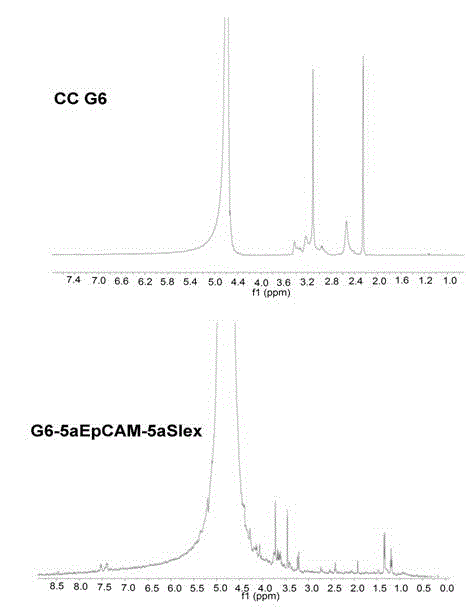
Functional nano material drug delivery system for identifying, capturing and restraining circulating tumor cells
InactiveCN104353082AEasy to prepareEasy to operateOrganic active ingredientsPowder deliveryCancer preventionCancer metastasis
The invention belongs to the field of application of a nano material coating technology in identifying, capturing and activity regulating of circulating tumor cells. For pre-warning and preventing of cancer metastasis, particularly, the invention relates to a functional nano material drug delivery system for identifying, capturing and restraining circulating tumor cells. The functional nano material drug delivery system consists of a central nano material carrier, a surface targeted antibody or an aptamer, and a drug for resisting cancers or preventing the cancer metastasis. The functional nano material drug delivery system disclosed by the invention can be used for in-vitro simulation or specific identification and capturing research of trace circulating tumor cells of a blood sample of a clinical patient, and also can be used for regulating the activities of the captured circulating tumor cells, so that the application prospect in pre-warning and preventing of the cancer metastasis is expanded.
Owner:FUZHOU UNIV
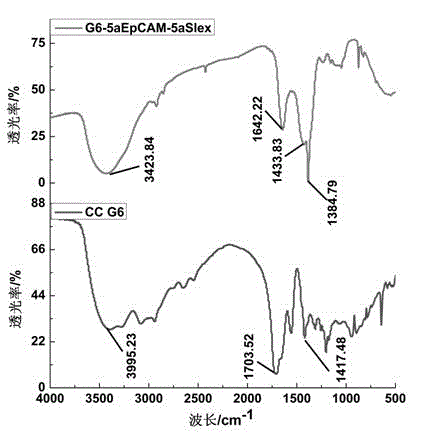
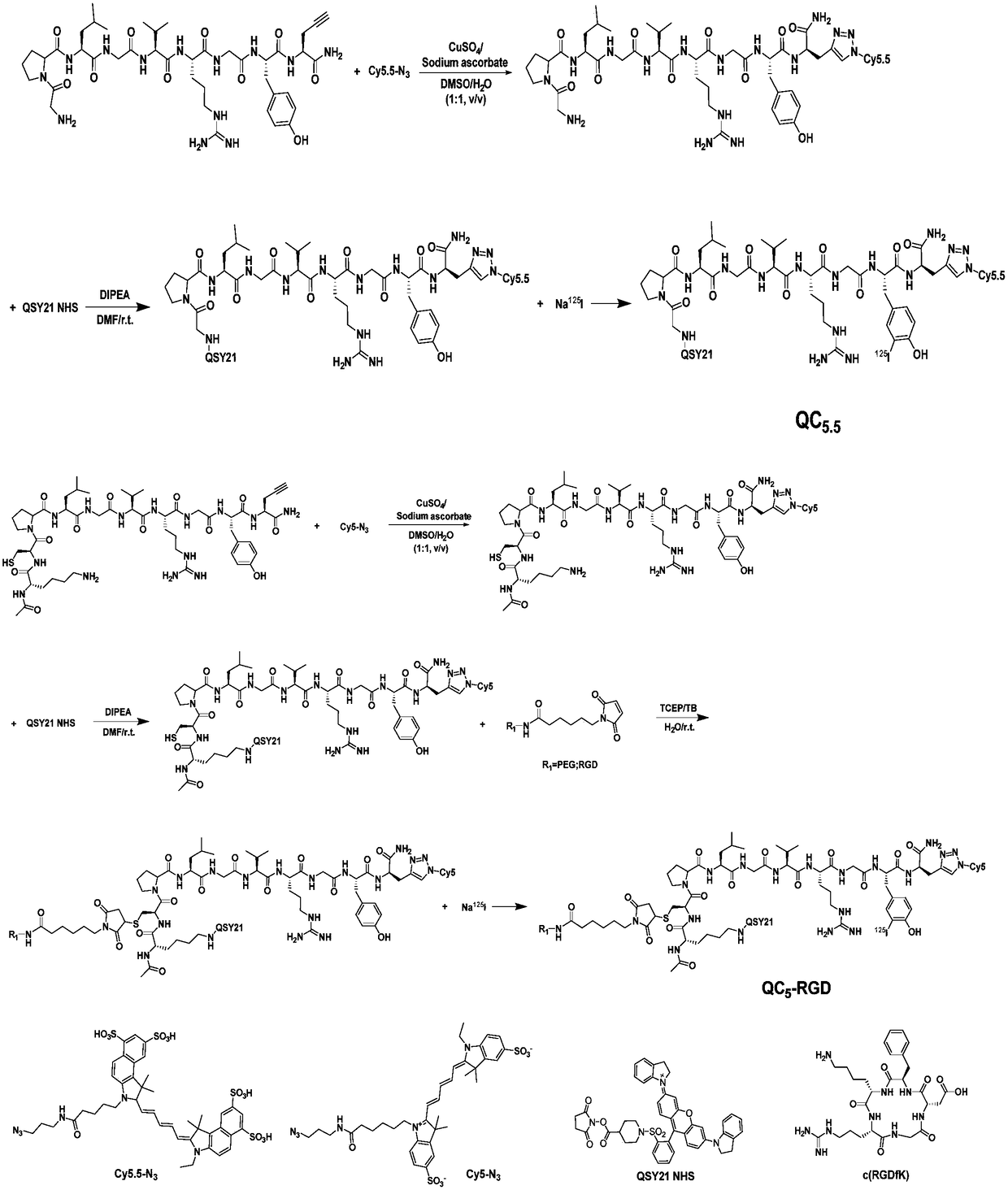
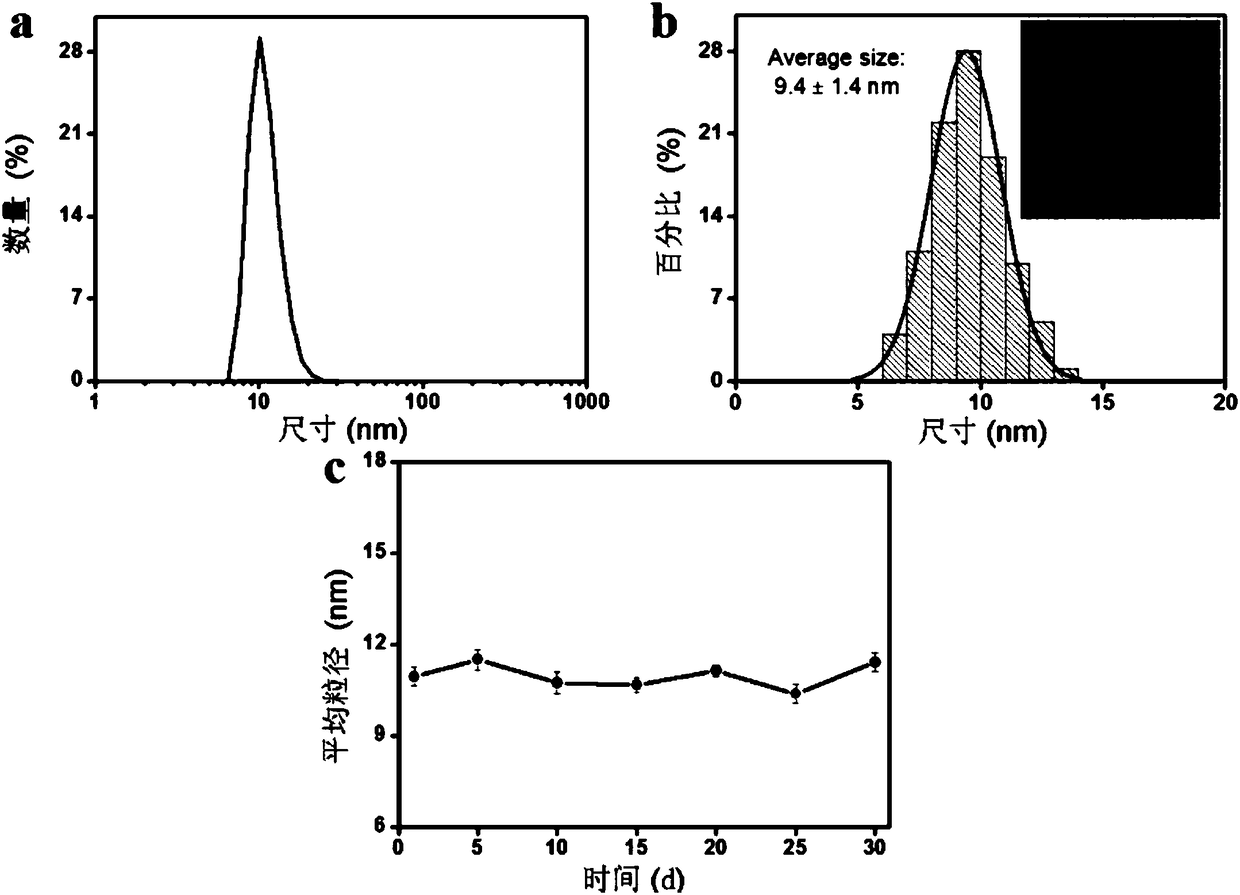
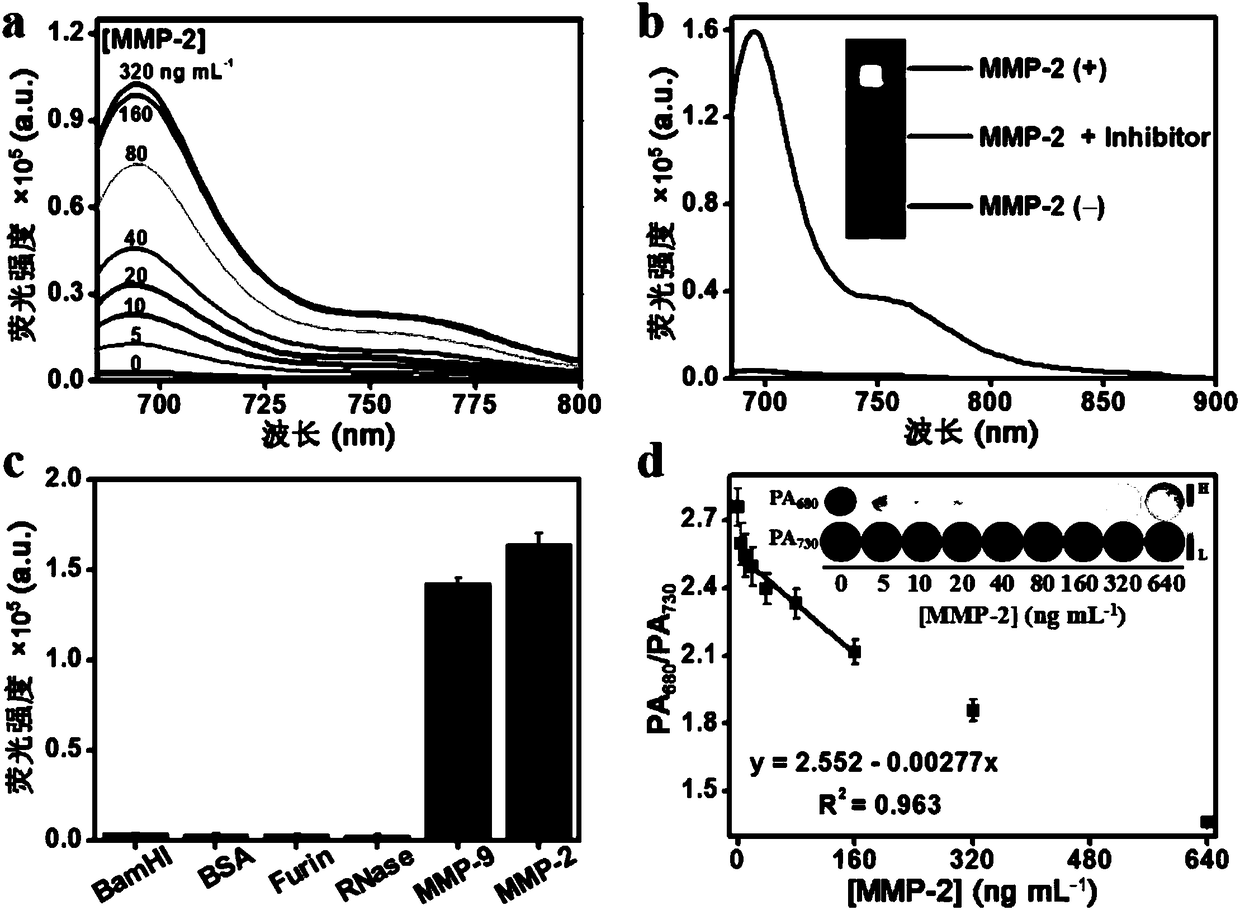
Matrix metalloproteinase-2 specific multi-modality molecular image probe and preparation method and application in preparation of tumor imaging agent thereof
ActiveCN108514647AImprove targetingImprove featuresGeneral/multifunctional contrast agentsEchographic/ultrasound-imaging preparationsSide chainTumor targeting
The invention discloses a matrix metalloproteinase-2 specific multi-modality molecular image probe preparation method and application thereof. The preparation method comprises the following steps: 1)preparing peptide substrates for matrix metalloproteinase-2(MMP-2) specific identification; 2) modifying near infrared fluorescent dye on peptide substrate; 3) modifying fluorescent quenching group onpeptide substrate; 4) connecting different molecular weight PEG or tumor targeting group RGD on peptide substrate modified with near infrared fluorescent dye and fluorescent quenching group; 5) labeling the nuclide on the above modified peptide substrate side chain tyrosine. Compared with the prior art, the matrix metalloproteinase-2 specific multi-modality molecular image probe preparation method has the advantages that: 1) by utilizing the specificity and responsiveness of the MMP-2 protease by the probe, so that the probe is selectively enriched at the tumor site so as to improve the targeting property of the probe to the tumor; 2) performing multi-modality imaging in a living body tumor by using advanced molecular imaging technology such as optical, opto-acoustic, SPECT and the like,improving the sensitivity and accuracy of tumor imaging, and finally achieving accurate positioning of the tumor; 3) providing a new train of thought and method for early diagnosis, process research and prognosis evaluation of tumor.
Owner:苏州智影特生物医药技术有限公司
QCM detection method for detecting lysozyme based on multiple signal amplification technologies and application
InactiveCN107419005AEnhanced signalEasy to operateMicrobiological testing/measurementAptamerBiotin-streptavidin complex
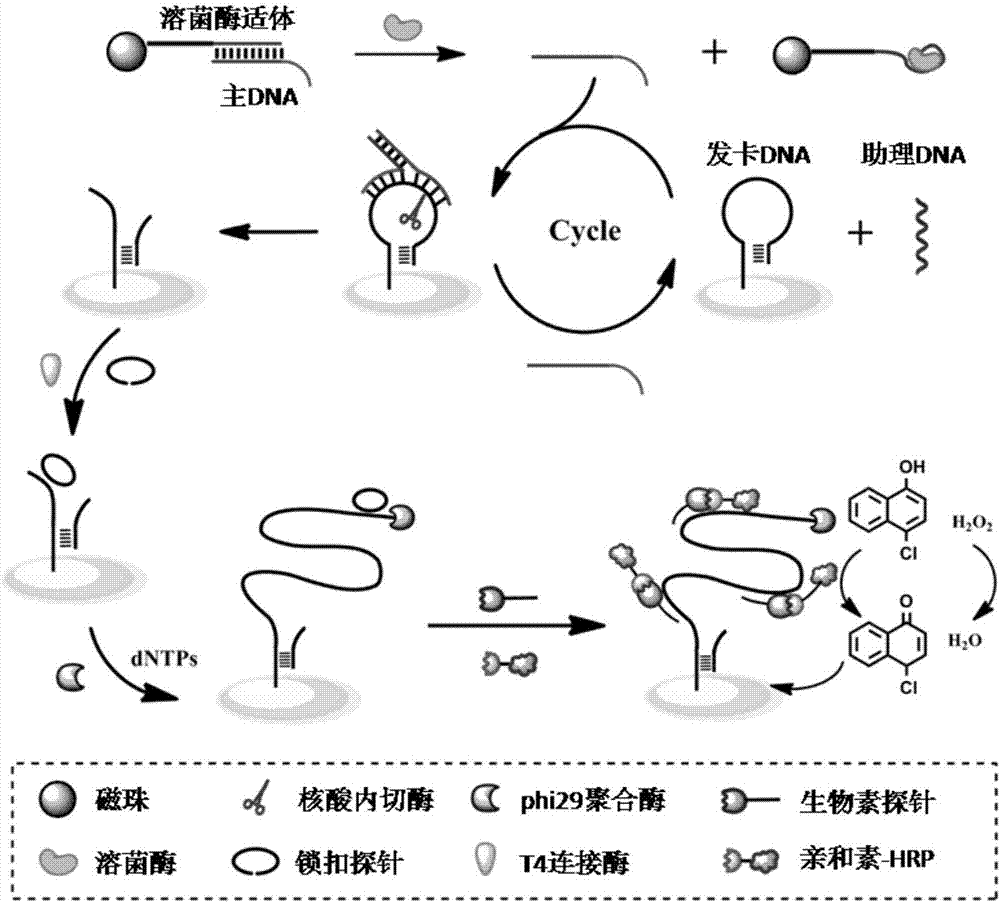
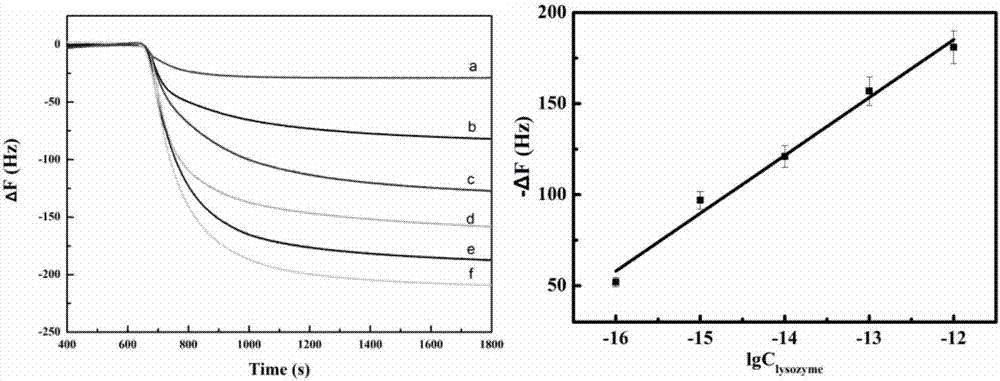
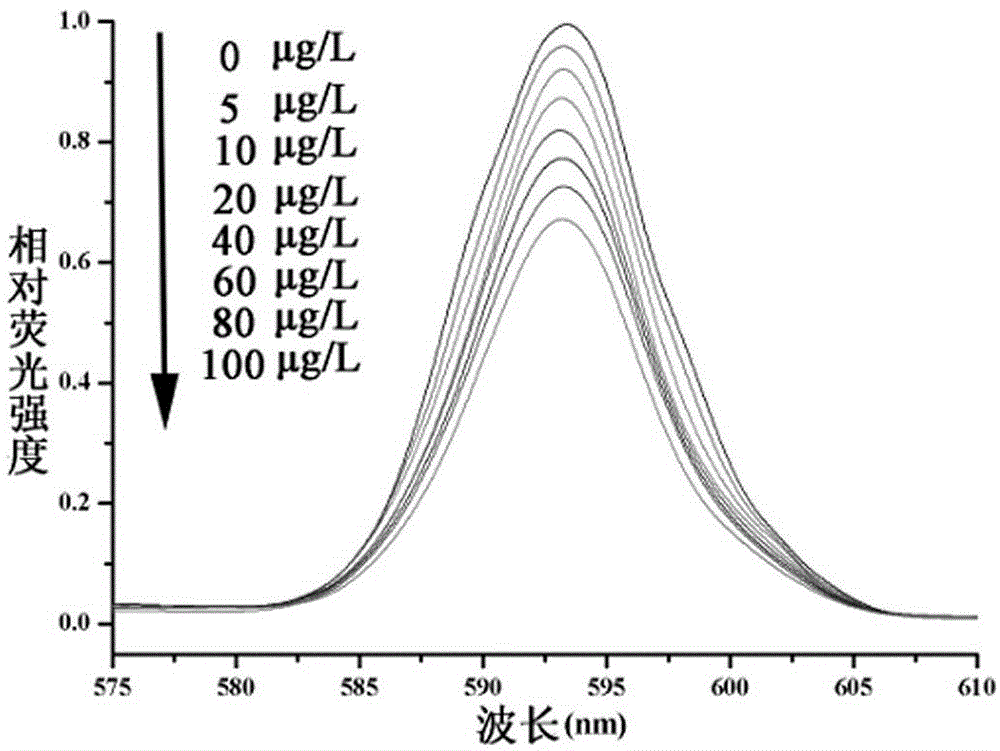
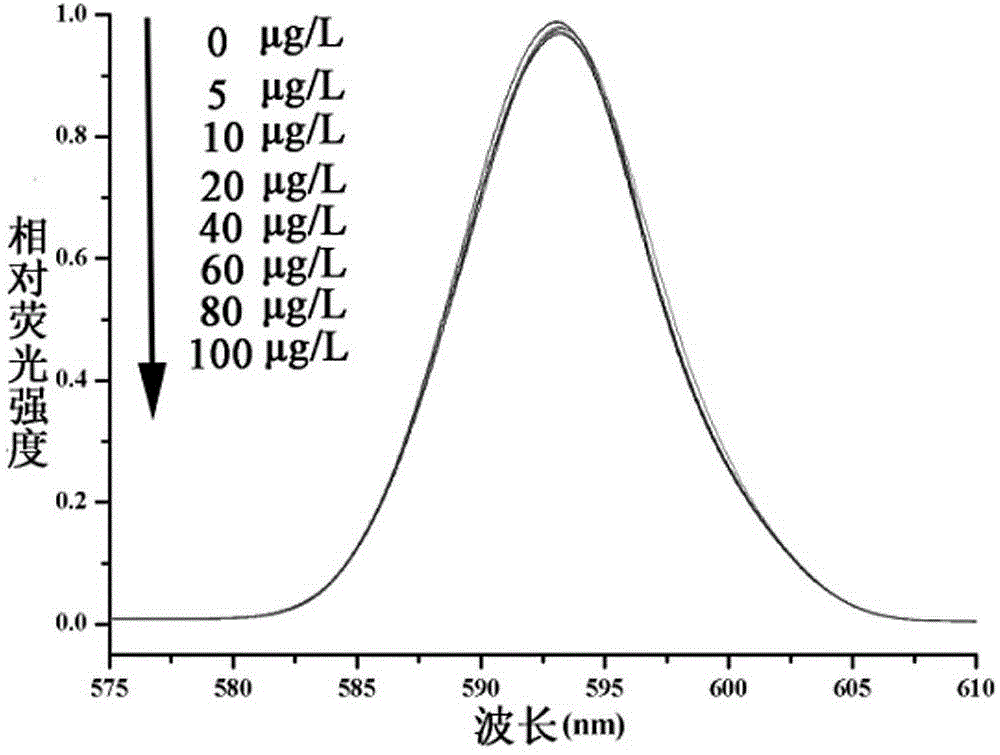
The invention discloses a QCM detection method for detecting lysozyme based on multiple signal amplification technologies and application. The QCM detection method comprises the following steps that DNA hybridizes with lysozyme aptamer partially in a complementary mode, and through the specific binding reaction of the lysozyme and the lysozyme aptamer, the DNA is released; a Y-shaped structure is formed by complementary hybrid of the released DNA with hairpin DNA and assistant DNA modified on a gold leaf, and under the action of restriction enzyme, the hairpin DNA is cut and opened through specific identification sites; under the action of DNA ligase and DNA polymerase, using locking-ring-shaped DNA as a template chain, polymerization growing along the opened hairpin DNA is carried out, and a single chain with a large number of repeated sequences is formed; a signal probe marked with biotin hybridizes with the generated repeated sequences in a complementary mode, and after binding with streptavidin marked by HRP, hydrogen peroxide is catalyzed to oxidize 4-chloro naphthol, and precipitation reaction is generated; and accordingly the chip surface quality is increased, and the high-sensitivity detection of the QCM to the lysozyme is realized.
Owner:QINGDAO UNIV
Production method of clenbuterol molecularly imprinted-upconversion luminescent material fluorescence probe
ActiveCN104818025AHigh selectivityIncreased sensitivityFluorescence/phosphorescenceLuminescent compositionsFluoProbesMolecularly imprinted polymer
The invention relates to a production method of a clenbuterol molecularly imprinted-upconversion luminescent material fluorescence probe. The probe is a novel fluorescence probe produced through combining an upconversion luminescent material YF3:Yb<3+>, Er<3+> as a carrier with a covalent bond technology prepared clenbuterol molecularly imprinted polymer. The probe combines the high specific identification performance of the molecularly imprinted polymer with the high sensitivity of the upconversion luminescent material. The production method comprises the following steps: preparing the upconversion luminescent material, preparing the clenbuterol molecularly imprinted-upconversion luminescent material fluorescence probe, and eluting clenbuterol molecules. The fluorescence probe produced in the invention emits visible light under the excitation of infrared light, and has the advantages of high sensitivity, strong specificity and small interference. The production method has the advantages of simplicity, good reappearance, and good application prospect in selective identification and detection of clenbuterol in a practical sample.
Owner:BOHAI UNIV
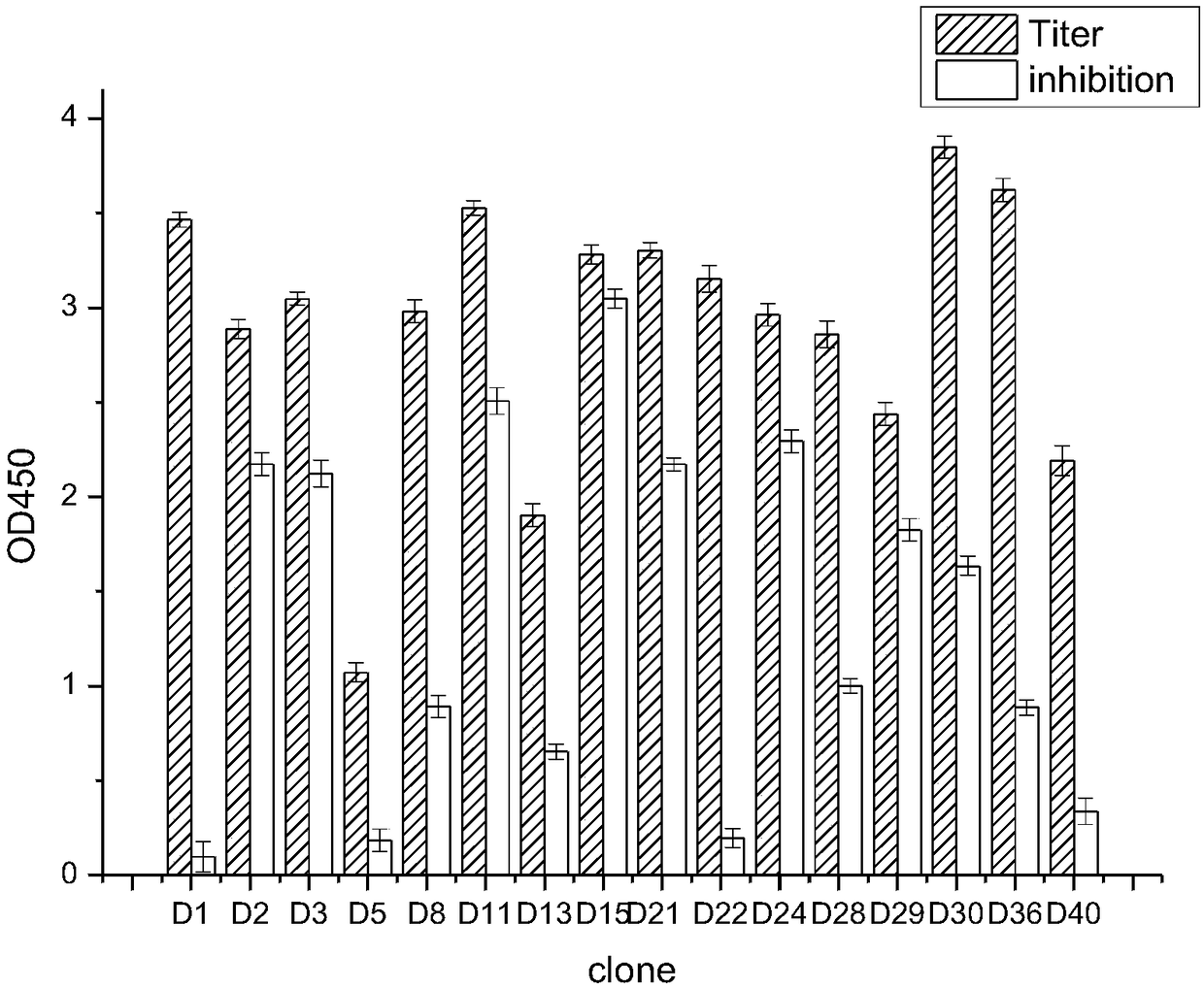
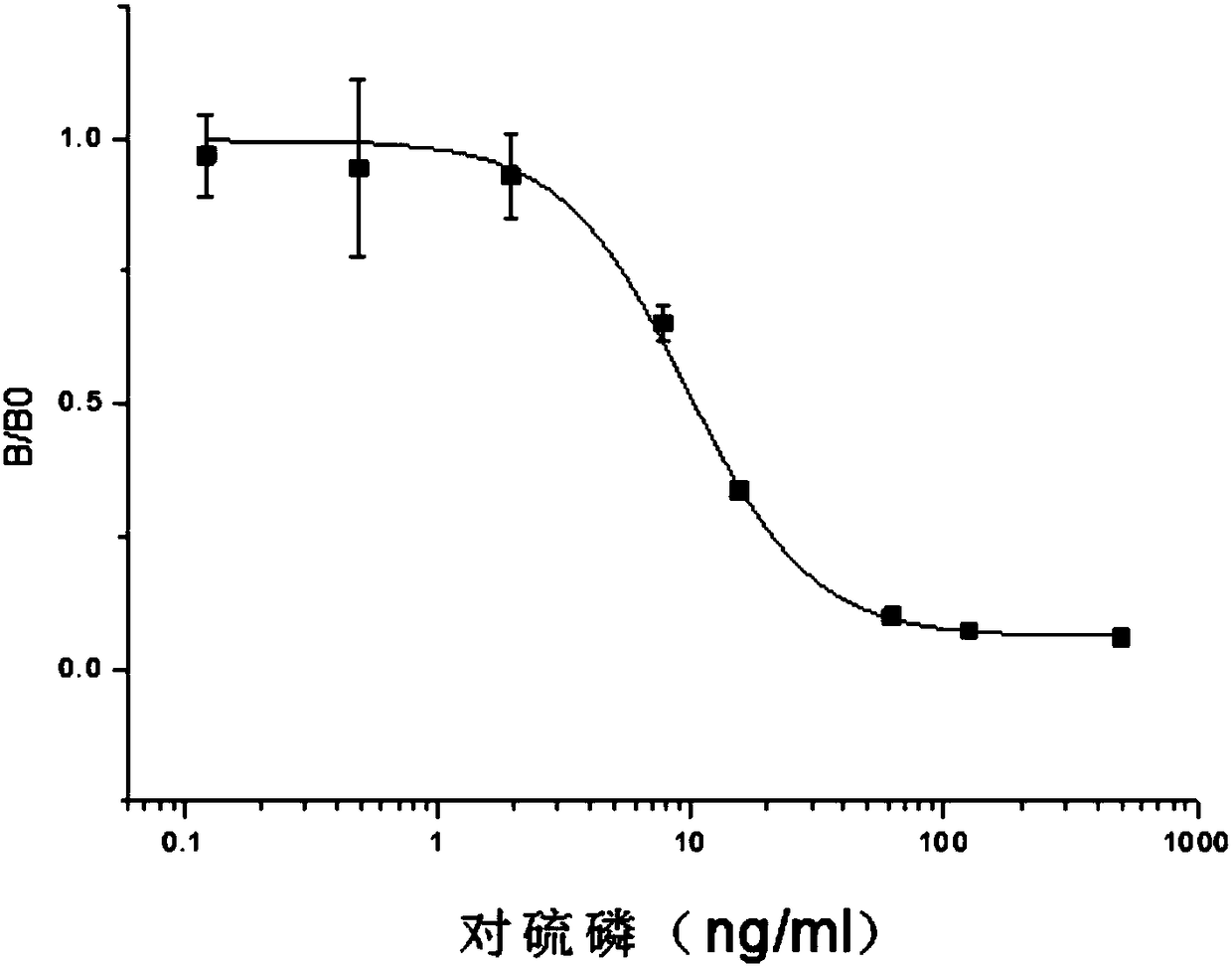
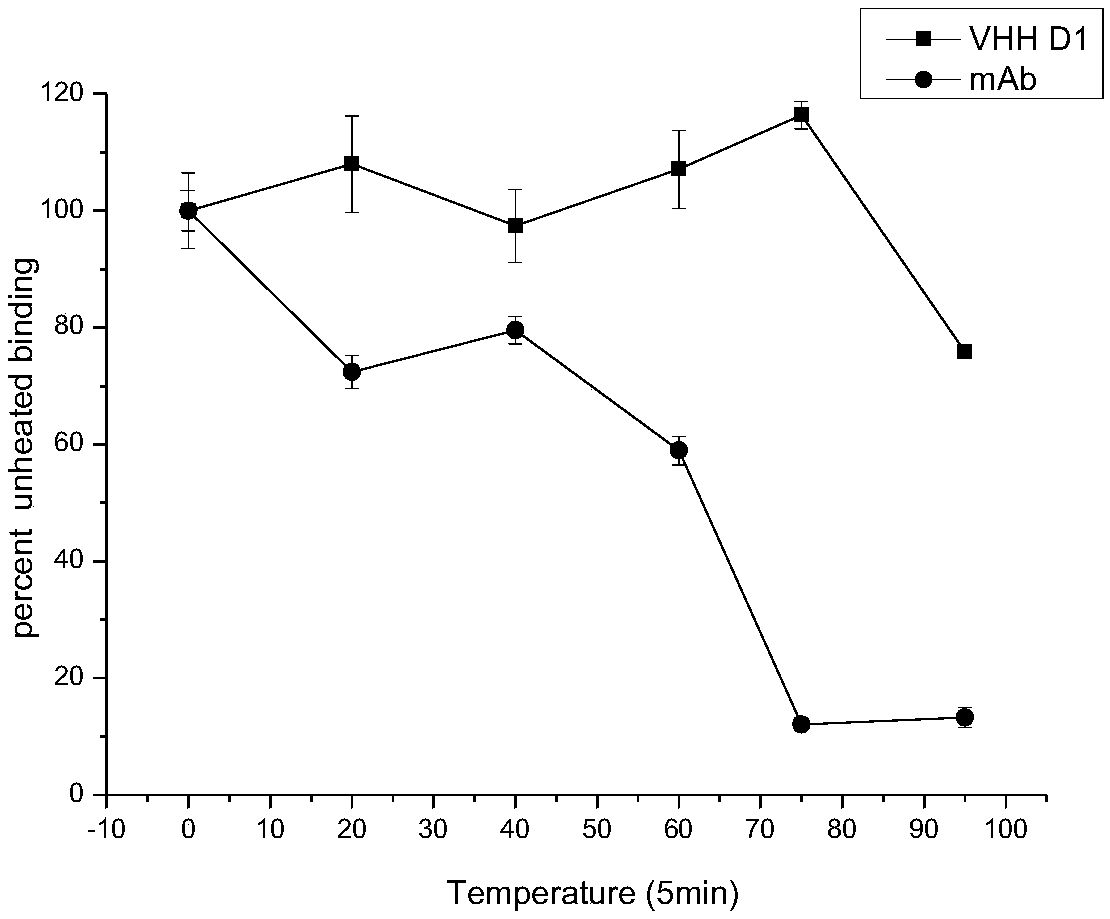
Enzyme linked immunosorbent assay kit for detecting diethoxy organophosphorus pesticide based on nano antibody
The invention discloses an enzyme linked immunosorbent assay kit for detecting a diethoxy organophosphorus pesticide based on a nano antibody. The enzyme linked immunosorbent assay kit contains the nano antibody as a detection antibody, and an amino acid sequence of the nano antibody is shown as SEQ ID NO.1. The kit is the enzyme linked immunosorbent assay kit, and the enzyme linked immunosorbentassay kit comprises an enzyme label plate containing organophosphorus pesticide antigen, a horseradish peroxidase-labelled secondary antibody solution, a diethoxy organophosphorus pesticide standard solution, substrate liquid, a substrate buffer solution, a stopping solution, and condensation washing liquid. The expression level of the diethoxy organophosphorus pesticide based on the nano antibodyis high, the stability is strong, the kit can be used for detecting a plurality of the diethoxy organophosphorus pesticides, realizes broad spectrum specific identification of the diethoxy organophosphorus pesticide, has the advantages of accurate detection result, good effect, good stability, simple operation, high sensitivity, and low cost, and can accurately detect the residues of the diethoxyorganophosphorus pesticide in an agricultural product.
Owner:SOUTH CHINA AGRI UNIV
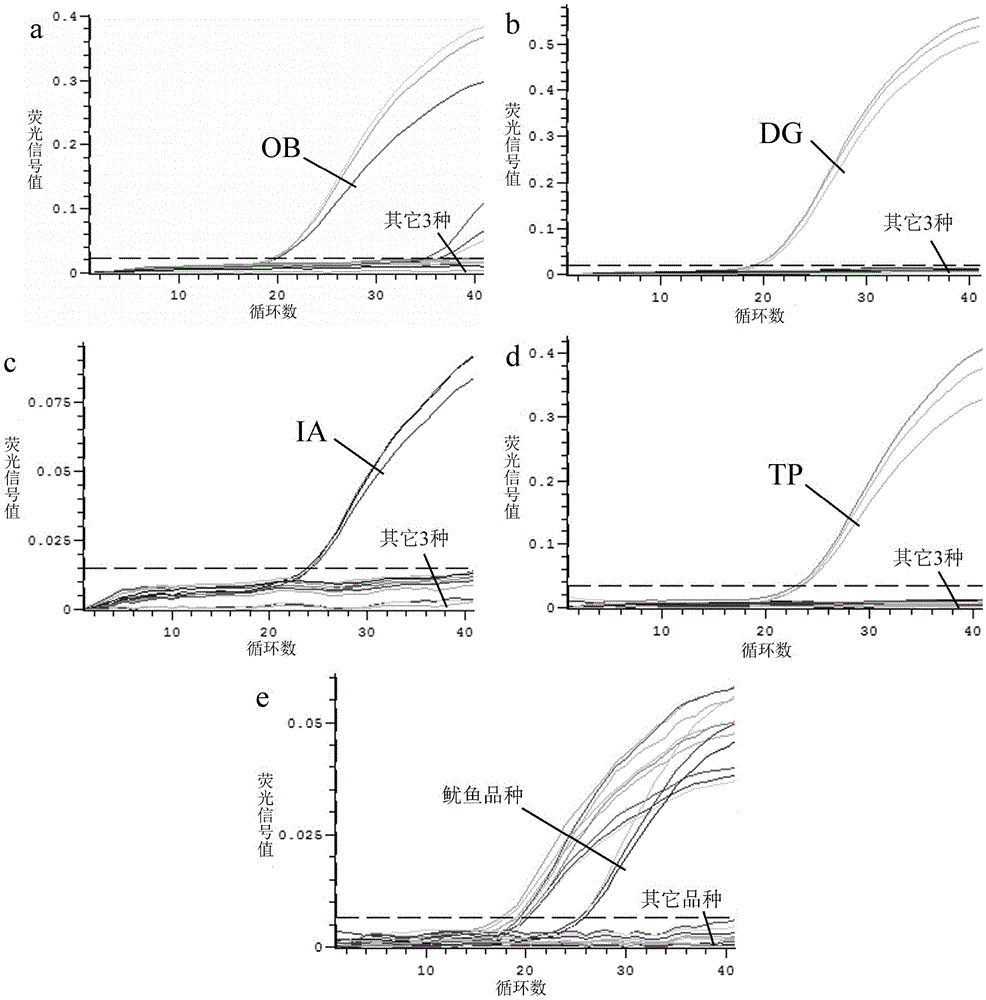
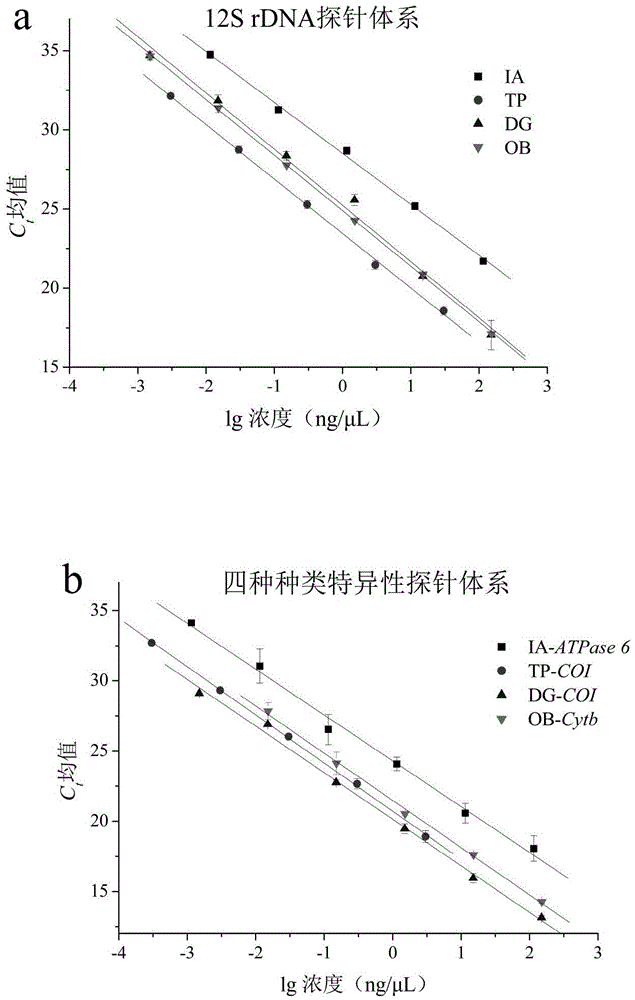
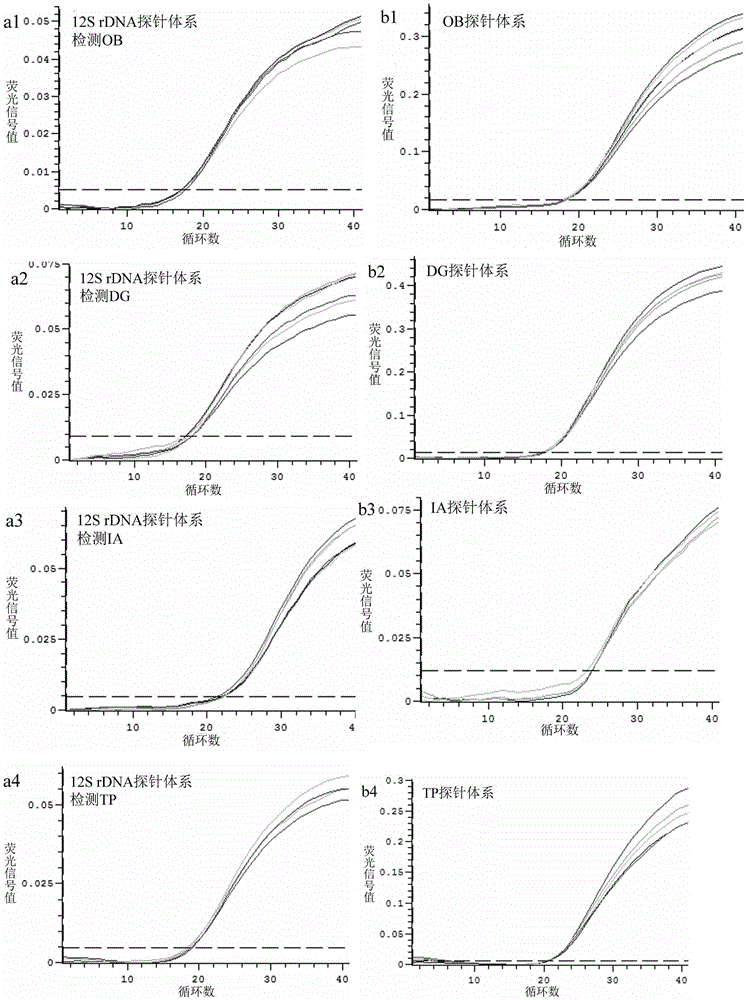
Method for identifying squids or highly processed product varieties thereof by virtue of fluorescent quantitative PCR
InactiveCN105525022AStrong specificityImprove accuracyMicrobiological testing/measurementDNA/RNA fragmentationFluorescenceBiology
The invention discloses a method for identifying squids or highly processed product varieties thereof by virtue of fluorescent quantitative PCR. The method comprises the following steps: (1) extracting genome DNA from a to-be-detected squid sample; (2) designing specific identification primer and probe for common squids, wherein the four common squids include Ommastrephes bartramii, dosidicus gigas, illex argentinus and todarodes pacificus; (3) setting a fluorescent quantitative PCR amplification system; and conducting real-time fluorescent PCR detection by virtue of the specific primer and probe by taking the genome DNA as an amplification template; and (4) judging in accordance with the result of the real-time fluorescent PCR detection. By virtue of the method disclosed by the invention, the squids and the highly processed product varieties thereof can be rapidly identified.
Owner:ZHEJIANG GONGSHANG UNIVERSITY
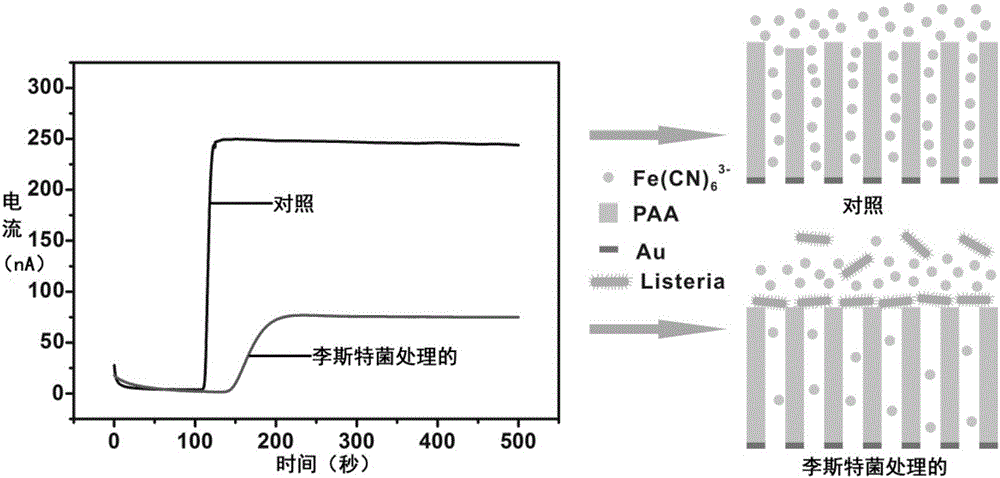
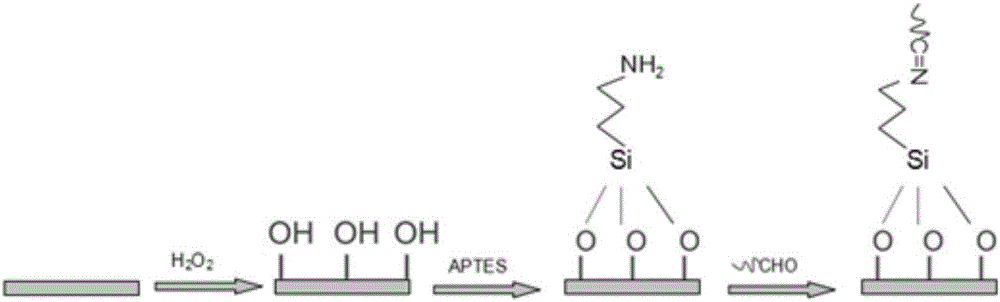
Novel high-sensitivity LM (listeria monocytogene) detection method based on aptamer modified porous alumina membrane
InactiveCN106018508ADetection has no effectHigh selectivityMaterial electrochemical variablesEscherichia coliPotassium ferricyanide
The invention relates to a fast ultra-sensitive LM (listeria monocytogene) detector based on nanochannel confinement characteristics and constructed according to the nature of specific identification between a target molecule and an aptamer of the target molecules. The invention further relates to a method for detecting the LM by taking potassium ferricyanide ions as probe ions and taking an LM DNA modified porous alumina membrane as an electrode for assembling a self-made electrolytic tank. When LM is present and the concentration is lower, a current increasing value is remarkably reduced with the increase of the concentration; a current change value is reduced with increase of the LM concentration; when the LM concentration is in a range of 100-1,250 CFU / mL, a linear relation is formed between the LM concentration and the current increasing value; when the LM concentration is higher than 1,500 CFU / mL, the current change value becomes stable. Therefore, the lowest detection limit of the detection method for the LM can reach 100 CFU / mL, the linear range is 100-1,250 CFU / mL, and the detection can be completed within 10 min; a 108 CFU / mL of escherichia coli and staphylococcus aureus control experiment indicates that the method has high selectivity on the LM.
Owner:GUANGDONG OCEAN UNIVERSITY
Fast method for detecting micro-organisms in food samples
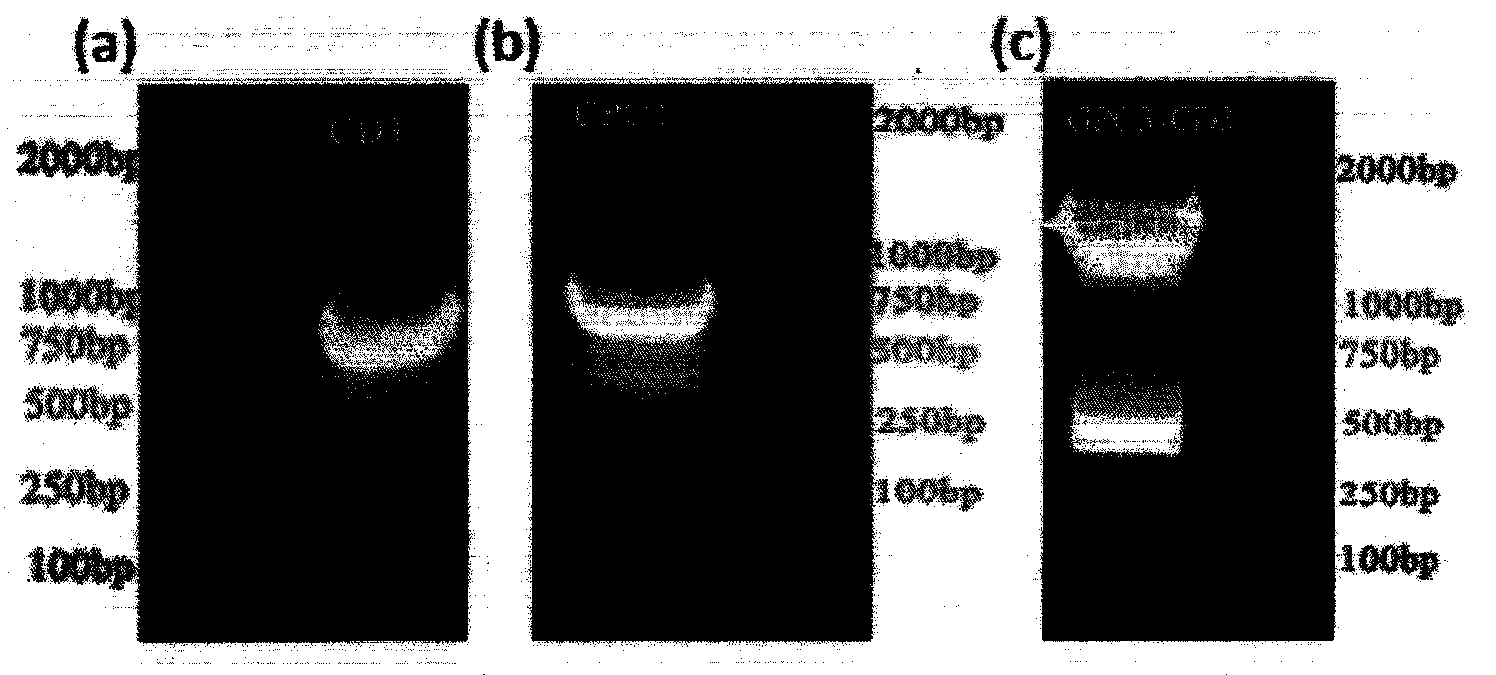
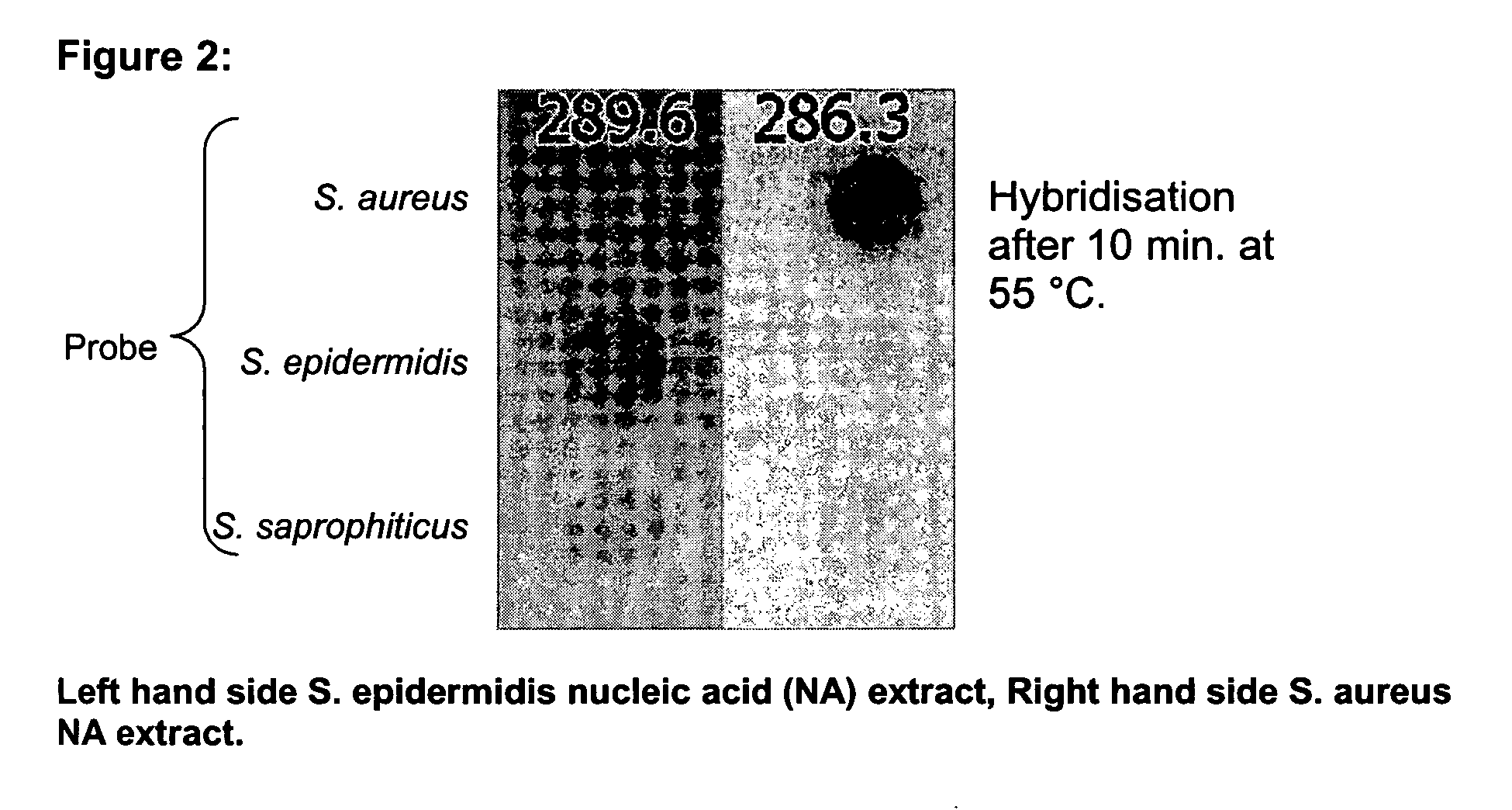
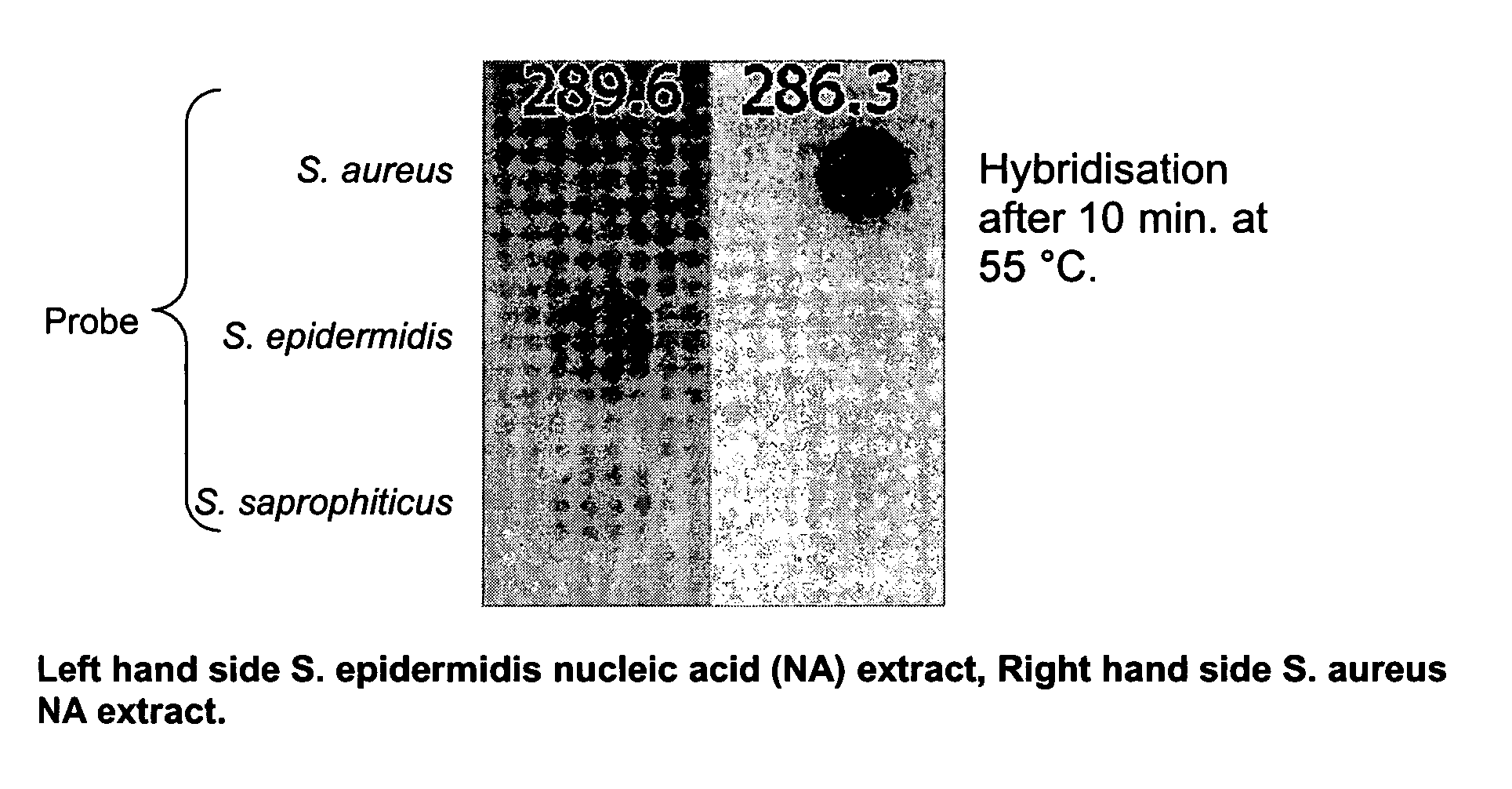
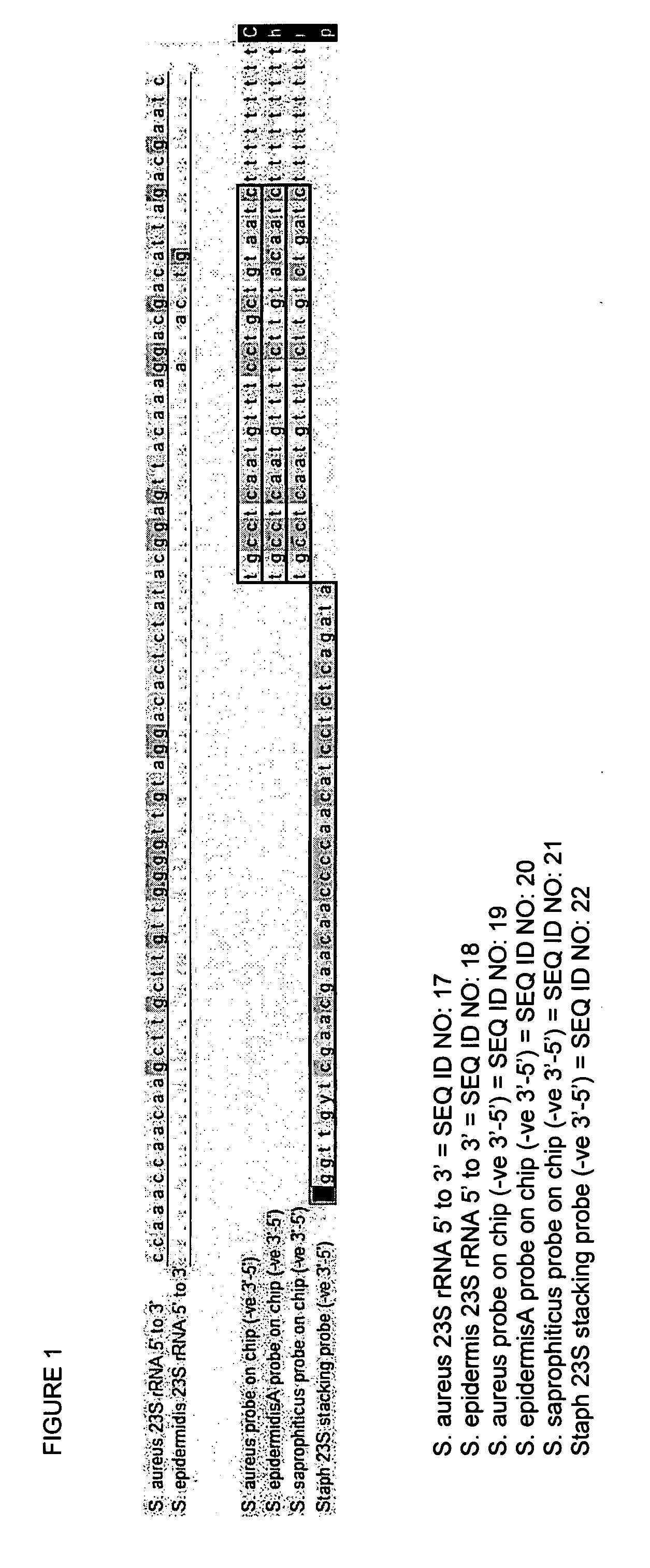
InactiveUS20060194223A1Increase flow rateLower overall flow resistanceMicrobiological testing/measurementBiological testingBiotechnologyGenomic Segment
The present invention relates to a specific method accomplishing fast and specific identification of contaminating micro-organisms in large amounts of food stuffs. A method has been developed based on random genome fragments or Zipcode oligonucleotides and DNA microarray technology that overcomes the disadvantages of whole-genome DNA-DNA hybridisation. In particular, the present invention provides a method for characterising micro-organisms possibly present in a sample, comprising the steps of collecting said micro-organisms if present, extracting nucleic acids from said micro-organisms, specifically amplifying said nucleic acids, thereby providing an amplified nucleic acid mixture comprising the target nucleic acid in amplified form, and analysing the amplified nucleic acid mixture, whereby the said micro-organisms if present are characterised. The present invention further relates to the use of filters, microarrays and amplification steps in said method as well as a kit comprising the same.
Owner:CHECK POINTS HLDG
Fluorescent probe for detection of cystatin C and construction method thereof
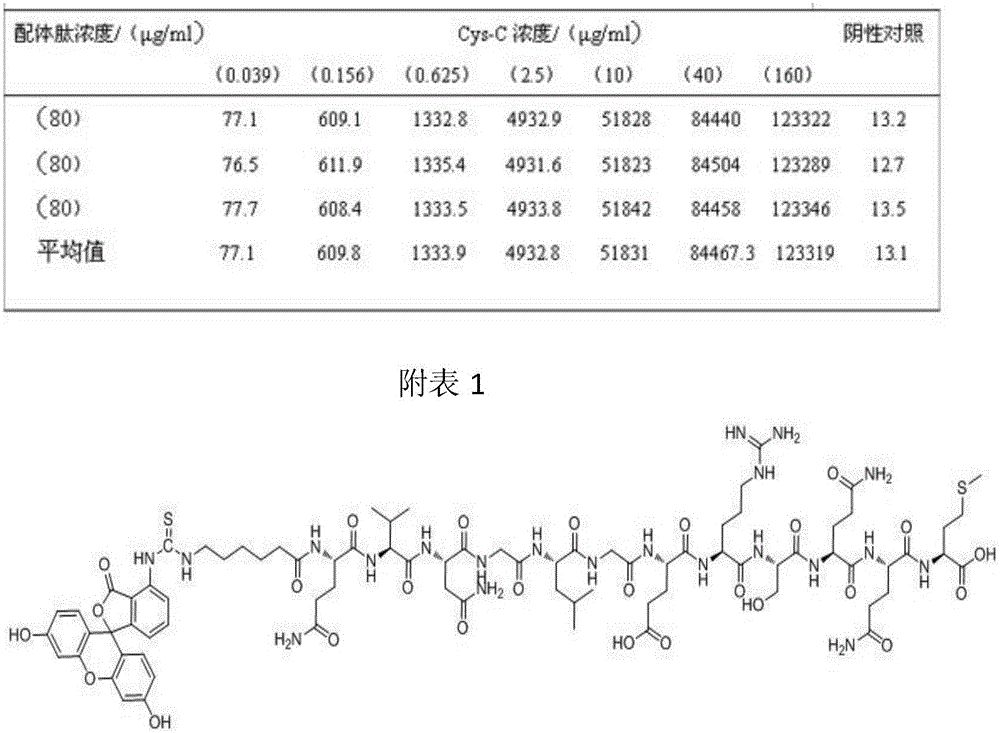
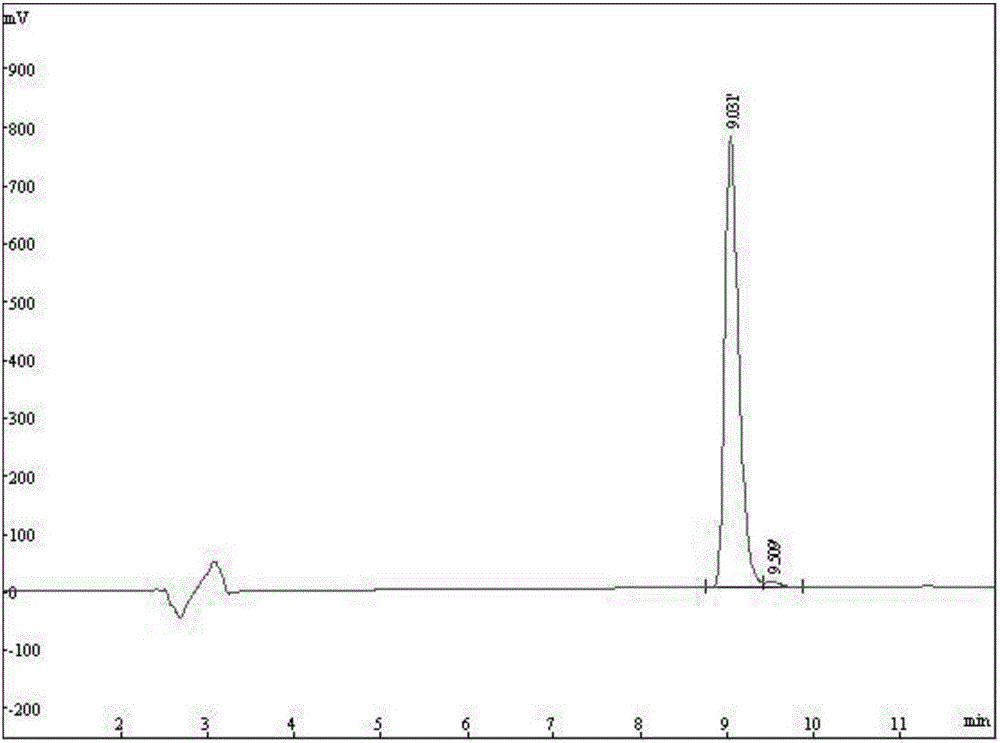
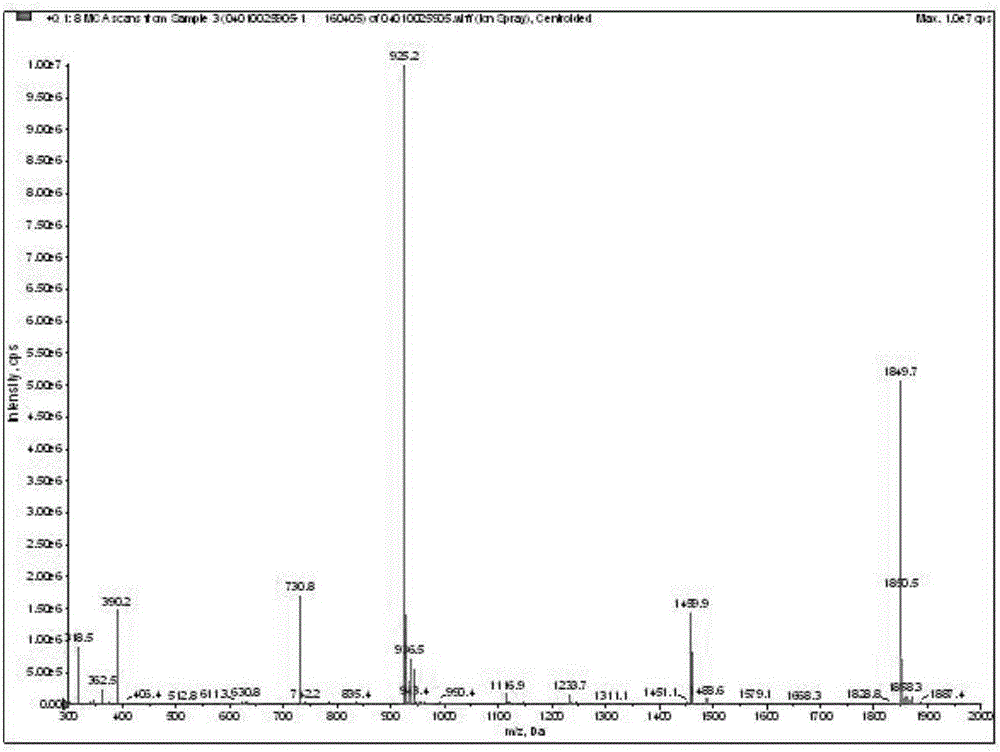
The invention discloses a fluorescent probe for detection of cystatin C and a construction method thereof, relates to the molecular biological and microbiological technical fields, and is characterized in that the construction method comprises the steps: with Cys-C as a target, screening to obtain a Cys-C specific-binding recombinant phage by using a phage surface random displayed 12-mer peptide library, extracting single stranded DNA of the recombinant phage, carrying out sequencing and sequence comparison, to obtain a sequence Gln-Val-Asn-Gly-Leu-Gly-Glu-Arg-Ser-Gln-Gln-Met of a Cys-C specific affinity ligand having the molecular weight of 1346.5, then carrying out solid phase polypeptide synthesis and fluorescence labeling to obtain the fluorescent probe FITC-Acp-Gln-Val-Asn-Gly-Leu-Gly-Glu-Arg-Ser-Gln-Gln-Met having the molecular weight of 1848.7 and specifically bond with the Cys-C. The fluorescent probe and the construction method thereof have the beneficial effects: 1, a brand new 'tool' is provided for specific identification and content detection of the Cys-C; and 2, the fluorescent ligand peptide probe not only can qualitatively identify the Cys-C, but also can quantitatively detect the content of the Cys-C in samples.
Owner:长春技特生物技术有限公司
Molecularly imprinted photonic crystal film for rapidly detecting lysozyme, and preparation method and application thereof
ActiveCN109900653AQuick checkObvious diffraction peak shiftScattering properties measurementsColor/spectral properties measurementsSpectral responseMicrosphere
The invention relates to a molecularly imprinted photonic crystal film for rapidly detecting lysozyme and belongs to the field of material chemistry and analysis and detection. The molecularly imprinted photonic crystal film is characterized in that the molecularly imprinted photonic crystal film which can realize specific identification and binding to lysozyme protein molecules and has the inverse opal structure is prepared by combining silica photonic crystal microspheres with the molecular imprinting technology. As the number of the bound lysozyme protein molecules increases, the macroporous structure on the film changes, the Bragg diffraction wavelength of the prepared molecularly imprinted photonic crystal film is made to realize red shift. According to the spectral response of the prepared molecularly imprinted photonic crystal gel membrane to the lysozyme, rapid detection of the concentration of the lysozyme in the sample can be realized. The molecularly imprinted photonic crystal film is advantaged in that properties of high sensitivity and selectivity and low detection limit are realized, and the molecularly imprinted photonic crystal film can be used as a biosensor used for rapidly detecting the lysozyme protein molecules in a biological sample.
Owner:SHANGHAI E K M BIOTECH
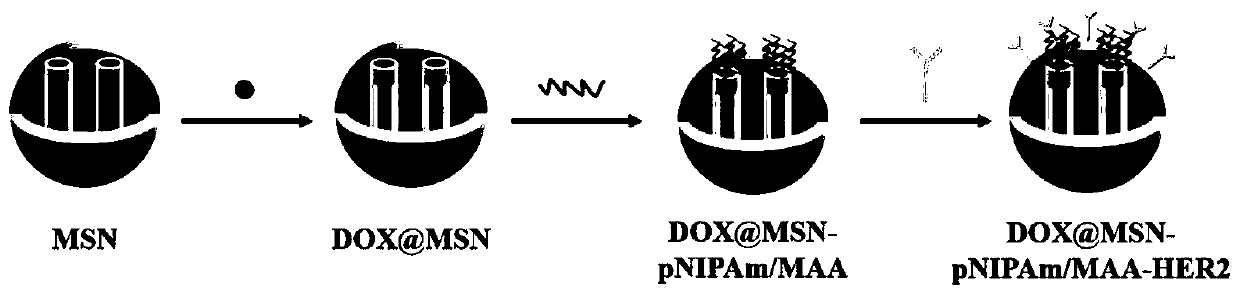
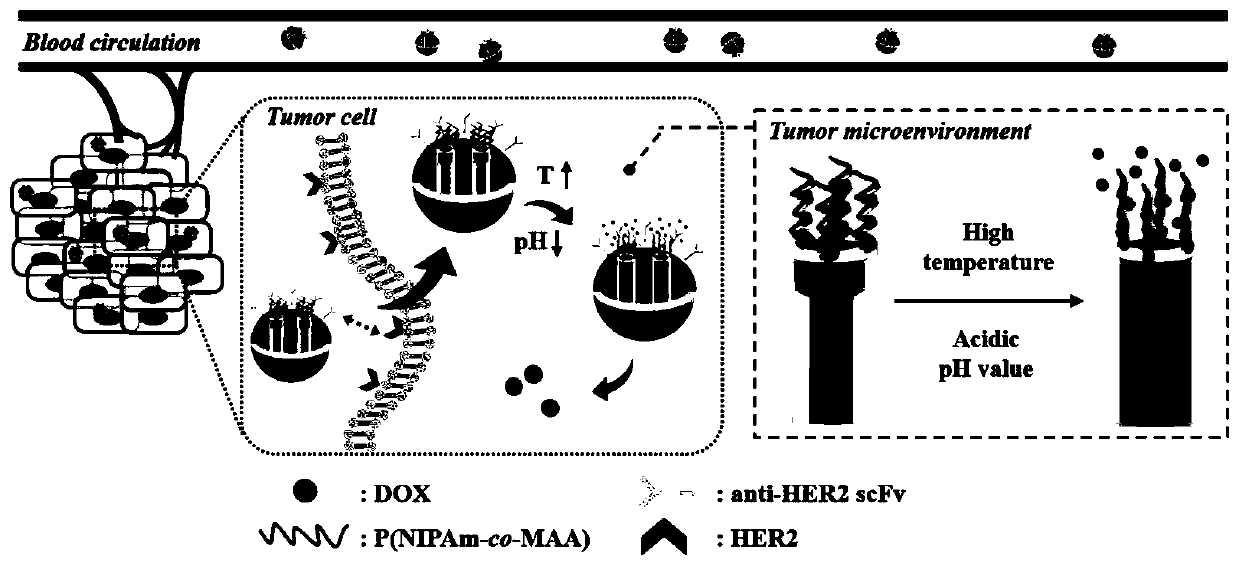
Tumor self-targeting multilevel response type mesoporous silicon drug delivery system and preparation method thereof
InactiveCN111249467AReduce premature leakageStable and controlled releaseOrganic active ingredientsPharmaceutical non-active ingredientsAntiendomysial antibodiesEngineering
The invention relates to a tumor self-targeting multilevel response type mesoporous silicon drug delivery system which includes mesoporous silicon nanoparticles. The mesoporous silicon nanoparticles are loaded with anticarcinoma drugs, and the surfaces of the mesoporous silicon nanoparticles are modified with polymer layers and low-toxicity tumor-targeting single-chain antibodies, so that the drugdelivery system has breast cancer cell targeting, temperature responsiveness and pH responsiveness. The invention further provides a corresponding preparation method. The tumor self-targeting multilevel response type mesoporous silicon drug delivery system has good biocompatibility, the specific targeting of HER2 highly expressing breast cancer cells and the controlled release of drug with temperature and pH dual response, and safe and controllable drug release can be achieved while tumor-specific identification is improved.
Owner:EAST CHINA UNIV OF SCI & TECH
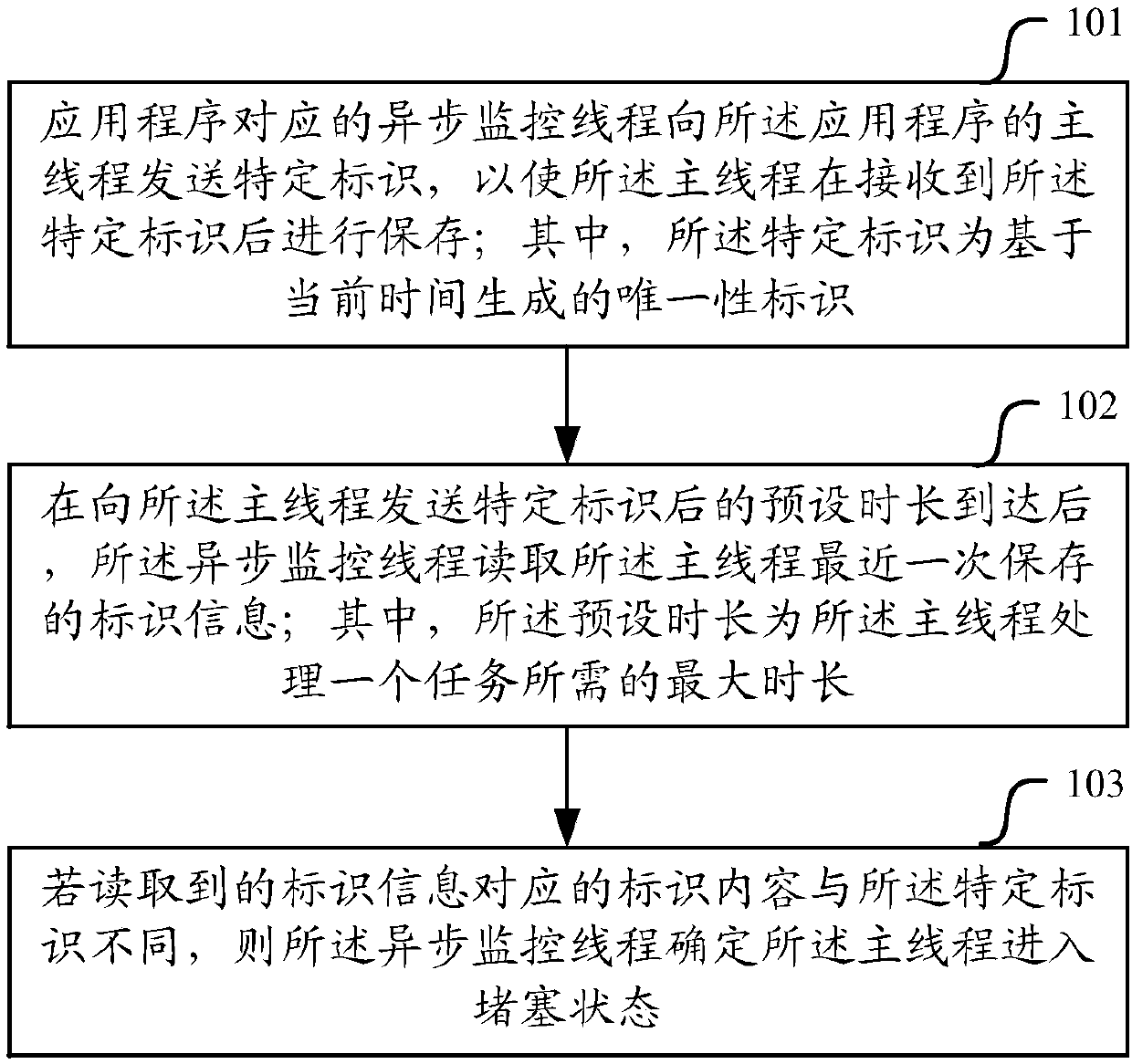
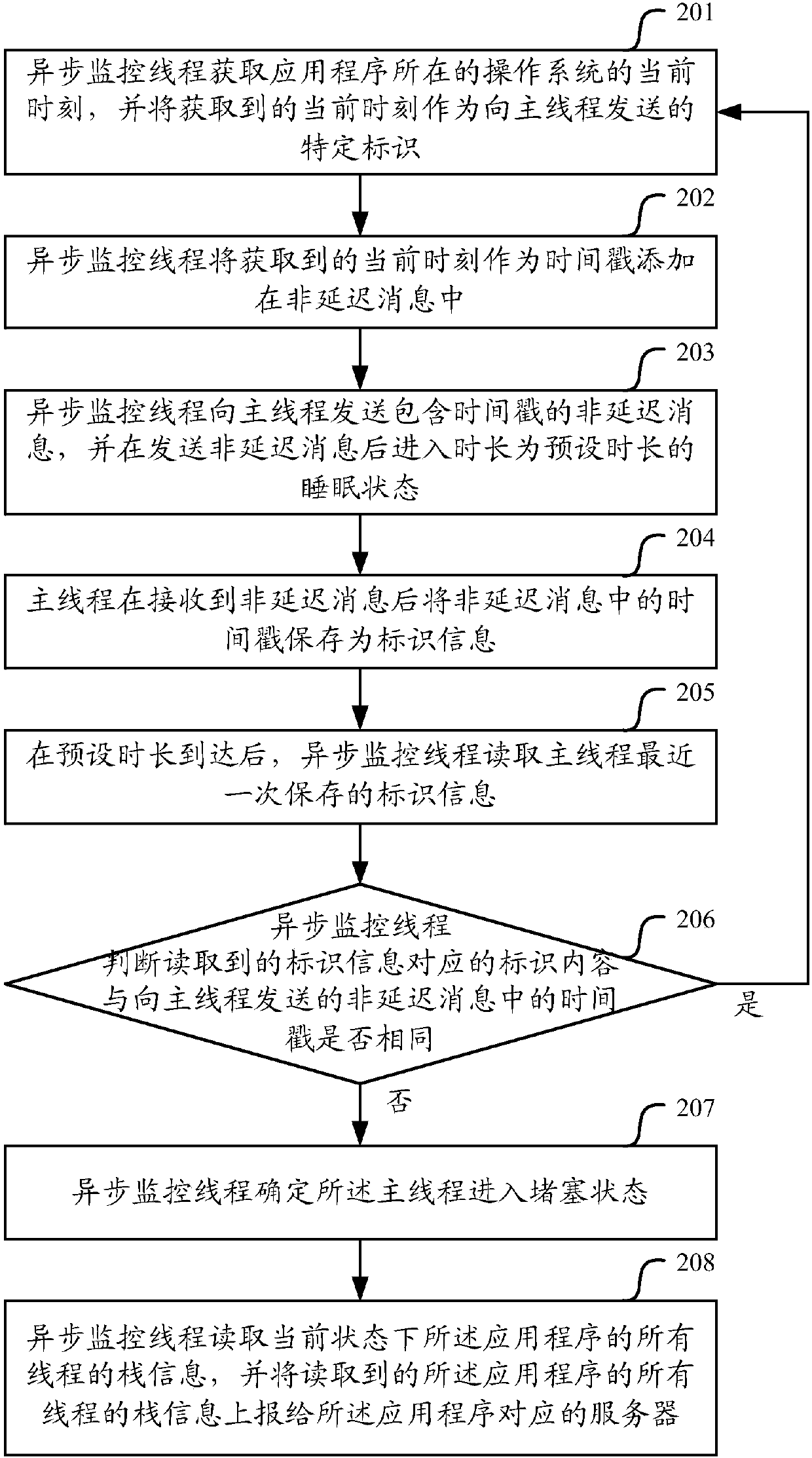
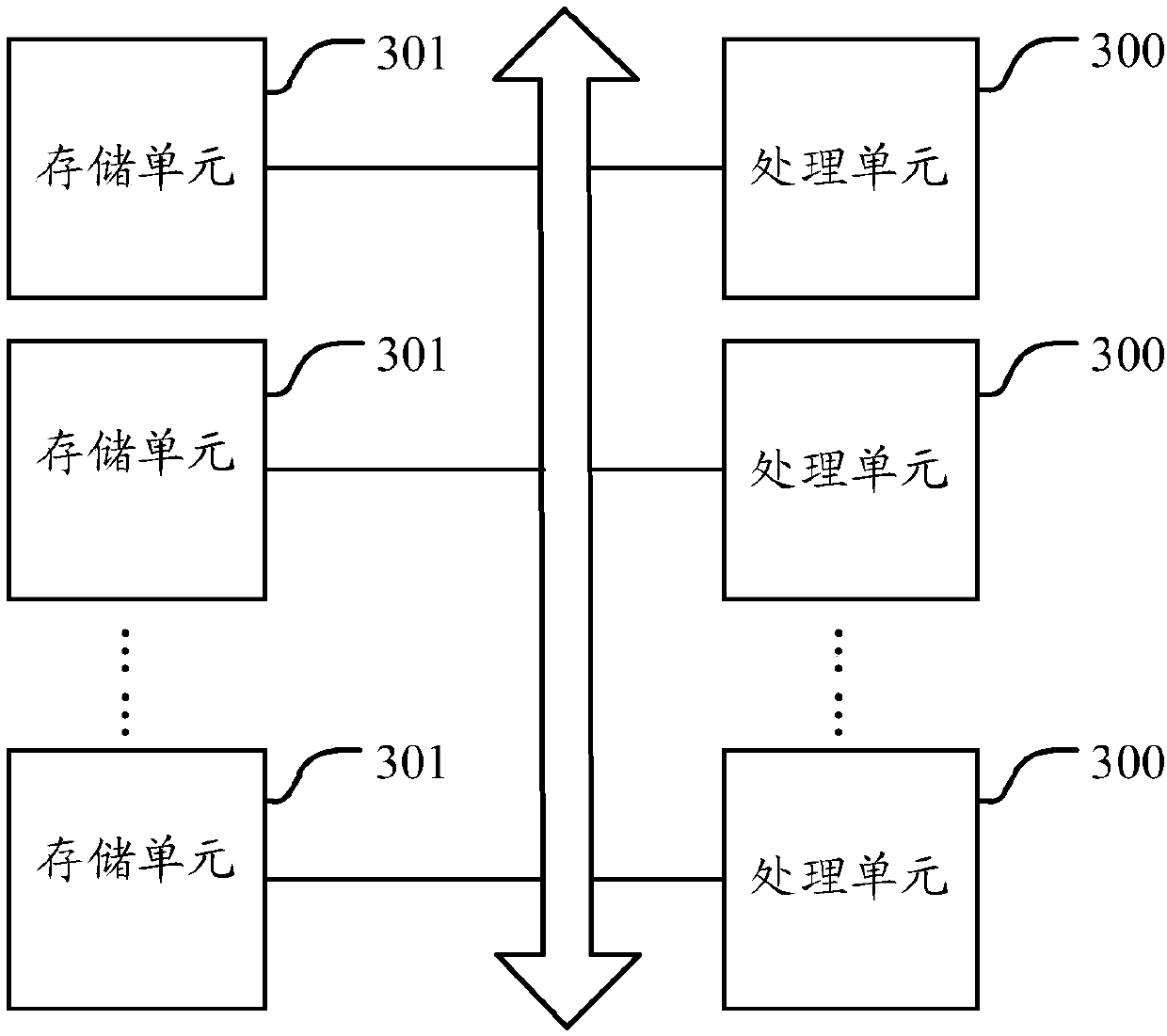
Monitoring method and device of main thread of application program
ActiveCN107729207AImprove the efficiency of health monitoringRealize monitoringHardware monitoringReal-time computingApplication procedure
The invention relates to the technical field of system application development, in particular to a monitoring method and device of a main thread of an application program. The monitoring method and device are used for solving the problem in the prior art that a monitoring method of the main thread of the application program is low in efficiency. According to the embodiment, an asynchronous monitoring thread sends a specific identification to the main thread, so that the main thread stores the specific identification after receiving the specific identification; after a preset duration for sending the specific identification to the main thread is reached, the asynchronous monitoring thread reads the last identification information stored by the main thread; if the identification content corresponding to the read identification information is different from the specific identification, it is determined that the main thread enters a congestion state. By means of the monitoring method and device of the main thread of the application program, since one asynchronous monitoring thread is set for each application program to be used for monitoring the running state of the main thread of thecorresponding application program, when the main thread is congested, the congestion can be detected within a short time, and thus the efficiency of monitoring the running state of the main thread isimproved.
Owner:HISENSE VISUAL TECH CO LTD
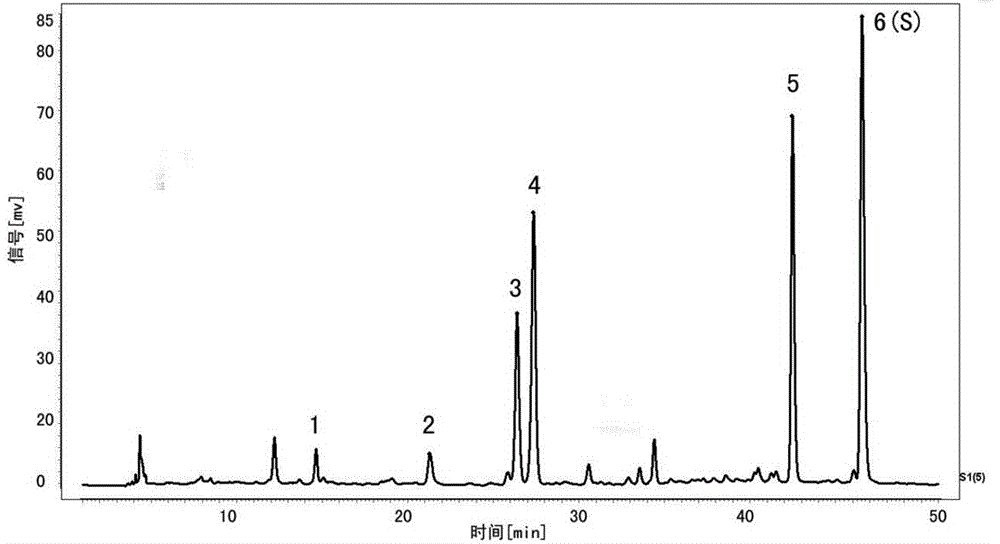
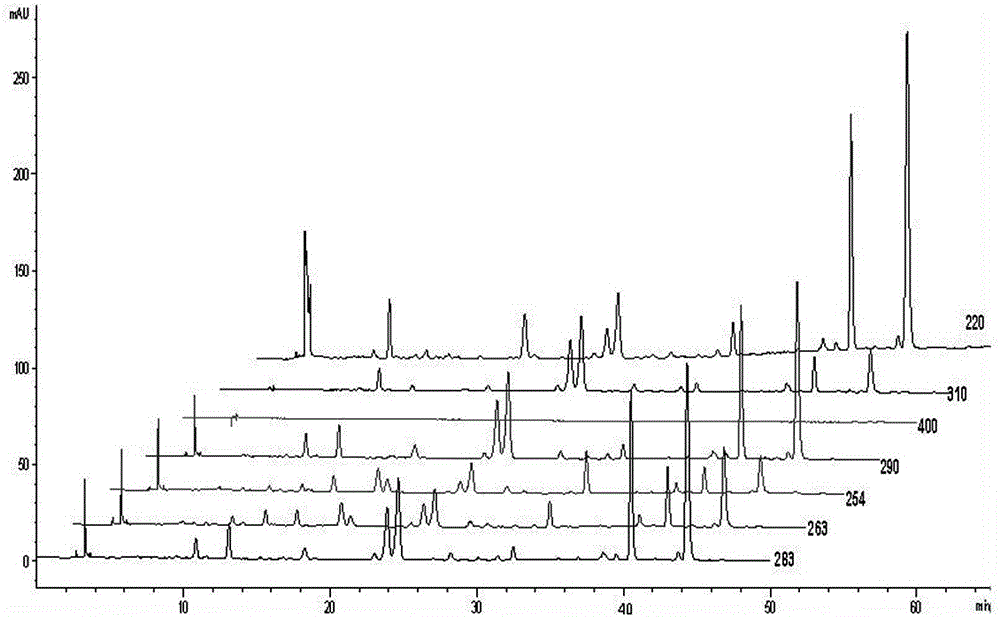
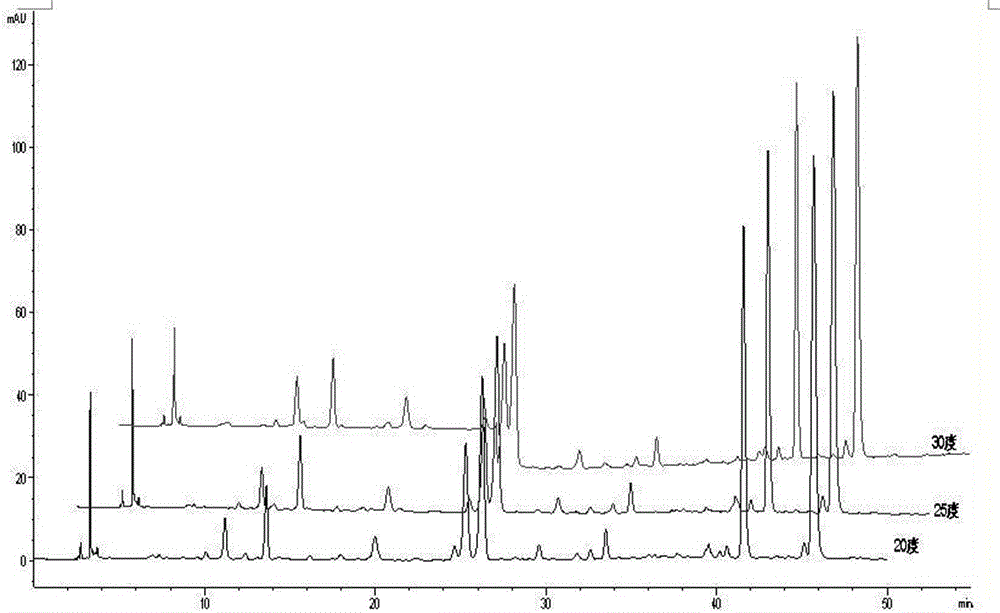
Burned Rhizoma drynariae formula granule characteristic spectrum and establishing method thereof
ActiveCN106053658AStrong specificityStrengthen the identification of specificityComponent separationTheoretical plateNaringin
The invention relates to the technical field of pharmaceutical analysis, in particular to a burned Rhizoma drynariae formula granule characteristic spectrum and an establishing method thereof. The establishing method includes that octadecylsilane chemically bonded silica is used as a filling agent, methanol is used as a mobile phase A, a 0.1v / v% phosphoric acid solution is used as a mobile phase B, and gradient elution is performed according to regulations in the following table; detection wavelength is 290 nm, column temperature is 20 DEG C, and flow speed is 1.0ml / min; number of theoretical plates should not be lower than 50000 if being calculated according to naringin peak; six characteristic peaks should be presented in a test piece characteristic spectrum, and relative retention time should be within + / -5% of specified values; the specified values are 0.299 (peak 1), 0.446(peak 2), 0.557(peak 3), 0.579(peak 4), 0.911(peak 5) and 1.00(peak 6). According to characteristics of formula granules, specific identification and multi-component integral quality control are enhanced, and the characteristic spectrum of the variety is established for comprehensive quality control.
Owner:SICHUAN NEO GREEN PHARMA TECH DEV
Specific molecular marker for panax ginseng and panax quinquefolius and identification method thereof
InactiveCN103923909AGood repeatabilityReduce length requirementsMicrobiological testing/measurementDNA/RNA fragmentationPanax stipuleanatusA-DNA
Belonging to the technical field of traditional Chinese medicine, the invention relates to identification methods of Chinese herbal medicines, in particular to a specific molecular marker for panax ginseng and panax quinquefolius and an identification method thereof. The invention provides a DNA molecular marker for specific identification of panax ginseng and panax quinquefolius. The DNA molecular marker is located on the (SSR) loci of DNA sequences Seq.No.1-5 respectively. 5 pairs of primers are designed according to the sequences to perform PCR (polymerase chain reaction) amplification on the DNA templates of panax ginseng and panax quinquefolius. PCR products for specific identification of panax ginseng and panax quinquefolius can be amplified respectively by each pair of the primers, the products with different lengths are detected by polyacrylamide gel electrophoresis and agarose gel electrophoresis, so that panax ginseng and panax quinquefolius can be rapidly identified. The method needs a small amount of samples, is not restricted by product shapes, and has low requirement for sample's DNA length. The PCR is stable and has good repeatability.
Owner:FUDAN UNIV
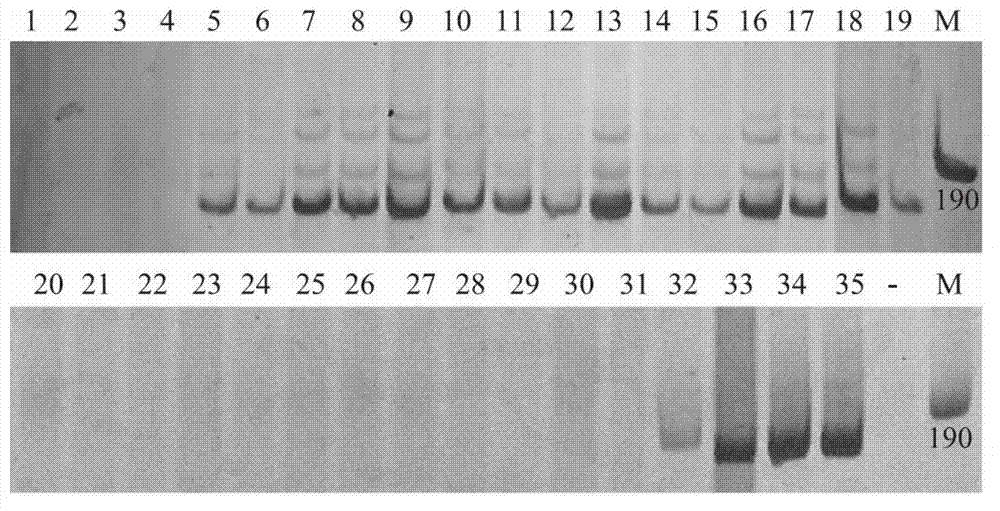
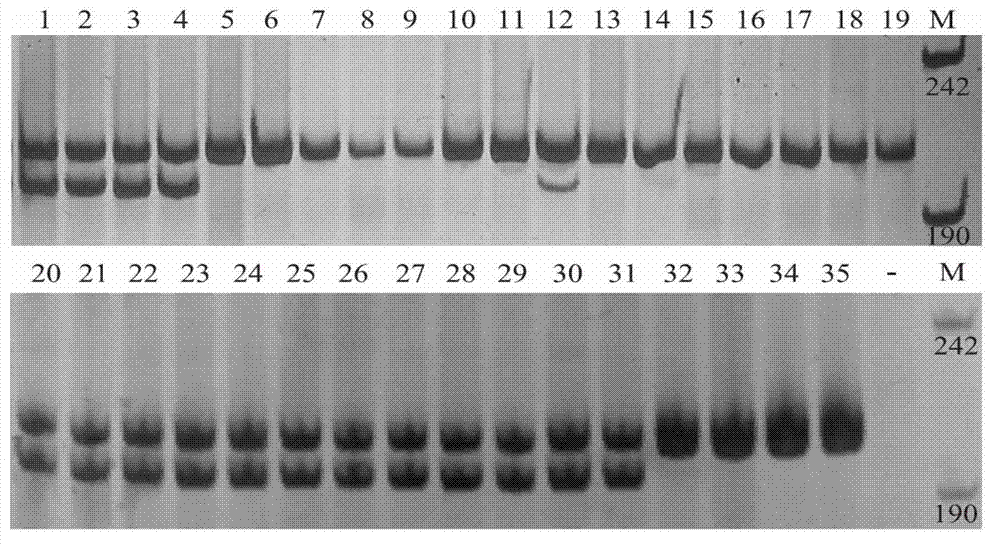
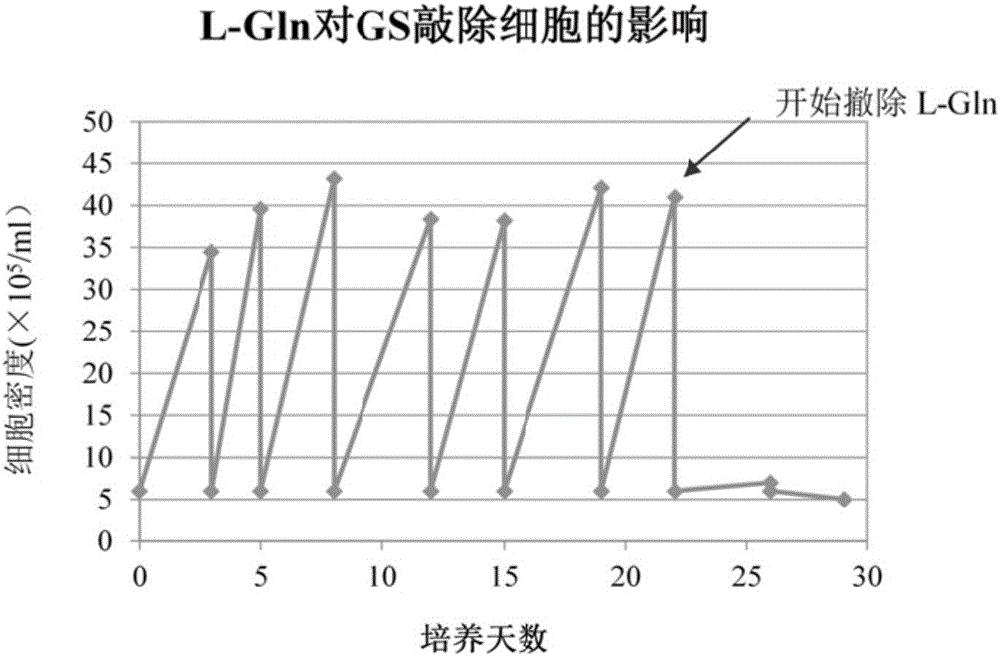
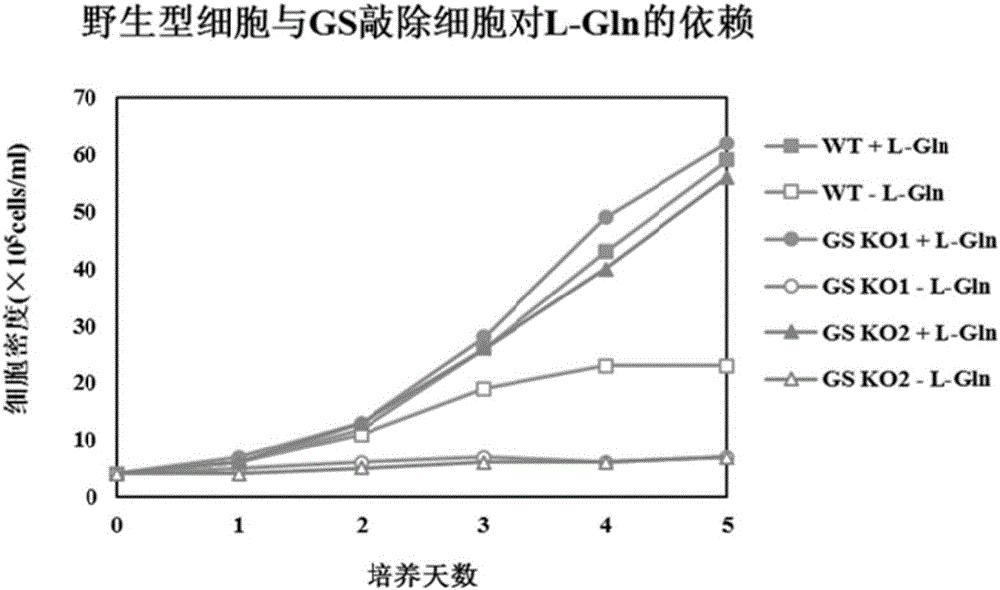
GS (glutamine synthetase) gene specific identification crRNA and application thereof
ActiveCN105950622AMixed knockouts are easily achievedImprove efficiencyNucleic acid vectorFermentationProtein targetWild type
The invention belongs to the field of cell and gene engineering, genetic modification and therapeutic recombinant protein industrial production, and particularly relates to GS (glutamine synthetase) gene specific identification crRNA and application thereof. DNA (deoxyribonucleic acid) sequences identified by the GS gene specific identification crRNA are selectively DNA sequences shown as one of sequences SEQ ID NO.1-SEQ ID NO.8. The GS gene specific identification crRNA and the application have the advantages that GS genes of diversified cells can be specifically identified by the GS gene specific identification crRNA, the GS gene specific identification crRNA is wide in applicability, and integral procedures can be implemented easily and efficiently; the target protein expression quantities of cells without the GS genes can be greatly increased on the basis of imported exogenous carriers as compared with wild host cells, and accordingly required-to-be-inputted labor, material resources and financial resources for screening protein expression cell strains can be reduced; the GS genes are knocked out, and accordingly the industrial production cost can be reduced; the cells without the GS are used as host target gene expression cells, other chemical substances can be omitted in procedures for producing the cells, and accordingly the production safety can be improved.
Owner:苏州晟济药业有限公司
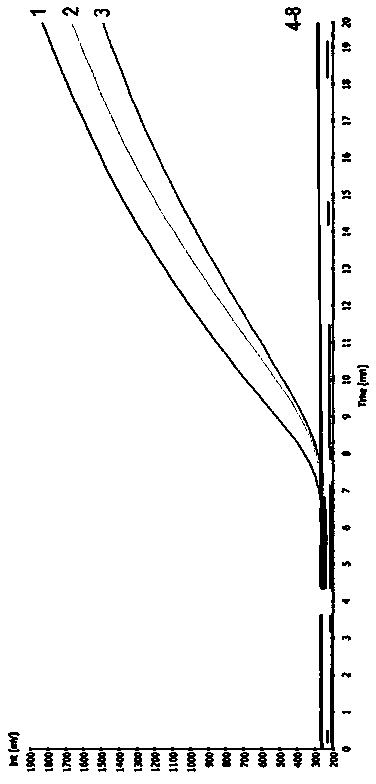
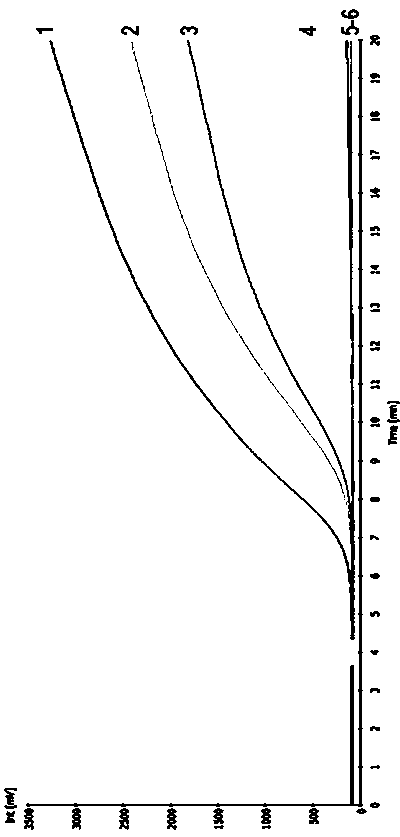
Specific identification for transgenic rice kefeng No.6 strain via adoption of RPA (Recombinase Ploymerase Amplification) technology
ActiveCN103525936AObvious amplification curveMicrobiological testing/measurementDNA/RNA fragmentationBiotechnologyGenetically modified rice
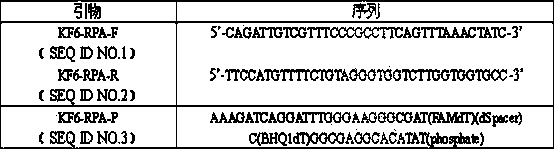
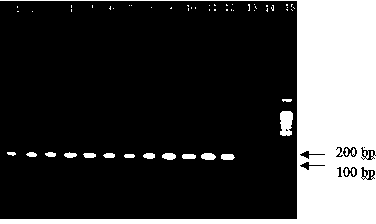
The invention firstly provides an RPA (Recombinase Ploymerase Amplification) detection method of identification of transgenic insect-resistant rice kefeng No.6 strain and in particular discloses a primer and probe combination used for indentifying transgenic insect-resistant rice kefeng No.6 via a recombinase ploymerase amplification technology; a sequence of a positive primer is expressed as SEQ ID No.1; a sequence of a negative primer is expressed as SEQ ID No. 2; a sequence of the probe is expressed as SEQ ID No.3. The invention also discloses a method for indentifying transgenic insect-resistant rice kefeng No.6. The method comprises the following steps: extracting DNA (Deoxyribose Nucleic Acid) of a sample to be detected as a template, performing RPA quick amplification and real-time fluorescence detection by using the primer; and proving that the detected sample contains components of the transgenic insect-resistant rice kefeng No.6 if an obvious amplification curve is obtained.
Owner:THE INST OF BIOTECHNOLOGY OF THE CHINESE ACAD OF AGRI SCI
Sugarcane genome endogenous reference gene acetolactate synthase gene PCR (polymerase chain reaction) primer sequences and amplification method
InactiveCN103409414AWide applicabilityBroad representationDNA preparationDNA/RNA fragmentationReference genesAcetolactate synthase
The invention discloses sugarcane genome endogenous reference gene acetolactate synthase gene PCR (polymerase chain reaction) primer sequences and an amplification method. The primer sequences are composed of ScALS-F and ScALS-R. The PCR amplification method comprises a reaction system and amplification conditions. The invention provides a genome endogenous reference gene for PCR detection and species specific identification of transgenic sugarcane, and meanwhile, provides insurance for detection precision and result accuracy of the transgenic sugarcane.
Owner:FUJIAN AGRI & FORESTRY UNIV
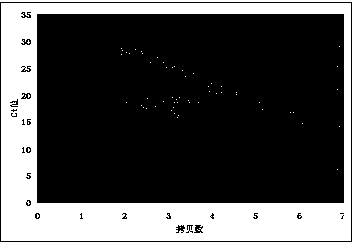
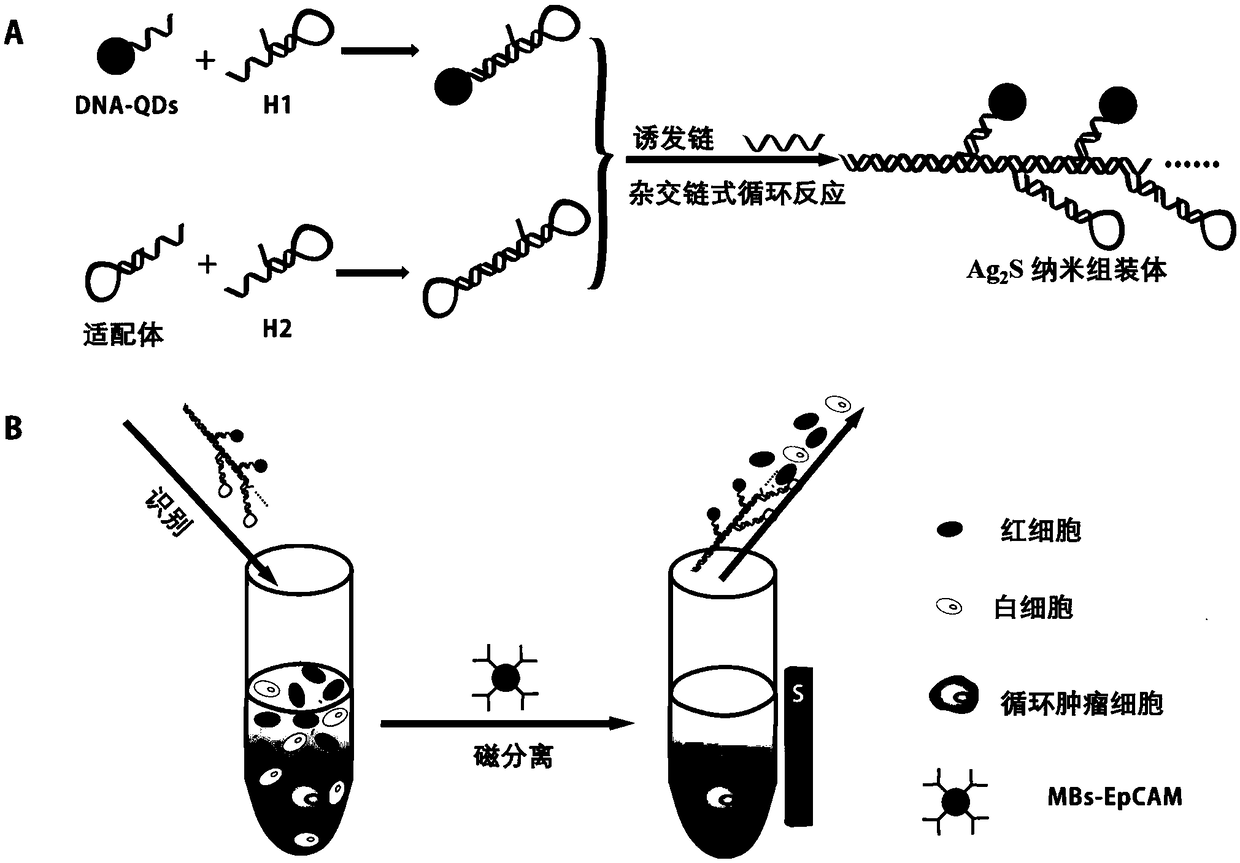
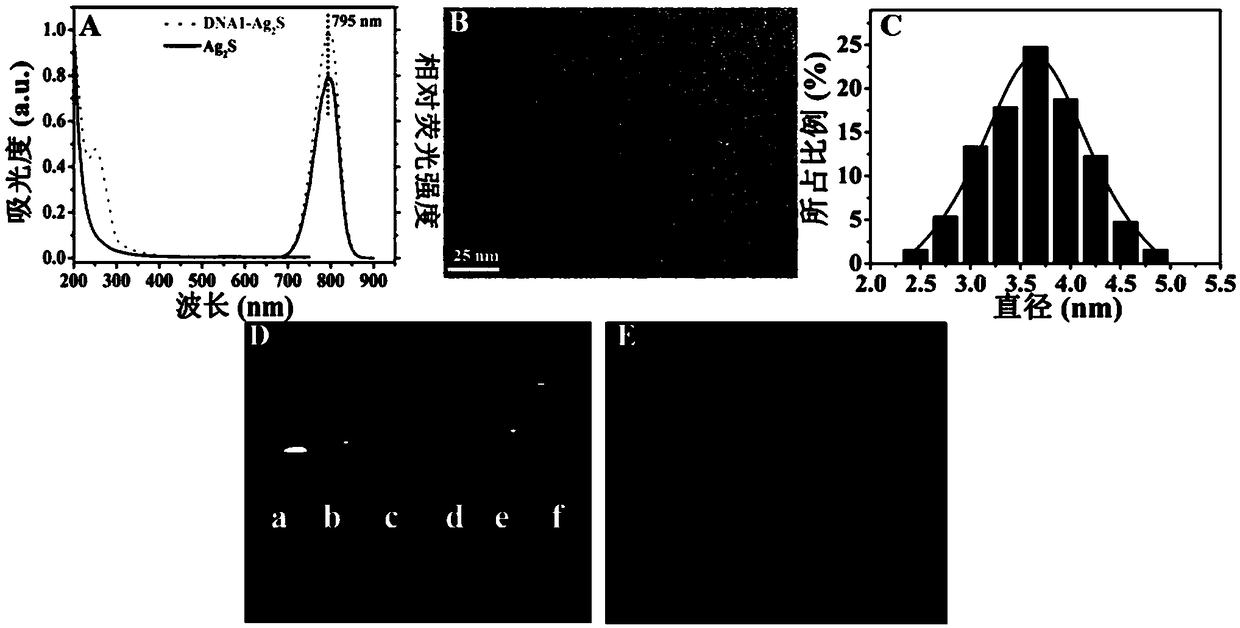
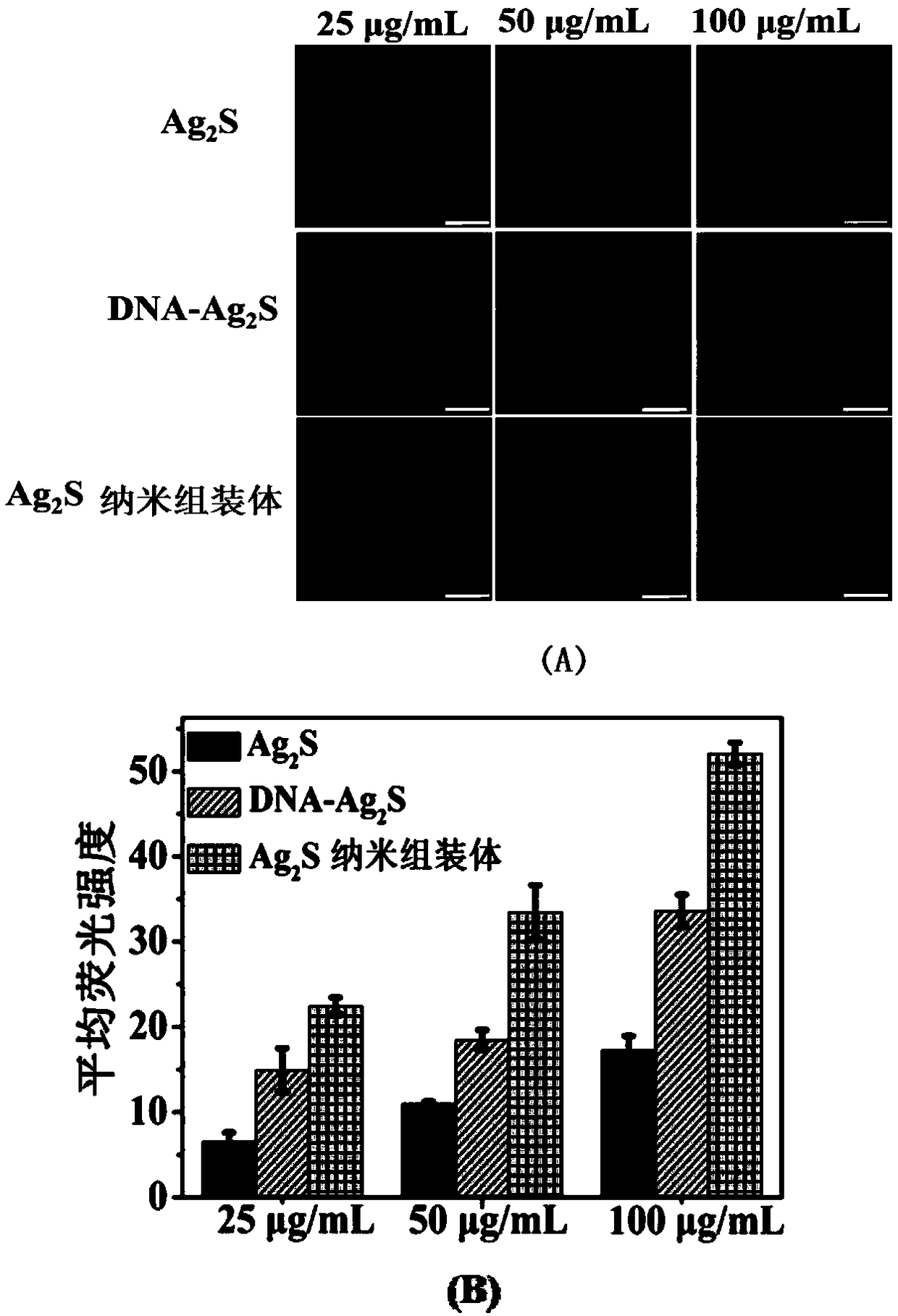
Near-infrared Ag2S nanometer assembly probe, and preparation method and applications thereof
ActiveCN108918477AAchieve early diagnosisHigh sensitivityInvestigating moving fluids/granular solidsFluorescence/phosphorescenceNanodotTumor cells
The invention provides a near-infrared Ag2S nanometer assembly probe. The near-infrared Ag2S nanometer assembly probe comprises a plurality of monomers M1 and M2 which are assembled based on HCR reaction; the monomer M1 is prepared through assembling of DNA-Ag2S nanodot and nucleic acid sequence such as H1 represented by SEQ ID NO.2, wherein the DNA sequence is represented by SEQ ID NO.1; the monomer M2 is prepared through assembling of an aptamer such as the aptamer represented by SEQ ID NO.3 and nucleic acid sequence such as H2 represented by SEQ ID NO.4. The invention also provides a preparation method of the near-infrared Ag2S nanometer assembly probe. The raw materials are easily available; the conditions are mild; normal temperature reaction is adopted; no heating is needed; productquality is stable. The invention also provides application method and applications of the near-infrared Ag2S nanometer assembly probe in detection of circulating tumor cells. The near-infrared Ag2S nanometer assembly probe is used in detection of circulating tumor cells, is high in specific identification capacity and sensitivity, can be used for further analysis of target cells in whole blood, and is promising in application prospect in cancer clinical diagnosis, postoperative prevention, and treatment.
Owner:EAST CHINA NORMAL UNIVERSITY
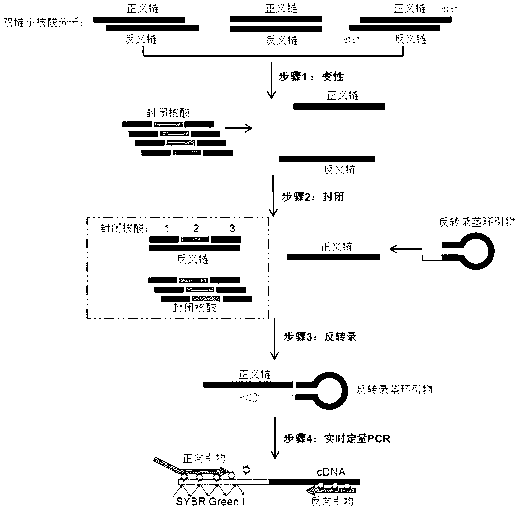
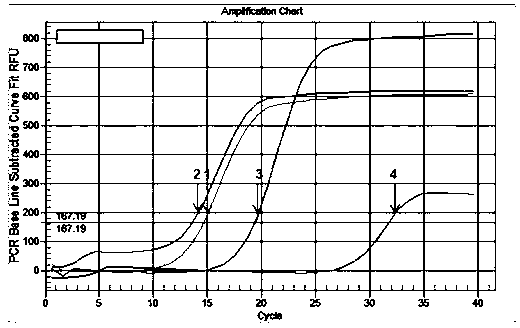
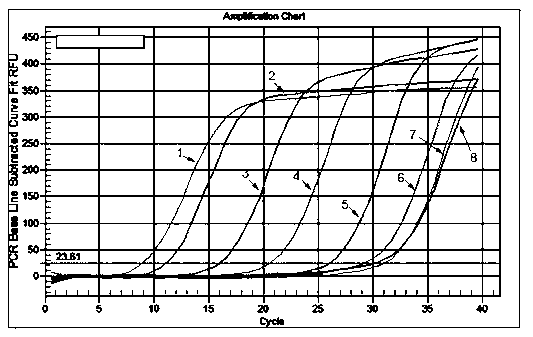
Small RNA detection method and use thereof
The invention relates to a small RNA detection method and a use thereof and belongs to the field of double-stranded small RNA detection. The small RNA detection method comprises the following steps of 1, designing multiple continuous or discontinuous closed RNAs which are complementary with one chain of a small RNA, a reverse transcription stem-loop primer which is complementary with an end 3' of the other chain of the small RNA, and a forward primer and a reverse primer for quantitative PCR, and 2, detecting the small RNA by denaturation, enclosing, inverse transcription and quantitative PCR. The small RNA detection method has simple processes, a low detection cost, a detection limit reaching a femtogram grade, high detection efficiency, strong singularity and high sensitivity. The small RNA detection method can be widely used for quantitative analysis, qualitative analysis and pharmacokinetic analysis of a small RNA drug, for amplification, specific identification and differentiation of a small RNA molecule, and especially for molecular diagnosis, small RNA drug metabolism and small RNA drug functional study.
Owner:BIOMICS BIOTECH
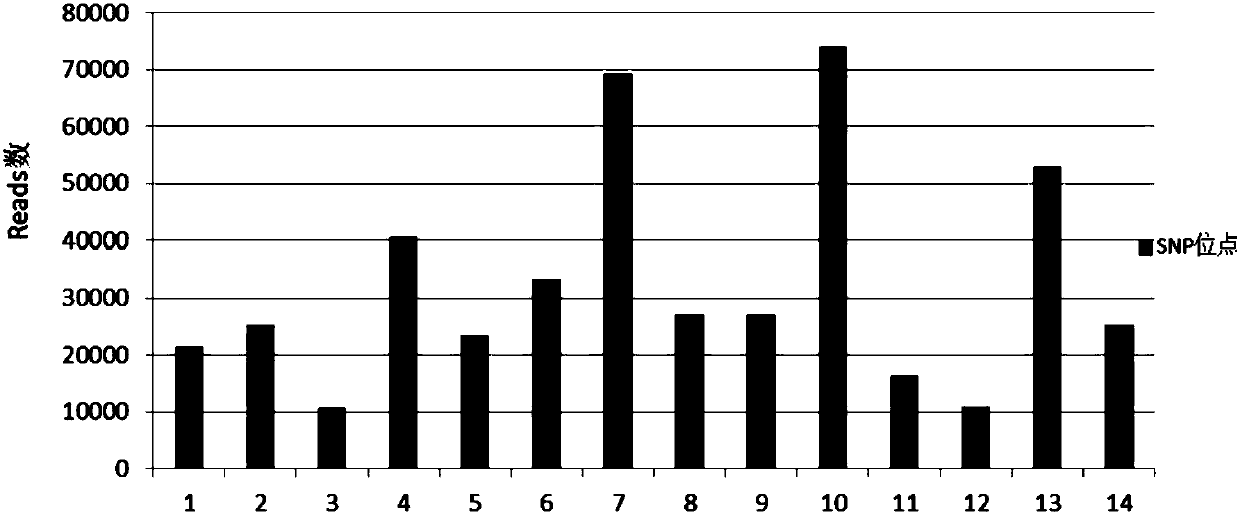
A method and a kit for detecting nucleic acid sample pollution and application of the method
ActiveCN108823296AGuarantee the quality of inspectionEasy to operateMicrobiological testing/measurementPolymorphism analysisBiology
The application discloses a method and a kit for detecting nucleic acid sample pollution and application of the method. The method for detecting the nucleic acid sample pollution includes the following steps: PCR amplification is performed on a nucleic acid sample to be measured by adopting a SNP detection primer, mononucleotide polymorphism of the nucleic acid sample to be measured is analyzed, whether pollution of different individual sources is existed in the nucleic acid sample to be measured or not is judged according to a mononucleotide polymorphism analysis result; and the SNP detectionprimer is composed of at least an analysis primer pair of 14 SNP sites. The specificity of each individual SNP is creatively utilized and is used as specific identification information of the nucleicacid sample to be detected, if SNP information of the detected nucleic acid sample to be measured conforms to the self SNP information, the result that the nucleic acid sample to be detected is not polluted is judged, if not, and the result that the nucleic acid sample to be detected has pollution is judged. Whether the nucleic acid sample to be measured exists pollution or not can be accuratelyjudged by the detection method.
Owner:BGI GENOMICS CO LTD +2
Water-soluble cationic polyelectrolyte with end group provided with fluorophore pyrene, and preparation method and application thereof
InactiveCN103012639AOrganic compound preparationMicrobiological testing/measurementFluoProbesCationic polyelectrolytes
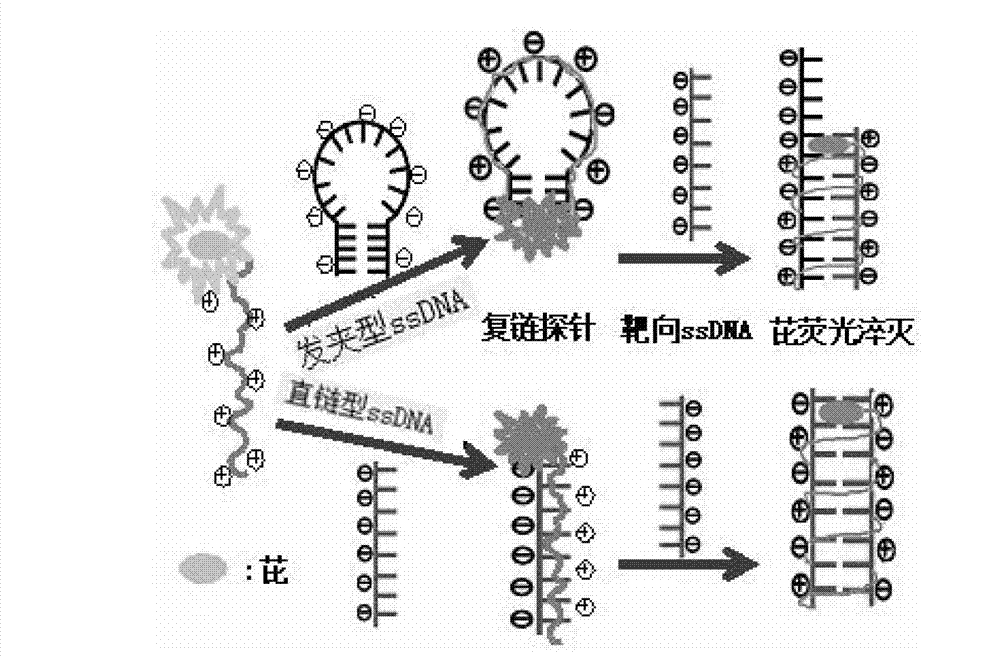
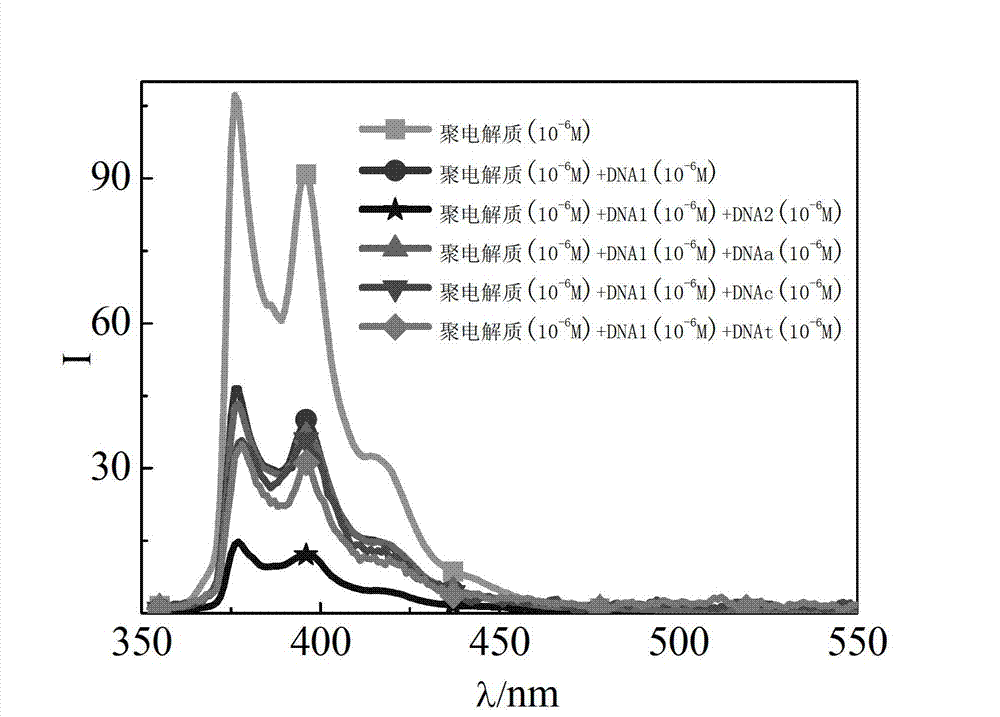
The invention relates to a fluorescent high-molecular polymer and a preparation method of the fluorescent high-molecular polymer, in particular relates to a water-soluble cationic polyelectrolyte with an end group provided with fluorophore pyrene, a preparation method of the water-soluble cationic polyelectrolyte, and an application of water-soluble cationic polyelectrolyte in the detection of base sequences of nucleic acid molecules. A novel multi-stranded fluorescent probe is formed by electrostatic adsorption and hydrophobic interaction between the water-soluble cationic polyelectrolyte with the end group provided with fluorophore pyrene and single-stranded oligomerization nucleic acid molecules (DNAs with a hairpin structure and a linear chain structure). When nucleic acid molecules with base sequence structures complementary with those of the fluorescent probe exist, hybridization is performed through Watson-Crick base pairing principle, as the fluorescent pyrene at the end group is inserted into a DNA double-helix structure, the quenching ability of the base to the pyrene fluorescence is enhanced, so that the fluorescence intensity of the pyrene is remarkably reduced compared with that of the pyrene acted on non-target nucleic acid molecules, therefore, the specific identification of the base sequence structures of the nucleic acid molecules can be realized through difference of fluorescence properties of polyelectrolyte with fluorophore pyrene.
Owner:UNIV OF SCI & TECH BEIJING
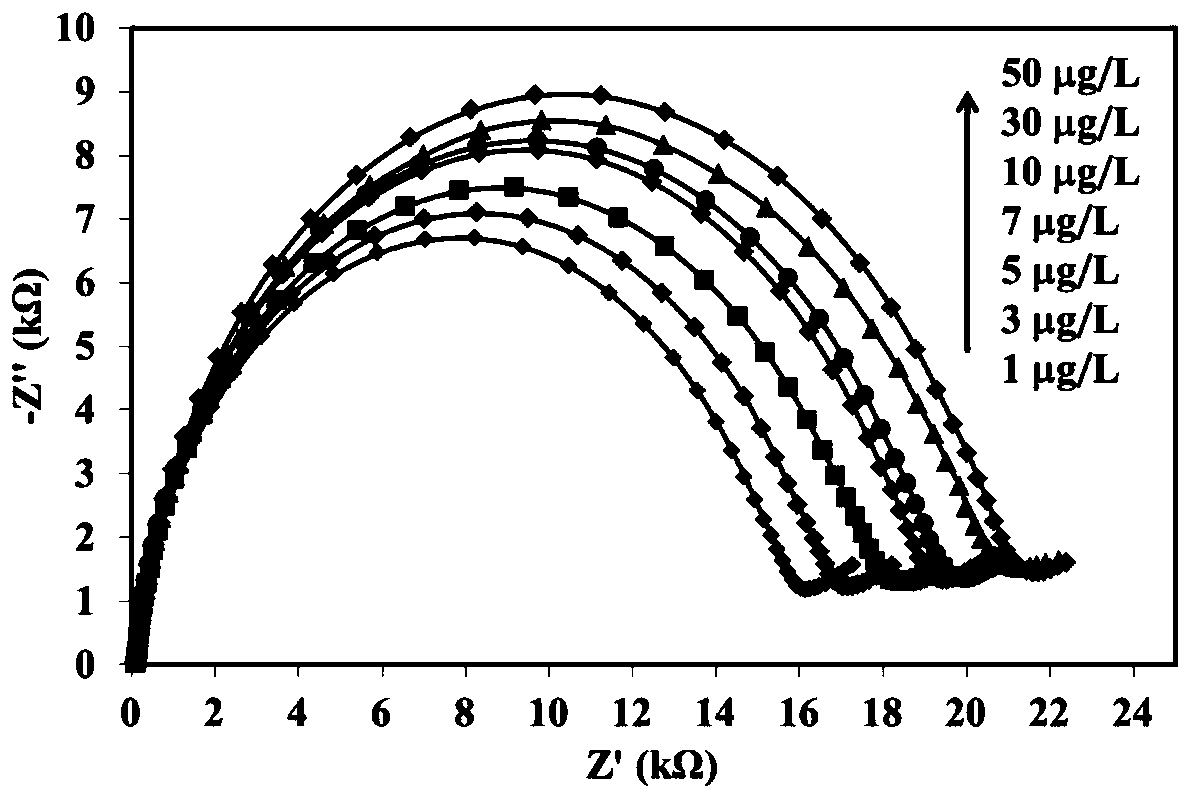
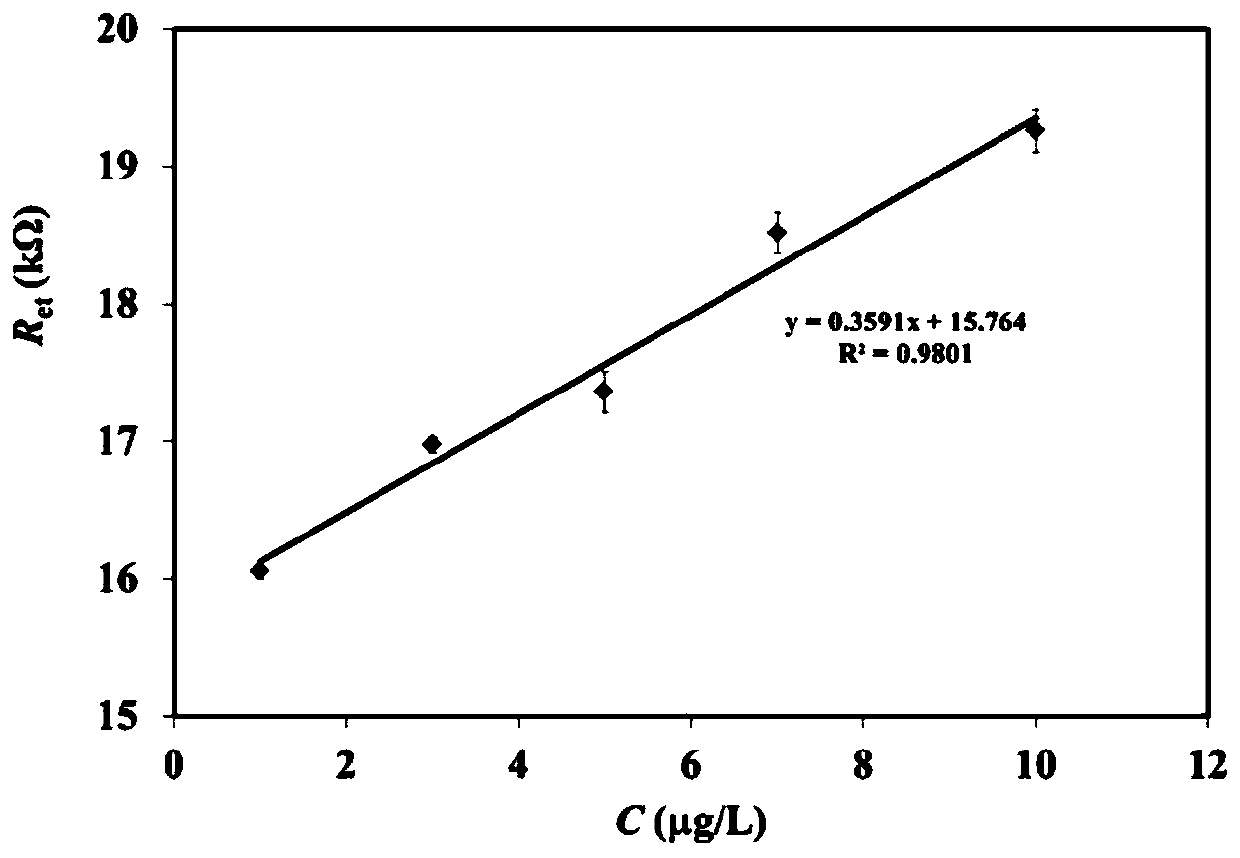
Electrochemical AC impedance biosensor for detecting MMP-14 and preparation method thereof
InactiveCN109916985AHigh selectivityHigh sensitivity detectionMaterial electrochemical variablesImpedance biosensorBuffer solution
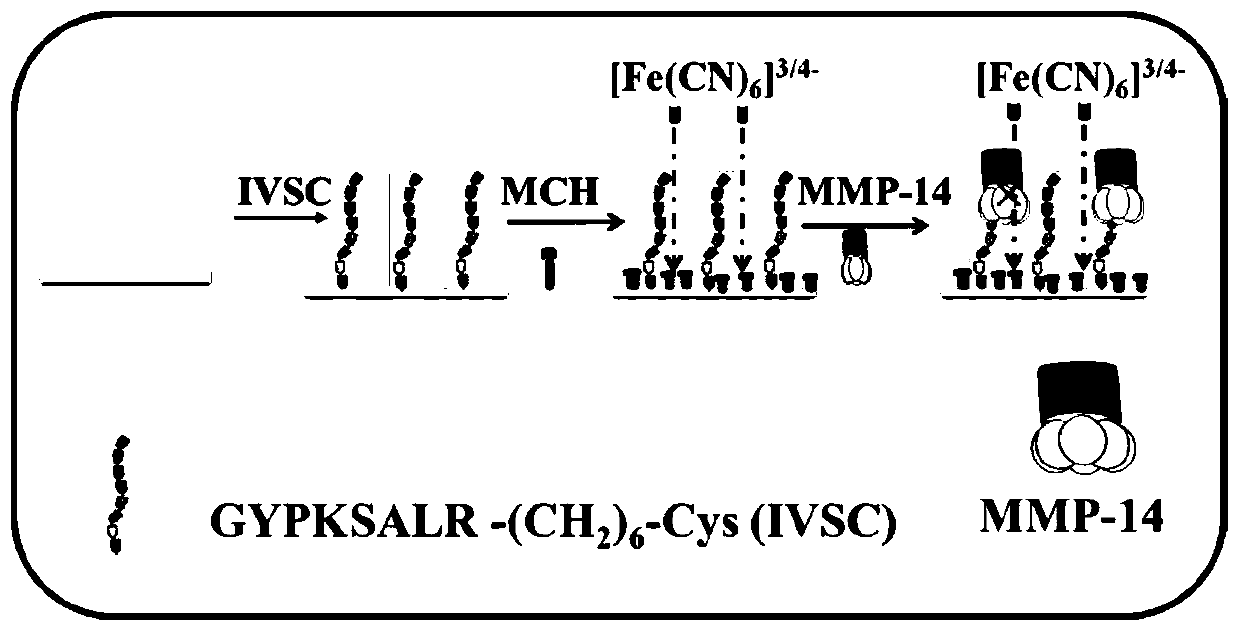
The invention provides an electrochemical AC impedance biosensor for detecting MMP-14 and a preparation method thereof. The polypeptide inhibitor is dissolved in the Tris-HCl buffer solution so as toobtain the MMP-14 specific identification probe solution, wherein that amino acid sequence of the polypeptide inhibitor is: GYPKSALR-(CH2) 6-Cys; the gold electrode is immersed into the MMP-14 specific identification probe solution to perform self-assembling modification and then rinsed by the Tris-HCl buffer solution so as to obtain the IVSC / GE electrode; and the MCH solution is dropped into theIVSC / GE electrode surface for sealing and the electrode surface is rinsed so as to obtain the electrochemical AC impedance biosensor for detecting MMP-14. The electrochemical AC impedance biosensor has high specificity.
Owner:NORTHWEST UNIV
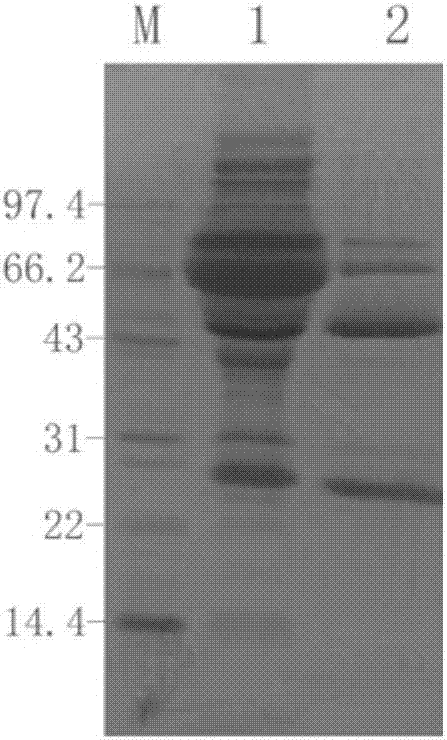
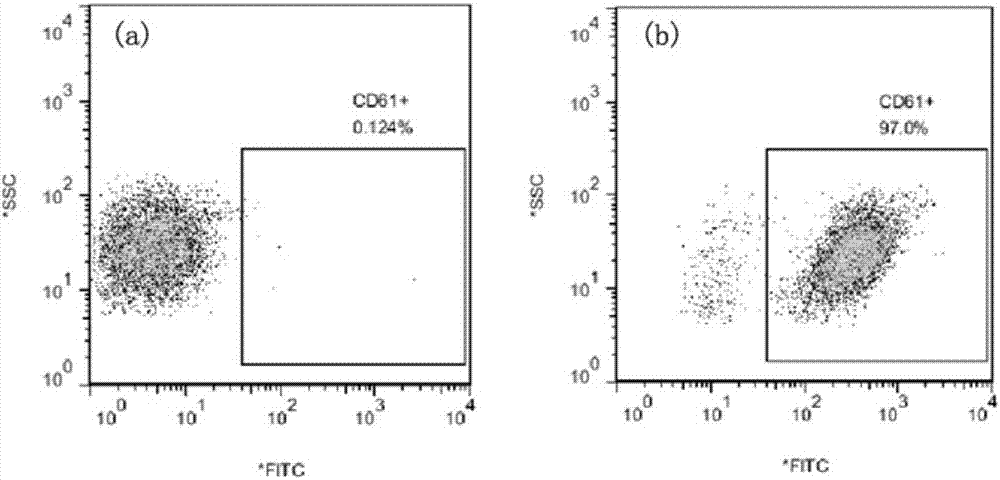
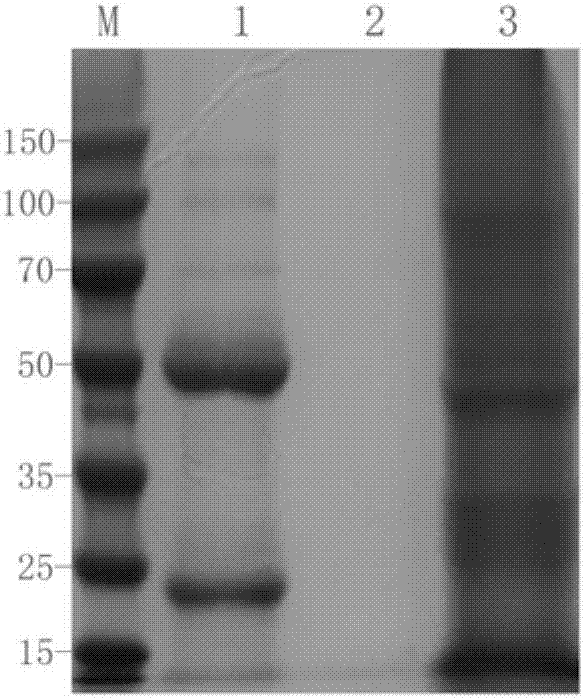
Mouse anti-human CD61 monoclonal antibody hybridoma cell line, monoclonal antibody and preparation method and application thereof and flow cytometry detection reagent
ActiveCN107058242AImmunoglobulins against cell receptors/antigens/surface-determinantsTissue cultureAntigenBALB/c
The invention discloses a mouse anti-human CD61 monoclonal antibody hybridoma cell line, a monoclonal antibody and a preparation method and application thereof and a flow cytometry detection reagent, and belongs to the technical field of biological engineering. According to the mouse anti-human CD61 monoclonal antibody hybridoma cell line, human peripheral blood leukocytes are used as an immunogen for immunizing Balb / c mice; a cell fusion technology is adopted and hybridoma cells are screened by utilizing an immunocytochemistry and a flow cytometry indirect immunofluorescent method; an antibody generated by monoclonal hybridoma cells is subjected to specific identification by applying an immunoprecipitation-mass-spectrometric method; Western Blot proves that one hybridoma cell line IID5G8 capable of stably secreting a mouse anti-human CD61 monoclonal antibody is obtained and a preservation number is CGMCC (China General Microbiological Culture Collection Center) No. 13300. The monoclonal antibody is prepared by adopting an animal in-vivo induction method; a heavy chain of the antibody is identified to be IgG1 and a light chain is identified to be kappa type; after the antibody is purified, an FITC (Fluorescein Isothiocyanate) label can be used for detecting antigen expression of human peripheral blood platelets CD61 by utilizing flow cytometry direct-labeling immunofluorescence.
Owner:HENAN UNIV OF CHINESE MEDICINE
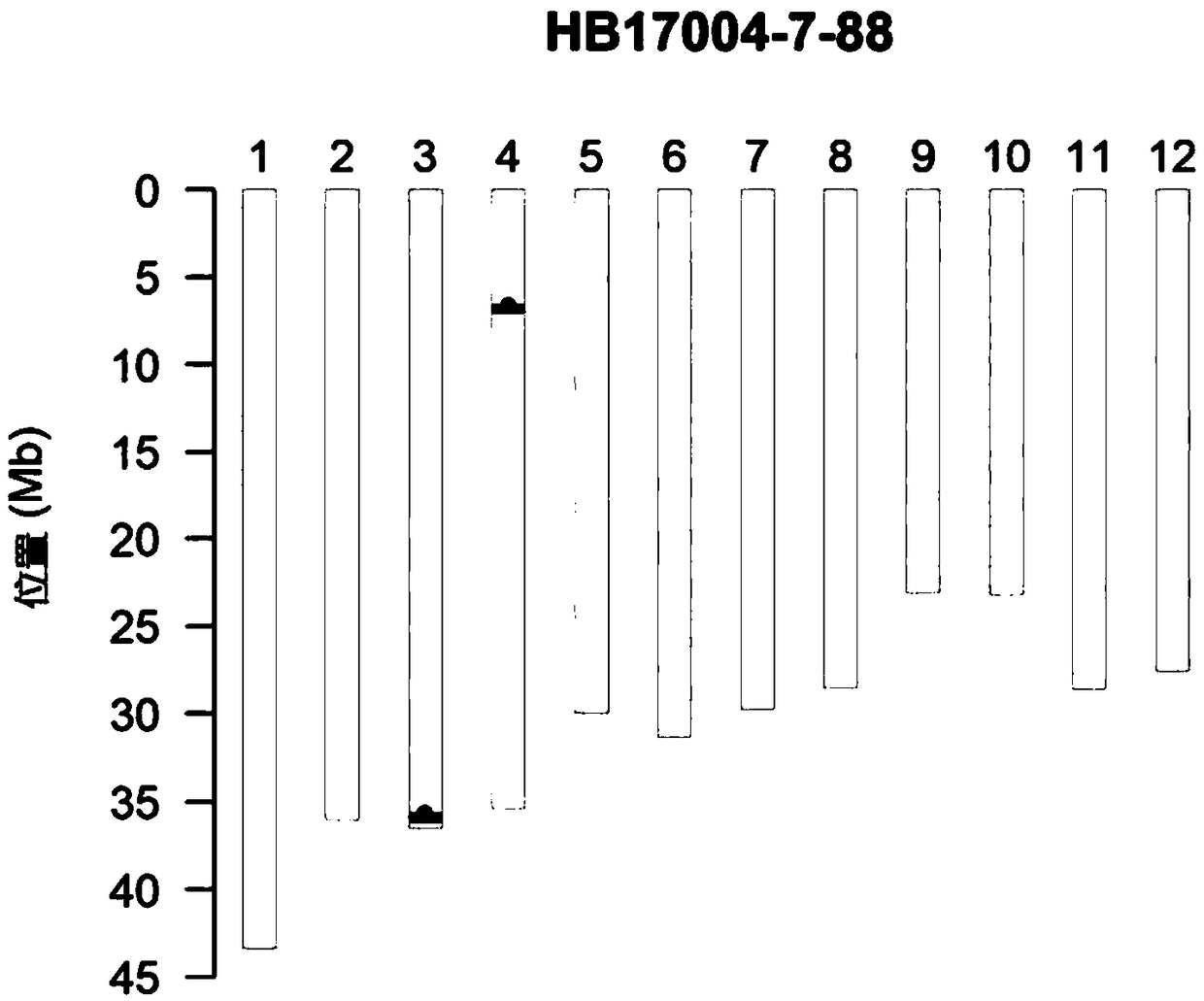
Recombinant nucleotide fragments Rec78801 and Rec78802, detection primer and application thereof
ActiveCN109486996AStrong specificityEasy to detectMicrobiological testing/measurementDNA/RNA fragmentationBiotechnologyMolecular identification
The embodiment of the invention relates to the field of molecular breeding and particularly relates to recombinant nucleotide fragments Rec78801 and Rec78802 obtained when a molecular breeding technology is utilized for selective breeding of rice plants containing genes resisting brown planthopper, a detection primer and an application thereof. The recombinant nucleotide fragments provided by theembodiment of the invention are selected from a sequence containing nucleotide at the 430-603 site of a sequence shown in SEQ ID NO:1, or a variant or a complementary sequence thereof; a sequence containing a sequence shown in SEQ ID NO:1, or a variant or a complementary sequence thereof; a sequence containing nucleotide at the 678-895 site of a sequence shown in SEQ ID NO:2, or a variant or a complementary sequence thereof; a sequence containing a sequence shown in SEQ ID NO:2, or a variant or a complementary sequence; and combination of the above fragments. The recombinant nucleotide fragments can be used as specific identification markers of major gene to be applied to molecular-marker selective breeding, and also can be used as specific molecular identification markers for identifyingrice materials.
Owner:WIN ALL HI TECH SEED CO LTD
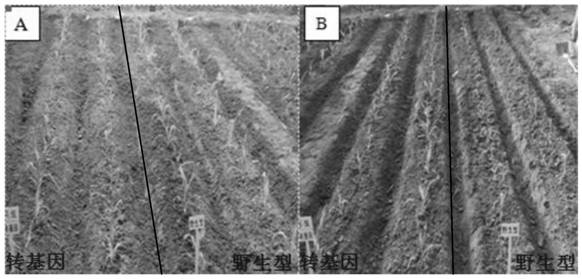
Herbicide-resistant corn HH2823 transformation event capable of efficiently utilizing nutrients and specific identification method and application of herbicide-resistant corn HH2823 transformation event
ActiveCN112280786AGenetic stabilityImproved agronomic traitsBacteriaMicrobiological testing/measurementBiotechnologyNucleotide
The invention belongs to the technical field of biology, and relates to a herbicide-resistant corn HH2823 corn transformation event for efficiently utilizing nutrients of AlHAK1 and bar genes and a specific identification method and application of the herbicide-resistant corn HH2823 corn transformation event. The transformation event takes a nucleotide sequence shown in SEQ ID NO.1 as a left wingsequence of an exogenous gene and takes a nucleotide sequence shown in SEQ ID NO.2 as a right wing sequence of the exogenous gene. According to the corn transformation event provided by the invention,exogenous gene specificity can be introduced into a corn strain, and the receptor is endowed with efficient utilization of corn potassium nutrition and herbicide glufosinate-ammonium resistance. Theexogenous gene can be stably inherited in the receptor corn; expression of the exogenous gene can improve the potassium absorption and utilization efficiency of the receptor corn. According to the detection method, molecular marker detection can be carried out by utilizing a transformation event, and the breeding work efficiency is improved.
Owner:DALIAN UNIV OF TECH
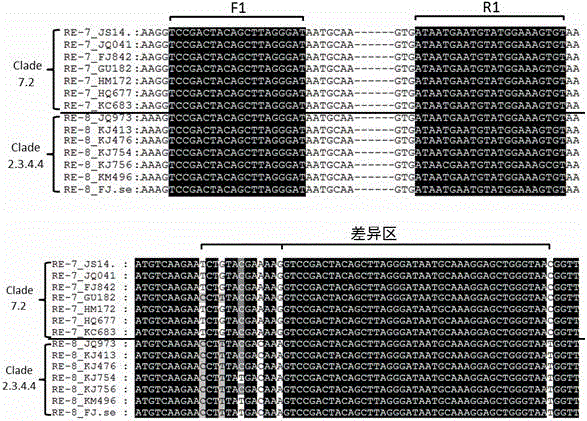
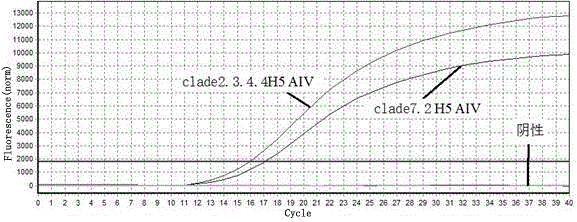
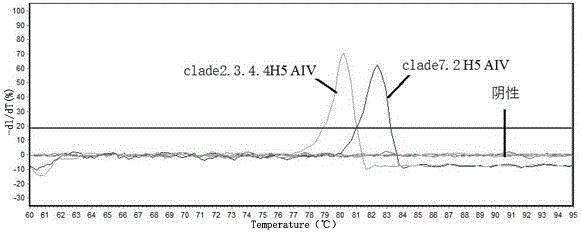
Real-time fluorescent quantitative PCR (Polymerase Chain Reaction) primers for distinguishing clade 7.2 type H5 AIV (H5 Subtype Avian Influenza Virus) from clade 2.3.4.4 type H5 AIV
ActiveCN106755586AStrong specificityIdentification method is simpleMicrobiological testing/measurementMicroorganism based processesAgricultural scienceNucleotide
The invention provides a group of real-time fluorescent quantitative PCR (Polymerase Chain Reaction) primers for distinguishing a clade 7.2 type H5 AIV (H5 Subtype Avian Influenza Virus) from a clade 2.3.4.4 type H5 AIV, and the real-time fluorescent quantitative PCR primers are designed by utilizing GC content difference existing in characteristic nucleotide sequences of HA (Hemagglutin) genes of the clade 7.2 type H5 AIV and the clade 2.3.4.4 type H5 AIV. The clade 7.2 type H5 AIV and the clade 2.3.4.4 type H5 AIV are effectively distinguished by utilizing the phenomena that both the real-time fluorescent quantitative PCR primers can be effectively amplified when real-time fluorescent quantitative PCR is carried out on the clade 7.2 type H5 AIV and the clade 2.3.4.4 type H5 AIV, but the real-time fluorescent quantitative PCR primers have melting temperature (Tm value) difference existing in melting curves generated after the real-time fluorescent quantitative PCR is carried out on the clade 7.2 type H5 AIV and the clade 2.3.4.4 type H5 AIV. By matching with a computer which is connected with a real-time fluorescent quantitative PCR instrument and utilizing analysis software of the real-time fluorescent quantitative PCR instrument, specific identification and diagnosis on infection situations of the clade 7.2 type H5 AIV and the clade 2.3.4.4 type H5 AIV can be achieved by just one group of primers. According to the real-time fluorescent quantitative PCR primers provided by the invention, an identification method is simple, and the efficiency and an accuracy rate are higher.
Owner:INST OF ANIMAL HUSBANDRY & VETERINARY FUJIAN ACADEMY OF AGRI SCI
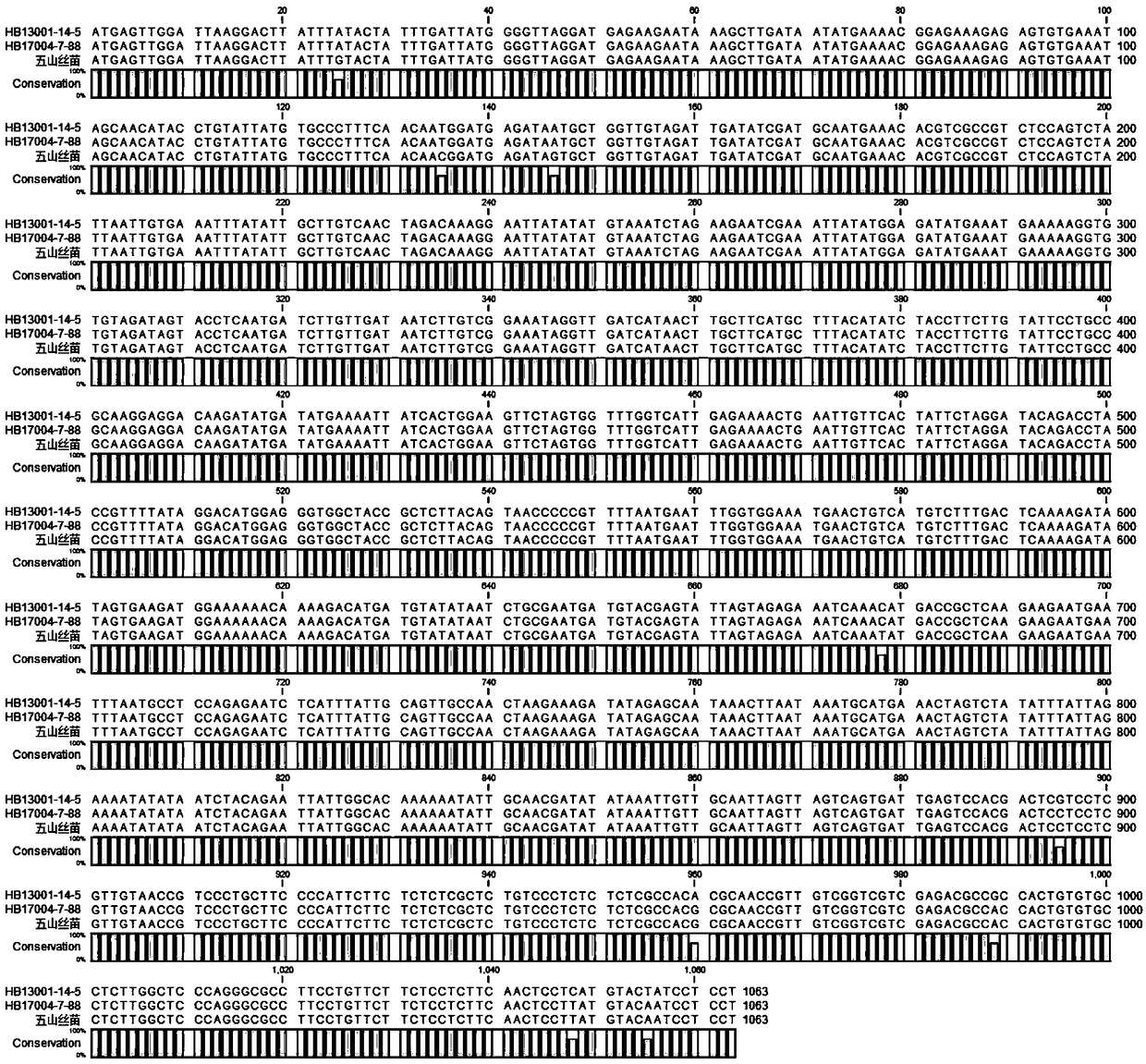
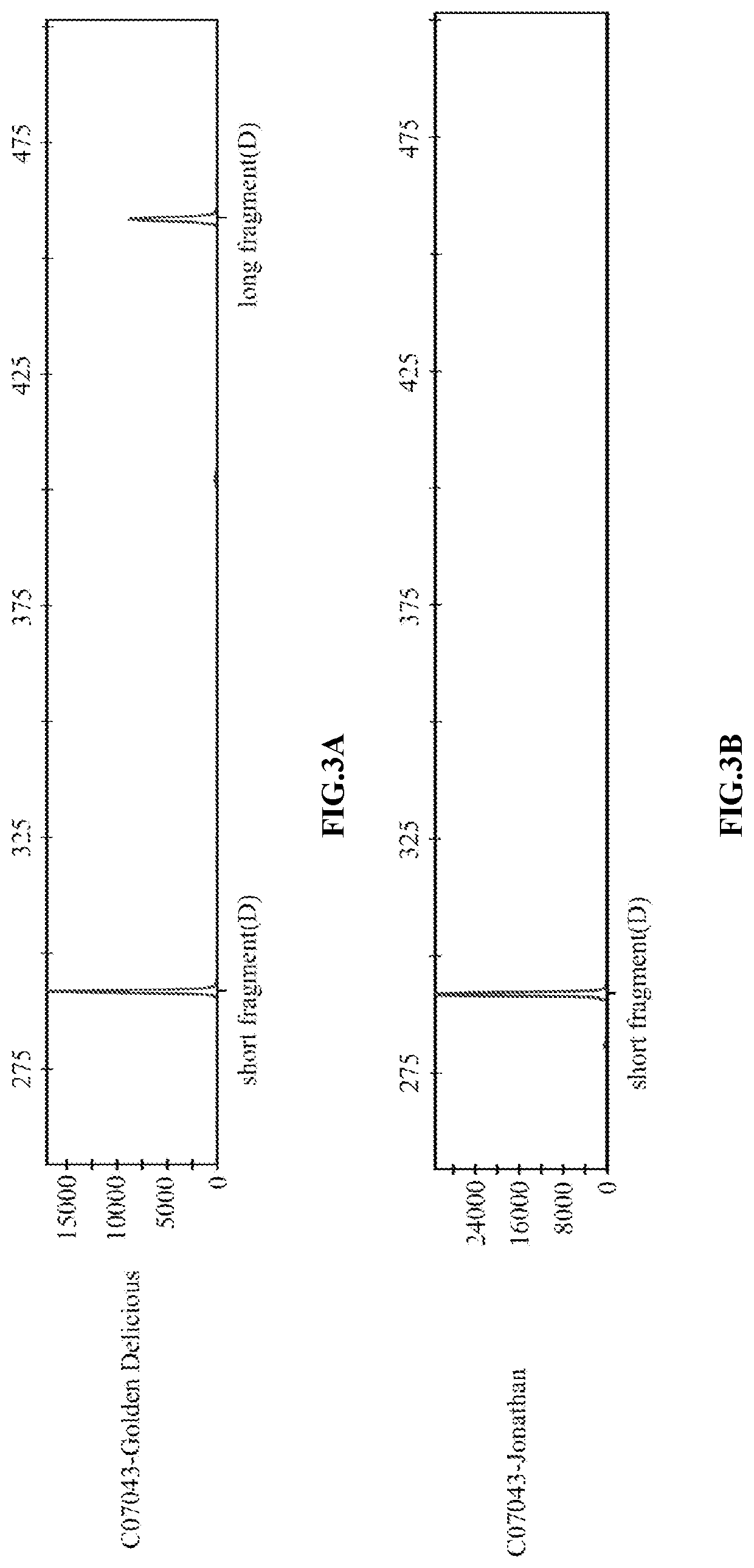
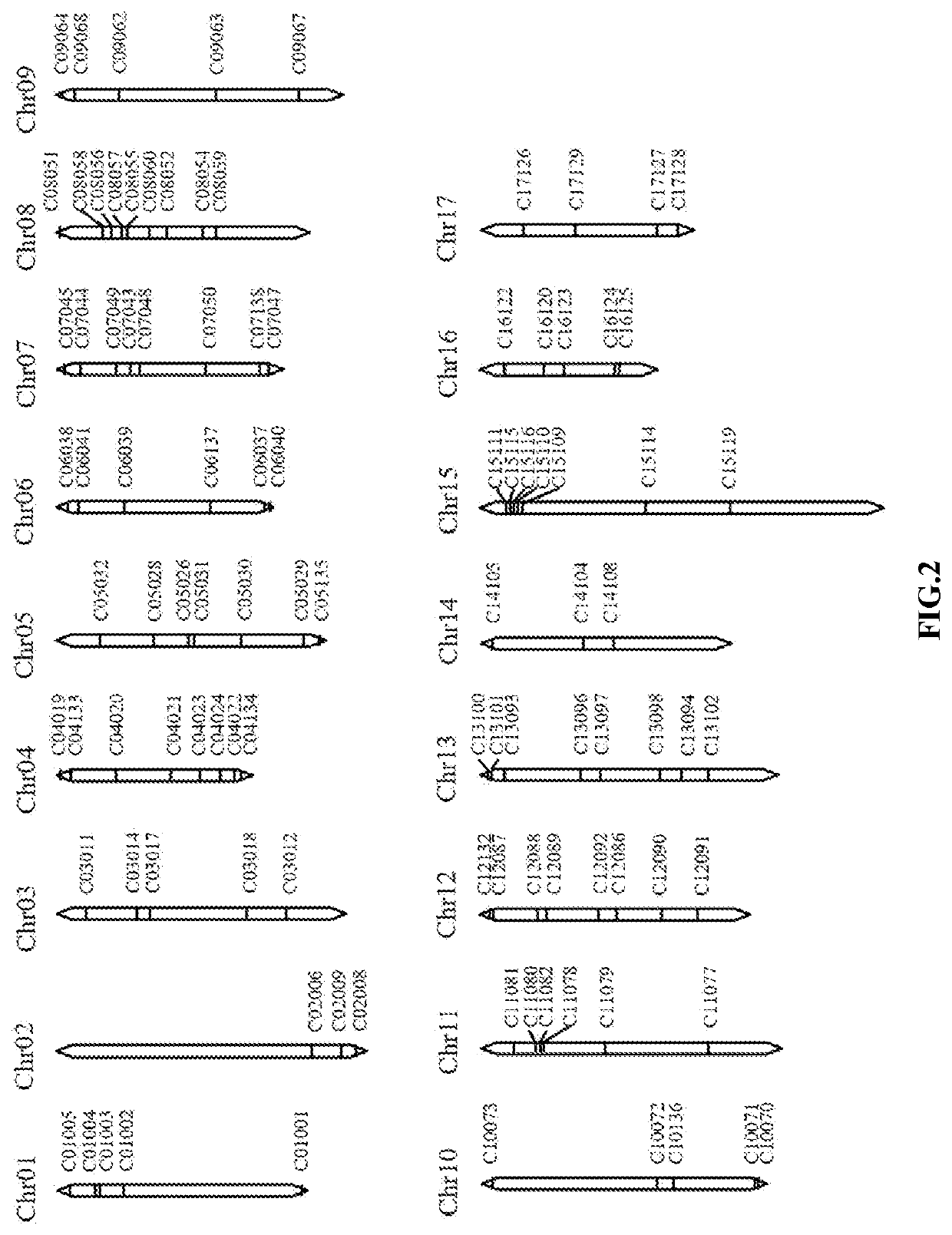
Indel marker genotype database of malus and use in identity fingerprinting of germplasm resources
PendingUS20220090213A1The process is stable and efficientManagementMicrobiological testing/measurementProteomicsBiotechnologyMultiplex
The disclosure provides an InDel marker genotype database of whole genome of Malus and use in specific identification of germplasm resources thereof, belonging to the technical field of molecular biology. The constructing method of the genotype database comprises: genotyping InDel markers of genomic DNA samples of Malus germplasm resources by using primer pairs and multiplex PCR; recording one line data for each Malus germplasm resource according to the genotype of InDel marker; collecting all germplasm resources records to obtain an InDel marker genotype database of the whole genome of Malus. The whole genome InDel marker genotype database of Malus provided by the disclosure can identify the specificity of apple germplasm simply, efficiently and stably, thus overcoming the limitation that DUS test is easily influenced by environment, and providing a powerful tool for the protection of new apple varieties and the management of germplasm resources.
Owner:CHINA AGRI UNIV
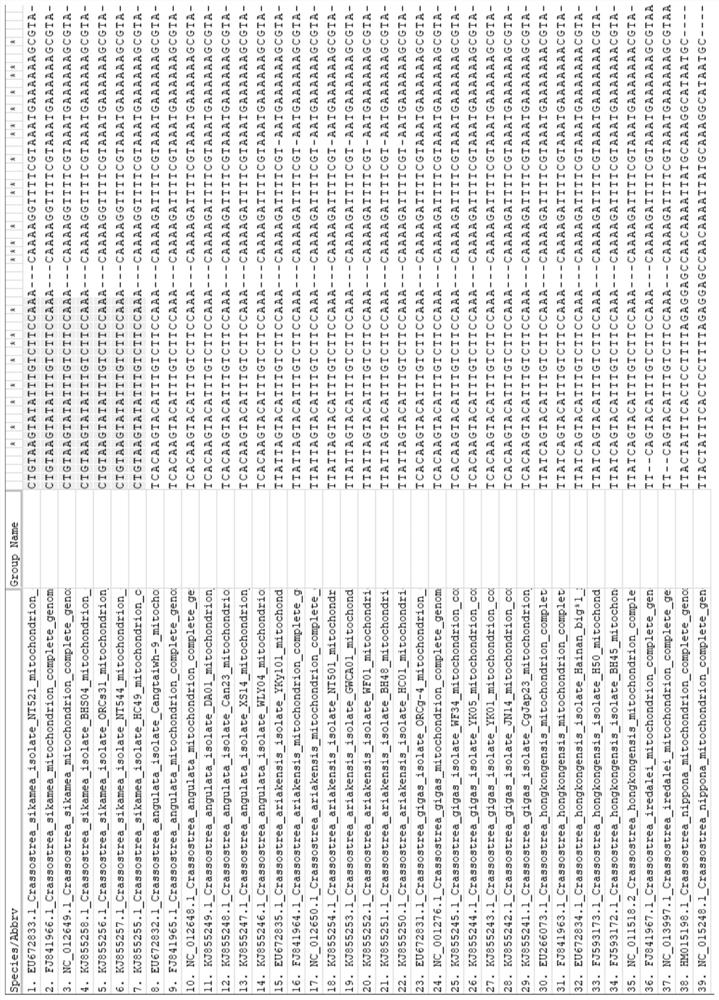
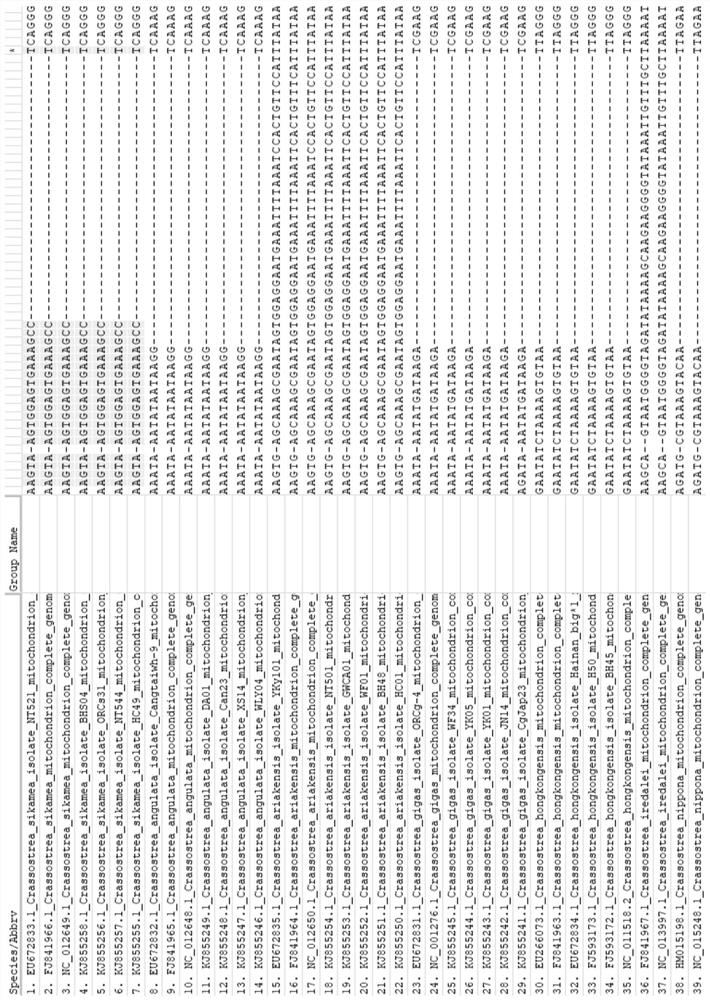
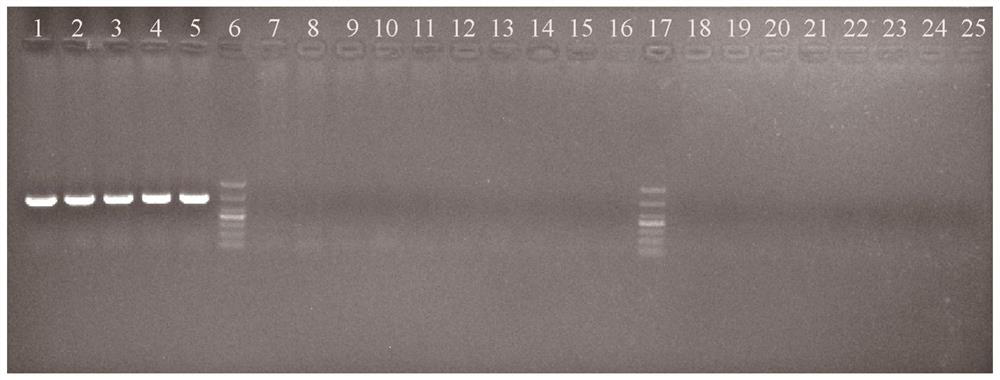
Characteristic sequence, specific identification primer and identification method for identifying crassostrea sikamea
ActiveCN112626238ARapid identificationHigh polymorphismMicrobiological testing/measurementDNA/RNA fragmentationBiotechnologyNucleotide
The invention discloses a characteristic sequence, specific identification primer and identification method for identifying crassostrea sikamea. The characteristic sequence is located in a mitochondrial non-coding region between glycine tRNA synthetase and proline tRNA synthetase of the crassostrea sikamea, and the characteristic sequence is a nucleotide sequence as shown in SEQ ID NO.1; a sequence of the specific identification primer comprises an upstream primer MNR-F: 5' -CTGTAAGTATATTTGTCTTCCA-3' ; and a downstream specific primer MNR-S-R: 5' -AGGCTTTCACTCCACTTACT-3' . According to the invention, the crassostrea sikamea can be effectively distinguished from other four drassostrea species, the length of the determined characteristic sequence of the crassostrea sikamea is 660+ / -5 bp, and whether the crassostrea sikamea exists or not can be judged according to the existence and the size of a specific band. In the field of crassostrea sikamea germplasm resource protection and crassostrea sikamea breeding, species identification can be conducted on parent crassostrea sikamea before seed breeding, and it is guaranteed that bred seeds are purebreds and are not affected by other species possibly hybridized with the bred seeds.
Owner:浙江万里学院宁海海洋生物种业研究院 +1
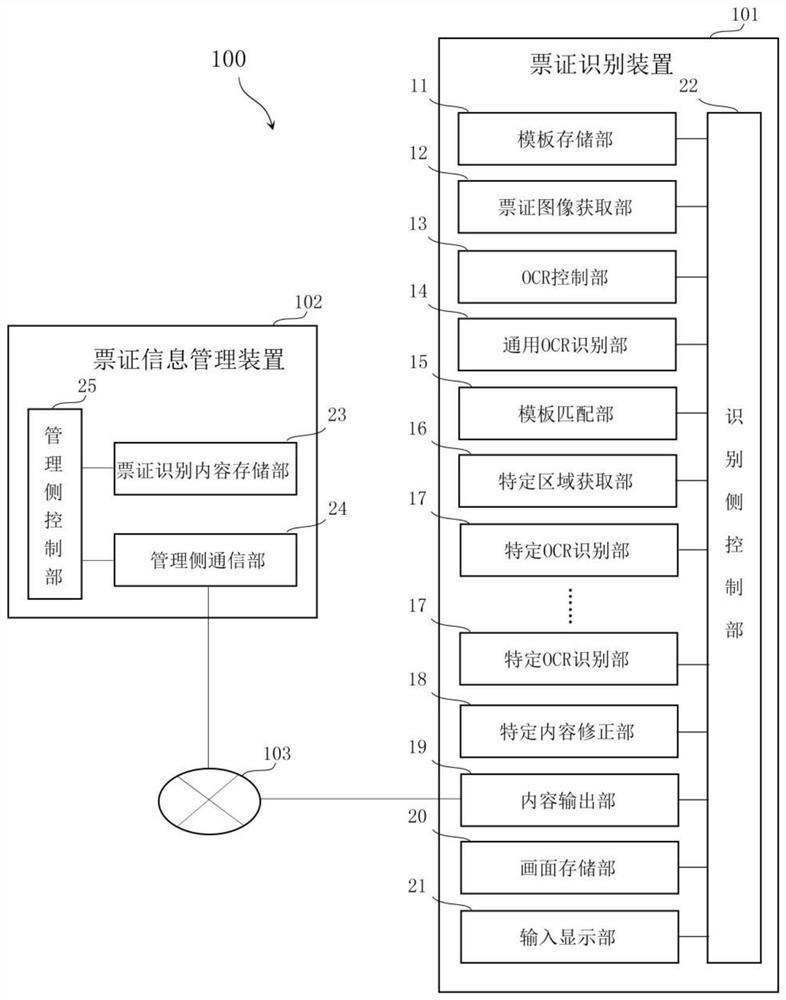
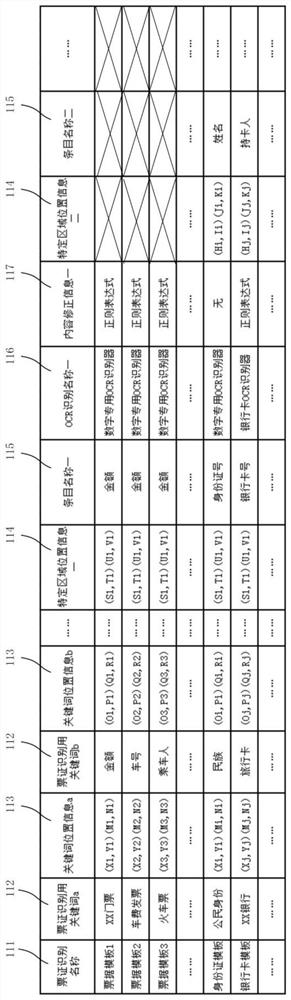
Ticket identification device and ticket information management system
PendingCN111860450AAccurate identificationConvenient statisticsOther databases queryingSpecial data processing applicationsTicketInformation control
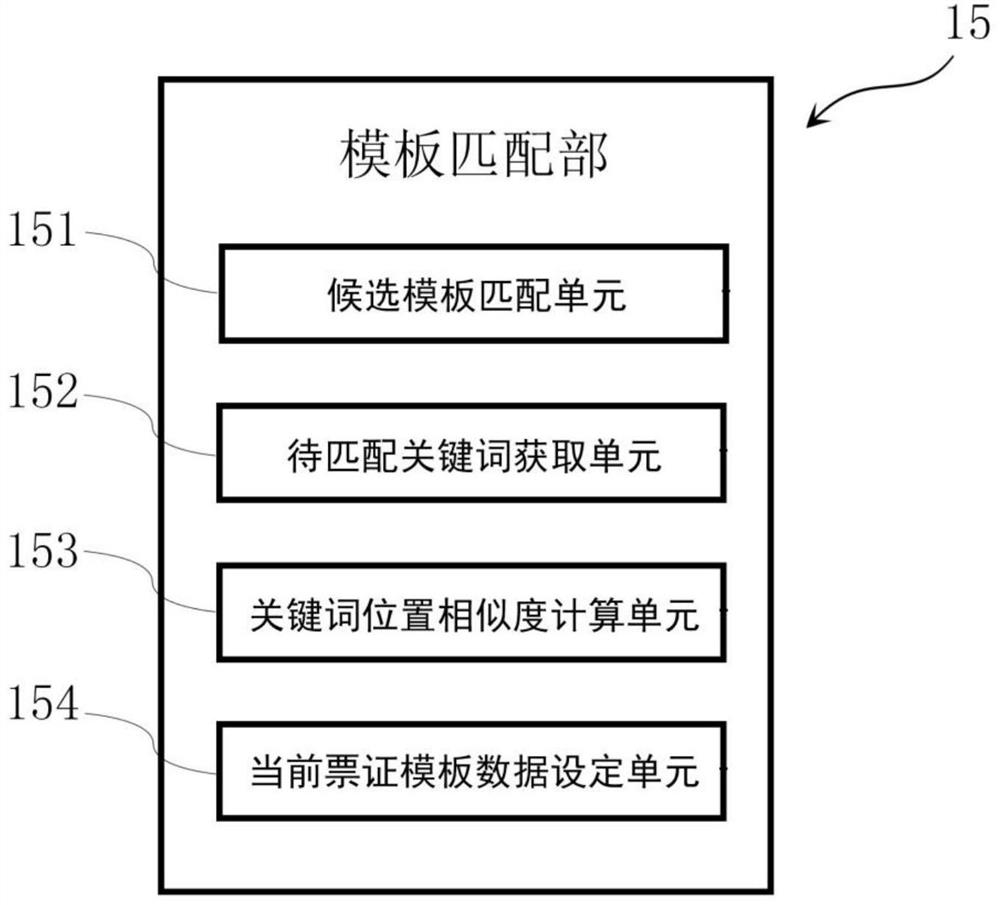
The invention provides a ticket identification device and a ticket information management system. The device is used for identifying contents recorded on a plurality of predetermined tickets and managing the identified contents, and is characterized in that the device comprises a template storage part, an OCR control part, a universal OCR identification part, a template matching part and at leastone specific OCR identification part, wherein the template storage part is used for storing ticket template data corresponding to a ticket template of each type of ticket; the ticket template data comprises specific area position information corresponding to a specific area needing to be recognized by a specific OCR recognition part in the ticket; the OCR control part controls the universal OCR identification part to carry out preliminary identification on the ticket image to be identified; the template matching part is used for matching ticket template data corresponding to the ticket to be identified as current ticket template data based on a preliminary identification result; and the OCR control part is used for controlling the specific OCR part to carry out specific identification on characters in the specific area so as to identify corresponding specific contents.
Owner:RICOH IMAGING TECH SHANGHAI CO LTD